Higher Order Programming to Mine Knowledge for a Modern Medical Expert System
Knowledge mining is the process of deriving new and useful knowledge from vast volumes of data and background knowledge. Modern healthcare organizations regularly generate huge amount of electronic data stored in the databases. These data are a valua…
Authors: Nittaya Kerdprasop, Kittisak Kerdprasop
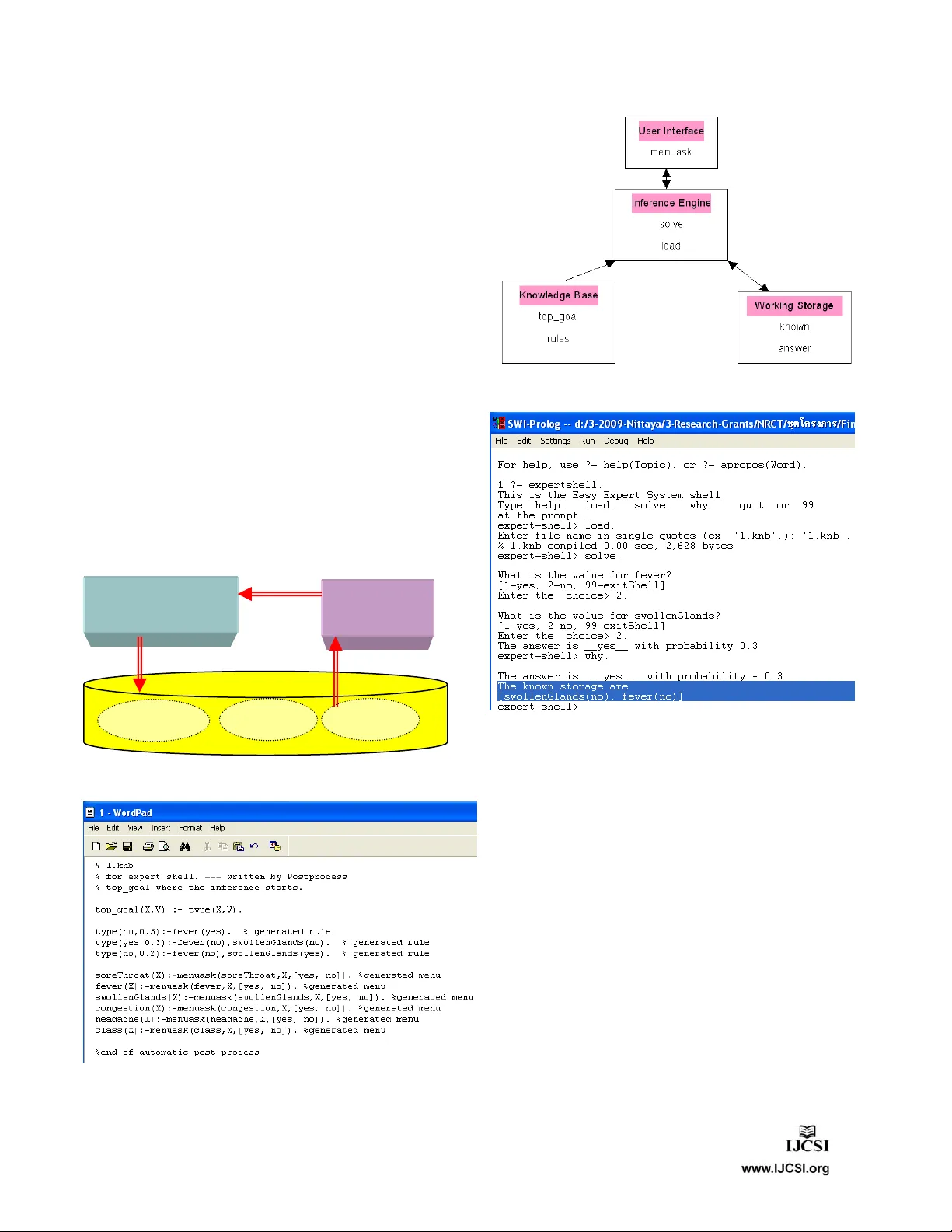
IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 64 Higher Order Programming to Mine Knowledge for a Modern Medical Expert System Nittaya Kerdprasop and Kittisak Kerdprasop Data Engineering and Knowledge Discovery (DEKD) Research Unit, School of Computer Engineerin g, Suranaree University of Techn ology, Nakhon Ratchasima 30000, Thailand Abstract Knowledge mining is the proce ss of deriving new and useful knowledge from vast volumes of data and background knowledge. Modern healthcare organizations regularly generate huge amount of electronic data stored in the databases. These data are a valuable resource for mining useful knowledge to help medical practitioners making appropriate and accurate decision on the diagnosis and treatment of diseases. In this paper, we propose the design of a novel medi cal expert system based on a logic-programming framework. The proposed system includes a knowledge-mining component as a repertoire of tools for discovering useful knowledge. The implementation of classification and association mining tools based on the higher order and meta-level progra mming schemes us ing Prolog has been presented to express the power of logic-based language. Such language also provides a pattern matching facility, which is an essential function for the de velopment of knowledge-intensive tasks. Besides the major goal of medical decision support, the knowledge discovered by our logic-based knowledge-mining component can also be deployed as background knowledge to pre-treatment data from other sources as well as to guard the data repositories against constraint violation. A framework for knowledge deployment is also presented. Keywords: Knowledge Mining, Association Mining, Decision- tree Induction, Higher-order Logic Programming, Medical Expert System. 1. Introduction Knowledge is a valuable asset to m ost organizations as a substantial source to support better decisions and thus t o enhance organizational competency. Researchers and practitioners in the area of knowledge management view knowledge in a broad sense as a state of m ind, an object, a process, an access to informa tion, or a capability [2, 13]. The term knowledge asset [24, 26] is used t o refer to any organizational int angible property relat ed to knowledge such as know-how, expertise, intellectual propert y. In clinical companies and computerized healthcare organizations knowledge assets include order set s, drug- drug interaction ru les, guidelines for practitioners, and clinical protocols [12] . Knowledge assets can be stored in data repositories either in implic it or explicit form . Explicit knowle dge can be managed through the exis ting tools available in the current database technology. Im plicit knowl edge, on the contrary, is harder to achieve and retrieve. Specific tools and suitable environments are needed to extract such knowledge. Implicit knowledge acquisition can be achieved through the availability of the knowledge-mining system. Knowledge mini ng is the discovery of hidden knowledge stored possibly in various fo rms and places in large data repositories. In health and m edical domai ns, knowledge has been discovered in different form s such as association rules, classification trees, clusteri ng means, trend or temporal patt erns [27]. The discovered knowledge facilitates expert decisi on support, diagnosis and prediction. It is t he current trend in the design and development of deci sion support system s [3, 16, 20, 31] to incorporate knowledge discovery as a tool to extract implicit information. In this paper we present the design of a m edical expert system and the im plementat ion of knowledge mi ning component. Medi cal data mi ning is an emerging area of computational intellig ence applied to au tomatically analyze electronic medical reco rds and health databases. The non-hypothesis driven analysis approach of data mining technology can induce knowledge from clini cal data repositories and health dat abases. Induced knowledge such as breast cancer recurre nce conditions or diabet es implication is important not only to increase accurate diagnosis and successful treat ment, but also to enhance safety and reduce medication-related errors. A rapid prototyping of the proposed system is demonstrated in the paper t o highlight the fact that hi gher order and meta-level programmi ng are suitable schemes to IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 65 implem ent a complex knowl edge-intensive system . For such a complicated sy stem program coding should be done declaratively at a high abstr action level to alleviate th e burden of programmers and t o ease reasoning about program sem antics. The rest of this paper is organized as follows. Section 2 provides some prelim inaries on two m ajor knowledge- mining tasks, i.e. classi fication and association m ining. Section 3 proposes the medi cal expert system design framework with t he knowledge-mini ng component. Running examples on me dical da ta set and the illustration on knowledge deploym ent are presented in Sect ion 4. Section 5 discusses related work and t hen conclusions are drawn in Section 6. The im plementat ion of knowledge- mining com ponent is presented i n the Appendix. 2. Preliminaries on Tree-based Classification and Association Mining Decision tree induction [21] is a popular me thod for mining knowledge from medical dat a and representing the result as a classifier tree. Popularit y is due to the fact that mining result in a form of decision tree is interpretability, which is more concern among m edical practitioners than a sophisticated method but l ack of understandability. A decision tree is a hierarchi cal structure with each node contains decision attri bute and node branches corresponding to different attri bute values of the decision node. The goal of building decision tree is to partition data with mixing classes down the tree until each leaf node contains data with pure class. In order to build a decision tree, we need to choose t he best attribute that contri butes the most t owards partitioning data to the purity groups. The m etric to measure attribute’s ability to partition data into p ure class is Info , which is the number of bi ts required to encode a data mixt ure. The metric Info of positive (p) and neg ative (n) data mixture can be calculates as: Inf o(P (p), P(n )) = P(p)log 2 P(p) P(n)log 2 P(n) . The symbols P(p) and P(n) are probabilities of pos itive and negative data instances, respectively. The symbol p represents number of pos itive data instances, an d n is the negative cases. To choose the best attribute we have to calculate information gain, which is th e yield we obtained from choosing that att ribute. The informati on gain calculation of data with two classes (positive and negative) is given as: Gain(Attribute) = Info{p/(p+n), n/(p+n)} i=1 to v {(p i +n i )/(p +n)} In fo{ p i /( p i +n i ), n i /( p i +n i) } . The information gain calculates yield on Info of data set before splitting and Info after choosing attribute with v splits. The gain value of each candidate attribute is calculated, and then the maxim um one has been chosen to be the decision node. The pr ocess of data partitioning continues until th e data subset has the same class label. Classification t ask based on decision-tree i nduction predicts the va lue of a target attribute or class, whereas association-m ining task is a generalizat ion of classification in that any attribute in th e data set can be a target attribute. Association m ining is the discovery of frequent ly occurred relationships or correlations between att ributes (or item s) in a database. Association mining problem can be decomposed as (1) find all sets of items that are frequent patterns, (2) use the frequent pattern s to generate rules. Let I = {i 1 , i 2 , i 3 , ... , i m } be a set of m items and DB = { C 1 , C 2 , C 3 , ..., C n } be a database of n cases and each case contains items in I . A pattern is a set of items that occur in a case. The number of items in a pattern is called the leng th of the pattern. To search for all valid patte rns of length 1 up to m in large database is comput ational expensive. For a set I of m different items, the search space of all distinct patterns can be as huge as 2 m -1. To reduce the size of the search space, the support measurement has been introduced [1]. The function support ( P ) of a pattern P is defi ned as a number of cases in DB containing P . Thus, support ( P ) = |{ T | T DB, P T }|. A pattern P is called frequent pattern if the support value of P is not less than a predefi ned minim um support threshold minS . It is the minS constraint that helps reducing the com putational com plexity of frequent patt ern generation. The minS met ric has an anti-m onotone property such that if the pattern co ntains an item that is not frequent, then none of the pattern’s supersets are frequent. This property helps reducing the search space of mining frequent patterns in algori thm Apriori [1] . In this paper we adopt this al gorithm as a basis for our implem entation of association m ining engine. 3. Medical Expert System Framework and the Knowledge Mining Engines 3.1 System Architecture Health information is normally distributiv e and heterogeneous. Hence, we design the me dical expert system (Figure 1) t o include data integration com ponent at the top level to coll ect data from dist ributed databases and also from docum ents in text form at. IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 66 Medical Expert System Data Integration Patient records Clinical data & Other documents Data warehouse Knowledge Mining Knowledge induced Evaluation knowledge background knowledge Knowledge Base OLAP Tools Request/query Knowledge inferring and reasoning Response Medical practitioner Fig. 1 Knowledge-mining com ponent and a m edical expert system framework. Double line ar rows are process flow, whereas the dash line arrows are data flow. The data integration com ponent has been designed to input and select data with natural l anguage processing. Data at this stage are to be stored in a warehouse to support direct querying (through OLAP tools) as well as to perform analyzing wit h knowledge mining engi ne. Knowledge base in our design stores both induced knowledge in whi ch its si gnificance has to be evaluated by the domain expert, and background knowle dge encoded from hum an experts. Knowledge i nferring and reasoning is the module interfacing with medical practitioners and physicians at the front-end and accessing knowledge base at the back-end. The focus of this paper is on t he implem entation of knowl edge-mining com ponent, which currently contains classificati on and association mi ning engine. 3.2 Classification Mining Tool Our classificati on mining engi ne is the impl ementation of decision-tree induction (ID3) al gorithm [21]. The steps in our implementation are presented as follows: Algorithm 1 Classificat ion mining engi ne Input: a data set formatt ed as Prolog clauses Output: a decision tree with node and edge structures (1) Initialization (1.1) Clear tem porary knowledge base (KB) by removing all information regardi ng the predicates node, edge and current_node (1.2) Set node counter = 0 (1.3) Scan data set to get inform ation about data attributes, positive instan ces, negative instances, total data instances (2) Building tree (2.1) Increment node counter (2.2) Repeat steps 2.2.1-2.2.4 u ntil there is no more attributes left for creat ing decision attribut es (2.2.1) Compute the Info value of each candidate attribute (2.2.2) Choose the attribute that yields mi nimum Info to be decision node (2.2.3) Assert edge and node informati on into the knowledge base (2.2.4) Split data instances along node branches (2.3) Repeat steps 2.1 and 2.2 until the lists of positive and n egative instances are empty (2.4) Output a tree structure that cont ains node and edge predicates The program source code is based on the synt ax of SWI prolog (www.swi-prolog.org). main :- init(AllAttr, EdgeList), % initialize node % and edge structures getNode(N), % get node sequence number create_edge(N, AllAttr, EdgeList), % recursively create tree print_model. % print tree model Classification m ining engine is composed of t wo files main and id3 . The main module (m ain.pl) calls initialization procedu re (init) and starts creating edges and nodes of the decision tree. The data (data.pl) to be used by main m odule to create deci sion tree is also in a format of Prolog file. The m ining engine induces data model of two classes: positive (class = yes) and neg ative (class = no). Binary classification is a typi cal task in medical dom ain. The code can be easily modified to classify data with more than two classes. 3.3 Association Mining Tool The implem entation of associat ion mining engi ne is based primaril y on the concept of higher-order Horn clauses. Such concept has been utili zed through the predicates maplist , include , and setof . IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 67 The extensive use of these predicates contributes significantly to program conciseness and the ease of program verification. The program produces frequent patterns as a set of co-occurring items. To generate a nice representation of association rule such as X => Y , the list L in the predicate association_mining has to be furt her processed. association_mining :- min_support(V), % set minimum support makeC1(C), % create candidate 1-itemset makeL(C,L), % compute large itemset apriori_loop(L,1). % recursively run apriori makeC1(Ans):- input(D), % input data as a list allComb(1, ItemList, Ans2), % make combination of itemset maplist(countSS(D),Ans2,Ans). % scan database and pass countSS % to maplist makeC(N, ItemSet, Ans) :- input(D), allComb(2, ItemSet, Ans1), maplist(flatten, Ans1, Ans2), maplist(list_to_ord_set, Ans2, Ans3), list_to_set(Ans3, Ans4), include(len(N), Ans4, Ans5) , % include is % also a higher-order predicate maplist(countSS(D), Ans5, Ans). % scan database to find: List+N 4. Running Examples and Knowledge Deployment To show the running exam ples of our program coding, we use the following simple medical data represented as a Prolog file. %% Data set: Allergy diagnosis % Symptoms of disease and their possible values attribute( soreThroat, [yes, no]). attribute( fever, [yes, no] ). attribute( swolle nGlands, [yes, no]). attribute( congestion, [y es, no]). attribute( headache, [yes, no]). attribute( class, [yes, no]). % Data instances instance(1, class=no, [soreThroat=yes, fever=ye s, swollenGlands=yes, congesti on=yes, headache=yes]). instance(2, class=yes, [soreThroat=no, fever=no, swollenGlands=no, congestion=y es, headache=yes]). instance(3, class=no, [soreThroat=yes, fever=ye s, swollenGlands=no, congestion=y es, headache=no]). … Data as shown are patient records suffering from allergy (class=yes). There are ten patient records in this simple data set: patient IDs 2, 6, and 8 are those who are suffering from allergy , whereas patient IDs 1, 3, 4, 5, 7, 9, 10 are suffering from other di seases but has shown some basic symptoms similar to allergy patients. To induce classification model for allergy patients from this data, we have to save this data set as a Prolog file (data.pl) and include this file name at the header declaration of the main program. By calling predicate main , the syst em should respond as true . At this moment we can view the tree model by cal ling list ing(node) , then l isting(edge) and get the following results. 1 ?- main. true. 2 ?- listing(node). :- dynamic user: node/2. user:node(1, [2, 6, 8]-[1, 3, 4, 5, 7, 9, 10]). user:node(2, []-[1, 3, 5, 9, 10]). user:node(3, [2, 6, 8]-[4, 7]). user:node(4, []-[4, 7]). user:node(5, [2, 6, 8]-[]). true. 3 ?- listing(edge). :- dynamic user: edge/3. user:edge(0, root-nil, 1). user:edge(1, fever-yes, 2). user:edge(1, fever-no, 3). user:edge(3, swollenGlands-y es, 4). user:edge(3, swollenGla nds-no, 5). true. The node and edge structures have the following form ats: node(nodeID, [Positive_Cases]-[Negative_Cases] ) edge(ParentNode, EdgeLabel, Chil dNode) The node structure is a tuple of nodeID and a m ixture of positive and negative cases represen ted as a list pattern: [Positive_Cases]-[Negative_Cases] . Node 0 is a special node, representing root node of the tree. Node 1 contains a mixture of t en patients, whereas node 5 is a pure group of allergy pat ients. The edges leadi ng from node 1 to node 5 capture the model of allergy patients. Therefore, th e classification result represents th e following data model: class(allergy) :- fever=no, swollenGlands=no. This model is represented as a Horn clause, thus, it provide flexib ility of including this clause as a rule to select data in other group of patients who are sufferi ng from throat infect ion. This kind of infection shows the same basic symptom s as allerg y; therefore, screening data with the above rule can help focusing onl y on throat infection cases. IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 68 Applying the same data set wi th association mi ning and setting mini mum support value = 50%, we got the following frequent patt erns: {fever=yes & class=no} {fever=yes & congestion=yes} {swollenGlands=no & congestion=y es} {congestion=yes & headache=yes} {congestion=yes & class=no} {fever=yes & congestion=yes & class=no} The first pattern can be interpreted as association rule as “ if patient has fever, that the patient does not suffer from allergy. ” This kind of rule can help accurately diagnosing patients with symptoms very close to allergy. Knowledge Deployment: E xample 1. We suggest that such discovered rules, aft er confirmi ng their correctness by hum an experts, can be added into the database system as trigger rules (Figure 2). The triggers guard database content against any updates t hat violates the rules. Any attempt to insert violating data will raise an error message to draw attention from the database administrat or. Such trigger rules are thus deployed as a tool to enforce database in tegrity checking. induced rules Trigger Generation Mining Component Component aggregated data Trigger rules Knowledge Data Fig. 2 The fram ework of knowledge depl oym ent as triggers in a medical database. Fig. 3 The content of automatically induced knowledge base. Fig. 4 Structure of a simple expert system shell with the induced knowledge base. Fig. 5 A snapshot of medical expert sy stem inductively created fr om the allergy data set. Knowledge Deployment: E xample 2. The induced knowledge once confirm ed by the domain expert can be added to the knowledge base of the expert system shell. We illustrate the knowledge base that automatically created from th e induced tree in Figure 3. This expert system shell has simple structure as diagrammatically shown in Figure 4. User can interact with the system through a line com mand as shown in Figure 5, in whi ch the user can ask for further explanati on by typing the ‘why ’ comm and. 5. Related Work In recent years we have witnessed increasing number of applications devising dat abase technology and m achine learning techniques to mine knowle dge from biom edicine, clinical and healt h data. Roddick et al [22] discussed t he two categories of m ining techniques appli ed over medical IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 69 data: explanatory and explorat ory. Explanatory m ining refers to techniques that are used for the purpose of confirmati on or ma king decisions. Exploratory mining is data investigation norm ally done at an early stage of data analysis in which an exact mining objective has not yet been set. Explanatory m ining in me dical data has been extensively studied in the past decade employing vari ous learning techniques. Bojarczuk et al [4] applied geneti c programm ing method wi th constrained syntax to discover classification rul es from me dical data sets. Thongkam et al [28] studied br east cancer survivability using AdaBoost algorithm. Ghazavi and Liao [9] proposed the idea of fuzzy modeling on sel ected features of medical data. Huang et al [11] introduced a system to apply mi ning techniques to discover rule s from health exam ination data. Then they em ployed a case-based reasoning to support the chronic disease diagnosis and treatments. The recent work of Zhuang et al [31] also com bined mining wit h case- based reasoning, but applied a different mining m ethod. They performed data clusteri ng based on self-organizing maps in order to facilitate decision support on solving new cases of pathology test orderi ng problem. Bi omedical discovery support systems ar e recently proposed by a number of researchers [5, 6, 10, 29, 30]. Som e work [20, 25] extended me dical databases to the level of data warehouses. Exploratory, as oppose to explanatory , is rarely applied to medical dom ains. Among t he rare cases, Nguyen et al [19] introduced knowledge visuali zation in the study of hepatitis patients. Palaniappan and Ling [20] applied th e functionality of OLAP tools to improve visualization in data analysis. It can be seen from the literature that most medical knowledge discovery system s have applied only some mining t echniques to discover hidden knowledge wit h the main purpose to support medical di agnosis [4, 14, 17] . Some researchers [3, 8, 15, 16] have exte nded the knowledge discovery aspect to the large scal e of a medical decision support system . Our work is also in the main stream of medical decision support system developm ent, but our m ethodology is different from those appeared in the literature. Th e system proposed in this paper is based on a logic-programm ing paradigm. The justifi cation of our logic-based system is that the closed form of Horn clauses that treats program in the same way as data facilitates fusion of knowledge learned from different sources, which is a normal setti ng in medical domain. Knowledge reuse can easily practice in this framework. The declarative style of our implementation also eases the future extension of the proposed medi cal support system to cover the concepts of higher-order mini ng [23], i.e. mining from the discovered knowledge, and constraint mining [7] , i.e. mining wi th some specified constrai nts to obtain relevant knowle dge. 6. Conclusions and Discussion Modern healthcare organizati ons generate huge amount of electronic data stored in hete rogeneous databases. Data collected by hospitals and clinics are not yet turn ed into useful knowledge due to the lack of efficient analys is tools. We thus propose a rapid prototy ping of automa tic mining t ools to induce knowledge from medical data. The induced knowledge is to be evaluat ed and integrated into the knowledge base of a medical expert sy stem. Discovered knowledge facilitates the reuse of knowledge base among decision-support appl ications withi n organizations that own het erogeneous clinical and health databases. Direct applicati on of the proposed system is for medical rel ated decision-m aking. Other indirect but obvious application of such knowl edge is to pre-process other data sets by groupi ng it into focused subset containing only rele vant data instances. The main contri bution of this work is our implem entation of knowledge mini ng engines based on the concept of higher-order Horn clauses using Prolog language. Hi gher- order programming has been original ly appeared in functional languages in whi ch functions can be passed as arguments to other functi ons and can also be returned from other functi ons. This style of program ming has soon been ubiquitous in several m odern programm ing languages such as Perl, PHP, and JavaScri pt. Higher order style of program mi ng has shown the outstanding benefits of code reuse and high level of abstracti on. This paper illustrates higher order prog rammi ng techniques in SW I- Prolog. The powerful feature of met a-level programm ing in Prolog facilitates the reuse of mining results rep resented as rules to be flexibly ap plied as conditional clauses in other applications. The plausible extensions of our current work are to add constraints int o the knowledge mini ng method in order t o limit the search space and theref ore yield the m ost relevant and timely knowledge, and due to the uniform representation of Prolog’s statements as a clausal form, mining from the previously mi ned knowledge should be implem ented naturally. We also pl an to extend our system to work with stream data that normally occur in modern medical organi zations. IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 70 Appendix The implem entation of knowl edge-mining component i s based on the concept of higher-order and m eta- programm ing styles. Hi gher-order programmi ng in Prolog refers to Horn clauses that can quantify over other predicate symbol s [18]. Meta-level program ming i s also another powerful feature of Prolog. Dat a and program in Prolog take the same representation al format; that is clausal form. Higher-order and meta-level clauses in the following source code are typed in bold face. /* Classification mining engine */ :- include('data.pl'). :- dynamic current_node/1, node/2, edge/3. main :- init(AllAttr, EdgeList), getNode(N), % get node sequence number create_edge(N, AllAttr, EdgeList), print_model. init(AllAttr, [root-nil/PB-NB]) :- retractall(node(_, _)), retractall(current_node(_)), retractall(edge(_, _, _)), assert(current_node( 0)) , findall(X, attribute(X, _), AllAttr 1) , delete(AllAttr1, class, AllAttr), findall(X 2 , instance(X 2 , class=yes, _), PB), findall(X 3 , instance(X 3 , class=no, _), NB). getNode(X) :- current_node(X), X1 is X+1, retractall(current_node(_)), assert(current_node(X1)). create_edge(_, _, []) :- !. create_edge(_, [], _) :- !. create_edge(N, AllAttr, EdgeList) :- create_nodes(N, AllAttr, EdgeList). create_nodes(_, _, []) :- !. create_nodes(_, [], _) :- !. create_nodes(N, AllAttr, [H1-H2/PB-NB|T]) :- getNode(N1), % get node sequence number N1 assert(edge(N, H 1- H 2 , N 1)) , % H1-H2 is % a pattern assert(node(N 1 , PB-NB)), % PB-NB is % a pattern append(PB, NB, AllInst), ((PB \== [], NB \== []) -> % if-condition % then clauses (cand_node(AllAttr, AllInst, AllSplit), best_attribute(AllSplit, [V, MinAttr, Split]), delete(AllAttr, MinAttr, Attr2), create_edge( N1, Attr2, Split)) ; % else clause true ), create_nodes(N, AllAttr, T). % % select best attribute to be a decision node % best_attribute([], Min, Min). best_attribute([H|T], Min) :- best_attribute(T, H, Min). best_attribute([H|T], Min0, Min) :- H = [V, _, _ ], Min0 = [V0, _, _ ], ( V < V0 -> Min1 = H ; Min1 = Min0), best_attribute(T, Min1, Min). % % generate candidate decision node % cand_node([], _, []) :- !. cand_node(_, [], []). cand_node([H|T], CurInstL, [[Val,H,SplitL] |OtherAttr]) :- info(H, CurInstL, Val, SplitL), cand_node(T, CurInstL, OtherAttr). % % compute Info of each candidate node % info(A, CurInstL, R, Split) :- attribute(A,L), maplist(concat 3( A,=), L, L 1) , suminfo(L1, CurInstL, R, Split). concat3(A,B,C,R) :- atom_concat(A,B,R1), atom_concat(R1,C,R). suminfo([],_,0,[]). suminfo([H|T], CurInstL, R, [Split | ST]) :- AllBag = CurInstL, term_to_atom(H1, H), findall(X 1 , (instance(X 1 , _, L 1) , member(X 1 , CurInstL), member(H 1 , L 1)) , BagGro), findall(X 2 ,(instance(X 2 , class=yes, L 2) , member(X 2 , CurInstL), member(H 1 , L 2)) , BagPos), findall(X 3 ,(instance(X 3 , class=no, L 3) , member(X 3 , CurInstL), member(H 1 , L 3)) , BagNeg), (H11= H22) = H1, length(AllBag, Nall), length(BagGro, NGro), length(BagPos, NPos), length(BagNeg, NNeg), Split = H11-H22/BagPos-BagNeg, suminfo(T, CurInstL, R1,ST), ( NPos is 0 *-> L1 = 0; L1 is (log(NPos/NGro)/log(2)) ), ( 0 is NNeg *-> L2 = 0; L2 is (log(NNeg/NGro)/log(2)) ), ( NGro is 0 -> R= 999; R is (NGro/Nall)* (-(NPos/NGro)* L1- (NNeg/NGro)*L2)+R1). /* ========================= */ /* Association mining engine */ IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 71 /* ========================= */ association_mining:- min_support(V), % set minimum support makeC1(C), % create candidate 1-itemset makeL(C,L), % compute large itemset apriori_loop(L,1). % recursively run apriori apriori_loop(L, N) :- % base case of recursion length(L) is 1,!. apriori_loop(L, N) :- % inductive step N1 is N+1, makeC(N1, L, C), makeL(C, Res), apriori_loop(Res, N1). makeC1(Ans):- input(D), % input data as a list, % e.g. [[a], [a,b]] % then make combination of itemset allComb(1, ItemList, Ans2), % scan database and pass countSS to maplist maplist(countSS(D),Ans2,Ans). makeC(N, ItemSet, Ans) :- input(D), allComb(2, ItemSet, Ans1), maplist(flatten, Ans1, Ans2), maplist(list_to_ord_set, Ans2, Ans3), list_to_set(Ans3, Ans4), include(len(N), Ans4, Ans5) , % include is also a % higher-order predicate maplist(countSS(D), Ans5, Ans). % scan database to find: List+N makeL(C, Res):- % for all large itemset creation % call higher-order predicates % include and maplist include(filter, C, Ans), maplist(head, Ans, Res). % % filter and head are for pattern matching of % data format % filter(_+N):- input(D), length(D,I), min_support(V), N>=(V/100)*I. head(H+_, H). % % an arbitrary subset of the set containing % given number of elements % comb(0, _, []). comb(N, [X|T], [X|Comb]) :- N>0, N1 is N-1, comb(N1, T, Comb). comb(N, [_|T], Comb) :- N>0, comb(N, T, Comb). allComb(N, I, Ans) :- setof(L, comb(N, I, L), Ans). countSubset(A, [], 0). countSubset(A, [B|X], N) :- not(subset(A, B)), countSubset(A, X, N). countSubset(A, [B|X], N) :- subset(A, B), countSubset(A, X, N1), N is N1+1. countSS(SL, S, S+N) :- countSubset(S, SL, N). Acknowledgments This work has been fully supporte d by research fund from Suranaree University of Technology granted t o the Data Engineering and Knowledge Discovery (DEKD) research unit. This research is also support ed by grants from the National Research Counci l of Thailand (NRCT) and the Thailand Research Fund (TRF). References [1] R. Agrawal, and R. Srikant, “Fast algorithm for mining association rules”, in: Proc. VLDB , 1994, pp.487-499. [2] M. Alavi, and D.E. Le idner, “Review: Knowledge management and knowledge management systems: Conceptual foundations and research issues”, MIS Quarterly , Vol.25, No.1, 2001, pp.107-136. [3] Y. Bedard et al., “Integrating GIS components with knowledge discovery technology for environmental health decision support”, Int. J Medical Informatics , Vol.70, 2003, pp.79-94. [4] C.C. Bojarczuk et al., “A constrained-syntax genetic programming system for discove ring classification rules: Application to medical data sets”, Artificial Intelligence in Medicine , Vol.30, 2004, pp.27-48. [5] C. Bratsas et al., “KnowBaSICS-M: An ontology-based system for semantic ma nageme nt of medical problems and computerised algorithmic solutions”, Computer Methods and Programs in Biomedicine , Vol.83, 2007, pp.39-51. [6] R. Correia et al., “Borboleta: A mobile telehealth system for primary homecare”, in: Proc. ACM Symposium on Applied Computing , 2008, pp.1343-1347. [7] L. De Raedt et al., “Constraint programming for itemset mining”, in: Proc. KDD , 2008, pp.204-212. [8] E. German et al., “An architecture for linking medical decision-support applications to clinical databases and its evaluation”, J. Biomedical Informatics , Vol.42, 2009, pp.203-218. [9] S. Ghazavi and T.W. Liao, “M edical data mining by fuzzy modeling with selected features”, Artificial Intelligence in Medicine , Vol.43, No.3, 2008, pp.195-206. [10] D. Hristovski et al., “Using literature-based discovery to identify disease candidate genes”, Int. J Medical Informatics , Vol.74, 2005, pp.289-298. IJCSI International Journal of Com puter Science Issues, Vol. 8, Issue 3, No. 1, May 2011 ISSN (Online): 1694-0814 www.IJCSI.org 72 [11] M.J. Huang et al., “Integra ting data mining with case-based reasoning for chronic diseases prognosis and diagnosis”, Expert Systems with Applications , Vol.32, 2007, pp.856-867. [12] N.C. Hulse et al., “Towards an on-demand peer feedback system for a clinical knowledge base: A case study with order sets”, J Biomedical Informatics , Vol.41, 2008, pp.152- 164. [13] N.K. Kakabadse et al., “F rom tacit knowledge to knowledge management: Leveraging invisible assets”, Knowledge and Process Management , Vol. 8, No. 3, 2001, pp.137-154. [14] E. Kretschmann et al., “A utomatic rule generation for protein annotation with the C4.5 data mining algorithm applied on SWISS-PROT”, Bioinformatics , Vol.17, No.10, 2001, pp.920-926. [15] P.-J. Kwon et al., “A study on the web-based intelligent self-diagnosis medical system”, Advances in Engineering Software , Vol.40, 2009, pp.402-406. [16] C. Lin et al., “A decision support system for improving doctors’ prescribing behavior”, Expert Systems with Applications , Vol.36, 2009, pp.7975-7984. [17] E. Mugambi et al., “Polynomial-fuzzy decision tree structures for classifying medical data”, Knowledge-Based System , Vol.17, No.2-4, 2004, pp.81-87. [18] G. Nadathur, and D. Miller, “Higher-order Horn clauses”, J ACM , Vol.37, 1990, pp.777-814. [19] D. Nguyen et al., “Knowledge visualization in hepatitis study”, in: Proc. Asia-Pacific Symposium on Information Visualization , 2006, pp.59-62. [20] S. Palaniappan, and C.S. Ling, “Clinical decision support using OLAP with data mining”, Int. J Computer Science and Network Security , Vol.8, No.9, 2008, pp.290-296. [21] J.R. Quinlan, “Induction of decision trees”, Machine Learning , Vol.1, 1986, pp.81-106. [22] J.F. Roddick et al., “Exploratory medical knowledge discovery: experiences and issues”, ACM SIGKDD Explorations Newsletter , Vol.5, No.1, 2003, pp.94-99. [23] J.F. Roddick et al., “Higher order mining”, ACM SIGKDD Explorations Newsletter , Vol.10, No.1, 2008, pp.5-17. [24] C.P. Ruppel, and S.J. Harrington, “Sharing knowledge through intranets: A study of organizational culture and intranet implementation”, IEEE Transactions on Professional Communication , Vol.44, No.1, 2001, pp.37-51. [25] T.R. Sahama, and P.R. Croll, “A data warehouse architecture for clinical data warehousing”, in: Proc. 12 th Australasian Symposium on ACSW Frontiers , 2007, pp.227- 232. [26] A. Satyadas et al., “Know ledge management tutorial: An editorial overview”, IEEE Transactions on Systems, Man and Cybernetics , Part C, Vol.31, No.4, 2001, pp.429-437. [27] A. Shillabeer, and J.F. Roddick, “Establishing a lineage for medical knowledge discovery”, in: Proc. 6 th Australasian Conf. on Data Mining and Analytics , 2007, pp.29-37. [28] J. Thongkam et al., “Breast cancer survivability via AdaBoost algorithms”, in: Proc. 2 nd Australasian Workshop on Health Data and Knowledge Management , 2008, pp.55- 64. [29] N. Uramoto et al., “A te xt-mining system for knowledge discovery from biomedical documents”, IBM Systems J , Vol.43, No.3, 2004, pp.516-533. [30] X. Zhou et al., “Text mining for clinical Chinese herbal medical knowledge discovery”, in: Proc. 8 th Int. Conf. on Discovery Science , 2005, pp.396-398. [31] Z.Y. Zhuang et al., “Combi ning data mining and case-based reasoning for intelligent decision support for pathology ordering by general practitioners”, European J Operational Research , Vol.195, No.3, 2009, pp.662-675. Nitt aya Kerd prasop is an associate professor at the school of computer engineering, Suranaree University of T echnology , Thailand. She received her B.S. in radiation techniques from Mahidol University , Thailand, in 1985, M.S. in computer science from the Prince of Songkla University , Thailand, in 1991 and Ph.D. in computer science from Nova Southeastern University , USA, in 1999. She is a member of IAENG , ACM, and IEEE Computer Society . Her research of interest includes Knowledge Discovery in Databases, Data Mining, Arti ficial Intelligence, Logic and Constraint Programming, Deduc tive and Active Databases. Kittisak Kerd prasop is an associate professor and the director of DEKD (Data Engineering and Know ledge Discovery ) research unit at the school of computer engineering, Suranaree University of T echnology , Thailand. He received his bachelor degree in Mathematics from Srinakarinwirot University , Thailand, in 1986, master degree in computer science from the Prince of Songkla University , Thailand, in 1991 and doctoral degree in computer science from Nova Southeastern University , USA, in 1999. His current research includes Data mi ning, Machine Learning, Artificial Intelligence, Logic and Functional Programming, Probabilistic Databases and Knowledge Bases.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment