Learning Dictionaries for Named Entity Recognition using Minimal Supervision
This paper describes an approach for automatic construction of dictionaries for Named Entity Recognition (NER) using large amounts of unlabeled data and a few seed examples. We use Canonical Correlation Analysis (CCA) to obtain lower dimensional embe…
Authors: Arvind Neelakantan, Michael Collins
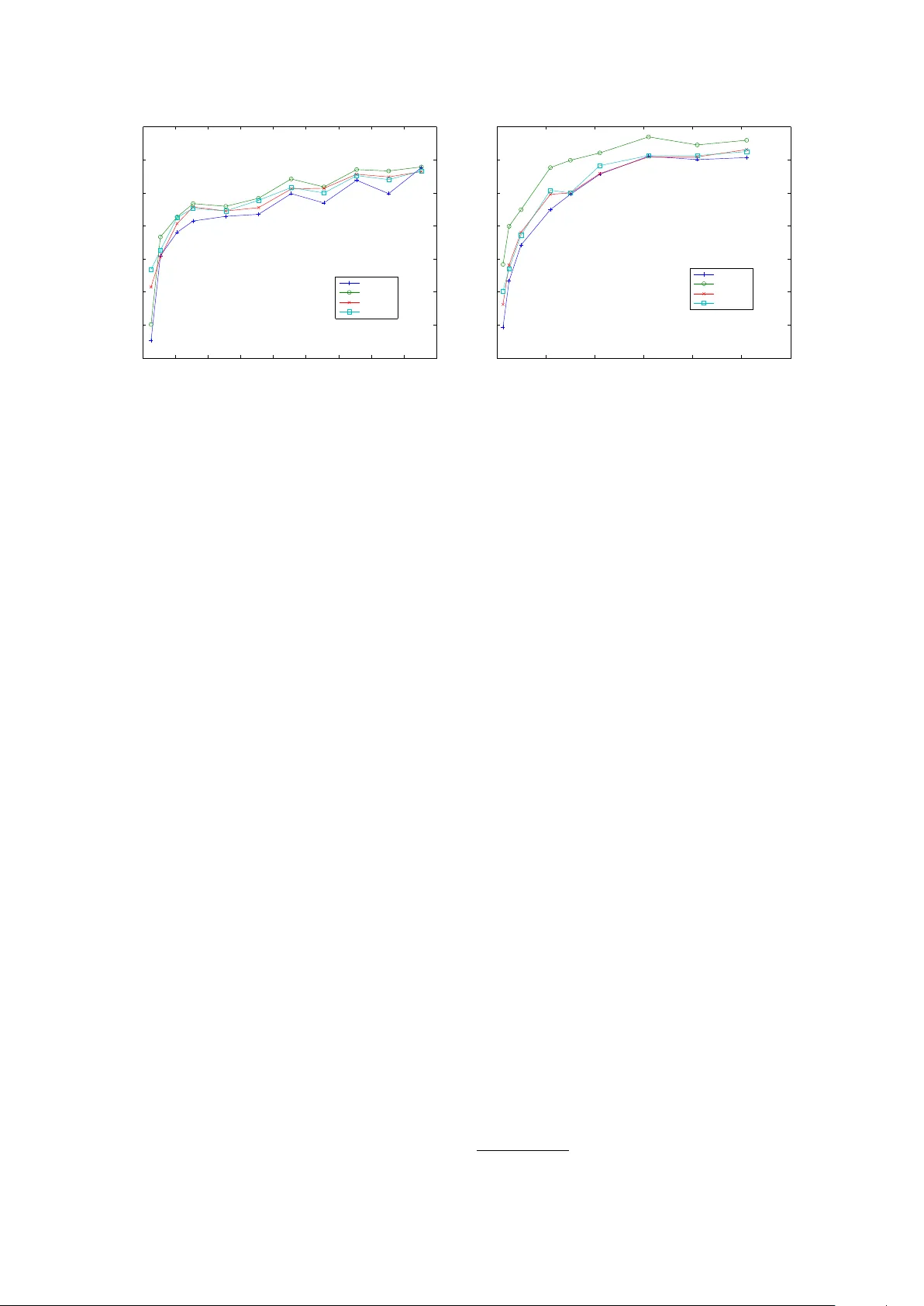
Learning Dictionaries f or Named Entity Recognition using Minimal Supervision Arvind Neelakantan Department of Computer Science Uni versity of Massachusetts, Amherst Amherst, MA, 01003 arvind@cs.umass.edu Michael Collins Department of Computer Science Columbia Uni versity Ne w-Y ork, NY 10027, USA mcollins@cs.columbia.edu Abstract This paper describes an approach for au- tomatic construction of dictionaries for Named Entity Recognition (NER) using large amounts of unlabeled data and a few seed examples. W e use Canonical Cor- relation Analysis (CCA) to obtain lower dimensional embeddings (representations) for candidate phrases and classify these phrases using a small number of labeled examples. Our method achiev es 16.5% and 11.3% F-1 score improvement over co-training on disease and virus NER re- specti vely . W e also show that by adding candidate phrase embeddings as features in a sequence tagger gi ves better perfor- mance compared to using word embed- dings. 1 Introduction Se veral works (e.g., Ratinov and Roth, 2009; Co- hen and Sarawagi, 2004) hav e sho wn that inject- ing dictionary matches as features in a sequence tagger results in significant gains in NER perfor- mance. Ho wev er , building these dictionaries re- quires a huge amount of human effort and it is of- ten difficult to get good co verage for many named entity types. The problem is more se vere when we consider named entity types such as gene, virus and disease, because of the large (and gro wing) number of names in use, the fact that the names are heavily abbre viated and multiple names are used to refer to the same entity (Leaman et al., 2010; Dogan and Lu, 2012). Also, these dictionaries can only be b uilt by domain experts, making the pro- cess very e xpensive. This paper describes an approach for automatic construction of dictionaries for NER using large amounts of unlabeled data and a small number of seed examples. Our approach consists of two steps. First, we collect a high recall, low preci- sion list of candidate phrases from the large unla- beled data collection for e very named entity type using simple rules. In the second step, we con- struct an accurate dictionary of named entities by removing the noisy candidates from the list ob- tained in the first step. This is done by learning a classifier using the lower dimensional, real-valued CCA (Hotelling, 1935) embeddings of the can- didate phrases as features and training it using a small number of labeled examples. The classifier we use is a binary SVM which predicts whether a candidate phrase is a named entity or not. W e compare our method to a widely used semi- supervised algorithm based on co-training (Blum and Mitchell, 1998). The dictionaries are first e valuated on virus (GENIA, 2003) and disease (Dogan and Lu, 2012) NER by using them directly in dictionary based taggers. W e also giv e results comparing the dictionaries produced by the two semi-supervised approaches with dictionaries that are compiled manually . The effecti veness of the dictionaries are also measured by injecting dictio- nary matches as features in a Conditional Random Field (CRF) based tagger . The results indicate that our approach with minimal supervision pro- duces dictionaries that are comparable to dictio- naries compiled manually . Finally , we also com- pare the quality of the candidate phrase embed- dings with w ord embeddings (Dhillon et al., 2011) by adding them as features in a CRF based se- quence tagger . 2 Background W e first giv e background on Canonical Correla- tion Analysis (CCA), and then give background on CRFs for the NER problem. 2.1 Canonical Correlation Analysis (CCA) The input to CCA consists of n paired observa- tions ( x 1 , z 1 ) , . . . , ( x n , z n ) where x i ∈ R d 1 , z i ∈ R d 2 ( ∀ i ∈ { 1 , 2 , . . . , n } ) are the feature represen- tations for the two views of a data point. CCA simultaneously learns projection matrices Φ 1 ∈ R d 1 × k , Φ 2 ∈ R d 2 × k ( k is a small number) which are used to obtain the lo wer dimensional represen- tations ( ¯ x 1 , ¯ z 1 ) , . . . , ( ¯ x n , ¯ z n ) where ¯ x i = Φ T 1 x i ∈ R k , ¯ z i = Φ T 2 z i ∈ R k , ∀ i ∈ { 1 , 2 , . . . , n } . Φ 1 , Φ 2 are chosen to maximize the correlation between ¯ x i and ¯ z i , ∀ i ∈ { 1 , 2 , . . . , n } . Consider the setting where we ha ve a label for the data point along with it’ s two views and ei- ther view is suf ficient to make accurate predic- tions. Kakade and Foster (2007) and Sridharan and Kakade (2008) gi ve strong theoretical guaran- tees when the lower dimensional embeddings from CCA are used for predicting the label of the data point. This setting is similar to the one considered in co-training (Collins and Singer, 1999) b ut there is no assumption of independence between the two vie ws of the data point. Also, it is an exact al- gorithm unlik e the algorithm gi ven in Collins and Singer (1999). Since we are using lower dimen- sional embeddings of the data point for prediction, we can learn a predictor with fe wer labeled exam- ples. 2.2 CRFs f or Named Entity Recognition CRF based sequence taggers have been used for a number of NER tasks (e.g., McCallum and Li, 2003) and in particular for biomedical NER (e.g., McDonald and Pereira, 2005; Burr Settles, 2004) because the y allo w a great deal of fle xibility in the features which can be included. The input to a CRF tagger is a sentence ( w 1 , w 2 , . . . , w n ) where w i , ∀ i ∈ { 1 , 2 , . . . , n } are words in the sentence. The output is a sequence of tags y 1 , y 2 , . . . , y n where y i ∈ { B, I, O } , ∀ i ∈ { 1 , 2 , . . . , n } . B is the tag giv en to the first word in a named entity , I is the tag gi ven to all words except the first w ord in a named entity and O is the tag gi ven to all other words. W e used the standard NER baseline fea- tures (e.g., Dhillon et al., 2011; Ratinov and Roth, 2009) which include: • Current W ord w i and its lexical features which include whether the word is capital- ized and whether all the characters are cap- italized. Prefix and suffixes of the word w i were also added. • W ord tokens in window of size two around the current word which include w i − 2 , w i − 1 , w i +1 , w i +2 and also the capital- ization pattern in the windo w . • Pre vious two predictions y i − 1 and y i − 2 . The effecti veness of the dictionaries are ev aluated by adding dictionary matches as features along with the baseline features (Ratinov and Roth, 2009; Cohen and Sara wagi, 2004) in the CRF tag- ger . W e also compared the quality of the candi- date phrase embeddings with the word-le vel em- beddings by adding them as features (Dhillon et al., 2011) along with the baseline features in the CRF tagger . 3 Method This section describes the two steps in our ap- proach: obtaining candidate phrases and classify- ing them. 3.1 Obtaining Candidate Phrases W e used the full text of 110,369 biomedical pub- lications in the BioMed Central corpus 1 to get the high recall, lo w precision list of candidate phrases. The advantages of using this huge collection of publications are obvious: almost all (including rare) named entities related to the biomedical do- main will be mentioned and contains more re- cent dev elopments than a structured resource like W ikipedia. The challenge howe ver is that these publications are unstructured and hence it is a dif- ficult task to construct accurate dictionaries using them with minimal supervision. The list of virus candidate phrases were ob- tained by extracting phrases that occur between “the” and “virus” in the simple pattern “the ... virus” during a single pass over the unlabeled doc- ument collection. This noisy list had a lot of virus names such as influenza , human immunodeficiency and Epstein-Barr along with phrases that are not virus names, like mutant , same , ne w , and so on. A similar rule like “the ... disease” did not gi ve a good cov erage of disease names since it is not the common way of ho w diseases are mentioned in publications. So we took a different approach 1 The corpus can be downloaded at http://www .biomedcentral.com/about/datamining to obtain the noisy list of disease names. W e col- lected ev ery sentence in the unlabeled data col- lection that has the word “disease” in it and ex- tracted noun phrases 2 follo wing the patterns “dis- eases like .... ”, “diseases such as .... ” , “diseases in- cluding .... ” , “diagnosed with .... ”, “patients with .... ” and “suf fering from .... ”. 3.2 Classification of Candidate Phrases Having found the list of candidate phrases, we no w describe how noisy words are filtered out from them. W e gather ( spelling , context ) pairs for e very instance of a candidate phrase in the unla- beled data collection. spelling refers to the can- didate phrase itself while context includes three words each to the left and the right of the candidate phrase in the sentence. The spelling and the con- text of the candidate phrase provide a natural split into two views which multi-vie w algorithms like co-training and CCA can exploit. The only super- vision in our method is to provide a few spelling seed examples (10 in the case of virus, 18 in the case of disease), for e xample, human immunodefi- ciency is a virus and mutant is not a virus. 3.2.1 A pproach using CCA embeddings W e use CCA described in the previous section to obtain lower dimensional embeddings for the candidate phrases using the ( spelling , context ) vie ws. Unlike previous works such as Dhillon et al. (2011) and Dhillon et al. (2012), we use CCA to learn embeddings for candidate phrases instead of all words in the vocab ulary so that we don’t miss named entities which hav e two or more words. Let the number of ( spelling , context ) pairs be n (sum of total number of instances of ev ery can- didate phrase in the unlabeled data collection). First, we map the spelling and conte xt to high- dimensional feature vectors. F or the spelling view , we define a feature for e very candidate phrase and also a boolean feature which indicates whether the phrase is capitalized or not. For the conte xt view , we use features similar to Dhillon et al. (2011) where a feature for e very word in the context in conjunction with its position is defined. Each of the n ( spelling , context ) pairs are mapped to a pair of high-dimensional feature vectors to get n paired observ ations ( x 1 , z 1 ) , . . . , ( x n , z n ) with x i ∈ R d 1 , z i ∈ R d 2 , ∀ i ∈ { 1 , 2 , . . . , n } ( d 1 , d 2 are the feature space dimensions of the spelling 2 Noun phrases were obtained using http://www .umiacs.umd.edu/ ∼ hal/T agChunk/ and conte xt view respectiv ely). Using CCA 3 , we learn the projection matrices Φ 1 ∈ R d 1 × k , Φ 2 ∈ R d 2 × k ( k << d 1 and k << d 2 ) and obtain spelling view projections ¯ x i = Φ T 1 x i ∈ R k , ∀ i ∈ { 1 , 2 , . . . , n } . The k-dimensional spelling view projection of any instance of a candidate phrase is used as it’ s embedding 4 . The k-dimensional candidate phrase embed- dings are used as features to learn a binary SVM with the seed spelling examples given in figure 1 as training data. The binary SVM predicts whether a candidate phrase is a named entity or not. Since the v alue of k is small, a small number of labeled examples are sufficient to train an accurate clas- sifier . The learned SVM is used to filter out the noisy phrases from the list of candidate phrases obtained in the pre vious step. T o summarize, our approach for classifying candidate phrases has the follo wing steps: • Input: n ( spelling , conte xt ) pairs, spelling seed examples. • Each of the n ( spelling , context ) pairs are mapped to a pair of high-dimensional fea- ture vectors to get n paired observations ( x 1 , z 1 ) , . . . , ( x n , z n ) with x i ∈ R d 1 , z i ∈ R d 2 , ∀ i ∈ { 1 , 2 , . . . , n } . • Using CCA, we learn the projection matri- ces Φ 1 ∈ R d 1 × k , Φ 2 ∈ R d 2 × k and ob- tain spelling view projections ¯ x i = Φ T 1 x i ∈ R k , ∀ i ∈ { 1 , 2 , . . . , n } . • The embedding of a candidate phrase is gi ven by the k-dimensional spelling view projec- tion of any instance of the candidate phrase. • W e learn a binary SVM with the candi- date phrase embeddings as features and the spelling seed examples gi ven in figure 1 as training data. Using this SVM, we predict whether a candidate phrase is a named entity or not. 3.2.2 A pproach based on Co-training W e discuss here briefly the DL-CoT rain algorithm (Collins and Singer, 1999) which is based on co- training (Blum and Mitchell, 1998), to classify 3 Similar to Dhillon et al. (2012) we used the method giv en in Halko et al. (2011) to perform the SVD computation in CCA for practical considerations. 4 Note that a candidate phrase gets the same spelling vie w projection across it’ s different instances since the spelling features of a candidate phrase are identical across it’ s in- stances. • Virus seed sp el ling examples – Virus Names : human imm uno deficiency , hepatitis C, influenza, Epstein-Barr, hepatitis B – Non-virus Names : mutan t, same, wild t yp e, paren tal, recombinan t • Disease seed sp el ling examples – Disease Names : tumor, malaria, breast cancer, cancer, IDDM, DM, A-T, tumors, VHL – Non-disease Names : cells, patients, study , data, expression, breast, BRCA1, protein, m utant 1 Figure 1: Seed spelling examples candidate phrases. W e compare our approach us- ing CCA embeddings with this approach. Here, two decision list of rules are learned simultane- ously one using the spelling vie w and the other using the context view . The rules using the spelling view are of the form: full-string=human immunodeficiency → V irus, full-string=mutant → Not a virus and so on. In the context vie w , we used bigram 5 rules where we considered all pos- sible bigrams using the context . The rules are of two types: one which giv es a positi ve label, for example, full-string=human immunodeficienc y → V irus and the other which gi ves a negati ve label, for example, full-string=mutant → Not a virus. The DL-CoT rain algorithm is as follows: • Input: ( spelling , context ) pairs for ev ery in- stance of a candidate phrase in the corpus, m specifying the number of rules to be added in e very iteration, precision threshold , spelling seed examples. • Algorithm: 1. Initialize the spelling decision list using the spelling seed e xamples giv en in fig- ure 1 and set i = 1 . 2. Label the entire input collection using the learned decision list of spelling rules. 3. Add i × m ne w context rules of each type to the decision list of context rules using the current labeled data. The rules are added using the same criterion as giv en in Collins and Singer (1999), i.e., among the rules whose strength is greater than the precision threshold , the ones which are seen more often with the corresponding label in the input data collection are added. 5 W e tried using unigram rules but they were very weak predictors and the performance of the algorithm was poor when they were considered. 4. Label the entire input collection using the learned decision list of context rules. 5. Add i × m new spelling rules of each type to the decision list of spelling rules using the current labeled data. The rules are added using the same criterion as in step 3. Set i = i + 1 . If rules were added in the pre vious iteration, return to step 2. The algorithm is run until no new rules are left to be added. The spelling decision list along with its strength (Collins and Singer, 1999) is used to construct the dictionaries. The phrases present in the spelling rules which gi ve a positiv e label and whose strength is greater than the precision thresh- old, were added to the dictionary of named enti- ties. W e found the parameters m and difficult to tune and they could significantly af fect the per- formance of the algorithm. W e giv e more details regarding this in the e xperiments section. 4 Related W ork Pre viously , Collins and Singer (1999) introduced a multi-view , semi-supervised algorithm based on co-training (Blum and Mitchell, 1998) for collect- ing names of people, org anizations and locations. This algorithm makes a strong independence as- sumption about the data and employs many heuris- tics to greedily optimize an objectiv e function. This greedy approach also introduces ne w param- eters that are often dif ficult to tune. In other works such as T oral and Mu ˜ noz (2006) and Kazama and T orisawa (2007) external struc- tured resources like W ikipedia hav e been used to construct dictionaries. Even though these meth- ods are fairly successful they suffer from a num- ber of dra wbacks especially in the biomedical do- main. The main drawback of these approaches is that it is very difficult to accurately disambiguate ambiguous entities especially when the entities are abbre viations (Kazama and T orisaw a, 2007). F or example, DM is the abbreviation for the disease Diabetes Mellitus and the disambiguation page for DM in W ikipedia associates it to more than 50 cat- egories since DM can be expanded to Doctor of Management , Dichr oic mirr or , and so on, each of it belonging to a different category . Due to the rapid growth of W ikipedia, the number of enti- ties that hav e disambiguation pages is growing fast and it is increasingly difficult to retrie ve the article we want. Also, it is tough to understand these ap- proaches from a theoretical standpoint. Dhillon et al. (2011) used CCA to learn word embeddings and added them as features in a se- quence tagger . They show that CCA learns bet- ter word embeddings than CW embeddings (Col- lobert and W eston , 2008), Hierarchical log-linear (HLBL) embeddings (Mnih and Hinton, 2007) and embeddings learned from man y other tech- niques for NER and chunking. Unlike PCA, a widely used dimensionality reduction technique, CCA is in v ariant to linear transformations of the data. Our approach is motiv ated by the theoreti- cal result in Kakade and Foster (2007) which is de veloped in the co-training setting. W e directly use the CCA embeddings to predict the label of a data point instead of using them as features in a sequence tagger . Also, we learn CCA embed- dings for candidate phrases instead of all words in the vocab ulary since named entities often contain more than one word. Dhillon et al. (2012) learn a multi-class SVM using the CCA word embed- dings to predict the POS tag of a word type. W e extend this technique to NER by learning a binary SVM using the CCA embeddings of a high recall, lo w precision list of candidate phrases to predict whether a candidate phrase is a named entity or not. 5 Experiments In this section, we gi ve experimental results on virus and disease NER. 5.1 Data The noisy lists of both virus and disease names were obtained from the BioMed Central corpus. This corpus w as also used to get the collection of ( spelling , context ) pairs which are the input to the CCA procedure and the DL-CoT rain algorithm de- scribed in the pre vious section. W e obtained CCA embeddings for the 100 , 000 most frequently oc- curring word types in this collection along with e very word type present in the training and de- velopment data of the virus and the disease NER dataset. These word embeddings are similar to the ones described in Dhillon et al. (2011) and Dhillon et al. (2012). W e used the virus annotations in the GE- NIA corpus (GENIA, 2003) for our experiments. The dataset contains 18,546 annotated sentences. W e randomly selected 8,546 sentences for train- ing and the remaining sentences were randomly split equally into dev elopment and testing sen- tences. The training sentences are used only for experiments with the sequence taggers. Previ- ously , Zhang et al. (2004) tested their HMM-based named entity recognizer on this data. For disease NER, we used the recent disease corpus (Dogan and Lu, 2012) and used the same training, devel- opment and test data split given by them. W e used a sentence segmenter 6 to get sentence segmented data and Stanford T okenizer 7 to tokenize the data. Similar to Dogan and Lu (2012), all the different disease categories were flattened into one single category of disease mentions. The dev elopment data was used to tune the h yperparameters and the methods were e valuated on the test data. 5.2 Results using a dictionary-based tagger First, we compare the dictionaries compiled us- ing different methods by using them directly in a dictionary-based tagger . This is a simple and informati ve way to understand the quality of the dictionaries before using them in a CRF-tagger . Since these taggers can be trained using a hand- ful of training examples, we can use them to build NER systems ev en when there are no labeled sen- tences to train. The input to a dictionary tagger is a list of named entities and a sentence. If there is an exact match between a phrase in the input list to the words in the gi ven sentence then it is tagged as a named entity . All other words are labeled as non-entities. W e ev aluated the performance of the follo wing methods for building dictionaries: • Candidate List : This dictionary contains all the candidate phrases that were obtained us- ing the method described in Section 3.1. The noisy list of virus candidates and disease can- didates had 3,100 and 60,080 entries respec- ti vely . 6 https://pypi.python.or g/pypi/text-sentence/0.13 7 http://nlp.stanford.edu/software/tokenizer .shtml Method V irus NER Disease NER Precision Recall F-1 Score Precision Recall F-1 Score Candidate List 2.20 69.58 4.27 4.86 60.32 8.99 Manual 42.69 68.75 52.67 51.39 45.08 48.03 Co-T raining 48.33 66.46 55.96 58.87 23.17 33.26 CCA 57.24 68.33 62.30 38.34 44.55 41.21 T able 1: Precision, recall, F- 1 scores of dictionary-based taggers • Manual : Manually constructed dictionaries, which requires a lar ge amount of human ef- fort, are employed for the task. W e used the list of virus names gi ven in Wikipedia 8 . Un- fortunately , abbreviations of virus names are not present in this list and we could not find any other more complete list of virus names. Hence, we constructed abbreviations by con- catenating the first letters of all the strings in a virus name, for e very virus name gi ven in the W ikipedia list. For diseases, we used the list of disease names gi ven in the Unified Medical Lan- guage System (UMLS) Metathesaurus. This dictionary has been widely used in disease NER (e.g., Dogan and Lu, 2012; Leaman et al., 2010) 9 . • Co-T raining : The dictionaries are con- structed using the DL-CoT rain algorithm de- scribed pre viously . The parameters used were m = 5 and = 0 . 95 as gi ven in Collins and Singer (1999). The phrases present in the spelling rules which gi ve a positi ve label and whose strength is greater than the preci- sion threshold, were added to the dictionary of named entities. In our experiment to construct a dictionary of virus names, the algorithm stopped after just 12 iterations and hence the dictionary had only 390 virus names. This was because there were no spelling rules with strength greater than 0 . 95 to be added. W e tried varying both the parameters b ut in all cases, the algo- rithm did not progress after a few iterations. W e adopted a simple heuristic to increase the cov erage of virus names by using the strength of the spelling rules obtained after the 12 th it- eration. All spelling rules that giv e a positi ve 8 http://en.wikipedia.org/wiki/List of viruses 9 The list of disease names from UMLS can be found at https://sites.google.com/site/fmchowdhury2/bioene x . label and which has a strength greater than θ were added to the decision list of spelling rules. The phrases present in these rules are added to the dictionary . W e pick ed the θ pa- rameter from the set [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9] using the de velopment data. The co-training algorithm for constructing the dictionary of disease names ran for close to 50 iterations and hence we obtained bet- ter cov erage for disease names. W e still used the same heuristic of adding more named en- tities using the strength of the rule since it performed better . • CCA : Using the CCA embeddings of the candidate phrases 10 as features we learned a binary SVM 11 to predict whether a candidate phrase is a named entity or not. W e consid- ered using 10 to 30 dimensions of candidate phrase embeddings and the regularizer was picked from the set [0.0001, 0.001, 0.01, 0.1, 1, 10, 100]. Both the regularizer and the num- ber of dimensions to be used were tuned us- ing the de velopment data. T able 1 gi ves the results of the dictionary based taggers using the different methods described abov e. As e xpected, when the noisy list of candi- date phrases are used as dictionaries the recall of the system is quite high but the precision is very lo w . The lo w precision of the W ikipedia virus lists was due to the heuristic used to obtain ab- bre viations which produced a few noisy abbrevia- tions b ut this heuristic was crucial to get a high re- call. The list of disease names from UMLS gives a lo w recall because the list does not contain many disease abbre viations and composite disease men- tions such as br east and ovarian cancer . The pres- 10 The performance of the dictionaries learned from w ord embeddings was very poor and we do not report it’ s perfor - mance here. 11 we used LIBSVM (http://www .csie.ntu.edu.tw/ ∼ cjlin/libsvm/) in our SVM experiments 0 1000 2000 3000 4000 5000 6000 7000 8000 9000 0.5 0.55 0.6 0.65 0.7 0.75 0.8 0.85 Number of Training Sentences F−1 Score Virus NER baseline manual co−training cca 0 1000 2000 3000 4000 5000 6000 0.45 0.5 0.55 0.6 0.65 0.7 0.75 0.8 F−1 Score Number of Training Sentences Disease NER baseline manual co−training cca 1 Figure 2: V irus and Disease NER F-1 scores for v arying training data size when dictionaries obtained from dif ferent methods are injected ence of ambiguous abbreviations affected the ac- curacy of this dictionary . The virus dictionary constructed using the CCA embeddings was very accurate and the false pos- iti ves were mainly due to ambiguous phrases, for example, in the phrase HIV replication , HIV which usually refers to the name of a virus is tagged as a RN A molecule. The accuracy of the disease dictionary produced using CCA embed- dings was mainly af fected by noisy abbreviations. W e can see that the dictionaries obtained us- ing CCA embeddings perform better than the dic- tionaries obtained from co-training on both dis- ease and virus NER even after improving the co- training algorithm’ s coverage using the heuristic described in this section. It is important to note that the dictionaries constructed using the CCA embeddings and a small number of labeled exam- ples performs competitiv ely with dictionaries that are entirely b uilt by domain experts. These re- sults show that by using the CCA based approach we can build NER systems that give reasonable performance e ven for difficult named entity types with almost no supervision. 5.3 Results using a CRF tagger W e did two sets of experiments using a CRF tag- ger . In the first experiment, we add dictionary fea- tures to the CRF tagger while in the second ex- periment we add the embeddings as features to the CRF tagger . The same baseline model is used in both the e xperiments whose features are described in Section 2.2. For both the CRF 12 experiments the re gularizers from the set [0.0001, 0.001, 0.01, 0.1, 1.0, 10.0] were considered and it was tuned on the de velopment set. 5.3.1 Dictionary F eatures Here, we inject dictionary matches as features (e.g., Ratinov and Roth, 2009; Cohen and Saraw agi, 2004) in the CRF tagger . Gi ven a dic- tionary of named entities, e very word in the input sentence has a dictionary feature associated with it. When there is an exact match between a phrase in the dictionary with the words in the input sen- tence, the dictionary feature of the first word in the named entity is set to B and the dictionary fea- ture of the remaining words in the named entity is set to I . The dictionary feature of all the other words in the input sentence which are not part of any named entity in the dictionary is set to O . The ef fectiv eness of the dictionaries constructed from v arious methods are compared by adding dictio- nary match features to the CRF tagger . These dic- tionary match features were added along with the baseline features. Figure 2 indicates that the dictionary features in general are helpful to the CRF model. W e can see that the dictionaries produced from our approach using CCA are much more helpful than the dictio- naries produced from co-training especially when there are fe wer labeled sentences to train. Simi- lar to the dictionary tagger e xperiments discussed 12 W e used CRFsuite (www .chokkan.org/software/crfsuite/) for our experiments with CRFs. 0 1000 2000 3000 4000 5000 6000 7000 8000 9000 0.5 0.55 0.6 0.65 0.7 0.75 0.8 0.85 F−1 Score Number of Training Sentences Virus NER baseline cca−word cca−phrase 0 1000 2000 3000 4000 5000 6000 0.45 0.5 0.55 0.6 0.65 0.7 0.75 0.8 Number of Training Sentences F−1 Score Disease NER baseline cca−word cca−phrase 1 Figure 3: V irus and Disease NER F-1 scores for varying training data size when embeddings obtained from dif ferent methods are used as features pre viously , the dictionaries produced from our ap- proach performs competitively with dictionaries that are entirely built by domain e xperts. 5.3.2 Embedding F eatures The quality of the candidate phrase embeddings are compared with word embeddings by adding the embeddings as features in the CRF tagger . Along with the baseline features, CCA-word model adds word embeddings as features while the CCA-phrase model adds candidate phrase em- beddings as features. CCA-word model is similar to the one used in Dhillon et al. (2011). W e considered adding 10, 20, 30, 40 and 50 di- mensional w ord embeddings as features for ev ery training data size and the best performing model on the de velopment data was pick ed for the e xper- iments on the test data. For candidate phrase em- beddings we used the same number of dimensions that was used for training the SVMs to construct the best dictionary . When candidate phrase embeddings are ob- tained using CCA, we do not hav e embeddings for words which are not in the list of candidate phrases. Also, a candidate phrase having more than one word has a joint representation, i.e., the phrase “human immunodeficiency” has a lower dimensional representation while the words “hu- man” and “immunodeficiency” do not have their o wn lower dimensional representations (assuming they are not part of the candidate list). T o ov er- come this issue, we used a simple technique to dif- ferentiate between candidate phrases and the rest of the words. Let x be the highest real valued can- didate phrase embedding and the candidate phrase embedding be a d dimensional real v alued vector . If a candidate phrase occurs in a sentence, the em- beddings of that candidate phrase are added as fea- tures to the first word of that candidate phrase. If the candidate phrase has more than one word, the other words in the candidate phrase are gi ven an embedding of dimension d with each dimension having the v alue 2 × x . All the other words are gi ven an embedding of dimension d with each di- mension having the v alue 4 × x . Figure 3 shows that almost always the candi- date phrase embeddings help the CRF model. It is also interesting to note that sometimes the word- le vel embeddings hav e an adverse affect on the performance of the CRF model. The CCA-phrase model performs significantly better than the other two models when there are fewer labeled sen- tences to train and the separation of the candidate phrases from the other words seems to have helped the CRF model. 6 Conclusion W e described an approach for automatic construc- tion of dictionaries for NER using minimal super- vision. Compared to the previous approaches, our method is free from ov erly-stringent assumptions about the data, uses SVD that can be solved ex- actly and achiev es better empirical performance. Our approach which uses a small number of seed examples performs competitively with dictionar- ies that are compiled manually . Acknowledgments W e are grateful to Alexander Rush, Ale xandre Passos and the anon ymous revie wers for their useful feedback. This work was supported by the Intelligence Advanced Research Projects Ac- ti vity (IARP A) via Department of Interior Na- tional Business Center (DoI/NBC) contract num- ber D11PC20153. The U.S. Go vernment is autho- rized to reproduce and distribute reprints for Gov- ernmental purposes notwithstanding any copy- right annotation thereon. The views and conclu- sions contained herein are those of the authors and should not be interpreted as necessarily represent- ing the official policies or endorsements, either e x- pressed or implied, of IARP A, DoI/NBC, or the U.S. Gov ernment. References [McCallum and Li2003] Andre w McCallum and W ei Li. Early Results for Named Entity Reco gnition with Conditional Random F ields, F eatur e Induction and W eb-Enhanced Le xicons. 2003. Conference on Nat- ural Language Learning (CoNLL). [Mnih and Hinton2007] Andriy Mnih and Geoffrey Hinton. Thr ee Ne w Graphical Models for Statistical Language Modelling. 2007. International Confer- ence on Machine learning (ICML). [T oral and Mu ˜ noz2006] Antonio T oral and Rafael Mu ˜ noz. A pr oposal to automatically build and maintain gazetteers for Named Entity Recognition by using W ikipedia. 2006. W orkshop On Ne w T ext W ikis And Blogs And Other Dynamic T ext Sources. [Blum and Mitchell1998] A vrin Blum and T om M. Mitchell. Combining Labeled and Unlabeled Data with Co-T raining . 1998. Conference on Learning Theory (COL T). [Burr Settles2004] Burr Settles. Biomedical Named Entity Recognition Using Conditional Random F ields and Rich F eatur e Sets. 2004. International Joint W orkshop on Natural Language Processing in Biomedicine and its Applications (NLPB A). [Hotelling1935] H. Hotelling. Canonical correlation analysis (cca) 1935. Journal of Educational Psy- chology . [Zhang et al.2004] Jie Zhang, Dan Shen, Guodong Zhou, Jian Su and Chew-Lim T an. Enhancing HMM-based Biomedical Named Entity Recognition by Studying Special Phenomena. 2004. Journal of Biomedical Informatics. [GENIA2003] Jin-Dong Kim, T omoko Ohta, Y uka T ateisi and Jun’ichi Tsujii. GENIA corpus - a semantically annotated corpus for bio-textmining . 2003. ISMB. [Kazama and T orisawa2007] Junichi Kazama and Ken- taro T orisawa. Exploiting W ikipedia as External Knowledge for Named Entity Recognition. 2007. Association for Computational Linguistics (A CL). [Sridharan and Kakade2008] Karthik Sridharan and Sham M. Kakade. An Information Theor etic F rame- work for Multi-view Learning. 2008. Conference on Learning Theory (COL T). [Ratinov and Roth2009] Lev Ratinov and Dan Roth. Design Challenges and Misconceptions in Named Entity Recognition. 2009. Conference on Natural Language Learning (CoNLL). [Collins and Singer1999] Michael Collins and Y oram Singer. Unsupervised Models for Named Entity Classification. 1999. In Proceedings of the Joint SIGD A T Conference on Empirical Methods in Nat- ural Language Processing and V ery Large Corpora. [Halko et al.2011] Nathan Halko, Per-Gunnar Martins- son, Joel A. Tropp. F inding structure with random- ness: Pr obabilistic algorithms for constructing ap- pr oximate matrix decompositions. 2011. Society for Industrial and Applied Mathematics. [Dhillon et al.2011] Paramv eer S. Dhillon, Dean Foster and L yle Ungar . Multi-V iew Learning of W ord Em- beddings via CCA. 2011. Advances in Neural In- formation Processing Systems (NIPS). [Dhillon et al.2012] Paramv eer Dhillon, Jordan Rodu, Dean Foster and L yle Ungar . T wo Step CCA: A new spectral method for estimating vector models of wor ds. 2012. International Conference on Machine learning (ICML). [Dogan and Lu2012] Rezarta Islamaj Dogan and Zhiy- ong Lu. An impr oved corpus of disease mentions in PubMed citations. 2012. W orkshop on Biomedical Natural Language Processing, Association for Com- putational Linguistics (A CL). [Leaman et al.2010] Robert Leaman, Christopher Miller and Graciela Gonzalez. Enabling Recogni- tion of Diseases in Biomedical T ext with Machine Learning: Corpus and Benchmark. 2010. W ork- shop on Biomedical Natural Language Processing, Association for Computational Linguistics (A CL). [Collobert and W eston 2008] Ronan Collobert and Ja- son W eston. A unified ar chitectur e for natural lan- guage pr ocessing: deep neural networks with mul- titask learning . 2008. International Conference on Machine learning (ICML). [McDonald and Pereira2005] Ryan McDonald and Fer - nando Pereira. Identifying Gene and Pr otein Men- tions in T e xt Using Conditional Random F ields. 2005. BMC Bioinformatics. [Kakade and Foster 2007] Sham M. Kakade and Dean P . Foster . Multi-view r e gression via canonical cor - r elation analysis. 2007. Conference on Learning Theory (COL T). [Cohen and Sarawagi2004] W illiam W . Cohen and Sunita Sarawagi. Exploiting Dictionaries in Named Entity Extraction: Combining Semi-Markov Extr ac- tion Pr ocesses and Data Integr ation Methods. 2004. Semi-Markov Extraction Processes and Data Inte- gration Methods, Proceedings of KDD.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment