Fitting birth-death processes to panel data with applications to bacterial DNA fingerprinting
Continuous-time linear birth-death-immigration (BDI) processes are frequently used in ecology and epidemiology to model stochastic dynamics of the population of interest. In clinical settings, multiple birth-death processes can describe disease traje…
Authors: Charles R. Doss, Marc A. Suchard, Ian Holmes
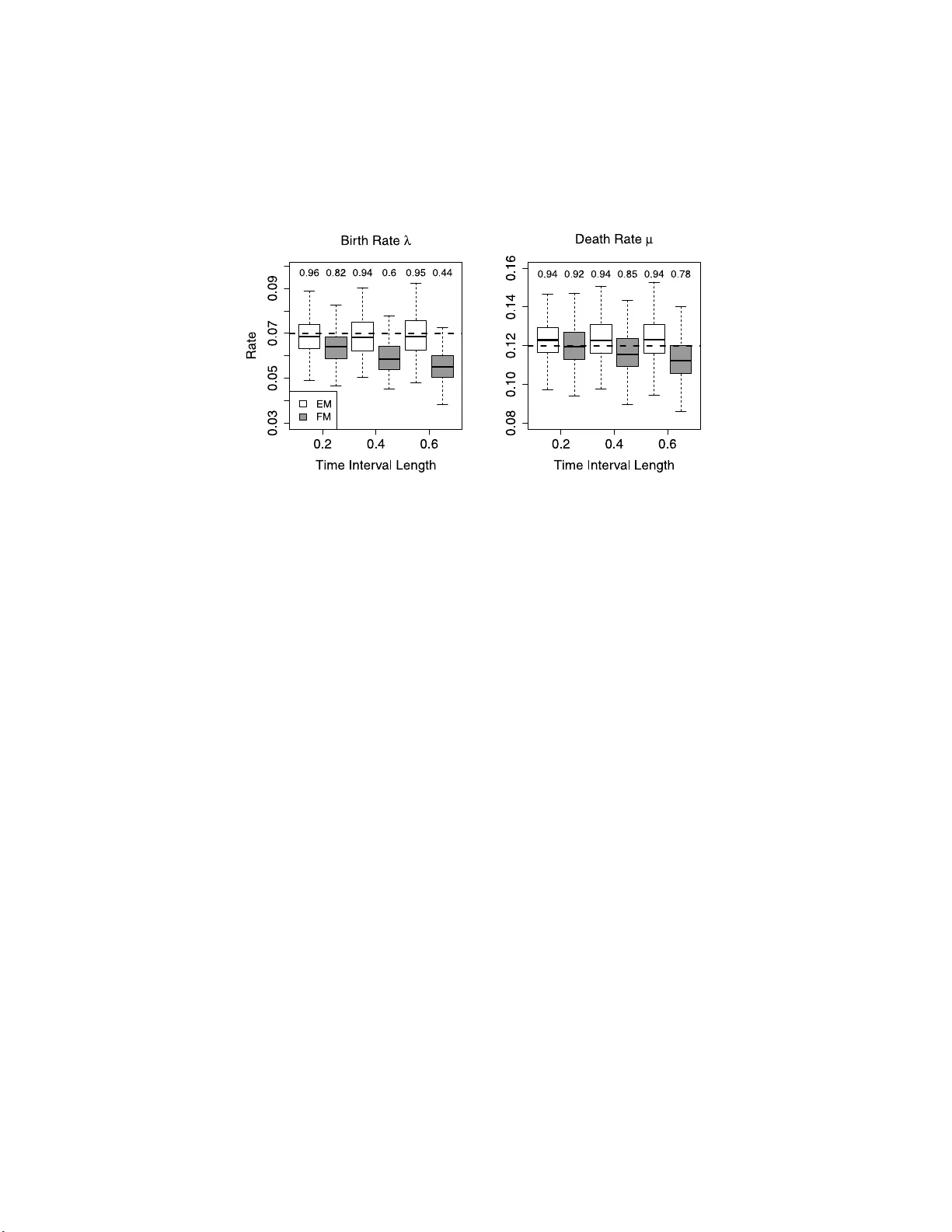
The Annals of Applie d Statistics 2013, V ol. 7, No. 4, 2315–233 5 DOI: 10.1214 /13-A OAS673 c Institute of Mathematical Statistics , 2 013 FITTING BIR TH–DEA TH PR OCES SES TO P ANEL D A T A W ITH APPLICA TIONS TO BA C TERIAL DNA FING ERPRINTING By Charles R. Doss ∗ , Marc A. Suchard 1 , 5 , † , Ian Holm e s 2 , ‡ , Midori Ka to-Maeda 3 , § and Vladimir N. Minin 4 , 5 , ∗ University of Washington, Se attle ∗ , University of California, L os Angeles † , University of California, Berkeley ‡ , and Univ e rsity of California, San F r ancisc o § Con tinuous-time linear birth–death-immigration (BDI ) pro cesses are frequently u sed in ecology and epidemiology to mod el stochastic dynamics of the p opulation o f interest. I n clinical settings, multiple birth–death pro cesses can describ e disease tra jectories of individu al patients, allo wing for estimation of the effects of individual cov ari- ates on the birth and death rates of the pro cess. Such estimation is usually accomplished by analyzing patient data collected at unevenly spaced time p oints, referred to as panel d ata in the biostatistics lit- erature. Fitting linear BDI pro cesses to panel data is a nontrivial optimization problem b ecause birth and death rates can b e functions of man y parameters related to the cov ariates of interest. W e prop ose a nov el exp ectation–maximization (EM) algorithm for fi tting linear BDI mo dels with cov ariates to panel data. W e derive a closed-form expression for the joint generating fun ct ion of some of the BDI pro- cess statistics and u se th is generating function to redu ce t he E-step of the EM algorithm, as well as calculation of the Fisher informa- tion, to one-dimensional in t egration. This analytical technique yields a computationally efficient and robust optimization algorithm that w e implemented in an open- source R pack age. W e apply our meth o d to DNA fingerprinting of Myc ob acterium tub er culosis , the causative agen t of tub erculosis, to study intrapatient time evolution of IS 6110 copy number, a genetic marker frequently used during estimation of epidemiological clusters of Myc ob acterium tub er culosis infections. Received December 2012; revised July 2013. 1 Supp orted by NI H Gran t R 01 GM0868 87. 2 Supp orted by NI H Gran t GM076705. 3 Supp orted by NI H Gran t A I034238. 4 Supp orted by th e UW Roya lty Research F un d. 5 Supp orted by NS F Gran t DMS- 08-56099. Key w or ds and phr ases. Missing data, EM algorithm, transp osable element, IS6110, tub erculosis. This is an electronic reprint of the or iginal ar ticle published by the Institute of Mathematical Statistics in The Annals of Applie d Statistics , 2013, V o l. 7, No. 4, 231 5–233 5 . This reprint differs from the or iginal in pa g ination and typo graphic deta il. 1 2 C. R. DOSS ET AL. Our analysis reveals previously und ocumented differences in IS 6110 birth–death rates among t hree ma jor lineages of Myc ob acterium tu- b er culosis , whic h has imp ortant implications for epidemiologists that use IS 6110 for DNA fingerprinting of Myc ob acterium tub er culosis . 1. In tro du ction. Linear b irth–death-immigration (BDI) pro cesses p ro- vide u seful buildin g b lo c ks for mo deling p opulation dynamics in ecolog y [Nee ( 2006 )], molecular ev olution [Th orne, Kishin o and F elsenstein ( 1991 )] and epid emiology [Gibson an d Rensh a w ( 1998 )], among m any other areas. Although Keiding ( 1975 ) has extensiv ely studied inference for fully ob- serv ed con tinuous-time BDI pro cesses, more often s uc h pro cesses are not observ ed completely , p osing challengi ng computational problems for statis- ticians. Here, w e u s e applied probab ility to ols to deve lop a new, efficien t implemen tation of the exp ectation–maximiza tion (EM) algorithm for fitting discretely obs er ved BDI pro cesses. W e are interested in situations where w e observe multiple ind ep endent con tinuous-time BDI tra jectories at fixed, p ossibly ir regularly s p aced, time p oint s. Su c h observ ations, called panel data, often arise in med ical applica- tions, with indep enden t BDI tra jectories corresp onding to some sto chastic pro cess recorded in different p atients und er stud y [Cr esp i, C um b erland and Blo w er ( 2005 )]. The birth and death rates can then b e mo deled as fu nctions of patient -sp ecific co v ariates. This mo deling fr amew ork is similar to th e u se of con tinuous-time Marko v c h ains (CTMCs) in m u lti-state disease p rogres- sion mo d els with a finite n um b er of states [Kalbfleisc h and Lawle ss ( 1985 )]. Although established metho ds for fitting finite state CT MC s to panel d ata exist [Kalbfleisc h and L awless ( 1985 ), Lange ( 1995 ), Jackson ( 2011 )], less atten tion has b een paid to infinite state-space pro cesses, such as BDI m o d- els. Outside of medical applications, estimating parameters of discretely ob- serv ed BDI mo dels is considered in the molecular evolutio n and bioinfor- matics literature [Thorne, K ishino and F elsenstein ( 1991 ), Holmes ( 2005 )]. F or example, Holmes ( 200 5 ) prop osed an E M algo rithm for discretely ob- serv ed BDI pr o cesses in the con text of fi nding the most optimal alignmen t of multiple genomic s equences. The author argues that th e E M algorithm’s simplicit y and robustness mak e this metho d attractiv e for large-scale bioin- formatics app lications. Un fortunately , imp lemen tation of the E M algorithm b y Holmes ( 2005 ) is applicable only to a ve r y restricted class of BDI pro- cesses. In this pap er, we deve lop a more general EM algorithm that applies to a large class of BDI mo dels and is n ot restricted to molecular ev olution applications. Computing exp ecta tions of the complete-data log-lik eliho o d, needed f or executing an EM algorithm, can b e c hallenging, esp ecially if the complete- data w er e generated by a conti n uous-time s to chastic pro cess. When the com- plete data are generated by a fi nite state-space CTMC, th ese exp ecta tions BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 3 can b e computed efficien tly [Lange ( 1995 ), Holmes and Rubin ( 2002 )]. Al- though th e BDI pro cess is also a CTMC, the infinite state-space of the pro cess prohibits us from using these computationally efficient metho d s. Holmes ( 2005 ) considers a BDI mo del w ith the immigration rate either zero or prop ortional to the birth rate. Und er this r estriction, th e complete-data lik eliho o d b elongs to the exp onen tial family , whic h means th at the complete- data log-lik eliho o d is a linear fun ction of sufficient statistics of the complete data. Making furth er stringent assu m ptions ab out the initial state of th e pro cess, Holmes ( 2005 ) compu tes exp ectations of these suffi cient statistics b y numerically solving a system of coupled nonlin ear ordinary differen tial equations (ODEs). W orking with this b irth–death-restricted immigratio n (BDRI) mo del, bu t without an y restrictio ns on the starting state of the pro cess, we d ev elop a new computationally efficien t metho d for computing the exp ected suffi cien t s tatistics. Ou r metho d com bin es ideas from Kend all ( 1948 ) and Lange ( 1982 ) and r educes computations of the exp ected s u ffi- cien t statistics to one-dimensional int egration, a computational task that is m uc h simpler than solving a system of n onlinear ODEs. W e d ev elop a similar integ ration metho d to compute the observed Fisher information ma- trix via Louis’ form ula [Louis ( 1982 )] and use this matrix for calculation of confi dence int erv als and sets. In add ition, wh en we ha ve multi ple BDRI tra jectorie s observed, w e allo w the bir th and death rates to b e fu nctions of tra jectory-specific co v ariates. W e first test our EM algorithm on s imulated data and then turn to a problem of estimating birth and death rates of the transp osable elemen t IS 6110 in M yc ob acterium tub er culosis , th e causativ e b acterial agen t of most tub erculosis (TB) in humans. M yc ob acterium tub er culosis genome carries m u ltiple IS 6110 copies th at get du p licated and deleted rapidly d uring repli- cation. Estimating IS 6110 cop y num b er birth (duplication) and death (loss) rates is an imp ortan t task in TB molecular epid emiology b ecause r esearc hers use IS 6110 copy num b er to group infected ind ividuals into epidemiological clusters [Small et al. ( 1994 )]. In the United States, the r esurgence of T B cases, attributed to signifi can t c h an ges in so cio economic factors, started in the late 1980s, with the n umber of TB cases reac h in g its p eak in 1991 and steadily d eclining since then [Cattamanc hi et al. ( 2006 )]. Since 1991, the Univ ersity of C alifornia, San F rancisco has b een maint aining a database of TB cases rep orted to the San F rancisco Department of Pu blic Health. T he database con tains demographic and certain clinical information as w ell as M. tub er culosis genot yp es (e.g., IS 6110 cop y n u m b er) f or eac h rep orted TB case [Jasmer et al. ( 1999 )]. Rosen b erg, Tsolaki and T anak a ( 2003 ) u sed a su b set of this database to estimate IS 6110 birth an d death rates. Th ese au th ors prop osed an approximat e lik eliho o d metho d to accomplish this estimation. W e revisit this problem using our EM algorithm and compare our results with the app ro ximation of Rosenberg, Tsolaki and T anak a ( 2003 ). F urther, 4 C. R. DOSS ET AL. w e examine differences in b irth and death r ates among three main lineages of M. tub er culosis and find that the East-Asian M. tub er culosis is ev olv- ing at a s low er rate than its Europ ean–American counterpart. This no vel finding h as serious implications on th e definition of ep idemiologic al clusters based on the IS 6110 copy num b er. T o inv estigate the p ossibility of s p urious effect of M . tub er culosis lineage on IS 6110 birth and death rates due to a confounding factor, we bu ild a more complicated mo del for birth and d eath rates. In addition to the lineage, we include M . tub er culosis d rug-resistance status and HIV infection status of eac h patien t as birth and d eath r ate co- v ariates. W e fi nd that after in clud ing these co v ariates, the lineage remains the only v ariable that significantly affects IS 6110 birth and death rates. 2. BDRI pro cess w ith co v ariates. W e start with m indep endent con ti- n u ous-time homoge neous linear BDRI pro cesses { X p,t } , for p = 1 , . . . , m , with corresp on d ing p er capita b ir th rates λ p ≥ 0, p er capita death rates µ p ≥ 0 and immigration rates ν p = β λ p , wh ere β ≥ 0 is a kno w n constan t. Assuming that eac h pro cess p h as c 1 co v ariates related to th e birth rates and c 2 co v ariates r elated to the d eath rates, colle cted in to ve ctors z ′ p,λ = ( z p,λ, 1 , . . . , z p,λ,c 1 ) ∈ R c 1 and z ′ p,µ = ( z p,µ, 1 , . . . , z p,µ,c 2 ) ∈ R c 2 , we mo del birth and death rates as log-linear fu nctions of these co v ariates: log λ p = z ′ p,λ γ λ and log µ p = z ′ p,µ γ µ , (1) where γ ′ λ = ( γ λ, 1 , . . . , γ λ,c 1 ) and γ ′ µ = ( γ µ, 1 , . . . , γ µ,c 2 ) are birth and death regression coefficients. Cov ariate v ectors z p,λ and z p,µ are assumed to b e kno wn and fixed for eve ry pro cess p . F or example, if eac h BDRI pro cess mo dels a disease related tra jectory for eac h p atient, then co v ariates are usually comp osed of patien t-sp ecific clinical and demographic inform ation (e.g., gender, medical history). W e assume th at we observ e the p th pro cess at n ( p ) + 1 distinct times, 0 = t p, 0 < t p, 1 < · · · < t p,n ( p ) . W e den ote our data vec tor b y Y = ( X 1 ,t 1 , 0 , . . . , X 1 ,t 1 ,n (1) , . . . , X m,t m, 0 , . . . , X m,t m,n ( m ) ) and the parameter v ector by γ = ( γ λ , γ µ ) ∈ R c 1 + c 2 . W e are in terested in com- puting the parameter maxim um lik eliho o d estimates (MLEs), ˆ γ = arg max γ l o ( Y ; γ ), where l o ( Y ; γ ) := m X p =1 n ( p ) − 1 X i =0 log p X p,t p,i ,X p,t p,i +1 ( t p,i +1 − t p,i ; λ p , µ p ) (2) is the ob s erv ed-data log -likeli ho o d and p i,j ( t ; λ, µ ) = P λ,µ ( X t = j | X 0 = i ), i, j = 0 , 1 , . . . , are the transition probabilities of the BDRI pro cess. These transition pr obabilities can b e calculated either u sing the generating function BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 5 deriv ed by Kendall ( 1948 ) or v ia the orthogonal p olynomial repr esen tation of Karlin and McG regor ( 1958 ). Despite the explicit algebraic nature of the orthogonal p olynomials, the latter metho d can b e n umerically uns table and the generating f u nction metho d is often preferred [S ehl et al. ( 2011 )]. Although one can maximize the lik eliho o d l o ( Y ; γ ) using standard off-the- shelf optimization algorithms, suc h generic algorithms can b e pr oblematic when the BDI rates are fu nctions of a high-dim en sional parameter vec tor, suc h as the vecto r of regression co efficien ts γ in our case. As an alternativ e to generic optimization, we dev elop an EM algorithm, kn o wn for its robu stness and abilit y to cop e with h igh-dimensional optimizati on [Dempster, Laird and Rub in ( 1977 )]. 3. EM algorithm for the BDRI pro cess. The complete d ata in our case consist of the BDRI tra jectories { X p,t } , observed con tinuously during the corresp onding interv als [0 , t p,n ( p ) ], p = 1 , . . . , m . Let X = { X p,t } t ∈ [0 ,t p,n ( p ) ] p =1 ,...,m b e the complete data and let l c ( X ; γ ) b e th e complete data log-lik eliho o d. The EM algorithm starts by initializing the parameter v ector to an arbitrarily c hosen v ector γ 0 . A t the k th iteration of th e algorithm we set γ k = arg max γ E γ k − 1 [ l c ( X ; γ ) | Y ] . (3) T o accomplish the ab o ve m aximization, w e need to b e able to ev aluate the exp ectation in ( 3 ) for an y vect or γ . T raditionally , a numerical pro cedure for computing suc h an exp ectatio n is called an E-step of the EM algo r ithm. The maximization of the exp ecta tion is called an M-step of the EM algo rithm. Belo w, w e d ev elop efficien t algorithms for implemen ting these E- and M- steps for the discretely observed BDRI pro cess. As is often the case, we will see that to compu te the needed exp ectations for all γ ∈ R c 1 + c 2 , w e need to compute only the exp ectations of certain statistics that do not dep end on γ . 3.1. E-step. Since our BDRI pro cess is a CTMC, the log-lik eliho o d of the complete data is l c ( X ; γ ) = − m X p =1 " ∞ X i =0 d p ( i )[ i ( λ p + µ p ) + ν p ] (4) + ∞ X i =0 ( n p i,i +1 log( iλ p + ν p ) + n p i,i − 1 log( iµ p )) # , where d p ( i ) is the tota l time sp en t by X p,t in state i and n p i,j is th e n um b er of jumps from state i to state j during th e inte rv al [0 , t p,n ( p ) ] [Guttorp ( 1995 )]. Replacing ν p with β λ p in the ab o v e equation, we arriv e at a m ore compact 6 C. R. DOSS ET AL. represent ation of the complete-data log-lik eliho o d : l c ( X ; γ ) = m X p =1 [ − R p,t p,n ( p ) ( λ p + µ p ) − t n ( p ) β λ p (5) + N + p,t p,n ( p ) log λ p + N − p,t p,n ( p ) log µ p ] + const , where the num b er of jum ps up N + p,t p,n ( p ) := P i ≥ 0 n p i,i +1 , the n um b er of jump s do wn N − p,t p,n ( p ) := P i ≥ 0 n p i,i − 1 , and th e total particle-time R p,t p,n ( p ) := Z t p,n ( p ) t 0 X s ds = ∞ X i =0 id p ( i ) for p = 1 , . . . , m , are the sufficien t statistics. Equ ation ( 5 ) sho ws that, for the E-step, the only exp ectations we need are E ˜ γ [ N + p,t p,n ( p ) | Y ], E ˜ γ [ N − p,t p,n ( p ) | Y ] and E ˜ γ [ R p,t p,n ( p ) | Y ] for all v alues ˜ γ . Using indep endence of the p BD RI pro cesses, the Marko v prop ert y and additivit y of exp ectations, we break the desired exp ectations in to su m s of exp ectations of the num b ers of jump s up and d o wn and the total particle time du ring eac h time interv al [ t p,k , t p,k +1 ], conditional on X p,t p,k and X p,t p,k +1 . By the h omogeneit y of eac h of the BDRI pro cesses, in order to complete th e E-step of the EM algorithm, we need to b e able to calculate U i,j ( t ) = U i,j ( t ; λ, µ ) = E( N + t | X 0 = i, X t = j ) , D i,j ( t ) = D i,j ( t ; λ, µ ) = E( N − t | X 0 = i, X t = j ) and (6) P i,j ( t ) = P i,j ( t ; λ, µ ) = E( R t | X 0 = i, X t = j ) for all n onnegativ e in tegers i and j . F ollo w in g Minin and Suc hard ( 2008 ), w e c h o ose to w ork with restricted momen ts ˜ U i,j ( t ) = ˜ U i,j ( t ; λ, µ ) = E( N + t 1 { X t = j } | X 0 = i ) , ˜ D i,j ( t ) = ˜ D i,j ( t ; λ, µ ) = E( N − t 1 { X t = j } | X 0 = i ) and (7) ˜ P i,j ( t ) = ˜ P i,j ( t ; λ, µ ) = E( R t 1 { X t = j } | X 0 = i ) , that w e can divide by trans ition probabilities p i,j ( t ) to reco v er the condi- tional exp ecta tions ( 6 ), U i,j ( t ) = ˜ U i,j ( t ) /p i,j ( t ) , D i,j ( t ) = ˜ D i,j ( t ) /p i,j ( t ) and (8) P i,j ( t ) = ˜ P i,j ( t ) /p i,j ( t ) . BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 7 In order to compu te the restricted momen ts, w e first consider th e joint gen- erating fu nction H i ( u, v , w , s, t ) := E( u N + t v N − t e − w R t s X t | X 0 = i ) , (9) where 0 ≤ u, v , s ≤ 1 and w ≥ 0 . Pa rtial d eriv ativ es of this function, ∂ H i ( u, 1 , 0 , s, t ) ∂ u u =1 = ∞ X j = 0 s j ∞ X n =0 n Pr i ( N + t = n, X t = j ) = ∞ X j = 0 ˜ U i,j ( t ) s j , ∂ H i (1 , v , 0 , s, t ) ∂ v v =1 = ∞ X j = 0 s j ∞ X n =0 n Pr i ( N − t = n, X t = j ) = ∞ X j = 0 ˜ D i,j ( t ) s j and (10) ∂ H i (1 , 1 , w , s , t ) ∂ w w =0 = − ∞ X j = 0 s j Z ∞ 0 x d Pr i ( R t ≤ x, X t = j ) = − ∞ X j = 0 ˜ P i,j ( t ) s j are p ow er series with co efficien ts ˜ U i,j ( t ), ˜ D i,j ( t ) and − ˜ P i,j ( t ), resp ectiv ely , for j = 0 , 1 , . . . , ∞ , wher e Pr i denotes pr obabilit y conditional on X 0 = i . W e will d enote these p o w er series by G + i ( t, s ) , G − i ( t, s ) and G ∗ i ( t, s ) , resp ectiv ely . If w e can compute G + i ( t, s ) , G − i ( t, s ) and G ∗ i ( t, s ) for ev ery p ossible t and s , then w e should b e able to reco v er co efficien ts of the corresp onding p ow er series via different iation or integ ration. Num erical ev aluation of the partial deriv ativ es ( 10 ) is s tr aigh tforwa r d if w e can compute finite differences of H i ( u, v , w , s, t ). Remark ably , H i ( u, v , w , s, t ) is a v ailable in closed form, as w e demonstrate in the theorem b elo w, so one can ev en obtain d er iv ativ es ( 10 ) analytically . Note that the theorem b elo w applies to a general linear BDI pro cess, n ot only to the BDRI pro cesses. Theorem 1. L et { X t } b e a line ar BDI pr o c ess with p ar ameters λ ≥ 0 , µ ≥ 0 and ν ≥ 0 . Over the interval [0 , t ] , let N + t b e the numb er of jumps up, N − t b e the nu mb er of jumps down and R t b e the total p article-time. Then H i ( u, v , w , s, t ) = E ( u N + t v N − t e − w R t s X t | X 0 = i ) satisfies the fol lowing p artial differ ential e quation: ∂ ∂ t H i = [ s 2 uλ − ( λ + µ + w ) s + v µ ] ∂ ∂ s H i + ν ( us − 1) H i , (11) 8 C. R. DOSS ET AL. subje ct to initial c ondition H i ( u, v , w , s, 0) = s i . The Cauchy pr oblem define d by e quation ( 11 ) and the initial c ond ition has a unique solution. When λ > 0 , the solution is H i ( u, v , w , s, t ) = α 1 − α 2 ( s − α 1 ) e − λ ( α 2 − α 1 ) ut / ( s − α 2 ) 1 − ( s − α 1 ) e − λ ( α 2 − α 1 ) ut / ( s − a 2) i (12) × α 1 − α 2 s − α 2 − ( s − α 1 ) e − λ ( α 2 − α 1 ) ut ν /λ e − ν (1 − uα 1 ) t , wher e α 1 = λ + µ + w − √ ( λ + µ + w ) 2 − 4 λµuv 2 λu and α 2 = λ + µ + w + √ ( λ + µ + w ) 2 − 4 λµuv 2 λu . When λ = 0 , the solution is H i ( u, v , w , s, t ) = se − ( µ + w ) t − v µ ( e − ( µ + w ) t − 1) µ + w i (13) × e ν u [ vµ − ( µ + w ) s ]( e − ( µ + w ) t − 1) / ( µ + w ) 2 + ν ( uv µ / ( µ + w ) − 1) t . Pr oof. Our pr o of, detailed in Ap p endix A, is a generalizatio n of Ken dall’s deriv ation of the generating function of X t [Doss et al. ( 2013 ), Kendall ( 1948 )]. Ha ving H i in closed form give s us access to fu nctions G + i , G − i and G ∗ i , so w e are left with the task of reco v ering co efficien ts of th ese p o we r series. On e w ay to accomplish this task is to differen tiate the p ow er series r ep eatedly , for example, ˜ U i,j ( t ) = 1 j ! ∂ j G + i ( s,t ) ∂ s j | s =0 . In App endix C, w e demonstrate that for the d eath-immigration mo del ( λ = 0 , ν 6 = 0 , µ 6 = 0) and the BDRI mo del considered b y Holmes ( 2005 ), these d er iv ativ es can b e found analytically [Doss et al. ( 2013 )]. In general, rep eate d d ifferen tiation of G + i , G − i and G ∗ i needs to b e done numerically , making this metho d impr actica l. Instead, w e extend G + i ( t, · ), G − i ( t, · ) and G ∗ i ( t, · ) to the b oundary of a unit circle in the complex plane by the c hange of v ariables s = e 2 π iz ( i in this con text is the imaginary num b er √ − 1, not th e initial state of the BDI pro cess). F or example, G + l ( t, e 2 π iz ) = ∞ X j = 0 ˜ U l,j ( t ) e 2 π ij z is a p eriod ic fu nction in z , whic h means that ˜ U l,j ( t ) are F ourier co efficien ts of this p eriodic f unction. Th erefore, we can u se the Riemann appr o ximation to the F ourier transform integral to obtain ˜ U l,j ( t ) = Z 1 0 G + l ( t, e 2 π is ) e − 2 π ij s ds ≈ 1 K K − 1 X k =0 G + l ( t, e 2 π ik/K ) e − 2 π ij k /K BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 9 for some suitably large K . The F ast F our ier T r ansform (FFT) [Henrici ( 1979 )] can b e applied to quic kly compute multiple F ourier co efficien ts [Lange ( 1982 ), Dorman, Sin sheimer and Lange ( 2004 ), Suc hard, Lange and Sin- sheimer ( 2008 )]. W e d o not, ho w ev er, u se the FFT in our algo rithm b ecause, for a particular time int erv al length t , w e almost alw ays n eed to compute ˜ U i,j ( t ), ˜ D i,j ( t ), ˜ P i,j ( t ) f or only one v alue of j . No w, we can put the p ieces together to compute E ˜ γ [ l c ( X ; γ ) | Y ]. As men- tioned ab o ve, N + p,t p,n ( p ) equals the sum of the n u m b er of ju m ps up o v er the disjoin t interv als [ t p,i − 1 , t p,i ), i = 1 , . . . , n ( p ). The Mark o v prop erty sa y s that the conditional exp ectations of the n um b er of jumps up of X p,t o ve r [ t p,i − 1 , t p,i ) giv en Y is equal to the conditional exp ectation of the num b er of jumps up ov er [ t p,i − 1 , t p,i ) giv en just X p,t p,i − 1 and X p,t p,i . Using similar logic for N − p,t p,n ( p ) and R p,t p,n ( p ) , this give s for p = 1 , . . . , m , E ˜ γ p [ N + p,t p,n ( p ) | Y ] = n ( p ) X i =1 U X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) , E ˜ γ p [ N − p,t p,n ( p ) | Y ] = n ( p ) X i =1 D X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) and (14) E ˜ γ p [ R p,t p,n ( p ) | Y ] = n ( p ) X i =1 P X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) , where log ˜ λ p = z ′ p,λ ˜ γ p,λ and log ˜ µ p = z ′ p,µ ˜ γ p,µ . Thus, by ( 5 ), ( 8 ) and ( 14 ), w e see that, up to an add itive constan t, E ˜ γ [ l c ( X ; γ ) | Y ] is equal to m X p =1 ( − t n ( p ) β λ p + n ( p ) X i =1 − ˜ P X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) p X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) ( λ p + µ p ) + ˜ U X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) p X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) log λ p + ˜ D X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) p X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) log µ p ) , where th e transition p robabilities p X p,t p,i − 1 ,X p,t p,i ( t p,i − t p,i − 1 ; ˜ λ p , ˜ µ p ) can b e calculate d b y u s ing the (kno w n) generating f unction for the BDI p ro cess, as is describ ed in Ap p endix A [Doss et al. ( 2013 )]. 10 C. R. DOSS ET AL. 3.2. M-step. T o complete the M-step for eac h iteration of the EM algo- rithm, we use a Newton–Raphson algorithm to maximize f ( γ ) = E ˜ γ [ l c ( X ; γ ) | Y ] . In eac h Newton–Raphson step, we u p date γ via the f ollo wing recursion: γ new = γ cur − [ H f ( γ cur )] − 1 ∇ f ( γ cur ) , where ∇ f ( γ cur ) is th e gradient v ector and H f ( γ cur ) is th e Hessian matrix of the fun ction f ( γ ). If we collect the observ ation times into a v ector T ′ = ( t 1 ,n (1) , . . . , t m,n ( m ) ), th e exp ectati ons of the su fficien t statistics into v ectors U ′ = ( E ˜ γ [ N + 1 ,t 1 ,n (1) | Y ] , . . . , E ˜ γ [ N + m,t m,n ( m ) | Y ]) , D ′ = ( E ˜ γ [ N − 1 ,t 1 ,n (1) | Y ] , . . . , E ˜ γ [ N − m,t m,n ( m ) | Y ]) , (15) P ′ = ( E ˜ γ [ R 1 ,t 1 ,n (1) | Y ] , . . . , E ˜ γ [ R m,t m,n ( m ) | Y ]) , and the pro cess-sp ecific birth and death rates in to v ectors λ ′ = ( λ 1 , . . . , λ m ) and µ ′ = ( µ 1 , . . . , µ m ) , then after d efining co v ariate matrices Z ′ λ = ( z 1 ,λ , . . . , z m,λ ) and Z ′ µ = ( z 1 ,µ , . . . , z m,µ ) , the gradient and the Hessian can b e compactly expressed in matrix form as ∇ f ( γ ) = ( Z ′ λ [ − diag( P + β T ) λ + U ] , Z ′ µ [ − diag( P ) µ + D ]) , (16) H f ( γ ) = − Z ′ λ diag( P + β T ) diag ( λ ) Z λ 0 0 − Z ′ µ diag( P ) diag ( µ ) Z µ , (17) whic h we sh o w in App endix B; see (S-4), (S-6) and (S-9) [Doss et al. ( 2013 )]. Notice that the algebraic separation of the birth and the death comp onen ts in the complete-data likel iho o d results in blo cks—co rresp ond ing to γ λ and γ µ —in the abov e form ulae. The fact that th e gradien t and Hessian of f ( γ ) is a v ailable analytically r esults in fast execution of Newton–Raphson u p dates. In our exp erience, the Newton–Raphson algorithm in our M-step con verges after only 3–5 iterations. Ho w ever, w e also note th at it is not critical to ac hiev e con vergence of this algorithm since ev en a single Newton–Raphson up d ate within the M-step is enough to guaran tee the u sual conv ergence prop erties of the EM algorithm [Lange ( 1995 )]. W e obtain the observ ed Fisher information via Louis’ form ula: ˆ I Y ( ˆ γ ) = E ˆ γ [ − H l c ( X ; ˆ γ ) | Y ] − E ˆ γ [ ∇ l c ( X ; ˆ γ ) ∇ l c ( X ; ˆ γ ) ′ | Y ] , where ∇ l c is the gradien t and H l c is the Hessian of the complete-data log- lik eliho o d [Louis ( 1982 )]. T his requ ires calculation of the conditional cross- pro du ct m eans, E [ N + t N − t | Y ], E[ N + t R t | Y ], E[ N − t R t | Y ], and the cond itional BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 11 T able 1 Summary statistics for the simulate d and M . tub er culosis IS6110 data V alue Simulated d ata IS6110 d ata Number of interv als 387 252 Average interv al length 5 0 . 35 Number of indiv id u als 100 196 Number of interv als with an increase 78 14 Average increase given an in crease 1 . 5 1 Number of interv als with a decrease 190 14 Average d ecrease give n a decrease 2 . 5 1 . 2 Number of interv als with no change 119 224 Mean starting state 5 . 5 11 Standard deviation of starting state 3 . 8 5 . 3 T otal length of time 1947 89 second moments of N + t , N − T and R t . The deriv ation of the information in terms of these moments is in App end ix B [Doss et al. ( 2013 )]. Th ese con- ditional second- and cross-momen ts, as well as P and D , can b e computed in analogous fash ion to U ab ov e, u sing th e join t generating f unction ( 12 ). W e use the inf orm ation matrix to compute approximat e standard errors of ˆ γ and use these s tandard errors together with asymptotic normalit y of maxim um lik eliho o d estimators to form confiden ce in terv als and s ets for our mo del p arameters. 4. Results. 4.1. Simulations. T o test our metho ds, w e sim u late data from the BDRI mo del with λ = 0 . 07, µ = 0 . 12 and β = 1 . 2, where β is assumed to b e kno wn, lea ving us with only tw o parameters to estimate: λ and µ . W e choose these parameters to resemble, but not exactly matc h, th e dynamics of our biologi- cal example, discus sed in the n ext su bsection. W e sim ulate 100 indep enden t pro cesses s tarting fr om initial states dr a wn uniformly b et ween 1 and 15. F rom eac h p ro cess we collect at lea st t wo observ ations. W e place obser v ation times un iformly b etw een 0 and 30. T able 1 giv es some summary statistics for the simulated data. W e test our EM algorithm and confidence in terv al calculations on these sim u lated data with initial parameter v alues of 0 . 2 for b oth λ and µ . W e considered other choi ces of starting v alues, but the algorithm was not sensi- tiv e to them. Notice that this is the simp lest parameterization of our BDRI mo del, w here b oth z λ and z µ are vec tors of ones. W e estimate 0 . 067 with a 95% confid ence in terv al of (0 . 052 , 0 . 081 ) for λ and 0 . 12 , (0 . 1 , 0 . 14) for µ , indicating that our algorithm su ccessfully r eco v ered these BDRI mo del p a- rameters. W e also condu ct a similar simulatio n study for the BDRI mo d el 12 C. R. DOSS ET AL. with cov ariates, successfully estimating parameters of this mo d el as w ell, but omit d etailed results of this sim u lation for b revit y . 4.2. Comp arison with the fr e quent monitoring metho d. W e compare our EM algorithm for computing the actual MLE to the frequent monitoring (FM) m etho d of Rosen b erg, Tsolaki and T anak a ( 2003 ) for computing the MLE of an app ro ximate lik eliho o d . In the FM metho d, Rosen b erg, Tsolaki and T anak a ( 2003 ) assume that if the starting and endin g v alues of the b irth– death p ro cess are equal for a particular in terv al, then no ju mps o ccur red in this interv al. F urther, if the difference b et w een the starting and ending v alues is − 1 or 1, then exactly one ju mp up or exactly one ju mp down m us t ha ve o ccurr ed, resp ecti v ely . The authors exclude all observed interv als, for whic h starting and ending v alues d iffer by more than one unit. Let i b e the starting state for an in terv al, t the length of th e int erv al and λ i = i ( λ + µ ). Then the corresp on d ing probabilities for the three p ossible ev ents are e − λ i u , iλ λ i (1 − e − λ i u ) and iµ λ i (1 − e − λ i u ), resp ectiv ely . Rosen b erg, Tsolaki and T anak a ( 2003 ) use this FM metho d to estimate r ates in w hat is effectiv ely a multi- state br anc hing p ro cess, but we will compare the tw o metho ds on our BDRI mo del with the immigration rate β constrained to b e 0. W e again simulate an un derlying BD p ro cess using λ = 0 . 07 and µ = 0 . 12. T o compare the tw o metho ds, w e generate thr ee different sets of data. In eac h set, w e generate observ ed states of the BD pro cess at a fixed constan t d istance dt apart. This distance v aries across the data sets, taking th e v alues 0 . 2 , 0 . 4 and 0 . 6, resp ectiv ely . W e rep eat this pro cedu re 200 times and compute birth and death rate estimates an d corresp ondin g 95% confi dence inte rv als u sing the EM algorithm and FM app ro ximation metho d. W e sh o w b ox plots of the resulting estimates for λ and µ in Figure 1 . As exp ected, th e FM estimates b eha v e reasonably when interv al lengths are small, bu t the appro ximation b ecomes p o or as w e increase the interv al length. Th e FM metho d alw a ys underestimates the parameters since the metho d effectiv ely und ercoun ts the n u m b er of unobserve d jumps in the BD p ro cess. W e also compute Mon te Carlo estimates of co verage probabilities of the tw o metho d s, shown ab o v e the b o x plots in Figure 1 . Not su rprisingly , co verage of the 95% confi dence in terv als computed u n der the pr op er BD mo del lik eliho o d are very close to the promised v alue of 0.95. In cont rast, the FM appro ximation-based 95% confidence in terv als con tain th e tru e parameter v alue less than 95% for all three simulatio n scenarios. 4.3. Myc ob acterium tub er culosis IS6110 tr ansp oson. W e apply our EM algorithm to estimation of birth and death rates of the transp oson IS 6110 in M . tub er culosis [McEvo y et al. ( 2007 )]. A transp oson, or transp osable el- emen t, is a genetic sequence that can duplicate, remov e itself and j ump to a BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 13 Fig. 1. Box plots of birth (left p anel) and de ath (right p anel) r ate estimates, obtaine d fr om 200 simulate d data sets using the EM al gorithm and fr e quent m oni toring (FM) m etho d. The true p ar ameter values, use d in data simulations, ar e marke d by the horizontal dashe d lines. Ab ove the b ox plots, we show Monte Carlo estimates of c over age pr ob abili ties of the 95% c onfidenc e intervals. new lo cation in the genome. IS 6110 is a transp oson that pla ys an imp ortant role in epid emiologi cal studies of tub erculosis. More sp ecifically , the num b er and lo cations of IS 6110 ele men ts in the M. tub er culosis form a genetic sig- nature or genot yp e of the m ycobacterium, allo win g epidemiologists to dr a w inference ab out disease transm ission when the same genot yp e is observe d among patien ts with activ e tub erculosis [v an Em b den et al. ( 1993 )]. Such genot ypic comparison can translate into meaningful epidemiological infer- ence only if the dynamics of I S 6110 ev olution are well understo o d . T here- fore, accurate estimation of rates of c hanges of IS 6110 -based genot yp es is critical f or using these genot yp es in epidemiological studies [T anak a and Rosen b erg ( 2001 )]. W e analyze data from an ongoing p opulation-based stud y that includes all tu b erculosis cases rep orted to the San F rancisco Departmen t of Public Health [Cattamanc h i et al. ( 2006 )]. O ur data includ e p atien ts with more th an one M . tub er culosis isolate from sp ecimens samp led more than 10 days apart and genotyped with IS 6110 restriction fragmen t length p olymorphism. W e ignore genomic lo cations of IS 6110 and assum e th at the transp oson coun ts are discretely observed realizatio ns of a BDRI pro cess, with no immigration ( β = 0); in particular, we assume th at p atients are n ot reinfected with a differen t strain of the bacteria in the p erio d b et we en observ ations. The th ir d column in T able 1 gives summary statistics for the d ata. W e fi rst u se a simple mo del with one single birth rate and one single death rate of the IS 6110 for all p atien ts. In th e analysis presented, w e s tart the EM algorithm w ith parameter gu esses of 0 . 05 and 0 . 05 for λ and µ , resp ectiv ely , and their MLEs are 0 . 0176 and 0 . 0207, resp ectiv ely . The start- 14 C. R. DOSS ET AL. ing v alues for the EM do n ot affect th ese r esults. Our estimate and 95% confidence in terv al for λ , 0 . 017 6 and (0 . 0 082 , 0 . 027), are consisten t with the corr esp onding quant ities, 0 . 0188 and (0 . 0085 , 0 . 0291), from Rosenb erg, Tsolaki and T anak a ( 2003 ). Although the authors’ confidence in terv al for µ , (0 . 0057 , 0 . 02 37), o v erlaps with ours, (0 . 011 , 0 . 031), our estimate for µ , 0 . 0207, is n oticeably higher than Rosenb erg, Tsolaki and T anak a’s ( 2003 ) estimate of 0 . 0147. Note from T able 1 that among the inte r v als with a d e- crease, the a verage coun t drop is b y more than 1; th ere are 3 interv als where IS 6110 coun ts drop by 2, wh ereas there are n o interv als that exp erience an increase by m ore than 1. Thus, we would exp ect our estimate for µ to in- crease o v er Rosen b erg, Tsolaki and T anak a’s ( 2003 ) appro ximation, wh er eas that of λ should b e similar b et ween the t w o metho ds. W e also p oint out that w e analyze an up dated version of the data analyzed by Rosenberg, Tsolaki and T anak a ( 2003 ). Moreo v er, Rosenberg, Tsolaki and T anak a ( 2003 ) use a slightl y m ore complicated mo del for IS 6110 evol ution, w hic h tak es int o accoun t shifts in transp oson lo cation. W e conclude that estimates of birth and d eath rates of IS 6110 do n ot v ary dramatically when estimation meth- o ds and d ata collection are altered. W e now tur n to more complicate d BDRI mo dels that ha ve not b een app lied b efore to the M. tub er culosis IS 6110 cop y n u m b er ev olution. These mo dels will tak e in to accoun t p oten tial d ep endence of IS 6110 b ir th and death rates on patien t-sp ecific co v ariates. 4.3.1. Myc ob acterium tub er culosis line age c omp arison. In addition to es- timation of the global b ir th and death rates, we separately estimate these parameters in eac h of the thr ee lineages of M . tub er culosis obs er ved in San F rancisco. Based on genomic sequence similarity , M. tub er culosis is divided into six main lineages: Eu r o-American, East-Asian, In do-Oceanic, East-African–Indian, W est-African I and W est-African I I [Gagneux et al. ( 2006 )]. I n our lineage-sp ecific analysis, w e consider 109 in dividuals infected with Euro-American (EU) lineage strains, 54 individuals infected with East- Asian (EA) lineage strains and 25 individuals in fected with Indo-Oceanic (IND) lineage strains. One simp le wa y to accommodate this lineage effect is to b uild a log-linear mo del for birth and death r ates with t w o categorica l co v ariates: log λ p = γ λ, 1 + γ λ, 2 EU p + γ λ, 3 IND p , log µ p = γ µ, 1 + γ µ, 2 EU p + γ µ, 3 IND p , where EU p = 1 if patien t p is inf ected with the EU strain and 0 otherwise, and IND p = 1 if patient p is infected with the IND strain and 0 otherwise. The in tercepts, γ λ, 1 and γ µ, 1 , corresp ond to birth and death of the EA strain. W e transform the co efficien ts ( γ λ, 1 , γ λ, 2 , γ λ, 3 ) and ( γ µ, 1 , γ µ, 2 , γ µ, 3 ) int o the M. tub er culosis lineage-sp ecific birth and death rates and show these estimates together with their corresp ond ing confidence in the fi r st column of Figure 2 . Most n otably , there app ears to b e a su bstan tial difference b etw een death rates of th e Euro-American and East-Asian lineages. W e r ep ort regression BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 15 Fig. 2. Point estimates and 95% c onfidenc e intervals for birth and de ath r ates of the IS6110 tr ansp osable element obtaine d by sep ar ately analyzing thr e e M. tub er culosis lin- e ages: Eur op e an–Americ an (EU), I ndo-Oc e anic (IND) and East Asian (EA) (lef tmost c olumn) and by fitting the l o g-line ar mo del with li ne age, drug r esistanc e and HIV status as c ovariates. F or the latter mo del, the estimate d r e gr ession c o efficients ar e tr ansforme d into four sets of line age-sp e cific birth and de ath r ates (last four c olum ns). co efficien ts on the m ultiplicativ e scale [e.g., exp( γ λ, 1 )] with their corresp ond- ing 95% confidence in terv als in the lineage mo del columns of T able 2 . In this table the highligh ted EU rate m ultiplier shows that the d eath rate of IS 6110 cop y num b er is estimated to b e app r o ximately ten times higher than the corresp onding d eath rate in the EA lineage. T he confidence in terv al of the EU r ate multi plier does not con tain one, indicating that EA an d EU lin eages ha ve different death r ates of the IS 6110 transp oson. Since this is a n o ve l result that h as implications for m onitoring tub ercu- losis with molecular genot yping, we examine the d ifference in death rates b et ween the three lineages m ore closely . More sp ecifically , w e add t wo bin ary co v ariates to our log-linear mo del: M. tub er culosis dru g resistance (DR) and HIV infection status of eac h patient (HIV + ). O ur n ew mo d el for birth and death rates b ecomes log λ p = γ λ, 1 + γ λ, 2 EU p + γ λ, 3 IND p + γ λ, 5 DR p + γ λ, 4 HIV + p , log µ p = γ µ, 1 + γ µ, 2 EU p + γ µ, 3 IND p + γ µ, 5 DR p + γ µ, 4 HIV + p , where DR p = 1 if patien t p is inf ected with a drug resistan t strain M . tub er- culosis and 0 otherwise, and HIV + p = 1 if patien t p is infected with HIV and 0 otherwise. P arameter estimates of this full mo del and their corresp onding 95% confidence in terv als are rep orted in th e full mo d el column s of T able 2 . The HIV infection and dr ug resistance app ear to hav e no effect on the bir th 16 C. R. DOSS ET AL. T able 2 R esults of the two lo g-line ar mo dels f or birth and de ath r ates of IS6110. The line age mo del includes only effe cts of M. tub er culosis line ages (EA, EU, I ND). The f ul l mo del c ombines the effe cts of line ages, H I V i nf e ction status ( HIV + ) and drug r esistanc e status (DR). The birth and de ath r ate m ultiplier estimates f or the EU l ine age ar e highlighte d in b old to indic ate that the c onfidenc e intervals for these p ar ameters ar e ab ove one Lineage mod e l F u ll model Coefficient MLE CIs MLE CIs EA birth rate, exp( γ λ, 1 ) 0 . 011 (0.003, 0.034) 0 . 012 (0.006, 0.025) EU multiplier, exp( γ λ, 2 ) 2 . 63 (0.689, 10.0) 3.2 (1.1, 9.4) IND multiplier, exp( γ λ, 3 ) 1 . 40 (0.229, 8.53) 1 . 7 (0.29, 9.7) DR multiplier, exp( γ λ, 4 ) – – 0 . 88 (0.36, 2.1) HIV + multipli er, exp( γ λ, 5 ) – – 0 . 61 (0.28, 1.3) EA death rate, exp( γ µ, 1 ) 0 . 00 4 (0.0005 , 0.028) 0 . 004 (0.000 5, 0.031) EU multiplier, exp( γ µ, 2 ) 9.32 (1.19, 72.8) 11 (1.2, 114) IND multiplier, exp( γ µ, 3 ) 5 . 40 (0.55 3, 52.6) 6 . 2 (0. 36, 1.1) DR multiplier, exp( γ µ, 4 ) – – 1 . 1 (0.52, 2.3) HIV + multipli er, exp( γ µ, 5 ) – – 0 . 64 (0.36 , 1.1) and d eath rates of IS 6110 transp oson. IS 6110 cop y num b er v ariatio n ma y ha ve an impact on functions of neigh b oring genes in the M . tub er culosis genome [Alonso et al. ( 2011 )]. Therefore, IS 6110 cop y num b er can p oten- tially int eract w ith other M. tub er culosis phenot yp es, su c h as dru g resistance and adaptation to HIV and anti viral treatmen t, with the help of selectio n [McEv o y et al. ( 2007 )]. Ho wev er, w e do not exp ect to see asso ciation b et w een IS 6110 copy num b er and M. tub er culosis ph enot yp es within one p atien t b e- cause selection is unlik ely to play a r ole on suc h a short time scale. Hence, w e view our estimated small effects of HIV in fection and drug resistance on IS 6110 cop y num b er as biologically p lausible. The EU lineage effect on the death rate remains s tatistica lly signifi can t ev en after con tr olling for th e t w o additional co v ariates. In terestingly , the EU lineage effect on the birth rate also b ecomes statisticall y significan t in the fu ll m o del. Effect sizes for b oth birth and death r ates incr ease and the confidence in terv als include larger v alues in the full mo del o v er the lineage-only mo del. This indicates that the full m o d el tend s to find more differences in r ates b et ween the lineages than the lineage-only mo del d o es. While more data are certainly needed to confirm th at EU lineage b irth rate effect is not 1, the full mo del ma y b e capturing information the s impler lineage-only mo d el do es not, whic h , in the face of limited data, is v aluable. F or pr actical considerations, th e fact that our most parameter r ich fu ll mo del results in significan t effects of EU lineage on IS 6110 birth and death rates su ggests that M. tub er c u losis lin- eage has to b e tak en into consideration when IS 6110 genot yp e data are used to unco v er th e h istory of M. tub er culosis transmission. BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 17 Fig. 3. L ow- vs high-c ount genotyp e analysis. Histo gr ams of simulate d numb ers of inter- vals and sums of interval lengths ar e plotte d for intervals with start ing values less than six and gr e ater or e qual to six. The vertic al dashe d lines i ndi c ate the observe d values of the four statistics. 4.3.2. IS6110 c ounts. The initial num b er of IS 6110 elemen ts is a p oten- tial confound er in our analysis b ecause patients infected with Eu r o-American and East-Asian differ drastically in the num b er of IS 6110 elemen ts at the b eginning of the obs erv ation p erio d . The isolate s f rom the Euro-American lineage hav e b etw een 2 and 17 IS 6110 elemen ts, with 41 out of 109 patien ts ha ving th e first r ecorded IS 6110 count less th an 6, wh ile IS 6110 counts v ary b et ween 6 and 22 for the East-Asian isolates. W arr en et al. ( 2002 ) suggest that I S 6110 genot yp es with few er than six elemen ts h a ve a ve ry lo w rate of c hange, b ecause in their data cases with n o observed c hanges in the geno- t yp e are dominated by suc h low-co unt genot yp es. How ev er, our birth–death mo del ve ry well predicts the conclusion of W arren et al. ( 2002 ) that lo w- coun t genot yp es ev olv e slo w er than high-count genot yp es. T o demonstrate this, we simulate 1000 data sets u sing our global birth and death rates and observ ed initial IS 6110 coun ts for eac h patien t. W e record th e n um b er of in terv als with equal s tarting and end ing v alues less than six, n 0 ,< 6 , and equal starting and en d ing v alues great er or equal to six, n 0 , ≥ 6 . W e also recorded the length su m of b oth kinds of in terv als: t 0 ,< 6 and t 0 , ≥ 6 . In our data, n obs 0 ,< 6 = 53 and n obs 0 , ≥ 6 = 171 with n obs 0 ,< 6 /t obs 0 ,< 6 = 4 . 6 > 2 . 8 = n obs 0 , ≥ 6 /t obs 0 , ≥ 6 , in agreement with W arr en et al.’s ( 2002 ) analysis. Histograms of sim u lated v alues of the four statistics, n 0 ,< 6 , n 0 , ≥ 6 , t 0 ,< 6 and t 0 , ≥ 6 , sho w n in Figure 3 , demonstrate that our birth–death mo del replicates well the obs er ved dy- namics of lo w -count and high-count IS 6110 genot yp es. W e conclude th at our d ata d o not pro vid e evidence th at ev olutionary dynamics of low-c oun t genot yp es d iffer fr om high-count genot yp e d y n amics. T herefore, it is un- lik ely th at a high p ercen tage of low-c oun t genot yp es in the Euro-American lineage isolates causes our estimated discrepancy b et w een death rates of Euro-American and East-Asian M. tub er culosis lineages. 18 C. R. DOSS ET AL. 5. Discussion. In this pap er w e present a n o ve l EM algorithm for fit- ting b irth–death pr o cesses to panel data. W e allo w logarithms of birth and death rates to b e linear com binations of individual-lev el co v ariates. Suc h birth–death m o dels with co v ariates sh are analogy with cov ariate-dep endent CTMC mo d els on fin ite state spaces—a w idely used class of mo dels in med- ical statistics [K albfl eisc h and La wless ( 1985 )]. T o our kno wledge, there is no established and well teste d metho d for fitting birth–death pro cesses, con- sidered in this p ap er, to panel data. W e hop e that by filling this void with our new EM algorithm, accompanied b y an op en-source R pack age DOBAD (a v ailable at http://c ran.r- project.o rg ), we will stim ulate statistical ap- plications of b ir th–death pro cesses, at least in the context of panel data. W e illus tr ate the ap p licabilit y of b irth–death mo dels by an alyzing th e evo - lutionary dynamics of the IS 6110 transp oson—an imp ortan t genetic marker that serves as a genetic signature of the M. tub er culosis b acterium. By b uild- ing realistic mo dels f or IS 6110 dynamics, we u n co v er differences in IS 6110 birth and death rates among ma j or lineages of M. tub er culosis , while con- trolling for other clinical co v ariates. This n o ve l r esult is imp ortant b ecause IS 6110 cop y n u m b er is u sed as a genetic marker to create DNA fi n gerprint s of M. tub er culosis usin g the restriction fragmen t length p olymorph ism tec h- nology [v an Em b den et al. ( 1993 ), Kato-Maeda, Metcalfe and Flores ( 2011 )]. Strains that ha ve th e same IS 6110 coun ts and in wh ic h the IS 6110 elemen t is lo cated in DNA fragmen ts of similar size are considered identica l. When suc h identical strains are found in communit y-based s tudies, the strains are clustered and patien ts carryin g these s tr ains are inferr ed to b elong to the same M. tub er culosis transmission chai n [K ato-Ma ed a, Metcalfe and Flores ( 2011 )]. Ho wev er, if some M . tub er c ulosis lineages ev olv e at muc h slo wer rates than others, as we disco ver in our analysis, then u sing the same n otion of similarity b et ween IS 6110 coun ts for these slo w-ev olving lineages could b e highly misleading. Therefore, we suggest that when u sing I S 6110 genot yp es, M. tub er culosis lineage effect should b e included explicitly in statistical p ro- to cols of estimating tub erculosis epid emiologi cal clusters. Although in our M. tub er culosis fingerpr in ting example we do not consider the p ossib ility of immigration, we include immigration in our metho dological dev elopments. More sp ecifically , our EM algorithm and the accompan ying soft ware pac k age allo w for immigration to o ccur at a rate prop ortional to the birth rate. W e ha ve t wo reasons f or includ ing this generalization. First, this limited f orm of imm igration complicates neither our mathematical devel op- men ts nor computational tractabilit y of the EM algorithm. S econd, incor- p orating immigration make s our EM algorithm more transferable to other domains of app lication of b irth–death pr o cesses. F or example, our metho d- ologic al deve lopmen ts directly apply to mo deling the evol ution of insertions and d eletions in m olecular sequences, wh ere immigration is n eeded to pre- v ent molecular sequences cont racting to length zero [Th orne, K ishino and BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 19 F elsenstein ( 1991 ), Holmes ( 2005 )]. Moreo v er, as w e sho w in App endix C, for this particular app licatio n, the E-step of our EM algorithm is a v ailable in closed form, eliminating the need for numerical in tegration [Doss et al. ( 2013 )]. Another example of p oten tial transferabilit y of our EM algorithm is for hidd en death-immigration mo dels for r ecurren t medical cond itions, su c h as that considered by Crespi, Cumb erland and Blo we r ( 2005 ). Although our EM algorithm d o es not apply dir ectly to the app lication these authors con- sider, b ecause the states of the immigration-death p ro cess are only partially observ ed at d iscrete time p oin ts, our mathematical results remain usefu l here. More sp ecifically , on e can use our mathematical dev elopments in th e con text of con tinuous-time hidden Marko v mo dels [Rob erts and Ephraim ( 2008 )] in order to dev elop an EM algorithm, akin to a classical Baum–W elc h algorithm [Baum et al. ( 1970 )]. As in the aforemen tioned insertion-deletion mo del, App endix C d emonstrates that the exp ectat ions of complete data sufficien t statistics for the death-immigration mo del are av ailable in closed form [Doss et al. ( 2013 )]. W e note that b ecause our Theorem 1 applies to general linear BDI mo dels, we are able to u se this th eorem to stu d y p rop er- ties of a death-immigration mo del, which is not a BDRI mo del—the main fo cus of this man uscript. Finally , w e wo uld lik e to p oin t out that the generating fu nctions deriv ed in Theorem 1 are u s eful n ot only for dev eloping EM algorithms for birth–death mo dels, bu t also f or probabilistic c h aracteriza tion of birth–d eath tra jectories in general. F or example, we are not a ware of analytic f ormulae for exp ecta- tions of the sufficient statistics that do n ot in volv e the endin g state of the pro- cess at time t : E( N + t | X 0 = i ) , E( N − t | X 0 = i ) an d E( R + t | X 0 = i ). T hese ex- p ectations, useful for p rediction purp oses, arise analytically from the gener- ating fun ctions in Theorem 1 [e.g., E( N + t | X 0 = i ) = ∂ H i ( u, 1 , 0 , 1 , t ) /∂ u | u =1 ]. Ac kn owledgmen t. W e thank P eter Guttorp for stim u lating discussions and for p oin ting u s to the work of Golinelli ( 2000 ). SUPPLEMENT AR Y MA TERIAL F u rther mathematical details (DOI: 10.121 4/13-A OAS673SUPP ; .p df ). App end ices referenced in S ections 2 and 5 are av ailable in the supp lemen tary material [Doss et al. ( 2013 )]. REFERENCES Alonso, H. , Aguilo, J. I. , S amper, S. , C aminero, J. A. , Campos-Herre ro, M. I. , Gicquel, B. , B r osch, R. , Mar t ´ ın, C. and Ot al, I. (2011). Deciphering the role of IS 6110 in a highly transmissible Myc ob acterium tub er culosis Beijing strain, GC1237. T ub er culosis 91 117–12 6. Baum, L. E. , Petrie, T. , Soules, G. and Weiss, N. (1970). A maximization technique occurring in the statistical analysis of probabilistic functions of Mark o v chains. Ann. Math. Statist. 41 164–171. MR0287613 20 C. R. DOSS ET AL. Ca tt amanchi, A. , Hopewell, P. C. , Gonzalez, L. C. , Osmond, D. H. , M asae , Ka w a- mura, L. , Daley, C. L. and Jasmer, R. M . (2006). A 13-year molecular epidemiolo g- ical analysis of tu b erculosis in San F rancisco. The International Journal of T ub er culosis and Lung D i se ase 10 297–304. Crespi, C. M. , Cumberland, W. G. and Blo wer, S . (2005). A queueing model for chronic recurrent conditions und er panel observ ation. Biometrics 61 193–198. MR2135860 Dempster, A. P. , Laird , N. M. and Rubin, D. B. (1977). Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. So c. Ser. B Stat. Metho dol. 39 1–38. MR0501537 Dorman, K. S. , Si n sheimer, J. S. and Lange, K. (2004). In the garden of branching processes. SIAM R ev. 46 202–229 (electronic). MR2114452 Doss, C. R. , S uchard, M. A. , Holmes, I. , Ka to-Maeda, M. and Mi n in, V. N. (2013). Supplement to “Fitting birth–death pro cesses to pan el d ata with applications to bac- terial DNA fingerprinting.” DOI: 10.1214 /13-AO AS673SUPP . Gagneux, S . , DeRiemer, K. , V an , T. , Ka to-Maeda, M. , de Jong, B. C. , Nara y anan, S. , Nicol, M. , Ni emann, S. , Kremer, K. , Guti e rrez, M. C. , Hil ty, M. , Hopewell, P. C. and Small, P. M. (2006). V ariable host-path ogen com- patibilit y in Myc ob acterium tub er culosis . Pr o c. Natl. A c ad. Sci. USA 103 2869–2873. Gibson, G. J. and Rensha w, E. (1998). Estimating parameters in sto chastic compart- mental mo d els u sing Marko v chain metho ds. IMA Journal of Mathematics Applie d in Me dicine & Biolo gy 15 19–40. Golinelli, D. (2000). Bay esian inference in hidden stochastic p opulation pro cesses. Ph.D. thesis, Univ. W ashington, Seattle, W A. Guttorp, P. (1995). Sto chastic M o deling of Scientific Data . Chapman & Hall, London. MR1358359 Henrici, P. (1979). F ast Fourier met h ods in compu t ational complex analysis. SIAM Re v. 21 481–527. MR0545882 Holmes, I. (2005). Using evo lutionary exp ectation maximization to estimate indel rates. Bioinformatics 21 2294–2300. Holmes, I. and Rubin, G. M. (2002). A n exp ectation maximization algorithm for training hidden substitution mo dels. Journal of Mol e cular Biolo gy 317 753–764. Ja ckson, C . H. ( 2011). Multi-state mo dels for panel data: The msm pack age for R . Journal of Statistic al Softwar e 38 1–29. Jasmer, R. M. , Hahn , J. A. , S m all, P. M. , Daley, C. L. , Behr, M . A. , Moss, A. R. , Creasman, J. M. , Sche cter, G. F. , P az, E. A. and Hopewell, P. C. (1999). A molecular epidemiologic analysis of tub erculosis trends in S an Francisco, 1991 –1997. Ann als of Internal Me dicine 130 971–978. Kalbfleisch, J. D. and La wless, J. F. (1985). The analysis of panel data un der a Mark o v assumption. J. Amer . Statist. Asso c. 80 863–871. MR0819585 Karlin, S. and McGr e gor, J. (1958). Linear growth birth and death pro cesses. J. Math. Me ch. 7 643–662. MR0098435 Ka to-Maeda, M. , Metcalfe, J. Z. and Flores, L. (2011). Genotyping of Mycobac- terium tub erculosis: Ap plication in epidemiologic studies. F utur e Micr obiol . 6 203–216. Keiding, N . (1975). Maximum likeli ho od estimation in the b irth -and-death process. Ann. Statist. 3 363–372. MR0362773 Kendall, D. G. (1948). On the generalized “birth-and- death” p rocess. Ann. Math. Statist. 19 1–15. MR0024091 Lange, K. (1982). Calculation of th e equilibrium distribution for a d eleterious gene by the finite F ourier transform. Biometrics 38 79–86. BIR TH–DEA TH PR OCESSES FOR P ANEL DA T A 21 Lange, K. (1995). A gradient algorithm lo cally equ iv alent to the EM algorithm. J. R. Stat. So c. Ser. B Stat. Metho dol. 57 425–437. MR1323348 Louis, T. A. (1982). Finding the ob served information matrix when using t h e EM algo- rithm. J. R. Stat. So c. Ser. B Stat. Metho dol. 44 226–233 . MR0676213 McEv o y, C. R. E. , F almer, A. A. , v an Pittius, N. C. G. , Victor, T. C. , v an Helden, P. D . and W arren, R. M. (2007). The role of IS 6110 in the evolution of Myc ob acterium tub er culosis . T ub er culosis 87 393–404. Minin, V . N. and Suchard , M. A . (2008). Counting lab eled transitions in continuous- time Marko v mo dels of evolution. J. Math. B i ol. 56 391–412 . MR2358440 Nee, S. (2006). Birth–death mo dels in macro evolution. Annual R eview of Ec olo gy, Evo- lution, and Systematics 37 1–17. Ro ber ts, W. J. J. and Ephraim, Y. ( 2008). An EM algorithm for ion-channel current estimation. IEEE T r ans. Signal Pr o c ess. 56 26–33. MR2439810 Ro senberg, N. A. , Tsolaki, A. G. and T anaka, M. M. ( 2003). Estimating change rates of genetic markers u sing serial samples: Applications to the transp oson I S 6110 in Mycobacterium tub ercu losis. The or etic al Population Bi olo gy 63 347–363. Sehl, M. , Zhou, H. , Sinshei mer, J. S . and Lange, K. L. (2011). Extinction mo dels for cancer stem cell therapy. Math. B i osci. 234 132–1 46. MR2907020 Small, P. M. , Hopewell, P. C. , Sin g h , S. P. , P az, A. , P arsonne t, J. , Ruston, D. C. , Schecter, G . F. , Daley, C. L. and Schoolnik, G. K. (1994). The epidemiology of tub erculosis in San Francisco. A p opulation-based study using conv entional and molecular metho ds. New England Journal of Me di cine 330 1703–1 709. Suchard, M. A . , Lange, K. and S insheimer, J. S. (2008). Efficiency of protein pro duc- tion from mRN A. J. Stat. The ory Pr act. 2 173–182. MR2524460 T anaka, M. M. and Rosenber g, N. A. (2001). Optimal estimation of transp osition rates of insertion sequences for molecular epid emiology. Stat. Me d. 20 2409–2420. Thorne, J. L. , Kishino, H. and Fe lsen ste i n, J. (1991). An evolutionary model for maximum lik eliho od alignment of DNA sequences. J. Mol. Evol. 33 114–124. v an Embden, J. D. , Ca ve, M. D. , Cra wf ord, J. T. , Dale, J. W. , Eisenach, K. D. , Gicquel, B. , Hermans, P. , Mar tin, C . , McAdam, R. , Shinni ck, T. M. et al. (1993). Strain id entification of Mycobacterium tub erculosis by DNA fingerprinting: Recommen- dations for a standardized met h odology . J. Cl in. Micr obiol. 31 406–409. W arren, R. M. , v an der Spuy, G. D. , Richardson, M. , Be yers, N . , B oo ysen , C. , Behr, M. A. and v an Helden , P. D. ( 2002). Evolution of the I S6110-based restric- tion fragment length p olymorphism p attern during the transmission of Mycobacterium tub erculosis. J. Clin. Micr obiol. 40 1277–12 82. C. R. Doss V. N. Minin Dep ar tment of St a tistics University of W ashington Sea ttle, W ashing ton 98195 USA E-mail: cdoss@u w. edu vminin@uw.edu M. A. Suchard Dep ar tments of Bioma thema tics, Biost a tistics and Human Genetics University of California, Los Angeles Los Angeles, California 9009 5 USA E-mail: msuc hard@ucla.edu I. Holmes Dep ar tment of Bioengineering and Biophysics Gradua te Group University of California, Berkeley Berkeley, California 9 4720 USA E-mail: ihh@ber k eley .edu M. Ka to-Maeda Dep ar tment of Medicine San Francisco General Hospit al University of California, San Francisco San Francisco, California 94143 USA E-mail: Midori.Kato-Maeda@ucsf.edu
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment