HYDRA: Hybrid Deep Magnetic Resonance Fingerprinting
Purpose: Magnetic resonance fingerprinting (MRF) methods typically rely on dictio-nary matching to map the temporal MRF signals to quantitative tissue parameters. Such approaches suffer from inherent discretization errors, as well as high computation…
Authors: Pingfan Song, Yonina C. Eldar, Gal Mazor
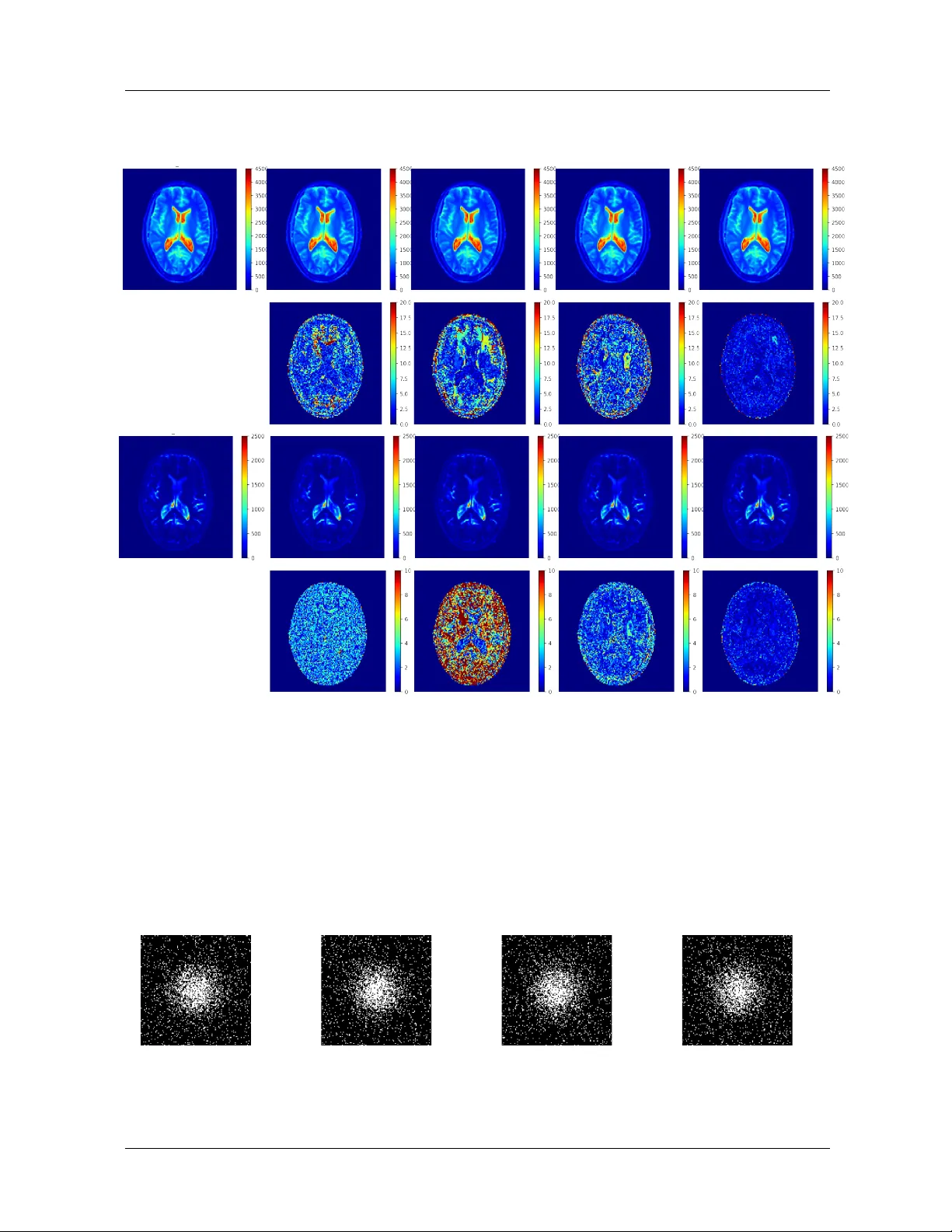
HYDRA: Hyb rid Deep Magnetic Resonance Fingerp rinting Pingfan Song 1 Y onina C. Elda r 2 Gal Mazo r 3 Miguel R. D. Ro drigues 4 Abstract Purp ose: Magnetic resonance fingerprin ting (MRF) metho ds typically rely on dictio- nary matc hing to map the temp oral MRF signals to quantitativ e tissue parameters. Suc h approac hes suffer from inheren t discretization errors, as well as high computa- tional complexity as the dictionary size grows. T o alleviate these issues, we prop ose a HYbrid Deep magnetic ResonAnce fingerprin ting approach, referred to as HYDRA. Metho ds: HYDRA in v olves tw o stages: a mo del-based signature restoration phase and a learning-based parameter restoration phase. Signal restoration is implemen ted using lo w-rank based de-aliasing tec hniques while parameter restoration is performed using a deep nonlo cal residual conv olutional neural netw ork. The designed netw ork is trained on syn thesized MRF data simulated with the Bloch equations and fast imaging with steady state precession (FISP) sequences. In test mo de, it takes a temp oral MRF signal as input and pro duces the corresp onding tissue parameters. Results: W e v alidated our approach on b oth syn thetic data and anatomical data generated from a health y sub ject. The results demonstrate that, in con trast to conv en- tional dictionary-matc hing based MRF tec hniques, our approach significan tly impro ves inference sp eed by eliminating the time-consuming dictionary matc hing op eration, and alleviates discretization errors b y outputting contin uous-v alued parameters. W e further a void the need to store a large dictionary , th us reducing memory requirements. Conclusions: Our approach demonstrates adv antages in terms of inference speed, accuracy and storage requiremen ts ov er comp eting MRF metho ds. Key w ords: Magnetic Resonance Fingerprinting, Quantitativ e Magnetic Resonance Imaging, Deep Learning, Nonlo cal Residual Conv olutional Neural Netw ork, Self- atten tion This work was supp orte d b y the Royal So ciety International Exchange Scheme IE160348, by the Eur op e an Union ’s Horizon 2020 gr ant ER C-BNYQ, by the Isr ael Scienc e F oundation gr ant no. 335/14, by ICor e: the Isr aeli Exc el lenc e Center ’Cir cle of Light’, by the Ministry of Scienc e and T e chnolo gy, Isr ael, by UCL Overse as R ese ar ch Scholarship (UCL-ORS) and by China Scholarship Council (CSC) and by EPSRC gr ant EP/K033166/1. 1 Dep artment of Ele ctr onic and Ele ctrical Engine ering, Imp erial Col le ge L ondon, UK 2 F aculty of Mathematics and Computer Scienc e, W eizmann Institute of Scienc e, Israel 3 Dep artment of Ele ctric al Engine ering, T e chnion – Isr ael Institute of T e chnolo gy, Isr ael 4 Dep artment of Ele ctr onic and Ele ctrical Engine ering, University Col le ge L ondon, UK p.song@imp erial.ac.uk, yonina.eldar@weizmann.ac.il, galmazor@te chnion.ac.il, m.r o drigues@ucl.ac.uk i Hyb rid Deep MRF page 1 I . In tro duction Magnetic Resonance Fingerprinting (MRF) 1 , 2 , 3 , 4 , 5 , 6 , 7 , 8 has emerged as a promising Quan- titativ e Magnetic Resonance Imaging (QMRI) approach, with the capability of providing m ultiple tissue’s intrinsic spin parameters sim ultaneously , such as the spin-lattice magnetic relaxation time (T1) and the spin-spin magnetic relaxation time (T2). Based on the fact that the resp onse from each tissue with resp ect to a given pseudo-random pulse sequence is unique, MRF exploits pseudo-randomized acquisition parameters to create unique temp oral signal signatures, analogous to a "fingerprint", for differen t tissues. A dictionary matc hing op eration is then p erformed to map an inquiry temp oral signature to the b est matching en try in a precomputed dictionary , leading to multiple tissue parameters directly . The temp oral signatures are generated b y v arying the acquisition parameters of a pseudo-random excitation pulse sequence, such as rep etition time (TR), time of echo (TE), and radio frequency flip angle (F A) o v er time. The dictionary is comp osed of a large n umber of en tries that are usually sim ulated b y the Bloch equations giv en pseudo-random pulse se- quences. Each entry represents a unique temporal signature asso ciated with a sp ecific tissue and its quantitativ e parameters, such as the T1 and T2 relaxation times. Th us, once the b est matc hing (i.e. most correlated) en try is found, it directly leads to multiple quantitativ e parameters sim ultaneously via a lo okup-table op eration. MRI physics and ph ysiological constrain ts mak e the MR scanning pro cedure time- consuming. T o shorten acquisition time, subsampling is commonly performed in k-space (a.k.a conjugate F ourier transform domain) in order to reduce the n umber of samples and accelerate imaging sp eed. How ev er, suc h k-space subsampling results in temp oral signatures that are corrupted by aliasing, blurring and noise. This hamp ers the accuracy asso ciated with estimation of the tissue parameters using a dictionary matc hing pro cedure. In order to alleviate the impact of suc h distortion and corruption, de-aliasing op erations are often exploited to restore cleaner signatures b efore p erforming signature-to-parameter mapping. Therefore, MRF reconstruction usually inv olv es t wo op erations: signature restoration and parameter restoration. Inspired b y the successful application of sparsit y-driven image pro cessing approac hes in MRI reconstruction 9 , 10 , 11 , 12 , sev eral works 3 , 4 , 5 , 6 suggest to incorp orate prior knowledge such as sparsit y and low-rank to atten uate distortion and corruption, impro ving the signature page 2 P . Song restoration p erformance, during the initial MRF reconstruction stage. This is then follow ed b y a dictionary matching op eration, p erforming mapping from purified temp oral signatures to tissue’s quan titative parameters. Ho wev er, suc h dictionary matching based signature- to-parameter mapping exhibits sev eral drawbac ks 13 , 14 . Since the sim ulated dictionary and lo okup-table con tain a finite n um b er of elemen ts, they can only co ver a limited n umber of discrete v alues for eac h type of tissue parameter. W e refer to the difference betw een a con tinuous-v alued tissue parameter and its closest a v ailable discrete v alue on a lattice as the discretization error. F or example, a pair of dictionary and lo okup-table that con tain 101 elemen ts will lead to a discretization error of maxim um 25 ms if they cov er the range of 0 ms - 5000 ms with a fixed interv al of 50 ms for a specific tissue parameter, e.g. T1. T o reduce the discretization error, a h uge dictionary that is comp osed of a large n um b er of en tries is needed to represent tissues with fine gran ularit y ov er the entire v alue range of target tissue parameters. Ho w ever, storing a large dictionary becomes prohibitiv ely memory-consuming, as the dictionary size and densit y often increase exp onen tially with the num b er of tissue parameters. Sp ecifically , the n umber of en tries in a dictionary will b e P s for s parameters eac h containing P v alues, since every com bination of these s parameters determines a sp ecific tissue whic h is characterized by a sp ecific signature. F or example, given T1, T2 relaxation times, i.e. s = 2 , if each of them con tains 1000 v alues, the dictionary will hav e 1000 2 en tries. In addition, finding the b est matc hing en try b ecomes computationally intense for a large dictionary , considerably limiting the inference sp eed. In this pap er, w e prop ose an alternativ e approac h to dictionary matc hing based on deep neural netw orks (a.k.a. deep learning) 15 , 16 , whic h w e refer to as HYDRA: HYbrid Deep mag- netic ResonAnce fingerprin ting. The motiv ation deriv es from the fact that a w ell designed and tuned deep neural net work is capable of approximating complex functions, leading to state-of-the-art results in a n umber of tasks suc h as image classification, sup er-resolution, sp eec h recognition, and more 17 , 18 , 19 , 20 , 21 , 22 , 23 . Recen t work 13 , 14 prop osed to exploit neural net works to replace the dictionary and lo okup-table used in con v entional MRF reconstruc- tion approaches. These prop osed neural net w orks suffer from t w o limitations: First, these approac hes are based on neural net work mo dels con taining only 3-la y ers, th us suffer from limited capacit y of capturing complex mapping functions. Second, these metho ds fo cused exclusiv ely on parameter restoration stage (the second stage in MRF reconstruction), but not on signature restoration (the first stage in MRF reconstruction). Therefore, these techniques Hyb rid Deep MRF page 3 rely on fully-sampled data instead of t ypically av ailable sub-sampled k-space data. Differen t from Cohen et al.’s fully-connected feed-forward neural net work 13 , and Hopp e et al.’s v anilla conv olutional neural net w ork (CNN) 14 , the prop osed HYDRA inv olv es both a signature restoration and a parameter restoration phase. Signature restoration is im- plemen ted using a lo w-rank based de-aliasing method adapted from Mazor et al. 6 while parameter restoration is implemented using a deep nonlo cal residual conv olutional neural net work developed for this purp ose. Our key contributions with resp ect to prior work are: • HYDRA is, to the b est of our kno wledge, the first deep net work approac h to combine mo del-based de-aliasing and learning-based parameter mapping. HYDRA eliminates the requiremen t for the memory and time-consuming dictionary matching operation, th us significantly improving inference speed without compromising on reconstruction p erformance. • A 1D nonlo cal residual con volutional neural net w ork is designed to capture the map- pings from temp oral MRF signals to tissu e parameters. Owing to residual learning and a self-attention mec hanism, our netw ork is deep er and more sophisticated than comp eting netw ork mo dels. This allo ws to capture complex parameter mappings more effectiv ely , and output contin uous parameters to alleviate discretization issues. • The designed net work is trained on synthesized MRF data sim ulated with the Blo c h equations, but is still applicable to anatomical data. This con tributes to eliminating the requiremen t for a large amount of real MRF data. • A lo w-rank based de-aliasing tec hnique is dev elop ed in order to tak e adv antage of temp oral similarit y for signature restoration. • The lo w-rank based signature restoration is organically com bined with the learning- based parameter restoration to ac hieve fast and accurate MRF reconstruction. Suc h strategy enables HYDRA to handle b oth fully-sampled k-space data and more imp or- tan tly sub-sampled k-space data. • A series of n umerical exp erimen ts are conducted to ev aluate the proposed approach on b oth syn thetic and anatomical data. The results demonstrate impro ved inference sp eed, accuracy and discretization errors o ver comp eting metho ds 1 , 2 , 3 , 4 , 5 , 6 , 13 , 14 . The rest of the pap er is organized as follows. In Section I I . , w e form ulate the MRF reconstruction problem, in tro duce related methods, and present our approac h, inv olving the page 4 P . Song use of a low-rank based signature restoration pro cedure together with a deep net work for parameter restoration. Section II I . is dev oted to exp erimen tal results, follow ed b y a discussion in Section IV . and a conclusion in Section V . . I I . Materials and Metho ds I I . A . The MRF Problem F orm ulation MRF data is comp osed of m ultiple frames sampled in k-space o v er time. A series of suc h frames are v ectorized and then stac k ed together along the temp oral dimension to construct a measurement matrix Y ∈ C Q × L , where Q is the num b er of k-space samples in eac h frame, and L is the n umber of frames. Due to k-space subsampling, ev ery column v ector Y : ,i represen ts a subsampled F ourier transform of a vectorized image frame X : ,i : Y = [ Y : , 1 , · · · , Y : ,L ] = [ F u { X : , 1 } , · · · , F u { X : ,L } ] , (1) where F u {·} denotes a subsampled 2D F ourier transform. Eac h column X : ,i represen ts a MR contrast acquired with RF sequence parameters: Θ T RE : ,i = [ T R i , T E i , F A i ] T , i ∈ [1 , L ] (2) where T R i and T E i denote the rep etition time and ec ho time, resp ectiv ely , and F A i denotes the flip angle of the RF pulse during sampling the i -th contrast. Ev ery ro w X j, : represen ts a temp oral signature, i.e. temp oral signal evolution of a sp ecific tissue at the j -th image pixel. The signature dep ends on the tissue’s relaxation times, such as T1 and T2, group ed as a ro w vector: Θ T 12 j, : = [ T 1 j , T 2 j ] , j ∈ [1 , N ] (3) where, N denotes the num b er of pixels in each image frame. Note that, j is the spatial index while i is the temp oral index throughout. Given RF sequence parameters Θ T RE , and parameters Θ T 12 j, : of a sp ecific tissue, its temp oral signature X j, : can b e deriv ed as: X j, : = f ( Θ T 12 j, : , Θ T RE ) (4) where f ( · ) denotes the Blo ch equations. This MR con trast matrix X is associated with the k-space measuremen ts Y p er column by the subsampled F ourier transform, and it is related to tissue parameters Θ T 12 p er ro w by the Blo ch equations, as illustrated in Fig. 1 . Hyb rid Deep MRF page 5 Giv en RF sequence parameters Θ T RE and k-space measurements Y , the goal of MRF reconstruction is to estimate the tissue parameters Θ T 12 . T ypically , the image stac k X is first reconstructed from Y , referred to as signature restoration, and then mapped to tissue parameters Θ T 12 via dictionary matc hing, referred to as parameter restoration 1 , 2 , 3 , 4 , 5 , 6 . This pro cess is illustrated in Fig. 2 . The dictionary is a collection of temp oral signatures that are usually simulated by the Blo c h equations for v arious t ypical tissues, giv en the pseudo-random RF pulse sequences and tissue parameters. Giv en an inquiry temporal signature, dictionary matching computes the inner pro duct b etw een the temp oral signature with each dictionary en try , selecting the en try in the dictionary exhibiting the highest correlation with the inquiry one as the b est matching signature. Once the b est en try is found, it directly leads to multiple tissue parameters, suc h as T1, T2, sim ultaneously , via searching a lo okup-table. Let LUT ∈ R K × 2 denote a lo okup-table comp osed of K tissues, eac h con taining 2 parameters, i.e., T1 and T2 relaxation times 1 . Let D ∈ C K × L denote the corresp onding dictionary simulated using Bloch equations giv en the RF sequence parameters Θ T RE , for- m ulated as D k, : = f ( LUT k, : , Θ T RE ) . Since eac h temporal signature D k, : is link ed with the k -th tissue’s parameters LUT k, : , the choice of a large dictionary size K can in principle pro vide enough granularit y to capture a range of possible tissue v alues. In conclusion, existing MRF reconstruction approaches in volv e tw o stages: signature restoration and parameter restoration, that can b e succinctly written as Θ T 12 j, : = g ( h ( Y ) j, : | Θ T RE ) , j ∈ [1 , N ] , (5) where the function X = h ( Y ) represen ts the signature restoration op eration suc h as sparsit y or lo w-rank based de-aliasing and denoising metho ds, whereas g ( X j, : | Θ T RE ) denotes the parameter restoration op eration, suc h as dictionary matching based metho ds 1 , 2 , 3 , 4 , 5 , 6 . Our approac h aims to p erform signature restoration via low-rank based de-aliasing and parameter restoration via a neural netw ork in order to achiev e improv ed MRF reconstruction p erformance. W e highligh t that our metho d only requires a sim ulated dictionary during net work training. Once the net w ork is trained, the dictionary is not needed an ymore. In 1 Note that the off resonance parameter, whic h app eared in the original MRF pap er 1 , has been omitted here, since the sequence used in our experiments is deriv ed from the FISP sequence, which is insensitive to off resonance effects 2 , 6 . I I.A. The MRF Problem F ormulation page 6 P . Song Algorithm 1 Original MRF metho d 1 Input: A set of subsampled k-space images: Y A pre-sim ulated dictionary: D An appropriate lo okup-table: LUT Output: Magnetic parameter maps: b T 1 , b T 2 Step 1. Restore signatures: b X : ,i = F H u { Y : ,i } , ∀ i Step 2. Restore parameters for ev ery j via dictionary matc hing: b k j = arg max k Re D D k, : , b X j, : E k D k, : k 2 2 , b T j 1 , b T j 2 = LUT ( b k j ) addition, our approach also eliminates a sim ulated dictionary for signature restoration, whic h is a k ey difference from FLOR 5 , 6 during signature restoration. I I . B . Previous Metho ds Dictionary Matc hing based MRF approac hes. The original MRF reconstruction al- gorithm 1 is based on dictionary matching, as presen ted in Algorithm 1 . It finds the best matc hing dictionary en try for the acquired temporal s ignature according to their inner prod- uct and then searc hes the lo okup-table to obtain corresp onding tissue parameters. Here, F H u {·} denotes the in v erse F ourier transform op erating on the zero filled k-space data where zeros are filled at the unknown frequencies and symbol Re h a , b i represen ts the real part of the inner pro duct of t wo vectors a and b . Exploiting the nature of signals, by using appropriate prior kno wledge, can often con- tribute to impro ved signal pro cessing p erformance. In this spirit, later w orks suggested to incorporate sparsit y in MRF reconstruction to further impro ve performance, inspired b y successful applications of sparsit y in MRI reconstruction 9 , 10 , 11 . Da vies et al. 3 prop osed BLo c h resp onse recov ery via Iterative Pro jection (BLIP) whic h exploits sparsity in the dic- tionary domain. BLIP consists of iterating b etw een t wo main steps: (a) a gradient step whic h enforces consistency with the measuremen ts, based on the Pro jected Landw eb er Al- gorithm (PLA) generalized from the iterative hard thresholding metho d; (b) a pro jection whic h matc hes each row of X to a single dictionary atom. Instead of exploiting sparsit y in I I.B. Previous Metho ds Hyb rid Deep MRF page 7 the dictionary domain, W ang et al. 4 suggested to lev erage sparsity in the w a velet domain of eac h imaging frame, X : ,i . They further replaced the Euclidean norm with the Mahalanobis distance for dictionary matc hing. Considering that adjacent MR image frames along the temp oral dimension should exhibit high resem blance, Mazor et al. 5 , 6 prop osed a magnetic resonance Fingerprin t with LOw-Rank prior for reconstructing the image stack and quan ti- tativ e parameters, referred to as FLOR, which ac hiev ed state-of-the-art p erformance. The algorithm, describ ed in Algorithm 2 , relies on t w o priors: a lo w rank prior on the the matrix X , and the fact that the ro ws of X lie in the column space of the dictionary D . Algorithm 2 FLOR 6 Input: A set of subsampled k-space images: Y ; A pre-simulated dictionary: D ; An appropriate lookup- table: LUT ; P arameters µ for gradient step and λ for regularization Output: Magnetic parameter maps: b T 1 , b T 2 Initialization: b X 0 = 0 , P = D † D , where D † is the pseudo-in verse of D . Step 1. Restore signatures via iterating un til conv ergence: • Gradien t step for every i : b Z t +1 : ,i = b X t : ,i − µF H u { F u { b X t : ,i } − Y : ,i } (6) where the sup erscript t represen ts the index of iterations. • Pro ject onto the dictionary subspace: [ U , S , V ] = svd( b Z t +1 P ) (7) where svd denotes the singular-v alue decomp osition op eration, and S = diag ( { σ j } ) is a rectangular diagonal matrix with singular v alues { σ j } on its diagonal. • Soft-threshold the non-zero singular v alues with λµ and reconstruct signatures b X t +1 : σ 0 j = max { σ j − λµ, 0 } , b X t +1 = US 0 V H (8) where S 0 = diag ( { σ 0 j } ) . Step 2. Restore parameters for ev ery j via dictionary matc hing: b k j = arg max k Re D D k, : , b X j, : E k D k, : k 2 2 , b T j 1 , b T j 2 = LUT ( b k j ) Learning-based MRF approaches. The ab ov e tec hniques all use dictionary matching I I.B. Previous Metho ds page 8 P . Song to p erform mapping from temp oral signatures to tissue parameters. Therefore, these meth- o ds suffer from drawbac ks suc h as discretization error, slo w inference sp eed and memory- consuming storage. In order to alleviate these issues, recent works 13 , 14 prop ose to exploit neural netw orks to replace dictionaries and lookup-tables used in con v entional MRF recon- struction approaches. Cohen et al. suggest a fully-connected feed-forw ard neural net work (FNN) 13 . Since the input la y er of the FNN is fully connected with the input temporal signature, the n um b er of neurons in the input la y er corresp onds to the length of the input temp oral signature. This mak es the net work structure less flexible, as a FNN net w ork trained on temporal signatures with a certain length is not applicable to temporal signatures with a differen t length. In addition, the fully-connected structure results in rapid increase in the n umber of parameters along with the gro wth of depth, making the net w ork more suscepti- ble to o v erfitting. Hopp e et al. 14 prop ose a 3-la y er v anilla CNN for parameter restoration. Both 13 and 14 fo cus exclusively on learning the signature-to-parameter mapping from a pair of dictionary and lookup-table sim ulated using the Blo ch equations. During the v alidation, they assume that clean temp oral signatures are a v ailable as input in to the trained netw orks. Ho wev er, since temp oral signatures obtained from k-space subsampled MRF data are al- w ays contaminated by aliasing and noise, their approac hes, when applied directly in suc h k-space subsampling situations, suffer from hea vy artifacts introduced during the signature restoration phase, leading to p o or p erformance. I I . C . Prop osed Metho ds The prop osed h ybrid deep magnetic resonance fingerprin ting (HYDRA) approach, summa- rized in Algorithm 3 , consists of tw o stages: signature restoration and parameter restoration, (see also ( 5 )). As illustrated in Fig. 3 , a lo w-rank based de-aliasing method is used to restore signatures, and then a 1D nonlo cal residual conv olutional neural netw ork is used to map eac h restored signature to corresp onding tissue parameters. In particular, given Θ T RE and k-space samples Y , in our prop osed approach, the func- tion X = h ( Y ) in ( 5 ) represen ts a signature restoration op eration using low-rank based de-aliasing tec hniques without requiring a dictionary . The function Θ T 12 j, : = g ( X j, : | Θ T RE ) in ( 5 ) represen ts a parameter restoration op eration that exploits a trained neural net work to map each restored signature X j, : to corresp onding tissue parameters Θ T 12 j, : directly . In the subsequen t sections, we provide a detailed description of b oth stages of our technique. I I.C. Prop osed Metho ds Hyb rid Deep MRF page 9 I I . C . 1 . Lo w-rank based signature restoration Since MRF data consists of m ultiple frames exhibiting temporal similarit y , the imaging con trasts matrix X is typically a low-rank matrix 6 . Therefore, h ( · ) leverages a low-rank prior for denoising and de-aliasing, form ulated as h ( Y ) = arg min X 1 2 P i k Y : ,i − F u { X : ,i }k 2 2 s.t. rank ( X ) < r (9) where the parameter r is the rank of the matrix, a fixed pre-c hosen parameter. Since t ypically r is not known in adv ance, we consider a relaxed regularized version: h ( Y ) = arg min X 1 2 X i k Y : ,i − F u { X : ,i }k 2 2 + λ k X k ∗ (10) where k X k ∗ denotes the n uclear norm 24 of X , defined as the sum of the singular v alues of X , and λ is the Lagrangian multiplier manually selected for balancing data fidelity and the rank. Problem ( 10 ) can b e solved using the incremental subgradien t pro ximal metho d 25 , similar as to FLOR 6 . The pro cedure for solving ( 5 ) is shown in Algorithm 3 . One of differences from FLOR 6 is the fact that w e remov ed the op eration of pro ject- ing the temp oral signal on to a dictionary . This allo ws to eliminate the requiremen t for a sim ulated dictionary in the signature restoration stage, whic h also alleviates the memory consumption issue. In addition, the computational complexity is reduced by N · L 2 floating- p oin t op erations in eac h iteration, where L is the dimension of a dictionary elemen t, and N is the n um b er of pixels in each image frame. On the other hand, the gained b enefits are at the price of requiring more iterations to conv erge. Another difference from FLOR 6 is that w e exploit a netw ork, instead of dictionary matc hing, for signature-to-parameter mapping. I I . C . 2 . Learning-based parameter restoration Once the imaging con trasts matrix X is reco v ered from the k-space samples Y , each temporal signature X j, : is input in to the trained netw ork for parameter restoration, form ulated as: Θ T 12 j, : = g ( X j, : | Θ T RE ) , j ∈ [1 , N ] (11) where g ( · ) denotes the trained net w ork, Θ T RE denotes the fixed RF sequence parameters. W e next describ e the net work structure, training and testing pro cedures. Net w ork structure. I I.C. Prop osed Metho ds page 10 P . Song Algorithm 3 Prop osed MRF reconstruction approach: HYDRA Input: A set of subsampled k-space images: Y The trained net work: g P arameters µ for gradient step and λ for regularization Output: Magnetic parameter maps b T 1 , b T 2 Initialization: b X 0 = 0 Step 1. Restore signatures via iterating un til conv ergence: • Gradien t step for every i , the same as ( 6 ). • P erform SVD: [ U , S , V ] = svd( b Z t +1 ) • Soft-threshold the non-zero singular v alues { σ j } of S with parameter λµ and reconstruct signatures b X t +1 , the same as ( 8 ). Step 2. Restore parameters for ev ery j via the trained netw ork: b T j 1 , b T j 2 = g ( b X j, : ) The prop osed netw ork has a 1D nonlo cal residual CNN arc hitecture with short-cuts for residual learning and nonlocal op erations for ac hieving a self-attention mec hanism. As illustrated in Fig. 3 , it starts with t w o 1D conv olutional lay ers b efore connecting with 4 residual / non-lo cal op eration blo c ks, and finally ends with a global-a verage-po oling la yer follo wed by a fully-connected lay er. Every residual blo c k is follo w ed b y a non-lo cal operation blo c k. F our such blo cks are interspersed with each other. Eac h residual blo ck contains a max-po oling la yer with stride 2, t wo conv olution lay ers and a shortcut that enforces the netw ork to learn the residual conten t. The filter size is set to be equal to 21 throughout con volutional lay ers. The n um b er of c hannels, a.k.a feature maps, in the first t wo con volutional lay ers is set to 16 and then is doubled in subsequen t four residual blocks until 128 in the final residual blo ck. The size of feature maps in the next blo c k halves in con trast with the previous one due to max-po oling. In this w a y , w e gradually reduce temp oral resolution while extract more features to increase con tent information. Inspired by the self-attention scheme and nonlo cal neural net works 26 , 27 , non-local op- erations are incorp orated into the designed netw ork to ac hieve the atten tion mec hanism, in order to capture long-range dependencies with few er la yers. In contrast to the progressive b eha vior of con v olutional operations that pro cess one lo cal neighborho o d at a time, the non- I I.C. Prop osed Metho ds Hyb rid Deep MRF page 11 lo cal op erations compute the resp onse at a p osition as a w eighted sum of the features at all p ositions in the feature maps. F ormally , the nonlo cal op eration is formulated as 27 : y i = 1 C ( x ) X ∀ j f ( x i , x j ) g ( x j ) . (12) Here, i is the index of an output p osition and j is the index that en umerates all p ossible temp oral p ositions, x is the input temp oral signal or its features and y is the output signal of the same size as x . A pairwise function f computes a scalar betw een i and all j to represen t the affinity relationship of these tw o p ositions. The unary function g computes a represen tation of the input signal at p osition j . The resp onse is normalized b y a factor C ( x ) . There exists a few instan tiations for function f and g . F or simplicity , the unary function g is c hosen as a linear embedding: g ( x j ) = W g ( x j ) , where W g is a weigh t matrix to b e learned. Regarding the affinit y matrix f , w e adopt the em b edded Gaussian to compute similarit y in an em b edding space, whic h is form ulated as: f ( x i , x j ) = e θ ( x i ) > φ ( x j ) . Here, θ ( x i ) = W θ x i and φ ( x j ) = W φ x j are tw o learned embeddings. The normalization factor is set as C ( x ) = P ∀ j f ( x i , x j ) . F or a given i , 1 C ( x ) f ( x i , x j ) b ecomes the softmax computation along the dimension j , whic h leads to y = softmax ( x > W > θ W φ x ) g ( x ) , whic h is the self- atten tion form 26 . The non-lo cal b ehavior in ( 12 ) is due to the fact that all p ositions ( j ) are considered in the op eration. As a comparison, a con volutional operation sums up the w eighted input in a lo cal neigh b orho o d 27 . It implies that the non-lo cal op eration directly captures long-range dep endencies in the temp oral dimension via computing interactions b et ween any t wo p oin ts, regardless of their p ositional distance. In this wa y , the netw ork is able to extract global features and tak e adv an tage of the full receptive field in each la yer. The global-av erage-p o oling la y er is used to av erage each feature map in order to in tegrate information in every channel for impro ved robustness to corrupted input data. This global- a verage-po oling la yer also reduces the n um b er of parameters significan tly , th us lessening the computation cost as w ell as prev enting ov er-fitting. The last fully-connected la y er outputs estimated parameters – T1 and T2 relaxation times. The designed netw ork con tains around 0.27 million parameters. The w eights are initialized using He-normal-distribution 28 . The max-norm k ernel constrain t 29 is exploited to regularize the w eigh t matrix directly in order to prev ent o v er-fitting. The designed netw ork can also b e adapted for v arious MRF sequences, suc h as the original MRF sequence – in v ersion-recov ery balanced steady state free-precession I I.C. Prop osed Metho ds page 12 P . Song (IR-bSSFP) sequence, that dep ends also on the in trinsic df parameter. It is p ossible to adjust the n umber of outputs to adapt to more parameters, such as proton density , B0. T o summarize, our netw ork is motiv ated and inspired by recent successful applications of conv olutional neural net w orks and v arian ts. Conv olutional neural netw orks hav e b een pro ved to b e a p o werful mo del to capture useful features from signals and images. By in tro ducing conv olution, local receptive field and weigh t sharing design, a CNN is capable of taking adv an tage of lo cal spatial coherence and translation inv ariance c haracteristics in the input signal, thus b ecome esp ecially w ell suited to extract relev ant information at a low computational cost 17 , 18 , 19 , 20 , 21 , 22 . On the other hand, the residual net work arc hitecture 18 , 19 pro vides an effective w a y to design and train a deep er mo del, since it alleviates the gradient v anishing or exploding problems by propagating gradients throughout the mo del via short- cuts, a.k.a skip connections. By leveraging non-lo cal operation based attention mechanism, neural net w orks are endo wed with capabilit y of extracting global features and capturing long-range dep endencies. Net w ork training. The designed netw ork is trained on a syn thesized dictionary D and corresp onding lo okup-table LUT to learn the signature-to-parameter mappings LUT k, : = g ( D k, : | Θ T RE ) . The training dataset is synthesized as follo ws. First, w e determine the range of tissue parameters. F or example, one ma y set T1 relaxation times to co ver a range of [1, 5000] ms and T2 relaxation times to co ver a range of [1, 2000] ms with an increment of 10 ms for b oth. Th us, the T1 and T2 v alues constitute a grid with dimension 500 × 200, in which each p oin t represen ts a sp ecific combination of T1 and T2 v alues, and hence characterizes a sp ecific tissue. P oin ts corresponding to T1 < T2 hav e been excluded as such combinations ha ve no ph ysical meaning. All the v alid points are s tac k ed together to generate a l ookup-table. F or instance, the ab ov e setting for T1 and T2 leads to a lo okup-table of dimension 80100 × 2 . The RF pulse sequences used in our w ork are fast imaging with steady state precession (FISP) pulse sequences with parameters that hav e b een used in previous publications in the field of MRF 2 , 6 , 8 . Giv en the lo okup-table and RF pulse sequences, dictionary entries can b e syn thesized by solving the Blo ch equations using the extended phase graph formalism 30 , 31 . When the training dataset is ready , the dictionary en tries are used as input signals and I I.C. Prop osed Metho ds Hyb rid Deep MRF page 13 corresp onding lookup-table entries serve as the groundtruth. All the dictionary entries are input in to the designed netw ork batc h b y batc h whic h outputs estimated parameters. The ro ot mean square errors (RMSE) of the outputs are calculated with respect to corresp onding groundtruth. The resulting RMSE loss is then backpropagated from the output la yer to the first la yer to up date the weigh ts and bias b y using Adam 32 as the optimization algorithm. More training details are pro vided in the subsequent exp eriment section. Once the training pro cedure is completed, given an inquiry signal ev olution X j, : , it is able to map such a time sequence directly to corresp onding tissue parameters, as formulated in ( 11 ), implying that no dictionary or lo okup-table are required during the inference. Since w e only need to store the trained net w ork whic h is a compact mo del, it consumes less memory than storing the dictionary and lo okup-table. W e emphasize that even though the net work is trained on a grid of tissue v alues, it is exp ected to capture the mapping function from temporal signatures to tissue parameters. Th us the trained net work is capable of outputting tissue v alues not existing in the grid of training v alues. Detailed results can b e found in Fig. 7 and T able 2 . This feature is fa vorable, as it implies that well designed and trained netw orks hav e an ability to o vercome discretization issues. The o verall procedures for solving ( 10 ) and ( 11 ) are sho wn in Algorithm 3 . I I I . Exp erimen tal Results In this section, we conduct a series of experiments to ev aluate our approach, comparing it with other state-of-the-art MRF metho ds 1 , 3 , 6 , 13 , 14 . The exp erimen ts are categorized in to a few t yp es: training, testing on syn thetic data, testing on anatomical data using v ariable densit y Gaussian sampling patterns and spiral sampling patterns at differen t sampling ratios and num b er of time frames, as described in T able 1 . F or the net work training, synthesized temp oral signatures, i.e. simulated dictionary en tries of D shown as Fig. 4 , are used as input signals and corresponding parameter v alues in the lo okup-table LUT serv e as the groundtruth. The prop osed net work is trained to capture the signature-to-parameter mappin gs. F or testing on synthetic data, syn thesized temp oral signatures in X are used as input signals and corresp onding parameter v alues in Θ T 12 serv e as the groundtruth. The aim is to test the parameter restoration performance page 14 P . Song only . F or testing on anatomical data, the k-space measuremen ts Y whic h are deriv ed from the F ourier transform (for Gaussian patterns) or non-uniform FFT (for spiral tra jectories) 33 of X , are used as input and corresp onding parameter v alues in Θ T 12 serv e as reference. When there is no k-space subsampling, the aim is to test the parameter restoration performance only . When there exists k-space subsampling, the aim is to test the ov erall p erformance, including b oth signature restoration and parameter restoration. More detailed descriptions are pro vided in each subsection. I I I . A . T raining As mentioned in Section I I . C . 2 . , the designed net work is trained on a pair of syn thesized dictionary D and lo okup-table LUT , simulated using Blo c h equations and FISP pulse se- quences 2 , 6 . The FISP pulse sequence used in our experiments was designed with parameters Θ T RE : ,i = [ T R i , T E i , F A i ] T , i ∈ [1 , L ] that ha ve b een used in previous publications in the field of MRF 2 , 6 , 8 . The ec ho time T E i w as constant of 2ms. The rep etition time T R i w as randomly v aried in the range of 11.5 - 14.5 ms with a Perlin noise pattern. All the flip angles F A i , i ∈ [1 , L ] constituted a sin usoidal v ariation in the range of 0 - 70 degrees to ensure smoothly v arying transien t state of the magnetization, as shown in Figure 5 . F or the range of tissue parameters, T1 relaxation times are set to co ver a range of [1, 5000] ms and T2 relaxation times to cov er a range of [1, 2000] ms with an increment of 10 ms for b oth. Suc h parameter ranges co ver the relaxation time v alues that can be commonly found in a brain scan 34 . All the v alid combinations of T1 and T2 v alues are stac ked together, generating a lo okup-table LUT of dimension K × 2 where K = 80100 . Giv en the lo okup-table and RF pulse sequences, dictionary entries are syn thesized b y solving the Blo ch equations using the extended phase graph formalism, leading to a dictionary of dimension K × L where L = 200 or 1000 is the n umber of time frames. When the training dataset is ready , the dictionary en tries are used as input signals and corresp onding lookup-table en tries serv e as the groundtruth to train the designed net work, as mentioned in Section I I . C . 2 . . The model w as trained for 50 epo chs. It takes around 30 seconds for running one ep o c h on a verage, th us around 25 min utes for completing 50 ep o chs, on a NVIDIA GeF orce GTX 1080 Ti GPU. In each training ep o c h, 20% of the training I I I.A. T raining Hyb rid Deep MRF page 15 samples are separated aside for v alidation dataset. The learning rate deca ys from 1e-2 to 1e-6 every 10 epo chs. Eac h batc h w as experimentally set to contain 256 time-sequences in order to balance the con vergence rate and w eights up dating rate w ell. F or comparison purp oses, w e also implemented Hopp e et al.’s CNN referring to 14 , and Cohen et al.’s FNN referring to 13 with the same structure and parameters as sp ecified in their pap ers. Then we use the same GPU and training dataset to train their netw orks with sp ecified learning rate and n umber of ep o chs until con vergence. W e adopt a few widely used metrics, suc h as ro ot mean square error (RMSE), signal- to-noise ratio (SNR) and peak signal-to-noise ratio (PSNR) to ev aluate the image quality quan titatively . The definitions of RMSE, SNR and PS NR are given as follows: RMSE = s k X − b X k 2 F N , (13) SNR = 20 log 10 k X k 2 F RMSE , (14) PSNR = 20 log 10 P eak V al RMSE , (15) where matrices X and b X denote the ground truth signal and its reconstructed v ersion, resp ectiv ely , N denotes the total num b er of elemen ts in the signal and k · k F denotes the F rob enius norm. Peak V al stands for the pixel p eak v alue in an image, e.g., 1 for a normalized signal. I I I . B . T esting on syn thetic dataset In this subsection, w e ev aluate the p erformance of HYDRA on a syn thetic testing dataset. The procedures of constructing a syn thetic testing dataset is similar to the construction of the training dataset: 500 differen t T1 v alues are randomly selected from 1 - 5000 ms, while 200 differen t T2 v alues are randomly selected from 1 - 2000 ms, using random p ermutation based on uniformly distributed pseudorandom num b ers. All the v alid com binations from the selected T1 and T2 v alues are stac ked together, generating a parameter matrix Θ T 12 of dimension 80000 × 2 with N = 80000 . The RF pulse sequences are the same as in the training stage. Given the parameter matrix and RF pulse sequences, input signal signatures are syn thesized b y solving the Blo c h equations using the extended phase graph formalism, leading to a signature matrix X of dimension N × L = 80000 × 200 , with each row representing I I I.B. T esting on synthetic dataset page 16 P . Song a temp oral signature corresp onding to a sp ecific com bination of T1 and T2 v alues. The signature matrix X and parameter matrix Θ T 12 constitute the syn thetic testing dataset, with X as input and Θ T 12 as the groundtruth. W e input the synthetic testing signatures X into Hopp e et al.’s CNN 14 , Cohen et al.’s FNN 13 , and the net work of HYDRA to compare the outputs with groundtruth T1 and T2 v alues in Θ T 12 . W e also compare with dictionary matc hing metho ds 1 , 2 , 3 , 4 , 5 , 6 whic h exploit the same dictionary D and lo okup-table LUT to find the b est matching entry for each sig- nature in X and then estimate parameter v alues by searc hing the lo okup-table. As sho wn in T able 2 , T able 3 , Fig. 6 and Fig. 7 , the estimated parameter v alues using the proposed net work obtained outstanding agreement with the groundtruth, yielding higher PSNR, SNR and smaller RMSE than the dictionary matc hing metho d 1 , 2 , 3 , 4 , 5 , 6 , as well as comp eting net- w orks 13 , 14 . In particular, to illustrate in detail ho w well neural net works tac kle the discretization issue inherent to dictionary matching, we sho w the testing p erformance on con tinuous-v alued T1, T2 parameters which ha ve small in terv als, e.g. 0.5ms, that is 20 times smaller than the training grid interv als 10ms, b etw een neigh b oring v alues in T able 3 . Since these v alues and their corresponding MRF signatures do not exist in the training dictionary and lo okup-table 2 , the dictionary matching methods report a T1 and T2 v alue – the closest discretized v alue presen t in the dictionary – that can b e quite distinct from the groundtruth. In contrast, the v arious neural netw ork approaches can p oten tially learn an underlying mapping from the temp oral signatures to the resp ectiv e T1 and T2 v alues, leading to estimates that are muc h closer to the groundtruth. In terestingly , our approach outp erforms previous netw orks 13 , 14 as sho wn in T able 3 and Fig. 6 . Eviden tly , neural net works demonstrate m uch b etter ro- bustness to discretization issues, leading to improv ed parameter restoration in comparison to dictionary based metho ds. Another impressive adv antage of HYDRA is the fast inference sp eed. HYDRA tak es only 8.2 s to complete the mapping op eration for eigh t y thousand temp oral signatures, that is, 53 × faster than dictionary matching. F urthermore, the inference sp eed of HYDRA is sub ject 2 As mentioned in the exp eriment setting in Sec tion I I I . A . , in the training dataset, T1 relaxation times are set to cov er a range of [1, 5000] ms and T2 relaxation times to cov er a range of [1, 2000] ms with an incremen t of 10 ms for b oth, that is, T1 v alues = {1, 11, 21, · · · , 4991}, and T2 v alues = {1, 11, 21, · · · , 1991}. I I I.B. T esting on synthetic dataset Hyb rid Deep MRF page 17 to the netw ork top ology . That is, once the net work structure is fixed, the complexity is fixed. In con trast, the complexit y of dictionary matching is limited b y the dictionary density . This implies that our adv antage will b e more prominent in comparison with comp eting techniques using a dictionary with higher densit y . I I I . C . T esting on anatomical dataset In this subsection, w e ev aluate our approach on an anatomical testing dataset. W e construct the dataset from brain scans that w ere acquired with GE Signa 3T HDXT scanner from a health y sub ject. 3 Since there are no groundtruth parameter v alues for the T1 and T2 param- eter maps, w e obtain gold standard data by acquiring F ast Imaging Emplo ying Steady-state A cquisition (FIEST A) and Sp oiled Gradient Recalled Acquisition in Steady State (SPGR) images, at 4 differen t flip angles (3 ◦ ,5 ◦ ,12 ◦ and 20 ◦ ), and implementing corrections 35 fol- lo wed b y DESPOT1 and DESPOT2 36 algorithms. The constructed gold standard T1, T2 parameter maps ha ve a dimension of 128 × 128 for eac h map, accordingly leading to a pa- rameter matrix Θ T 12 of size 16384 × 2 by stac king vectorized T1, T2 maps together. Based on the parameter matrix Θ T 12 and pre-defined RF pulse sequences, w e generate temp oral signatures using Bloch equations, the same mechanism as generating the syn thetic testing dataset, leading to a signature matrix X of dimension N × L = 16384 × 200 . The signature matrix X and parameter matrix Θ T 12 constitute the anatomical testing dataset, with X as input and Θ T 12 as the gold standard reference. Note that, since the gold standard T1, T2 maps exhibit spatial structures in the image domain, the resulting signature matrix X can be regarded as a stack of L = 200 vectorized image frames, where eac h frame exhibits sp ecific spatial structures. Therefore, it makes sense to p erform F ourier transform and k-space subsampling for eac h column of X to get k-space measurements Y . This is the k ey difference b etw een the anatomical dataset and the syn thetic dataset. W e first explore the case with full k-space sampling in order to ev aluate the parameter restoration performance of HYDRA. Then, we consider situations with k-space subsampling in order to ev aluate both the signature restoration and the parameter restoration performance of HYDRA. 3 The exp eriment pro cedures in volving human sub jects describ ed in this pap er w ere appro ved b y the Institutional Review Board of T el-A viv Sourasky Medical Center, Israel. I I I.C. T esting on anatomical dataset page 18 P . Song I I I . C . 1 . F ull k-space sampling In the first case, the fully-sampled k-space measurements Y , derived from the F ourier trans- form of X , are used as input to obtain the estimated Θ T 12 . This is equiv alent to inputting X in to the netw ork of HYDRA, or performing dictionary matching based on X directly , since the in verse F ourier transform of the fully-sampled measuremen ts Y is exactly the same as X . The aim is to test the parameter restoration p erformance only . In the experiment, cor- resp onding parameter v alues in Θ T 12 serv e as the gold standard reference. F or comparison, dictionary matching metho ds 1 , 2 , 3 , 4 , 5 , 6 exploit the same dictionary D and lo okup-table LUT as in our training stage to find the b est matching entry and estimate parameter v alues for eac h signature in X . Visual and quan titativ e results are sho wn in Fig. 8 , Fig. 9 and T able 4 . It can be seen that our basic version of HYDRA outp erforms dictionary matc hing 1 , 2 , 3 , 4 , 5 , 6 , yielding b etter visual and quan titativ e p erformance, e.g., 7.9 dB SNR gains for T2 mapping. The RMSE of T2 mapping is also reduced to 2.498 from 6.252, accordingly . Our nonlo cal v ersion of HYDRA ac hieves ev en better p erformance, leading to 10 dB SNR gains with RMSE as small as 1.86. This is o wing to the adv antage that the trained net work is a pow erful function appro ximator, which is able to output well-estimated parameter v alues based on learn t mapping functions, ev en though these v alues do not exist in the training dictionary and lo okup-table. In con trast, dictionary matching only matc hes signatures to discrete parameters existing in the training dataset. In other w ords, if there are no exact matc hing dictionary elemen t and parameter v alues for an inquiry MRF signature, it will find adjacen t v alues as appro ximations, thus introducing discretization error. On the other hand, the adv an tage of HYDRA o ver dictionary matc hing on T1 mapping is not as significant as on T2 mapping quan titatively . But the visual impro vemen ts are eviden t. A similar trend is observ ed when comparing our netw ork with competing netw orks suc h as Hoppe et al.’s CNN 14 and Cohen et al.’s FNN 13 . In addition, HYDRA tak es around 2 s to accomplish the mapping for 16384 signatures, 40 × faster than dictionary matc hing 1 , 2 , 3 , 4 , 5 , 6 . I I I . C . 2 . k-space subsampling using Gaussian patterns In k-space subsampling situations, the developed lo w-rank based de-aliasing method is ap- plied to restore the signature matrix X from the measuremen ts matrix Y . Then, the re- I I I.C. T esting on anatomical dataset Hyb rid Deep MRF page 19 constructed X is used as input in to the net work for parameter mapping to obtain the cor- resp onding tissue parameter v alues. In the exp erimen ts, the sub-sampling factor β is set to b e 70% and 15%. F or β = 15%, 15% k-space data is acquired by a series of 2D random Gaussian sampling patterns, sho wn in Fig. 10 , leading to a k-space measuremen t matrix Y of size Q × L = 2458 × 200 . Similarly , β = 70% giv es rise to a k-space measurement matrix Y of size Q × L = 11469 × 200 . A larger λ enforces low er rank for the restored signature matrix X to strengthen the de-aliasing effect, while a smaller λ encourages X to ha ve a sub- sampled F ourier transform that approximates the k-space measuremen ts matrix Y b etter. Therefore, w e tried a range of v alues from 1 to 20 for λ and exp erimentally select the best one λ = 5 . Since the lo w-rank based signature restoration inv olves gradien t descen t steps, a larger step size µ accelerates gradien t descent sp eed, but tends to result in oscillation or ev en div ergence, while a smaller µ leads to a slow er conv ergence. W e exp erimen tally find that µ = 1 gives a go o d balance. The same k-space measuremen ts Y are also used b y dictionary- matc hing based metho ds 1 , 3 , 6 for comparison, and the same signature restoration approach is used to con v ert Y onto X for learning based metho ds 13 , 14 . The aim is to ev aluate the o verall p erformance on b oth signature restoration and parameter restoration. Quan titative performance is shown in T able 5 . Note that the adv antage of learning- based metho ds o ver dictionary matching degrades when the subsampling factor increases. This is due to the fact that the restored signatures from highly subsampled k-space data exhibit deviations and distortions, th us leading to p o orer input for the trained netw orks. In spite of this, the prop osed approac h outp erforms the dictionary matching based metho ds 1 , 3 with significant gains, and also yields b etter or comparable p erformance as the state-of-the- art metho ds FLOR 6 , CNN 14 and FNN 13 . In addition, it tak es around 23s for lo w-rank based signature restoration and less than 3s for net work based parameter restoration. Th us, the total time cost is around 26s, almost 4.8 × faster than FLOR 6 . F urthermore, the sp eed of our metho d is 60 × faster than FLOR 6 for parameter restoration. W e compared the p erformance with/without nonlo cal op erations in our developed net- w ork. The results in T able 4 and 5 show that the prop osed netw ork with nonlo cal op erations based self-atten tion scheme outp erforms the basic coun terpart. In particular, the nonlocal v ersion achiev es 6 dB gains in terms of SNR ov er the basic version for T2 mapping. Such significan t improv ement demonstrates the b enefits of capturing long-range dep endencies and global features using the nonlo cal op eration based atten tion scheme. I I I.C. T esting on anatomical dataset page 20 P . Song W e also inv estigated the performance with resp ect to the n umber of time frames. In particular, we increased L from 200 to 1000 and kept other exp erimen t settings the same as b efore. The quantitativ e results are sho wn in T able 6 . It is noticed that giv en more time frames, all the metho ds show b etter p erformance. Moreov er, the p erformance of learning- based metho ds, including CNN 14 , FNN 13 and HYDRA, improv e more than model-based tec hniques 1 , 3 , 6 . In particular, our approac h outp erforms comp eting algorithms quan titativ ely in terms of PSRN, SNR, and RMSE, as well as demonstrates visual adv antage, as shown in Figure 11 and Figure 12 . I I I . C . 3 . k-space subsampling using spiral tra jectories W e carried out additional exp erimen ts with widely used non-Cartesian sampling patterns – v ariable densit y spiral tra jectories 6 , 37 . A set of spiral tra jectories used in the exp eriments are sho wn in Figure 13 . They ha v e F OV of 24 and rotation angle difference of 7.5 degrees betw een an y tw o adjacent spirals to spread out the alias artifacts. Giv en suc h spiral tra jectories, data were subsampled to acquire 1488 k-space samples in each time frame, leading to a subsampling ratio of 9% whic h is defined by the n umber of acquired samples in the k-space domain divided b y the n umber of pixels in a frame. This setting closely matc hes the original MRF pap er 1 where eac h single spiral tra jectory samples 1450 k-space p oints (leading to a subsampling ratio around 9%) and any t wo adjacent spiral tra jectories hav e a rotation angle of 7.5 degrees. In the case of spiral subsampling, during the signature restoration, SP arse Uniform Re- Sampling (SPURS) algorithm 38 w as exploited to implement non uniform F ourier transform b et ween k-space domain and image domain, as SPURS has pro ved to achiev e smaller appro x- imation errors while maintaining lo w computational cost comparing with other resampling metho ds, suc h as nonuniform-FFT algorithm 33 and regularized Blo ck Uniform ReSampling (rBURS) 39 . In the exp erimen ts, 1000 densit y v ariable spiral tra jectories were used for k-space subsampling, leading to 1000 time frames. The quan titativ e and qualitative reconstruction results demonstrate that HYDRA outp erforms comp eting metho ds with smaller estimation errors, as sho wn in T able 7 , Figure 15 and Figure 14 . I I I.C. T esting on anatomical dataset Hyb rid Deep MRF page 21 IV . Discussion IV . A . Relation to previous w orks Our lo w-rank based signature restoration metho d is adapted from FLOR 6 b y removing the op eration of pro jecting the temp oral signal on to a dictionary . Th us, the signature restoration do es not require a sim ulated dictionary , and sa ves computational cost. Although recen t w orks 13 , 14 exploit neural net works to p erform parameter mapping, replacing dictionaries and lo okup-tables used in conv entional MRF reconstruction approac hes, our tec hnique is differen t from these metho ds 13 , 14 . W e design a deep nonlo cal residual CNN for capturing signature-to- parameter mapping which is organically combined with lo w-rank based de-aliasing tec hniques for signature restoration. In this wa y , our algorithm can bypass some of the issues asso ciated with other tec hniques: (1) The input dimension issue. The proposed approac h can ingest temp oral signatures with differen t lengths without the need to c hange the structure of the net work. This is due to the fact that w e rely on conv olutional neural net works (CNNs) rather than fully-connected neural net w orks (FNNs) such as the mo del used in 13 . (2) The k-space subsampling issue. The prop osed approach inv olves a hybrid of a neural netw ork with a lo w-rank based de-aliasing approac h. Thus it is able to deal with correlations b oth o ver time and space via exploiting low-rank regularization and conv olution operation. This enables our work to handle k-space subsampling situations. (3) The complex mappings issue. By exploiting a residual netw ork structure, our metho d can b e successfully extended to deep er lev els and th us obtain a better capacit y to learn complex signature-to-parameter mapping functions. (4) Distortion and corruption issue. Due to the subsampling in k-space, the restored temp oral signatures suffer from lo cal distortion and corruption. Suc h deviation ma y lead to performance degeneration in the second stage. By incorporating non-lo cal op erations in the netw ork design, our metho d is able to capture global features and find most relev ant comp onen ts for inference, thereb y reducing in terference of lo cal distortion and corruption. IV . B . Computational complexit y HYDRA in volv es tw o main stages: the lo w-rank based signature restoration stage and the net work based parameter restoration stage. Even though the time cost for parameter restora- tion is longer than previous methods 13 , 14 , the time cost in the this stage is only a small fraction of the total time consumption, as the computational complexit y is dominated b y page 22 P . Song the signature restoration stage. In other words, the computational burden of HYDRA lies in the SVD calculation in the first stage. Hence, fast SVD metho ds can b e employ ed to dramatically impro ve the efficiency of signature restoration. IV . C . Mo del storage requiremen ts Regarding the storage requirement (in double precision), HYDRA needs only 2.1 megab ytes to store the netw ork with 0.5 million parameters, while it requires 108 megab ytes to store a sim ulated dictionary of size 80100 × 200 and 551 megabytes for size 80100 × 1000 . Note that the dictionary volume will gro w exp onentially with the n umber of parameters, but the space required for storing a net work is not strictly limited b y the dictionary density once the top ology of the net w ork is fixed, thus significantly alleviating the storage burden inherent to the exp onen tial growth of multi-dimensional dictionaries. IV . D . Impact of pro viding con tin uous T1/T2 v alues Pro viding contin uous T1/T2 v alues is an adv an tage of neural net work based parameter mapping ov er dictionary matc hing. This prop erty may find promising applications in some practical scenarios, for exampling, monitoring sensitive c hanging of pathology condition o ver time, suc h as m ultiple sclerosis 40 , 41 , stroke 42 , and treatment resp onses 43 , 44 , where the differences in T1 and T2 v alues betw een health y and diseased tissues or b etw een disease stages could b e v ery small 45 . On the other hand, to fulfil this p otential of net w ork based MRF tec hniques, prerequisites on the accuracy and precision of MRI measurements are needed. T aking T1/T2 quan tification as an example, ev en for the in v ersion recov ery spin ec ho (IR-SE) / multiple single-echo spin ec ho MRI sequences whic h are considered as the gold standard for T1/T2 quantification, there exist v ariations of 2% - 9% on the measured relaxation times 45 . Suc h anatomical measurement uncertainties and model imp erfections ma y weak en the adv an tage and clinical impact of providing contin uous T1/T2 v alues using net work based MRF techniques to some exten t. Therefore, impro ving the accuracy of gold standard approaches in the future w ould con tribute to making the most of the p otential of neural net works in the MRF domain. IV.C. Mo del sto rage requirements Hyb rid Deep MRF page 23 V . Conclusion W e prop osed a h ybrid deep MRF approac h which combines lo w-rank based signature restora- tion with learning-based parameter restoration. In our approac h, a low-rank based de- aliasing metho d is used to restore clean signatures from subsampled k-space measuremen ts. Then, a 1D deep nonlo cal residual CNN is dev elop ed for efficient signature-to-parameter mapping, replacing the time-consuming dictionary matc hing op eration in conv en tional MRF tec hniques. Our approac h demonstrates adv an tages in terms of inference sp eed, accuracy and storage requirements o ver comp eting MRF metho ds as no dictionary is needed for recov ery . References 1 D. Ma, V. Gulani, N. Seib erlich, K. Liu, J. L. Sunshine, J. L. Duerk, and M. A. Grisw old, Magnetic resonance fingerprin ting, Nature 495 , 187 (2013). 2 Y. Jiang, D. Ma, N. Seib erlich, V. Gulani, and M. A. Griswold, MR fingerprinting using fast imaging with steady state precession (FISP) with spiral readout, Magnetic resonance in medicine 74 , 1621–1631 (2015). 3 M. Davies, G. Puy , P . V andergheynst, and Y. Wiaux, A compressed sensing framew ork for magnetic resonance fingerprin ting, SIAM Journal on Imaging Sciences 7 , 2623–2656 (2014). 4 Z. W ang, H. Li, Q. Zhang, J. Y uan, and X. W ang, Magnetic Resonance Fingerprinting with compressed sensing and distance metric learning, Neuro computing 174 , 560–570 (2016). 5 G. Mazor, L. W eizman, A. T al, and Y. C. Eldar, Low rank magnetic resonance fin- gerprin ting, in Engine ering in Me dicine and Biolo gy So ciety (EMBC), 2016 IEEE 38th A nnual International Confer enc e of the , pages 439–442, IEEE, 2016. 6 G. Mazor, L. W eizman, A. T al, and Y. C. Eldar, Low-rank magnetic resonance finger- prin ting, Medical physics 45 , 4066–4084 (2018). 7 C. Liao, B. Bilgic, M. K. Manhard, B. Zhao, X. Cao, J. Zhong, L. L. W ald, and K. Set- page 24 P . Song somp op, 3D MR fingerprin ting with accelerated stac k-of-spirals and h ybrid sliding- windo w and GRAPP A reconstruction, Neuroimage 162 , 13–22 (2017). 8 X. Cao, C. Liao, Z. W ang, Y. Chen, H. Y e, H. He, and J. Zhong, Robust sliding-window reconstruction for A ccelerating the acquisition of MR fingerprinting, Magnetic resonance in medicine 78 , 1579–1588 (2017). 9 M. Lustig, D. L. Donoho, J. M. Santos, and J. M. Pauly , Compressed sensing MRI, IEEE signal pro cessing magazine 25 , 72–82 (2008). 10 L. W eizman, Y. C. Eldar, and D. Ben Bashat, Compressed sensing for longitudinal MRI: An adaptiv e-weigh ted approach, Medical physics 42 , 5195–5208 (2015). 11 L. W eizman, Y. C. Eldar, and D. Ben Bashat, Reference-based MRI, Medical ph ysics 43 , 5357–5369 (2016). 12 Y. C. Eldar, Sampling The ory: Beyond Band limite d Systems , Cambridge Univ ersit y Press, 2015. 13 O. Cohen, B. Zh u, and M. S. Rosen, MR fingerprinting deep reconstruction net work (DR ONE), Magnetic resonance in medicine 80 , 885–894 (2018). 14 E. Hopp e, G. Körzdörfer, T. Würfl, J. W etzl, F. Lugauer, J. Pfeuffer, and A. Maier, Deep Learning for Magnetic Resonance Fingerprinting: A New Approac h for Predicting Quan titative P arameter V alues from Time Series, Stud Health T echnol Inform 243 , 202–206 (2017). 15 Y. LeCun, Y. Bengio, and G. Hin ton, Deep learning, N ature 521 , 436 (2015). 16 I. Goo dfellow, Y. Bengio, A. Courville, and Y. Bengio, De ep le arning , v olume 1, MIT press Cam bridge, 2016. 17 A. Krizhevsky , I. Sutskev er, and G. E. Hin ton, Imagenet classification with deep conv o- lutional neural netw orks, in A dvanc es in neur al information pr o c essing systems , pages 1097–1105, 2012. 18 K. He, X. Zhang, S. Ren, and J. Sun, Deep residual learning for image recognition, in Pr o c. IEEE Conf. Comput. Vision Pattern R e c o g , pages 770–778, 2016. REFERENCES Hyb rid Deep MRF page 25 19 K. He, X. Zhang, S. Ren, and J. Sun, Iden tit y mappings in deep residual net w orks, in Pr o c. Eur. Conf. Comput. Vision , pages 630–645, Springer, 2016. 20 C. Dong, C. C. Lo y , K. He, and X. T ang, Image sup er-resolution using deep con volutional net works, IEEE T rans. Pattern Anal. Mach. Intell. 38 , 295–307 (2016). 21 J. Kim, J. Kw on Lee, and K. Mu Lee, Deeply-recursiv e conv olutional netw ork for image sup er-resolution, in Pr o c. IEEE Conf. Comput. Vision Pattern R e c o g , pages 1637–1645, 2016. 22 J. Gehring, M. Auli, D. Grangier, D. Y arats, and Y. N. Dauphin, Conv olutional sequence to sequence learning, arXiv preprint arXiv:1705.03122 (2017). 23 G. Hinton et al., Deep neural netw orks for acoustic mo deling in sp eec h recognition: The shared views of four researc h groups, IEEE Signal pro cessing magazine 29 , 82–97 (2012). 24 J.-F. Cai, E. J. Candès, and Z. Shen, A singular v alue thresholding algorithm for matrix completion, SIAM Journal on Optimization 20 , 1956–1982 (2010). 25 S. Sra, S. No wozin, and S. J. W righ t, Optimization for machine le arning , Mit Press, 2012. 26 A. V aswani, N. Shazeer, N. Parmar, J. Uszk oreit, L. Jones, A. N. Gomez, Ł. Kaiser, and I. P olosukhin, Atten tion is all y ou need, in A dvanc es in neur al information pr o c essing systems , pages 5998–6008, 2017. 27 X. W ang, R. Girshick, A. Gupta, and K. He, Non-lo cal neural netw orks, in Pr o c e e dings of the IEEE Confer enc e on Computer Vision and Pattern R e c o gnition , pages 7794–7803, 2018. 28 K. He, X. Zhang, S. Ren, and J. Sun, Delving deep in to rectifiers: Surpassing h uman- lev el p erformance on imagenet classification, in Pr o c. IEEE Int. Conf. Comput. Vision , pages 1026–1034, 2015. 29 N. Sriv astav a, G. Hin ton, A. Krizhevsky , I. Sutskev er, and R. Salakh utdinov, Drop out: a simple w ay to prev ent neural net works from o verfitting, The Journal of Mac hine Learning Researc h 15 , 1929–1958 (2014). REFERENCES page 26 P . Song 30 J. Hennig, Ec ho es – ho w to generate, recognize, use or a void them in MR-imaging se- quences. P art I: F undamen tal and not so fundamen tal prop erties of spin ec ho es, Concepts in Magnetic Resonance 3 , 125–143 (1991). 31 M. W eigel, Extended phase graphs: dephasing, RF pulses, and echoes-pure and simple, Journal of Magnetic Resonance Imaging 41 , 266–295 (2015). 32 D. P . Kingma and J. Ba, Adam: A metho d for sto chastic optimization, arXiv preprint arXiv:1412.6980 (2014). 33 J. A. F essler and B. P . Sutton, Nonuniform fast F ourier transforms using min-max in terp olation, IEEE T rans. Sig. Pro c. 51 , 560–574 (2003). 34 J. V ymazal, A. Righini, R. A. Brooks, M. Canesi, C. Mariani, M. Leonardi, and G. P ez- zoli, T1 and T2 in the brain of health y sub jects, patien ts with P arkinson disease, and patien ts with multiple system atroph y: relation to iron conten t, Radiology 211 , 489–495 (1999). 35 G. Liberman, Y. Louzoun, and D. Ben Bashat, T1 mapping using v ariable flip angle SPGR data with flip angle correction, Journal of Magnetic Resonance Imaging 40 , 171–180 (2014). 36 S. C. Deoni, T. M. Peters, and B. K. Rutt, High-resolution T1 and T2 mapping of the brain in a clinically acceptable time with DESPOT1 and DESPOT2, Magnetic Resonance in Medicine: An Official Journal of the In ternational So ciet y for Magnetic Resonance in Medicine 53 , 237–241 (2005). 37 J. H. Lee, B. A. Hargrea ves, B. S. Hu, and D. G. Nishim ura, F ast 3D imaging us- ing v ariable-density spiral tra jectories with applications to limb p erfusion, Magnetic Resonance in Medicine: An Official Journal of the In ternational So ciet y for Magnetic Resonance in Medicine 50 , 1276–1285 (2003). 38 A. Kiperwas, D. Rosenfeld, and Y. C. Eldar, The SPURS algorithm for resampling an irregularly sampled signal on to a cartesian grid, IEEE transactions on medical imaging 36 , 628–640 (2017). REFERENCES Hyb rid Deep MRF page 27 39 D. Rosenfeld, New approach to gridding using regularization and estimation theory , Magnetic Resonance in Medicine: An Official Journal of the In ternational So ciety for Magnetic Resonance in Medicine 48 , 193–202 (2002). 40 F. Manfredonia, O. Ciccarelli, Z. Khaleeli, D. J. T ozer, J. Sastre-Garriga, D. H. Miller, and A. J. Thompson, Normal-app earing brain t1 relaxation time predicts disabilit y in early primary progressiv e multiple sclerosis, Archiv es of neurology 64 , 411–415 (2007). 41 K. P apadop oulos, D. J. T ozer, L. Fisniku, D. R. Altmann, G. Davies, W. Rashid, A. J. Thompson, D. H. Miller, and D. T. Chard, TI-relaxation time c hanges ov er fiv e years in relapsing-remitting multiple sclerosis, Multiple Sclerosis Journal 16 , 427–433 (2010). 42 J. Bernarding, J. Braun, J. Hohmann, U. Mansmann, M. Ho ehn-Berlage, C. Stapf, K.- J. W olf, and T. T olxdorff, Histogram-based characterization of health y and isc hemic brain tissues using m ultiparametric MR imaging including apparen t diffusion co efficient maps and relaxometry , Magnetic Resonance in Medicine: An Official Journal of the In ternational So ciety for Magnetic Resonance in Medicine 43 , 52–61 (2000). 43 P . M. McSheeh y , C. W eidensteiner, C. Cannet, S. F erretti, D. Laurent, S. Ruetz, M. Stumm, and P . R. Allegrini, Quantified tumor T1 is a generic early-resp onse imag- ing biomark er for chemotherap y reflecting cell viabilit y , Clinical Cancer Researc h 16 , 212–225 (2010). 44 C. W eidensteiner, P . R. Allegrini, M. Stic k er-Jantsc heff, V. Romanet, S. F erretti, and P . M. McSheeh y , T umour T 1 c hanges in viv o are highly predictiv e of response to c hemotherapy and reflect the num b er of viable tumour cells–a preclinical MR study in mice, BMC cancer 14 , 88 (2014). 45 Y. Jiang, D. Ma, K. E. Keenan, K. F. Stupic, V. Gulani, and M. A. Griswold, Re- p eatabilit y of magnetic resonance fingerprinting T1 and T2 estimates assessed using the ISMRM/NIST MRI system phan tom, Magnetic resonance in medicine 78 , 1452–1457 (2017). REFERENCES page 28 P . Song Figure 1: Relationship b et w een k ey v ariables. The MR con trast matrix X is asso ciated with the k-space measuremen ts Y p er column b y the subsampled F ourier transform. It is related to tissue parameters Θ T 12 p er row b y the Blo c h equations. Giv en Θ T RE and Y , the image stac k X is commonly first reconstructed from Y , referred to as signature restoration, and then mapp ed to tissue parameters Θ T 12 via dictionary matching, referred to as parameter restoration. Figure 2: Parameter restoration using dictionary matc hing. Giv en an inquiry temporal signature, dictionary matc hing computes its inner pro duct with eac h dictionary en try , and selects the most correlated one with the highest inner pro duct as the b est matc hing signature. Once the b est matching entry is found, it directly leads to multiple tissue parameters, such as T1, T2, sim ultaneously , via searching a lo okup-table. REFERENCES Hyb rid Deep MRF page 29 1 6 f i l t e r s T 1 m a p F u l l y - c o n n e c t e d l a y e r P a r a m e t e r r e s t o r a t i o n u s i n g 1 D n o n l o c a l r e s i d u a l C N N … T r a i n i n g s i g n a t u r e s i . e . d i c t i o n a r y e n t r i e s S i g n a l e v o l u t i o n i n a n i m a g e s t a c k … … … k e r n e l s i z e = 2 1 a v e r a g e p o o l i n g … T r a i n i n g T e s t i n g … k - s p a c e s a m p l i n g S i g n a t u r e r e s t o r a t i o n u s i n g l o w - r a n k d e - a l i a s i n g L o o k u p - t a b l e D i c t i o n a r y R e s i d u a l b l o c k R e s i d u a l b l o c k N o n l o c a l b l o c k N o n l o c a l b l o c k 1 6 f i l t e r s 3 2 f i l t e r s 3 2 f i l t e r s 1 2 8 f i l t e r s 1 2 8 f i l t e r s … I n p u t m a x p o o l i n g s h o r t c u t R e s i d u a l b l o c k … B l o c h … … … … … T 2 m a p s o f t m a x L e a r n e d A f f i n i t y M a t r i x f ( x i , x j ) N o n l o c a l o p e r a t i o n b l o c k 1 x 1 C o n v o p e r a t i o n g ( x ) ϕ ( x ) θ ( x ) s h o r t c u t s h o r t c u t Figure 3: Diagram of the prop osed MRF reconstruction approac h. During the training stage, syn thesized dictionary entries are used as training signatures to train the designed 1D nonlo cal residual CNN un til the outputs approximate parameter v alues in LUT well. In this w ay , the net work captures the signature-to-parameter mapping. During the testing stage, a lo w-rank based algorithm is used to restore the image stack, a matrix containing signatures in ro ws, from k-space measurements. Then the restored signatures are input in to the trained net work to obtain corresp onding tissue parameters directly . T able 1: Brief description of exp eriment types and settings. Exp erimen t Settings T raining Input: D , size K × L = 80100 × 200 . Groundtruth: LUT , size 80100 × 2 . k-space subsampling factor β : not av ailable. T esting on syn thetic data Input: X , size N × L = 80000 × 200 . Groundtruth: Θ T 12 , size 80000 × 2 . k-space subsampling factor β : not av ailable. T esting on anatomical data Input: Y , size Q × L = 16384 β × 200 or 16384 β × 1000 . Reference: Θ T 12 , size N × 2 = 16384 × 2 . k-space subsampling factor β : 70%, 15% using Gaussian patterns. T esting on anatomical data Input: Y , size Q × L = 16384 β × 1000 . Reference: Θ T 12 , size N × 2 = 16384 × 2 . k-space subsampling factor β : 9% using spiral tra jectories. REFERENCES page 30 P . Song (a) (b) Figure 4: Synthetic MRF temp oral signatures with 200 time frames. (a) T emp oral signatures corresponding to parameter v alues {(T1, T2)} ms = {(800,40),(800,60),(800,80),(800,100)} ms. (2) T emp oral signatures corresp onding to parameter v alues {(T1, T2)} ms = {(400,80),(600,80),(800,80),(1000,80)} ms. Figure 5: FISP pulse sequence parameters. All the flip angles (F A) constituted a sinusoidal v ariation in the range of 0 - 70 degrees to ensure smo othly v arying transient state of the magnetization. The rep etition time (TR) w as randomly v aried in the range of 11.5 - 14.5 ms with a P erlin noise pattern. T able 2: T esting on synthetic dataset. Comparing parameter restoration p erformance, in terms of PSNR, SNR, RMSE and correlation co efficien t. Dict. Match. CNN 14 FNN 13 Prop osed T1 / T 2 T1 / T2 T1 / T2 T1 / T2 PSNR (dB) 59.15 / 52.31 62.96 / 49.64 58.97 / 54.96 79.30 / 72.99 SNR (dB) 55.23 / 47.15 59.05 / 44.49 55.06 / 49.81 75.38 / 67.83 RMSE (ms) 5.515 / 4.847 3.554 / 6.591 5.63 / 3.57 0.542 / 0.448 CorrCo ef 1.00 / 1.00 1.00 / 1.00 1.00 / 1.00 1.00 / 1.00 time cost (s) 464.10 2.87 1.58 8.2 REFERENCES Hyb rid Deep MRF page 31 T able 3: T esting on syn thetic dataset in volving detailed T1 / T2 examples that are not on the training grid and their interv als are muc h smaller than the training grid in terv als. D.M. denotes dictionary matc hing. T1 and T2 errors are defined as the difference betw een estimated v alues and groundtruth v alues. T1 Estimation T1 Errors T ruth D.M. 14 13 Ours D.M. 14 13 Ours 1005.0 1001.0 1002.9 1009.3 1004.8 -4.0 -2.1 4.3 -0.2 1005.5 1001.0 1003.3 1010.0 1005.3 -4.5 -2.3 4.5 -0.2 1006.0 1011.0 1003.6 1010.6 1005.8 5.0 -2.4 4.6 -0.2 1006.5 1011.0 1004.1 1011.2 1006.3 4.5 -2.5 4.7 -0.2 1007.0 1011.0 1004.5 1011.8 1006.8 4.0 -2.5 4.8 -0.3 RMSE - - - - 4.4 2.3 4.6 0.2 T2 Estimation T2 Errors T ruth D.M. 14 13 Ours D.M. 14 13 Ours 505.0 501.0 513.6 504.3 505.2 -4.0 8.6 -0.7 0.2 505.5 511.0 514.1 504.8 505.7 5.5 8.6 -0.7 0.2 506.0 501.0 514.7 505.3 506.2 -5.0 8.6 -0.7 0.2 506.5 511.0 515.2 505.9 506.8 4.5 8.7 -0.6 0.3 507.0 511.0 515.7 506.4 507.3 4.0 8.7 -0.6 0.3 RMSE - - - - 4.6 8.6 0.7 0.2 Dictionary Matc hing CNN 14 FNN 13 HYDRA Figure 6: T esting on syn thetic dataset in volving detailed T1 / T2 examples that are not on the training grid and their interv als are muc h smaller than the training grid in terv als. Dictionary matc hing finds best adjacen t v alues from the dictionary , i.e. 1001, 1011 for T1, and 501, 511 for T2. In contrast, o wing to the captured mapping functions, neural net works output con tinuous v alues. Proposed HYDRA leads to the smallest deviations and bias. T able 4: T esting on anatomical dataset with full k-space sampling. Comparing parameter restoration p erformance, in terms of PSNR, SNR, RMSE and correlation co efficien t. Dict. Match. CNN 14 FNN 13 Prop osed basic Proposed nonlo cal T1 / T 2 T1 / T2 T1 / T2 T1 / T2 T1 / T2 PSNR (dB) 56.64 / 52.04 54.06 / 49.88 54.53 / 54.36 56.59 / 60.01 56.47 / 62.56 SNR (dB) 42.20 / 27.81 39.63 / 25.66 40.09 / 30.07 42.15 / 35.76 42.03 / 38.32 RMSE (ms) 6.623 / 6.252 8.912 / 8.015 8.45 / 4.78 6.661 / 2.498 6.76 / 1.86 CorrCo ef 1.00 / 1.00 1.00 / 1.00 1.00 / 1.00 1.00 / 1.00 1.00 / 1.00 time cost (s) 84.56 0.69 0.41 1.6 2.1 REFERENCES page 32 P . Song (a) T1, T2 estimations using dictionary matching. (b) T1, T2 estimations using CNN 14 . (c) T1, T2 estimations using FNN 13 . (d) T1, T2 estimations using HYDRA. Figure 7: T esting on the synthetic dataset for comparing parameter restoration performance. Subfig. (a) - (d) sho w the results using dictionary matc hing 1 , 2 , 3 , 4 , 5 , 6 , FNN 13 , CNN 14 and HYDRA. In each subfigure, the left figure compares the estimated T1 or T2 v alues (marked with red dot) with groundtruth v alues (marked with blue line), and the righ t figure shows the deviations of the estimation from the groundtruth. Parameter mapping performance of HYDRA is m uch b etter than comp eting methods, in the entire v alue range of T1 and T2 parameters, resulting in smaller deviations. T able 5: T esting on anatomical dataset with k-space subsampling ratio 70% and 15% using Gaussian patterns and 200 time frames. k-space subsampling factor β = 70% Ma et al. 1 BLIP 3 FLOR 6 CNN 14 FNN 13 Proposed basic nonlocal PSNR (dB) 23.69 / 38.17 45.67 / 47.84 50.11 / 50.85 49.71 / 45.48 50.15 / 51.08 50.79 / 51.59 49.87 / 57.57 SNR (dB) 8.73 / 13.84 31.28 / 23.49 35.67 / 26.48 35.26 / 21.19 35.70 / 26.67 36.34 / 27.19 35.42 / 33.30 RMSE (ms) 294.32 / 30.87 23.42 / 10.14 14.01 / 7.17 14.71 / 13.31 13.99 / 6.98 12.99 / 6.57 14.44 / 3.31 time cost (s) 72.88 75.70 85.35 23.72 23.53 24.85 26.3 k-space subsampling factor β = 15% Ma et al. 1 BLIP 3 FLOR 6 CNN 14 FNN 13 Proposed basic nonlocal PSNR (dB) 27.94 / 32.84 35.45 / 39.25 44.95 / 46.11 43.74 / 35.98 45.03 / 45.90 45.23 / 44.44 45.39 / 51.32 SNR (dB) 13.50 / 8.61 20.99 / 14.58 30.51 / 21.89 29.23 / 12.26 30.58 / 21.32 30.76 / 19.78 30.91 / 26.99 RMSE (ms) 180.3 / 57.03 76.01 / 27.25 25.46 / 12.37 29.27 / 39.73 25.21 / 12.68 24.65 / 15.00 24.20 / 6.79 time cost (s) 106 112.8 121.7 24.54 24.36 25.67 27.31 REFERENCES Hyb rid Deep MRF page 33 (a) T1, T2 estimations using dictionary matching. (b) T1, T2 estimations using CNN 14 . (c) T1, T2 estimations using FNN 13 . (d) T1, T2 estimations using HYDRA. Figure 8: T esting on the anatomical dataset with full k-space sampling for comparing parameter restoration p erformance. Subfig. (a) - (d) sho w the results using dictionary matc hing 1 , 2 , 3 , 4 , 5 , 6 , FNN 13 , CNN 14 and HYDRA. Eac h subfigure s ho ws the deviations of the estimation from the reference. Parameter mapping p erformance using HYDRA outp erforms competing methods significan tly , resulting in smaller deviations. The p erformance is also v erified b y quantitativ e metrics, as shown in T able 4 . T able 6: T esting on anatomical dataset with k-space subsampling ratio 15% using Gaussian patterns and 1000 time frames. Ma et al. 1 BLIP 3 FLOR 6 CNN 14 FNN 13 Prop osed PSNR (dB) 27.53 / 33.28 35.50 / 39.10 50.90 / 50.04 41.96 / 39.21 52.62 / 49.86 52.32 / 52.79 SNR (dB) 13.09 / 9.05 21.06 / 14.87 36.44 / 25.65 27.44 / 15.05 38.17 / 25.43 37.86 / 28.35 RMSE (ms) 189.09 / 54.21 75.53 / 27.74 12.83 / 7.87 35.91 / 27.37 10.52 / 8.04 10.89 / 5.74 T able 7: T esting on anatomical dataset with k-space subsampling ratio 9% using spiral tra jectories and 1000 time frames. Ma et al. 1 BLIP 3 FLOR 6 CNN 14 FNN 13 Proposed PSNR (dB) 26.66 / 30.44 29.35 / 39.47 39.32 / 44.60 35.68 / 27.74 40.26 / 44.70 41.45 / 45.41 SNR (dB) 12.22 / 6.21 15.03 / 15.22 24.88 / 20.38 21.45 / 4.48 25.84 / 20.30 27.02 / 21.04 RMSE (ms) 209.01 / 75.18 153.37 / 26.57 48.67 / 14.72 73.96 / 102.57 43.65 / 14.55 38.08 / 13.41 REFERENCES page 34 P . Song T1/T2 Reference Dictionary Matc hing SNR = 42.20/27.81 dB CNN 14 SNR = 39.63/25.66 dB FNN 13 SNR = 40.09/30.07dB HYDRA SNR = 42.03/38.32 dB Figure 9: Visual results of testing on anatomical dataset with full k-space sampling for comparing parameter restoration p erformance. T op tw o rows corresp ond to T1 maps and residual errors while b ottom t wo ro ws corresp ond to T2 maps and residual errors. Prop osed HYDRA results in comparable p erformance for T1 mapping and yields m uch better p erformance for T2 mapping, obtaining 10dB higher SNR gains than comp eting dictionary-matching based metho ds 1 , 2 , 3 , 4 , 5 , 6 . HYDRA also outperforms previous netw orks, suc h as CNN b y Hopp e et al. 14 and FNN b y Cohen et al. 13 . Figure 10: A series of Gaussian patterns used for k-space subsampling. REFERENCES Hyb rid Deep MRF page 35 (a) T1, T2 estimations using Ma et al. 1 . (b) T1, T2 estimations using BLIP 3 (c) T1, T2 estimations using FLOR 6 (d) T1, T2 estimations using CNN 14 . (e) T1, T2 estimations using FNN 13 . (f ) T1, T2 estimations using HYDRA. Figure 11: T esting on the anatomical dataset with k-space subsampling factor 15% using Gaussian patterns and 1000 time frames. Subfig. (a) - (f ) sho w the results using Ma et al. 1 , BLIP 3 , FLOR 6 , CNN b y Hopp e et al. 14 , FNN b y Cohen et al. 13 and HYDRA. REFERENCES page 36 P . Song (a) T1 e stimation (top row) and residual errors (bottom ro w). (b) T2 estimation (top row) and residual errors (bottom ro w). BLIP 3 SNR = 21.06/14.87 dB FLOR 6 SNR = 36.44/25.65 dB CNN 14 SNR = 27.44/15.05 dB FNN 13 SNR = 38.17/25.43 dB HYDRA SNR = 37.86/28.35 dB Figure 12: Visual results of testing on anatomical dataset with k-space subsampling factor 15% using Gaussian pattern with L = 1000 . Comparison b et ween BLIP 3 , FLOR 6 , CNN b y Hopp e et al. 14 , FNN b y Cohen et al. 13 and HYDRA. Figure 13: A series of spiral tra jectories for k-space subsampling. REFERENCES Hyb rid Deep MRF page 37 (a) T1, T2 estimations using Ma et al. 1 . (b) T1, T2 estimations using BLIP 3 (c) T1, T2 estimations using FLOR 6 (d) T1, T2 estimations using CNN 14 . (e) T1, T2 estimations using FNN 13 . (f ) T1, T2 estimations using HYDRA. Figure 14: T esting on the anatomical dataset with k-space subsampling factor 9% using spiral tra jectories and 1000 time frames. Subfig. (a) - (f ) sho w the results using Ma et al. 1 , BLIP 3 , FLOR 6 , CNN b y Hopp e et al. 14 , FNN b y Cohen et al. 13 and HYDRA. REFERENCES page 38 P . Song (a) T1 Reference (left) and T2 Reference (right). (b) T1 estimation (top row) and residual errors (bottom ro w). (c) T2 estimation (top row) and residual errors (bottom ro w). BLIP 3 SNR = 15.03/15.22 dB FLOR 6 SNR = 24.88/20.38 dB CNN 14 SNR = 21.45/4.48 dB FNN 13 SNR = 25.84/20.30 dB HYDRA SNR = 27.02/21.04 dB Figure 15: Visual results of testing on anatomical dataset with k-space subsampling factor 9% using spiral tra jectories with L = 1000 . Comparison b et ween BLIP 3 , FLOR 6 , CNN b y Hopp e et al. 14 , FNN b y Cohen et al. 13 and HYDRA. REFERENCES
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment