Colon-Bench: An Agentic Workflow for Scalable Dense Lesion Annotation in Full-Procedure Colonoscopy Videos
Early screening via colonoscopy is critical for colon cancer prevention, yet developing robust AI systems for this domain is hindered by the lack of densely annotated, long-sequence video datasets. Existing datasets predominantly focus on single-clas…
Authors: Abdullah Hamdi, Changchun Yang, Xin Gao
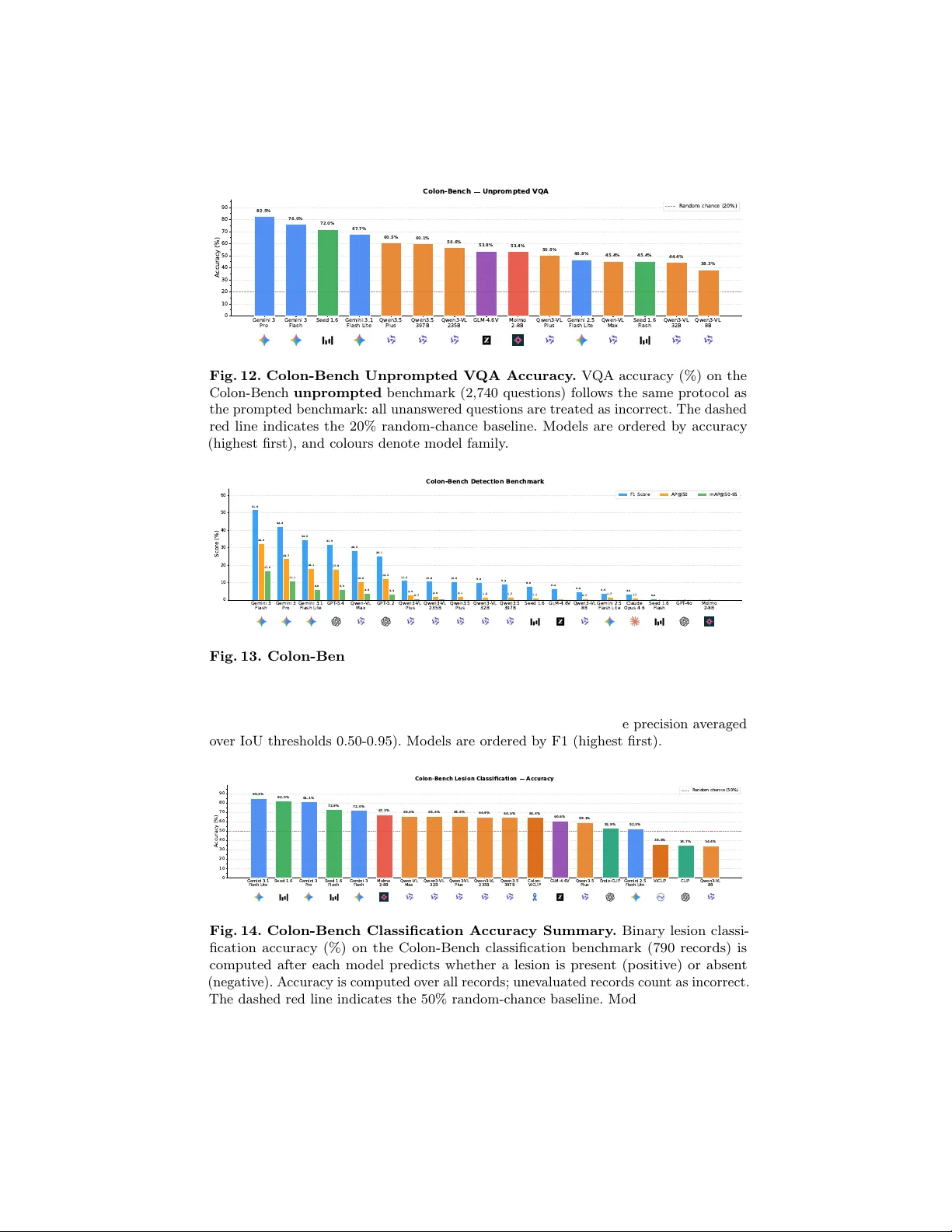
Colon-Benc h: An Agen tic W orkflo w for Scalable Dense Lesion Annotation in F ull-Pro cedure Colonoscop y Videos Ab dullah Hamdi, Changch un Y ang, and Xin Gao King Abdullah Universit y of Science and T ec hnology (KAUST) {abdullah.hamdi@kaust.edu.sa} Abstract. Early screening via colonoscopy is critical for colon cancer prev ention, y et developing robust AI systems for this domain is hin- dered by the lac k of densely annotated, long-sequence video datasets. Existing datasets predominantly fo cus on single-class p olyp detection and lack the ric h spatial, temp oral, and linguistic annotations required to ev aluate modern Multimo dal Large Language Mo dels (MLLMs). T o address this critical gap, we introduce Colon-Bench, generated via a no vel m ulti-stage agentic workflo w. Our pip eline seamlessly integrates temp oral prop osals, b ounding-box tracking, AI-driv en visual confirmation, and h uman-in-the-lo op review to scalably annotate full-pro cedure videos. The resulting verified b enchmark is unprecedented in scop e, encompassing 528 videos, 14 distinct lesion categories (including polyps, ulcers, and bleeding), ov er 300,000 b ounding b o xes, 213,000 segmentation masks, and 133,000 words of clinical descriptions. W e utilize Colon-Bench to rigorously ev aluate state-of-the-art MLLMs across lesion classification, Op en-V ocabulary Video Ob ject Segmen tation (O V-VOS), and video Visual Question Answering (VQA). The MLLM results demonstrate sur- prisingly high lo calization p erformance in medical domains compared to SAM-3. Finally , we analyze common VQA errors from MLLMs to in tro duce a nov el "colon-skill" prompting strategy , improving zero-shot MLLM p erformance by up to 9.7% across most MLLMs. The dataset and the code are a v ailable at h ttps://ab dullahamdi.com/colon- b enc h Keyw ords: colonoscop y · lesion detection · p olyps · MLLMs. 1 In tro duction Colon cancer is the second leading cause of cancer death worldwide [ 23 ], and many of these deaths are prev entable with early screening and remov al of precancerous lesions; ho wev er, colonoscopy remains exp ensiv e, inv asive, and logistically di fficult to arrange, requiring sp ecialized clinicians and often long pro cedures (up to t wo hours) that increase facilit y and staffing costs. Automated AI analysis promises to ease these burdens, but lesion discov ery is a needle-in-a-haystac k problem: lesions are sparse and frequently obscured b y motion blur, o cclusion, 2 H. Abdullah et al. T able 1. Comparison of Colonoscopy Datasets. Relativ e to prior datasets that emphasize single-class p olyp detection or anatomical segmentation, Colon-Bench pro- vides a broader lesion taxonomy and richer sup ervision. It combines long-sequence co verage with dense b o xes, masks, and clinical text, enabling multi-task ev aluation across detection, video segmentation, and language-based understanding. Attribute K v asir-SEG [ 14 ] SUN [ 19 ] PolypGen [ 1 ] REAL-Col. [ 3 ] CAS-Col. [ 22 ] Colon-Bench (Ours) Y ear 2020 2021 2023 2024 2025 2026 F ocus Segmentation Detection Segmen tation Detection Anatomy Lesion, Video Seg., & T ext Number of Videos N/A N/A N/A 60 78 528 T otal F rames/Images 1,000 158,690 6,282 2,757,723 1,961,100 464,035 Lesion Classes 1 (Polyp) 1 (Polyp) 1 (Polyp) 1 (P olyp) N/A 14 (Bleeding, Polyps, ...) Bounding Boxes N/A 158,690 6,282 351,264 N/A 300,132 Segmentation Masks 1,000 N/A 6,282 N/A Y es 213,067 Language Descriptions N/A N/A N/A N/A N/A Y es (133k W ords) debris, sto ol, or fluids, and b y camera contact with the colon wall, making dense visual analysis challenging. Moreov er, manual dense annotation of colonoscopy videos is labor-intensiv e and inconsisten t, whic h motiv ates our work on a scalable, affordable pip eline for dense colonoscop y video annotation, facilitating AI research, ev aluation, and applications in colonoscopy . T o address the inherent visual c hallenges of colonoscopy videos, recent meth- o ds hav e prop osed sp ecialized architectures optimized for pixel-lev el precision and subtle lesion identification [ 27 , 17 ]. F urthermore, b ecause colonoscopy pro ce- dures are inherently long, efficient spatiotemp oral analysis is critical; approaches utilizing state-space mo dels [ 24 ] and token merging [ 26 ] hav e b een introduced to capture long-range dep endencies while reducing computational redundancy . Man y methods rely on self-supervised learning with large-scale pretraining [ 15 ] or utilize a small num ber of annotated image examples with text [ 13 ], b ounding b o xes [ 7 , 16 ], or segmen tation masks [ 14 , 7 ]. These arc hitectural and synthetic w ork arounds underscore the critical gap in the field that our work addresses: the need for a comprehensiv e, densely annotated, long-sequence dataset to ground the spatiotemp oral analysis of real colonoscopies. REAL-COLON [ 3 ] introduced 60 long sequences of colonoscopy procedures. These colonoscopies are lab eled with 351,264 b ounding b oxes on p olyps using commercial softw are. CAS-COLON [ 22 ] introduced a colonoscopy anatomical temp oral classification (which part of the colon is each image) with a total of 78 videos totaling 9 hours and around 2 million images. Other colonoscopy datasets are either small in size for comprehensive training and ev aluations [ 18 ], or they co ver large image datasets but are limited in the scop e of tasks and t yp es of les ions that they inv estigate [ 14 , 19 , 18 , 1 ]. As T able 1 shows, our Colon- Benc h offers a dense annotation scheme for lesions (with text descriptions and segmen tation mask trac klets) across a large v ariety of video clips and provides comprehensiv e ev aluation b enc hmarks (including Video Question Answering, lesion clip classification, and Op en-V o cabulary Video Segmentation) for MLLMs as a general to ol for colonoscopy video understanding. The rapid evolution of Large Language Mo dels (LLMs) [ 20 , 11 ] and foun- dational vision-language mo dels [ 21 ] has driven the developmen t of Multi- mo dal Large Language Mo dels (MLLMs) capable of complex visual reasoning Title S uppressed Due to Excessive Length 3 Fig. 1. Colon-Benc h Annotation Pip eline. Overview of the m ulti-stage agen tic w orkflow used to build the dataset, from VLM prop osal generation through verification, trac king with spatial annotations, AI confirmation, and clinician review. The figure highligh ts how successive filters reduce false p ositives while adding dense spatial and textual lab els for the retained windows, all easily verified b y a physician at the end. The accepted annotations are then used to create a comprehensive Colon-Benc h for MLLMs co vering multiple video tasks: Visual Quesion Answering (VQA), binary lesion classification, and Op en-V o cabulary Video Ob ject Segmentation (OV-V OS). [ 2 , 8 , 10 , 12 , 11 ]. While these MLLMs demonstrate remark able zero-shot compre- hension in general domains, their proficiency in in terpreting long, o ccluded, and highly noisy medical sequences remains largely untested. This underscores the critical need for our prop osed Colon-Bench to systematically ev aluate how w ell these adv anced video-language mo dels understand complex colonoscopy pro- cedures. Recent works sho w strong results with agentic MLLM pip elines for long-form videos [ 9 , 25 ] and for hybrid automatic labeling with human v erification for large medical image datasets [ 6 ], motiv ating our pip eline in Fig. 1 . W e prov ide a new annotated dataset of long sequences with dense annotations. It includes the first Op en-V o cabulary Video Ob ject Segmentation (OV-V OS) dataset in colonoscop y that inv olv es mask tracking of lesions even with occluding frames. Our con tributions can b e summarized as follows: Con tributions: (i) W e in tro duce a nov el, m ulti-stage agen tic workflo w for the dense annotation of full-pro cedure colonoscopy videos. By combining temp o- ral prop osals, b ounding-box trac king, AI-driv en confirmation with visual cues, and an efficient human-in-the-loop review in terface, our pip eline dramatically reduces manual lab or while yielding high-quality spatial, temp oral, and tex- tual annotations. (ii) W e present a comprehensive, m ulti-task video b enc hmark ( Colon-Bench ) for colonoscop y understanding. Spanning 14 distinct lesion cate- gories (including polyps, bleeding, and ulcers), the dataset comprises ov er 464k frames, 300k b ounding b o xes, 213k segmentation masks, and 133k words of 4 H. Abdullah et al. T able 2. Annotation Pip eline Stages. W e show Colon-Bench annotations after each ma jor processing stage. As automated filters and human review progressively discard false-p ositiv e windows (reducing total frames), the temp oral precision, F1 score, and sp ecificit y of the automatically identified lesions demonstrably increase on the surrogate p olyp lab els of REAL-COLON [ 3 ] (p olyps are one type of lesion in Colon-Benc h). Initial VLM + v erification + Cued AI Final Human- Metric Proposals Agent Filtering Confirmation Agent Curated Annots. T otal Windows 1,325 903 597 528 T otal F rames 826,763 648,440 492,606 464,035 Duration (Hours) 22.97 18.01 13.68 12.89 T otal Bounding Boxes - - 314,408 300,132 T otal Seg. Masks - - 227,343 213,067 T otal Clinical W ords - - 145,515 133,289 Precision 30.9 36.2 53.1 55.4 Recall 67.5 68.7 48.8 48.6 F1 Score 42.4 47.4 50.8 51.8 Specificity 78.6 82.9 93.9 94.4 clinical descriptions, all v erified b y humans and an exp erienced surgeon. It facili- tates four rigorous ev aluation tasks: binary lesion classification, Op en-V o cabulary Video Ob ject Segmentation (OV-V OS), and t wo difficulty tiers of Video Vi- sual Question Answering (VQA). (iii) W e conduct a large-scale ev aluation of colonoscop y visual reasoning and localization in state-of-the-art MLLMs (e.g., Gemini 3, Qw en, Seed, GPT), fo cused on lesions. By analyzing error patterns across MLLMs, we prop ose a nov el “colon-skill” that distills cross-mo del error patterns into structured textual guidance, which w e sho w improv es most MLLM p erformance by up to 9.7% without any additional training (only adding the text skill to the prompt improv es performance). 2 Metho dology 2.1 Agen tic W orkflow for Colonoscopy Annotation Automatic annotations with quality control. As illustrated in Fig. 1 , our dense colonoscopy annotations w ere constructed by applying a m ulti-stage agen tic pip eline to 60 video sequences from the REAL-COLON dataset [ 3 ]. As detailed in T able 2 , an initial vision-language mo del iden tified 1,325 candidate lesion windo ws (22.97 hours). Successiv e v erification filtering agent, bounding-b ox trac king, cued AI confirmation agen t (using an o verla y of the box on the lesion as cues), and a final h uman review progressively filtered this set to isolate high-quality detections. By observing the dataset state at each stage (T able 2 ), w e demonstrate that automated AI stages provide the principal filtering gains on the surrogate p olyp lab els of REAL-COLON [ 3 ] (p olyps are one t yp e of lesion in Colon-Bench, but act as a quality control). Initial video verification follow ed by EdgeT AM [ 28 ] trac king (efficien t SAM-based tracking and segmentation) and AI confirmation sim ultaneously established the spatial annotations (yielding ov er 314k initial b ounding boxes) and radically pruned weak temp oral b oundaries. The recent Title S uppressed Due to Excessive Length 5 Sessile P olyp Bleeding Ulcer Erythematous SSL T umour / LST P edunculated P olyp Angioectasia Diverticulum Mucosal Abn. Cr ohn's Hemor r hoid P arasites Others 0 60 120 180 240 300 360 420 F r equency 411 252 160 112 86 85 72 55 51 51 7 5 4 1 Lesion Category Distribution Fig. 2. Lesion Category Distribution. Long-tailed lesion category distribution in Colon-Benc h, highlighting the div ersity of lesions. success of the Gemini-3 vision mo del for medical domains [ 4 ] has prompted the use of Gemini-3 as one to ol in the annotation pip eline. F or prop osals, video v erification agents, and b ounding b o xes, Gemini-2.5-flash-lite, Gemini-3-pro, and Gemini-3-flash w ere used, resp ectively , balancing performance and cost (T able 3 ). Human review. Human review served as the final quality gate. T o enable efficien t review at scale, the pipeline pre-rendered short clips with spatial ov erla ys in to an interactiv e web interface. A reviewing physician rejected only 69 of the 597 presented windows (11.6%), demonstrating strong agreemen t with the AI filters. Ultimately , the pip eline retained 528 curated windows (39.8% retention) spanning 464,035 frames. The final dataset yields a dense, multi-modal b enc hmark comprising 300,132 b ounding b o xes, 213,067 segmentation masks, and ov er 133k w ords of verified clinical text (a veraging 252.4 words p er windo w). Lesion t yp es. The dataset spans 14 lesion categories identified through m ulti- lab el keyw ord matching o ver clinician-v erified text fields (Fig. 2 ). The distribution is heavily long-tailed: sessile p olyps dominate (411 windo ws), follo wed by bleed- ing (252), ulcers (160), and erythematous lesions (112). Because Colon-b ecnh descriptions are free-form text (one windo w can hav e multiple lesions), a single review window ma y con tribute to more than one category . 2.2 Colon-Benc h Ev aluation Benchmark Filtering. Colon-Bench is a multi-task video b enchmark for colonoscopy un- derstanding comprising 1,597 clips from 60 patients (955,126 frames), spanning fiv e tasks: binary classification (790 clips), detection (272 clips, 61,538 p er-frame b ounding b oxes), instance segmentation (264 clips, 57,550 p er-frame masks), and V QA at t wo difficult y tiers. The binary lesion classes and segmentation masks follo w directly from Sec. 2.1 to establish the first colonoscopy Open-V ocabulary Video Ob ject Segmentation (O V-VOS) benchmark. V QA (prompted & unprompted) . The pr ompte d VQA split contains 1,485 fiv e-choice questions ov er 499 clips whose videos include b ounding-box ov erla ys 6 H. Abdullah et al. T able 3. Colon-Benc h Com bined Results. Summary of mo del p erformance across the four b enc hmark tasks: visual prompted and unprompted video Visual Question Answ ering (VQA) accuracy (with visual box prompts and without them), binary lesion classification precision/recall/F1, and open-vocabulary video segmentation IoU/Dice. The video segemtnation is based on 3 box detections for each MLLM prompting the same EdgeT AM track er [ 28 ]. Best scores p er metric are sho wn in b old . V QA Accuracy Lesion Cls. Segmen tation Model Prompted Unprompted Precision Recall F1 IoU Dice CLIP [ 21 ] - - 34.5 100 51.3 - - ViCLIP [ 25 ] - - 34.8 98.5 51.4 - - Endo-CLIP [ 13 ] - - 41.9 95.2 58.2 - - Colon-ViCLIP [ 25 ] - - 49.0 84.2 62.0 - - SAM 3 [ 5 ] - - - - - 2.5 2.9 GPT-4o - - - - - 0.5 0.8 Claude Opus 4.6 - - - - - 16.1 20.5 GPT-5.2 - - - - - 30.7 36.5 GPT-5.4 - - - - - 34.5 41.1 Qwen3-VL 8B 32.9 38.3 34.4 100 51.2 10.4 13.1 Seed 1.6 Flash 38.1 45.4 94.2 24.3 38.6 2.6 3.5 Qwen-VL Max 39.1 45.4 0.0 0.0 0.0 25.6 29.6 Qwen3-VL 32B 39.3 44.4 0.0 0.0 0.0 12.7 15.9 Qwen3.5 Plus 44.3 60.5 36.2 24.6 29.3 16.7 21.0 Molmo 2-8B 46.1 53.4 52.9 46.7 49.6 2.6 3.8 Qwen3-VL 235B 46.7 56.6 22.2 0.7 1.4 13.6 16.9 Gemini 2.5 F. Li. 47.8 46.8 42.3 95.9 58.7 19.9 24.3 Qwen3.5 397B 49.0 60.1 10.0 0.4 0.7 16.6 21.0 GLM-4.6V 55.7 53.8 46.5 94.1 62.2 12.5 16.1 Seed 1.6 62.9 72.0 85.0 58.7 69.4 12.6 16.1 Gemini 3.1 F. Li. 69.2 67.7 72.6 90.8 80.7 37.4 43.4 Gemini 3 Flash 76.6 76.0 55.3 97.1 70.5 48.3 54.7 Gemini 3 Pro 78.6 82.5 66.1 93.0 77.3 45.0 51.3 on confirmed lesion windows; the unpr ompte d split con tains 2,740 questions o ver 918 clips rendered from ra w frames, fully encompassing all detection and segmen tation videos. F or instance, an unprompted question for the clip in Fig. 3 first row asks: “How is the morpholo gy and anatomic al lo c ation of the identifie d lesion describ e d?” with distractors spanning p edunculated, depressed, and flat lesions at v arious anatomical sites; the correct answer is (D) a sessile p olyp on a haustr al fold . More examples are sho wn in the App endix . Debiasing Colon-Bench. F or eac h clip, three fiv e-wa y MCQs are generated with Gemini-3, cov ering lesion identification, clinical characteristics, and tem- p oral reasoning. T o mitigate text-only shortcuts, we apply tw o-stage debiasing: (i) adversarial distractors are regenerated in a separate LLM call receiving only the stem and correct answ er; (ii) a blind text-only stress-test identifies and reverts an y newly introduced bias. Post-debiasing blind accuracy is 44 . 6% (prompted) and 37 . 1% (unprompted) vs. the 20% random baseline, with residual margins reflecting dataset priors rather than surface cues. Title S uppressed Due to Excessive Length 7 Input GT GPT- 5.2 Opus 4.6 Gemini 3 Flash Qw en 3.5-Plus Qw en VL-Max Qw en 3 VL- 235B SAM 3 Fig. 3. Qualitative Comparisons. The first row shows detection ( Sessile Polyp ); the remaining rows show segmentation ( Sessile Polyp & Erythematous R e gion ). Columns sho w the input, ground truth, and mo del outputs. SAM-3 underperforms compared to mo dern MLLMs when paired with a prompted track er suc h as EdgeT AM [ 28 ]. Baselines. Metho ds rep orted in this w ork include a set of Multimo dal Large Language Mo dels (MLLMs) such as different v ariants of GPT s, Gemini, Qwen, Seed, GLM, and Molmo [ 20 , 2 , 8 , 10 , 12 , 11 ]. Also w e rep ort (for some corresp onding b enc hmarks) SAM-3 [ 5 ], CLIP [ 21 ], Video CLIP (ViCLIP) [ 25 ] and its fine-tuned v ersion on Colon-Bench text (Colon-CLIP), and a recent mo del Endo-CLIP [ 13 ]. Mo dles that do not hav e video API supp ort ( E.g. GPT-series) are only evluated on the segemntation task. All models in segem tnation (Except SAM-3) are based on 3-frame box detections prompting EdgeT AM trac ker [ 28 ]. 2.3 Colon-Benc h Results T able 3 reveals a clear p erformance hierarch y across all four Colon-Bench tasks. Gemini 3 Pro and Gemini 3 Flash consistently dominate, achieving the highest V QA accuracy on b oth prompted and unprompted splits and the b est segmen ta- tion quality (IoU/Dice of 45-48%/51-55%), while Gemini 3.1 Flash Lite leads in classification F1 (80.7%). Among op en-weigh t mo dels, Seed 1.6 is the strongest o verall, ranking third in V QA and delivering comp etitive classification F1 (69.4%) with the highest precision among full-b enc hmark mo dels (85.0%). The Qwen family shows mixed b ehavior: larger v arian ts (Qw en3.5 397B, Qwen3-VL 235B) p erform w ell on V QA yet nearly collapse on classification (R < 1%, F1 < 2%), sug- gesting they default to the ma jorit y (negative) b enign class rather than detecting lesion presence. F or segmentation, p erformance drops sharply b ey ond the top three mo dels. Overall, our Colon-Bench demonstrates how these strong MLLM mo dels can b e used for lesion detection in short clips, b eating sp ecialized mo dels lik e Endo-CLIP [ 13 ] by 30%. F or the task of op en-v o cabulary lesion segmentation in video clips, an MLLM like GPT-5.4 (when com bined with b o x-promptable EdgeT AM track er [ 28 ]) b eats SAM-3 [ 5 ] b y 32.0% mIoU (see Fig. 4 ). 8 H. Abdullah et al. Gemini 3 Flash Gemini 3 P r o Gemini 3.1 Flash Lite GPT -5.4 GPT -5.2 Qwen- VL Max Gemini 2.5 Flash Lite Qwen3.5 397B Claude Opus 4.6 Qwen3- VL 235B Qwen3- VL 32B Seed 1.6 Qwen3- VL 8B Molmo 2-8B S AM 3 GPT -4o 0 10 20 30 40 50 60 Scor e (%) 48.3 45.0 37.4 34.5 30.7 25.6 19.9 16.6 16.1 13.6 12.7 12.6 10.4 2.6 2.5 0.5 54.7 51.3 43.4 41.1 36.5 29.6 24.3 21.0 20.5 16.9 15.9 16.1 13.1 3.8 2.9 0.8 Colon-Bench Segmentation Benchmark Mean IoU Mean Dice Fig. 4. Multi-Mo dal Large Language Mo dels for Lesion Segmen tation. Video ob ject segmentation results on the 272-video b enc hmark, where MLLMs [ 20 , 2 , 8 , 10 , 12 , 11 ] first lo calize the target lesion with 3 b ounding b o xes and a EdgeT AM track er [ 28 ] propagates masks. 3 Analysis and Insigh ts 3.1 Colon-Skill for MLLMs T o test whether Multimo dal Large Language Mo dels (MLLMs) b enefit from structured domain knowledge at inference time, we use a tw o-stage skill-extraction and prompt-augmentation pip eline. First, we collect p er-mo del VQA predictions on the full benchmark and stratify errors by lesion category (Fig. 2 ), retaining only questions that a ma jorit y of mo dels answ er incorrectly . These shared failure cases, along with their question/answ er con text and category metadata, are fed to a fron tier large language mo del whic h synthesises a concise, natural-language Colon-Skil l : morphological cues, common confusion traps, and a decision chec klist tailored to colonoscopic VQA. Second, we prep end this Colon-Skill to every VQA prompt and re-ev aluate all mo dels under iden tical conditions, yielding consistent impro vemen ts up to +9 . 7% (Fig. 5 ). This suggests that distilling cross-mo del error patterns in to structured textual guidance is an effective, training-free wa y to improv e MLLM p erformance on sp ecialised medical VQA tasks, particularly when mo dels can in tegrate the added context. 3.2 Ablation Study Effect of frame count on segmen tation. As shown in Fig. 6 , increasing the num b er of input frames for detection yields steady gains in downstream segmen tation quality . F or instance, expanding the context from 1 to 7 frames b oosts the mean IoU of Gemini 3 Flash from 43.1% to 54.4% (mDice: 48.8% to 61.5%), and similarly improv es Qwen-VL Max from 19.7% to 33.7% mIoU. Ho wev er, this comes with a significant cost as these are p er-windo w detections. W e pick 3 equally-spaced frames p er window as a go o d trade-off in all of our segmen tation results in this work. T emporal context in V QA. Ev aluating temp oral context on the prompted V QA split sho wed that full video generally outperforms single-frame inputs, Title S uppressed Due to Excessive Length 9 though the degree of impact is mo del-dependent. Restricting the input to a single frame reduced accuracy for Qwen3.5 397B ( − 7 . 0 %), Qwen-VL Max ( − 3 . 4 %), and Gemini 3 Flash ( − 2 . 5 %). This highlights the imp ortance of using videos instead of images used in previous p olyp detection b enc hmarks. Gemini 3 Flash Qwen3.5 397B Qwen3- VL 235B Qwen3- VL Plus Qwen3- VL 32B Qwen3- VL 8B 0 20 40 60 80 A ccuracy (%) 76.6% 49.0% 46.7% 41.2% 39.3% 32.9% 78.5% 58.7% 50.1% 46.1% 43.5% 38.8% +1.8 pp +9.7 pp +3.4 pp +4.9 pp +4.2 pp +5.9 pp Baseline vs Colon-Skill Prompted VQA R andom chance (20%) Baseline W ith Colon-Skill Fig. 5. Colon-Skill on Prompted VQA. Skill-augmented prompting on the prompted V QA split, where a distilled error-based skill context yields consistent accuracy im- pro vemen ts for higher-capacit y models. 1 2 3 5 7 Detected frames count 10 20 30 40 50 Mean IoU (%) Mean IoU vs Detection F rames Count Gemini 3 Flash Qwen- VL Max Seed 1.6 Fig. 6. Ablation Study on Lesion Segmentation. W e ablate the num b er of de- tection frames p er windwo used in the segmentation b enc hmark, sho wing diminishing impro vemen t in do wnstream segmen tation quality . 4 Conclusions W e pres en ted Colon-Bench, a nov el agentic workflo w and a comprehensiv e multi- task b enchmark for full-pro cedure colonoscopy video understanding. By scalably 10 H. Abdullah et al. and densely annotating 14 distinct lesion categories with masks, bounding b o xes, and clinical text, we demonstrated that state-of-the-art MLLMs excel at high-level colonoscop y reasoning and spatial grounding with engineered context and agen tic w orkflows. F uture w ork will fo cus on optimizing these agent-driv en pip elines for real-time deploymen t in clinical settings, low ering the barrier to affordable and accurate colon cancer screening. A ckno wledgmen ts. W e thank Dr. Jamal Hamdi, a surgeon with ov er 30 y ears of exp erience, who reviewed the accuracy of the Colon-Benc h annotations together with the authors. This work was supp orted by the King Ab dullah Universit y of Science and T ec hnology (KAUST) Center of Excellence for Smart Health. A dditional support came from the KA UST Ibn Rushd Postdoctoral F ello wship Program. License. Colon-Bench is in tended strictly for academic research, and an y form of commercial use is prohibited. Copyrigh t for all videos is retained b y their owners [ 3 ]. The dataset as a whole is licensed under the Creative Commons A ttribution (CC BY) license, consistent with the licensing of the original REAL-COLON dataset [ 3 ]. References 1. Ali, S., Jha, D., Ghatw ary , N., Realdon, S., Cannizzaro, R., Salem, O.E., Lamarâo, D.V., Da Roza, C., Riegler, M.A., Halvorsen, P .: A multi-cen tre polyp detection and segmen tation dataset for generalisabilit y assessment. Scientific Data 10 (1), 75 (2023) 2 2. Bai, S., Cai, Y., Chen, R., Chen, K., Chen, X., Cheng, Z., Deng, L., Ding, W., Gao, C., Ge, C., et al.: Qwen3-vl technical rep ort. arXiv preprint (2025) 3 , 7 , 8 3. Biffi, C., Antonelli, G., Bernhofer, S., Hassan, C., Hirata, D., Iwatate, M., Maieron, A., Salv agnini, P ., Cherubini, A.: Real-colon: A dataset for dev eloping real-w orld ai applications in colonoscopy . Scien tific Data 11 (1), 539 (2024). https://doi.org/10. 1038/s41597- 024- 03359- 0 2 , 4 , 10 , 13 , 16 4. Bose, K., Kumar, A., Soundarara jan, R., Mudgil, P ., Ralmilay , S., Dutta, N., Singhal, M., Kumar, A., Sen, S., Patra, A., et al.: Multi-rads syn thetic radiology rep ort dataset and head-to-head b enchmarking of 41 op en-weigh t and proprietary language mo dels. arXiv preprint arXiv:2601.03232 (2026) 5 5. Carion, N., Gustafson, L., Hu, Y.T., Debnath, S., Hu, R., Suris, D., Ryali, C., Alw ala, K.V., Khedr, H., Huang, A., et al.: Sam 3: Segment anything with concepts. arXiv preprin t arXiv:2511.16719 (2025) 6 , 7 6. Cham b on, P ., Delbrouc k, J.B., Sounack, T., Huang, S. C., Chen, Z., V arma, M., T ruong, S.Q., Chuong, C.T., Langlotz, C.P .: Chexp ert plus: Augmenting a large c hest x-ray dataset with text radiology reports, patient demographics and additional image formats. arXiv preprint arXiv:2405.19538 (2024) 3 7. Choudh uri, A., Gao, Z., Zheng, M., Planche, B., Chen, T., W u, Z.: P olypseg- trac k: Unified foundation model for colonoscopy video analysis. In: Medical Image Computing and Computer Assisted Interv ention – MICCAI 2025. Springer (2025) 2 8. Deitk e, M., Clark, C., et al.: Molmo and pixmo: Op en weigh ts and op en data for state-of-the-art vision-language mo dels. In: Pro ceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) (2025) 3 , 7 , 8 Title S uppressed Due to Excessive Length 11 9. F an, Y., Ma, X., W u, R., Du, Y., Li, J., Gao, Z., Li, Q.: Videoagen t: A memory- augmen ted multimodal agent for video understanding. In: European Conference on Computer Vision. pp. 75–92. Springer (2024) 3 10. Ge, Y., Zhao, Y., Zhu, Z., et al.: Planting a seed of vision in large language mo del. arXiv preprin t arXiv:2307.08041 (2023) 3 , 7 , 8 11. Gemini T eam: Gemini 1.5: Unlo cking multimodal understanding across millions of tok ens of con text. arXiv preprin t arXiv:2403.05530 (2024) 2 , 3 , 7 , 8 12. GLM T eam: Glm-4: Open multilingual multimodal c hat lms (2024), https://gith ub. com/THUDM/GLM- 4 3 , 7 , 8 13. He, Y., Zhu, Y., F u, P ., Y ang, R., Chen, T., W ang, Z., Li, Q., Zhou, P ., Y ang, X., W ang, S.: Endo-clip: Progressive self-supervised pre-training on raw colonoscopy records. In: Medical Image Computing and Computer Assisted Interv ention – MICCAI 2025. Springer (2025), early accepted. A v ailable at: 2505.09435 2 , 6 , 7 14. Jha, D., Smedsrud, P .H., Riegler, M.A., Halvorsen, P ., de Lange, T., Johansen, D., Johansen, H.D.: K v asir-seg: A segmen ted p olyp dataset. In: MultiMedia Modeling. Lecture Notes in Computer Science, v ol. 11962, pp. 451–462. Springer, Cham (2020). h ttps://doi.org/10.1007/978- 3- 030- 37734- 2_37 2 15. Jong, M.R., Bo ers, T.G.W., Kusters, K.H.J., Jasp ers, T.J.M., v an der Putten, J.A., de With, P .H.N., v an der Sommen, F., de Gro of, A.J., Bergman, J.J.G.H.M., et al.: Gastronet-5m: A m ulticen ter dataset for developing foundation models in gastrointestinal endoscopy . Gastro en terology 170 (1), 174–187 (2026). https: //doi.org/10.1053/j.gastro.2025.07.030 2 16. Li, K., F athan, M.I., Patel, K., Zhang, T., Zhong, C., Bansal, A., Rastogi, A., W ang, Z., W ang, G.: Colonoscopy polyp detection and classification: Dataset creation and comparativ e ev aluations. PLoS ONE 16 (8), e0255809 (2021) 2 17. Li, S., Ren, Y., Y u, Y., Li, H.: A survey of deep learning algorithms for colorectal p olyp segmentation. Neuro computing (2024) 2 18. Ma, Y., Chen, X., Cheng, K., Li, Y., Sun, B.: Ldp olypvideo benchmark: A large- scale colonoscopy video dataset of div erse p olyps. In: Medical Image Computing and Computer Assisted Interv ention–MICCAI 2021: 24th International Conference. pp. 387–396. Springer (2021) 2 19. Misa wa, M., Kudo, S.e., Mori, Y., Maeda, Y., Kataok a, S., Ogata, N., Ichimasa, K., Kudo, T., Mori, K., Ishigaki, T.: Developmen t of a computer-aided detection system for colonoscopy and a publicly accessible large colonoscopy video database (with video). Gastrointestinal Endoscopy 93 (4), 960–967 (2021) 2 20. Op enAI: Gpt-4 tec hnical rep ort (2023) 2 , 7 , 8 21. Radford, A., Kim, J.W., Hallacy , C., Ramesh, A., Goh, G., Agarw al, S., Sastry , G., Ask ell, A., Mishkin, P ., Clark, J., et al.: Learning transferable visual mo dels from natural language sup ervision. In: ICML. pp. 8748–8763. PMLR (2021) 2 , 6 , 7 22. Song, Y., Zhang, Z., W ang, R., Zhong, L., Cai, C., Chen, J., Zhou, Y., W ang, X., Li, Z., Y ang, L., Li, Z., Y an, H., Zhang, Q., Qian, D., Li, X.: Cas-colon: A comprehensiv e colonoscop y anatomical segmen tation dataset for artificial intelligence developmen t. Scien tific Data 12 (1), 1382 (2025). https://doi.org/10.1038/s41597- 025- 05588- 3 2 23. Sung, H., F erla y , J., Siegel, R.L., Lav ersanne, M., So erjomataram, I., Jemal, A., Bray , F.: Global cancer statistics 2020: Glob ocan estimates of incidence and mortalit y w orldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians 71 (3), 209–249 (2021). https://doi.org/10.3322/caac.21660 1 24. Tian, Q., Liao, H., Huang, X., Y ang, B., Lei, D., Ourselin, S., Liu, H.: Endomamba: An efficient foundation mo del for endoscopic videos via hierarc hical pre-training. In: 12 H. Abdullah et al. Medical Image Computing and Computer Assisted Interv ention – MICCAI 2025. Springer (2025) 2 25. W ang, Y., He, Y., Li, Y., Li, K., Y u, J., Ma, X., Li, X., Chen, G., Chen, X., W ang, Y., Luo, P ., Liu, Z., W ang, Y., W ang, L., Qiao, Y.: Intern vid: A large-scale video-text dataset for m ultimo dal understanding and generation. In: The T welfth In ternational Conference on Learning Represen tations (ICLR). Op enReview.net (2024), https://openreview.net/forum?id=MLBdi W u4F w 3 , 6 , 7 26. W ang, Z., Liu, C., Dou, Q., Zhang, S., et al.: Improving foundation model for endoscop y video analysis via representation learning on long sequences. IEEE Journal of Biomedical and Health Informatics pp. 3526–3536 (2025). https://doi. org/10.1109/JBHI.2025.3349925 2 27. Zeng, Q., Xie, Y., Lu, Z., Lu, M., W u, Y., Xia, Y.: Segment together: A v ersatile paradigm for semi-sup ervised medical image segmentation. IEEE T ransactions on Medical Imaging 44 (7), 2948–2959 (2025). https://doi.org/10.1109/TMI.2025. 3360447 2 28. Zhou, C., Zhu, C., Xiong, Y., Suri, S., Xiao, F., W u, L., Krishnamo orthi, R., Dai, B., Loy , C.C., Chandra, V., et al.: Edgetam: On-device track anything mo del. In: Pro ceedings of the Computer Vision and Pattern Recognition Conference. pp. 13832–13842 (2025) 4 , 6 , 7 , 8 , 22 Title S uppressed Due to Excessive Length 13 A A dditional Dataset Details A.1 Annotation Pip eline The b enchmark w as constructed through a multi-stage filtering pip eline applied to 60 video sequences from the REAL-colon dataset [ 3 ]. A vision-language mo del iden tified 1,325 candidate lesion windows spanning 22.97 hours of video. Successiv e verification agent filtering, AI confirmation agen t with cued video from the tracking, and human review retained 528 windo ws (39.8%) cov ering 464,035 frames (12.89 hours) across 59 sequences (T able 5 ). The largest reduction o ccurred at verification filtering (422 windows remo ved), follo wed by AI confirmation (306 windo ws), with human review removing only 69 windows (11.6%), indicating strong agreement with automated filtering. Curated windo ws av erage 878.9 frames compared to 414-509 for rejected ones, and carry richer annotations av eraging 252.4 w ords p er windo w v ersus 145-190 for rejected windows (T able 6 ). The curated set comprises 314,408 b ounding b o xes, 227,343 segmentation masks, and 145,515 words of clinical text (T able 4 ). Windo w duration distributions across pip eline stages are shown in Fig. 8 . A.2 V alidating the Annotations The agentic pip eline pro duced 597 lesion videos, each reviewed b y a clinician. Of these, 528 w ere accepted (88.4% success rate), totalling 464,035 frames. W e ablate each pip eline stage on the full REAL-colon dataset b ox annotations for p olyps (T able 7 , Fig. 7 ). V erification filtering, in whic h a a video agen t reviews the candidate window and confirms whether a lesion is present, yields the largest single-step precision gain (+7.3%) with minimal recall loss. T racking restricts p ositiv e predictions to frames with actual b ounding b oxes, sharply increasing precision (+10.2%) at the cost of recall ( − 17.2%). AI confirmation, where a second video agent re-examines the track ed window with b ounding-b o x o verla ys to v alidate the lo calised lesion, remov es 34% of remaining windo ws and improv es precision b y +6.7% and F1 by +2.0pp. Human review provides a further marginal but consistent improv emen t (+2.4% precision, +1.0% F1), while spatial metrics remain nearly unchanged, indicating that the rejected windows had minimal impact on localisation quality . A.3 Colon-Benc h Suite of Ev aluation Benchmarks Colon-Benc h is a multi-task video b enc hmark for colonoscopy understanding spanning five tasks: binary lesion classification, lesion detection, instance seg- men tation, and visual question answering at t wo difficult y levels (prompted and unprompted). The b enchmark comprises 1,597 unique video clips from 60 patien t sequences totalling 955,126 frames (T able 9 ). Classification co vers 790 clips (518 lesion-free, 272 lesion-p ositive), while detection and segmentation use the 272 and 264 lesion-p ositive clips resp ectiv ely , providing 61,538 p er-frame b ounding b oxes and 57,550 p er-frame masks. The prompted VQA split contains 1,485 fiv e-choice 14 H. Abdullah et al. T able 4. Curated Colon-Benc h Dataset Statistics. Windows that passed all filtering stages (Confirmed, n = 528 ) are compared against those rejected during h uman review (Rejected, n =69 ). Metric Confirmed (528) Rejected (69) T otal (597) Sequences cov ered 59 37 60 T otal frames 464,035 28,571 492,606 A vg. frames / window 878.9 414.1 - Duration (hours, 10 fps) 12.89 0.79 13.68 T otal b ounding b o xes 300,132 14,276 314,408 A vg. bb o xes / windo w 568.4 206.9 - Bb o x co verage (%) 64.7 50.0 - T otal segmen tation masks 213,067 14,276 227,343 Windo ws with masks 465 / 528 69 / 69 534 / 597 Mask cov erage (%) 45.9 50.0 - A vg. words - initial desc. 162.8 88.6 - A vg. words - v erified desc. 55.5 53.0 - A vg. words - confirmation note 34.2 35.7 - A vg. words - all text combined 252.4 177.2 - T otal w ords (all text) 133,289 12,226 145,515 questions ov er 499 clips, while the unprompted split con tains 2,740 questions ov er 918 clips and fully encompasses all detection and segmentation videos, enabling direct comparison of spatial lo calisation against op en-ended clinical reasoning (see T able 8 for examples). A.4 Benc hmark F ormation and Blind T ests F or each clip, three five-w a y multiple-c hoice questions are generated using Gemini 3 Flash, co vering lesion identification, clinical c haracteristics, and temp oral reasoning. The pr ompte d set uses videos with b ounding-box and mask ov erlays on confirmed lesion windows, while the unpr ompte d set uses raw frames and additionally includes non-lesion windows. Both sets undergo structural v alidation and are randomis ed with a fixed seed for reproducibility . T o mitigate text-only exploitabilit y , we apply tw o-stage debiasing: (i) for ev ery question, four adv ersarial distractors are regenerated in a separate LLM call receiving only the question stem and correct answer, pro ducing length- and style-matc hed alternatives with re-randomised option p ositions; (ii) a blind text-only stress-test identifies cases where debiasing in tro duced new textual shortcuts, and these questions revert to their original formulation. After this pro cedure, blind-only accuracy is 44.6% (prompted) and 37.1% (unprompted) on Gemini-3 Flash versus the 20% random baseline, with residual margins attributable to skew ed lesion-type distributions rather than surface-lev el cues (see Fig. 2 ). Title S uppressed Due to Excessive Length 15 T able 5. Colon-Benc h Filtering F unnel Overview. Each row shows the n umber of windows, total frames, and video duration at successive pip eline stages. Dropp ed ro ws indicate windo ws remov ed by the corresp onding filter. Pip eline Stage Windo ws F rames Hours Retention (%) VLM Detection Prop osals + Merging 1,325 826,763 22.97 100.0 − Rejected by V erification Agent − 422 − 178,323 − 4.95 - + V erification Agent Filtering 903 648,440 18.01 68.2 − rejected by AI confirmation agent − 306 − 155,834 − 4.33 - + Cu ed AI Confirmation Agent 597 492,606 13.68 45.1 − Hu man Review Rejected − 69 − 28,571 − 0.79 - Final Curated Set 528 464,035 12.89 39.8 T able 6. Colon-Benc h Annotations Stage-Wise Rejection Characteristics. Statistics include windo w duration (in frames at 10 fps) and text description lengths (in w ords). At earlier stages, only a subset of text fields is av ailable. V erification AI Confirmation Human Review Final Curated Rejected (422) Rejected (306) Rejected (69) Confirmed (528) Sequences 60 56 37 59 T otal frames 178,323 155,834 28,571 464,035 A vg. frames/window 422.6 509.3 414.1 878.9 Median frames/window 339.5 417.0 259.0 599.0 Duration (hours) 4.95 4.33 0.79 12.89 T ext descriptions (avg. wor ds p er window) Initial desc. 84.9 101.3 88.6 162.8 V erified desc. 60.4 53.8 53.0 55.5 Confirmation note - 34.4 35.7 34.2 All text combined 145.3 189.5 177.2 252.4 T otal words 61,326 57,976 12,226 133,289 B A dditional Results P er-task results are rep orted in T ables 10 (V QA), 12 (classification), 13 (detec- tion), and 11 (segmen tation), with corresponding visualisations in Figs. 11 , 12 (V QA), Fig. 14 and 15 (classification), and Fig. 13 (detection). Qualitativ e detection and segmen tation examples are shown in Figs. 9 and 10 . 16 H. Abdullah et al. T able 7. Colon-Benc h Pip eline Stage Ablation on REAL-colon [ 3 ]. T emp oral metrics are frame-level detection rates. Spatial metrics (av ailable only after tracking) rep ort b ounding-box F1 at IoU thresholds 0.25, 0.50, 0.75 and the mean IoU of matc hed b o xes. T emp oral (F rame-level) Spatial (Bounding Box) Pipeline Stage Prec. Rec. F1 Spec. F1@.25 F1@.50 F1@.75 mIoU VLM Detection Agent 30.9 67.5 42.4 78.6 - - - - + Windo w Merging 28.9 69.9 40.9 75.6 - - - - + V erification Agent Filtering 36.2 68.7 47.4 82.9 - - - - + Spatial Lo calization + T racking 46.4 51.5 48.8 91.6 34.2 27.7 15.1 34.4 + AI Confirmation Agent 53.1 48.8 50.8 93.9 34.2 27.8 15.3 35.2 + Human Review 55.4 48.6 51.8 94.4 34.2 27.8 15.3 35.2 T able 8. Colon-Benc h Example VQA Items. Correct answ ers are b olded . Unprompted (r aw vide o; same clip as Fig. 9 / Fig. 10 ) Q: How is the morpholo gy and anatomic al lo c ation of the identifie d lesion describ e d? (A) Pedunculated p olyp on a rectal fold (B) Depressed lesion on the cecal base (C) Semi-p edunculated polyp in the rectum (D) Sessile p olyp on a haustral fold (E) Flat lesion on the splenic flexure Prompted (vide o with bb ox / mask overlays) Q: At the 1.0 s mark, which clinic al finding is first cle arly visible? (A) Flat mucosal lesion (Paris IIb) (B) Sessile p olyp (Paris Is) (C) Sub epithelial bulge (D) No abnormality detected (E) Pedunculated p olyp VLM Detection Agent + W indow Mer ging + V erification Agent F iltering + Spatial L ocalization + T racking + AI Confir mation Agent + Human R eview 0 10 20 30 40 50 60 70 80 Scor e (%) 30.9 28.9 36.2 46.4 53.1 55.4 67.5 69.9 68.7 51.5 48.8 48.6 42.4 40.9 47.4 48.8 50.8 51.8 Precision / Recall Pipeline Stage Ablation T emporal Detection Metrics P r ecision R ecall F1 Fig. 7. P olyp-Based Colon-Bench Pip eline Ablation. T emp oral frame-lev el Preci- sion, Recall, and F1 are rep orted at each pro cessing stage, from ra w VLM detection through human review. Annotated v alues are p ercen tages from the p olyp annotations of REAL-COLON[ 3 ]. Title S uppressed Due to Excessive Length 17 0 60 120 180 240 300 360 420 480 540 600 660 720 W indow duration (seconds) 0 50 100 150 200 Number of windows Window Length Distribution Across Pipeline Stages VLM Detection + Mer ging (n=1325) + V erification Agent F iltering (n=903) + AI Confir mation Agent (n=597) + Human R eview (n=528) Fig. 8. Window Duration Distributions Across Stages. Ov erlaid distributions of lesion window durations (seconds at 10 FPS) are sho wn at successive pip eline filtering stages: VLM-detected, after verification, after AI confirmation with visual b o x ov erla ys, and the final human-curated set. Eac h stage is a strict subset of the previous one. T able 9. Colon-Bench T ask Statistics. F rame counts are deriv ed from the name mapping using per-clip start/end frame indices. Bounding b oxes and masks are p er- frame annotations. VQA questions are 5-c hoice MCQ with approximately 3 questions p er video. Statistic Classification Detection Segmentation V QA Prompted VQA Unprompted T otal clips / questions 790 272 264 1,485 2,740 Unique videos 790 272 264 499 918 T otal frames 241,996 92,311 87,631 367,572 538,115 A vg. clip length (frames) 306 339 332 737 586 T otal b ounding boxes - 61,538 - - - T otal mask annotations - - 57,550 - - MCQ choices p er question - - - 5 5 A vg. questions p er video - - - 3.0 3.0 Unique patient sequences 60 57 57 59 60 L esion / No-lesion split 272 / 518 - - - - 18 H. Abdullah et al. T able 10. Colon-Bench VQA Benchmark Accuracy . Accuracy (%) is rep orted on Colon-Benc h Prompted (1485 questions) and Unprompted (2740 questions), computed as correct answers divided by total questions; unansw ered questions coun t as incorrect. Best results p er b enchmark are sho wn in bold . Mo del Prompted Accuracy (%) Unprompted Accuracy (%) Qw en3-VL 8B 32.9 38.3 Seed 1.6 Flash 38.1 45.4 Qw en-VL Max 39.1 45.4 Qw en3-VL 32B 39.3 44.4 Qw en3-VL Plus 41.2 50.5 Qw en3.5 Plus 44.3 60.5 Molmo 2-8B 46.1 53.4 Qw en3-VL 235B 46.7 56.6 Gemini 2.5 Flash Lite 47.8 46.8 Qw en3.5 397B 49.0 60.1 GLM-4.6V 55.7 53.8 Seed 1.6 62.9 72.0 Gemini 3.1 Flash Lite 69.2 67.7 Gemini 3 Flash 76.6 76.0 Gemini 3 Pro 78.6 82.5 Title S uppressed Due to Excessive Length 19 T able 11. Colon-Benc h Segmentation Benc hmark Results. Video ob ject seg- men tation results (%) are rep orted on the Colon-Benc h segmentation b enc hmark (272 videos). Eac h mo del first detects the target lesion via b ounding-b o x prompting, then a SAM-based track er propagates the mask across the tracking window. Mean IoU and Mean Dice are computed o ver all ev aluated frames. Best results p er metric are shown in bold . Mo del mIoU mDice GPT-4o 0.5 0.8 SAM 3 2.5 2.9 Seed 1.6 Flash 2.6 3.5 Molmo 2-8B 2.6 3.8 Qw en3-VL 8B 10.4 13.1 GLM-4.6V 12.5 16.1 Seed 1.6 12.6 16.1 Qw en3-VL 32B 12.7 15.9 Qw en3-VL 235B 13.6 16.9 Claude Opus 4.6 16.1 20.5 Qw en3.5 397B 16.6 21.0 Qw en3.5 Plus 16.7 21.0 Gemini 2.5 Flash Lite 19.9 24.3 Qw en3-VL Plus 20.4 24.6 Qw en-VL Max 25.6 29.6 GPT-5.2 30.7 36.5 GPT-5.4 34.5 41.1 Gemini 3.1 Flash Lite 37.4 43.4 Gemini 3 Pro 45.0 51.3 Gemini 3 Flash 48.3 54.7 20 H. Abdullah et al. T able 12. Colon-Bench Classification Benc hmark Results. Binary lesion clas- sification results (%) are rep orted on the Colon-Bench classification b enc hmark (790 records). Each mo del receives a video clip and predicts whether a lesion is present (p ositiv e) or absent (negative). Accuracy is computed ov er all records; unev aluated records count as incorrect. Precision, Recall, and F1 are rep orted for the p ositive (lesion-presen t) class. Best results p er metric are sho wn in bold . Mo del A cc. Prec. Rec. F1 Qw en3-VL 8B 34.4 34.4 100.0 51.2 CLIP 34.7 34.5 100.0 51.3 ViCLIP 35.8 34.8 98.5 51.4 Gemini 2.5 Flash Lite 52.3 42.3 95.9 58.7 Endo-CLIP 52.9 41.9 95.2 58.2 Qw en3.5 Plus 59.1 36.2 24.6 29.3 GLM-4.6V 60.6 46.5 94.1 62.2 Colon- Vi CLIP 64.4 49.0 84.2 62.0 Qw en3.5 397B 64.6 10.0 0.4 0.7 Qw en3-VL 235B 64.9 22.2 0.7 1.4 Qw en-VL Max 65.6 0.0 0.0 0.0 Qw en3-VL 32B 65.6 0.0 0.0 0.0 Qw en3-VL Plus 65.6 0.0 0.0 0.0 Molmo 2-8B 67.3 52.9 46.7 49.6 Gemini 3 Flash 72.0 55.3 97.1 70.5 Seed 1.6 Flash 72.9 94.2 24.3 38.6 Gemini 3 Pro 81.1 66.1 93.0 77.3 Seed 1.6 82.0 85.0 58.7 69.4 Gemini 3.1 Flash Lite 85.1 72.6 90.8 80.7 Title S uppressed Due to Excessive Length 21 T able 13. Colon-Benc h Detection Benchmark Results. Ob ject detection results (%) are rep orted on the Colon-Bench detection b enchmark (272 videos). Precision, Recall, and F1 are computed at IoU ≥ 0.50. AP@50 is the av erage precision at IoU ≥ 0.50; mAP@50-95 a verages AP across IoU thresholds 0.50-0.95. Mean IoU is the a verage in tersection-ov er-union of matc hed predicted and ground-truth b o xes. Best results p er metric are shown in b old . Mo del Prec. Rec. F1 AP@50 mAP@50-95 mIoU Molmo 2-8B 0.2 0.1 0.1 0.0 0.0 53.2 GPT-4o 0.6 0.1 0.2 0.0 0.0 55.4 Seed 1.6 Flash 1.6 0.5 0.8 0.0 0.0 55.7 Claude Opus 4.6 3.5 3.7 3.6 1.1 0.1 59.5 Gemini 2.5 Flash Lite 2.5 9.5 3.9 1.7 0.4 61.8 Qw en3-VL 8B 6.6 3.8 4.8 0.7 0.2 62.6 GLM-4.6V 6.6 6.2 6.4 0.5 0.1 61.6 Seed 1.6 10.4 6.6 8.0 1.1 0.2 59.7 Qw en3.5 397B 9.4 8.9 9.2 1.7 0.4 62.9 Qw en3-VL 32B 14.4 7.6 9.9 1.5 0.4 62.1 Qw en3.5 Plus 10.7 10.3 10.5 2.1 0.4 61.5 Qw en3-VL 235B 15.7 8.2 10.8 2.0 0.5 62.9 Qw en3-VL Plus 10.0 12.7 11.2 2.9 0.7 64.0 GPT-5.2 26.1 24.2 25.1 12.2 3.3 66.1 Qw en-VL Max 39.0 22.2 28.3 10.6 3.8 71.4 GPT-5.4 33.2 30.6 31.9 17.9 5.9 68.4 Gemini 3.1 Flash Lite 36.8 32.5 34.5 18.1 6.0 69.9 Gemini 3 Pro 42.2 42.5 42.3 23.7 11.1 76.3 Gemini 3 Flash 50.9 52.7 51.8 32.6 17.0 79.8 22 H. Abdullah et al. Input GT GPT-5.2 Opus 4.6 Gemini 3 Flash Qw en 3.5-Plus Qw en VL-Max Qw en 3 VL-235B Fig. 9. Colon-Bench Qualitativ e Detection Comparison. Each row shows a differen t endoscopy video frame: (1) mass , (2) sessile p olyp , (3) p e dunculate d p olyp , (4) ulc er , (5) p e dunculate d p olyp , (6) sessile p olyp . Columns sho w the raw input, ground- truth bounding b o x, and predicted b ounding b oxes from eac h mo del. Bounding b o xes are dra wn in red. These detections are used in the segmentation b enchmark using 3 b o x detections as prompts for the EdgeT AM track er [ 28 ]. Title S uppressed Due to Excessive Length 23 Input GT GPT- 5.2 Opus 4.6 Gemini 3 Flash Qw en3.5 - Plus Qw en VL - Max Qw en3- VL- 235B SAM 3 Fig. 10. Colon-Bench Qualitative Segmentation Comparison. Eac h ro w shows a different endoscopy video frame: (1) erythematous r e gion , (2) lip oma , (3) sessile p olyp , (4) er osion , (5) ulc er , (6) erythematous re gion . Columns show the ra w input, ground-truth mask ov erla y , and predicted mask o verla ys from each mo del (including SAM 3). Masks are rendered as semi-transparen t purple o verla ys. Gemini 3 P r o Gemini 3 Flash Gemini 3.1 Flash Lite Seed 1.6 GLM-4.6V Qwen3.5 397B Gemini 2.5 Flash Lite Qwen3- VL 235B Molmo 2-8B Qwen3.5 Plus Qwen3- VL Plus Qwen3- VL 32B Qwen- VL Max Seed 1.6 Flash Qwen3- VL 8B 0 10 20 30 40 50 60 70 80 90 A ccuracy (%) 78.6% 76.6% 69.2% 62.9% 55.7% 49.0% 47.8% 46.7% 46.1% 44.3% 41.2% 39.3% 39.1% 38.1% 32.9% Colon-Bench Prompted VQA R andom chance (20%) Fig. 11. Colon-Bench Prompted V QA Accuracy . V QA accuracy (%) on the Colon- Benc h prompted b enc hmark (1,485 questions) is computed as correct answers divided b y the total num ber of questions so that unanswered questions count as incorrect. The dashed red line indicates the 20% random-chance baseline. Mo dels are ordered b y accuracy (highest first), and colours denote mo del family . 24 H. Abdullah et al. Gemini 3 P r o Gemini 3 Flash Seed 1.6 Gemini 3.1 Flash Lite Qwen3.5 Plus Qwen3.5 397B Qwen3- VL 235B GLM-4.6V Molmo 2-8B Qwen3- VL Plus Gemini 2.5 Flash Lite Qwen- VL Max Seed 1.6 Flash Qwen3- VL 32B Qwen3- VL 8B 0 10 20 30 40 50 60 70 80 90 A ccuracy (%) 82.5% 76.0% 72.0% 67.7% 60.5% 60.1% 56.6% 53.8% 53.4% 50.5% 46.8% 45.4% 45.4% 44.4% 38.3% Colon-Bench Unprompted VQA R andom chance (20%) Fig. 12. Colon-Benc h Unprompted VQA Accuracy . VQA accuracy (%) on the Colon-Benc h unprompted benchmark (2,740 questions) follo ws the same proto col as the prompted b enc hmark: all unanswered questions are treated as incorrect. The dashed red line indicates the 20% random-chance baseline. Mo dels are ordered by accuracy (highest fi rst), and colours denote mo del family . Gemini 3 Flash Gemini 3 P r o Gemini 3.1 Flash Lite GPT -5.4 Qwen- VL Max GPT -5.2 Qwen3- VL Plus Qwen3- VL 235B Qwen3.5 Plus Qwen3- VL 32B Qwen3.5 397B Seed 1.6 GLM-4.6V Qwen3- VL 8B Gemini 2.5 Flash Lite Claude Opus 4.6 Seed 1.6 Flash GPT -4o Molmo 2-8B 0 10 20 30 40 50 60 Scor e (%) 51.8 42.3 34.5 31.9 28.3 25.1 11.2 10.8 10.5 9.9 9.2 8.0 6.4 4.8 3.9 3.6 0.8 32.6 23.7 18.1 17.9 10.6 12.2 2.9 2.0 2.1 1.5 1.7 1.1 0.7 1.7 1.1 17.0 11.1 6.0 5.9 3.8 3.3 0.7 Colon-Bench Detection Benchmark F1 Scor e AP@50 mAP@50-95 Fig. 13. Colon-Benc h Detection Metric Breakdo wn. Ob ject detection results on the Colon-Bench (272 videos) ev aluate eac h mo del’s ability to lo calise lesions via b ounding-box prediction. These detections are used in the main segmentation b enchmark. Three metrics are rep orted for eac h mo del: F1 Score (primary sort key), AP@50 (a verage precision at IoU ≥ 0 . 50 ), and mAP@50-95 (mean av erage precision a veraged o ver IoU thresholds 0.50-0.95). Mo dels are ordered by F1 (highest first). Gemini 3.1 Flash Lite Seed 1.6 Gemini 3 P r o Seed 1.6 Flash Gemini 3 Flash Molmo 2-8B Qwen- VL Max Qwen3- VL 32B Qwen3- VL Plus Qwen3- VL 235B Qwen3.5 397B Colon- V iCLIP GLM-4.6V Qwen3.5 Plus Endo -CLIP Gemini 2.5 Flash Lite V iCLIP CLIP Qwen3- VL 8B 0 10 20 30 40 50 60 70 80 90 A ccuracy (%) 85.1% 82.0% 81.1% 72.9% 72.0% 67.3% 65.6% 65.6% 65.6% 64.9% 64.6% 64.4% 60.6% 59.1% 52.9% 52.3% 35.8% 34.7% 34.4% Colon-Bench Lesion Classification Accuracy R andom chance (50%) Fig. 14. Colon-Bench Classification Accuracy Summary . Binary lesion classi- fication accuracy (%) on the Colon-Bench classification b enc hmark (790 records) is computed after each model predicts whether a lesion is present (p ositiv e) or absent (negativ e). Accuracy is computed ov er all records; unev aluated records coun t as incorrect. The dashed red line indicates the 50% random-chance baseline. Mo dels are ordered b y accuracy (highest first), and colours denote mo del family . Title S uppressed Due to Excessive Length 25 Gemini 3.1 Flash Lite Gemini 3 P r o Gemini 3 Flash Seed 1.6 GLM-4.6V Colon- V iCLIP Gemini 2.5 Flash Lite Endo -CLIP V iCLIP CLIP Qwen3- VL 8B Molmo 2-8B Seed 1.6 Flash Qwen3.5 Plus Qwen3- VL 235B Qwen3.5 397B 0 10 20 30 40 50 60 70 80 90 100 Scor e (%) 72.6 66.1 55.3 85.0 46.5 49.0 42.3 41.9 34.8 34.5 34.4 52.9 94.2 36.2 22.2 10.0 90.8 93.0 97.1 58.7 94.1 84.2 95.9 95.2 98.5 100.0 100.0 46.7 24.3 24.6 0.7 80.7 77.3 70.5 69.4 62.2 62.0 58.7 58.2 51.4 51.3 51.2 49.6 38.6 29.3 1.4 0.7 Colon-Bench Lesion Classification Precision / Recall / F1 P r ecision R ecall F1 Scor e Fig. 15. Colon-Benc h Classification Precision,Recall,F1. Binary lesion classi- fication Precision , Recall , and F1 Score (%) are rep orted on the Colon-Bench classification benchmark (790 records) for the p ositive (p olyp-presen t) class. Mo dels are ordered by F1 (highest first). 26 H. Abdullah et al. C A dditional Analysis C.1 Ablation Study W e study four ablations with tabulated results in T abs. 14 - 16 . The detection frame count ablation shows steady gains in downstream segmentation qualit y , with Gemini 3 Flash improving mIoU/mDice as frames rise (T ab. 14 ). The temp oral con text ablation for binary lesion classification compares video vs. single-frame inputs and finds temp oral context is generally helpful, esp ecially for Seed 1.6, while Gemini 3 Flash is slightly stronger on frames (T ab. 17 ). The b ounding-box annotation ablation for V QA shows only marginal changes, indicating limited dep endence on explicit b o xes (T ab. 15 ). The temp oral context ablation for prompted VQA shows small, mo del-dep enden t shifts, with most mo dels benefiting from video but a few showing minor gains on frames (T ab. 16 ). T able 14. Colon-Benc h Ablation: Detection F rames for Segmentation. Each column pair rep orts Mean IoU and Mean Dice for the given num ber of detection frames. Best results p er frame count are sho wn in b old . Mo del 1 F rame 2 F rames 3 F rames 5 F rames 7 F rames mIoU mDice mIoU mDice mIoU mDice mIoU mDice mIoU mDice Seed 1.6 10.1 12.4 9.3 11.8 13.0 16.6 13.9 17.9 14.7 19.2 Qw en-VL Max 19.7 22.6 20.2 23.4 25.2 29.3 31.3 36.3 33.7 39.4 Gemini 3 Flash 43.1 48.8 46.0 52.2 49.0 55.2 53.6 60.3 54.4 61.5 T able 15. Colon-Benc h Ablation: Bounding-Box Effect on VQA. “With bbox” uses annotated videos containing polyp bounding boxes, while “Without bb ox” uses frame-generated videos of the same lesion windows without annotations (499 matched clip windo ws). ∆ = Without − With. Best results per condition are shown in b old . Mo del With bb o x W/o bbox ∆ Qw en-VL Max 39.1 37.2 -1.8 Gemini 2.5 Flash Lite 48.5 47.8 -0.7 Qw en3.5 397B 49.0 48.1 -0.9 Gemini 3 Flash 76.6 77.2 +0.6 C.2 Colon-Skill for MLLMs W e inv estigate whether MLLMs b enefit from structured domain knowledge at inference time. W e first collect per-mo del V QA predictions and stratify errors by lesion category (Figs. 17 and 18 ), retaining questions that a ma jority of mo dels Title S uppressed Due to Excessive Length 27 T able 16. Colon-Bench Ablation: T emp oral Context for Prompted VQA. “Video” uses the full video clip and “F rame” uses a single representativ e frame ov er 1485 questions. ∆ = F rame − Video. Best results per condition are shown in b old . Mo del Video F rame ∆ Qw en-VL Max 37.5 34.1 -3.4 Gemini 2.5 Flash Lite 46.7 48.0 +1.3 Qw en 3.5 397b 49.6 42.6 -7.0 Gemini 3 Flash 76.8 74.3 -2.5 T able 17. Colon-Bench Ablation: T emp oral Context for Classification. “Video” uses the full video clip and “F rame” uses a single representativ e frame. ∆ = F rame − Video. Best results p er condition are sho wn in b old . A ccuracy (%) F1 Score (%) Mo del Video F rame ∆ Video F rame ∆ Qw en VL Max 65.6 49.6 -15.9 0.0 0.0 0.0 Gemini 3 Flash 69.1 70.8 +1.6 68.6 69.5 +0.9 Seed 1.6 82.4 60.0 -22.4 69.9 60.8 -9.1 answ er incorrectly . A frontier LLM then synthesises these shared failure cases into a concise natural-language Colon-Skil l comprising morphological cues, common confusion traps, and a decision chec klist. This skill context is prep ended to ev ery V QA prompt at inference time. A cross nine MLLMs, skill-augmented prompting yields gains of up to +9.7% (prompted) and +9.0% (unprompted), with the effect most pronounced for higher- capacit y mo dels (T able 18 , Figs. 19 and 20 ). Smaller mo dels suc h as Molmo-2-8B sho w marginal or slightly negativ e deltas, suggesting sufficient mo del capacity is needed to in tegrate the additional context. 28 H. Abdullah et al. T able 18. Colon-Skill V QA Gains. Effect of skill-context prompting on VQA accuracy (%) across Unprompted (2740 questions) and Prompted (1485 questions) splits. Baseline uses the default prompt; With Skill app ends a distilled skill context. ∆ is th e absolute c hange in percentage p oints. Best results p er split are shown in bold . Unprompted Prompted Mo del Baseline w/ Skill ∆ Baseline w/ Skill ∆ Gemini 3 Flash 76.0 78.9 +3.0 76.6 78.5 +1.8 Qw en 3.5 Plus 60.5 58.6 -1.9 44.3 45.0 +0.7 Qw en 3.5 397B 60.1 63.8 +3.8 49.0 58.7 +9.7 Qw en 3 VL 235B 56.6 60.5 +3.9 46.7 50.1 +3.4 Molmo 2 8B 53.4 51.2 -2.2 46.1 44.4 -1.8 Qw en 3 VL Plus 50.5 56.6 +6.1 41.2 46.1 +4.9 Qw en VL Max 45.4 53.1 +7.7 39.1 40.8 +1.8 Qw en 3 VL 32B 44.4 49.6 +5.2 39.3 43.5 +4.2 Qw en 3 VL 8B 38.3 47.3 +9.0 32.9 38.8 +5.9 Title S uppressed Due to Excessive Length 29 Colon-Skill ## Universal Anti-Error Rules * **Do not hallucinate stalks:** The most common morphological error is calling a sessile (broad-based) polyp "pedunculated ." If the lesion sits flatly on a haustral fold without a distinct, narrowed fibrous tether, it is a sessile or flat- elevated lesion. * **Correctly interpret NBI (Narrow-Band Imaging):** Under NBI, healthy mucosa appears greenish/cyan. Adenomas typically appear brownish with regular pit patterns, while hyperplastic or sessile serrated lesions (SSLs) appear pale or whitish. Do not confuse white-light colors with NBI colors. * **Differentiate holes from masses:** Models consistently mistake diverticula (dark, hollow outpouchings) for depressed flat lesions or dark polyps. If a finding is a perfectly circular dark shadow with smooth margins, it is a hole/ pocket, not a lesion. * **Be conservative with size estimates:** Models routinely overestimate diminutive lesions. Without tool reference, subtle , dome-shaped nodules on folds are usually 3-5mm (diminutive). Lesions occupying a large portion of the lumen are >15 mm. * **Accurately identify interventions:** * *Water jet:* Used for irrigation/clearing mucus or blood (not for coagulation). * *Needle catheter:* Injects fluid to create a blue submucosal cushion (lifts flat lesions). * *Cold/Hot Snare:* Wire loop used for Endoscopic Mucosal Resection (EMR) or polypectomy. ## Lesion Morphology Cues by Category * **Sessile Polyps (Paris Is / IIa):** Broad base, typically situated on the crest of a haustral fold. Often pale, isochromatic (matches mucosa color), or slightly yellowish. Surface is smooth or subtly granular. * **Sessile Serrated Lesions (SSL):** Flat, highly subtle, pale/isochromatic lesions. Key visual signatures include a " cloud-like" surface texture, indistinct/blurred borders, and a distinct "mucus cap" (adherent yellow/white debris). * **Angioectasia / Angiodysplasia:** Sits completely flush with the mucosa. Appears as a bright, cherry-red, "fern-like" or stellate pattern of tortuous, dilated submucosal blood vessels. * **Pedunculated Polyps (Paris Ip):** Features a distinct, thick or thin stalk attached to the colonic wall. The head is usually bulbous, multi-lobulated, and noticeably redder (erythematous) than the stalk. * **Ulcers:** Deep or shallow punched-out depressions. Look for white/yellow necrotic slough or fibrin exudate in the center, surrounded by raised, rolled, and erythematous margins. * **Diverticula:** Distinct, dark, circular pockets with smooth, well-defined rims. Often seen in clusters. ## Common Confusion Traps and How to Resolve Them * **Trap: Angioectasia vs. Suction Artifact vs. Dieulafoy.** Models overwhelmingly fail here. *Resolution:* Suction artifacts are tiny, non-specific red petechial spots. Dieulafoy/ulcers involve tissue defects or active arterial spurts. Angioectasia is a flat network of *fern-like*, branching dilated vessels without a mucosal defect. * **Trap: Diverticulum vs. Depressed Lesion (Paris IIc).** *Resolution:* A diverticulum is a true anatomical hole (deep black center, hollow). A Paris IIc lesion is a mucosal depression with a visible base, often showing irregular margins and altered pit patterns. * **Trap: Mischaracterizing pale flat lesions.** Models frequently call pale, mucus-covered flat lesions "ulcers" or " tumors." *Resolution:* If a flat lesion is pale with a "cloud-like" surface and lacks rolled margins or central necrosis, it is an SSL, not an ulcer or malignant tumor. * **Trap: "Lobulated" vs. "Smooth" surface confusion.** *Resolution:* "Smooth" dome-shaped pale nodules are typically diminutive hyperplastic or small sessile polyps. "Lobulated" or "cerebriform" (brain-like) surfaces with redder hues indicate adenomatous polyps. * **Trap: Mistaking anatomical folds for masses.** *Resolution:* Prominent haustral folds curve predictably around the lumen. Do not label a normal fold as a "sessile mass" unless there is a localized change in color, vascularity, or elevation (nodularity). ## Fast VQA Decision Checklist 1. **Assess the geometry:** Is it a mass (protruding), a defect (depressed/ulcerated), a hole (diverticulum), or a flat vascular anomaly? 2. **Evaluate the attachment:** If it’s a polyp, is it draped over/broadly attached to a fold (sessile) or hanging by a tether (pedunculated)? 3. **Check the lighting mode:** Is the image under white light (pink/red hues) or NBI/BLI (green/brown/ cyan hues)? Adjust color descriptions accordingly. 4. **Examine the surface & margins:** Is the surface smooth, cloud-like, granular, or lobulated? Are th e margins sharp, rolled, or indistinct? 5. **Identify active tools or residue:** Are there white/yellow strings (mucus/stool), active red oozing (bleeding), a wire loop (snare), or blue fluid (submucosal injection)? Fig. 16. Colon-Skill Prompt Context. The colon-skill SKILL.md file is used as con text augmentation for MLLMs in the VQA b enc hmarks. The skill is extracted b y analysing error patterns across lesion categories and examples of failure modes. 30 H. Abdullah et al. SSL P edunculated P olyp Mucosal Abn. Ulcer Diverticulum Erythematous Sessile P olyp P arasites T umour / LST Bleeding Others Angioectasia Hemor r hoid Cr ohn's 0.0 0.5 1.0 1.5 2.0 2.5 Nor malized Er r or R ate (model mean = 1.0) Normalized Error Rate by Lesion Category Unprompted VQA Cr oss-model avg Gemini 3 Flash Qwen- VL Max Molmo 2 8B GLM-4.6V Fig. 17. Error P atterns: Unprompted Split. Normalized error rate by lesion category (1.0 = mo del av erage). Mucosal Abn. Hemor r hoid SSL P edunculated P olyp Cr ohn's Bleeding Angioectasia Ulcer Erythematous Sessile P olyp T umour / LST Diverticulum P arasites Others 0.0 0.5 1.0 1.5 2.0 2.5 Nor malized Er r or R ate (model mean = 1.0) Normalized Error Rate by Lesion Category Prompted VQA Cr oss-model avg Gemini 3 Flash Qwen- VL Max Molmo 2 8B GLM-4.6V Fig. 18. Error Patterns: Prompted Split. Normalized error rate b y lesion category (1.0 = mo del av erage). Title S uppressed Due to Excessive Length 31 Gemini 3 Flash Qwen3.5 397B Qwen3- VL 235B Molmo 2-8B Qwen3.5 Plus Qwen3- VL Plus Qwen3- VL 32B Qwen- VL Max Qwen3- VL 8B 0 20 40 60 80 A ccuracy (%) 76.6% 49.0% 46.7% 46.1% 44.3% 41.2% 39.3% 39.1% 32.9% 78.5% 58.7% 50.1% 44.4% 45.0% 46.1% 43.5% 40.8% 38.8% +1.8 pp +9.7 pp +3.4 pp -1.8 pp +0.7 pp +4.9 pp +4.2 pp +1.8 pp +5.9 pp Baseline vs Colon-Skill Prompted VQA R andom chance (20%) Baseline W ith Colon-Skill Fig. 19. Colon-Skill Effect: Prompted V QA. Effect of skill con text on VQA p erformance ( pr ompte d split). Gemini 3 Flash Qwen3.5 Plus Qwen3.5 397B Qwen3- VL 235B Molmo 2-8B Qwen3- VL Plus Qwen- VL Max Qwen3- VL 32B Qwen3- VL 8B 0 20 40 60 80 A ccuracy (%) 76.0% 60.5% 60.1% 56.6% 53.4% 50.5% 45.4% 44.4% 38.3% 78.9% 58.6% 63.8% 60.5% 51.2% 56.6% 53.1% 49.6% 47.3% +3.0 pp -1.9 pp +3.8 pp +3.9 pp -2.2 pp +6.1 pp +7.7 pp +5.2 pp +9.0 pp Baseline vs Colon-Skill Unprompted VQA R andom chance (20%) Baseline W ith Colon-Skill Fig. 20. Colon-Skill Effect: Unprompted V QA. Effect of skill context on VQA p erformance ( unpr ompte d split).
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment