Ranking relations using analogies in biological and information networks
Analogical reasoning depends fundamentally on the ability to learn and generalize about relations between objects. We develop an approach to relational learning which, given a set of pairs of objects $\mathbf{S}=\{A^{(1)}:B^{(1)},A^{(2)}:B^{(2)},\ldo…
Authors: Ricardo Silva, Katherine Heller, Zoubin Ghahramani
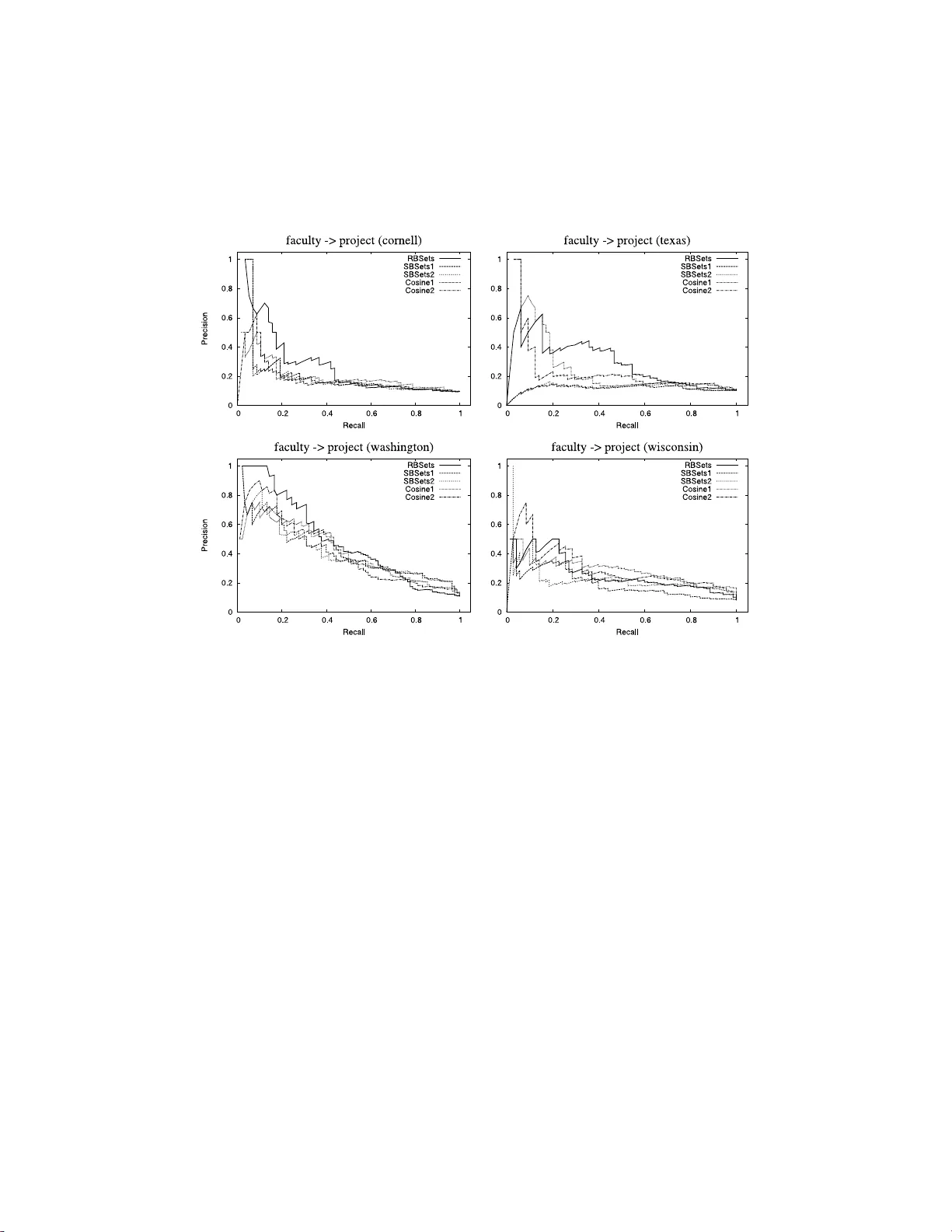
The Annals of Applie d Statistics 2010, V ol. 4, No. 2, 615–644 DOI: 10.1214 /09-A OAS321 c Institute of Mathematical Statistics , 2 010 RANKING RELA TIONS USING ANALOGIES IN BIOLOGICAL AND INF ORMA TION NE TW ORKS 1 By Ricardo Sil v a, Ka therine Heller, Zoubin Ghahraman i and Edo ard o M. Air old i University Co l le g e L ondon, University o f Cambridge, University o f Cambr idge and Harvar d Univ e rsity Analogical reasoning dep ends fundamentally on the abilit y to learn and generalize ab out relations b etw een ob jects. W e develop an approac h to relational learning which, given a set of p airs of ob jects S = { A (1) : B (1) , A (2) : B (2) , . . . , A ( N ) : B ( N ) } , measures how well oth er pairs A : B fit in with the set S . Ou r work addresses th e follo wing question: is the relation b etw een ob jects A and B analogous to th ose relations found in S ? Suc h qu estions are particularly relev ant in in- formation retriev al, where an inv estigator might wa nt to searc h for analogous pairs of ob jects that matc h the query set of interest. There are many w ays in whic h ob jects can b e related, making the task of measuring analogies very c h allenging. Ou r approach combines a sim- ilarit y measure on function spaces with Bay esian analysis t o pro duce a ranking. It req uires data contai ning features of the ob jects of inter- est and a link matrix sp ecifying whic h relationships exist; no further attributes of such relationships are necessary . W e illustrate the p o- tential of our metho d on tex t analysis and information netw orks. A n application on discov ering functional interactio ns b etw een pairs of proteins is discussed in detail, where w e sh ow that ou r approac h can w ork in p ractice even if a small set of protein p airs is provided. 1. Con tribu tion. Man y universit y admission exams, such as the Ameri- can Sc h olastic Assessmen t T est (SA T) and Graduate Record Exam (GRE), ha ve h istorically included a section on analogical reasoning. A protot ypical analogica l reasoning q u estion is as follo ws: doctor : hosp ital : (A) sports fa n : stadium Received May 2009; revised Nov ember 2009. 1 Supp orted in part by NSF Grant D MS-09-07009, by NIH Grant R01 GM096193, and by th e Gatsby Charitable F oundation. Key wor ds and phr ases. Netw ork analysis, Bay esian inference, v ariational approxima- tion, ran k ing, information retriev al, data integr ation, Sac char omyc es c er evisiae . This is a n e le ctronic repr int of the orig inal a rticle published by the Institute of Ma thematical Statistics in The Annals of A pplie d Statistics , 2010, V ol. 4, No. 2, 61 5 –644 . This reprint differs from the or iginal in pagina tion and typogra phic detail. 1 2 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI (B) cow : farm (C) professor : coll ege (D) criminal : jail (E) food : grocery st ore The examinee h as to answer whic h of the fi v e pairs b est matc hes the relation implicit in doctor : ho spital . Although all candidate pairs h a ve some t yp e of relation, pair prof essor : colle ge seems to b est fi t the n otion of ( pr ofession , plac e of work ), or the “w orks in” relation implicit b etw een do ctor and hospital. This problem is non trivial b ecause measuring the similarit y b etw een ob- jects directly is not an appropriate wa y of disco vering analogies, as exten- siv ely discussed in th e cognitiv e science literature. F or ins tance, the analogy b et w een an electron spinning around the nucleus of an atom and a planet or- biting aroun d the Sun is not justified b y isolated, nonrelational, comparisons of an electron to a planet, and of an atomic nucleus to the Sun [Gentner ( 1983 )]. Disco vering the underlyin g relationship b etw een the elemen ts of eac h pair is k ey in determining analogies. 1.1. Applic ations. This p ap er concerns p ractical pr oblems of data an al- ysis where analogies, implicitly or n ot, play a role. On e of our motiv ations comes from the bioPIXIE 2 pro ject [My ers et al. ( 2005 )]. bioPIXIE is a to ol for exploratory analysis of protein–protein in teractions. Proteins ha ve multi- ple f u nctional r oles in the cell, for example, regulating m etab olism and r egu- lating cell cycle, among others. A protein often assumes different fun ctional roles while in teracting w ith d ifferen t p roteins. When a molecular biologist exp erimenta lly observe s an interac tion b et wee n t wo proteins, for example, a binding eve n t of { P i , P j } , it migh t not b e clear wh ich function that particu- lar inte raction is con tribu ting to. T he bioPIXIE system allo ws a molecular biologist to inpu t a set S of p roteins that are b elieve d to h av e a particu- lar functional r ole in common, and generates a list of other proteins that are deduced to pla y the same role. Evidence for such pr ed ictions is pro- vided b y a v ariet y of sources, such as the expr ession lev els for the genes that encod e the p roteins of interest and their cellular lo calizatio n. Another imp ortant source of inform ation bioPIXIE tak es adv anta ge of is a matrix of relationships, in dicating which proteins interac t according to some b io- logica l criterion. Ho w ev er, we do not n ecessarily kn ow whic h interact ions corresp ond to which f u nctional roles. The application to pr otein interacti on n et wo rks that w e dev elop in Sec- tion 5 shares some of the f eatures and m otiv ations of bioPIXIE. Ho wev er, w e aim at providing more detailed information. Our input set S is a smal l 2 http://pix ie.prince ton.edu/p ixie/ . RANKI N G RELA TIONS USIN G A N ALOGIES 3 set of p airs of proteins th at are p ostulated to all pla y a common r ole, and w e wan t to r ank other p airs P i : P j according to how similar they are with resp ect to S . The goal is to automatically r eturn pairs that corresp ond to analogous interac tions. T o use an analogy itself to explain our p ro cedure, recall the SA T example that op ened this section. Th e pair of words doct or : hospit al present ed in the S A T question pla y the r ole of a protein–protein interactio n and is th e smallest p ossible case of S , that is, a single pair. Th e fiv e c hoices A – E in the SA T qu estion corresp ond to other observed protein–protein inte ractions w e wa nt to m atch with S , that is, other p ossib le pairs. Since m u ltiple v alid answ ers are p ossible, w e rank them according to a similarit y metric. In the application to protein inte ractions, in Section 5 , we p erform thousands of queries and we ev aluate the go o dness of the resulting rankings according to m u ltiple gold stand ard s, widely accepted by molecular and cellular biologists [Ash b urner et al. ( 2000 ); Kanehisa and Goto ( 2000 ); Mewes et al. ( 2004 )]. The general problem of in terest in this pap er is a pr actica l p roblem of in- formation r etrieval [Manning, Ragha v an and Sch¨ utze ( 2008 )] for exploratory data analysis: give n a qu ery set S of link ed pairs, which other pairs of ob jects in my relational database are link ed in a similar w a y? W e apply this analysis to cases where it is not kno w n ho w to explicitly describ e th e differen t classes of relations, b ut go o d mo dels to pr ed ict the existenc e of relationships are a v ailable. In S ection 4 we consider an app lication to information retriev al in text do cuments for illustrativ e purp oses. Give n a s et of pairs of w eb pages whic h are related by s ome hyp er lin k, w e wo uld lik e to fin d other p airs of pages that are link ed in a similar wa y . In in f ormation net work settings, the prop osed metho d could b e useful, for instance, to answer qu eries for en- cyclop edia p ages relating scien tists and their ma jor disco ve ries, to searc h for analog ous concepts, or to ident ify the absence of analogous concepts, in Wikip edia. F rom an ev aluation p ersp ectiv e, this app lication domain pro- vides an example where large scale ev aluation is more straightfo rw ard than in the biological setting. In this pap er we in tro duce a metho d for ranking relations based on the Ba y esian similarit y criterion u nderlying Bayesian sets , a metho d originally prop osed b y Ghahramani and Heller ( 2005 ) and r eview ed in S ection 2 . In con trast to Ba ye s ian sets, ho wev er, our metho d is tailored to dr a wing analo- gies b et ween pairs of ob jects. W e also pr o vide sup p lemen tary material with a Jav a implemen tation of our metho d , and instructions on ho w to rebuild the exp erimen ts [Silv a et al. ( 2010 )]. 1.2. R elate d work. T o giv e an idea of the t yp e of data w hic h our metho d is useful for analyzing, consider the metho ds of T urney and Littman ( 2005 ) for automatically solving S A T problems. Their analysis is b ased on a large corpus of do cuments extracted from the W orld Wide W eb. Relatio ns b e- 4 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI t wee n tw o wo r ds W i and W j are charac terized by their join t co-o curren ce with other relev ant words (such as particular prep ositions) within a small windo w of text. This defines a set of features for eac h W i : W j relationship, whic h can then b e compared to other pairs of w ord s using some notion of similarit y . Unlik e in this application, ho w ever, there are often no (or very few) explicit features for the relationships of in terest. Instead we need a metho d for defining similarities using features of the ob jects in eac h rela- tionship, wh ile at th e same time a vo iding the mistake of d irectly comparing ob jects ins tead of relations. One of the earliest appr oac hes f or d etermining analogica l similarit y was in tro duced b y Rum elhart and Abrahamson ( 1973 ). In their pap er, one is initially giv en a set of p airwise distances b et ween ob jects (sa y , by the su b- jectiv e j udgemen t of a group of p eople). Su c h distances are used to emb ed the given ob jects in a latent space via a multidimensional scaling appr oac h. A related pair A : B is then r epresen ted as a vecto r connecting A and B in the laten t space. Its s imilarit y w ith r esp ect to another pair C : D is defin ed b y comparing the d ir ection and magnitude of the corresp onding v ectors. Our approac h is pr obabilistic instead of geometrical, and op erates directly on the ob ject features instead of pairwise d istances. W e will f o cus solely on ranking p airwise relations. Th e idea can b e ex- tended to more complex relations, but we will not pursu e this here. Ou r approac h is d escrib ed in detail in S ection 3 . Finally , the p robabilistic, geometrical and logical approac h es applied to analogica l r easoning p r oblems can b e seen as a t yp e of relational data anal- ysis [D ˇ zeroski an d La vr a ˇ c ( 2001 ); Getoor and T ask ar ( 2007 )]. In p articular, analogica l reasoning is a p art of the more general problem of generating la- ten t relationships from relational data. Seve ral app r oac hes for this problem are discuss ed in Section 6 . T o the b est of our knowledge, h o wev er , m ost ana- logica l reasoning applications are interesting p ro ofs of concept that tac k le am bitious pr oblems suc h as plannin g [V eloso and Carb onell ( 1993 )], or are motiv ated as mo dels of cognition [Gentner ( 1983 )]. Our goal is to create an off-the-shelf m etho d for practical exploratory data analysis. 2. A review of probabilistic information retriev al an d the Bay esian sets metho d. The goal of information r etriev al is to provide d ata p oints (e.g., text do cuments, images, m ed ical records) that are judged to b e relev ant to a particular query . Queries can b e defin ed in a v ariet y of wa ys and, in general, they do n ot sp ecify exactly w hic h records should b e presente d. In p ractice, retriev al metho ds r ank data p oints according to some measure of similarit y with resp ect to the qu er y [Manning, Ragha v an and Sc h ¨ utze ( 2008 )]. Al- though queries can, in pr actice, consist of an y piece of information, for the purp oses of th is p ap er we will assume that q u eries are sets of ob jects of the same type w e w ant to retriev e. RANKI N G RELA TIONS USIN G A N ALOGIES 5 Probabilities can b e exploited as a measure of similarit y . W e will briefly review one standard probabilistic framew ork for information retriev al [Man- ning, Raghav an and Sc h ¨ utze ( 2008 ), Chapter 11]. Let R b e a binary r andom v ariable rep r esen ting whether an arbitrary d ata p oin t X is “relev an t” f or a giv en query set S ( R = 1) or not ( R = 0). Let P ( ·|· ) b e a generic probability mass function or densit y function, with its m eanin g giv en by the cont ext. P oints are rank ed in decreasing order by the follo w in g criterion: P ( R = 1 | X, S ) P ( R = 0 | X, S ) = P ( R = 1 | S ) P ( R = 0 | S ) P ( X | R = 1 , S ) P ( X | R = 0 , S ) , whic h is equiv alent to r an k in g p oints by the expression log P ( X | R = 1 , S ) − log P ( X | R = 0 , S ) . (2.1) The challenge is to define w hat form P ( X | R = r , S ) should assume. It is not pr actical to collect lab eled data in adv an ce wh ic h, for ev ery p ossible class of queries, will give an estimate for P ( R = 1 | X , S ): in general, one cannot anticipat e w hic h classes of q u eries will exist. Instead, a v ariet y of approac hes hav e b een dev elop ed in the literature in order to d efine a su itable instan tiation of ( 2.1 ). These include a metho d th at b uilds a classifier on-the- fly using S as elements of the p ositiv e class R = 1, and a r andom subset of data p oin ts as the negativ e class R = 0 [e.g., T u rney ( 2008b )]. The Ba y esian sets metho d of Ghahramani and Heller ( 2005 ) is a state- of-the-art pr obabilistic metho d for r an k in g ob jects, partially inspir ed b y Ba y esian p s yc hological m o dels of generalization in human cognition [T enen- baum and Griffiths ( 2001 )]. In this setup the ev ent “ R = 1 ” is equated with the ev ent that X and the elemen ts of S are i.i.d. p oin ts generated b y the same mo d el. The even t “ R = 0” is the ev ent by whic h X and S are generated b y t wo indep end en t mo dels: one for X and another for S . T he parameters of all mo d els are r andom v ariables that ha v e b een integ r ated out, with fixed (and common) h yp erparameters. Th e result is the in stan tiation of ( 2.1 ) as log P ( X | S ) − log P ( X ) = log P ( X , S ) P ( X ) P ( S ) , (2.2) the Ba yesian sets sc or e function by whic h we ran k p oint s X giv en a qu ery S . T he right-hand s id e w as rearranged to provide a more intuitiv e graph ical mo del, shown in Figure 1 . F rom this graph ical mo del in terpretation we can see that th e score fun ction is a Ba y es factor comparing tw o mo d els [Kass and Raftery ( 1995 )]. In the next section we describ e h o w the Ba yesia n sets metho d can b e adapted to defin e analogical similarit y in the biological and information net works settings w e consider, and why su c h mo difications are necessary . 3. A m o del of Ba yesian analogical similarit y for relations. T o d efine an analogy is to define a measure of s im ilarity b etw een structures of relate d ob jects. In our s etting, we need to measure th e sim ilarity b etw een p airs of 6 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI Fig. 1. In or der to sc or e how wel l an arbitr ary element X fits in with query set S = { X 1 , X 2 , . . . , X q } , the Bayesian sets metho dolo gy c omp ar es the mar ginal likeliho o d of the m o del in (a) , P ( X, S ) , against the mo del in (b) , P ( X ) P ( S ) . In (a) , the r andom p ar ameter ve ctor Θ i s given a prior define d by the (fixe d) hyp erp ar ameter α . The same (la- tent) p ar ameter ve ctor is shar e d by the query set and the new p oint. In (b) , the p ar ameter ve ctor Θ that gener ates X i s di ffer ent fr om the one that gener ates the query set. ob jects. The k ey asp ect that distinguishes our approac h from others is that w e fo cus on the similarit y b et wee n functions that map p airs to links, r ather than fo cus in g on the similarit y b etw een the fe atur es of ob jects in a candid ate pair and the features of ob jects in the query pairs. As an illustration, consider an analogic al reasoning question f r om a S A T - lik e exam wh ere for a giv en p air (sa y , water : river ) we h a ve to c h o ose, out of 5 pairs, the one that b est matc h es the typ e of relation imp licit in suc h a “query .” In th is case, it is reasonable to sa y c ar : highway would b e a b etter matc h than (the somewhat nonsens ical) so da : o c e an , sin ce cars fl o w on a high wa y , and so do es w ater in a rive r. Notice that if w e w ere to measure the similarit y b et ween obje c ts in stead of r elations , so da : o c e an would b e a muc h closer p air, since so da is similar to water , and o c e an is similar to river . Nev ertheless, it is legitimate to infer relational similarit y from individual ob ject features, as summarized by Gen tner and Medina ( 1998 ) in their “kind w orld h yp othesis.” What is needed is a mec h anism b y whic h ob ject features should b e wei gh ted in a particular relational similarity problem. W e p ostu- late that, in analogical reasoning, similarit y b etw een features of ob j ects is only meaningful to the extent by wh ic h suc h features are u seful to p r edict the existence of the relationships. Our app roac h can b e describ ed as follo ws . L et A and B represent ob- ject spaces. T o sa y that an int eraction A : B is analogo us to S = { A (1) : B (1) , A (2) : B (2) , . . . , A ( N ) : B ( N ) } amounts to implicitly definin g a measure of sim- ilarit y b et ween the pair A : B and th e set of pairs S , where eac h qu ery item A ( k ) : B ( k ) corresp onds to some pair A i : B j . How ev er, this similarit y is not directly derived from the similarit y of the in f ormation con tained in the d is- tribution of ob jects themselv es, { A i } ⊂ A , { B i } ⊂ B . Rather, the similarit y RANKI N G RELA TIONS USIN G A N ALOGIES 7 b et w een A : B and the set S is defin ed in terms of th e similarit y of the func- tions map p ing the pairs as b eing linke d . Eac h p ossible fu nction captures a differen t p ossible relationship b et ween the ob jects in the pair. Ba yes ian anal ogical reaso ning formula tion. Consider a sp ace of laten t fun ctions in A × B → { 0 , 1 } . Assume that A and B are t wo ob jects classified as linked by some unknown function f ( A, B ) , that is, f ( A, B ) = 1. W e wan t to quanti fy h o w similar th e function f ( A, B ) is to the fu n ction g ( · , · ), which classifies all pairs ( A i , B j ) ∈ S as b eing link ed, that is, w here g ( A i , B j ) = 1. The similarit y s h ould dep end on the observ ations { S , A, B } and our prior distr ibution o ve r f ( · , · ) and g ( · , · ). F u n ctions f ( · ) and g ( · ) are u nobserve d, hence the need for a prior that will b e used to int egrate o v er the function s p ace. Our similarit y metric will b e d efi ned using Ba y es factors, as explained next. 3.1. Analo gy in function sp ac es via lo gistic r e gr ession. F or simplicit y , w e w ill consider a family of latent functions that is parameterized b y a finite-dimensional ve ctor: the logistic regression fun ction with multiv ariate Gaussian p riors for its parameters. F or a particular pair ( A i ∈ A , B j ∈ B ), let X ij = [Φ 1 ( A i , B j ) Φ 2 ( A i , B j ) · · · Φ K ( A i , B j )] T b e a p oint on a feature sp ace defin ed by the mapp ing Φ : A × B → ℜ K . This feature space mapping compu tes a K -dimensional v ector of attributes of the p air that ma y b e p oten tially relev an t to pred icting the relation b et ween the ob jects in the pair. Let L ij ∈ { 0 , 1 } b e an indicator of the existence of a link or relation b et wee n A i and B j in the d atabase. L et Θ = [ θ 1 , . . . , θ K ] T b e th e p arameter v ector for our logistic regression mo del suc h that P ( L ij = 1 | X ij , Θ) = logistic(Θ T X ij ) , (3.1) where logistic( x ) = (1 + e − x ) − 1 . W e no w apply the same score fu nction underlying the Ba yesian sets metho dology explained in Section 2 . Ho w ever, instead of comparing ob j ects b y marginalizing o ver the parameters of their feature distributions, we com- pare functions for lin k in d icators by marginalizing o ver the p arameters of the fu nctions. Let L S b e the v ector of link indicators for S : in fact, eac h L ∈ L S has the v alue L = 1, ind icating that ev ery pair of ob jects in S is linked. Consider the follo w ing Ba ye s factor: P ( L ij = 1 , L S = 1 | X ij , S ) P ( L ij = 1 | X ij ) P ( L S = 1 | S ) . (3.2) 8 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI Fig. 2. The sc or e of a new data p oint { A i , B j } is given by the Bayes factor that c omp ar es mo dels (a) and ( b) . No de α r epr esents the hyp erp ar ameters f or Θ . In (a) , the gener ative mo del is the same for b oth the new p oint and the query set r epr esente d i n the r e ctangle. Notic e that our c onditioning set S of p airs mi ght c ontain r ep e ate d i nstanc es of a same p oint, that is, some A or B might app e ar multiple times in differ ent r elations, as il lustr ate d by no des with multiple outgoing e dges. In (b) , the new p oi nt and the query set do not shar e the same p ar ameters. This is an adaptation of equation ( 2.2 ) wh er e r elev ance is defin ed n o w b y whether L ij and L S w ere generated b y th e same mo del, for fixed { X ij , S } . In one sense, this is a discriminativ e Ba yesia n sets mo d el, where w e pre- dict links instead of mo deling j oint ob ject features. Since w e are integrat ing out Θ , a prior for this parameter vec tor is needed. T he graph ical mo d els corresp ondin g to this Ba y es factor are illustrated in Figure 2 . Th us, eac h pair ( A i , B j ) is ev aluated with resp ect to a query set S b y the score function giv en in ( 3.2 ), rewritten after taking a logarithm and droppin g constan ts as score( A i , B j ) = log P ( L ij = 1 | X ij , S , L S = 1) (3.3) − log P ( L ij = 1 | X ij ) . The exact d etails of our pro cedur e are as follo ws. W e are giv en a relational database ( D A , D B , L AB ). Dataset D A ( D B ) is a s ample of ob jects of t yp e A ( B ). Relationship table L AB is a binary matrix mo deled as generated from a logistic regression mo del of link existence. A qu ery pro ceeds according to the follo w ing steps: 1. the user selects a set of pairs S th at are lin k ed in the database, w here the p airs in S are assumed to ha ve some relation of interest; 2. the sys tem p erf orms Ba yesia n inference to obtain the corresp on d ing p os- terior distrib ution for Θ, P (Θ | S , L S ), giv en a Gaussian p rior P (Θ) ; 3. the system iterates thr ough all link ed pairs, compu ting the follo wing for eac h pair: P ( L ij = 1 | X ij , S , L S = 1) = Z P ( L ij = 1 | X ij , Θ) P (Θ | S , L S = 1) d Θ . RANKI N G RELA TIONS USIN G A N ALOGIES 9 Fig. 3. Gener al fr amework of the pr o c e dur e: first, a “prior” over p ar ameters Θ for a link classifier is define d empiric al ly using li nke d and unl inke d p airs of p oints (the dashe d e dges indic ate that cr e ating a prior empiric al ly is optional, but in pr actic e we r ely on this metho d). Given a query set S of linke d p airs of inter est, the system c omputes the pr e dic- tive likeliho o d of e ach l inke d p air D ( i ) ∈ D + and c omp ar es it to the c onditional pr e di ctive likeliho o d, given the query. This defines a me asur e of simil arity with r esp e ct to S by which al l p airs in D + ar e sorte d. P ( L ij = 1 | X ij ) is similarly computed by integ rating o ve r P (Θ). All pairs are present ed in decreasing order according to the score in equation ( 3.3 ). The in tegral pr esen ted ab o v e do es not hav e a closed form u la. Because computing the in tegrals b y a Mon te Carlo metho d for a large n umb er of pairs would b e un reasonable, w e use a v ariational appro ximation [Jord an et al. ( 1999 ); Airoldi ( 2007 )]. Figure 3 presents a summary of the app roac h. The suggested setup scales as O ( K 3 ) with the f eature space d im en sion, due to the matrix inv ersions necessary for (v ariational) Ba ye sian logistic regression [Jaakk ola an d Jordan ( 2000 )]. A less p recise appro ximation to P (Θ | S , L S ) can b e imp osed if the d imensionalit y of Θ is to o high. Ho w ever, it is imp ortan t to p oin t ou t that once the initial integral P (Θ | S , L S ) is appro ximated, eac h s core function can b e computed at a cost of O ( K 2 ). Our analogical reasoning formulation is a relational mo del in that it mo d - els the presence and absence of int eractions b etw een ob j ects. By conditioning on the link indicators, the similarit y score b et w een A : B and C : D is alw ays a function of pairs ( A, B ) and ( C , D ) that is n ot in general decomp osable as similarities b et ween A and C , and B and D . 3.2. Comp arison with Bayesian sets and sto chastic b lo ck mo dels. T he mo del presented in Figure 2 is a c onditiona l in dep enden ce mo del for rela- tionship indicators, that is, give n ob ject features and p arameters, the entries of L D are indep endent. Ho we v er, the entries in L D are in general mar g i nal ly 10 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI dep end ent. Since this is a mo del of relationships give n ob ject attributes, w e call the mo del introd uced here th e r e lational Bayesian sets mo del . Our approac h h as some similarit y to the so-called sto chastic blo ck mo dels . These m o dels w ere develo p ed four decades ago in the net work literature to quan tify the notion of “structural equiv alence” by means of blo c ks no des that instan tiate s imilar connectivit y patterns [Lorrain and White ( 1971 ); Holland and Leinhard t ( 1975 )]. Mo dern sto chastic b lo c k mo del approac hes, in statistics and mac hine learnin g, bu ild on these seminal wo rks b y in tro- ducing the disco very of th e blo ck stru cture as part of the mo d el search strat- egy [Fien b erg, Mey er and W asserman ( 1985 ); No wic k i and Snijd er s ( 2001 ); Kemp et al. ( 2006 ); Xu et al. ( 2006 ); Airoldi et al. ( 2005 , 2008 ); Hoff ( 2008 )]. The observed features in our approac h, X ij , effectiv ely p la y the same role as th e laten t ind icators in sto chastic blo ck mo dels. 3 Since X ij is observ ed, there is no need to int egrate ov er th e feature sp ace to obtain the p osterior distribution of Θ. T h is compu tational efficiency is particularly r elev ant in information retriev al and exploratory data an alysis, where users exp ect a relativ ely short resp onse time. As an alternativ e to our relational Ba y esian sets appr oac h, consider the follo wing direct mo dification of the standard Bay esian sets formulatio n to this problem: m erge th e data sets D A and D B in to a single data set, cre- ating for eac h pair ( A i , B j ) a ro w in the database with an extra b inary indicator of relationship existence. Create a joint mo d el for pairs by u s ing the marginal mo dels for A and B and treating different ro ws as b eing in- dep end ent. Th is ignores the fact that the r esulting merged data p oin ts are not really i.i.d. u nder such a mo del, b ecause th e same ob ject migh t app ear in m u ltiple relations [D ˇ zeroski and Lavra ˇ c ( 2001 )]. The mo del also fails to capture the dep end ency b etw een A i and B j that arises fr om conditioning on L ij , even if A i and B j are marginally in d ep endent. Nevertheless, heuris- tically th is approac h can s ometimes p r o duce go o d resu lts, and for sev eral t yp es of p r obabilit y families it is v ery computationally efficient. W e ev aluate it in Section 4 . 3.3. Choic e of fe atur es and r elational discrimination. Ou r setup assumes that the feature sp ace Φ pr o vides a reasonable classifier to predict the ex- istence of link s . Useful predictiv e f eatures can also b e generated automati- cally with a v ariet y of algorithms [e.g., the “structural logistic r egression” of P op escul and Ungar ( 2003 )]. S ee also D ˇ zeroski and La vraˇ c ( 2001 ). Jensen and Neville ( 2002 ) d iscuss s hortcomings of metho d s for automated feature selection in relational classification. 3 In a sto chastic blo ck model, typically each ob ject has a single feature η indicating membership to some latent class. F or a p air A i , B j , the corresp onding feature vector X ij w ou ld b e ( η A , η B ). RANKI N G RELA TIONS USIN G A N ALOGIES 11 W e also assum e feature spaces are the same for all p ossible com binations of ob jects. This allo ws for comparisons b et ween, for example, cells from d if- feren t sp ecies, or w eb pages from differen t web domains, as long as f eatures are generated b y the same fun ction Φ( · , · ). In general, we wo u ld like to relax this requ ir emen t, but for the problem to b e w ell-defin ed, features from the differen t spaces must b e related s omeho w. A h ierarc hical Ba y esian form u - lation for linking different feature sp aces is on e p ossibility wh ich might b e treated in a fu ture w ork . 3.4. Priors. Th e c h oice of prior is based on the observed d ata, in a w ay that is equiv alen t to the c h oice of priors used in the original formulati on of Ba y esian sets [Ghahramani and Heller ( 2005 )]. Let b Θ b e the maxim u m like li- ho o d estimator of Θ using the r elational database ( D A , D B , L AB ). Since th e n u m b er of p ossible pairs gro ws at a quadratic r ate with the num b er of ob- jects, w e do not use the whole d atabase for maximum likel iho o d estimation. Instead, to get b Θ, we use all linked pairs as mem b ers of the “p ositive ” class ( L = 1 ), and su bsample unlinked pairs as memb ers of the “negativ e” class ( L = 0). W e subsample by sampling eac h ob ject uniformly at random from the r esp ectiv e data sets D A and D B to get a new pair. Sin ce link m atrices L AB are usually v er y spars e, in pr actice, this will almost alwa y s pr o vide an unlinked p air. Sections 4 and 5 pr o vide more details. W e use the p rior P (Θ) = N ( b Θ , ( c b T ) − 1 ), where N ( m , V ) is a n ormal of mean m an d v ariance V . Matrix b T is the empirical s econd m omen ts matrix of the lin k ed ob ject features, although a different choi ce m ight b e adequate for d ifferen t applications. Constant c is a smo othing parameter set by the user. I n all of our exp eriments we set c to b e equal to the num b er of p ositiv e pairs. A go o d choic e of c might b e imp ortan t to obtain maxim um p erf or- mance, but we lea v e this issue as future work. W ang et al. ( 2009 ) p resen t some s ensitivit y analysis r esu lts for a particular application in text analysis. Empirical priors are a sen s ible c h oice, since this is a retriev al, not a pr edic- tiv e, task. Basically , the entire data set is the p opulation, from whic h p r ior information is obtained on p ossib le quer y sets. A data-dep end en t pr ior based on the p opulation is imp ortant for an approac h suc h as Ba ye sian sets, since deviances from th e “a v erage” b ehavio r in the data are usefu l to discriminate b et w een subp opulations. 3.5. On c ontinuous and multivariate r elations. Although w e fo cu s on measuring similarity of qualitativ e relationships, the same idea could b e ex- tended to c ontinuous (or ordinal) measures of relationship, or relationships where eac h L ij is a vec tor. F or instance, T ur n ey and Littman ( 2005 ) m ea- sure relations b etw een words by their co-o ccurrences on the neigh b orho o d of sp ecific keyw ords, su c h as the frequency of t w o w ord s b eing connected by 12 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI a sp ecific prep osition in a large b o dy of text do cum en ts. Sev eral similarit y metrics can b e defin ed on this v ector of contin uous r elationships. Ho wev er, giv en data on word features, one can easily mo d ify our ap p roac h by sub- stituting the logistic regression compon ent with some multiple regression mo del. 4. Ranking h yp erlinks on the web. In th e follo wing app lication w e con- sider a collection of web pages from sev er al univ ers ities: the W ebKB col- lection, where relations are give n by h yp erlinks [Crav en et al. ( 1998 )]. W eb pages are classified as b eing of type c ourse , dep artment , faculty , pr oje ct , staff , student or other . Do cuments come from four u niv ersities ( Cornel l , T exas , Washington and Wisc onsin ). W e are int erested in reco v ering pairs of web pages { A, B } where web page A has a link to we b page B . Notice that the relationship is asymmetric. Differen t t yp es of web p ages im p ly dif- feren t t yp es of links. F or instance, a faculty w eb p age linking to a pr oje ct w eb page constitutes a t yp e of link. Th e analogical reasoning task here is simplified if w e assume eac h w eb p age ob ject has a single role (i.e., exactly one out of the pre-defined t yp es { c ourse , dep artment , faculty , pr oje ct , staff , student , other } ), and therefore a p air of web pages implies a u n ique t yp e of relationship. The web page typ es are for ev aluation pur p oses only , as w e explain later: we will not pr o vide th is information to the m o del. Our main stand ard of comparison is a “flattened Ba yesia n sets” algo- rithm (whic h we will call “standard Ba yesian s ets,” SBSets , in constrast to the relational mo del, RBSets ). Usin g a multiv ariate indep endent Bernoulli mo del as in the original pap er [Ghahramani and Heller ( 2005 )], w e merge link ed w eb page pairs into single ro w s, and th en app ly the original algorithm directly to the merged data. It is clear that data p oints are not indep en d en t an ymore, b ut th e SBSe ts algorithm assumes this is the case. Ev aluating this algorithm serves th e pu rp ose of b oth measuring the loss of not treating relational d ata as suc h , as well as the limitations of ev aluating the similarit y of pairs th r ough mo dels for the marginal pr obabilities of A and B instead of mo d els for the predictiv e fun ction P ( L ij | X ij ). Binary data w as extracted from this database using the same metho dol- ogy as in Ghahramani and Heller ( 2005 ). A total of 19,450 binary v ariables p er ob ject are generated, where eac h v ariable ind icates whether a word f r om a fixed dictionary app ears in a giv en do cu m en t more frequently than the a v- erage. T o a void in tro ducing extra appr o ximations into RBSets , we redu ced the dimensionalit y of the original representa tion using singular v alue decom- p osition, obtaining 25 measur es p er ob ject. In this exp eriment ob jects are of th e s ame typ e, and therefore, dimen- sionalit y . The feature vect or X ij for eac h pair of ob jects { A i , B j } consists of the V features for ob ject A i , th e V features of ob ject B j , and m ea- sures Z = { Z 1 , . . . , Z V } , where Z v = ( A i v × B j v ) / ( | A i | × k B j k ), k A i k b eing RANKI N G RELA TIONS USIN G A N ALOGIES 13 the Euclidean norm of the V -dimensional repr esen tation of A i . W e also add a constant v alue (1) to the feature set as an interce pt term for the logistic regression. F eature set Z is exactly the one used in the cosine distance mea- sure, 4 a common and practical measure widely used in information retriev al [Manning, Ragha v an and Sc h ¨ utze ( 2008 )]. This feature space also h as the imp ortant adv antag e of scaling w ell (linearly) with th e n u m b er of v ariables in th e database. Moreo ver, adopting su c h features will mak e ou r compar- isons fairer, sin ce we ev aluate ho w w ell cosine d istance itself p erforms in our task. Notice that our c h oice of X ij is suitable for asymmetric relation- ships, as natur ally o ccurs in the domain of w eb page links. F or symmetric relationships, f eatures such as | A i v − B j v | could b e used instead. In order to s et the empirical pr ior, w e sample 10 “negativ e” p airs for eac h “p ositiv e” one, and weig h t them to reflect the p r op ortion of linke d to unlinked pairs in the database. Th at is, in the W ebKB study we use 10 negativ es for eac h p ositiv e, and we count eac h negativ e case as b eing 350 cases replicated. W e p er f orm sub s ampling and reweigh ting in ord er to b e able to fit the database in the memory of a d esktop computer. Ev aluation of the significance of retrieve d items often relies on su b jectiv e assessmen ts [Ghahramani and Heller ( 2005 )]. T o simplify our study , we will fo cus on particular setup s where ob jectiv e measures of success are defined. T o ev aluate the gain of our m o del ov er comp etitors, we will use the follo w- ing setup . In the first query , we are giv en all pairs of web pages of the t yp e student → c ourse from three of the lab eled universities, and ev aluate ho w relations are r ank ed in the fourth unive rsit y . Because we kno w class lab els for th e w eb pages (while the algorithm do es not), we can use the classes of the returned pairs to lab el a hit as b eing “relev ant” or “irr elev an t.” W e lab el a p air ( A i , B j ) as relev ant if and only if A i is of t yp e student and B j is of t yp e c ourse , and A i links to B j . This is a very stringent criterion, since other t yp es of relations could also b e v alid (e.g., staff → c ourse app ears to b e a r easonable matc h). How ev er, this facilitate s ob jectiv e comparisons of algorithms. Also, the other cla ss con tains man y t yp es of pages, whic h allo ws f or p ossibilities suc h as a student → “hobby” pair. Suc h p airs migh t b e hard to ev aluate (e.g., is that particular hobby demanding or chall enging in a similar w a y to coursewo rk?). As a compromise, w e omit all pages f r om the category other in order to b etter clarify differences b etw een algorithms. 5 Precision/recall curv es [Manning, Ragha v an and Sc h ¨ utze ( 2008 )] for the student → c ourse qu eries are shown in Figure 4 . There are four queries, 4 The cosine similarit y measure b etw een tw o items corresp onds t o th e sum of the fea- tures in Z . 5 As an ex treme example, querying s tudent → c ourse pairs from the wisc onsin u niver- sit y return ed student → other pairs at the top four. How ever, these other p ages were for some reason course p ages—suc h as http://www .cs.wisc. edu/ ~ markhill/c s752.html . 14 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI Fig. 4. R esults for student → c ourse r elationships. eac h corresp onding to a searc h ov er a sp ecific universit y giv en all v alid stu- dent → c ourse pairs from the other three. There are four algorithms on eac h ev aluation: the standard Ba yesia n sets with the original 19,450 binary v ariables for eac h ob ject, p lus another 19,45 0 binary v ariables, eac h cor- resp ond in g to the pro d uct of th e resp ectiv e v ariables in the original p air of ob jects ( SBSe ts1 ); the standard Ba yesian sets with the original bin ary v ariables only ( SBSet s 2 ); a standard cosine distance measur e o v er the 25- dimensional representat ion ( Cosine 1 ) for eac h page, with pairs b eing give n b y the com bined v ector of 50 features; a cosine distance measure using the ra w 19,450-dimensional binary for eac h do cum ent ( Cosine 2 ); our approac h, RBSets . In Figure 4 RBSe t s demonstrates consisten tly su p erior or equal p recision- recall. Although SBSets p erforms w ell when asked to r etriev e only student items or only c ourse items, it falls short of detecting what features of stu- dent a nd c ourse are r elev ant to predict a link. The discriminativ e mo del within RBS ets con veys this information thr ough the link parameters. W e also d id an exp eriment with a qu ery of type faculty → pr oje ct , sh o wn in Figure 5 . Th is time results b et ween algorithms we re closer to eac h other. T o mak e d ifferences more eviden t, w e adopt a sligh tly differen t measur e of RANKI N G RELA TIONS USIN G A N ALOGIES 15 Fig. 5. R esults for faculty → pr oje ct r elationships. success: w e coun t as a 1 hit if the p air retriev ed is a faculty → pr oje ct p air, and coun t as a 0.5 hit for pairs of t yp e student → pr oje ct an d staff → pr oje ct . Notice this is a muc h harder query . F or instance, the structur e of the pr oje ct w eb pages in the texas group w as qu ite distinct from the other unive rsities: they are mostly v ery short, b asically con taining links for mem b er s of the pro ject and other pro ject w eb p ages. Although the pr ecision/recall curve s con v ey a global picture of the p er- formance of eac h algorithm, they migh t not b e a completely clear w ay of ranking app roac hes for cases where cur v es in tersect at sev er al p oints. In order to su mmarize algorithm p erf ormances with a sin gle statistic, w e com- puted the area und er eac h p recision/recall curv e (with linear inte rp olation b et w een p oin ts). Resu lts are giv en in T able 1 . Numb ers in b old indicate the largest area under the curve . Th e d omin ance of RBSets should b e clear. 5. Ranking pr otein in teractions. T h e bu dding y east is a unicellular or- ganism that has b ecome a de-facto mo del organism for the study of molec- ular an d cellular biology [Botstein, Ch ervitz and Cherr y ( 1997 )]. There are ab out 6000 proteins in th e bud ding yea s t, whic h in teract in a num b er of w ays [C herry et al. ( 1997 )]. F or instance, proteins bind together to form 16 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI T able 1 Ar e a under the pr e ci sion/r e c al l curve for e ach algorithm and query Student → course F aculty → pro j ect C1 C2 RB SB1 SB2 C1 C2 R B S B1 SB2 Cornell 0.87 0.82 0.87 0.82 0.80 0.19 0.18 0.24 0.18 0.18 T exas 0.62 0.32 0.77 0.55 0.54 0.24 0.21 0.29 0.12 0.12 W ashington 0.69 0.31 0.76 0.67 0.64 0.40 0.42 0.47 0.40 0.40 Wisconsin 0.77 0.72 0.88 0.75 0.73 0.28 0.30 0.26 0.1 9 0.21 protein complexes, the p hysical u nits that carry out most functions in the cell [Kr ogan et al. ( 2006 )]. In r ecen t ye ars, signifi cant resources hav e b een directed to collect exp erimental evidence of physical p r oteins b inding, in an effort to infer and catalogue protein complexes and their m ultifaceted f unc- tional roles [e.g., Fields and Song ( 1989 ); Itˆ o et al. ( 2000 ); Uetz et al. ( 2000 ); Ga vin et al. ( 2002 ); Ho et al. ( 2002 )]. Currently , ther e are four main sour ces of in teractions b etw een pairs of proteins that target pr oteins lo calized in differen t cellular compartment s with v ariable degrees of success: (i) litera- ture curated inte ractions [Reguly et al. ( 2006 )], (ii) yea st t wo-h ybrid (Y2H) in teraction assa ys [Y u et al. ( 2008 )], (iii) p rotein fragmen t complemen tation (PCA) in teraction assays [T arasso v et al. ( 2008 )], and (iv) tandem affinity purification (T AP) interact ion assa ys [Ga vin et al. ( 2006 ); K rogan et al. ( 2006 )]. These collections in clud e a total of ab out 12,292 p rotein in terac- tions [Jen sen and Bork ( 2008 )], although th e num b er of such in teractions is estimated to b e b etw een 18,000 [Y u et al. ( 2008 )] and 30,000 [v on Mering et al. ( 2002 )]. Statistical metho d s hav e b een dev elop ed for analyzing many asp ects of this large protein in teraction netw ork, including de-noising [Bernard , V aughn and Hartemink ( 2007 ); Airoldi et al. ( 2008 )], fu nction pr ediction [Nabiev a et al. ( 2005 )] and id entificatio n of b inding motifs [Banks et al. ( 2008 )]. 5.1. Overvie w of the analysis. W e consider m u ltiple f u nctional catego - rization systems f or the proteins in buddin g yeast. F or ev aluation p urp oses, w e use individu al proteins’ fun ctional annotations cu r ated by the Munich Institute for Pr otein Sequencing [MIPS, Mew es et al. ( 2004 )], those b y the Ky oto En cyclop edia of Genes and Genomes [KEGG, Kanehisa and Goto ( 2000 )] and those by the Gene Onto logy consortium [GO, Ashburner et al. ( 2000 )]. W e consider m u ltiple collections of physic al p rotein in teractions that enco de alternativ e seman tics. Physica l protein-to-protein inte ractions in the MIPS curated collection measure physic al b inding ev ents obs er ved exp er i- men tally in Y2H and T AP exp erim ents, wh ereas physical protein-to-protein in teractions in the K EGG curated collection measure a num b er of d ifferent mo des of int eractions, including phosp orelation, meth ylation and physica l RANKI N G RELA TIONS USIN G A N ALOGIES 17 T able 2 Col le ction of data sets use d to gener ate pr otein-sp e cific fe atur es No. Measurements d escription Data sources 1. Expression microarra ys Gasc h et al. ( 2000 ); Brem et al. ( 2005 ); Primig et al. ( 2000 ); Yvert et al. ( 2003 ) 2. Synthetic genetic interactions Breitkreutz, Stark and Tye rs ( 2003 ); SGD 3. Cellular lo calization Huh et al. ( 2003 ) 4. T ranscription factor binding sites Harbison et al. ( 2004 ); TRANS F AC 5. Sequence similarities Altsch ul et al. ( 1990 ); Zhu and Zhang ( 1999 ) binding, all taking place in the conte x t of a sp ecific signaling p ath wa y . So we ha ve th r ee p ossible functional annotation databases (MIPS, KEGG and GO) and t wo p ossible link matrices (MIPS and KEGG), which can b e com bin ed. Our exp eriment al pip eline is as follo ws: (i) Pic k a database of f u nctional annotations, sa y , MIPS, and a collection of interact ions, sa y , MIPS (again). (ii) Pic k a pair of categ ories, M 1 and M 2 . F or instance, tak e M 1 to b e c y- toplasm (MIPS 40.03) and M 2 to b e cytoplasm ic and nucle ar de gr adatio n (MIPS 06.13.01 ). (iii) Sample, un iformly at random and without replace- men t, a set S of 15 int eractions in the c h osen collection. (iv) Rank other in teracting p airs 6 according to the s core in equation ( 3.3 ) and , for compar- ison purp oses, according to three other ap p roac hes to b e describ ed in S ec- tion 5.1.4 . (v) The p r o cess is rep eated for a large n u m b er of p airs M 1 × M 2 , and 5 different qu ery sets S are generated for eac h pair of categories. (vi) Calculate an ev aluation metric for eac h query and eac h of the four scores, and rep ort a comparative summary of the results. 5.1.1. Pr otein-sp e cific fe atur es. The protein-sp ecific features w ere gen- erated usin g the data sets su mmarized in T able 2 and an additional d ata set [Qi, Bar-Joseph and Klein-Seetharaman ( 2006 )]. Twen t y gene expr ession attributes w ere obtained fr om the data set pro cessed b y Qi, Bar-Joseph and Klein-Seetharaman ( 2006 ). Eac h gene expression attribute for a protein pair P i : P j corresp onds to the correlation co efficient b et ween the expression lev- els of corresp onding genes. Th e 20 different attributes are obtained f r om 20 differen t exp erimen tal conditions as measured by microarra ys. W e did not use pairs of p roteins fr om Qi et al. for which w e did not h a ve d ata in the data sets listed in T able 2 . This r esulted in approximat ely 6000 p ositive ly link ed data p oin ts for the MIPS n et wo rk an d 39,000 f or KEGG. W e generated another 25 pr otein–protein gene exp ression features from the data in T able 2 using the same p ro cedure based on correlation co effi- 6 The p ortion of ranked list that is relev ant for ev aluation purp oses is limited to a subset of th e protein–protein interactions. More details are given in Section 5.1.3 . 18 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI cien ts. Th is giv es a total of 45 attributes, corresp onding to the main data set used in our relational Ba yesian sets runs. Another data set was generated using the remaining (i.e., nonmicroarray) features of T able 2 . Suc h features are b inary and highly sparse, with most en tries b eing 0 for the m a j orit y of linked p airs. W e remov ed attributes for whic h we had fewer th an 20 link ed pairs with p ositiv e v alues according to the MIPS net work. The total num b er of extra bin ary attributes was 16. Sev eral measurements we re missin g. W e imp uted missin g v alues for eac h v ariable in a particular d ata p oin t by using its empirical a ve rage among the observ ed v alues. Giv en the 45 or 61 attributes of a giv en pair { P i , P j } , we app lied a nonlinear tran s formation where w e normalize the v ector b y its Euclidean norm in order to obtain our feature table X . 5.1.2. Calibr ating the prior for Θ . W e initial ly fit a logistic regression classifier u sing a maxim um lik eliho o d estimation (MLE) and our data, ob- taining the estimate b Θ. Ou r choice of co v ariance matrix b Σ for Θ is defin ed to b e a rescaling of a squared norm of the d ata: ( b Σ) − 1 = X T POS X POS , (5.1) where X POS is the m atrix con taining th e pr otein–protein features only of the linked pairs u sed in the MLE computation. 5.1.3. Evaluation metrics. As in the W ebKB exp erimen t, w e p rop ose an ob jectiv e measure of ev aluation that is used to compare differen t algorithms. Consider a quer y set S , and a rank ed r esp onse list R = { R 1 , R 2 , R 3 , . . . , R N } of protein–protein pairs. Ev ery elemen t of S is a p air of pr oteins P i : P j suc h that P i is of class M i and P j is of class M j , where M i and M j are classes from either MIPS, KE GG or Gene Ontolog y . In general, proteins b elong to m u ltiple classes. This is in con trast with the W ebKB exp erimen t, where, according to our w eb page categ orization, there w as only one p ossible t yp e of relationship for eac h pair of we b pages. Th e retriev al algorithm that generates R do es not receiv e any inform ation concerning the MIPS, KEGG or GO taxonom y . R starts with the link ed p rotein pair that is judged most similar to S , follo w ed by the other pr otein pairs in the p opulation, in decreasing order of similarit y . Eac h algorithm has its own measure of similarit y . The ev aluation criterion for eac h algorithm is as f ollo ws: as b efore, w e generate a precision-recall curve and calculate the area und er the curve (A UC). W e also calculate the prop ortion (TOP10), among the top 10 ele- men ts in eac h ranking, of pairs that matc h the original { M 1 , M 2 } selection (i.e., a “correct” P i : P j is one where P i is of class M 1 and P j of class M 2 , or vice-v ersa. Notice that eac h p rotein b elongs to multiple classes, so b oth RANKI N G RELA TIONS USIN G A N ALOGIES 19 conditions migh t b e satisfied.) Sin ce a researc her is only like ly to lo ok at the top ranked p airs, it mak es sense to define a m easure that uses only a sub set of the r anking. AUC and TOP10 are our tw o ev aluation measures. The original classes { M 1 , M 2 } are kno w n to the exp erimente r but not kno wn to the algorithms. As in the W ebKB exp eriment , our criterion is rather stringent, in the sense th at it requires a p erfect matc h of eac h R I with th e MIPS, KEGG or GO cat egorizati on. T here are sev eral wa y s by whic h a pair R I migh t b e analogous to the relation im p licit in S , and they do n ot need to agree with MIPS, GO or K EGG. Still, if we are willing to b eliev e that these standard categ orization systems capture functional or- ganizatio n of p roteins at some lev el, this must lead to asso ciation b et ween catego ries giv en to S and relev an t subp opulations of protein–protein inter- actions similar to S . Therefore, th e corresp onding A UC and TOP10 are useful tools for comparing differen t algorithms even if the actual measures are like ly to b e p essimistic for a fixed algorithm. 5.1.4. Comp eting algorithms. W e compare our metho d against a v arian t of it and tw o similarit y metrics wid ely used for information r etriev al: 1. The cosine score [Manning, Ragha v an and Sch ¨ utze ( 2008 )], denoted by cos . 2. The n earest neighbor score, d enoted by nns . 3. The r elational m aximum lik eliho o d sets score, denoted by mls . The nearest neigh b or score measures the minimum Euclidean distance b e- t wee n R I and any individual p oin t in S , for a giv en query set S and a given candidate p oin t R I . The relational m axim um lik eliho o d sets is a v ariation of RBS ets where we initially samp le a subset of the un link ed p airs (10,000 p oint s in our setup) and, for eac h query S , we fit a logistic regression mo del to obtain the parameter estimate Θ MLE S . W e also u s e a logistic regression mo del fit to the whole data set (the same one u sed to generate th e p rior for RBSe t s ), giving the estimate Θ MLE . A new score, analogous to ( 3.3 ), is giv en by log P ( L ij = 1 | X ij , Θ MLE S ) − log P ( L ij = 1 | X ij , Θ MLE ), that is, w e do not integ r ate out the parameters or use a prior, b ut in stead the mo dels are fixed at their r esp ectiv e estimates. Neither cos or nns can b e interpreted as measures of analo gical simi- larit y , in the s en se that they do n ot tak e in to account h ow the protein p air features X con tribu te to th eir int eraction. 7 It is true that a d irect measure of analogica l similarit y is n ot theoretically requir ed to p er f orm wel l according to our (nonanalogical) ev aluation metric. How ev er, w e w ill see that th er e are pr actica l adv an tages in doing so. 7 As a consequence, none uses negative data. Another consequence is th e n ecessit y of mod eling the input space that generates X , a difficult task given th e dimensionalit y and the continuous nature of the features. 20 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI 5.2. R esults on the MIP S c ol le ction of physic al inter actions. F or this batc h of exp eriments, we u se the MIPS n et wo rk of p rotein–protein inter- actions to defi ne the relationships. In the initial exp eriment, w e selected queries f rom all combinatio ns of MIPS classes f or which there we re at least 50 lin ked pairs P i : P j in the net work that satisfied the c hoice of classes. Eac h query set cont ained 15 pairs. After remo vin g the MIPS-categorize d proteins for w h ic h w e had no feature data, we ended up with a total of 6125 pro- teins and 7788 p ositiv e int eractions. W e set the prior for RBSets usin g a sample of 225,842 pairs lab eled as having no in teraction, as selected by Qi, Bar-Joseph and Klein-Seetharaman ( 2006 ). F or eac h ten tativ e query set S of catego ries { M 1 , M 2 } , we scored and rank ed pairs P ′ i : P ′ j suc h that b oth P ′ i and P ′ j w ere connected to some pro- tein app earing in S by a path of n o more than tw o steps, according to the MIPS n et work. The reasons for the fi ltering are t wo -fold: to increase the computational p erf orm ance of th e rankin g sin ce few er pairs are scored; and to minimize the c hance that u ndesirable pairs would app ear in th e top 10 rank ed p airs. T en tativ e qu eries w ould n ot b e p erformed if after filtering we obtained fewe r than 50 p ossible correct matc h es. T rivial queries, where filter- ing resu lted only in p airs in the same class as the query , were also discarded. The resulting num b er of u nique pairs of categorie s { M 1 , M 2 } wa s 931 classes of in teractio ns. F or eac h pair of categories, we sampled our qu er y set S 5 times, generating a total of 4655 rankings p er algorithm. W e run t wo t yp es of exp eriment s. In one ve rsion, w e giv e to RBS ets the data conta in ing only the 45 (con tinuous) microarray measurements. In the second v ariation, we provide to RBSe t s all 61 v ariables, includin g the 16 sparse binary indicators. How ev er, w e n oticed that th e addition of the 16 binary v ariables hurts RBS ets considerably . W e conjecture that one reason migh t b e th e d egradation of the v ariational app r o ximation. In cluding the binary v ariables hardly c hanged the other three metho ds, so we c ho ose to use the 61 v ariable data set for the other metho ds. 8 T able 3 summarizes the results of this exp eriment. W e s ho w the n um b er of times eac h metho d wins according to b oth the A UC and TO P10 criteria. The num b er of win s is presen ted as divided by 5, the num b er of rand om sets generated for eac h query t yp e { M 1 , M 2 } (notice these n u m b ers do not need to add up to 931, since ties are p ossible). Moreo ver, we also p resen ted “smo othed” v ersions of this statistic, where we count a m etho d as the winner for any giv en { M 1 , M 2 } category if, for the group of 5 queries, the metho d obtains the b est result in at least 3 of the sets. The motiv ation is to smo oth 8 W e also p erformed an exp eriment (n ot included ) where only the contin uous attributes w ere used by the other method s. The advan tage of RBSets still increased, slightly (by a 2% margin against the cosine distance metho d). F or this reason, w e analyze the most p essimistic case. RANKI N G RELA TIONS USIN G A N ALOGIES 21 T able 3 Numb er of times e ach metho d wins when querying p airs of MI PS classes using the M IPS pr otein–pr otein i nter action network. The first two c olum ns, # A U C and # T OP10 , c ount the numb er of times the r esp e ctive metho d obtains the b est sc or e ac c or ding to the AUC and TOP10 me asur es, r esp e ctively, among the 4 appr o aches. This is divide d by the numb er of r eplic ations of e ach query typ e (5). The last two c olumns, # AUC.S and # TOP10.S , ar e “smo othe d” versions of this statistic: a metho d is de clar e d the winner of a r ound of 5 r eplic ations if it obtains the b est sc or e in at l e ast 3 out of the 5 r epli c ations. The top table shows the r esults when only the c ontinuous variables ar e use d by RB S ets , and in the b ottom table when the discr ete variables ar e also given to RBSets Metho d #A UC #TOP10 #A UC.S #TOP10.S (a) COS 240 294 219 277 NNS 42 122 28 75 MLS 105 270 52 198 RBSets 542 556 578 587 (b) COS 314 356 306 340 NNS 75 146 62 111 MLS 273 329 246 272 RBSets 267 402 245 387 out the extra v ariabilit y added by th e p articular set of 15 protein pairs f or a fixed { M 1 , M 2 } . Th e prop osed r elational Ba yesian s ets metho d is the clear winner according to all measures wh en we s elect only the con tinuous v ari- ables. F or th is reason, for the r est of this section all analysis and exp eriments will consider only this case. T able 4 displays a pairwise comparison of the metho ds. In this table w e sho w ho w often the ro w metho d p erforms b etter than the column metho d, among those trials wher e there was n o tie. Again, RBSe t s domin ates. T able 4 Pairwise c omp arison of metho ds ac c or ding to the AUC and TOP10 criterion. Each c el l shows the pr op ortion of the trials wher e the metho d in the r esp e ctive r ow wins over the metho d in the c olumn, ac c or ding to b oth criteria. In e ach c el l, the pr op ortion is c alculate d with r esp e ct to the 4655 r ankings wher e no tie happ ene d A UC TOP10 COS NNS MLS RBSets COS NNS MLS RBSets COS – 0.67 0.43 0 .30 – 0.70 0.46 0.30 NNS 0.32 – 0.18 0.06 0 .29 – 0.25 0.11 MLS 0.56 0.81 – 0 .25 0.53 0.74 – 0.28 RBSets 0.69 0.93 0.74 – 0.69 0.88 0.71 – 22 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI T able 5 Distribution acr oss al l queries of the numb er of hits in the top 10 p ai rs, as r anke d by e ach algorithm. The mor e skewe d to the right, the b etter. Notic e that using GO c ate gories doubles the numb er of zer o hits for RB S ets 0 1 2 3 4 5 6 7 8 9 10 Proportion of t op hits using MIPS categories and links sp ecified by the MIPS database COS 0.12 0.15 0.12 0.10 0.08 0.07 0.06 0.05 0.04 0.07 0.08 NNS 0.29 0.16 0.14 0.10 0.06 0.05 0.03 0.03 0.03 0.03 0.02 MLS 0.12 0.12 0.12 0.10 0.09 0.08 0.07 0.06 0.07 0.06 0.07 RBSets 0.04 0.08 0.09 0.09 0.09 0.08 0.09 0.07 0.09 0.08 0.14 Proportion of t op hits u sing GO categories and links sp ecified by the MIPS database COS 0.12 0.13 0.11 0.10 0.11 0.09 0.06 0.06 0.04 0.06 0.06 NNS 0.53 0.23 0.07 0.02 0.02 0.02 0.04 0.01 0.00 0.00 0.01 MLS 0.16 0.11 0.12 0.10 0.08 0.08 0.08 0.06 0.05 0.06 0.05 RBSets 0.09 0.09 0.10 0.10 0.08 0.08 0.06 0.08 0.08 0.07 0.12 Another us efu l summary is the d istr ibution of correct hits in th e top 10 rank ed element s across queries. This provi des a measure of the d ifficult y of the p roblem, b esides the r elativ e p erformance of eac h algorithm. In T able 5 w e show the p rop ortion of correct hits among the top 10 f or eac h algorithm for our queries using MIPS categorization and also GO categorizat ion, as explained in the next section. Ab out 14% of the time, all pairs in the top 10 pairs rank ed by RBS ets were of the in tended typ e, compared to 8% of the second b est approac h. 5.2.1. Changing the c ate gorization system. A v ariation of th is exp er i- men t was p erformed where the pr otein categorizatio n s do not come from the same family as the link n et wo rk, that is, where we used th e MIPS net work but not the MIPS catego rization. Ins tead we p erformed queries according to the Gene Onto logy cat egories. S tarting from 150 pre-selected GO cat- egories [Mye rs et al. ( 2006 )], we once again generated unord ered category pairs { M 1 , M 2 } . A total of 179 queries, with 5 replications eac h (a total of 895 rankin gs), were generated and th e results summ arized in T able 6 . T able 6 Numb er of times e ach metho d wins when querying p ai rs of GO classes using the MIPS pr otein–pr otein i nter action network. Columns # AUC , # T OP10 , # AUC.S and # TOP10.S ar e define d as in T able 3 Metho d #A UC #TOP10 #A UC.S #TOP10.S COS 58 73 58 72 NNS 1 10 0 4 MLS 26 55 13 38 RBSets 93 105 101 110 RANKI N G RELA TIONS USIN G A N ALOGIES 23 Fig. 6. Distribution of the c over age of valid p airs in the M I PS network, ac c or di ng to our gener ate d query sets. R esults ar e br oken into the two c ate gorization systems (MIPS and GO) use d in this exp eriment. This is a more challenging scenario for our app roac h, whic h is optimized with resp ect to MIPS. Still, we are able to outp erf orm other approac hes. Differences are less dramatic, but consistent. In the pairwise comparison of RBSets against th e second b est metho d , cos , ou r metho d wins 62% of the time by the TOP10 criterion. 5.2.2. The r ole of filtering. In b oth exp erimen ts with the MIPS net work, w e filtered candidates by examinin g only a subset of the proteins linke d to the elemen ts in the query set by a path of n o more than t w o pr oteins. It is relev ant to ev aluate ho w m u c h co v erage of eac h category pair { M 1 , M 2 } w e obtain by th is n eigh b orho o d selection. F or eac h query S , we calculate the prop ortion of pairs P i : P j of th e same catego rization { M 1 , M 2 } suc h that b oth P i and P j are included in the n eigh- b orho o d. Figure 6 s h o ws the resulting distribu tions of su c h prop ortions (from 0 to 100%): a histogram for the MIPS searc h and a histogram for the GO searc h. Despite the s mall neigh b orho o d , cov erage is large. F or the MIPS catego rization, 93% of the queries resulted in a co ve r age of at least 75% (with 24% of the queries resulting in p erfect cov erage). Although fil- tering implies that some v alid pairs will neve r b e ranked, the gain obtained b y redu cing false p ositive s in the top 10 ranked pairs is considerable (results not sh o wn) across all metho ds, and the computational gain of redu cing the searc h space is particularly relev ant in exploratory data analysis. 5.3. R esults on the KE GG c ol le ction of signaling p athways. W e rep eat the same exp erimental setup, no w us in g the KEGG net work to defin e the protein–protein in teractions. W e selected p roteins from the KEGG catego- 24 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI T able 7 Numb er of times e ach metho d wins when querying p airs of KEGG classes using the KEGG pr otein–pr otein inter action network. Columns # AUC , # TOP10 , # AUC.S and # TOP10.S ar e define d as in T able 3 Metho d #A UC #TOP10 #A UC.S #TOP10.S COS 159 575 134 507 NNS 30 305 17 227 MLS 290 506 199 431 RBSets 1042 1091 1107 1212 rization system for w hic h we had data av ailable. A total of 6125 p r oteins w ere selected. The KE GG net work is muc h more dense than MIPS. A total of 38,961 p ositiv e pairs and 226,18 8 negativ e links w ere used to generate our empirical prior. Ho we v er, since the K EGG net work is m uc h more dense than MIPS, we filtered our candidate pairs by allo wing on ly pr oteins that are dir ectly linked to the pr oteins in the qu er y set S . Ev en under this restriction, w e are able to obtain high co v erage: the n eigh b orho o d of 90% of the queries included all v alid pairs of the same category , and essentia lly all quer ies included at least 75% of the p airs falling in the same category as the query set. A total of 1523 p ossible pairs of categories (7615 queries, consid er in g the 5 replications) w ere generated. Results are summarized in T able 7 . Again, it is eviden t that RBSe ts dominates other metho d s. I n the pairwise comparison against cos , RB- Sets w ins 76% of the times according to the TOP10 criterion. Ho w ever, the ranking problem in the KEGG net work w as muc h harder than in the MIPS net work (according to our au tomated nonanalogical criterion). W e b eliev e that the reason is that, in KEGG, th e simple fi ltering sc heme h as muc h less influence as reflected by the high co v erage. Th e distribution of th e n u m b er of hits in the top 10 ranke d items is sho w n in T able 8 . Despite th e success of R BSets r elativ e to the other algorithms, there is ro om f or impr o veme n t. T able 8 Distribution acr oss al l queries of the numb er of hits in the top 10 p ai rs, as r anke d by e ach algorithm. T he mor e skewe d to the right, the b etter 0 1 2 3 4 5 6 7 8 9 10 Proportion of t op hits using K EGG categories and links sp ecified by t h e KEGG database COS 0.56 0.21 0.08 0.03 0.02 0.01 0.01 0.01 0.01 0.01 0.01 NNS 0.89 0.03 0.04 0.01 0.00 0.00 0.00 0.00 0.00 0.00 0.00 MLS 0.57 0.21 0.08 0.04 0.02 0.01 0.01 0.00 0.00 0.00 0.00 RBSets 0.29 0.24 0.16 0.09 0.06 0.03 0.02 0.01 0.03 0.02 0.01 RANKI N G RELA TIONS USIN G A N ALOGIES 25 6. More related work. Th ere is a large literature on analogical reasoning in artificial in telligence and p syc h ology . W e refer to F r enc h ( 2002 ) for a sur- v ey , and to more recent pap ers on clustering [Marx et al. ( 2002 )], prediction [T u rney an d Littman ( 2005 ); T urney ( 2008a )] and dimensionalit y r eduction [Memisevic and Hin ton ( 2005 )] as examples of other applications. Classical approac hes f or p lann ing ha v e also exploited analogical similarities [V eloso and Carb onell ( 1993 )]. Nonprobabilistic similarit y functions b etw een relational structures ha v e also b een dev elop ed for the p urp ose of d eriving ke rnel matrices, suc h as those required by sup p ort v ector machines. Borgw ardt ( 2007 ) pro v id es a comprehensiv e survey and state-of-the-art metho ds. It wo u ld b e inte r esting to adapt suc h metho ds to problems of analogical reasoning. The graphical m o del formula tion of Geto or et al. ( 2002 ) incorp orates mo dels of link existence in r elational d atabases, an id ea u sed explicitly in Section 3 as the fi rst step of our problem form ulation. In the clustering literature, th e probabilistic approac h of Kemp et al. ( 2006 ) is motiv ated by principles similar to those in our formulat ion: the idea is th at there is an infinite mixture of su bp opu lations that generates the observed relations. O ur problem, how ev er, is to retriev e other elemen ts of a su bp opulation describ ed b y elemen ts of a query set, a goal th at is closer to the classical paradigm of analogica l reasoning. As discussed in Section 3.2 , our mo del can b e interpreted as a type of blo c k mo del [Kemp et al. ( 2006 ); Xu et al. ( 2006 ); Airoldi et al. ( 2008 )] with observ able f eatures. Link indicators are ind ep endent giv en the ob ject fea- tures, which might n ot actually b e the case for particular choice s of feature space. In theory , blo ck mo dels sidestep this issu e b y learning all the neces- sary latent features that account for link dep end ence. An im p ortan t future extension of our w ork w ou ld consist of tractably mo d eling the residual link asso ciation that is n ot accoun ted for by our observed features. Disco v erin g analogies is a sp ecific task within th e general problem of gen- erating latent relationships f r om relational data. Some of the first formal metho ds for disco ve ring laten t r elationships from m u ltiple data sets w er e in- tro duced in the literature of in ductiv e logic p rogramming, suc h as the in v erse resolution metho d [Muggleton ( 1981 )]. A more recen t prob ab ilistic m etho d is discussed b y Kok and Domingos ( 2007 ). Dˇ zeroski and La vraˇ c ( 2001 ) and Getoor and T ask ar ( 2007 ) pro vid e an o v erview of r elational learning metho ds from a data mining and mac h ine learnin g p er s p ectiv e. A particularly activ e sub fi eld on laten t relationship generation lies within text analysis r esearc h. F or instance, S teph ens et al. ( 2001 ) d escrib e an ap- proac h for discov er in g relations b et w een genes giv en MEDLINE abstracts. In the cont ext of information retriev al, Cafarella, Banko and Etzioni ( 2006 ) describ e an application of recent unsup ervised in formation extraction meth- o ds: relati ons generated from unstructured text do cuments are used as a 26 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI prepro cessing step to build an in d ex of web pages. In analogical reasoning applications, our metho d has b een used b y others for question-answ ering analysis [W ang et al. ( 2009 )]. The idea of measuring the similarit y of tw o data p oin ts based on a p redic- tiv e function has app eared in th e literature on matc hin g for causal inf erence. Supp ose we are give n a mo d el for predicting an outcome Y giv en a treatmen t Z and a set of p oten tial confoun ders X . F or simplicit y , assume Z ∈ { 0 , 1 } . The goal of m atc hing is to fin d, for eac h data p oin t ( X i , Y i , Z i ), the closest matc h ( X j , Y j , Z j ) according to the confoun ding v ariables X . In p rinciple, an y clustering criterion could b e used in this task [Gelman and Hill ( 2007 )]. The p rop ensity score criterion [Rosenbaum ( 2002 )] m easur es the similarit y of tw o feature ve ctors X i and X j b y comparing the predictions P ( Z i = 1 | X i ) and P ( Z j = 1 | X j ). If the conditional P ( Z = 1 | X ) is give n by a logistic re- gression mo d el with parameter v ector Θ, Gelman and Hill ( 2007 ) s u ggest measuring the d ifference b et wee n X T i Θ and X T j Θ. Wh ile this is not th e same as comparing t wo pr ed ictiv e fun ctions as in our framew ork, the core idea of using pr edictiv e functions to define similarit y remains. A pr eliminary v ersion of this pap er app eared in the pro ceedings of the 11th Int ernational Conf erence on Artificial Intelli gence and Statistics [Silv a, Heller and Ghahramani ( 2007 )]. 7. Conclusion. W e ha v e presented a f ramew ork for p erforming analogi- cal reasoning within a Ba y esian data analysis formulat ion. There is of course m u c h more to analogical reasoning th an calculati ng the similarit y of related pairs. As future w ork, w e will consider hierarc hical mod els that could in principle compare r elational stru ctures (suc h as protein complexes) of d if- feren t sizes. In p articular, the literature on graph k ernels [Borgw ardt ( 2007 )] could provide ins ights on develo ping efficient similarit y metrics w ithin our probabilitistic framew ork. Also, we w ould like to com bine the pr op erties of the m ixed-mem b ership sto c hastic blo ck mo del of Airoldi et al. ( 2008 ), where ob jects are clustered in to multiple roles according to the relationship matrix L AB , with our frame- w ork where relationship in d icators are conditionally ind ep endent giv en ob- serv ed features. Finally , we would lik e to consider th e case where multiple relationship matrices are a v ailable, allo wing for the comparison of relational structures with multiple t yp es of ob jects. Muc h remains to b e done to create a complete analogical r easoning sys- tem, bu t the d escrib ed app r oac h h as immediate applications to information retriev al and exploratory data analysis. Ac kn o wledgment s. W e w ould lik e to thank the anon ymous r eview ers and the ed itor for sev eral suggestions that impro ved the p r esen tation of this pap er, and for add itional relev ant referen ces. RANKI N G RELA TIONS USIN G A N ALOGIES 27 SUPPLEMENT AR Y MA TERIAL Supp lemen t: Ja v a implemen tation of the Relational Ba yesia n Sets m etho d (DOI: 10.121 4/09-A OAS321SUPP ; .zip). W e pr o vide complete s ource co de for our metho d, and instr uctions on how to r ebuild our exp erimen ts. With the co de it is also p ossible to test v ariations of our q u eries, analyzing the sensitivit y of the results to different query sizes and initialization of the v ariational optimizer. REFERENCES Airoldi, E. M. (2007). Getting started in probabilistic graphical mo d els. PL oS Compu- tational Biol o gy 3 e252. Airoldi, E. M. , Blei , D. M. , Xing, E. P. and Fienberg, S. E. (2005). A latent mixed- membership mod el for relational data. In Workshop on Link D i sc overy: Issues, Ap- pr o aches and Applic ations, in Conjunction With the 11th International ACM SIGKDD Confer enc e . Chicago, IL. Airoldi, E. M. , Blei, D. M. , Fienberg, S. E. and Xing, E. P. (2008). Mixed mem- b ership sto chastic blo ckmodels. J. Mach. L e arn. R es. 9 1981–201 4. Al tschul, S. F. , Gish, W. , Miller, W. , Myers, E. W. and Li pman, D. J. (1990). Basic lo cal alignment search to ol. Journal of Mol e cular Biolo gy 215 403–410. Ashburner, M. , Ba ll, C. A. , Blake, J. A. , Botstein, D. , Butler, H. , Cherr y, J. M. , Da vis, A. P. , Dol inski, K. , Dwight, S. S. , Eppig, J. T. , Harris, M. A. , Hill, D. P. , Issel-T ar ver, L. , Kasarskis, A. , Lewis, S. , Ma tese, J. C. , Richa rd son, J. E. , Ringw ald, M . , R ubinand, G. M. and Sherlock, G. (2000). Gene ontol ogy: Tool for the u nification of b iology . The gene ontology consortium. Natur e Genetics 25 25–29. Banks, E. , Na bi e v a, E. , Peterson, R. and Singh , M. (2008). NetGrep: F ast netw ork sc hema searches in intera ctomes. Genome Biol o gy 24 1473–1480. Bernard, A . , V a ug h n, D. S. and Har temink, A. J. (2007). R econstructing the top ol- ogy of protein complexes. In Re se ar ch in Computational Mol e cular Bi olo gy 2007 (RE- COMB07) ( T. Sp eed and H . Huan g, eds.). L e ctur e Notes i n Bioinformatics 4453 32–46. Springer, Berlin. Borgw ardt, K. (2007). Graph kernels. Ph.D. thesis, Lu dwig-Maximilians-Univ. Munic h. Botstein, D. , Cher vitz, S. A. and Cherr y, J. M. (1997). Y east as a mo del organism. Scienc e 277 1259–1260. Breitkreutz, B. J. , St ark, C. and Tye rs, M . (2003). The GRID: The General Rep os- itory for I nteractio n Datasets. Genome Biolo gy 4 R23. Brem, R. B. , Storey, J. D. , W hittle, J. and Krugl y ak, L. (2005). Genetic interac- tions b etw een p olymorph isms that affect gene expression in yeast. Natur e 436 701–703. Caf arella, M. , Banko, M. and Etzioni, O. (2006). Relational w eb search. T echnical rep ort 2006-04-02, Univ. W ashington, D ept. Computer S cience and En gineering. Cherr y, J. M . , B all, C. , W e ng, S. , Juvik, G. , S chmidt, R. , Adler, C. , Dunn, B. , Dw ight, S. , R iles, L. , Mor ti mer, R. K. and Botstein, D. (1997). Genetic and physical maps of saccharom yces cerevisiae. Natur e 387 67–73. Cra ven, M. , DiP asquo, D. , Freit ag, D. , McCallum, A. , Mitchell, T. , Nigam, K. and Sla tter y, S. (1998). Learning to ext ract symbolic k now ledge from the W orld Wide Web. In Pr o c e e dings of AAAI’98 509–516. MIT Press, Cam b rid ge, MA. D ˇ zerosk i, S. and La vra ˇ c, N. (2001). R elational Data Mi ni ng . Springer, Berlin. Fields, S. and Song, O. (1989). A nove l genetic sy stem to detect protein–protein inter- actions. Natur e 340 245–246. 28 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI Fienberg, S. E. , Mey er, M . M. and W asserm an, S. (1985). Statistical analysis of multiple so ciometric relations. J. Amer . Statist. Asso c. 80 51–67. French, R. (2002). The computational mod eling of analogy-making. T r ends in Co gnitive Scienc es 6 200–205. Gasch, A. P. , S p ellman, P. T. , Kao, C. M . , Carmel-Harel, O. , Eisen, M. B . , Storz, G. , Botstein, D. and Bro wn, P. O. (2000). Genomic ex pression programs in the resp onse of yeast cells to environmen tal changes. Mol e cular Bi olo gy of the Cel l 11 4241–42 57. Ga vin, A.-C . , B ¨ osche, M. , Krause, R. , Grand i , P. , Marzioch, M. , B auer, A. , Schul tz, J. , Rick, J. M. , Michon, A.-M. , Crucia t, C.-M. , Remor, M. , H ¨ ofer t, C. , Schelde r , M . , Braj enovic, M. , Ruffner, H. , Me rino, A. , Klei n, K. , Dickson, D. , Hudak, M. , Rudi, T. , Gnau, V. , Bauch, A. , B astuck, S. , Huhse , B. , Leutwein, C. , H eur tier, M . -A. , Copley, R. R. , Edelmann, A. , Que rfur th, E. , R ybin, V . , Drewes, G. , R aida, M . , Bouwmeester, T. , Bork, P. , Seraph i n, B. , Kuster, B. , Neubauer, G. and Sup er ti-Furga, G. (2002). F unctional organization of th e yeast proteome by systematic analysis of protein complexes. N atur e 415 141–147. Ga vin, A.-C . , Aloy, P. , G randi, P. , Kra use , R. , Boesche, M. , Marzioch, M. , Rau, C. , Jensen, L. J. , Bastuck, S. , D ¨ umpelfeld, B . , Edelmann, A. , Heur tier, M. , Hoffman, V. , Hoefer t, C. , Klein, K. , Hudak, M. , Michon, A. , Schelder, M. , S chirle, M. , Re m or, M. , Rudi, T. , Hooper, S. , Bauer, A. , Bouwmeester, T. , C asari, G. , Drewes, G. , Neubauer, G. , Rick, J. M . , Kuster, B. , Bork, P. , Russell , R. B. and S u per ti-Furga, G. (2006). Proteome survey reveals mo dularity of th e yeas t cell machinery . Natur e 440 631–636. Gelman, A. and Hill, J. (2007). Data Analysis Using Multilevel/Hier ar chic al Mo dels . Cam b rid ge U n iv. Press. Gentner, D. (1983). Structure- mapping: A th eoretical framew ork for analogy . Co gnitive Scienc e 7 155–170. Gentner, D. and Medi na, J. (1998). Similarit y and the developmen t of rules. Co gnition 65 263–297. Getoor, L. and T askar, B. (2007). Intr o duction to Statistic al R elational L e arning . MIT Press, Cambridge, MA. MR2391486 Getoor, L. , Friedma n , N. , Koller, D. and T askar, B. (2002). Learning probabilistic mod els of link structu re. J. Mach. L e arn. R es. 3 679–707. MR1983942 Ghahramani , Z. and Heller, K. A . (2005). Ba yes ian sets. A dvanc es in Neur al Infor- mation Pr o c essing Systems 18 435–44 2. Harbison, C. T. , Gordon, D. B. , Lee, T. I. , Rinaldi, N. J. , Macisaa c, K. D. , Danf ord, T. W . , Hannett, N. M. , T agne, J. B. , Rey nolds, D. B. , Yoo, J. , Jennings, E. G. , Zeitlinge r, J. , Pokholo k, D. K. , Kellis, M. , Rolfe, P. A. , T akusaga w a, K. T. , Land er, E. S. , Gifford, D. K. , Fraenkel, E. and Young, R. A. (2004). T ranscriptional regulatory cod e of a euka ryoti c genome. Na- tur e 431 99–104. Ho, Y. , Gruhler, A. , Heilbut, A. , Bad e r, G. D. , Moore, L. , Adams, S.- L. , Millar, A. , T a ylor, P. , Benn e tt, K. , Bouti li e r, K. , Y ang, L. , Wol t- ing, C. , Donaldson, I. , Schan dorff, S. , Shewnara n e, J. , Vo, M. , T aggar t, J. , Goudreaul t, M. , Muska t, B. , A lf arano, C. , Dew ar, D. , Lin, Z. , Micha l- ick o v a , K. , Willems, A. R ., Sassi, H. , Ni elsen, P. A. , R asmussen, K. J. , Andersen, J. R. , Johanse n, L. E. , Hansen, L. H. , Jespersen, H. , Podtele- jniko v, A . , Nie lsen, E. , Cra wfo rd, J. , Poulsen, V. , Sørense n, B. D. , He n - drickson, R. C. , Ma tthiesen, J. , Gle e son, F. , P a wson, T. , Moran, M. F. , Duro cher, D. , Mann, M. , Hogue, C. W. V. , Fige y s, D. an d Tyers, M. (2002). RANKI N G RELA TIONS USIN G A N ALOGIES 29 Systematic identification of protein complexes in saccharom yces cerevisiae by mass sp ectrometry . Natur e 415 180–183. Hoff, P. D. (2008). Mo deling homophily and sto chastic equiva lence in symmetric rela- tional d ata. A dvanc es i n Neur al Information Pr o c esing Systems 20 657–664. Holland, P. W. and Leinha rdt, S. (1975). Lo cal stru cture in so cial netw orks. In So ci- olo gic al Metho dolo gy (D . Heise, ed .) 1–45. Jossey-Bass, New Y ork. Huh, W. K. , F al vo, J. V. , G e rke, L. C. , Carrol l, A. S . , Howso n, R. W. , Weiss- man, J. S. and O’Shea E. K. (2003). Global analysis of protein localization in bu d ding yeas t. Natur e 425 686–691. It ˆ o, T. , T ashiro, K. , Mut a, S. , Oza w a, R. , Chiba, T. , Nishiz a w a, M. , Y am amoto, K. , Kuhara, S. and Sakaki, Y. (2000). T ow ard a protein–protein interaction map of the bu dding yeast: A compreh en sive system to examine tw o-hybrid interactio n s in all p ossible combinations b etw een the yeast proteins. Pr o c. Natl. A c ad. Sci. 97 1143–114 7. Jaakk ola, T. and Jordan, M . (2000). Bay esian p arameter estimation via vari ational metho d s. Stat. Comput. 10 25–37. Jensen, D. and Neville, J. (2002). Link age and autocorrelation cause feature selection bias in rela tional learning. In Pr o c. 19th International Confer enc e on M achine L e arning . Morgan Kaufmann, San F rancisco. Jensen, L. J. and Bork, P. (2008). Bio chemis try: N ot comparable, b ut complementa ry . Scienc e 322 56–57. Jord an, M. , Gha h ramani, Z. , Jaakkol a, T. and Saul, L. (1999). I ntro duction to v ariational meth ods for graphical mo dels. Machine L e arning 37 183–233. Kanehisa, M. and G oto, S. (2000). K EGG: Kyoto encyclop edia of genes and genomes. Nucleic A cids R ese ar ch 28 27–30. Kass, R. and Rafte r y, A. (1995). Ba yes factors. J. Amer. Statist. Asso c. 90 773–795. Kemp, C. , Tenen ba um, J. , G riffths, T . , Y am ada, T. and Ueda, N . (2006). Learning systems of concepts with an infinite relational mo del. In Pr o c e e dings of AAAI ’06 . MIT Press, Cambridge, MA. Ko k, S. and Domi ngos, P. ( 2007). S tatistical predicate inv ention. I n 24th I nternational Confer enc e on Machine L e arning 12 93–104. Omnipress, Madison, WI. Kro g an, N. J. , C agney, G. , Yu, H . , Z h ong, G. , Guo, X. , Igna tchenko, A. , Li, J. , Pu, S. , Da tt a, N. , Ti kuisis, A. P. , Punna, T. , Peregrin-Al v arez, J. M . , Shales, M. , Zhang, X. , Da vey, M. , R obinson, M. D. , P a ccan a ro , A. , Bra y, J. E. , Sheung, A. , Bea ttie, B. , Richard s, D. P. , Ca nadien, V. , Lalev, A. , Mena, F. , Wong, P. , St aro stine, A . , Canete, M. M. , Vlasblom, J. , Wu, S. , Orsi, C. , Collins, S. R. , Chand ran, S . , Ha w, R. , Rilstone, J. J. , Ga ndi, K. , Thomp- son, N. J. , Musso, G. , St. Onge, P. , Ghan ny, S. , M. Lam, H. Y. , Butland, G. , Al t af-Ul, A. M. , Kana y a, S. , Shila tif ard, A. , O’She a, E. , Weissman, J. S. , In- gles, C . J. , Hug h es, T. R. , P arkinson, J. , Gerste i n, M. , Wod ak, S. J. , Emili, A. and Greenbla tt, J. F. (2006). Global landscape of protein complexes in the yeast Saccharo m yces Cerevisiae. Natur e 440 637–64 3. Lorrain, F. and Wh ite, H. C. (1971). Structu ral equiv alence of individuals in social netw orks. Journal of Mathematic al So ciolo gy 1 49–80. Manning, C. , R agha v an, P. and Sch ¨ utze, H. (2008). Intr o duction to Information R e- trieval . Cambridge Univ. Press. Marx, Z. , Dagan, I. , Buhma nn, J. and Sh a mir, E. (2002). Coupled clustering: A meth o d for detecting structural correspond ence. J. Mach. L e arn. R es. 3 747–780 . MR1983945 Memisevi c, R. and Hinton, G. (2005). Multiple relational embed ding. In 18th NIPS . V ancouver, BC. 30 SIL V A, HELLER, GHAHRAMA NI AN D AIROLDI Mewes, H . et al. (2004). MIPS: A nalysis and annotation of p roteins from whole genome. Nucleic A cids R ese ar ch 32 D41–D44. Muggleton, S. (1981). Inv erting t h e resolution principle. Machine Intel ligenc e 12 93– 104. Myers, C. , Robso n , D. , Wible, A. , Hibbs, M . , Chiriac, C. , Theesfe ld, C. , Dolin- ski, K. and Tro y anska y a, O. (2005). D isco very of b iologica l netw orks from diverse functional genomic data. Genome Biolo gy 6 R114.1–R114 .16. Myers, C. L. , Barre t, D. A. , Hibbs, M. A. , Huttenhower, C. and Tro y an- ska y a, O. G. (2006). Finding function: An ev aluation framew ork for functional ge- nomics. BMC Genomics 7 187. Nabiev a, E. , Jim, K. , Agar w al, A. , Chaz elle, B. and S ingh. M. (2005). Whole- proteome prediction of protein function via graph-theoretic analysis of in teraction maps. Bioinformatics 21 i302–i310. No wicki, K. and Snijde rs, T. A. B. (2001). Estimation and pred iction for sto chastic blockstructures. J. Amer. Statist. Asso c. 96 1077–10 87. MR1947255 Popescul, A. and Unga r, L. H. (2003). Stru ctural logistic regression for link analysis. In Multi-R elational Data Mining Workshop at KDD-2003 92–106 . ACM Press, New Y ork . Primig, M. , W illiams, R. M. , Winzeler, E. A. , Tevzadz e, G . G . , Conw a y, A. R. , Hw ang, S. Y. , D a vis, R. W. and Esposito, R. E. ( 2000). The core meiotic transcrip- tome in budding yeasts. Natur e Genetics 26 415–423. Qi, Y. , B ar-Joseph, Z. and Klein-Se etharaman, J. (2006). Ev aluation of different biological data and computational classification metho ds for use in protein in teraction prediction. Pr oteins: Structur e, F unction, and Bioinformatics 63 490–500. Regul y, T., B reitkreutz, A., B oucher, L., Bre itkreutz, B.-J., Hon, G. , My- ers, C., P arsons, A., Friesen, H ., Oughtred, R., Tong, A., St ark, C., Ho, Y., Botstein, D., Andre ws, B., Boone, C., Tro y ansky a, O., Ideker, T., Dolin- ski, K., B a t ada, N . and T yers, M. (2006). Comprehensive cu ration and analysis of global interaction netw orks in saccharom yces cerevisiae. Journal of Biolo gy 5 11. Ro senbaum, P. (2002). Observational Studies . Springer, Berlin. MR1899138 Rumelhar t, D. and Abrahamson, A. (1973). A mod el for analogica l reasoning. Co gnitive Psycholo gy 5 1–28. SGD. Saccharom y ces genome d atabase. Ava ilable at ftp://ftp.y eastgenome.org/yea st/ . Sil v a, R. , Heller, K. A. and Ghahraman i , Z. (2007). Analogical reasoning with re- lational Ba yesi an sets. In 11th International Conf er enc e on Artificial Intel ligenc e and Statistics, AIST A TS . San Juan. Sil v a, R. , Heller, K. A . , Gha hramani, Z. and Airoldi, E. M . (2010). Su p plement to: “Rankin g relations using analogies in biological and information n etw orks.” D OI: 10.1214 /09-A OAS321SUPP . Stephens, M. , P alakal, M . , Mukhop adhy a y, S. , Raj e, R. and Most af a, J. (2001). Detecting gene relations from MEDLINE abstracts. In Pr o c e e dings of the Si xth Annual Pacific Symp osium on Bio c om puting 483–496 . W orld Scientific, Singap ore. T arasso v, K., Me ssier, V., Land r y , C. R. , Radinovic, S., Molina, M. M. S., Shames, I., Malitska y a, Y., Vogel, J., Bussey, H. and Michnick, S . W. (2008). An in vivo map of the yeas t protein interactome. Scienc e 320 1465–147 0. Tenenbaum, J. and Griffi ths, T. (2001). Generalization, similarit y , and Bay esian in- ference. Behavior al and Br ain Scienc es 24 629–641. TRANSF A C. T ranscription factor database. Av ailable at http://w ww.gene-regulatio n. com/ . Turney, P. (2008a). The laten t relation mapping engine: Algorithm an d exp eriments. J. Art ificial I ntel li genc e R es. 33 615–655. RANKI N G RELA TIONS USIN G A N ALOGIES 31 Turney, P. (2008b). A uniform approach to analogies, synonyms, anton yms, and asso cia- tions. In Pr o c e e dings of the 22nd International Confer enc e on Computational Linguistics (COLING-08) 905–91 2. Asso ciation for Computational Linguistics, Stroudsburg, P A . Turney, P. and Littman, M. (2005). Corpus-based learning of an alogies and semantic relations. M achine L e arning 60 251–278. Uetz, P. , Gi ot, L. , Cagney, G. , Mansfi eld, T. A. , Judson, R. S. , Knight, J. R. , Lockshon, D. , Nara y an, V. , Sriniv asan, M. , Pochar t, P. , Qureshi-Emili, A. , Li, Y. , Godwin, B. , C ono ve r, D. , Kalbfleisch, T. , Vi j a y adamodar, G. , Y ang, M. , Johnston , M. , Fie lds, S. and Rothber g, J. M. (2000). A comprehen- sive analysis of protein–protein interactions in saccharom yces cerevisiae. Natur e 403 623–627 . Veloso, M. and Carbonell, J. (1993). Deriv ational analogy in PR ODIGY: Au tomating case acqu isition, storage an d utilization. M achine L e arning 10 249–278. vo n Mering, C. , Krause, R. , Snel, B. , Cornell, M. , Oliver, S. G. , Fields, S. and Bork, P. (2002). Comparativ e assessment of large-scale data sets of protein–protein intera ctions. Natur e 417 399–403. W ang, X.-J. , Tu, X. , Feng, D. and Zhang, L. (2009). R anking community answ ers by mo deling question–answer relationships via analogical reasoning. In Pr o c e e dings of the 32nd Annual ACM SIGIR Confer enc e on R ese ar ch & Development on Inf ormation R etrieval . A ssociation for Computing Machinery , New Y ork. Xu, Z. , Tresp, V. , Yu, K. and Krie gel, H.-P. (2006). In finite hidden relational models. In Pr o c e e dings of the 22nd Confer enc e on Unc ertainty in Art ificial Intel l igenc e . Morgan Kaufmann, San F rancisco, CA. Yu, H., Braun, P., Yildi ri m, M. A., Lem m ens, I., Venka tesan, K., Saha lie, J., Hirozane-Kishika w a, T. , Ge breab, F., Li, N., Simonis, N., Hao, T., Rual, J.-F., Dricot, A ., V az que z, A., Murra y, R. R., Simon, C. , T ardivo, L., T am, S., Svrzikap a, N., F an, C., de Smet, A.-S., Motyl, A., Hudson, M. E., P ark, J., Xin, X., Cu sick, M. E., Moore, T., Boone, C . , Sny der, M., Roth, F. P., Barabasi, A.-L., T a vern ier, J., Hill, D. E. and V idal, M. (2008). High-q uality binary protein interaction map of the yeast interactome netw ork. Scienc e 322 104–11 0. Yver t, G. , Brem, R. B. , Whittle, J. , Akey, J. M. , Fo ss, E. , Smith, E. N. , Mack elprang, R . and Krugl y ak, L. (2003). T rans-acting regulatory v ariation in sacc h aromyces cerevisiae and the role of transcription factors. Natur e Genetics 35 57– 64. Zhu, J. and Zhang, M . Q. ( 1999). SCPD: A p romoter database of the yeast S acc h a- rom yces cerevisiae. Bioinf ormatics 15 607–61 1. R. Sil v a University College London Gower S treet London, WC1E 6 BT United Kingdom E-mail: ricardo@stats.ucl.ac.uk K. Heller Z. Ghah ra mani University of Camb ridge Trumpington S treet Cambridge, CB2 1PZ United Kingdom E-mail: heller@gatsb y .ucl.ac.uk zoubin@eng.cam.ac.uk E. M. Airoldi Har v ard University 1 Oxford street Cambridge, Massachusetts 02138 USA E-mail: airoldi@fas.harv ard.edu
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment