Automating Early Disease Prediction Via Structured and Unstructured Clinical Data
This study presents a fully automated methodology for early prediction studies in clinical settings, leveraging information extracted from unstructured discharge reports. The proposed pipeline uses discharge reports to support the three main steps of…
Authors: Ane G Domingo-Aldama, Marcos Merino Prado, Alain García Olea
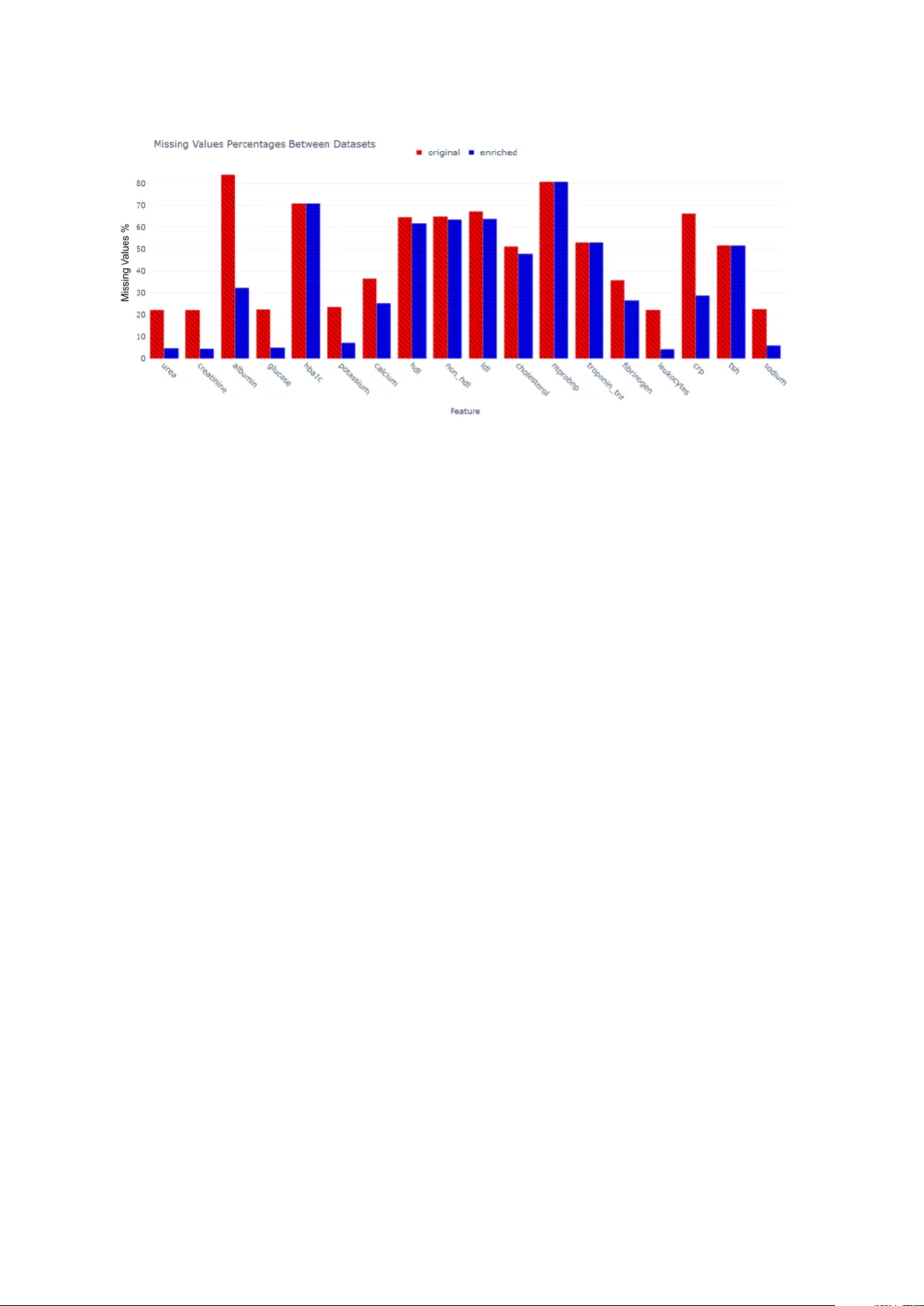
Automating Early Disease Prediction Via Structured and Unstructured Clinical Data A utomatizaci´ on de la pr e dic ci´ on tempr ana de enferme dades me diante datos cl ´ ınic os estructur ados y no estructur ados Ane G. Domingo-Aldama 1 , Marcos Merino Prado 1 , Alain Garc ´ ıa Olea 2 , Josu Goik o etxea 1 , Koldo Go jenola 1 , Aitzib er A tutxa 1 1 Univ ersity of the Basque Country (EHU), 2 Basurto Universit y Hospital ane.garciad@eh u.eus, mmerinoprado@gmail.com, alain.garciaolea@osakidetza.eus, josu.goik o etxea@eh u.eus, k oldo.go jenola@ehu.eus, aitzib er.atuc ha@eh u.eus Abstract: This study presents a fully automated metho dology for early prediction studies in clinical settings, lev eraging information extracted from unstructured dis- c harge rep orts. The prop osed pip eline uses disc harge rep orts to support the three main steps of early prediction: cohort selection, dataset generation, and outcome la- b eling. By pro cessing disc harge rep orts with natural language processing tec hniques, w e can efficiently identify relev an t patien t cohorts, enrich structured datasets with additional clinical v ariables, and generate high-quality labels without manual in ter- v ention. This approac h addresses the frequen t issue of missing or incomplete data in co dified electronic health records (EHR), capturing clinically relev an t information that is often underrepresented. W e ev aluate the metho dology in the context of pre- dicting atrial fibrillation (AF) progress ion, sho wing that predictiv e mo dels trained on datasets enric hed with disc harge rep ort information achiev e higher accuracy and correlation with true outcomes compared to mo dels trained solely on structured EHR data, while also surpassing traditional clinical scores. These results demon- strate that automating the in tegration of unstructured clinical text can streamline early prediction studies, improv e data qualit y , and enhance the reliability of predic- tiv e mo dels for clinical decision-making. Keyw ords: Early Prediction, A trial Fibrillation Progression, Electronic Health Records, Natural Language Pro cessing. Resumen: Este estudio presen ta una metodolog ´ ıa totalmente automatizada para estudios de predicci´ on temprana en entornos cl ´ ınicos, aprov echando la informaci´ on extra ´ ıda de informes de alta hospitalaria no estructurados. El pro ceso propuesto utiliza los informes de alta para respaldar los tres pasos principales de la predicci´ on temprana: selecci´ on de cohortes, generaci´ on de conjuntos de datos y etiquetado de resultados. Mediante el procesamiento de los informes de alta con t ´ ecnicas de pro ce- samien to del lengua je natural, p o demos iden tificar de manera eficiente las cohortes de pacientes relev an tes, enriquecer los conjuntos de datos estructurados con v ari- ables cl ´ ınicas adicionales y generar etiquetas de alta calidad sin interv enci´ on man- ual. Este enfoque ab orda el principal problema de los registros m ´ edicos electr´ onicos (RME) co dificados que son los datos faltantes o incompletos, capturando infor- maci´ on cl ´ ınicamen te relev an te que a menudo est´ a infrarrepresentada. Ev aluamos la meto dolog ´ ıa en el contexto de la predicci´ on de la progresi´ on de la fibrilaci´ on au- ricular (F A), demostrando que los mo delos predictivos en trenados con conjuntos de datos enriquecidos con informaci´ on de informes de alta logran una ma y or precisi´ on y correlaci´ on con los resultados reales en comparaci´ on con los mo delos en trenados ´ unicamen te con datos estructurados de RME, al tiemp o que sup eran las puntua- ciones cl ´ ınicas tradicionales. Estos resultados demuestran que la automatizaci´ on de la integraci´ on de texto cl ´ ınico no estructurado puede agilizar los estudios de predicci´ on temprana, mejorar la calidad de los datos y aumentar la fiabilidad de los mo delos predictivos para la toma de decisiones cl ´ ınicas. P alabras cla ve: Predicci´ on T emprana, Progresi´ on de la Fibrilaci´ on Auricular, His- torias Cl ´ ınicas Electr´ onicas, Pro cesamien to del Lengua je Natural. 1 Intr o duction Early diagnosis (ED) has garnered significan t atten tion in b oth Artificial Intelligence (AI) and medicine due to its transformative p oten- tial in p ersonalized medicine. By identifying a condition at its initial stages, clinicians can in tervene so oner, preven t complications, and tailor treatmen ts to each patient which leads to impro ved outcomes and reduced health- care costs. The success of these ED mo dels hea vily depends on the diversit y , v olume, and gran ularity of the data a v ailable (Ng et al., 2016; Alzubi, W atzlaf, and Sheridan, 2021). Mo dern AI systems for p ersonalized and predictiv e medicine predominan tly employ mac hine learning (ML) mo dels that rely ex- clusiv ely on structured, tabular data (in- cluding: demographic information, lab ora- tory measurements, co ded diagnoses, etc.). This reliance has led to an increasing dep en- dence on structured electronic health records (EHRs) (Ristevski and Chen, 2018). EHRs, no w cen tral to healthcare systems worldwide, pro vide a rich rep ository of both structured and unstructured patient information, play- ing a crucial role in clinical decision-making (Holmes et al., 2021; Alzubi, W atzlaf, and Sheridan, 2021). Despite the ric h information con tained in structured EHRs, they present notable chal- lenges, primarily stemming from the man- ual annotation and con version of medical data in to standardized structured and dis- crete fields. This pro cess is prone to errors, missing v alues, and inconsistencies, which can significan tly impact data reliabilit y and the ov erall quality of the predictive mo dels built up on this information (Garcia Olea et al., 2021; Botsis et al., 2010; Alzubi, W atzlaf, and Sheridan, 2021). Efforts to standardize medical co ding with systems lik e ICD-10, OPCS, and SNOMED ha ve mitigated some c hallenges. Ho wev er, no universal guidelines exist regarding the lev el of detail required for clinical do cumen- tation. Consequently , not all information rel- ev ant to sp ecific prediction tasks is captured in structured EHRs, often necessitating extra man ual annotation (a time-consuming pro- cess that in tro duces v ariabilit y and potential h uman error). F urthermore, ED studies require a cohort selection pro cess to identify patients suitable for the task. This step t ypically depends on structured EHR data or, when key informa- tion is missing, on manual review of clinical records. As a result, the pro cess b ecomes la- b or intensiv e, slo ws down research w orkflows, and in tro duces v ariabilit y and p oten tial h u- man error (Chen et al., 2025; Jin et al., 2024). In this context, the presen t study intro- duces a metho dology that automates the key steps required for early prediction tasks, in- cluding cohort selection, dataset generation, and patient lab eling. The approac h combines structured tabular data deriv ed from EHRs with semi-structured disc harge rep orts in free-text format, pro cessed through a natu- ral language processing (NLP) pip eline. This metho dology reduces the need for man ual an- notation while simultaneously impro ving the qualit y and completeness of structured tabu- lar data b y leveraging information extracted from disc harge rep orts to mitigate missing v alues or co dification errors. The primary con tribution of this w ork lies in the design of this end-to-end metho dology and data gen- eration pip eline, rather than in prop osing a no vel predictive mo deling architecture. Sp ecifically , we fo cus on the progression of atrial fibrillation (AF) within one month to t wo years after the initial episo de (see Fig- ure 1). The early prediction of AF progres- sion in this time-windo w helps arrhythmia sp ecialists determine whether rhythm con trol therapies are appropriate for a patient based on their individual risk. In this study , AF progression encompasses all atrial fibrillation subt yp es: paro xysmal, p ersisten t, and p er- manen t 1 . All metho dological steps are ev alu- ated in this con text, and the resulting predic- tiv e mo dels are compared against traditional clinical scores for AF progression. T o this end, the research is guided b y the follo wing core questions: • RQ1 : Can the combination of struc- tured EHR data and information ex- tracted from discharge rep orts enhance and automate the dev elopment of ED mo dels? • RQ2 : Can free-text disc harge rep orts pro cessed with NLP techniques impro ve the qualit y and completeness of struc- tured tabular data derived from EHRs? 1 F or paro xysmal AF, progression is defined as the recurrence of the arrhythmia, whereas for p ersisten t and p ermanen t AF, it corresponds to the ongoing presence of the condition. Figure 1: Clinical scenario of AF pro- gression. • RQ3 : Can automatically generated silv er annotations ac hieve p erformance comparable to gold-standard annota- tions while significantly reducing manual effort? • RQ4 : Do es the prop osed metho dology pro duce predictiv e mo dels that outp er- form existing clinical scores for AF pro- gression prediction? 2 R elate d Work Disease prediction (Sharma et al., 2022; Xie, Y u, and Lv, 2021; Y u et al., 2023; Mishra and T arar, 2020) has attracted considerable at- ten tion in b oth the fields of AI and medicine due to its p oten tial to rev olutionize ED and p ersonalized medicine (Awwalu et al., 2015; Shah, 2018; Sc hork, 2019; Ullah, Akbar, and Y annarelli, 2020). T o carry out these studies three steps are necessary: cohort selection, dataset generation and lab eling. F or the cohort selection step, recen t w ork has explored automatic metho ds based on rule driv en systems (Vydiswaran et al., 2019) as w ell as NLP approaches that use discriminativ e language models (Soni and Rob erts, 2021) or even generativ e large lan- guage mo dels (Guan et al., 2023). Our metho d is related to rule based strategies but incorp orates a h ybrid mo dule designed to im- pro ve generalizabilit y , addressing a common limitation of traditional rule based cohort se- lection systems (Stubbs et al., 2019). Regarding the dataset generation step, the quality of EHR data is a critical factor in ensuring the reliability and accuracy of predictiv e mo dels, particularly in the context of medical research. Numerous studies ha v e addressed the common c hallenges associated with EHR data qualit y , as well as metho ds for assessing and improving it (Cruz-Correia et al., 2009; Lewis et al., 2023; F eder, 2018; T erry et al., 2019). F or instance, (Jetley and Zhang, 2019) suggest lev eraging infor- mation extraction tec hniques or manual re- view of clinical notes to address gaps in struc- tured data. This approach aligns with the hy- p othesis of the present pro ject, whic h aims to utilize similar tec hniques for extracting rele- v ant information from free-text discharge re- p orts to supplement and impro v e the quality of structured EHR data. The detrimental impact of missing v alues in clinical contexts is widely recognized. Nu- merous studies emphasize that missing data in tro duce uncertaint y and bias into predic- tiv e mo dels, presen ting a significant challenge in this field (Sterne et al., 2009; Kahale et al., 2020; Ibrahim, Ch u, and Chen, 2012). Con- sequen tly , implemen ting strategies to reduce the incidence of missing v alues can effectiv ely mitigate the limitations of curren t predictive mo dels. The lab eling of instances is a critical comp onen t of ED pip elines, y et producing high qualit y gold standard lab els demands extensiv e exp ert in v olvemen t and manual re- view. This makes the process time consum- ing, costly , and difficult to scale. As a re- sult, the use of automatically generated sil- v er standard lab els has b ecome increasingly prev alent in ED research. Although these la- b els are imperfect, prior studies hav e sho wn that they can pro vide sufficient signal to train effectiv e mo dels (W agholik ar et al., 2020; Mc- Da vid et al., 2013). In the particular con text of AF pro- gression prediction , researc h in this do- main t ypically fo cuses on t wo primary sce- narios: the prediction of new-onset AF and the recurrence of AF following therap eutic in- terv entions. AI-driv en mo dels hav e demon- strated remark able success in predicting in- ciden t AF, often outp erforming conv en tional metho ds (Sion tis e t al., 2020). These mo dels dra w on div erse data sources such as clin- ical records, cardiac imaging, and electro- ph ysiological data, including the use of EHRs (Tseng and Noseworth y , 2021; Nadara jah et al., 2021; Hulme et al., 2019; Tiw ari et al., 2020). Notably , (Sung et al., 2022) dev el- op ed a ML model to predict the risk of newly detected AF p ost-strok e, incorp orating both structured v ariables and unstructured clini- cal text pro cessed through NLP . Most existing studies on AF progression fo cus on the p ost-catheter ablation setting. As review ed b y (F an et al., 2023), ML metho ds hav e shown strong p erformance in this con text. These approac hes often en- hance predictive accuracy b y com bining clin- ical v ariables with additional echocardiogra- ph y inputs (Knech t et al., 2024; Zhou et al., 2022), features extracted from electro cardio- grams (ECGs) (Qiu et al., 2024), or anatom- ical data from computed tomography (Liu et al., 2024; Brahier et al., 2023). In addition, despite all the adv ances in the field of AF progression prediction, a signifi- can t gap remains in accurately predicting AF progression following an initial ev ent. This underexplored time windo w is clinically im- p ortan t, as it can supp ort first-con tact ph ysi- cians with patient reference and cardiologists or arrh ythmologists in making more informed treatmen t decisions. Consequently , our study systematically compares traditional clinical scores with our generated mo dels using au- tomatically obtained datasets. 3 R esour c es The present researc h has b een p ossible thanks to the following resources kindly pro- vided b y the Basque Public Healthcare Sys- tem (Osakidetza): • Semi-structured disc harge rep orts in Spanish : 1 . 2 × 10 6 disc harge rep orts from 2015 to 2020 (see Figure 2). Figure 2: Example of a semi-structured disc harge rep ort. • Co dified structured data : Clini- cal data for each patien t, enco ded by healthcare professionals using standard- ized co ding systems and stored in the Osakidetza Business Intelligence (OBI) platform (see Figure 3). Figure 3: Example of the co dified struc- tured data. 4 Exp erimental setup The exp erimen tal setup fo cuses on the con- text of AF progression and is divided in three steps, the cohort selection for patien ts ap- propriate for the pro ject, the tabular dataset generation step and the automatic lab eling step of AF progression (See Figure 4). 4.1 Automatic Cohort selection T o study AF progression, w e iden tified pa- tien ts with AF onset, defined as their first do cumen ted AF episo de with no prior his- tory . Given the errors encoun tered in (Gar- cia Olea et al., 2021), w e implemen ted a dual v erification approach in tegrating both struc- tured and unstructured data sources. P atients were initially selected via struc- tured EHR data (OBI system), then v al- idated through clinical reports using a t wo-step NLP approac h: first with a fine-tuned EriBER T a enco der-only language mo del (De la Iglesia et al., 2025), then a reg- ular expression-based to ol for qualit y control. Both to ols are presen ted in (Garc ´ ıa-Olea et al., 2025). 4.2 T abular Dataset Generation and Enric hmen t A total of 85 clinical features w ere selected for this sp ecific task, encompassing demo- graphic data (7 features), patient history (35 features), lab oratory results (18 features), pro cedures and their outcomes (7 features), treatmen ts (16 features), and AF-related v ariables, including AF type and the AF pro- gression status, whic h serves as the target la- b el. These features include established AF risk factors and general clinical markers aimed at facilitating the prediction of AF progression. Figure 4: End to end o verview of the prop osed metho dology . The pipeline starts with automatic cohort selection, follo w ed b y dataset generation by com bining structured EHR data with information extracted from clinical rep orts. An NLP mo dule performs automatic lab eling, and the resulting silver and gold datasets are used to train and ev aluate a T abPFN mo del for AF progression prediction. While most of them are av ailable and co dified within the EHR, several k ey risk factors, suc h as left atrial size and even the AF progression status, are not represented in the structured co ding system. Ho wev er, some of this information can b e found in disc harge rep orts, consequen tly it is imp ortan t to retrieve this information to en- sure the qualit y of the predictive mo dels and to reduce the amoun t of manual annotation needed. The tabular generation pip eline inv olv ed three k ey steps (see Figure 5): 1. Extract and structure clinical in- formation from disc harge rep orts using the R ep ort2V e ctor (R2V) pip eline. This three-step NLP pro- cess detailed in (Garc ´ ıa-Olea et al., 2025) consists of: (a) section identifica- tion, which distinguishes b et w een differ- en t parts of the rep ort (e.g., past med- ical history vs. curren t episo de) (de la Iglesia et al., 2023); (b) medical en tit y recognition, whic h extracts relev ant clin- ical men tions such as symptoms, diag- noses, and pro cedures, along with nega- tion detection to differen tiate b etw een confirmed and ruled-out conditions; and (c) regular expression matching to cap- ture sp ecific patterns of interest. This step returns a table with the 84 predic- tiv e clinical v ariables of interest for AF progression extracted from the discharge rep orts of the patient. 2. Pro cess structured EHR data from the OBI system using the Struc- tur e d2V e ctor (S2V) mo dule. The same 84 predictive features obtained in the previous step are no w extracted from the OBI system for each patien t and or- ganized in to a tabular format. 3. Merge b oth data sources into uni- fied patient-lev el v ectors using the V e ctorMer ger . The final tabular data con tains the co dified information from the OBI system enric hed with the in- formation extracted from the disc harge rep orts. 4.3 Automatic Lab eling The automatic lab eling process for determin- ing AF progression status consists of the fol- lo wing steps: 1. Each patient rep ort is pro cessed fol- lo wing the same approac h as the R2V to ol (using a section identification mo d- ule, a medical entit y recognition and negation c omponent, and regular expres- sions). This process extracts the AF sta- tus for each consultation date, catego- rizing it as AF episo de, return to sinus rh ythm, or no information. As a result, the complete history of the arrh ythmia can b e reconstructed for eac h patient. 2. Using the first documented AF episo de as the onset p oint, subsequen t consulta- tions are examined for men tions of new AF episo des or returns to sin us rhythm. AF progression is defined as the o ccur- rence of a new AF episo de betw een one mon th and t wo years after the initial di- agnosis. Based on this rule, three lab els are assigned: • AF Progression (1): Explicit men- tion of new AF episode after its on- AF ONSET COHORT CODIFIED DA T A DISCHARGE REPORTS CODIFIED DA T A VECTORS R2V S2V HYBRID VECTOR id Debut Date urea creatinina ... 10/1/2019 1 0.66 33 ... VM P A TIENT WITH AF DEBUT id Date urea creatinina ... 10/1/2019 ... 1 Null 33 ... id Date urea creatinina ... 22/1/2019 ... 1 0.66 40 ... REPORT VECTORS Figure 5: Overview of the vector generation pro cess. F or eac h patien t in the AF on- set cohort, all discharge rep orts (free text) and co dified data (structured data stored in the Business Intelligence system) are collected and pro cessed using the R ep ort2V e ctor (R2V) and Structur e d2V e ctor (S2V) to ols, resp ectiv ely . Eac h to ol generates a corresp onding set of vectors, whic h are then merged by the V e ctorMer ger (VM) to ol to pro duce a patient-specific vector that in tegrates b oth sources of clinical information. set. • No Progression (0): Do cumen ted si- n us rhythm or non-AF ECG find- ings after the onset. • Excluded (-1): Cases without suffi- cien t evidence to determine progres- sion status. 4.4 Prediction of AF Progression T o ev aluate the proposed dataset generation metho dology , we also apply it to its original purp ose: predicting atrial fibrillation (AF) progression within one month to tw o years after onset. In this study , w e emplo y a tabular founda- tion mo del, also referred to as a Large T abu- lar Mo del (L TM) (v an Breugel and v an der Sc haar, 2024). Sp ecifically , w e use T abPFN (Hollmann et al., 2025) due to its ability to efficien tly handle complex, small-scale tabu- lar datasets while leveraging prior knowledge through pretraining. During preliminary e xperimentation, sev- eral ML arc hitectures were ev aluated, in- cluding Supp ort V ector Machines, Random F orests, and X GBoost. How ev er, due to the high dimensionalit y of the predictiv e features and the substan tial prop ortion of missing v alues, these mo dels did not yield satisfac- tory results and were subsequen tly discarded. The experiments also explored v arious miss- ing v alue imputation strategies (such as mean and median imputation, as w ell as logistic regression–based imputation) none of which outp erformed the T abPFN p erformance. Ad- ditionally , data prepro cessing tec hniques in- cluding feature standardization, feature se- lection, and different sampling strategies (un- dersampling, ov ersampling, SMOTE, and T omek). Moreov er, T abPFN incorp orates in- ternal data prepro cessing strategies (includ- ing handling missing v alues and scale normal- ization) as w ell as an integrated hyperparam- eter optimization routine, which we utilized. W e ev aluate p erformance differences b e- t ween t w o datasets: the original dataset de- riv ed from co dified EHR information, and the enric hed dataset that incorp orates features extracted from discharge rep orts. In addition, we compare mo del p erfor- mance when using automatically generated silv er-standard annotations with that ob- tained using gold-standard labels man ually annotated by a cardiologist. The c haracter- istics of the dataset used in these experiments are summarized in T able 1. 4.4.1 Ev aluation Mo del p erformance w as assessed using b oth accuracy and the Matthews Correlation Co- efficien t (MCC). While accuracy measures the prop ortion of correct predictions, it can b e misleading in datasets with im balanced classes. MCC provides a more robust ev alua- tion by incorporating all elemen ts of the con- Size % P ositives T r ain-Silver 1023 65.40% T r ain-Gold 541 66.17% T est 278 64.03% T able 1: Characteristics of the dataset. The second column indicates the num ber of patien ts and the last column the p ercen tage of AF progression. fusion matrix (true p ositiv es, true negativ es, false p ositives, and false negatives) yielding v alues b etw een –1 (complete disagreemen t) and 1 (p erfect agreement). In medical prediction tasks, MCC is es- p ecially v aluable because it ev aluates the qualit y of predictions for b oth classes sim ul- taneously . Unlik e the F1-score, whic h fo- cuses solely on the p ositiv e class, MCC cap- tures the balance b etw een correctly identify- ing patien ts with and without the condition. This distinction is crucial in clinical con texts, where recognizing true negatives is as imp or- tan t as detecting true positives to preven t unnecessary interv en tions, costs, and patient distress. Consequen tly , MCC offers a more comprehensiv e and reliable measure of model p erformance in healthcare applications. Moreo ver, to ev aluate the clinical rele- v ance of our predictiv e mo dels, we com- pared them with established clinical scores for AF progression: CHADS2-V ASc 2 (Lip et al., 2010), HA TCH 3 (De V os et al., 2010), and APPLE 4 (Kornej et al., 2015). As these scores produce numerical v alues rather than direct classifications, a threshold of ≥ 2 was applied to conv ert them into bi- nary outcomes. The scores w ere calculated using the generated tabular data and the for- m ulas from their original publications. 5 R esults and Discussion In this section, we present the results of the prop osed metho dology , analyzing its impact on the prop ortion of missing v alues in the final dataset as w ell as the differences b e- t ween the automatic and gold-standard an- notations. W e also rep ort the outcomes of the AF progression prediction exp erimen ts p erformed using the differen t versions of the dataset. 2 Used for stroke risk stratification in AF patients. 3 Used for AF onset prediction. 4 Used for AF recurrence risk prediction after catheter ablation. 5.1 Dataset Enric hmen t The enriched dataset using the information of disc harge rep orts improv es considerably the amoun t of missing v alues and general recall of v ariables (see Figure 6). The comparison b et w een the original and enric hed datasets highligh ts the significan t impact of incorp o- rating information extracted from discharge rep orts. As shown in the figure 6, the prop or- tion of missing v alues in the original dataset (blue bars) is notably reduced in the enric hed dataset (red bars) across most lab oratory v ariables, indicating a substan tial improv e- men t in data completeness. V ariables suc h as albumin, CRP , and NT-proBNP (which orig- inally presen ted high lev els of missingness) sho w a marked increase in data a v ailabilit y after enric hmen t. This suggests that the in- tegration of textual information helps recov er clinically relev ant details often absen t from co ded records. The reco very of past medical history fea- tures increased substan tially , rising from an av erage of 2.62% p ositiv es in the orig- inal dataset to 14.25% p ositives in the en- ric hed v ersion (an absolute gain of 11.63 p er- cen tage p oin ts). Among all feature cate- gories, treatmen t-related v ariables b enefited the most from the enric hment pro cess. Their mean recall rose from 1.73% in the original dataset to 45.05% in the enriched dataset. F urthermore, certain clinically relev an t v ariables with high predictive v alue were completely missing from the original co di- fied dataset, as they are not recorded within the OBI system. F or example, the left atrial size is a demonstrated predictor for AF recur- rence and through the use of NLP extraction from disc harge rep orts, this feature ac hiev ed a recall of 41.8%, which is remark able given that not all patien ts hav e this information do cumen ted in their discharge summaries. Missing v alues in EHRs can arise from m ultiple sources beyond human error. They ma y result from data in tegration issues across systems, v ariations in clinical do cumen tation practices, or patient-related factors suc h as refusal or missed tests. System design lim- itations, such as non-mandatory fields also migh t con tribute, as do temp oral gaps when data is pending or historical records are un- a v ailable. Ov erall, the enrichmen t pro cess not only enhances data completeness but also impro ves the representativ eness of clinical v ariables, providing a stronger foundation for do wnstream predictive mo deling. 5.2 Automatic Lab eling The automatic lab eling approac h for pa- tien ts’ AF progression status demonstrated an accuracy of 0.82 relative to the manu- ally annotated test set. Achieving an accu- racy of 0.82 is notable, esp ecially considering that the manual annotations incorp orate the full sp ectrum of patien t information, includ- ing electro cardiograms and other details that ma y not be fully do cumen ted in the disc harge rep orts. This means our metho dology is able to capture and lab el patien t AF progression status accurately despite the p ossible incom- pleteness of the discharge data. 5.3 AF Progression Prediction T o further assess the utility of our metho d- ology and its application in future studies of early prediction of diseases w e p erformed three exp eriments for AF progression predic- tion. The three exp erimen ts in volv e different v ersions of the generated dataset to ev alu- ate how the prop osed metho dology impacts the results of AF progression. The first ex- p erimen t uses the original dataset (1023 in- stances) extracted solely from co dified EHRs with the silv er annotation. The second ex- p erimen t uses the enriched dataset (1023 in- stances), whic h combines co dified informa- tion and discharge rep ort data along with our silv er-automatic annotation. Finally , the third exp eriment uses a smaller subset of the enriched dataset (541 instances) that has b een man ually annotated with gold-standard lab els by a cardiologist. As explained in section 4.4, w e use the T abPFN architecture for the three exp eri- men ts. The results are av ailable in T able 2. Dataset Accuracy MCC Original-Silver 0.65 0.11 Enriche d-Silver 0.66 0.20 Enriche d-Gold 0.66 0.21 T able 2: Results obtained in the AF pro- gression exp erimen ts. Using the original dataset with silver an- notations derived solely from co dified EHRs, the mo del ac hieved a mo derate accuracy of 0.65 and a low MCC of 0.11, indicating lim- ited predictiv e p o wer when relying exclu- siv ely on structured EHR data. Incorp orat- ing disc harge rep ort information in the en- ric hed dataset with silver-automatic anno- tations led to an impro vemen t in accuracy (0.66) and doubled the MCC (0.20), suggest- ing that integrating unstructured clinical text adds relev an t information. In medical predic- tion tasks, this increase is particularly mean- ingful, as MCC reflects the o verall reliabil- it y of the mo del across all p ossible outcomes. Doubling the MCC indicates a substantial impro vemen t in the mo del’s ability to cor- rectly iden tify b oth patien ts at risk and those not at risk, whic h is crucial in clinical settings where misclassification can lead to missed di- agnoses or unnecessary treatments. Finally , using the manually annotated gold-standard subset resulted in a similar ac- curacy of 0.66, with a slight increase in MCC to 0.21. Although this subset contains almost half the num b er of instances, the higher- qualit y annotations improv e the correlation b et w een predictions and true lab els. How- ev er, man ual annotation is time-consuming and requires exp ert kno wledge. Therefore, an automatic metho d that achiev es compa- rable p erformance, even if it requires more instances, remains adv an tageous. The predictive mo dels consistently out- p erform the traditional clinical scores in b oth accuracy and MCC (see T able 3). Both CHADS2-V ASc and HA TCH achiev ed an ac- curacy of 0.60 with very lo w MCC v alues (–0.0052 and 0.0832, resp ectiv ely). APPLE p erformed worse, with an accuracy of 0.48 and an MCC of 0.05. A CC MCC CHADS2-V ASc 0.6043 -0.0052 HA TCH 0.6043 0.0832 APPLE 0.4820 0.0510 T able 3: Results obtained in the AF progression exp eriments b y the clinical scores. These results highligh t that data-driven predictiv e mo dels, even with automatic anno- tations, can capture patterns in AF progres- sion that traditional clinical scores fail to de- tect. While clinical scores are useful for quick risk stratification, their limited consideration of patien t-sp ecific information and simplified scoring rules result in lo wer predictive p er- formance. In con trast, integrating structured EHR data with textual discharge rep orts al- Figure 6: Amount of missing v alues. Difference in p ercen tage b et ween the original and enric hed datasets. The bar plots illustrate the recov ery of features that were absen t in the original co dified dataset but retrieved from the information contained in the discharge rep orts. lo ws the mo dels to leverage a ric her set of features, yielding more reliable predictions, particularly for imbalanced outcomes like AF progression. This suggests that such mo d- els could serve as v aluable decision-supp ort to ols, complementing or enhancing existing clinical scores. 6 Conclusions This study provides evidence that discharge rep orts, when pro cessed with NLP tech- niques, can pla y a significant role in supp ort- ing early prediction tasks and enhancing clin- ical data quality . Addressing eac h research question: R Q1: Can the com bination of struc- tured EHR data and information ex- tracted from disc harge rep orts en- hance and automate the dev elopmen t of early disease prediction mo dels? The study demonstrates that integrating struc- tured EHR data with NLP extracted in- formation from disc harge rep orts b oth fa- cilitates automation of the data prepara- tion pip eline and yields measurable impro ve- men ts in mo del p erformance and data com- pleteness. The prop osed pip eline automates cohort selection, tabular feature generation and outcome lab eling b y combining codified EHR vectors with features extracted from free text. Practically , this automation re- duces the need for man ual case review and man ual lab el assignment, streamlining the end to end process required to build early prediction mo dels. In the AF progression ex- p erimen ts the enriched dataset that merged co dified data with disc harge rep ort features pro duced a higher MCC than the co dified dataset alone (MCC 0.20 versus 0.11) while accuracy rose sligh tly from 0.65 to 0.66. Au- tomatic lab eling via the pip eline also pro- duced a high agreement with manual anno- tation, with an automatic lab eling accuracy of 0.82 on the manually annotated test set. These results indicate that the in tegrated approac h b oth automates previously manual steps and supplies additional predictive sig- nal for the mo del. R Q2: Can free-text discharge re- p orts pro cessed with NLP techniques impro v e the quality and completeness of structured tabular data deriv ed from EHRs The integration of NLP-extracted in- formation with existing tabular data sub- stan tially reduces missingness and increases recall for many clinically relev an t v ariables that are underrepresen ted in co dified fields. The article rep orts a clear reduction in the prop ortion of missing v alues for many lab- oratory and categorical features after en- ric hment with disc harge rep ort information. Lab oratory v ariables that originally had high missingness, such as albumin, C reactiv e protein and NT proBNP , show ed marked increases in av ailability . Similarly , recall for past medical history and treatment fea- tures rose noticeably in the enriched dataset. The V ectorMerger approach pro duces uni- fied patient vectors where features absent from the co dified system are reco v ered from text, improving completeness and represen- tativ eness. This adresses the gaps caused b y system limitations, do cumen tation v ari- abilit y , or patien t-related factors, the en- ric hment pro cess enhances data completeness and representativ eness, providing a more re- liable foundation for predictive mo delling. R Q3: Can automatically generated silv er annotations ac hieve p erformance comparable to gold-standard annota- tions while significantly reducing man- ual effort? Automatically generated sil- v er annotations achiev e performance close to gold-standard annotations in mo del ev alua- tion and greatly reduce man ual effort. The similarit y of the enriched-silv er and enriched- gold results, esp ecially in MCC, indicates that silver annotations can approach gold p erformance in this problem setting. Giv en comparable mo del p erformance and that man ual gold lab eling is costly and time con- suming, silv er lab eling p ermits larger train- ing sets and faster iteration. R Q4: Do es the prop osed metho d- ology pro duce predictive mo dels that outp erform existing clinical scores for AF progression prediction? Mo dels trained with the enriched dataset outp er- form traditional clinical scores for AF pro- gression on b oth accuracy and MCC. Across the exp erimen ts, data driven models consis- ten tly outp erformed the clinical scores con- sidered. The T abPFN model trained on the enric hed dataset achiev ed accuracy 0.66 and MCC 0.20. By con trast, CHADS2 V ASc and HA TCH each had accuracy appro ximately 0.60 with v ery lo w or near zero MCC v alues. The APPLE score performed w orse with ac- curacy 0.48. These results indicate that ML mo dels capture predictive patterns that sim- plified clinical scores do not. Ov erall, this study highligh ts the p oten- tial of combining structured EHR data with NLP-extracted information from discharge rep orts to enhance early prediction tasks. Suc h an approach not only supp orts more robust and comprehensive early prediction mo dels for AF progression but also lays the groundw ork for applying similar methodolo- gies to other diseases, ultimately contribut- ing to more informed and data-driv en clinical decision-making. 7 F utur e Work The current study was limited to AF pro- gression; applying the same pip eline to other c hronic and acute conditions will b e essen- tial to assess its generalizability and robust- ness. In addition, external v alidation using datasets from differen t hospitals and health- care systems will help ev aluate the metho d- ology under diverse do cumen tation practices and data standards. Expanding the study to m ultiple institutions and inv olving a larger n umber of exp ert annotators w ould further strengthen the robustness and external v a- lidit y of the prop osed framework. How ev er, generating high qualit y exp ert lab els is a time consuming and resource in tensive pro cess, whic h was b ey ond the scop e of the present w ork. Therefore, broader exp ert inv olv emen t and multi cen ter v alidation are imp ortant di- rections for future research. F uture work will also explore the incor- p oration of adv anced NLP approac hes, in- cluding large language mo dels (LLMs), in to the existing pipeline. The rapid ev olution of these arc hitectures offers promising opp ortu- nities to further enhance the accuracy of fea- ture extraction and automatic lab eling. In the sp ecific context of AF progression prediction, future research could inv estigate m ultimo dal approac hes that pro cess struc- tured tabular data and unstructured clini- cal text jointly when estimating patient risk. Moreo ver, in tegrating temp oral information from longitudinal patien t histories ma y im- pro ve the mo deling of disease dynamics and lead to more precise and clinically meaningful predictions. A cknow le dgements This w ork has b een partially sup- p orted b y the HiTZ Cen ter and the Basque Gov ernmen t, Spain (Research group funding IT1570-22) as well as by MCIN/AEI/10.13039/5011 00011033 Span- ish Ministry of Universities, Science and Inno v ation b y means of the pro jects: EDHIA PID2022-136522OB-C22 (also supp orted b y FEDER, UE). A. G. Domingo-Aldama has been funded b y the Predo ctoral T raining Program for Non-PhD Researc h Personnel grant of the Basque Go vernmen t (PRE 2024 1 0224). A. Garc ´ ıa Olea has b een funded b y BioBizk aia gran t under the co de BB/I/PMIR/24/001 Biblio gr af ´ ıa Alzubi, A. A., V. J. W atzlaf, and P . Sheri- dan. 2021. Electronic health record (ehr) abstraction. Persp e ctives in he alth infor- mation management , 18(Spring). Aww alu, J., A. G. Garba, A. Ghazvini, and R. A tuah. 2015. Artificial intelligence in p ersonalized medicine application of ai al- gorithms in solving p ersonalized medicine problems. International Journal of Com- puter The ory and Engine ering , 7(6):439. Botsis, T., G. Hartvigsen, F. Chen, and C. W eng. 2010. Secondary use of ehr: data qualit y issues and informatics opp or- tunities. Summit on tr anslational bioin- formatics , 2010:1. Brahier, M. S., F. Zou, M. Ab dulk areem, S. Ko chi, F. Migliarese, A. Thomaides, X. Ma, C. W u, V. Sandfort, P . J. Bergquist, et al. 2023. Using machine learning to enhance prediction of atrial fibrillation recurrence after catheter abla- tion. Journal of A rrhythmia , 39(6):868– 875. Chen, H., X. Li, X. He, A. Chen, J. McGill, E. C. W ebber, H. Xu, M. Liu, and J. Bian. 2025. Enhancing patien t-trial matc hing with large language mo dels: A scoping review of emerging applications and ap- proac hes. JCO Clinic al Canc er Informat- ics , 9:e2500071. Cruz-Correia, R. J., P . P . Ro drigues, A. F re- itas, F. C. Almeida, R. Chen, and A. Costa-P ereira. 2009. Data qual- it y and integration issues in electronic health records. In Information disc overy on ele ctr onic he alth r e c or ds . Chapman and Hall/CR C, pages 73–114. De la Iglesia, I., A. S´ anc hez-F reire, O. Urquijo-Dur´ an, A. Barrena, and A. Atutxa. 2025. Eriberta priv ate sur- passes her public alter ego: Enhancing a bilingual pretrained enco der with limited priv ate medical data. Pr o c esamiento del L enguaje Natur al , 75:283–296. de la Iglesia, I., M. Viv´ o, P . Cho cr´ on, G. de Maeztu, K. Go jenola, and A. Atutxa. 2023. An op en source corpus and automatic to ol for section iden tification in spanish health records. J. Biome d. Informatics , 145:104461. De V os, C. B., R. Pisters, R. Nieuwlaat, M. H. Prins, R. G. Tieleman, R.-J. S. Coe- len, A. C. v an den Heijk an t, M. A. Al- lessie, and H. J. Crijns. 2010. Progression from paroxysmal to persistent atrial fib- rillation: clinical correlates and prognosis. Journal of the Americ an Col le ge of Car- diolo gy , 55(8):725–731. F an, X., Y. Li, Q. He, M. W ang, X. Lan, K. Zhang, C. Ma, and H. Zhang. 2023. Predictiv e v alue of machine learning for recurrence of atrial fibrillation after catheter ablation: A systematic review and meta-analysis. R eviews in Car diovas- cular Me dicine , 24(11):315. F eder, S. L. 2018. Data quality in electronic health records research: quality domains and assessmen t metho ds. Western journal of nursing r ese ar ch , 40(5):753–766. Garcia Olea, A., J. Ormaetxe Mero- dio, A. Atutxa Salazar, I. Diez Gon- zalez, I. F ernandez De La Prieta, M. Maeztu Rada, E. Amuriza De Luis, K. Ugedo Alzaga, U. Idiazabal Ro driguez, I. Pereiro Lili, et al. 2021. The role of con- gestiv e heart failure at atrial fibrillation onset in the data entry errors of electronic health records. In EUROPEAN JOUR- NAL OF HEAR T F AILURE , volume 23, pages 303–304. WILEY 111 RIVER ST, HOBOKEN 07030-5774, NJ USA. Garc ´ ıa-Olea, A., A. G. Domingo-Aldama, M. Merino, K. Go jenola, J. Goikoetxea, A. Atutxa, and J. M. Ormaetxe. 2025. The application of deep learning to ols on medical rep orts to optimize the input of an atrial-fibrillation-recurrence predic- tiv e model. Journal of Clinic al Me dicine , 14(7):2297. Guan, Z., Z. W u, Z. Liu, D. W u, H. Ren, Q. Li, X. Li, and N. Liu. 2023. Cohort- gpt: An enhanced gpt for participan t re- cruitmen t in clinical study . Hollmann, N., S. M ¨ uller, L. Puruck er, A. Kr- ishnakumar, M. K¨ orfer, S. B. Hoo, R. T. Sc hirrmeister, and F. Hutter. 2025. Ac- curate predictions on small data with a tabular foundation mo del. Natur e , 637(8045):319–326. Holmes, J. H., J. Beinlic h, M. R. Boland, K. H. Bowles, Y. Chen, T. S. Co ok, G. Demiris, M. Draugelis, L. Fluharty , P . E. Gabriel, et al. 2021. Why is the electronic health record so c hallenging for researc h and clinical care? Metho ds of information in me dicine , 60(01/02):032– 048. Hulme, O. L., S. Khurshid, L.-C. W eng, C. D. Anderson, E. Y. W ang, J. M. Ashburner, D. Ko, D. D. McMan us, E. J. Benjamin, P . T. Ellinor, et al. 2019. Developmen t and v alidation of a prediction mo del for atrial fibrillation using electronic health records. JA CC: Clinic al Ele ctr ophysiol- o gy , 5(11):1331–1341. Ibrahim, J. G., H. Chu, and M.-H. Chen. 2012. Missing data in clinical studies: is- sues and metho ds. Journal of clinic al on- c olo gy , 30(26):3297–3303. Jetley , G. and H. Zhang. 2019. Elec- tronic health records in is research: Qual- it y issues, essential thresholds and reme- dial actions. De cision Supp ort Systems , 126:113137. Jin, Q., Z. W ang, C. S. Floudas, F. Chen, C. Gong, D. Brack en-Clark e, E. Xue, Y. Y ang, J. Sun, and Z. Lu. 2024. Matc hing patients to clinical trials with large language mo dels. Natur e c ommuni- c ations , 15(1):9074. Kahale, L. A., A. M. Khamis, B. Diab, Y. Chang, L. C. Lop es, A. Agarw al, L. Li, R. A. Mustafa, S. Koujanian, R. W aziry , et al. 2020. Poten tial impact of missing outcome data on treatment effects in sys- tematic reviews: imputation study . bmj , 370. Knec ht, S., J. Cyriac, P . Badertsc her, P . Kri- sai, V. Sc hlageter, S. Ossw ald, M. Zell- w eger, M. Kuhne, and C. Sticherling. 2024. Machine learning for outcome pre- diction of atrial fibrillation recurrence af- ter catheter ablation. Eur op ac e , 26(Sup- plemen t 1):euae102–556. Kornej, J., G. Hindricks, M. B. Sho e- mak er, D. Husser, A. Arya, P . Sommer, S. Rolf, P . Saa v edra, A. Kanagasundram, S. Patric k Whalen, et al. 2015. The apple score: a no v el and simple score for the pre- diction of rhythm outcomes after catheter ablation of atrial fibrillation. Clinic al R e- se ar ch in Car diolo gy , 104:871–876. Lewis, A. E., N. W eiskopf, Z. B. Abrams, R. F oraker, A. M. Lai, P . R. Pa yne, and A. Gupta. 2023. Electronic health record data qualit y assessmen t and to ols: a systematic review. Journal of the A meric an Me dic al Informatics Asso cia- tion , 30(10):1730–1740. Lip, G. Y., R. Nieuwlaat, R. Pisters, D. A. Lane, and H. J. Crijns. 2010. Refin- ing clinical risk stratification for predict- ing stroke and throm b o em b olism in atrial fibrillation using a nov el risk factor-based approac h: the euro heart survey on atrial fibrillation. Chest , 137(2):263–272. Liu, C.-M., W.-S. Chen, S.-L. Chang, Y.- C. Hsieh, Y.-H. Hsu, H.-X. Chang, Y.- J. Lin, L.-W. Lo, Y.-F. Hu, F.-P . Ch ung, et al. 2024. Use of artificial intelligence and i-score for prediction of recurrence b efore catheter ablation of atrial fibrilla- tion. International Journal of Car diolo gy , 402:131851. McDa vid, A., P . K. Crane, K. M. Newton, D. R. Crosslin, W. McCormic k, N. W e- ston, K. Ehrlich, E. Hart, R. Harrison, W. A. Kukull, et al. 2013. Enhancing the pow er of genetic asso ciation studies through the use of silver standard cases deriv ed from electronic medical records. PloS one , 8(6):e63481. Mishra, J. and S. T arar. 2020. Chronic disease prediction using deep learning. In A dvanc es in Computing and Data Scienc es: 4th International Confer enc e, ICA CDS 2020, V al letta, Malta, April 24– 25, 2020, R evise d Sele cte d Pap ers 4 , pages 201–211. Springer. Nadara jah, R., J. W u, A. F. F rangi, D. Hogg, C. Co wan, and C. Gale. 2021. Predict- ing patient-lev el new-onset atrial fibril- lation from p opulation-based nationwide electronic health records: protocol of find- af for dev eloping a precision medicine pre- diction mo del using artificial in telligence. BMJ op en , 11(11):e052887. Ng, K., S. R. Steinhubl, C. DeFilippi, S. Dey , and W. F. Stewart. 2016. Early de- tection of heart failure using electronic health records: practical implications for time before diagnosis, data diversit y , data quan tity , and data density . Cir cula- tion: Car diovascular Quality and Out- c omes , 9(6):649–658. Qiu, Y., H. Guo, S. W ang, S. Y ang, X. Peng, D. Xiay ao, R. Chen, J. Y ang, J. Liu, M. Li, et al. 2024. Deep learning-based multi- mo dal fusion of the surface ecg and clin- ical features in prediction of atrial fibril- lation recurrence follo wing catheter abla- tion. BMC Me dic al Informatics and De ci- sion Making , 24(1):225. Ristevski, B. and M. Chen. 2018. Big data analytics in medicine and health- care. Journal of inte gr ative bioinformat- ics , 15(3):20170030. Sc hork, N. J. 2019. Artificial intelli- gence and p ersonalized medicine. Pr e ci- sion me dicine in Canc er ther apy , pages 265–283. Shah, V. 2018. Next-generation artifi- cial intelligence for p ersonalized medicine: Challenges and innov ations. INTERNA- TIONAL JOURNAL OF COMPUTER SCIENCE AND TECHNOLOGY , 2(2):1– 15. Sharma, D. K., M. Chatterjee, G. Kaur, and S. V avilala. 2022. Deep learning applica- tions for disease diagnosis. In De ep le arn- ing for me dic al applic ations with unique data . Elsevier, pages 31–51. Sion tis, K. C., X. Y ao, J. P . Pirruccello, A. A. Philippakis, and P . A. Nosew orth y . 2020. Ho w will mac hine learning inform the clin- ical care of atrial fibrillation? Cir culation r ese ar ch , 127(1):155–169. Soni, S. and K. Rob erts. 2021. Patien t co- hort retriev al using transformer language mo dels. In AMIA annual symp osium pr o- c e e dings , v olume 2020, page 1150. Sterne, J. A., I. R. White, J. B. Carlin, M. Spratt, P . Royston, M. G. Kenw ard, A. M. W o o d, and J. R. Carp en ter. 2009. Multiple imputation for missing data in epidemiological and clinical research: p o- ten tial and pitfalls. Bmj , 338. Stubbs, A., M. Filannino, E. So ysal, S. Henry , and ¨ O. Uzuner. 2019. Co- hort selection for clinical trials: n2c2 2018 shared task track 1. Journal of the Americ an Me dic al Informatics Asso- ciation , 26(11):1163–1171. Sung, S.-F., K.-L. Sung, R.-C. Pan, P .-J. Lee, and Y.-H. Hu. 2022. Automated risk assessmen t of newly detected atrial fib- rillation p oststrok e from electronic health record data using machine learning and natural language pro cessing. F r ontiers in Car diovascular Me dicine , 9:941237. T erry , A. L., M. Stew art, S. Cejic, J. N. Mar- shall, S. de Lusignan, B. M. Chesworth, V. Chev endra, H. Maddo cks, J. Shadd, F. Burge, et al. 2019. A basic mo del for assessing primary health care electronic medical record data quality . BMC me dic al informatics and de cision making , 19:1–11. Tiw ari, P ., K. L. Colb orn, D. E. Smith, F. Xing, D. Ghosh, and M. A. Rosenberg. 2020. Assessment of a mac hine learning mo del applied to harmonized electronic health record data for the prediction of inciden t atrial fibrillation. JAMA network op en , 3(1):e1919396–e1919396. Tseng, A. S. and P . A. Nosew orthy . 2021. Prediction of atrial fibrillation using ma- c hine learning: a review. F r ontiers in Physiolo gy , 12:752317. Ullah, M., A. Akbar, and G. G. Y annarelli. 2020. Applications of artificial in telligence in early detection of cancer, clinical diag- nosis and p ersonalized medicine. v an Breugel, B. and M. v an der Sc haar. 2024. Wh y tabular foundation models should be a research priority . Pr o c e e dings of the 41st International Confer enc e on Machine L e arning , 235:48976–48993, 21–27 Jul. Vydisw aran, V. V., A. Stra yhorn, X. Zhao, P . Robinson, M. Agarw al, E. Bagazinski, M. Essiet, B. E. Iott, H. Jo o, P . Ko, et al. 2019. Hybrid bag of approaches to char- acterize selection criteria for cohort iden- tification. Journal of the A meric an Me di- c al Informatics Asso ciation , 26(11):1172– 1180. W agholik ar, K. B., H. Estiri, M. Murph y , and S. N. Murphy . 2020. Polar lab eling: sil- v er standard algorithm for training disease classifiers. Bioinformatics , 36(10):3200– 3206. Xie, S., Z. Y u, and Z. Lv. 2021. Multi-disease prediction based on deep learning: a sur- v ey . Computer Mo deling in Engine ering & Scienc es , 128(2):489–522. Y u, Z., K. W ang, Z. W an, S. Xie, and Z. Lv. 2023. Popular deep learning algorithms for disease prediction: a review. Cluster Computing , 26(2):1231–1251. Zhou, X., K. Nak am ura, N. Sahara, T. T ak- agi, Y. T o yoda, Y. Enomoto, H. Hara, M. Noro, K. Sugi, M. Moroi, et al. 2022. Deep learning-based recurrence prediction of atrial fibrillation after catheter abla- tion. Cir culation Journal , 86(2):299–308.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment