MRI-to-CT synthesis using drifting models
Accurate MRI-to-CT synthesis could enable MR-only pelvic workflows by providing CT-like images with bone details while avoiding additional ionizing radiation. In this work, we investigate recently proposed drifting models for synthesizing pelvis CT i…
Authors: Qing Lyu, Jianxu Wang, Jeremy Hudson
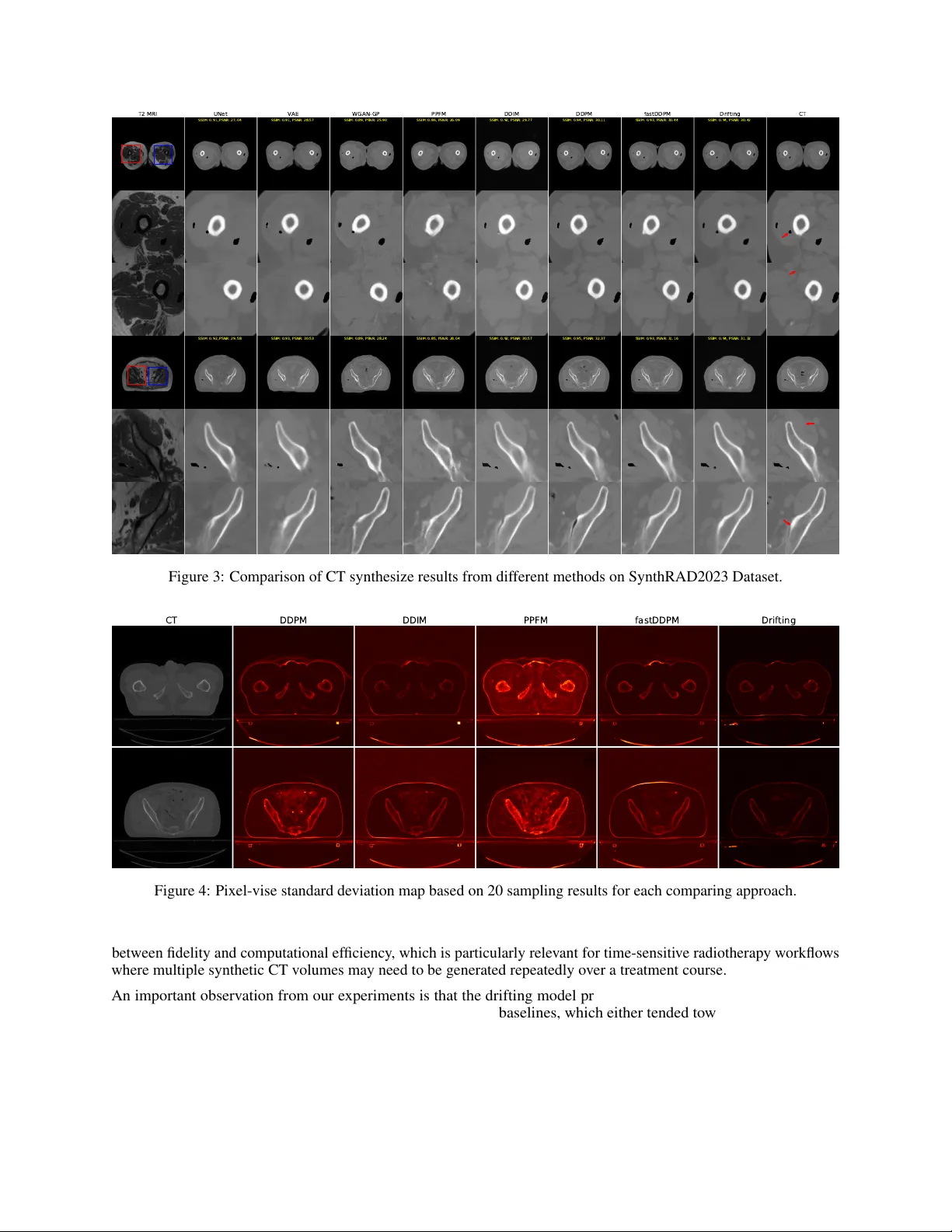
M R I - T O - C T S Y N T H E S I S U S I N G D R I F T I N G M O D E L S Qing L yu, Jer emy Hudson, Chirstopher T . Whitlow Department of Radiology and Biomedical Imaging Y ale School of Medicine New Ha ven, CT {qing.lyu, jeremy.hudson, christopher.whitlow}@yale.edu Jianxu W ang, Ge W ang Department of Biomedical Engineering Rensselaer Polytechnic Institute T roy , NY {wangj68, wangg6}@email A B S T R AC T Accurate MRI-to-CT synthesis could enable MR-only pelvic workflows by pro viding CT -like images with bone details while a voiding additional ionizing radiation. In this work, we in vestigate recently proposed drifting models for synthesizing pelvis CT images from MRI and benchmark them against con volutional neural networks (UNet, V AE), a generativ e adversarial netw ork (WGAN-GP), a physics-inspired probabilistic model (PPFM), and diffusion-based methods (FastDDPM, DDIM, DDPM). Experiments are performed on two complementary datasets: Gold Atlas Male Pelvis and the SynthRAD2023 pelvis subset. Image fidelity and structural consistency are e valuated with SSIM, PSNR, and RMSE, complemented by qualitativ e assessment of anatomically critical regions such as cortical bone and pelvic soft-tissue interfaces. Across both datasets, the proposed drifting model achiev es high SSIM and PSNR and low RMSE, surpassing strong diffusion baselines and con ventional CNN-, V AE-, GAN-, and PPFM-based methods. V isual inspection shows sharper cortical bone edges, improv ed depiction of sacral and femoral head geometry , and reduced artifacts or over -smoothing, particularly at bone–air–soft tissue boundaries. Moreov er , the drifting model attains these gains with one-step inference and inference times on the order of milliseconds, yielding a more fav orable accuracy–ef ficiency trade-off than iterativ e diffusion sampling while remaining competitiv e in image quality . These findings suggest that drifting models are a promising direction for fast, high-quality pelvic synthetic CT generation from MRI and warrant further in vestigation for downstream applications such as MRI-only radiotherapy planning and PET/MR attenuation correction. K eywords Medical Image Synthesis · Drifting Models · Computed T omography · Magnetic Resonance Imaging · Artificial Intelligence 1 Introduction Medical imaging often requires both Computed T omography (CT) and Magenetic Resonance Imaging (MRI) modalities for comprehensiv e clinical assessment—CT provides Hounsfield unit (HU) electron density information essential for radiotherapy dose calculation, while MRI of fers superior soft-tissue contrast for tumor delineation and diagnosis. Acquiring both modalities increases cost, radiation exposure, and introduces misalignment between scans [ 1 , 2 , 3 ]. In addition, CT acquisition may be constrained by practical factors such as equipment av ailability and maintenance burden, leading to v ariability in access and leaving some clinical or research cohorts without CT coverage [ 4 , 5 ]. These limitations, together with the relativ e rarity of high-quality paired and well-registered CT–MRI datasets, make CT synthesis from MRI clinically important because it can provide CT -like quantitati ve surrogates for CT -dependent tasks (e.g., attenuation correction and MRI-only radiotherapy workflo ws) when CT is unav ailable or undesirable [6, 7] Running T itle for Header Cross-modality synthesis—generating one modality from the other—has been a major research direction, but e xisting approaches face notable limitations [ 7 , 8 , 9 , 10 ]. Early strategies include atlas-based registration and patch-based learning, while more recent deep learning solutions span supervised con volutional neural netw orks (e.g., U-Net v ariants) and generative modeling frameworks [ 11 , 12 , 13 , 14 , 15 , 16 ]. Generativ e approaches such as V AEs and GANs can improv e realism and sharpness but may suf fer from blurring (V AEs) or training instability and hallucinated structures (GANs), especially near bone–air–soft tissue interf aces [ 17 , 18 , 19 , 20 , 21 , 22 , 23 , 24 , 25 , 26 , 27 ]. Diffusion models hav e shown strong fidelity and robustness, yet their iterati ve sampling can impose substantial inference-time costs that limit clinical practicality , moti vating interest in faster generati ve alternativ es [28, 29, 30]. Drifting models are a recent family of generativ e methods that learn a continuous “drift” (vector -field) dynamics to transform a simple source distrib ution into the target image distribution, making them naturally suited to conditional synthesis tasks such as MRI-to-CT by learning how an input-conditioned representation should ev olve toward a CT -like output [ 31 ]. Compared with dif fusion models, drifting models can be implemented with fe wer integration steps and therefore offer faster inference in practice while targeting comparable perceptual fidelity . Relative to GAN-based synthesis, which may be sensiti ve to training instability and can produce visually plausible yet unfaithful structures, drifting models provide an appealing balance between realism and controllability for anatomy-preserving medical translation. In this study , we in vestig ate recently emerged drifting models for pelvic MRI-to-CT synthesis and compare them against representativ e baselines including conv olutional neural network (CNN), variational autoencoder (V AE), GAN, and dif fusion models. W e conduct experiments on tw o datasets—SynthRAD2023 and Gold Atlas Male Pelvis—and ev aluate performance using structural similarity index (SSIM) and peak signal-to-noise ratio (PSNR), alongside qualitative assessment in anatomically critical regions. Our key contributions are: (1) modification of original drifting models and adding MRI as conditions for controllable CT synthesis, (2) a systematic benchmark of drifting models against established synthesis families on two complementary pelvic datasets, and (3) e vidence that drifting models can achieve high-quality synthetic CT generation with notably fast inference, offering a fa vorable accuracy–ef ficiency trade-off for downstream use. 2 Related Studies 2.1 Atlas/registration-based CT synthesis approaches Early sCT pipelines typically rely on aligning one or more atlas MR–CT pairs to a tar get MRI and then fusing the warped CTs into a pseudo-CT ; these methods are conceptually appealing because they encode prior anatomical correspondences, but they can be sensitive to registration quality and population mismatch. Uh et al. is a representative early study in this vein, demonstrating the feasibility of MRI-based radiotherapy treatment planning by generating pseudo-CT via atlas registration and then assessing its suitability for dose calculation [ 12 ]. Extending the atlas concept toward PET/MR workflo ws, Burgos et al. focus on head-and-neck attenuation correction and describe an iterativ e multi-atlas strategy , underscoring that reliable pseudo-CT generation is also critical when CT is una vailable for attenuation mapping in PET/MR [ 13 ]. Building directly on multi-atlas synthesis for radiotherapy , Guerreiro et al. e valuate a multi-atlas sCT approach for MRI-only treatment planning in head-and-neck and prostate cases, comparing it to a b ulk-density assignment alternativ e and showing that multi-atlas synthesis can achieve clinically small dose differences while improving automatic bone delineation [ 11 ]. In parallel, Roy et al. highlight a complementary perspecti ve: rather than treating synthesis only as the final deliv erable, they use image synthesis to enable improved MR-to-CT brain registration, illustrating ho w synthesis can reduce multimodal mismatch during alignment [ 15 ]. Lee et al. further refine the atlas pipeline by combining multi-atlas fusion with patch-based refinement, aiming to correct local details that global registration-and-fusion may miss [16]. 2.2 V AE/GAN-based approaches As CT synthesis shifted from atlas/re gistration pipelines to generativ e modeling, V AE- and GAN-based approaches became prominent because they enable direct MRI-to-CT image translation with substantially faster inference and improv ed representation power . In broad terms, V AE-based methods emphasize stable optimization and anatomically faithful reconstructions through probabilistic latent representations, whereas GAN-based methods tend to deliver sharper , more realistic-looking CT textures but often require additional constraints to mitigate hallucinations and preserve geometry in unpaired or weakly paired training. On the V AE side, Iyer et al. propose an MRI-to-CT synthesis framework for pediatric cranial imaging b uilt on a V AE. They further incorporate cranial suture segmentations as an auxiliary supervised signal, and report high structural similarity for sCT along with strong skull and suture segmentation performance on an in-house pediatric dataset [17]. 2 Running T itle for Header On the GAN side, much of the progress is dri ven by adding explicit mechanisms to improve structural reliability under imperfect pairing. Lei et al. introduce dense cycle-consistent GANs for MRI-only sCT generation, using cycle- consistency to reduce dependence on strictly paired datasets while encouraging anatomically plausible translation [ 20 ]. Y ang et al. extend this line with a structure-constrained CycleGAN by adding a structure-consistency loss based on the modality-independent neighborhood descriptor , explicitly tar geting geometric distortions that can arise in unconstrained CycleGAN mappings [ 24 ]. For head-and-neck radiotherapy , Liu et al. present a multi-cycle GAN that strengthens cyclic constraints to stabilize learning and better preserve anatomy in this challenging region [ 27 ]. In thoracic imaging, Matsuo et al. de velop an unsupervised attention-based framew ork for chest MRI to CT translation, reflecting the need for stronger guidance when motion and low-signal re gions make structure preserv ation difficult [ 25 ]. Gong et al. further push constraint design with cycleSimulationGAN, combining channel-wise attention with structural-similarity constraints—including a contour mutual information loss—to improve detail recovery and structural retention for unpaired head-and-neck MR-to-CT synthesis [26]. Sev eral works also emphasize clinical readiness and ev aluation beyond image similarity . W ang et al. apply GAN-based MRI-to-CT generation for intracranial tumor radiotherapy planning, illustrating a clinically oriented deployment focused on planning feasibility [ 21 ]. Hsu et al. study sCT generation for MRI-guided adaptive radiotherapy in prostate cancer , where models must produce reliable sCT repeatedly as anatomy e volves o ver the treatment course [ 22 ]. Lemus et al. reinforce that clinical acceptance should be assessed with task-based endpoints—such as dose metrics, D VH agreement, and gamma analysis—rather than relying solely on pixel-wise similarity [ 23 ]. Finally , Skandarani et al. provide an empirical study of GANs for medical image synthesis, offering practical insight into GAN behavior and common ev aluation pitfalls that are important when interpreting and comparing GAN-based sCT pipelines [19]. 2.3 Diffusion-based approaches Diffusion-based synthesis replaces one-shot generation with a learned conditional denoising process, which often improv es fidelity and reduces some GAN-specific failure modes at the cost of iterative sampling. L yu and W ang explores MRI-to-CT con version using dif fusion and score-matching models, representing an early effort to apply dif fusion-based models to CT synthesis [ 30 ]. Pan et al. presents an approach using a 3D transformer -based denoising diffusion model with a Gaussian forward noise process on CT and a re verse denoising process conditioned on MRI, reporting improv ed image metrics and dosimetric agreement with generation in minutes [ 28 ]. Özbey et al. proposes unsupervised medical image translation with adversarial dif fusion models, using a conditional diffusion translation mechanism plus adversarial/c ycle-style constraints to enable unpaired cross-modal translation while accelerating sampling through larger rev erse steps [ 29 ]. Li et al. proposes FDDM, an unsupervised MR → CT translation method that separates anatomy con version from appearance generation and uses frequency-decoupled diffusion with dual re verse paths for lo w/high frequency components, reporting impro ved fidelity and anatomical accuracy on brain and pelvis MR → CT translation benchmarks [18]. 3 Methodology 3.1 Dataset 3.1.1 Gold Atlas Male Pelvis Dataset The Gold Atlas Male Pelvis Dataset is a public pelvic MRI-CT dataset comprising 19 male patients, where each case includes a planning CT , a T1-weighted MRI, a T2-weighted MRI, and a deformably registered CT volume [ 32 ]. The data were acquired at three Swedish radiotherap y departments using site-specific clinical protocols and dif ferent scanners, which makes the dataset suitable for validating cross-site MRI-to-CT synthesis methods. For CT , the reported scanners were Siemens Somatom Definition AS+, T oshiba Aquilion, and Siemens Emotion 6, with slice thicknesses of 3.0, 2.0, and 2.5 mm , respectiv ely , and in-plane pixel sizes of approximately 0.98 × 0.98, 1.0 × 1.0, and 0.98 × 0.98 mm 2 . For MRI, the dataset includes both T1-weighted and T2-weighted acquisitions; the T1-weighted scans were acquired with slice thicknesses of 3.0, 2.0, and 2.5 mm and in-plane pixel sizes ranging from 0.875 × 0.875 to 1.1 × 1.1 mm 2 , while the T2-weighted scans used 2.5 mm slice thickness and in-plane pixel sizes from 0.875 × 0.875 to 1.1 × 1.1 mm 2 . The original dataset article reports site-specific acquisition settings rather than a single common image matrix for all subjects, reflecting the heterogeneous multi-center design. 3.1.2 SynthRAD2023 Dataset SynthRAD2023 is a large multi-center benchmark for synthetic CT generation in radiotherapy , containing 540 paired MRI-CT sets and 540 paired cone beam CT -CT sets across brain and pelvis anatomies [ 33 ]. In this work, we use the MRI-to-CT pelvis subset (T ask 1 pelvis), which contains 270 paired MRI-CT scans. This subset was collected from 3 Running T itle for Header two centers: Center A acquired pelvic MRI scans on Philips Ingenia 1.5T/3.0T systems using 3D T1-weighted spoiled gradient echo imaging, with in-plane pixel spacing between 0.94 and 1.14 mm , rows ranging from 400 to 528, and columns ranging from 103 to 528; Center C acquired scans on Siemens MA GNETOM A v anto, Skyra, and V ida 3T systems using 3D T2-weighted SP A CE imaging, with in-plane pixel spacing between 1.17 and 1.30 mm and matrix size 288 × 384. The paired CT scans were reconstructed at 512 × 512 in-plane size, with pixel spacing of 0.77-1.37 mm at Center A and 0.98-1.17 mm at Center C, and slice thicknesses of 1.5 to 3.0 mm and 2.0 to 3.0 mm , respectiv ely . 3.2 Image prepr ocessing All MRI and CT volumes were first resampled to an isotropic spatial resolution of 1 × 1 × 1 mm 3 . After resampling, each slice was centrally cropped or zero padded to ensure a uniform in-plane image size of 512 × 512. This step standardizes the field of vie w across subjects and simplifies mini-batch training. Finally , intensity normalization was performed independently for each scan so that inter-subject intensity scale variations were reduced before network training. Let x denote an input volume and T ( · ) the preprocessing operator; then the final network input can be written as x prep = T ( x ) , (1) where T ( · ) includes isotropic resampling, central cropping or zero padding, and per-scan intensity normalization. 3.3 Drifting Model Drifting models reformulate generativ e modeling as a train-time distribution ev olution problem rather than an iterative test-time denoising process. Instead of progressively refining a noisy sample during inference, the model learns during optimization ho w the generator distribution should mo ve tow ard the target data distribution. As a result, the transport process is absorbed into training, while inference remains a one-step mapping from MRI to synthetic CT [31]. Let m denote an MRI input, c the corresponding target CT , and ϵ ∼ p ( ϵ ) random noise. A conditional generator f θ produces a synthetic CT ˆ c based on conditional image m and noise ϵ ˆ c = f θ ( m, ϵ ) . (2) In the spirit of drifting models, adapted from the paper’ s unconditional pushforward formulation, the proposed conditional output distribution q θ ( · | m ) is expressed as follo ws q θ ( · | m ) = [ f θ ( m, · )] # p ( ϵ ) . (3) T o simplify the drifting formulation, we compute the drifting loss directly in the image domain rather than in a learned feature space. Let ˆ c denote a generated CT image, let c + denote a real CT sample from the tar get distribution, and let c − denote a generated CT sample from the current model distribution. The drifting field is therefore defined directly on image intensities, so the generator is trained to move each synthesized CT tow ard nearby real CT images while repelling it from other generated CT images. This follows the original ra w-space drifting formulation, but is adapted here to conditional MRI-to-CT synthesis. For a generated sample ˆ c , the general drifting field is written as V p,q (ˆ c ) = E c + ∼ p ( ·| m ) E c − ∼ q ( ·| m ) K ( ˆ c, c + , c − ) . (4) where K ( · , · , · ) is a kernel-like function describing interactions among three sample points. The drift field in drifting models is built from attraction to ward data samples and repulsion from generated samples, with anti-symmetry ensuring zero drift at equilibrium when the two distrib utions match: V + p (ˆ c ) = 1 Z p (ˆ c ) E c + ∼ p ( ·| m ) k (ˆ c, c + )( c + − ˆ c ) , (5) V − q (ˆ c ) = 1 Z q (ˆ c ) E c − ∼ q ( ·| m ) k (ˆ c, c − )( c − − ˆ c ) , (6) where the normalization factors are Z p (ˆ c ) = E c + ∼ p ( ·| m ) k (ˆ c, c + ) , Z q (ˆ c ) = E c − ∼ q ( ·| m ) k (ˆ c, c − ) . (7) k ( · , · ) is the pairwise similarity kernel k ( a, b ) = exp( − 1 τ ∥ a − b ∥ 2 ) . (8) 4 Running T itle for Header The final drifting field is defined as V p,q (ˆ c ) = V + p (ˆ c ) − V − q (ˆ c ) . (9) At equilibrium state, the generator output should satisfy the fixed-point relation f ˆ θ ( m, ϵ ) = f ˆ θ ( m, ϵ ) + V p,q ˆ θ f ˆ θ ( m, ϵ ) . (10) This motiv ates the training-time update f θ i +1 ( m, ϵ ) ← f θ i ( m, ϵ ) + V p,q θ i ( f θ i ( m, ϵ )) . (11) W e conv ert this update rule into the drifting loss L drift = E m,ϵ h ∥ f θ ( m, ϵ ) − sg( f θ ( m, ϵ ) + V p,q θ ( f θ ( m, ϵ ))) ∥ 2 2 i , (12) where sg( · ) denotes the stop-gradient operator . For paired MRI-to-CT synthesis, an additional vox el-level reconstruction term can be included to preserve anatomical correspondence: L total = λ drift L drift + λ 1 ∥ ˆ c − c ∥ 1 , (13) where λ drift and λ 1 control the balance between distribution-le vel drifting and vox el-wise fidelity . 3.4 Implementation Details W e split both Gold Atlas and SynthRAD2023 datasets based on 70%/10%/20% rule to create training, validation, and testing subsets, respecti vely . The conditional generation in the drifting model is a UNet-lik e model. For drifting loss computation, 16 positiv e and negati ve samples are created by randomly cropping 64 × 64 patches from real image and generated images, respectiv ely . In (13) , λ drif t and λ 1 are empirically set to 1 and 10, respectiv ely . Adam optimizer is used and the learning rate is set to 0.0001. The training loss is recorded and monitored during the training process. When the training loss does not decrease more than 1% for the most recent 20 epochs, the early stop triggers and the training model is sav ed for the later inference. All experiments are conducted using Pytorch via a single Nvidia H200 GPU. 4 Results T o ensure a fair comparison, we benchmark the proposed MRI - conditioned drifting model against a diverse set of representativ e baselines, including con volutional encoder–decoder architectures (UNet, V AE) [ 34 , 35 ], an adversar- ial model (WGAN - GP) [ 36 ], a physics - inspired probabilistic framew ork (PPFM) [ 37 ], and several diffusion - based approaches (FastDDPM, DDIM, DDPM) [ 38 , 39 , 40 ]. All models are implemented with a comparable number of trainable parameters and similar backbone capacity , so that performance dif ferences primarily reflect the underlying generativ e mechanism and conditioning strategy rather than model size. DDPM inference is based on a 1000-step denoising, while DDIM and fastDDPM is implemented via a 20-step sampling. PPFM inference results are made by a 18-step sampling process. 4.1 Results on Gold Atlas Male Pelvis dataset On the Gold Atlas dataset, the proposed drifting model outperforms all competing methods except DDPM in terms of SSIM, PSNR, and RMSE. Quantitati vely , it attains high SSIM and PSNR and low RMSE, surpassing the dif fusion-based models such as FastDDPM, PPFM, and DDIM, as well as non - diffusion methods such as UNet, V AE, and WGAN - GP . The only exception is DDPM, which obtains the highest SSIM and PSNR and the lowest RMSE. V isually , Figure 1 shows that the proposed drifting model better preserv es fine pelvic structures and soft - tissue boundaries, with clearer organ details and reduced o ver - smoothing compared with UNet/V AE and fe wer noise - induced artifacts than WGAN - GP and PPFM. Compared with other dif fusion models, drifting produces sharper cortical bone edges and more accurate Hounsfield unit transitions, which align more closely with the reference CT . A representativ e example in Figure 1 illustrates these trends: UNet and V AE yield overly smooth bone–soft - tissue transitions, WGAN - GP introduces artifacts that hinder image quality , and DDIM/PPFM/fastDDPM produce good results with clear tissue boundary but lower image quality when comparing with DDPM. Among all approaches, DDPM demonstrates the best ability to clearly demonstrate bon e details and tissue boundary . As for the proposed drifting model, it can bring about comparable results with DDIM/fastDDPM and present clear tissue boundary and bone structure. T able 1 presented quantitativ e comparison of synthetic CT from different approaches. It can be found that the proposed drifting model slightly ov ercome DDIM and fastDDPM but not as good as DDPM. 5 Running T itle for Header T2 MRI SSIM: 0.81, PSNR : 29.01 Unet SSIM: 0.81, PSNR : 28.77 V AE SSIM: 0.81, PSNR : 29.31 WGAN- GP SSIM: 0.82, PSNR : 29.83 DDPM SSIM: 0.80, PSNR : 28.93 DDIM SSIM: 0.78, PSNR : 26.96 PPFM SSIM: 0.80, PSNR : 29.27 fastDDPM SSIM: 0.83, PSNR : 30.28 Drif ting CT SSIM: 0.87, PSNR : 30.32 SSIM: 0.88, PSNR : 30.24 SSIM: 0.88, PSNR : 30.11 SSIM: 0.92, PSNR : 31.97 SSIM: 0.87, PSNR : 30.56 SSIM: 0.78, PSNR : 28.29 SSIM: 0.87, PSNR : 30.84 SSIM: 0.92, PSNR : 31.53 Figure 1: Comparison of CT synthesize results from different methods on Gold Atlas Dataset. T able 1: Quantitative comparison of v arious methods Gold Atlas SynthRAD2023 SSIM PSNR RMSE SSIM PSNR RMSE UNet 0.835 ± 0.030 29.53 ± 1.44 0.041 ± 0.003 0.935 ± 0.012 28.74 ± 1.68 0.040 ± 0.017 V AE 0.840 ± 0.039 29.26 ± 1.74 0.043 ± 0.003 0.931 ± 0.013 29.29 ± 1.87 0.043 ± 0.012 WGAN-GP 0.853 ± 0.035 29.64 ± 1.01 0.045 ± 0.002 0.909 ± 0.007 27.68 ± 1.69 0.045 ± 0.021 PPFM 0.823 ± 0.054 27.08 ± 2.25 0.084 ± 0.019 0.871 ± 0.028 27.18 ± 2.05 0.048 ± 0.023 FastDDPM 0.865 ± 0.041 30.58 ± 1.38 0.039 ± 0.003 0.948 ± 0.015 29.83 ± 1.75 0.033 ± 0.010 DDIM 0.854 ± 0.042 30.13 ± 1.42 0.042 ± 0.004 0.946 ± 0.021 30.12 ± 1.64 0.035 ± 0.013 DDPM 0.889 ± 0.045 31.34 ± 1.64 0.031 ± 0.003 0.969 ± 0.012 31.51 ± 1.84 0.024 ± 0.008 Drifting 0.869 ± 0.043 30.56 ± 1.23 0.035 ± 0.002 0.948 ± 0.009 30.82 ± 1.69 0.030 ± 0.011 4.2 Inference speed comparison Figure 2 summarizes the trade - off between inference time and image quality for all competing methods on the Gold Atlas dataset. The proposed drifting model achieves one of the lowest inference times (on the order of ms) while simultaneously obtaining the highest SSIM and PSNR, forming the most f av orable position in both “time vs SSIM” and “time vs PSNR” plots. DDPM reaches competitive SSIM/PSNR b ut requires longer sampling, shifting it to ward slower inference, whereas DDIM and F astDDPM reduce runtime at the cost of noticeable drops in SSIM and PSNR. Con ventional generativ e baselines such as V AE and WGAN - GP do not close the quality gap to drifting despite similar or higher computational cost. 4.3 Results on SynthRAD2023 dataset On the more div erse SynthRAD2023 dataset, the proposed drifting model ag ain yields superior quantitative performance, achie ving high SSIM and PSNR with low RMSE. Compared to UNet, V AE, WGAN - GP , and PPFM, drifting substantially improv es structural fidelity and intensity accurac y , and it further impro ves o ver DDIM and FastDDPM despite their strong baseline performance. Qualitativ e examples in Figure 3 sho w that drifting better captures pelvic bone geometry 6 Running T itle for Header 1 0 2 1 0 3 1 0 4 time (ms) 0.82 0.83 0.84 0.85 0.86 0.87 0.88 0.89 SSIM Drif ting DDPM DDIM WGAN- GP V AE UNet PPFM F astDDPM (a) Infer ence time vs SSIM 1 0 2 1 0 3 1 0 4 time (ms) 27 28 29 30 31 PSNR (dB) Drif ting DDPM DDIM WGAN- GP V AE UNet PPFM F astDDPM (b) Infer ence time vs PSNR Figure 2: Comparison of result image quality versus inference time. and soft - tissue contrast under varying anatomies and acquisition conditions, whereas competing methods either ov er-smooth high-contrast regions or introduce spurious texture in lo w-dose areas. Figure 3 also demonstrates that the drifting model maintains robust performance across dif ferent patients, with consistent depiction of sacral curv ature, femoral heads, and rectal filling patterns that closely match the ground - truth CT . Even in challenging cases with atypical positioning or metallic implants, drifting introduces fewer streaks and unnatural intensity transitions than GAN - based and other dif fusion baselines, supporting its generalizability be yond the controlled Gold Atlas cohort. 4.4 In vestigation on model uncertainty Although inference di versity can be adjusted by method-specific sampling parameters such as temperature in (8) for the drifting model or σ t in DDPM-based methods, we believ e that uncertainty in vestigation remains necessary . A controllable div ersity parameter does not by itself explain how sensiti ve a model is to stochastic perturbations under a fixed inference configuration, whereas repeated-sampling analysis directly measures the v ariability of the generated result for the same MRI input. Therefore, in this study , we ev aluate epistemic uncertainty using pixel-wise standard deviation maps obtained from 20 repeated samplings with dif ferent random noises, as shown in Figure 4. Under this unified protocol, the standard deviation map serves as a practical indicator of sampling stability: a lower and more spatially confined variance implies that the model is less sensiti ve to random initialization and thus more reliable at inference time. Figure 4 presents qualitativ e comparisons of model uncertainty for two representativ e cases. Across the compared methods, the standard de viation maps re veal clear differences in the spatial distrib ution. PPFM e xhibits broader highlighted regions with a relatively elev ated global uncertainty level, indicating stronger sensitivity to stochastic sampling, whereas DDPM shows a comparatively more constrained pattern but still retains noticeable uncertainty in sev eral structures. In comparison, the proposed drifting model shows a distinct uncertainty pattern relati ve to the baseline methods. Its standard deviation map highlights fe wer or more localized regions, suggesting impro ved control of sampling variability and more stable inference beha vior under repeated random initialization. 5 Discussion In this work, we systematically ev aluated a conditional drifting model for pelvic MRI-to-CT synthesis on two comple- mentary public datasets, Gold Atlas and SynthRAD2023, and compared it against con v olutional, variational, adversarial, and dif fusion-based baselines. Across both cohorts, the drifting model consistently achie ved competiti ve or superior image quality as measured by SSIM, PSNR, and RMSE, while also delivering fast inference times on the order of milliseconds per slice. These results indicate that drifting-based generativ e modeling provides a favorable balance 7 Running T itle for Header T2 MRI SSIM: 0.91, PSNR : 27.04 UNet SSIM: 0.91, PSNR : 28.57 V AE SSIM: 0.89, PSNR : 25.90 WGAN- GP SSIM: 0.86, PSNR : 26.09 PPFM SSIM: 0.92, PSNR : 29.77 DDIM SSIM: 0.94, PSNR : 30.11 DDPM SSIM: 0.93, PSNR : 30.44 fastDDPM SSIM: 0.94, PSNR : 30.42 Drif ting CT SSIM: 0.92, PSNR : 29.58 SSIM: 0.93, PSNR : 30.53 SSIM: 0.89, PSNR : 28.24 SSIM: 0.85, PSNR : 28.04 SSIM: 0.92, PSNR : 30.57 SSIM: 0.95, PSNR : 32.37 SSIM: 0.93, PSNR : 31.16 SSIM: 0.94, PSNR : 31.32 Figure 3: Comparison of CT synthesize results from different methods on SynthRAD2023 Dataset. CT DDPM DDIM PPFM fastDDPM Drif ting Figure 4: Pixel-vise standard de viation map based on 20 sampling results for each comparing approach. between fidelity and computational efficienc y , which is particularly relev ant for time-sensitiv e radiotherapy workflows where multiple synthetic CT volumes may need to be generated repeatedly o ver a treatment course. An important observation from our experiments is that the drifting model preserves pelvic bone geometry and soft- tissue boundaries more f aithfully than CNN, V AE, and GAN baselines, which either tended to ward ov er-smoothing or introduced artifacts in high-contrast regions. On Gold Atlas, qualitativ e inspection showed sharper cortical bone edges, clearer rectal wall delineation, and fewer checkerboard or noise-like patterns than WGAN-GP and PPFM, aligning with the superior SSIM/PSNR and lo wer RMSE reported for the drifting model. On the more heterogeneous SynthRAD2023 pelvis subset, the drifting model remained rob ust across patients and acquisition protocols, maintaining accurate depiction of sacrum curv ature, femoral heads, and rectal filling patterns ev en in challenging cases with atypical positioning or metallic implants. T ogether , these findings suggest that the attraction–repulsion mechanism in drifting 8 Running T itle for Header models can improve structural consistenc y at anatomy-critical interfaces compared with purely vox el-wise or adversarial objectiv es. Compared with diffusion-based methods, the proposed approach occupies an interesting point in the design space. Prior diffusion models for MRI-to-CT synthesis have demonstrated high fidelity and strong dosimetric agreement, but at the cost of iterative sampling that can require seconds to minutes per volume. In our study , DDPM achiev ed strong SSIM and PSNR but incurred substantially higher inference time, whereas accelerated variants such as DDIM and FastDDPM narrowed the runtime gap at the expense of noticeable drops in image quality . By contrast, the drifting model internalizes the transport dynamics into training, allowing one-step inference conditioned on MRI while still matching or surpassing dif fusion baselines in both similarity metrics and qualitative sharpness. This accuracy–ef ficiency trade-off suggests that drifting models may serve as a pragmatic alternativ e when clinical throughput is a primary constraint, for example in MRI-only planning or adaptiv e radiotherapy scenarios where same-session sCT generation is desirable. Our results should also be interpreted in the context of broader w ork on MRI-to-CT synthesis and generative medical imaging. T raditional atlas-based and registration-dri ven methods encode anatomical priors but can be brittle to registration errors and population mismatch, particularly in multi-center settings such as SynthRAD2023. V AE- and GAN-based framew orks have improv ed realism and flexibility b ut often require sophisticated structural constraints and careful training to a void blurring or hallucina ted structures. Recent diffusion approaches mitigate some GAN failure modes and provide strong baselines, b ut practicality can be an issue for integration into routine clinical pipelines. The drifting frame work, operating directly in image space with a kernel-based attraction–repulsion field, of fers an alternati ve that is simple to condition on MRI, yields deterministic one-step mapping at test time, and empirically reduces the tendency to ward over -smoothing or texture artifacts around bone–air–soft tissue interfaces. These characteristics make drifting models a promising candidate to complement, rather than replace, e xisting generative paradigms in synthetic CT research. Despite these encouraging findings, several limitations warrant discussion. First, our experiments are restricted to pelvic MRI-to-CT synthesis and rely solely on image similarity metrics (SSIM, PSNR, RMSE) and qualitati ve anatomical assessment, without explicit ev aluation on downstream clinical tasks such as dose calculation, DVH agreement, or attenuation correction performance. Prior work has emphasized that task-based endpoints are critical for judging clinical readiness; thus, future studies should integrate radiotherapy planning and PET/MR attenuation experiments to quantify the dosimetric and functional impact of drifting-based sCT . Second, we focused on paired supervised training and did not explore unpaired or semi-supervised regimes, which are highly rele vant gi ven the scarcity of well-registered MRI–CT pairs and the availability of large unpaired datasets. Extending drifting models to unpaired translation, potentially by combining them with cycle or structural consistency constraints, could further broaden their applicability . Third, our implementation uses a relatively standard UNet-like backbone and a simple image-domain drifting field with a fixed Gaussian kernel and globally chosen loss weights. It is plausible that more expressi ve architectures (e.g., transformer or hybrid CNN–transformer backbones) or adapti ve, spatially varying k ernels could further enhance performance, particularly in regions with complex anatomy or lar ge intensity heterogeneity . Additionally , we did not perform an extensi ve hyperparameter search across all baselines; while we attempted to match model capacity and training protocols, some performance dif ferences may partly reflect implementation details or optimization choices. Finally , all experiments were conducted on a single high-end GPU, and we did not systematically analyze memory consumption, scalability to 3D or 4D (time-resolved) volumes, or behavior under domain shifts such as different scanners, sequences, or institutions beyond those included in the two public datasets. Future work will therefore include se veral directions. From a methodological perspectiv e, incorporating anatomy-aware or frequency-decoupled losses into the drifting field, as well as e xploring multi-modal conditioning (e.g., joint T1/T2, or MRI plus prior se gmentations), may improve fine-detail reconstruction and robustness across varying acquisition protocols. From an e valuation standpoint, we plan to conduct comprehensi ve task-based studies in radiotherapy planning and PET/MR attenuation, including comparisons of dose metrics, gamma analysis, and clinical acceptability ratings between drifting-based sCT and ground-truth CT . Finally , extending the proposed frame work beyond the pelvis to other anatomies such as brain, head-and-neck, and thorax, and assessing its performance under domain adaptation scenarios, will be essential to understand how well drifting models generalize in realistic multi-center clinical settings. 6 Conclusion W e presented a conditional drifting model for pelvic MRI-to-CT synthesis and benchmarked it against con volutional, variational, adversarial, and diffusion-based approaches on the Gold Atlas Male Pelvis and SynthRAD2023 pelvis datasets. Across both cohorts, the drifting model achie ved consistently high SSIM and PSNR together with low RMSE, while preserving fine pelvic bone structures and soft-tissue boundaries with fewer artifacts than competing 9 Running T itle for Header methods. Compared with diffusion-based baselines, the proposed approach offers a more fa vorable accuracy–ef ficiency trade-off, attaining comparable or superior image quality with substantially reduced inference time through one-step generation. These properties make drifting models attractiv e for integration into time-critical MR-only workflows, including radiotherapy planning and attenuation correction, where reliable and fast synthetic CT generation is required. Future work will e xtend this framework to additional anatomies and task-based ev aluations, and in vestigate unpaired training and architecture refinements to further enhance robustness and clinical readiness. References [1] Norbert J Pelc. Recent and future directions in ct imaging. Annals of biomedical engineering , 42(2):260–268, 2014. [2] Carlo Liguori, Giulia Frauenfelder , Carlo Massaroni, Paola Saccomandi, Francesco Giurazza, Francesca Pitocco, Riccardo Marano, and Emiliano Schena. Emerging clinical applications of computed tomography . Medical Devices: Evidence and Researc h , pages 265–278, 2015. [3] David J Brenner and Eric J Hall. Computed tomography—an increasing source of radiation exposure. New England journal of medicine , 357(22):2277–2284, 2007. [4] Guy Frija, Ivana Blaži ´ c, Donald P Frush, Monika Hierath, Michael Kawooya, Lluis Donoso-Bach, and Boris Brklja ˇ ci ´ c. Ho w to improve access to medical imaging in lo w-and middle-income countries? EClinicalMedicine , 38, 2021. [5] Benjamin T Burdorf. Comparing magnetic resonance imaging and computed tomography machine accessibility among urban and rural county hospitals. J ournal of Public Health Researc h , 11(1):jphr–2021, 2022. [6] Zakaria Shams Siam, Md Y ounus Akon, Israt Jahan Munmun, Abdullah Al-Amin, Md Abdus Salam, and Ishtiak Al Mamoon. A paired ct and mri dataset for adv anced medical imaging applications. Data in Brief , 61:111768, 2025. [7] Jens M Edmund and T ufve Nyholm. A revie w of substitute ct generation for mri-only radiation therapy . Radiation Oncology , 12(1):28, 2017. [8] Mohamed A Bahloul, Saima Jabeen, Sara Benoumhani, Habib Abdulmohsen Alsaleh, Zehor Belkhatir , and Areej Al-W abil. Advancements in synthetic ct generation from mri: A re view of techniques, and trends in radiation therapy planning. Journal of Applied Clinical Medical Physics , 25(11):e14499, 2024. [9] Sanuwani Dayarathna, Kh T ohidul Islam, Sergio Uribe, Guang Y ang, Munaw ar Hayat, and Zhaolin Chen. Deep learning based synthesis of mri, ct and pet: Revie w and analysis. Medical imag e analysis , 92:103046, 2024. [10] Mahmoud Ibrahim, Y asmina Al Khalil, Sina Amirrajab, Chang Sun, Marcel Breeuwer , Josien Pluim, Bart Elen, Gökhan Ertaylan, and Michel Dumontier . Generati ve ai for synthetic data across multiple medical modalities: A systematic revie w of recent developments and challenges. Computers in biology and medicine , 189:109834, 2025. [11] F Guerreiro, Ninon Burgos, A Dunlop, K W ong, I Petkar , C Nutting, K Harrington, S Bhide, K Ne wbold, D Dearnaley , et al. Ev aluation of a multi-atlas ct synthesis approach for mri-only radiotherapy treatment planning. Physica Medica , 35:7–17, 2017. [12] Jinsoo Uh, Thomas E Merchant, Y imei Li, Xingyu Li, and Chiaho Hua. Mri-based treatment planning with pseudo ct generated through atlas registration. Medical physics , 41(5):051711, 2014. [13] Ninon Burgos, M Jorge Cardoso, Marc Modat, Shonit Punwani, David Atkinson, Simon R Arridge, Brian F Hutton, and Sébastien Ourselin. Ct synthesis in the head & neck region for pet/mr attenuation correction: an iterativ e multi-atlas approach. EJNMMI physics , 2(Suppl 1):A31, 2015. [14] Y asheng Chen and Hongyu An. Attenuation correction of pet/mr imaging. Magnetic Resonance Imaging Clinics , 25(2):245–255, 2017. [15] Snehashis Roy , Aaron Carass, Amod Jog, Jerry L Prince, and Junghoon Lee. Mr to ct registration of brains using image synthesis. In Pr oceedings of SPIE , v olume 9034, pages spie–org, 2014. [16] Junghoon Lee, Aaron Carass, Amod Jog, Can Zhao, and Jerry L Prince. Multi-atlas-based ct synthesis from con ventional mri with patch-based refinement for mri-based radiotherapy planning. In Medical Imaging 2017: Image Pr ocessing , v olume 10133, pages 434–439. SPIE, 2017. [17] Krithika Iyer , Austin T app, Athelia P aulli, Gabrielle Dickerson, Syed Muhammad Anwar , Natasha Lepore, and Marius George Linguraru. Mri-to-ct synthesis with cranial suture segmentations using a variational autoencoder framew ork. arXiv pr eprint arXiv:2512.23894 , 2025. 10 Running T itle for Header [18] Y unxiang Li, Hua-Chieh Shao, Xiaoxue Qian, and Y ou Zhang. Fddm: unsupervised medical image translation with a frequency-decoupled dif fusion model. Mac hine Learning: Science and T ec hnology , 6(2):025007, apr 2025. [19] Y oussef Skandarani, Pierre-Marc Jodoin, and Alain Lalande. Gans for medical image synthesis: An empirical study . Journal of Imaging , 9(3):69, 2023. [20] Y ang Lei, Joseph Harms, T onghe W ang, Y ingzi Liu, Hui-Kuo Shu, Ashesh B Jani, W alter J Curran, Hui Mao, T ian Liu, and Xiaofeng Y ang. Mri-only based synthetic ct generation using dense cycle consistent generative adversarial networks. Medical physics , 46(8):3565–3581, 2019. [21] Chun-Chieh W ang, Pei-Huan Wu, Gigin Lin, Y en-Ling Huang, Y u-Chun Lin, Y i-Peng Chang, and Jun-Cheng W eng. Magnetic resonance-based synthetic computed tomography using generativ e adversarial networks for intracranial tumor radiotherapy treatment planning. Journal of personalized medicine , 12(3):361, 2022. [22] Shu-Hui Hsu, Zhaohui Han, Jonathan E Leeman, Y ue-Houng Hu, Raymond H Mak, and Atchar Sudhyadhom. Synthetic ct generation for mri-guided adaptiv e radiotherapy in prostate cancer . F rontier s in Oncology , 12:969463, 2022. [23] Olga M Dona Lemus, Y i-Fang W ang, Fiona Li, Sachin Jambawalikar , David P Horowitz, Y uanguang Xu, and Cheng-Shie W uu. Dosimetric assessment of patient dose calculation on a deep learning-based synthesized computed tomography image for adapti ve radiotherapy . Journal of Applied Clinical Medical Physics , 23(7):e13595, 2022. [24] Heran Y ang, Jian Sun, Aaron Carass, Can Zhao, Junghoon Lee, Jerry L Prince, and Zongben Xu. Unsupervised mr- to-ct synthesis using structure-constrained cycleg an. IEEE tr ansactions on medical imaging , 39(12):4249–4261, 2020. [25] Hidetoshi Matsuo, Mizuho Nishio, Munenobu Nogami, Feibi Zeng, T akako Kurimoto, Sandeep Kaushik, Florian W iesinger, Atsushi K Kono, and T akamichi Murakami. Unsupervised-learning-based method for chest mri– ct transformation using structure constrained unsupervised generativ e attention networks. Scientific reports , 12(1):11090, 2022. [26] Changfei Gong, Y uling Huang, Mingming Luo, Shunxiang Cao, Xiaochang Gong, Shenggou Ding, Xingxing Y uan, W enheng Zheng, and Y un Zhang. Channel-wise attention enhanced and structural similarity constrained cycle gan for effecti ve synthetic ct generation from head and neck mri images. Radiation Oncology , 19(1):37, 2024. [27] Y anxia Liu, Anni Chen, Hongyu Shi, Sijuan Huang, W anjia Zheng, Zhiqiang Liu, Qin Zhang, and Xin Y ang. Ct synthesis from mri using multi-c ycle gan for head-and-neck radiation therapy . Computerized medical ima ging and graphics , 91:101953, 2021. [28] Shaoyan Pan, Elham Abouei, Jacob W ynne, Chih-W ei Chang, T onghe W ang, Richard LJ Qiu, Y uheng Li, Junbo Peng, Justin Roper , Pretesh Patel, et al. Synthetic ct generation from mri using 3d transformer-based denoising diffusion model. Medical Physics , 51(4):2538–2548, 2024. [29] Muzaffer Özbe y , Onat Dalmaz, Salman UH Dar, Hasan A Bedel, ¸ Saban Özturk, Alper Güngör , and T olga Cukur . Unsupervised medical image translation with adversarial diffusion models. IEEE T ransactions on Medical Imaging , 42(12):3524–3539, 2023. [30] Qing L yu and Ge W ang. Con version between ct and mri images using dif fusion and score-matching models. arXiv pr eprint arXiv:2209.12104 , 2022. [31] Mingyang Deng, He Li, T ianhong Li, Y ilun Du, and Kaiming He. Generativ e modeling via drifting. arXiv pr eprint arXiv:2602.04770 , 2026. [32] T ufve Nyholm, Stina Svensson, Sebastian Andersson, Joakim Jonsson, Maja Sohlin, Christian Gustafsson, Elisabeth Kjellén, Ludvig P . Muren, Hans von der Maase, Jinyi W ang, Sofie Ceberg, and Adalsteinn Gunnlaugsson. Mr and ct data with multiobserv er delineations of or gans in the pelvic area – part of the gold atlas project. Medical Physics , 45(3):1295–1300, 2018. [33] Adrian Thummerer , Erik van der Bijl, Arthur Galapon Jr , Joost J. C. V erhoeff, Johannes A. Langendijk, Stefan Both, Cornelis A. T . van den Berg, and Matteo Maspero. Synthrad2023 grand challenge dataset: Generating synthetic ct for radiotherapy . Medical Physics , 50(7):4664–4674, 2023. [34] Olaf Ronneberger, Philipp Fischer, and Thomas Brox. U-Net: Con volutional Netw orks for Biomedical Image Segmentation. arXiv e-prints , page arXiv:1505.04597, May 2015. [35] Diederik P Kingma and Max W elling. Auto-Encoding V ariational Bayes. arXiv e-prints , page December 2013. 11 Running T itle for Header [36] Ishaan Gulrajani, Faruk Ahmed, Martin Arjovsk y , V incent Dumoulin, and Aaron C Courville. Improv ed training of wasserstein gans. In Advances in neural information pr ocessing systems , volume 30, 2017. [37] Dennis Hein, Staff an Holmin, T imothy Szczykuto wicz, Jonathan S. Maltz, Mats Danielsson, Ge W ang, and Mats Persson. PPFM: Image Denoising in Photon-Counting CT Using Single-Step Posterior Sampling Poisson Flo w Generativ e Models. IEEE T ransactions on Radiation and Plasma Medical Sciences , 8(7):788–799, January 2024. [38] Hongxu Jiang, Muhammad Imran, T eng Zhang, Y uyin Zhou, Muxuan Liang, Kuang Gong, and W ei Shao. F ast- DDPM: Fast Denoising Dif fusion Probabilistic Models for Medical Image-to-Image Generation. arXiv e-prints , page arXiv:2405.14802, May 2024. [39] Jiaming Song, Chenlin Meng, and Stefano Ermon. Denoising Dif fusion Implicit Models. arXiv e-prints , page arXiv:2010.02502, October 2020. [40] Jonathan Ho, Ajay Jain, and Pieter Abbeel. Denoising Diffusion Probabilistic Models. arXiv e-prints , page arXiv:2006.11239, June 2020. 12
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment