Learning a dynamic four-chamber shape model of the human heart for 95,695 UK Biobank participants
The human heart is a sophisticated system composed of four cardiac chambers with distinct shapes, which function in a coordinated manner. Existing shape models of the heart mainly focus on the ventricular chambers and they are derived from relatively…
Authors: Qiang Ma, Qingjie Meng, Yicheng Wu
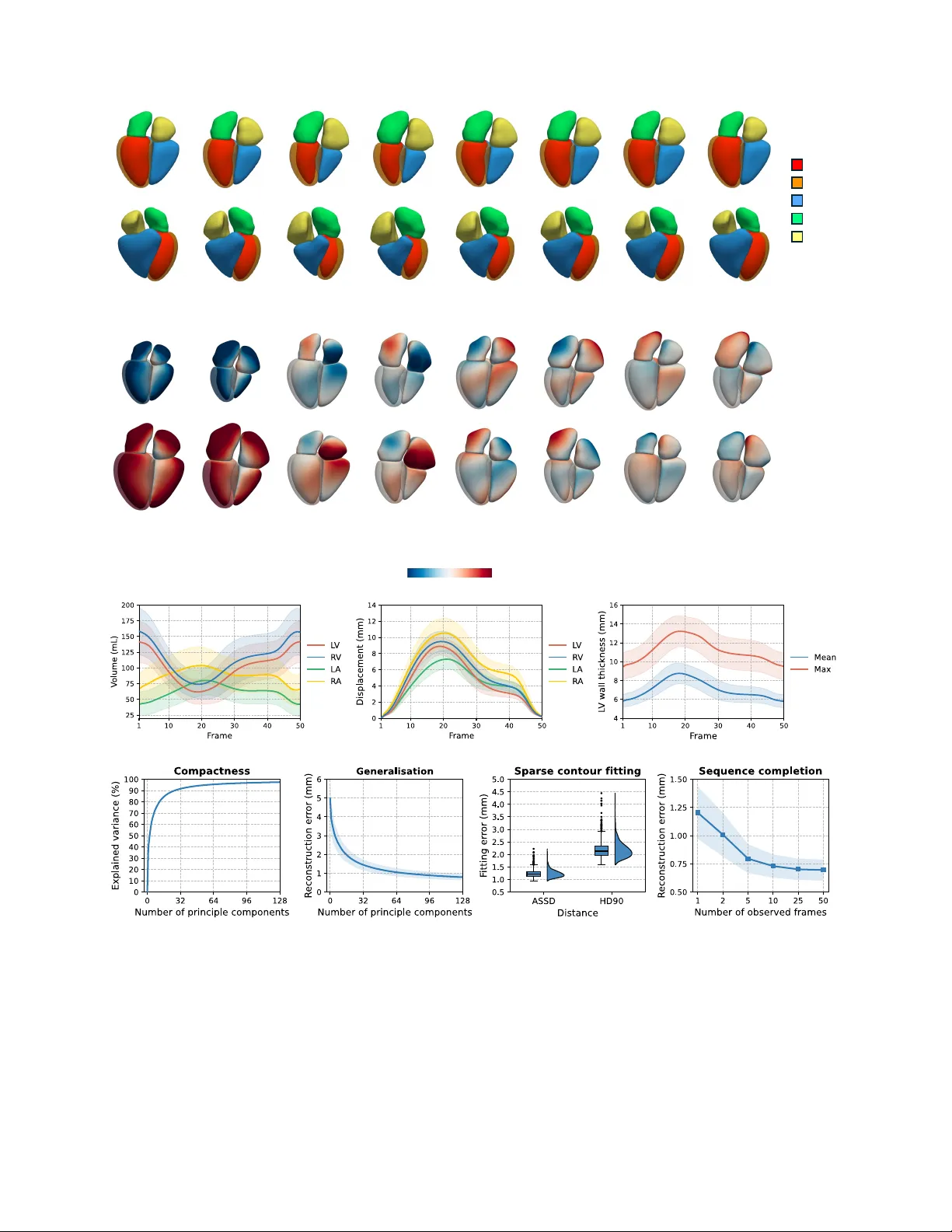
Lear ning a dynamic f our -chamber shape model of the human heart f or 95,695 UK Biobank participants Qiang Ma 1,2,* , Qingjie Meng 2,3 , Yicheng W u 1,2 , Shuo W ang 4,5 , Mengyun Qiao 2,6 , Steven Niederer 7 , Declan P . O’Regan 8 , P aul M. Matthews 1,9,10 , and W enjia Bai 1,2,5 1 Depar tment of Brain Sciences, Imperial College London, London, UK 2 Depar tment of Computing, Imperial College London, London, UK 3 School of Computer Science, Univ ersity of Birmingham, Bir mingham, UK 4 Digital Medical Research Center , School of Basic Medical Sciences, Fudan University , Shanghai, China 5 Data Science Institute, Imperial College London, London, UK 6 Depar tment of Mechanical Engineering, University College London, London, UK 7 National Hear t and Lung Institute, Imperial College London, London, UK 8 MRC Laboratory of Medical Sciences, Imperial College London, London, UK 9 UK Dementia Research Institute, Imperial College London, London, UK 10 Rosalind F ranklin Institute, Harwell Science and Inno vation Campus , Didcot, UK * q.ma20@imperial.ac.uk Abstract The human heart is a sophisticated system composed of four cardiac chambers with distinct shapes, which function in a coordinated manner . Existing shape models of the heart mainly focus on the ventricular chambers and they are deri v ed from relati vely small datasets. Here, we present a spatio-temporal (3D+t) statistical shape model of all four cardiac chambers, learnt from a large population of nearly 100,000 participants from the UK Biobank. A deep learning-based pipeline is developed to reconstruct 3D+t four-chamber meshes from the cardiac magnetic resonance images of the UK Biobank imaging population. Based on the reconstructed meshes, a 3D+t statistical shape model is learnt to characterise the shape v ariations and motion patterns of the four cardiac chambers. W e reveal the associations of the four-chamber shape model with demographics, anthropometrics, cardiov ascular risk factors, and cardiac diseases. Compared to conv entional image-derived phenotypes, we validate that the four-chamber shape-deriv ed phenotypes significantly enhance the performance in downstream tasks, including cardiov ascular disease classification and heart age prediction. Furthermore, we demonstrate the ef fectiv eness of shape-deriv ed phenotypes in novel applications such as heart shape retriev al and heart re-identification from longitudinal data. T o facilitate future research, we will release the learning-based mesh reconstruction pipeline, the four-chamber cardiac shape model, and return all deriv ed four -chamber meshes to the UK Biobank. Understanding the variations of cardiac anatomy and function, as well as their associations with disease risks and clinical outcomes remains one of the central challenges in cardiov as- cular research. The av ailability of large-scale cardiac imaging datasets in recent years, such as the UK Biobank Imaging Study 1 – 4 , provides an unprecedented opportunity to perform computational modelling of the cardiac shape and function at a population scale. Owing to excellent soft tissue contrast and non-ionising radiation, cine cardiac magnetic resonance (CMR) imaging employed by the UK Biobank study enables non-in v asi ve and dynamic imaging to capture the cardiac shape and motion 5 – 7 . Phenotypes deriv ed from CMR images, such as the chamber volume, ejection fraction, and myocar- dial wall thickness, provide global quantitative description of the cardiac structure and function, serving as important biomarkers for cardiovascular research 8 , 9 . Howe ver , these phenotypes may overlook the detailed spatio-temporal (3D+t) characterisation of the cardiac shape and motion, which has been demonstrated to capture richer information compared to the global description of image-derived phenotypes 10 – 13 . This motiv ates the cardiac imaging community to dev elop computational models to describe the 3D+t shape and motion of the heart 14 – 19 , which can facilitate a range of clinical and research tasks, including disease risk prediction 12 , clinical intervention planning 20 , 21 , biomechanical analysis 22 , haemo- dynamic simulation 23 , 24 , electrophysiological modelling 25 – 27 , cardiac digital twin modelling 28 – 31 , etc. Current computational shape modelling of the heart faces two major challenges. The first challenge is that existing car - diac shape models are mostly learnt from data at a relativ ely small scale. This may owe to both the lack of population- lev el cardiac imaging datasets and the absence of computa- tional algorithms that can efficiently extract 3D shape from cardiac imaging data. Statistical shape models (SSMs) of the heart hav e been constructed using datasets ranging from dozens to thousands of subjects 10 , 32 – 35 . W ith the av ailabil- ity of UK Biobank population imaging data, cardiac shape modelling is currently being extended to tens of thousands participants 14 , 16 , 19 . In order to extract cardiac shapes from CMR images, guide-point model and atlas-based approaches hav e been explored in the past 34 , 36 . In recent years, a grow- ing number of deep learning-based models ha ve been dev el- oped for cardiac mesh reconstruction, which substantially improv ed the computational efficienc y in processing large- scale datasets 14 , 15 , 37 – 41 . The second challenge is that existing cardiac shape models primarily focus on the v entricles of the heart 15 , 16 , 19 , instead of cov ering all the four chambers including the left v entricle (L V), right ventricle (R V), left atrium (LA), and right atrium (RA). As one of the most widely used imaging modalities for cardiov ascular research, cine CMR captures both cardiac shape and motion. Howe ver , the standard CMR imaging pro- tocols typically do not provide a full co verage of the left and right atria 5 – 7 . The two atria are captured only in a fe w long- axis view 2D image slices. The sparse coverage of the atria poses a technical challenge for estimating their 3D shapes. Al- though computed tomography (CT) imaging can cov er all four cardiac chambers, it is cautiously used for only specific patient studies due to radiation b urden 17 , 42 – 44 . T o characterise both shape and motion of four cardiac chambers, existing work trained deep neural networks to learn four-chamber shape pri- ors from CT datasets 43 and completed sparse multi-view CMR segmentations into 3D dense four -chamber volumes 45 – 48 . Xia et al. b uilt a four-chamber SSM by aligning a cardiac atlas to manually delineated contours and introduced MSCI-Net for four-chamber mesh reconstruction 14 . Howe ver , neither approaches enforced motion consistency across the cardiac cy- cle. Recent studies learnt 3D+t cardiac shape models from 4D CT 17 or MR 18 datasets. Howe ver , these studies were limited to the left atrium and trained on small-scale datasets only . Here, we present HeartSSM , a 3D+t statistical four- chamber shape model of the human heart learnt from a population-lev el CMR dataset from the UK Biobank Imag- ing Study with nearly 100,000 participants, the lar gest of its kind. T o perform efficient population-scale shape modelling and address the sparse co verage problem of the atria, we de- velop a fast deep learning-based pipeline for four-chamber cardiac shape reconstruction from multi-vie w cine CMR im- ages. The pipeline consists of sev eral stages, including CMR motion correction, 3D cardiac label completion, and 3D+t mesh reconstruction. W e employ the pipeline to reconstruct time-resolved four -chamber meshes for 95,695 UK Biobank participants after data cleaning, thereby curating a large-scale data resource for the shape modelling study . Based on the re- constructed meshes, we construct the HeartSSM shape model using incremental principle component analysis (PCA) to address the computational cost challenge 49 – 51 . W e perform comprehensiv e ev aluation of the deep learn- ing analysis pipeline with respect to motion correction, label completion, and mesh reconstruction performance. For the HeartSSM shape model, we conduct correlation analyses of learnt principle components and vertex-wise shape v ariations with participant characteristics. W e calculate shape-derived phenotypes and compare them to con ventional image-deriv ed phenotypes 9 in multiple downstream tasks, including cardio- vascular disease classification 19 and heart age prediction 52 . Finally , we demonstrate the usability of the HeartSSM shape model in two novel tasks, namely heart shape retriev al and longitudinal heart re-identification. W e envisage that the pro- posed deep learning-based mesh reconstruction pipeline, the curated 3D+t four-chamber shape dataset, and the de veloped cardiac shape model will provide a po werful foundation for future research in machine learning, 3D shape modelling, and cardiov ascular studies. Results Perf ormance of 3D+t cardiac four -chamber mesh recon- struction pipeline W e first provide an overvie w of the deep learning-based mesh reconstruction pipeline (Fig. 1 a), which takes CMR images and segmentations 8 as input and reconstructs 3D+t meshes of the four-chamber heart as output (see detailed descrip- tion in Methods). The pipeline consists of three components: 1) CMR motion correction; 2) 3D four-chamber label com- pletion; 3) 3D+t mesh reconstruction. First, an intensity- based motion correction algorithm 53 is employed and adapted to mitigate respiratory motion artefacts and multi-vie w mis- alignment in CMR images and se gmentations. Then, a label completion U-Net (LC-U-Net) is applied to sparse multi- view CMR segmentations, including short-axis and long- axis vie ws, to generate dense 3D+t segmentations of four chambers 45 – 48 (Supplementary Fig. 1). Finally , we dev elop a segmentation-to-mesh netw ork, HeartFFDNet , which tak es 3D+t four-chamber segmentations as input, estimates multi- scale and multi-frame free-form deformations (FFDs), and warps a template mesh into motion consistent 3D+t meshes of four cardiac chambers (Fig. 1 b). HeartFFDNet provides a consistent mesh representation of four-chamber heart so that the predicted meshes share the same verte x connecti vity across all time frames and across dif ferent subjects. The pro- posed deep learning pipeline is computationally efficient and only requires 2.87 seconds (s) to perform motion correction (0.32 s), label completion (2.4 s), and mesh reconstruction (0.15 s) for each CMR image sequence. W e e valuate the performance of each component of the cardiac four-chamber mesh reconstruction pipeline (Fig. 1 a). T o ev aluate the motion correction performance, we calcu- late the overlap of multi-view CMR segmentations at their intersection using the Dice coef ficient, and the intensity con- sistency of multi-vie w CMR images at their intersection using the Pearson correlation coeffi cient ( r ). Fig. 2 a reports the Dice coef ficient and the Pearson correlation coef ficient before and after motion correction. For 95,695 CMR images with segmentations, the motion correction algorithm yields signifi- cantly impro vement, with Dice increasing from 0 . 810 ± 0 . 077 to 0 . 859 ± 0 . 038 (paired t -test, p < 10 − 307 ) and Pearson’ s r 2 a b RA LA RV LV M LV LC -U- Net Label complet ion 2D+t sparse segmentat ion 3D+t dense segmentat ion HeartFFDNe t Mesh reconstruction 3D+t meshes 95,695 UK B iobank CMR seque nces SAX 4CH 2CH Motion c orrectio n Original Corrected … … … Template segmentat ion Loss … 3D+t segmentation Template mesh Initial mesh Multi -scale FFD 3D+t point clou ds Multi - frame FFD 3D+t meshes … … … … 3D+t segmentation SAX 4CH 2CH t=1 t=8 t=15 t=22 t=29 t=36 t=43 t=50 c Fig. 1 | The deep learning-based pipeline f or 3D+t cardiac four -chamber mesh reconstruction. a , The processing pipeline consists of three components: motion correction for multi-view CMR images and se gmentations, label completion from 2D+t multi-view sparse se gmentations to 3D+t dense segmentations of the four chambers via a label completion U-Net (LC-U-Net), and finally 3D+t four-chamber mesh reconstruction with HeartFFDNet. b , The architecture of HeartFFDNet. HeartFFDNet learns multi-scale and multi-frame free-form deformations (FFDs) from input 3D+t segmentations, warping a template mesh into four-chamber meshes across all time frames of a cardiac cycle. c , Qualitative visualisation of reconstructed 3D+t cardiac four-chamber meshes o verlaid on multi-vie w CMR images. SAX: short-axis vie w; 4CH: long-axis four-chamber vie w; 2CH: long-axis two-chamber vie w . 3 a b c d e Fig. 2 | Perf ormance of the 3D+t cardiac four -chamber mesh reconstruction pipeline. a , Performance of motion correction measured by segmentation o verlap (Dice) at the intersections of multi-vie w CMR segmentations and intensity consistenc y (Pearson’ s r ) at the intersections of multi-vie w CMR images. b , Performance (Dice and ASSD) of 3D cardiac four-chamber label completion. c , Geometric errors (ASSD and HD90) of 3D+t four-chamber meshes of the proposed HeartFFDNet compared to DeepMesh and HybridVNet. d , T emporal consistency error and c ycle consistency error of 3D+t four -chamber meshes reconstructed by HeartFFDNet compared to baseline methods. e , Geometric errors (ASSD and HD90) of the 3D+t meshes reconstructed by HeartFFDNet av eraged across four cardiac chambers, e valuated on randomly selected test samples and participants with cardiov ascular diseases. increasing from 0 . 796 ± 0 . 091 to 0 . 861 ± 0 . 054 (paired t -test, p < 10 − 307 ) after motion correction. For 3D cardiac label completion, we trained a LC-U- Net 45 – 48 to complete the 3D cardiac four-chamber se gmenta- tion from 2D sparse multi-view segmentations. Since there are no ground-truth 3D segmentations for UK Biobank CMR images, we curate a dataset containing 3D four-chamber se g- mentations of CT images 54 – 60 (see dataset information in Methods) for training and ev aluation of the LC-U-Net. Fig. 2 b presents the Dice score and the absolute symmetric surf ace dis- tance (ASSD) measured between the completed and ground- truth 3D four -chamber segmentations. The results show that the Dice scores of all four chambers are abov e 0.91, with the lo west performance observed for the RA (Dice, 0 . 918 ± 0 . 028 ; ASSD, 1 . 376 ± 0 . 483 mm ). The trained LC-U-Net is applied to the sparse 2D+t CMR segmentations from UK Biobank dataset to generate 3D+t dense four -chamber segmentations as illustrated in Fig. 1 a. For 3D+t mesh reconstruction, we ev aluate the geomet- ric accuracy of the 3D+t four-chamber meshes predicted by HeartFFDNet. W e extract 2D+t four-chamber contours from ground-truth UK Biobank CMR segmentations 8 with manual quality control and measure the distance between 2D+t contours and 3D+t meshes generated by HeartFFD- Net, in terms of the uni-directional ASSD and the 90th per- centile of Hausdorff distance (HD90). Fig. 2 c and Supple- mentary T able 3 compare the performance of HeartFFDNet to two recent baseline methods, DeepMesh 15 and Hybrid- 4 a c d b t=1 t=8 t=15 t=22 t=29 t=36 t=43 t=50 +2 SD - 2 SD - 6 mm 6 mm PC1 (ED) PC1 (ES) PC4 (ES) PC4 (ED) PC3 (ES) PC3 (ED) PC2 (ED) PC2 (ES) LV (epi ) RV LA RA LV (endo ) Fig. 3 | The 3D+t statistical shape model of the f our -chamber heart. a , The mean shapes (anterior and posterior view) of 3D+t four-chamber hearts deri ved from HeartSSM. b , The shape v ariations captured by the first four principle components (PC) of HeartSSM, with ± 2 standard de viation (SD) from the mean shape. The colour maps indicate the signed distances to the mean shape along normal directions. Red colour indicates inflation compared to the mean shape, while blue colour indicates contraction. Only end-diastolic (ED) and end-systolic (ES) frames are visualised. c , The mean (line) and standard deviation (shaded area) of cardiac four-chamber v olumes, displacements, and left ventricle w all thickness (mean and max) calculated from the HeartSSM fitted four-chamber meshes, displayed across all time frames and a veraged o ver 95,695 UK Biobank participants. d , Performance of the HeartSSM. From left to right: the compactness of HeartSSM; the generalisation ability of HeartSSM on unseen cardiac shapes; performance of HeartSSM on 2D+t sparse contour fitting; performance of HeartSSM on sequence completion with partially observed frames. The shaded areas indicate ± 1 standard deviation. 5 VNet 40 , on the test set. The results show that HeartFFD- Net achiev es significantly better geometric accuracy (ASSD, 1 . 426 ± 0 . 202 mm ; HD90, 2 . 605 ± 0 . 476 mm ) than DeepMesh (paired t -test; ASSD, 1 . 478 ± 0 . 249 mm , p = 3 . 5 × 10 − 74 ; HD90, 2 . 697 ± 0 . 561 mm , p = 2 . 2 × 10 − 49 ) and HybridVNet (paired t -test; ASSD, 1 . 579 ± 0 . 262 mm , p = 1 . 1 × 10 − 242 ; HD90, 2 . 913 ± 0 . 568 mm , p = 1 . 5 × 10 − 211 ). An example of reconstructed 3D+t cardiac four-chamber meshes o verlaid on multi-view CMR images is visualised in Fig. 1 c, showing that the reconstructed meshes align well with the anatomical boundaries of the four chambers on different CMR vie ws. In addition to geometric accuracy , we ev aluate the tem- poral and cycle consistenc y of predicted 3D+t four -chamber meshes. The temporal consistency is measured by the tempo- ral Laplacian error across all time frames of a cardiac cycle ( T = 50 ) to assess the coherence between consecutive frames. The cycle consistency is measured by the distance between the meshes at the starting frame and the end frame to assess whether the cardiac mesh returns to its original shape after a full cardiac cycle. Fig. 2 d demonstrates the temporal con- sistency error ( 0 . 150 ± 0 . 016 mm ) and cycle consistency error ( 0 . 284 ± 0 . 052 mm ) of HeartFFDNet, which outperforms both baseline methods (paired t -test, p < 10 − 183 ). Furthermore, we examine the generalisability of HeartFFD- Net for different subgroups within the UK Biobank popula- tions with different cardio vascular diseases (determined using ICD-10 code), including the dilated cardiomyopathy (I42.0), hypertrophic cardiomyopathy (I42.1, I42.2), pulmonary hy- pertension (I27.0, I27.2), atrial fibrillation and flutter (I48), and heart failure (I50). Please see Supplementary T able 2 for detailed data information. The mesh reconstruction errors (ASSD and HD90) are reported in Fig. 2 e and Supplementary T able 4 for different disease groups. The mean geometric errors of the diseased cohorts are within + 1 . 3 standard de vi- ations (ASSD, 1 . 688 mm ; HD90, 3 . 224 mm ) of the geometric errors on a test set of randomly selected samples, demonstrat- ing the generalisability of HeartFFDNet on participants with cardiov ascular diseases. 3D+t statistical shape model of four cardiac chambers W e construct a 3D+t SSM of the four cardiac chambers, named as HeartSSM . Due to the high computational cost, HeartSSM is learnt by incremental PCA 49 – 51 , using 64,000 3D+t four-chamber meshes generated by our learning-based pipeline as training data. HeartSSM captures the variations of both cardiac shape and motion in a large population, modelled by a mean 3D+t dynamic mesh sequence plus a linear combi- nation of principle components (PCs). Fig. 3 a visualises the mean 3D+t heart shapes at different time frames, demonstrat- ing the contraction and relaxation process of the four cham- bers across a cardiac cycle. Fig. 3 b visualises the first four PCs of HeartSSM with ± 2 standard de viation (SD) at the end- diastolic (ED) and end-systolic (ES) frames (see Supplemen- tary Fig. 5-7 for additional PCs and time frames). The colour map indicates the vertex-wise signed distance of a shape com- pared to the mean shape, measured along the normal direction of each vertex. Red colour indicates inflation compared to the mean shape and blue colour indicates contraction. Fig. 3 b shows that PC1 controls the size of the whole heart, PC2 mainly affects the shape of the RA, and other PCs characterise different modes of shape v ariations. W e employ HeartSSM to reconstruct 3D+t four -chamber meshes of 95,695 UK Biobank subjects and encode each of the 3D+t mesh sequence into a 128-dimensional shape descriptor . Based on the reconstructed meshes, we compute the four-chamber volumes, displace- ments, and L V wall thickness (mean and max) av eraged ov er all subjects across a cardiac cycle (Fig. 3 c). W e assess the representation capacity of HeartSSM using four metrics 61 , 62 in terms of compactness, generalisation, ca- pacity for fitting sparse 2D+t contours and for completing 3D+t sequences with missing frames (Fig. 3 d). Regarding compactness, HeartSSM explains over 90% of the variance with the first 27 PCs and 97.45% of the variance with all 128 PCs. The generalisation ability is ev aluated by fitting the HeartSSM to unseen test 3D+t cardiac meshes from the UK Biobank. W e measure the reconstruction error by com- puting the mean squared error between the test 3D+t shapes and the shapes reconstructed by the HeartSSM. Fig. 3 d shows that the reconstruction error ( 0 . 788 ± 0 . 170 mm ) is relati vely low when using 128 PCs. W e examine the fitting ability of HeartSSM using test subjects for sparse contour fitting, i.e. , fitting 3D+t meshes to sparse 2D+t four-chamber contours, and for sequence completion, i.e. , fitting 3D+t meshes to a mesh sequence with missing time frames. Qualitativ e il- lustrations of contour fitting and sequence completion are provided in Supplementary Fig. 3 and 4. By optimising the weights of the PCs, HeartSSM accurately fits onto 2D+t sparse contours with relativ ely low distance errors (Fig. 3 d; ASSD, 1 . 238 ± 0 . 154 mm ; HD90, 2 . 198 ± 0 . 346 mm ). For missing sequence completion, HeartSSM reconstructs 3D+t cardiac meshes effecti vely from e ven a single observed frame (Fig. 3 d; ASSD, 1 . 204 ± 0 . 232 mm ). More accurate fitting can be achiev ed with only 10 observed frames (Fig. 3 d; ASSD, 0 . 730 ± 0 . 104 mm ). Associations of f our -chamber shapes with demographics, anthropometrics and diseases Using 3D+t four-chamber meshes reconstructed by HeartSSM, we calculate phenotypes characterising the heart structure and function, including the ventricular ED volumes (EDV) and ES volumes (ESV), atrial maximum volumes (MAXV) and minimum volumes (MINV), stroke volumes (SV), ejection fractions (EF), L V myocardial mass (L VM), as well as L V wall thickness (L VWT). W e refer to these phenotypes as shape- derived phenotypes , in contrast to the image-deri ved pheno- types estimated directly from CMR image se gmentations 8 , 9 . T o interpret the cardiac shape variations captured by the PCs, we explore the correlations between the first 12 PC weights (capturing > 80% variance) of the HeartSSM with shape- deriv ed phenotypes, sex, age, weight, height, body mass index 6 a b - 3 mm 3 mm Sex 50 - 59 years -2 mm 2 mm Male Female All 60 - 69 years 70 - 79 years Sex Age c Heart fa ilure Hypertension Atrial fibrillation and flutter BMI Age - 0.5 0.5 - 0.7 0.7 - 0. 3 0. 3 - 0.3 0.3 - 0. 2 0. 2 - 1.0 1.0 Fig. 4 | Corr elation analysis of cardiac f our -chamber shape model. a , Correlations of PCs with four-chamber phenotypes, demographics, anthropometrics, and cardiov ascular risk factors. Only significant correlations are visualised ( p < 0 . 05). b , Mean cardiac four-chamber shapes and shape v ariations a veraged across all time frames for dif ferent sex and age groups. The colour maps depict the signed distances along surface normals to the mean cardiac shape of all participants. Red colour indicates inflation compared to the mean shape, while blue colour indicates contraction. c , Correlations of verte x-wise shape v ariations with se x, age, BMI, hypertension, atrial fibrillation and flutter , and heart f ailure. The colour maps visualise significant Pearson correlation coefficients after Bonferroni correction ( p < 0 . 05 / n , n = 27 , 034 denoting the number of all vertices). (BMI), diastolic blood pressure (DBP), systolic blood pres- sure (SBP), vigorous physical activities (P A), smoke, alcohol, high cholesterol, and diabetes 9 , 16 . The Pearson correlation coefficients are presented in Fig 4 a. As expected, the first PC, which is negati vely correlated to the size of the hearts (Fig. 3 b), sho ws negati ve correlation with L V mass, L V wall thickness, four-chamber volumes, sex, weight, and height. Positiv e correlations are observ ed between the first PC with four-chamber ejection fractions. W e compare the shape variations between different sex and age groups in a cohort of 62,778 UK Biobank partici- pants with av ailable demographic and disease information. 7 a b Fig. 5 | P erf ormance for cardio vascular disease classification and chr onological age prediction. a, Binary classification performance for six cardiac diseases using dif ferent features, including image-deri ved phenotypes, shape-deri ved phenotypes, and HeartSSM shape descriptor , with or without confounders (sex, age, weight, height, and BMI). The R OC A UC scores for disease classification using logistic regression and XGBoost models are reported. The symbol * indicates a significant difference (tw o-sided DeLong test, p < 0 . 05 ). b, Chronological age prediction performance using different features, including image-deri ved phenotypes, shape-deri ved phenotypes, 3D four -chamber (4CH) HeartSSM shape descriptor , 3D+t bi-ventricular (2CH+t) HeartSSM shape descriptor , and 3D+t four -chamber (4CH+t) HeartSSM shape descriptor . The mean absolute error, Pearson’ s r , and coefficient of determination R 2 for age prediction using CatBoost model are reported. The symbol * indicates statistically significant difference (paired t -test, p < 0 . 05). W e compute the mean cardiac four-chamber shape for each group, av eraged across all subjects of each group and across all T = 50 time frames, as shown in Fig. 4 b . The colour maps represent the shape v ariations compared to the mean cardiac shape of all subjects. Furthermore, we perform correlation analysis between the shape variation of each verte x and three clinical factors including se x, age, and BMI, as well as three disease types including hypertension (I10-I15), atrial fibrilla- tion and flutter (I48), and heart failure (I50). At each vertex of the cardiac four-chamber meshes, we calculate the Pearson correlation coefficient ( r ) between its mean signed distance ov er all frames and each of the abov e six factors. The correla- tion maps are visualised in Fig. 4 c. Fig. 4 b and c demonstrate that male hearts are generally larger than female hearts, and the ventricle volumes sho w a decreasing trend with increas- ing age. The correlation maps of shape variation with BMI and hypertension in Fig. 4 c exhibit similar patterns across all four chambers. For atrial fibrillation and flutter , we observe a stronger correlation (two-sided t -test, p < 10 − 307 ) on the two atria ( r = 0 . 108 † , † av eraged across all vertices) compared to the two ventricles ( r = 0 . 052 † ). The positive correlation shows that the enlar gement of the atria is positi vely associated with atrial fibrillation and flutter 63 , 64 . There is a stronger pos- itiv e correlation (two-sided t -test, p < 10 − 307 ) between the left heart shape variation with heart f ailure ( r = 0 . 063 † ) than the right heart shape variation ( r = 0 . 019 † ) 65 – 67 . 8 Shape features f or cardiovascular disease classification HeartSSM characterises each 3D+t four -chamber meshes us- ing a 128-dimensional shape descriptor , i.e. , the weights of the PCs. W e in vestigate whether the shape derived-phenotypes and HeartSSM shape descriptor provide discriminative fea- tures for cardiov ascular disease classification 19 . W e train two types of classification models, logistic regression and XGBoost 68 for the binary classification of six cardiov ascular diseases including hypertension (I10-I15), angina pectoris (I20), acute myocardial infraction (I21), chronic ischaemic heart disease (I25), atrial fibrillation and flutter (I48), and heart failure (I50). The classifiers are trained with differ - ent features, including confounders (age, sex, weight, height, BMI), image-deriv ed phenotypes, shape-deri ved phenotypes, and HeartSSM shape descriptor . Fig. 5 a and Supplementary T able 5 report the disease classification performance, evalu- ated using the area under the receiver operating characteristic curve (R OC A UC) in four-fold cross validation. The clas- sification performance with different features are compared using the two-sided DeLong test 19 , 69 . Across all six cardiac disease classification tasks, shape-deri ved phenotypes achie ve consistently higher A UCs than image-deriv ed phenotypes, regardless of the inclusion of confounders and the choice of classifiers. HeartSSM shape descriptor further improves classification performance compared to shaped-deri ved phe- notypes. Combining both confounders and HeartSSM shape descriptors as features yields the best performance for classi- fying all six diseases. Shape features f or chronological age prediction W e assess the effecti veness of shape-deri ved phenotypes and HeartSSM shape descriptor on chronological age prediction task 52 , using a set of 9,135 healthy UK Biobank partici- pants without reported diseases. A CatBoost regressor 70 is trained for age prediction using dif ferent features, including image-deriv ed phenotypes, shape-deri ved phenotypes, and HeartSSM shape descriptor . The prediction performance is e valuated by the mean absolute error (MAE), Pearson correla- tion coefficient ( r ), and coefficient of determination ( R 2 ). Fig 5 b compares the performance when different features are used. It sho ws that shape-deriv ed phenotypes (MAE, 5 . 538 ± 3 . 946 ; r = 0 . 378 ; R 2 = 0 . 141 ) outperform image-deri ved phenotypes (MAE, 5 . 628 ± 3 . 958 ; r = 0 . 350 ; R 2 = 0 . 120 ) with statistical significance (paired t -test, p = 1 . 0 × 10 − 5 ). HeartSSM shape descriptors leads to substantial improv ement of performance. T o demonstrate the advantage of the 3D+t four-chamber (4CH+t) HeartSSM model, we compare its performance with two variants, namely a 3D four-chamber (4CH) HeartSSM without motion and a 3D+t bi-ventricular (2CH+t) HeartSSM model. The 3D four-chamber HeartSSM is constructed us- ing ED and ES frames only without intermediate frames, whereas 3D+t bi-ventricular HeartSSM is constructed with- out the two atria. W e compare the performance of the shape descriptors of three HeartSSM models for the age prediction task. Fig. 5 b shows that the 3D+t four-chamber (4CH+t) HeartSSM (MAE, 4 . 249 ± 3 . 211 ; r = 0 . 689 ; R 2 = 0 . 473 ) significantly outperforms the 3D+t bi-ventricular (2CH+t) HeartSSM (MAE, 4 . 326 ± 3 . 205 ; r = 0 . 681 ; R 2 = 0 . 461 ; paired t -test, p = 2 . 4 × 10 − 5 ) and the 3D four-chamber (4CH) HeartSSM without motion information (MAE, 4 . 468 ± 3 . 308 ; r = 0 . 653 ; R 2 = 0 . 426 ; paired t -test, p = 2 . 7 × 10 − 28 ) for chronological age prediction. This demonstrates the effecti ve- ness of utilising both shape and motion information of all four cardiac chambers. Population-le vel cardiac shape r etriev al W e examine the representation ability of the HeartSSM shape descriptor for cardiac shape retriev al 71 , 72 . Giv en a query HeartSSM shape descriptor , we retriev e top K subjects with the most similar descriptors in the UK Biobank imaging pop- ulation and assess the retriev al precision, defined as the per- centage of the retriev ed subjects sharing the same clinical factor as the query subject. W e calculate the precision for six clinical factors, including se x, age, weight, height, BMI, and hypertension. All factors are represented as categorical vari ables as illustrated in Fig. 6 a. For comparison, we also use shape-deriv ed phenotypes and image-deri ved phenotypes as query vectors for the shape retriev al task, and include random retriev al as a baseline. Fig. 6 b and Supplementary T able 6 report the top- K precision for dif ferent clinical factors and dif- ferent K parameters. Each precision value is calculated using 5,000 queries. HeartSSM shape descriptor achiev es the high- est retriev al precision compared to shape-derived phenotypes and image-deriv ed phenotypes for all six clinical factors. Longitudinal re-identification of cardiac shapes Besides cardiac shape retriev al, we conduct experiments on longitudinal re-identification of cardiac four-chamber shapes. In our dataset, 4,134 participants have two CMR imaging scans with a time gap of 2 . 66 ± 1 . 09 years 4 . Shape-based lon- gitudinal heart re-identification is a particularly challenging task, as the planning of the CMR scan such as the long-axis view position may v ary substantially between the two scans, as illustrated in Fig. 6 c. Using the HeartSSM shape descrip- tor of a participant at the first time point, we retriev e top K subjects with the most similar descriptors at the second time point and assess the re-identification accuracy . W e re- port the top- K recall in Fig. 6 d and Supplementary T able 7, i.e. , the percentage of subjects that are re-identified within K retriev als. For the HeartSSM shape descriptor, we use 12/17/27/58 numbers of PCs that explain 80/85/90/95% of variance to represent the primary shape and motion patterns for longitudinal re-identification. Interestingly , we found that the reduced number of PCs results in better retriev al perfor- mance (Fig. 6 d, right). Using the HeartSSM shape descriptor (12 PCs, 80% v ariance) as the query vector achie ves 49 . 01% recall for K = 200 , which significantly outperforms the re- identification result of image-derived phenotypes ( 31 . 37% recall@200; paired t -test, p = 7 . 1 × 10 − 63 ). 9 a b c d First imaging Second imaging Fig. 6 | P erf ormance on cardiac shape retrie val and longitudinal r e-identification. a , Sex, age, weight, height, BMI and hypertension are categorised into dif ferent groups. b , T op- K precision for shape retriev al using random retriev al, image-derived phenotypes, shape-deriv ed phenotypes, and HeartSSM shape descriptor as query vectors. c , Long-axis four-chamber vie w CMR images of an example subject, compared between the first and second imaging scans. d , T op- K recall for longitudinal re-identification using random retriev al, image-derived phenotypes, shape-deri ved phenotypes, and HeartSSM shape descriptor as query vectors. The performance is also compared across HeartSSM shape descriptors with different number of PCs that explain dif ferent ratios (%) of v ariance. Discussion In this study , we present learning-based methods for mesh re- construction and shape modelling of the cardiac four chambers using a population-lev el imaging dataset, the UK Biobank. Our work mak es fiv e-fold contrib utions. First, we proposed an efficient deep learning-based pipeline for 3D+t cardiac four-chamber mesh reconstruction and motion tracking. Sec- ond, by employing the pipeline, we created a large-scale data resource of time-resolved four-chamber cardiac meshes for the UK Biobank imaging population, which comprises of nearly 100,000 participants in their mid-to-late ages. Third, we constructed HeartSSM, a population-scale shape model capturing both shape v ariations and motion patterns of cardiac four chambers. Fourth, we re vealed the correlation of cardiac four-chamber shapes with demographics, anthropometrics, cardiov ascular risk factors, and cardiac diseases. Finally , com- pared to con ventional image-deriv ed phenotypes, we demon- strated that shape-deriv ed phenotypes and HeartSSM shape descriptor achiev e significant improvement for cardiac dis- ease diagnosis, age prediction, cardiac shape retriev al, and longitudinal re-identification. The deep learning-based mesh reconstruction pipeline in- tegrates CMR respiratory motion correction, 3D cardiac label completion via LC-U-Net 45 – 48 , and 3D+t mesh re- construction by HeartFFDNet. The HeartFFDNet learns FFDs for joint four-chamber mesh reconstruction and mo- tion tracking, achie ving superior geometric accuracy (ASSD, 1 . 426 ± 0 . 202 mm ; HD90, 2 . 605 ± 0 . 476 mm ) as well as tem- poral ( 0 . 150 ± 0 . 016 mm ) and cycle ( 0 . 284 ± 0 . 052 mm ) con- sistency compared with state-of-the-art baseline methods 15 , 40 . The generalisation ability of HeartFFDNet has been verified on UK Biobank participants with a range of cardiovascular diseases. The pipeline is memory and time efficient, support- ing training and inference on a single 12GB NVIDIA 3080 GPU card with 2.87 seconds of inference time. The mesh representation of the four-chamber heart provides accurate estimation of shape-deriv ed phenotypes, in particular for the two atria. Moreover , the reconstructed meshes share the same verte x connecti vity across all time frames and across all UK 10 Biobank participants, enabling population-scale analysis of cardiac structure and function. The statistical shape model, HeartSSM, was constructed based on the reconstructed 3D+t cardiac four-chamber meshes of 95,696 UK Biobank participants. 128 PCs of HeartSSM explained 97 . 45% variance of the cardiac shape and motion distribution, summarising complex geometry and dynamics of the heart. HeartSSM exhibited outstanding generalisability on unseen shapes ( 0 . 788 ± 0 . 170 mm ) as well as remarkable performance on sparse contour fitting and missing sequence completion. HeartSSM encoded a cardiac four-chamber mesh sequence into a 128-dimensional shape descriptor , serving as an informativ e feature vector for do wnstream applications. The association studies of PCs with four-chamber shape- deriv ed phenotypes and participant characteristics linked HeartSSM to meaningful cardiac structural and functional traits, enhancing the interpretability of the shape model. W e measured verte x-wise correlation of shape variations with sex, age, BMI, hypertension, atrial fibrillation and flutter , and heart failure. Our findings re vealed physiologically plausible correlations aligned with prior kno wledge. More specifically , the heart size was highly associated with sex and age as ex- pected 9 , 16 . The correlation maps of BMI and hypertension displayed similar spatial distrib utions. Atrial fibrillation and flutter showed stronger associations with atrial shape varia- tions than the two ventricles. Heart failure was more strongly correlated with the shape variations of the left heart, which is typically affected by the disease 65 – 67 . The representation ability and reliability of shape-deri ved phenotypes and HeartSSM shape descriptor were ev aluated on cardiac disease classification 19 , chronological age pre- diction 52 , cardiac shape retriev al 71 , 72 , and longitudinal re- identification tasks. Experiments v alidated that shape-deriv ed phenotypes consistently outperformed image-deri ved pheno- types in these tasks. The HeartSSM shape descriptor , which characterises both shape and motion distributions of the hearts, provided ef fectiv e features and resulted in notable improve- ment for all downstream applications. Howe ver , this study has several limitations. First, the deep learning-based pipeline reconstructs 3D+t cardiac four - chamber meshes from se gmentation maps and does not cap- ture sub-voxel intensity patterns within CMR images. As a consequence, the quality of the reconstructed four-chamber meshes is affected by the accuracy of the input automated CMR se gmentations 8 . Second, 3D cardiac label completion is learnt from four-chamber se gmentation maps on multiple CT datasets 54 – 60 , which may be dif ferent from the UK Biobank population. Third, due to the sparsity of the CMR images, there is no av ailable 3D+t ground truth from UK Biobank to directly ev aluate the reconstructed four-chamber meshes. Therefore, we introduced surrogate metrics to ev aluate the geometric accuracy of the reconstructed 3D+t meshes against sparse 2D+t contours from quality-controlled CMR segmenta- tions 8 . Lastly , in the association study of verte x-wise shape variations, we only performed correlation analysis without confounders (sex, age, weight, height, BMI, etc.), as incorpo- rating these v ariables via linear regression for each vertex is computationally intractable. In summary , we have reconstructed the 3D+t four-chamber meshes of the heart for 95,695 UK Biobank participants by dev eloping a deep learning-based pipeline. W e have built a 3D+t four-chamber HeartSSM model to characterise the distribution of cardiac shape and motion. The phenotypes deriv ed from four -chamber meshes and the HeartSSM shape descriptor demonstrate superior performance across multiple downstream tasks. The developed methods, along with the curated population-level cardiac shape and motion data re- source, will benefit future research in machine learning and cardiov ascular research, facilitating deeper understanding of cardiac structure and motion and offering new insights into cardiov ascular diseases. Methods UK Biobank CMR data In this study , we mainly use the UK Biobank CMR dataset 1 – 4 to learn the 3D+t four-chamber cardiac shape model. 102,521 participants (UK Biobank field ID 53) attended the imaging visit to one of the four imaging centres (Cheadle, Ne wcastle, Reading, and Bristol). After MRI safety screening and exclud- ing participants with claustrophobia, multi-view CMR images (short-axis view , long-axis two-chamber view , long-axis four- chamber view) are a v ailable for 95,695 participants after data cleaning. Non-imaging participant characteristics, including demographics, anthropometrics, and health information, are av ailable for 62,778 participants for this study . Among the cohort, 4,134 participants hav e a second CMR imaging scan, approximately 3 years after the first scan. The detailed data information is summarised in Supplementary T able 1. Respiratory motion correction T o alleviate the respiratory motion and misalignment between multi-view CMR images, we adapt an intensity-based al- gorithm to correct in-plane motion artefacts 53 . F or short- axis view , long-axis two-chamber view , and long-axis four- chamber vie w images, we estimate the in-plane displacements ( ∆ x , ∆ y ) that minimise the mean absolute intensity difference at the intersections of multi-view images. Meanwhile, we impose a Laplacian smoothness re gularisation on the image intensities between consecutiv e short-axis vie w slices. Let I 2ch : R 2 × [ 1 , T ] → R , I 4ch : R 2 × [ 1 , T ] → R denote long-axis two-chamber and four-chamber vie w image se- quences with T time frames, and let I d sax : R 2 × [ 1 , T ] → R denote the d -th slice of the short-axis view sequences with D slices and d = 1 , ..., D . The aim is to find in-plane dis- placements ( ∆ x 2ch , ∆ y 2ch ) , ( ∆ x 4ch , ∆ y 4ch ) , ( ∆ x d sax , ∆ y d sax ) to op- timise a motion correction objectiv e. Given the displacements, 11 the motion corrected images are formulated as: ˜ I 2ch ( x , y , t ) : = I 2ch ( x + ∆ x 2ch , y + ∆ y 2ch , t ) , ˜ I 4ch ( x , y , t ) : = I 4ch ( x + ∆ x 4ch , y + ∆ y 4ch , t ) , ˜ I d sax ( x , y , t ) : = I d sax ( x + ∆ x d 4ch , y + ∆ y d 4ch , t ) , (1) and the objectiv e function is formulated by: L = 1 T T ∑ t = 1 ∑ ( x , y ) ∈ 2ch ∩ 4ch ˜ I 2ch ( x , y , t ) − ˜ I 4ch ( x , y , t ) + D ∑ d = 1 ∑ ( x , y ) ∈ 2ch ∩ ( sax ) d ˜ I d sax ( x , y , t ) − ˜ I 2ch ( x , y , t ) + D ∑ d = 1 ∑ ( x , y ) ∈ 4ch ∩ ( sax ) d ˜ I d sax ( x , y , t ) − ˜ I 4ch ( x , y , t ) + 1 2 D − 1 ∑ d = 2 ∑ ( x , y ) ˜ I d − 1 sax ( x , y , t ) + ˜ I d + 1 sax ( x , y , t ) − 2 ˜ I d sax ( x , y , t ) . (2) The objectiv e function consists of the mean absolute intensity difference at the intersections of multi-view CMR images and the Laplacian regularisation terms for each slice of the short-axis view images. W e estimate the displacements by minimising objective function ( 2 ) using Adam optimiser 73 with a learning rate 0.1 for 1,000 epochs, and subsequently perform motion correction. The performance is ev aluated us- ing the Pearson correlation coefficient of image intensities and the Dice coefficient of se gmentation maps at the intersections of multi-view CMR images. LC-U-Net for 3D cardiac label completion W e train a LC-U-Net to learn shape priors of cardiac four chambers from dense 3D CT segmentations for label comple- tion. W e collect n = 1 , 575 four-chamber heart se gmentation maps of CT images from multiple data sources: ImageCAS 54 ( n = 880 ), T otalSegmentator 55 ( n = 558 ), WHS++ 56 , 57 ( n = 32 ), the Left Atrium Segmentation Challenge 58 ( n = 25 ), the Coronary Artery Stenosis Detection Challenge 59 ( n = 48 ), and the Coronary Centerline Extraction Challenge 60 ( n = 32 ). For the ImageCAS dataset, the four-chamber heart segmen- tation maps are generated automatically by the heartcham- bers_highr es model of T otalSegmentator v2 55 follo wing man- ual quality control. The four -chamber se gmentations of all CT images follo w a consistent segmentation protocol, including L V , L VM, R V , LA, and RA label classes. All 3D CT seg- mentations are rescaled and rigidly transformed to an atlas space 74 at 2 mm resolution, cropped to a fixed size ( L , W , H ) with L = W = 96 and H = 128 . The 1,575 CT segmenta- tions are divided into subsets of 945/157/473 data for train- ing/validation/test. Note that the original CT images are not needed for cardiac label completion. As depicted in Supplementary Fig. 1, we simulate sparse multi-view segmentations based on the dense 3D four - chamber segmentations following the settings in previous work 45 – 48 . T o address motion artefacts that are not fully re- solved in the motion correction step, we introduce random in- plane displacements to the simulated short-axis view segmen- tations, follo wing a Gaussian distrib ution ∆ x , ∆ y ∼ N ( 0 , σ ) with a standard deviation σ = 2 mm . Both dense 3D and sparse multi-view se gmentations are scaled by a random fac- tor drawn from a uniform distribution U ([ 0 . 8 , 1 . 2 ]) . Using the simulated sparse multi-vie w segmentations as input, the LC-U-Net is trained to predict the dense 3D four-chamber segmentations for label completion. The mean squared error loss between the completed and original four-chamber seg- mentation maps is minimised using the Adam optimiser with a learning rate of 10 − 4 and a batch size of 1 for 200 epochs. The dropout rate for each layer of the LC-U-Net is set to 0 . 2 during training. Once trained, LC-U-Net is applied to com- plete the sparse 2D+t multi-view CMR segmentations into 3D+t dense cardiac four -chambers segmentations for 95,695 UK Biobank participants. Four -chamber heart template Based on the four-chamber heart shapes from 3D CT segmen- tations 54 – 60 , we construct a four -chamber heart template that consists of a template segmentation and a template mesh. First, the four-chamber se gmentation template is created by av erag- ing 945 CT segmentation maps in the training set, which hav e been rescaled and rigidly aligned to an atlas space 74 . Then, we compute the con ve x hull meshes for fi ve cardiac structures: L V endocardium (L V -endo), L V endocardium (L V -epi), R V , LA, and RA. These con ve x hull meshes are smoothed and remeshed as initial surfaces for a deformable model 75 , which deforms the conv ex hull meshes (L V -endo, L V -epi, R V , LA, RA) along surface normal directions to match the template segmentation, guided by the signed distance function deriv ed from the template segmentation. The L V -epi surface is pro- duced by inflating the L V -endo surface, so that the L V -epi and L V -endo meshes have corresponding vertices with the same connectivity , facilitating the measurement of L V myocardial wall thickness. The template mesh has 27,034 v ertices in total, with 6,141/6,141/5,696/4,305/4,751 vertices for L V -endo/L V - epi/R V/LA/RA respectively . Supplementary Fig. 2 visualises each structure of the cardiac four-chamber template mesh. HeartFFDNet for dynamic mesh r econstruction As illustrated in Fig. 1 b, HeartFFDNet learns FFDs 76 from input 3D+t dense segmentations of the four chambers and deforms a template mesh into 3D+t four-chamber meshes for an individual heart. Gi ven 3D+t four-chamber segmen- tations of size ( T , L , W , H ) and a template segmentation as input, HeartFFDNet employs a U-Net architecture 77 to pre- dict B-spline FFDs at two scales, which have control grids of sizes ( L / 16 , W / 16 , H / 16 , 3 ) and ( L / 8 , W / 8 , H / 8 , 3 ) . The four-chamber template mesh is warped by predicted multi- scale FFDs into an initial mesh for the follo wing mesh de- formation and motion tracking. Then, HeartFFDNet pre- dicts FFDs of size ( L / 4 , W / 4 , H / 4 , 3 ) for each of the T time 12 frames, which deform the initial mesh into a 3D+t dynamic sequence of four-chamber meshes. T o train HeartFFDNet, we formulate a loss function to min- imise the distance between the deformed 3D+t four-chamber meshes predicted by HeartFFDNet and the pseudo ground- truth 3D+t point clouds, which are derived from 3D+t seg- mentations using the marching cubes algorithm 78 . W e denote the deformed 3D+t four-chamber meshes as M t = { M c t } c ∈ C , C = { L V endo , L V epi , R V , LA , RA } , (3) where M c t denotes the mesh for structure c at time frame t = 1 , ..., T . Each mesh is represented as M c t = ( V c t , E c , F c ) , where V , E , F define the sets of vertices, edges and faces. Since the mesh of each structure has the same connectivity across all time frames, the edge set E c , the face set F c , and the number of vertices | V c | do not depend on the frame t . The loss function consists of a reconstruction loss L recon , an edge loss L edge , a curvature loss L curv , a temporal consis- tency loss L temp , and a cycle consistency loss L cycle . Giv en the pseudo ground-truth 3D+t point clouds P c t ⊂ R 3 for each structure c ∈ C , the reconstruction loss L recon is defined by the ASSD between predicted 3D+t four-chamber meshes and point clouds: L recon = 1 T T ∑ t = 1 ∑ c ∈ C 1 | V c t | ∑ v ∈ V c t min p ∈ P c t ∥ v − p ∥ + 1 | P c t | ∑ p ∈ P c t min v ∈ V c t ∥ v − p ∥ (4) T o regularise the smoothness of the meshes, we use an edge loss to minimise the standard deviation of edge length: L edge = 1 T T ∑ t = 1 ∑ c ∈ C 1 | E c | ∑ ( i , j ) ∈ E c e c t , i , j − ¯ e c t 2 1 2 (5) where e c t , i , j : = ∥ v c t , i − v c t , j ∥ is the edge length between the verte x i and j , and ¯ e c t : = 1 | E c | ∑ ( i , j ) ∈ E c e c t , i , j is the mean edge length for structure c and frame t . The curv ature loss L curv encourages all meshes to have the identical curv ature distribution as the template mesh, en- forcing the shape correspondence across all subjects and time frames. For the i -th verte x v c t , i ∈ V c t of structure c at frame t , its mean curvature H c t , i ∈ R is estimated by 79 , 80 , H c t , i = − 1 2 ∆ v c t , i · n c t , i ≈ − 1 2 ( L v c t ) i · n c t , i , (6) where n c t , i ∈ R 3 is the unit normal vector at vertex v c t , i , ∆ is the Laplacian-Beltrami operator 79 , 80 , L ∈ R | V c |×| V c | is the graph Laplacian matrix, and v c t ∈ R | V c |× 3 is the vectorised representation of all vertices v c t , i of the structure c at time frame t . Let H c t ∈ R | V c | denote the curv atures of all v ertices of structure c at frame t , and H c 0 ∈ R | V c | denote the curvatures of the template mesh. The curv ature loss L curv is defined by , L curv = 1 T T ∑ t = 1 ∑ c ∈ C 1 − r ( H c t , H c 0 ) , (7) where r ( H c t , H c 0 ) ∈ [ − 1 , 1 ] is the Pearson correlation coeffi- cient between the curvatures of the reconstructed meshes and the template mesh. The temporal consistency loss L temp regularises the defor - mation between two consecuti ve frames, which is defined as, L temp = 1 T − 1 T − 1 ∑ t = 1 ∑ c ∈ C 1 | V c | | V c | ∑ i = 1 ∥ v c t + 1 , i − v c t , i ∥ . (8) The cycle consistenc y loss L cycle is formulated as, L cycle = ∑ c ∈ C 1 | V c | | V c | ∑ i = 1 ∥ v c T , i − v c 1 , i ∥ , (9) which minimises the distance between the meshes at the start- ing ( t = 1 ) and end ( t = T ) frame of a cardiac cycle, encourag- ing the heart to return to its original shape after one complete cycle 48 . The final loss function is a combination of the loss terms defined as: L all = L recon + λ edge · L edge + λ curv · L curv + λ temp · L temp + λ cycle · L cycle , (10) where the weights λ edge = 0 . 5 , λ curv = 1 . 0 , λ temp = 0 . 1 , and λ cycle = 0 . 2 are empirically chosen. The loss function is min- imised using the Adam optimiser with a learning rate 10 − 4 and a batch size of 1 for 200 epochs, using 600 training and 100 validation subjects from the UK Biobank dataset. The performance of HeartFFDNet is ev aluated using 1,000 test subjects with multi-vie w 2D+t four -chamber contours deri ved from quality-controlled CMR segmentations. Please see Sup- plementary T able 2 for detailed data information. Learning the HeartSSM shape model W e learn a 3D+t cardiac four-chamber statistical shape model, named as HeartSSM, from the UK Biobank imaging popula- tion using incremental PCA 49 – 51 . In order to perform PCA, the 3D+t cardiac four -chamber meshes reconstructed by the deep learning-based pipeline are rigidly aligned to the space of the template mesh. Then, we vectorise all | V | = 27 , 034 ver - tices of each 3D+t four-chamber mesh sequence with T = 50 time frames into a shape vector v ∈ R 3 T | V | . Due to memory limitation, it is computationally intractable to apply standard PCA 81 – 83 to mesh sequences from a lar ge population. Giv en a training set with N subjects, standard PCA explicitly con- structs a co v ariance matrix for N vectorised shapes v of dimen- sionality 3 T | V | , which amounts to a quadratic memory cost of O ( N 2 T 2 | V | 2 ) . T o address the memory limitation, HeartSSM is constructed using incremental PCA 49 – 51 for N = 64 , 000 training subjects (see Supplementary T able 2), which updates 13 the mean shape and the PCs progressiv ely using mini-batches. W e employ a batch size of 128 for incremental PCA and set the number of PCs to M = 128. After training, HeartSSM approximates each 3D+t four- chamber shape vector v by , v ≈ ¯ v + Pw , (11) where ¯ v ∈ R 3 T | V | is the mean shape vector capturing both shape and motion of the four chambers, P ∈ R 3 T | V |× M denotes M = 128 orthogonal PCs learnt from the incremental PCA, and w ∈ R M contains the weights of the PCs. Consequently , the cardiac shape variation and motion pattern for each of the 95,695 UK Biobank participants can be represented by the low-dime nsional vector w , which is also called the HeartSSM shape descriptor . For any unseen test shape vector v , the HeartSSM shape descriptor w can be estimated by , w = P ⊤ ( v − ¯ v ) . (12) The generalisation ability of HeartSSM is ev aluated by fitting 10,000 unseen test four-chamber mesh sequences (Fig. 3 d). HeartSSM can fit onto 3D+t cardiac meshes with missing spatial or temporal information, such as spare contour fitting (Supplementary Fig. 3) or missing sequence completion (Sup- plementary Fig. 4). Gi ven incomplete meshes, we optimise the HeartSSM shape descriptor w to minimise the distance between the reconstructed meshes and incomplete meshes, which can be either sparse contours or a mesh sequence with missing time frames. W e use the Adam optimiser with 0.05 learning rate (500 iterations) for sparse contour fitting and 0.1 learning rate (200 iterations) for missing sequence com- pletion. The fitting ability of HeartSSM is ev aluated using 1,000 test subjects with manual quality control. Please see Supplementary T able 2 for detailed test data information. Correlation analysis of shape v ariation For the cardiac four-chamber mesh of each subject, we as- sess vertex-wise shape variation by comparing the individ- ual subject mesh to the population mean mesh deriv ed from HeartSSM. F or each vertex v t ∈ R 3 at frame t , the shape v aria- tion is quantified by the signed distance s t ∈ R from the vertex v t to the corresponding vertex ¯ v t on the mean mesh, along the normal direction ¯ n t at the verte x ¯ v t , i.e. , s t : = ⟨ v t − ¯ v t , ¯ n t ⟩ . (13) A positi ve shape v ariation ( s t > 0 ) indicates that the indi vid- ual four-chamber mesh expands outw ards along the surface normal direction relativ e to the mean mesh. T o summarise the shape variation across a cardiac cycle, we compute the mean shape variation s = 1 T ∑ T t = 1 s t at each vertex across all time frames. Subsequently , we perform correlation analysis of vertex-wise shape variation against each attribute y (sex, age, BMI, hypertension, atrial fibrillation and flutter , and heart failure). For each vertex, we compute the Pearson correlation coefficient between the shape v ariation s and the attributes y for 62,778 UK Biobank participants. Cardiovascular disease classification W e perform classification for six types of cardiovascular diseases: hypertension ( n = 13 , 453 ), angina pectoris ( n = 2 , 606 ), acute myocardial infraction ( n = 1 , 395 ), chronic ischaemic heart disease ( n = 3 , 940 ), atrial fibrillation and flutter ( n = 2 , 730 ), and heart failure ( n = 804 ). Please see Supplementary T able 1 for detailed data information. W e compare classification performance across different features: confounders (age, sex, weight, height, BMI), image-deri ved phenotypes with/without confounders, shape-deri ved pheno- types with/without confounders, and HeartSSM shape de- scriptor with/without confounders. W e include 17 image- deriv ed or shape-deriv ed phenotypes: left ventricular myocar - dial mass (L VM), ventricular ED and ES v olumes (L VED V , L VESV , R VED V , R VESV), atrial maximum and minimum volumes (LAMAXV , L VMINV , RAMAXV , RAMINV), four- chamber stroke v olumes (L VSV , R VSV , LASV , RASV), and four-chamber ejection fractions (L VEF , R VEF , LAEF , RAEF). W e employ two classifiers, logistic regression and XGBoost 68 , with default hyperparameters e xcept that the learning rate of XGBoost is set to 0.05. The classifiers are trained using four- fold cross validation on 62,778 UK Biobank participants with av ailable demographic and disease information. Chronological age pr ediction For chronological age prediction 52 , we train a CatBoost model 70 for age regression using four-fold cross validation on 9,135 healthy UK Biobank participants, with 0.05 learn- ing rate and 1,000 iterations. W e compare dif ferent features, including image-deriv ed phenotypes, shape-deri ved pheno- types, and HeartSSM shape descriptors. Apart from the proposed HeartSSM constructed using 3D+t four-chamber meshes, called HeartSSM (4CH+t), we also construct two variants of HeartSSM, using 3D four-chamber meshes of ED and ES frames only without motion, called HeartSSM (4CH), and using 3D+t bi-ventricular meshes without two atria, called HeartSSM (2CH+t), respectiv ely . Cardiac shape retrie val In the shape retriev al task, each of the UK Biobank partici- pants is represented by a shape-based feature vector x , which can be image-deriv ed phenotypes, shape-deriv ed phenotypes, or HeartSSM shape descriptor . Each participant is assigned to a group y according to sex, age, weight, height, BMI, or hy- pertension disease (Fig. 6 a). W e randomly select M = 5 , 000 query subjects ( x q , y q ) . The aim of shape retriev al is to retriev e a subject ( x r , y r ) from the UK Biobank population, based on the query vector x q , and assess whether the retrie ved subject belongs to the same group as the query subject, i.e . , y r = y q . For each query vector x q i , i = 1 , ..., M , we run KNN algorithm to retriev e K nearest neighbours x r i k , k = 1 , ..., K , from all participants except for the query subject. Then, the top- K precision is measured by Precision@K = 1 M M ∑ i = 1 1 K K ∑ k = 1 1 { y r i k = y q i } × 100% , (14) 14 where the indicator function 1 { a = b } = 1 if and only if a = b . Shape-based longitudinal re-identification In the longitudinal re-identification task, we utilise a subset of the UK Biobank population, which contains 4,134 participants with tw o CMR imaging scans 2 . 66 ± 1 . 09 years apart. W e use the query vector x ( 2 ) i of the second CMR scan to re-identify the i -th subject from the population, based on the feature vector x ( 1 ) i deri ved from the first CMR scan. W e employ KNN algorithm to retrieve K subjects x ( 1 ) i k , and the top- K recall is defined as Recall@K = 1 M M ∑ i = 1 K ∑ k = 1 1 { i k = i } × 100% , (15) where i k = i means the subject is successfully re-identified within K retrie vals. Data a vailability Raw imaging data and non-imaging participant char- acteristics are av ailable from UK Biobank to ap- prov ed researchers via a standard application process at https://www .ukbiobank.ac.uk/register -apply . Code a vailability The code for this study is publicly av ailable at the Github repository: https://github .com/m-qiang/HeartSSM . References 1. Collins, R. What makes UK Biobank special? The Lancet 379 , 1173–1174 (2012). 2. Sudlow , C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Medicine 12 , e1001779 (2015). 3. Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Natur e 562 , 203–209 (2018). 4. Littlejohns, T . J., Sudlow , C., Allen, N. E. & Collins, R. UK Biobank: opportunities for cardiov ascular research. Eur. Hear. J. 40 , 1158–1166 (2019). 5. Pennell, D. J. Cardiovascular magnetic resonance. Circu- lation 121 , 692–705 (2010). 6. Petersen, S. E. et al. UK Biobank’ s cardiov ascular mag- netic resonance protocol. J. Cardio vasc. Magn. Reson. 18 , 8 (2016). 7. Kramer , C. M. et al. Standardized cardiov ascular mag- netic resonance imaging (CMR) protocols: 2020 update. J. Car diovasc. Magn. Reson. 22 , 17 (2020). 8. Bai, W . et al. Automated cardiov ascular magnetic reso- nance image analysis with fully con volutional networks. J. Car diovasc. Magn. Reson. 20 , 65 (2018). 9. Bai, W . et al. A population-based phenome-wide associ- ation study of cardiac and aortic structure and function. Nat. Medicine 26 , 1654–1662 (2020). 10. Y oung, A. A. & Frangi, A. F . Computational cardiac atlases: from patient to population and back. Exp. Physiol. 94 , 578–596 (2009). 11. Puyol-Anton, E. et al. A multimodal spatiotemporal cardiac motion atlas from MR and ultrasound data. Med. Image Analysis 40 , 96–110 (2017). 12. Bello, G. A. et al. Deep-learning cardiac motion analysis for human survi v al prediction. Nat. Mach. Intell. 1 , 95– 104 (2019). 13. Duchateau, N., King, A. P . & De Craene, M. Machine learning approaches for myocardial motion and deforma- tion analysis. F r ont. Car diovasc. Medicine 6 , 190 (2020). 14. Xia, Y . et al. Automatic 3D+t four-chamber CMR quan- tification of the UK Biobank: integrating imaging and non-imaging data priors at scale. Med. Image Analysis 80 , 102498 (2022). 15. Meng, Q., Bai, W ., O’Regan, D. P . & Rueckert, D. DeepMesh: Mesh-based cardiac motion tracking using deep learning. IEEE T ransactions on Med. Imaging 43 , 1489–1500 (2023). 16. Burns, R. et al. Genetic basis of right and left ventricular heart shape. Nat. Commun. 15 , 9437 (2024). 17. Sørensen, K. et al. Spatio-temporal neural distance fields for conditional generativ e modeling of the heart. In Inter- national Confer ence on Medical Image Computing and Computer-Assisted Intervention , 422–432 (2024). 18. Galazis, C. et al. High-resolution maps of left atrial displacements and strains estimated with 3D cine MRI using online learning neural networks. IEEE T ransactions on Med. Imaging (2025). 19. Qiao, M. et al. A personalized time-resolved 3D mesh generativ e model for un veiling normal heart dynamics. Nat. Mach. Intell. 1–12 (2025). 20. Prakosa, A. et al. Personalized virtual-heart technol- ogy for guiding the ablation of infarct-related ventricular tachycardia. Nat. Biomed. Eng. 2 , 732–740 (2018). 21. Boyle, P . M. et al. Computationally guided personal- ized targeted ablation of persistent atrial fibrillation. Nat. Biomed. Eng. 3 , 870–879 (2019). 22. Nordsletten, D., Niederer, S., Nash, M., Hunter , P . & Smith, N. Coupling multi-physics models to cardiac mechanics. Pr og. Biophys. Mol. Biol. 104 , 77–88 (2011). 23. T aylor, C. A., Fonte, T . A. & Min, J. K. Computational fluid dynamics applied to cardiac computed tomography for nonin vasiv e quantification of fractional flo w reserve: scientific basis. J. Am. Coll. Car diol. 61 , 2233–2241 (2013). 15 24. K ong, F . & Shadden, S. C. Learning whole heart mesh generation from patient images for computational simula- tions. IEEE T ransactions on Med. Imaging 42 , 533–545 (2022). 25. T rayanov a, N. A. Whole-heart modeling: applications to cardiac electrophysiology and electromechanics. Cir c. Res. 108 , 113–128 (2011). 26. Roney , C. H. et al. Predicting atrial fibrillation recur- rence by combining population data and virtual cohorts of patient-specific left atrial models. Circ. Arrhythmia Electr ophysiol. 15 , e010253 (2022). 27. Li, L., Camps, J., Rodriguez, B. & Grau, V . Solving the in verse problem of electrocardiography for c ardiac digital twins: A surve y . IEEE Rev. Biomed. Eng. (2024). 28. Niederer , S. A., Lumens, J. & Trayano v a, N. A. Com- putational models in cardiology . Nat. Rev. Cardiol. 16 , 100–111 (2019). 29. Corral-Acero, J. et al. The ‘Digital T win’ to enable the vision of precision cardiology . Eur. Hear. J. 41 , 4556– 4564 (2020). 30. Niederer , S. A., Sacks, M. S., Girolami, M. & W illcox, K. Scaling digital twins from the artisanal to the industrial. Nat. Comput. Sci. 1 , 313–320 (2021). 31. Qian, S. et al. Dev eloping cardiac digital twin popula- tions powered by machine learning pro vides electrophys- iological insights in conduction and repolarization. Nat. Car diovasc. Res. 4 , 624–636 (2025). 32. Frangi, A. F ., Rueckert, D., Schnabel, J. A. & Niessen, W . J. Automatic construction of multiple-object three- dimensional statistical shape models: Application to car- diac modeling. IEEE T ransactions on Med. Imaging 21 , 1151–1166 (2003). 33. Lötjönen, J., Kivistö, S., K oikkalainen, J., Smutek, D. & Lauerma, K. Statistical shape model of atria, v entricles and epicardium from short-and long-axis MR images. Med. Image Analysis 8 , 371–386 (2004). 34. Bai, W . et al. A bi-ventricular cardiac atlas built from 1000+ high resolution MR images of healthy subjects and an analysis of shape and motion. Med. Image Analysis 26 , 133–145 (2015). 35. Suinesiaputra, A. et al. Statistical shape modeling of the left ventricle: myocardial infarct classification challenge. IEEE J. Biomed. Heal. Informatics 22 , 503–515 (2017). 36. Y oung, A. A., Cowan, B. R., Thrupp, S. F ., Hedley , W . J. & Dell’Italia, L. J. Left ventricular mass and v olume: fast calculation with guide-point modeling on MR images. Radiology 216 , 597–602 (2000). 37. Meng, Q. et al. MulV iMotion: Shape-aw are 3D myocar- dial motion tracking from multi-vie w cardiac MRI. IEEE T ransactions on Med. Imaging 41 , 1961–1974 (2022). 38. Beetz, M., Banerjee, A., Ossenberg-Engels, J. & Grau, V . Multi-class point cloud completion networks for 3D cardiac anatomy reconstruction from cine magnetic reso- nance images. Med. Image Analysis 90 , 102975 (2023). 39. Deng, Y . et al. ModusGraph: automated 3D and 4D mesh model reconstruction from cine CMR with impro ved ac- curacy and efficienc y . In International Conference on Medical Image Computing and Computer-Assisted Inter- vention , 173–183 (2023). 40. Gaggion, N. et al. Multi-view h ybrid graph con v olutional network for volume-to-mesh reconstruction in cardiov as- cular MRI. Med. Image Analysis 103630 (2025). 41. Qiao, M. et al. Mesh4D: A motion-aware multi-vie w variat ional autoencoder for 3D+t mesh reconstruction. In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , 343–353 (2025). 42. Hoogendoorn, C. et al. A high-resolution atlas and statis- tical model of the human heart from multislice CT. IEEE T ransactions on Med. Imaging 32 , 28–44 (2012). 43. SCO T -HEAR T In vestigators, T . CT coronary angiogra- phy in patients with suspected angina due to coronary heart disease (SCOT -HEAR T): an open-label, parallel- group, multicentre trial. The Lancet 385 , 2383–2391 (2015). 44. Rodero, C. et al. Linking statistical shape models and simulated function in the healthy adult human heart. PLoS Comput. Biol. 17 , e1008851 (2021). 45. Xu, H. et al. Whole heart 3D shape reconstruction from sparse vie ws: lev eraging cardiac computed tomography for cardiov ascular magnetic resonance. In International Confer ence on Functional Imaging and Modeling of the Heart , 255–264 (2023). 46. Xu, Y . et al. Improv ed 3D whole heart geometry from sparse CMR slices. In International W orkshop on Sta- tistical Atlases and Computational Models of the Heart , 43–52 (2024). 47. Muffoletto, M. et al. Evaluation of deep learning estima- tion of whole heart anatomy from automated cardiov as- cular magnetic resonance short-and long-axis analyses in UK Biobank. Eur. Hear. J ournal-Car diovascular Imag- ing 25 , 1374–1383 (2024). 48. Ma, Q. et al. CardiacFlow: 3D+t Four-Chamber Cardiac Shape Completion and Generation via Flo w Matching. In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , 89–99 (2025). 49. W eng, J., Zhang, Y . & Hwang, W .-S. Candid covariance- free incremental principal component analysis. IEEE T ransactions on P attern Analysis Mach. Intell. 25 , 1034– 1040 (2003). 50. Zhao, H., Y uen, P . C. & Kwok, J. T . A novel incremental principal component analysis and its application for face 16 recognition. IEEE T ransactions on Syst. Man, Cybern. 36 , 873–886 (2006). 51. Brand, M. Fast low-rank modifications of the thin singular value decomposition. Linear Algebr. its Appl. 415 , 20–30 (2006). 52. Shah, M. et al. Environmental and genetic predictors of human cardiov ascular ageing. Nat. Commun. 14 , 4941 (2023). 53. V illard, B., Zacur , E., Dall’Armellina, E. & Grau, V . Cor- rection of slice misalignment in multi-breath-hold cardiac MRI scans. In International W orkshop on Statistical At- lases and Computational Models of the Heart , 30–38 (2016). 54. Zeng, A. et al. ImageCAS: A large-scale dataset and benchmark for coronary artery segmentation based on computed tomography angiography images. Comput. Med. Imaging Graph. 109 , 102287 (2023). 55. W asserthal, J. et al. T otalSegmentator: robust segmen- tation of 104 anatomic structures in CT images. Radiol. Artif. Intell. 5 , e230024 (2023). 56. Zhuang, X. et al. Evaluation of algorithms for multi- modality whole heart segmentation: an open-access grand challenge. Med. Image Analysis 58 , 101537 (2019). 57. Zhuang, X. Multi variate mixture model for myocar- dial segmentation combining multi-source images. IEEE T ransactions on P attern Analysis Mach. Intell. 41 , 2933– 2946 (2019). 58. T obon-Gomez, C. et al. Benchmark for algorithms se g- menting the left atrium from 3D CT and MRI datasets. IEEE T ransactions on Med. Imaging 34 , 1460–1473 (2015). 59. Kiri ¸ sli, H. et al. Standardized ev aluation frame work for ev aluating coronary artery stenosis detection, stenosis quantification and lumen segmentation algorithms in com- puted tomography angiography . Med. Image Analysis 17 , 859–876 (2013). 60. Metz, C. et al. Coronary centerline extraction from CT coronary angiography images using a minimum cost path approach. Med. Phys. 36 , 5568–5579 (2009). 61. Davies, R., T wining, C. & T aylor, C. Statistical models of shape: Optimisation and evaluation (Springer Science & Business Media, 2008). 62. Booth, J., Roussos, A., Zafeiriou, S., Ponniah, A. & Dun- away , D. A 3D morphable model learnt from 10,000 faces. In IEEE Confer ence on Computer V ision and P at- tern Recognition , 5543–5552 (2016). 63. W o ˙ zako wska-Kapłon, B. Changes in left atrial size in patients with persistent atrial fibrillation: a prospectiv e echocardiographic study with a 5-year follow-up period. Int. J. Car diol. 101 , 47–52 (2005). 64. Gupta, D. K. et al. Left atrial structure and function in atrial fibrillation: ENGAGE AF-TIMI 48. Eur. Hear. J. 35 , 1457–1465 (2014). 65. Friedberg, M. K. & Redington, A. N. Right versus left ventricular f ailure: differences, similarities, and interac- tions. Cir culation 129 , 1033–1044 (2014). 66. UK, N. A.-A. et al. 2016 ESC guidelines for the diagnosis and treatment of acute and chronic heart failure. Eur. Hear. J. 37 , 2129–2200 (2016). 67. Bozkurt, B. et al. Universal definition and classification of heart failure: a report of the heart failure society of America, heart failure association of the European So- ciety of Cardiology , Japanese Heart Failure Society and Writing Committee of the Uni versal Definition of Heart Failure. J. Car diac F ail. 27 , 387–413 (2021). 68. Chen, T . & Guestrin, C. XGBoost: A scalable tree boost- ing system. In Pr oceedings of the 22nd ACM SIGKDD International Confer ence on Knowledge Discovery and Data Mining , 785–794 (2016). 69. Sun, X. & Xu, W . Fast implementation of DeLong’ s algo- rithm for comparing the areas under correlated recei ver operating characteristic curves. IEEE Signal Pr ocess. Lett. 21 , 1389–1393 (2014). 70. Prokhorenkov a, L., Guse v , G., V orobev , A., Dorogush, A. V . & Gulin, A. CatBoost: unbiased boosting with categorical features. Adv. Neural Inf. Pr ocess. Syst. 31 (2018). 71. T angelder, J. W . & V eltkamp, R. C. A survey of content based 3D shape retriev al methods. Pr oc. Shape Model. Appl. 145–156 (2004). 72. Chang, A. X. et al. ShapeNet: An information-rich 3D model repository . arXiv pr eprint (2015). 73. Kingma, D. P . & Ba, J. Adam: A method for stochastic optimization. In International Conference on Learning Repr esentations (2015). 74. Zhuang, X. & Shen, J. Multi-scale patch and multi- modality atlases for whole heart segmentation of MRI. Med. Image Analysis 31 , 77–87 (2016). 75. Schuh, A. et al. A deformable model for the reconstruc- tion of the neonatal cortex. In IEEE International Sympo- sium on Biomedical Imaging , 800–803 (2017). 76. Sederberg, T . W . & Parry , S. R. Free-form deformation of solid geometric models. In Pr oceedings of the 13th An- nual Confer ence on Computer Graphics and Interactive T echniques , 151–160 (1986). 77. Ronneberger , O., Fischer , P . & Brox, T . U-net: Con v olu- tional networks for biomedical image segmentation. In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , 234–241 (2015). 17 78. Lorensen, W . E. & Cline, H. E. Marching cubes: A high resolution 3D surface construction algorithm. In Seminal Graphics: Pioneering Efforts that Shaped the F ield , 347–353 (1998). 79. Nealen, A., Igarashi, T ., Sorkine, O. & Alexa, M. Lapla- cian mesh optimization. In Pr oceedings of the 4th Interna- tional Confer ence on Computer Graphics and Interactive T echniques in Australasia and Southeast Asia , 381–389 (2006). 80. Do Carmo, M. P . Differ ential geometry of curves and surfaces (Courier Dov er Publications, 2016). 81. Abdi, H. & Williams, L. J. Principal component analysis. W iley Interdiscip. Rev. Comput. Stat. 2 , 433–459 (2010). 82. Gewers, F . L. et al. Principal component analysis: A natural approach to data exploration. ACM Comput. Surv. 54 , 1–34 (2021). 83. Greenacre, M. et al. Principal component analysis. Nat. Rev. Methods Primers 2 , 100 (2022). Acknowledgements This research was conducted using the UK Biobank Re- source under Application Number 18545. W e thank all UK Biobank participants and staff. This work is supported by the EPSRC grants (EP/W01842X/1, EP/Z531297/1) and the BHF New Horizons Grant (NH/F/23/70013). S.N. is sup- ported by the National Institutes of Health (R01-HL152256), the European Research Council (PREDICT -HF 864055), the British Heart Foundation (RG/20/4/34803), the EPSRC (EP/X012603/1 and EP/P01268X/1), the T echnology Mis- sions Fund under the EPSRC (EP/X03870X/1), and the Alan T uring Institute. D.P .O. is supported by the Medical Re- search Council (MC_UP_1605/13), the NIHR Imperial Col- lege Biomedical Research Centre, and the British Heart Foun- dation (RG/F/24/110138, RE/24/130023, CH/F/24/90015). P .M.M. acknowledges generous personal support from the Edmond J. Safra Foundation and Lily Safra, an NIHR Senior In vestig ator A ward, Rosalind Franklin Institute, and the UK Dementia Research Institute, which is funded predominantly by the UKRI Medical Research Council. Competing interests S.N. has recei ved research funding from GSK, ANSYS and Synopsis. D.P .O. has consulted for Bayer A G and Bristol- Myers-Squibb, and received grants from Bayer and Calico Labs. P .M.M. has received consultanc y or speaker fees from Roche, Merck, Biogen, Rejuv eron, Sangamo, Nodthera and Nov artis. P .M.M. has recei ved research or educational funds from Biogen, Nov artis, Merck and GlaxoSmithKline. The remaining authors declare no competing interests. 18
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment