Building evidence-based knowledge graphs from full-text literature for disease-specific biomedical reasoning
Biomedical knowledge resources often either preserve evidence as unstructured text or compress it into flat triples that omit study design, provenance, and quantitative support. Here we present EvidenceNet, a framework and dataset for building diseas…
Authors: Chang Zong, Sicheng Lv, Si-tu Xue
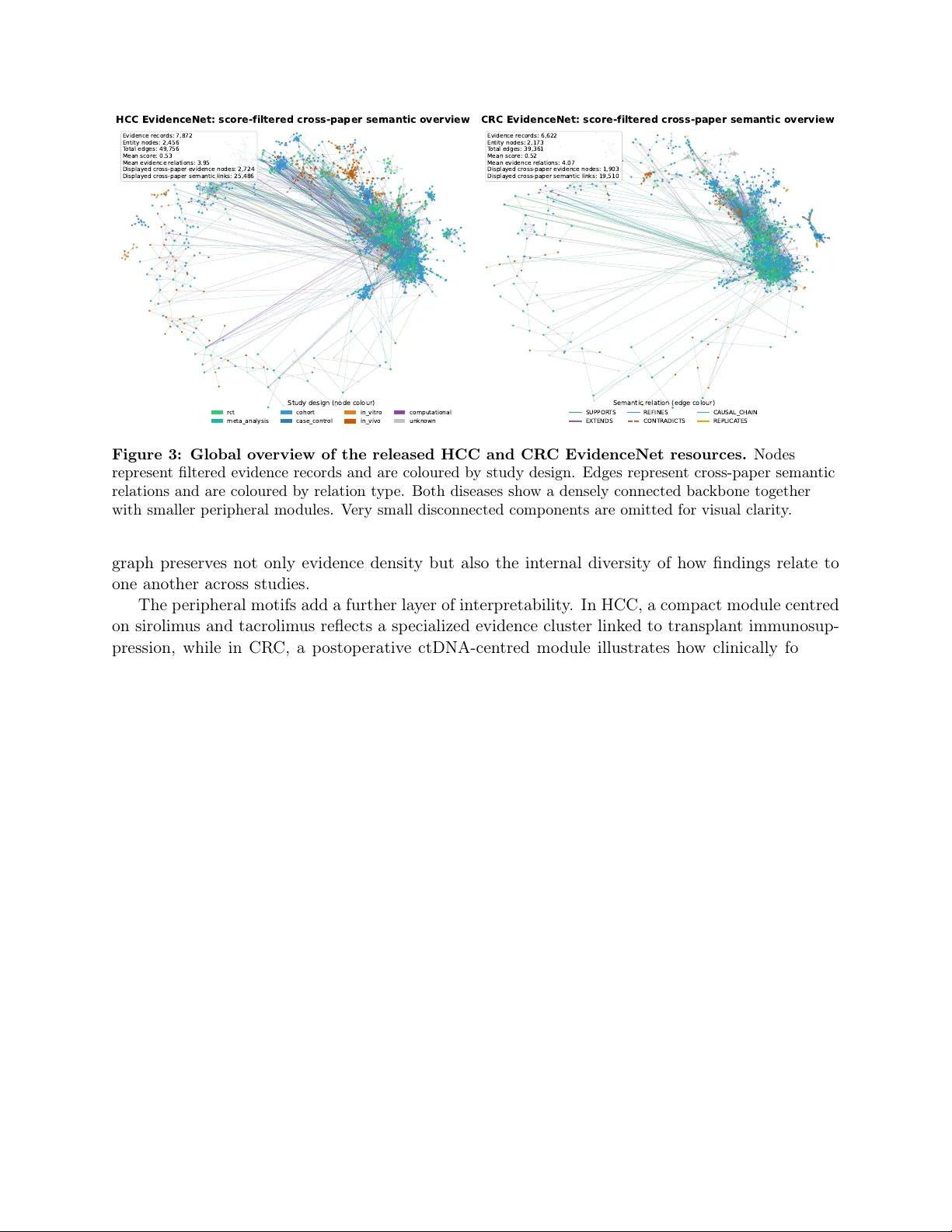
Building evidence-based kno wledge graphs from full-text literature for disease-sp ecific biomedical reasoning Chang Zong 1 , ∗ , Sic heng Lv 1 , Si-tu Xue 2 , Huilin Zheng 1 , Jian W an 3 , Lei Zhang 1 1 Sc ho ol of Computer Science and T echnology , Zhejiang Univ ersity of Science and T echnology , Hangzhou, China 2 Institute of Medicinal Biotec hnology , Chinese A cademy of Medical Sciences & P eking Union Medical College, Beijing, China 3 Zhejiang Key Lab oratory of Biomedical In telligen t Computing T ec hnology , Hangzhou, China *Corresp ondence: zongc hang@zust.edu.cn Abstract Biomedical kno wledge resources often either preserv e evidence as unstructured text or com- press it into flat triples that omit study design, prov enance, and quantitativ e supp ort. Here w e presen t EvidenceNet , a framew ork and dataset for building disease-sp ecific knowledge graphs from full-text biomedical literature. EvidenceNet uses a large language mo del (LLM)-assisted pip eline to extract exp erimen tally grounded findings as structured evidence no des, normalize biomedical entities, score evidence qualit y , and connect evidence records through typed seman- tic relations. W e release tw o resources: EvidenceNet-HCC with 7,872 evidence records, 10,328 graph no des, and 49,756 edges, and EvidenceNet-CR C with 6,622 records, 8,795 nodes, and 39,361 edges. T ec hnical v alidation sho ws high comp onen t fidelity , including 98.3% field-lev el ex- traction accuracy , 100.0% high-confidence en tity-link accuracy , 87.5% fusion integrit y , and 90.0% seman tic relation-type accuracy . In downstream ev aluation, EvidenceNet improv es in ternal and external retriev al-augmented question answ ering and retains structural signal for future link prediction and target prioritization. These results establish EvidenceNet as a disease-sp ecific resource for evidence-a ware biomedical reasoning and hypothesis generation. Keyw ords: evidence-based knowledge graph, large language mo dels, biomedical literature mining, disease-sp ecific reasoning 1 Bac kground & Summary The realization of precision medicine requires more than access to biomedical literature. It also requires w ays to organize that literature in to computable evidence [1–3]. Scien tific output contin ues to grow rapidly , but the ability to synthesize findings across pap ers remains limited [4–6]. Muc h of the relev an t kno wledge remains embedded in free text and is therefore difficult for computational systems to query , compare, and reuse [7, 8]. Curren t efforts to structure biomedical knowledge mainly rely on general-purp ose knowledge graphs (KGs) such as PrimeK G [9], Hetionet [10], and T arKG [11]. These resources aggregate millions of facts in to compact triples. This represen tation is p o werful, but it omits m uch of the con- text that determines how a finding should b e interpreted. In evidence-based medicine, a biological asso ciation is rarely absolute. Its meaning depends on the study p opulation, interv ention details, disease stage, and exp erimen tal design [12–14]. A treatmen t may show b enefit in vitro y et fail in a 1 phase I I trial [15, 16]. A pathw a y may promote tumour progression in one setting but suppress it in another [17, 18]. Flattening these findings into simple edges remov es the PICO (Population, In terven tion, Com- parison, Outcome) structure that supp orts assessmen t of relev ance and reliabilit y [13, 19, 20]. This loss of con text limits the v alue of conv en tional K Gs for tasks suc h as clinical decision supp ort [21, 22] and mechanism-guided drug discov ery [23–25], where prov enance and evidential strength matter as m uch as the claim itself. General KGs also mix many diseases in to a shared top ology . That breadth is useful, but it can blur the disease-sp ecific structure needed to study conditions such as hepato- cellular carcinoma (HCC) and colorectal cancer (CRC) [26–30]. Large language mo dels (LLMs) pro vide a practical w ay to reco ver this missing structure from full-text articles [31–34]. Compared with earlier NLP systems that fo cus mainly on named-entit y recognition [35, 36], mo dern LLMs can interpret longer scientific arguments and extract ric her ex- p erimen tal con text [37–39]. Recent work shows strong p erformance in zero-shot or weakly sup er- vised information extraction, esp ecially when schemas are complex and relations are con text de- p enden t [40–43]. LLMs are also increasingly used for on tology alignment and retriev al-augmen ted biomedical reasoning [38, 44–49]. Against this background, w e present EvidenceNet , a framework and dataset for constructing disease-sp ecific, evidence-centric knowledge graphs from biomedical literature. EvidenceNet treats the exp erimen tally grounded finding, rather than the entit y alone, as the primary graph unit. This design preserves prov enance, study context, and quantitativ e supp ort suc h as p -v alues or hazard ratios. It also keeps the graph fo cused on a sp ecific disease landscape, making lo cal top ology more in terpretable for downstream reasoning. EvidenceNet uses an automated pip eline to transform op en-access articles in to computable ev- idence. In this release, the corpus is restricted to disease-sp ecific, recen t PubMed-indexed arti- cles with accessible full text so that eac h extracted record can b e grounded in article-level context rather than abstract-only statemen ts. F ull-text pap ers are segmen ted into candidate evidence spans, con verted into structured PICO-style records, normalized against biomedical v o cabularies, scored for evidence qualit y , and integrated into a seman tic graph. The resulting resource supp orts both evidence-orien ted interpretation and graph-based analysis. T o clearly p osition EvidenceNet within the current landscap e, T able 1 contrasts our approac h with general biomedical KGs and raw literature databases across key dimensions. F eature General KGs Literature Databases EvidenceNet (Ours) Scop e Univ ersal (All diseases) Univ ersal Disease-Sp ecific (e.g., HCC, CRC) Data Unit Static T riple (Sub ject-Predicate-Ob ject) Do cumen t / Abstract Evidence No de (PICO Context) Con textual Depth Lo w (Relation type only) High (Unstructured text) High & Structured (Study design, p -v alues, cohorts) Reasoning Mo de Structural (P ath-finding, Embedding) Seman tic (Keyw ord/V ector searc h) Dual (Structural + Deep Semantic) T able 1: Comparison of EvidenceNet with existing biomedical kno wledge resources. EvidenceNet pro vides curated disease-sp ecific resources for HCC and CRC. EvidenceNet-HCC con tains 7,872 evidence records and a graph with 10,328 no des and 49,756 edges. EvidenceNet- CR C con tains 6,622 evidence records and a graph with 8,795 no des and 39,361 edges. Unlik e 2 traditional K Gs, these resources preserv e study-lev el seman tics and supp ort b oth semantic and structural reasoning. W e v alidated EvidenceNet across four complemen tary dimensions: • Comp onen t fidelity . T argeted audits show high reliabilit y for evidence extraction, entit y normalization, evidence fusion, and semantic relation typing. • Con text-aw are biomedical QA. By leveraging the PICO-ric h structure of evidence no des, EvidenceNet improv es internal and external retriev al-augmen ted question answ ering o v er base- line retriev al settings. • Scien tific discov ery link prediction. Multi-metho d graph exp erimen ts show robust reco v- ery of future entit y links, supp orting the biological v alidit y of the evidence-centric top ology . • Prosp ectiv e therap eutic target disco very . In a time-sliced scenario, EvidenceNet priori- tizes emerging HCC and CR C targets that the general T arKG misses, demonstrating practical utilit y for hypothesis generation. These results show that EvidenceNet serv es as a disease-sp ecific resource for biomedical reasoning and disco very , rather than only as a data rep ository . 2 Metho ds 2.1 Ov erview of the EvidenceNet w orkflo w EvidenceNet is a set of disease-sp ecific, evidence-centric biomedical knowledge graphs constructed from full-text literature. The w orkflow comprises four stages, namely Data Pr epr o c essing , LLM- Driven Evidenc e Extr action , Normalization and Sc oring , and Inte gr ation and Gr aph Construction (Fig. 1). The central design principle is to represen t each exp erimentally grounded finding as an explicit evidence no de , rather than collapsing the literature into direct sub ject–predicate–ob ject triples. This representation preserves prov enance, study context, and quan titative supp ort, making it w ell suited to downstream evidence-based biomedical reasoning [20, 50]. Giv en a disease-sp ecific corpus D = { d 1 , . . . , d N } , each article d i is con verted into a set of structured evidence ob jects E i = { e i 1 , . . . , e im } . These ob jects are in tegrated in to the directed graph defined in equation (1). G = ( V E ∪ V T , R ) , (1) where V E denotes evidence no des, V T denotes normalized biomedical entit y nodes, and R denotes evidence–en tity and evidence–evidence relations. The resulting resource is designed to preserv e b oth the depth of individual studies and the top ology required for graph-based analysis. 3 Phase 1: Data Preprocessing PDF File Metadata Fetcher Pap er Title CrossRef OpenAlex DOI IF Date … PDF Par se r Heuristic Filt ering (Remove Irrelev ant Secti ons) Chunking (Window Size + Overlap) Phase 2: LLM - Driven Evidence Ex traction List of Chunks with Metadata High - Per f or ma nc e LL M Evidence Extract or Te x t C h u n k Evidence Data Model Few - Shot Prompting Extracted V alues (PICO Element s, Named Entities, Study Design, Stati stics …) Raw Objects Intra - Document Agg regation Deduplication (Remove Exact the Same Evidence) Evidence Object Phase 3: Normalization & Scoring Entity Norma lizer Syn ony m Mapping ( via Dictionary) Sym bol St an dar di za ti on ( via Suffix Stripping) Fuzzy Matching (via Similarity Threshold) Evidence Scor er Fac to rs (Impact Factor, Stud y Design, Statistics Significance, Sample Size, LLM Confidence) Composite Confidence Score (via Weighted Sum) Scored Evidence Phase 4: Integration & Graph Construc tion Global Dedupli cator Existing Graph Evidence Connect or Fingerprint Matching Semantic Deduplicating Attri bute Merging Rel ati ons hip Description (Support, Contradict, Extend, …) LLM - base d Detection Evidence Filtering ( Shared Entities, Semantic Similarity ) Rule - based Detection Evidence Graph Manual Audit LLM Check Linked Evidence Figure 1: W orkflow of EvidenceNet for constructing an evidence knowledge graph. 1. Literature Example Titl e: “A Comprehensive Review of Sequencing and Combination Strategies of T ar geted Agents in Metas -tatic Colorectal Cancer ” Journal: The Oncologist Ye a r: 2018 DOI: 10.1634/theoncologist.2017 - 0203 “In FIRE- 3 (N5592),patients were randomized 1:1 to cetuximab plus FOLFIRI versus bevacizumab plus FOLFIRI, and objective response rate (O RR ) was assessed as the primary endpoint. Although no statistic al differe nces in ORR (62.0% v s. 58.0%; p = .18) or median progression - free survival (PFS; 10.0 vs.10.3 months; hazard ratio [HR] =1.06 [95% con fidence interval confidence interval (Cl), 0.88 - 1.26]; p = .55) were observed, median OS favored the cetux imab arm (28.7 vs. 25.0 months; HR = 0.77 [ 95% Cl, 0.62 - 0.96]; p = . 017)." Evidence node evidence_id: EVID - 266DDD68D593 target_disease: CRC study_desig n: rct study_obj ect: patients with RAS-w ildtype metastatic colorectal cancer intervention: cetuxima b plus FOLFIRI comparison: bevaci zumab plus FOLFIRI phenotype: median overall survival favored the t t cetuximab arm despite no statistical d differences in ORR or PFS bio_mechanis m: differential benef it of anti - EGFR therapy by tumor sidedness Key statistics: ORR: 62.0% vs 58.0%, p = .18 PFS: 10.0 vs 10.3 months, HR = 1.06, 95% CI 0.88 - 1.26, p = .55 OS: 28.7 vs 25.0 months, HR = 0.77, 95% CI 0.62 - 0.96, p = .017 composite_sco re: 0.6275 grade_lev el: B llm_confidence : 0.86 Cetuximab → TC546137 Irinotecan → TC777499 5- fluoropyrimidine → TC8191 42 RAS → P08647 "Clinical management of metastatic colorectal cancer in the era of precision medicine" (2022) EVID - 266DDD68D593 Cetuximab (Drug) 5-fluoropyrimidine (Drug) Irinotecan (Drug) RAS ( (Gene) "Exploring and bypassing resistance to targeted therapies in colorectal cancer" (2025) Evidence: “Monoclonal antibod ies for targeted th erapy in colorectal cancer.” (2010) LINKED_TO LINKED_TO LINKED_TO LINKED_TO SUPPORTS REFINES EXTENDS 2. Structured Evidence 3. Graph & Schema Evidence-entity: LINKED_TO Evidence-evidence: SUPPORTS, CONTRADICTS, REFINES, EXTENDS, REPLICATES, CAUSAL_CHAI N Figure 2: Compact ov erview of the EvidenceNet transformation pro cess and graph sc hema. F rom left to right, the figure traces a colorectal cancer example based on the FIRE-3 comparison of cetuximab plus F OLFIRI versus b ev acizumab plus FOLFIRI from source text to a structured evidence record and then to its graph represen tation. The schema highlights the principal evidence attributes retained during extraction and the t wo relation families used during graph integration, namely evidence–en tity LINKED_TO edges and typed evidence–evidence semantic relations. Fig. 2 complemen ts the stage-lev el workflo w in Fig. 1 by illustrating how a comparative treatmen t statemen t is conv erted into a graph-native evidence ob ject. The example is drawn from a colorectal cancer rep ort summarizing the FIRE-3 trial, in whic h cetuximab plus FOLFIRI is compared with b ev acizumab plus F OLFIRI. The extracted record retains pro venance, trial context, response and surviv al outcomes, and quality metadata. In the graph lay er, the corresp onding evidence no de 4 links to normalized en tities such as cetuximab, irinotecan, fluorop yrimidine, and RAS through LINKED_TO edges, while typed evidence–evidence relations, including SUPPOR TS, EXTENDS, and REFINES, p osition the finding within the broader b o dy of related evidence. This compact view clarifies ho w quantitativ e literature statements are carried into an interpretable graph structure. 2.2 Data prepro cessing The first stage standardizes full-text PDF articles for evidence mining. The current release op erates on disease-sp ecific corp ora assem bled from recen t PubMed-indexed articles with av ailable full text. Eac h article is asso ciated with bibliographic metadata, including DOI, title, journal, year, citation coun t, and journal-lev el impact indicators. Missing metadata are supplemented from public biblio- graphic resources. DOI resolution is p erformed first, follow ed by title-based reco very when needed. Automatically retriev ed metadata are merged with man ually curated v alues, with man ually supplied en tries treated as the preferred source. The full text is then parsed and segmen ted in to canonical scientific sections. References and other lo w-information sections are excluded from do wnstream evidence extraction. This section- a ware design reflects the structure of biomedical articles, in which exp erimen tally relev ant conten t is concen trated in a limited p ortion of the full text. Eac h article is further divided in to o verlapping text c hunks. F or a do cumen t d , this stage pro duces the ordered set shown in equation (2). C ( d ) = { c 1 , c 2 , . . . , c M } , (2) where each ch unk c j retains its section lab el. Overlap reduces b oundary effects and helps preserve exp erimen ts describ ed across adjacen t spans. This prepro cessing step increases evidence density in the input represen tation and improv es downstream language-mo del efficiency . 2.3 LLM-driv en evidence extraction In the second stage, eac h retained text c hunk is con verted in to structured candidate evidence using a LLM. Chunks unlikely to contain exp erimen tal findings are first filtered out. The remaining ch unks are pro cessed indep enden tly , enabling scalable extraction across large disease-sp ecific corp ora. Extraction follo ws a structured sc hema based on the PICO framew ork. The sc hema captures study ob ject, interv en tion, comparison, outcomes, biological mechanism, phenotype, study design, clinical stage, and quantitativ e attributes such as p -v alues, sample size, and fold change [19]. The LLM is constrained to extract only explicitly stated findings, separate distinct exp eriments in to atomic evidence units, and retain grounded supp orting text. Background statements and descrip- tions of prior w ork are excluded. This design reduces unsupp orted inference and improv es extraction fidelit y [34]. F or each ch unk c j , the extraction mo del f θ pro duces a set of candidate evidence ob jects as in equation (3). ˆ E j = f θ ( c j ) . (3) Ch unk-level outputs are then aggregated at the do cumen t level according to equation (4). ˆ E ( d ) = Agg M [ j =1 ˆ E j . (4) Aggregation remo ves exact duplicates and fills missing attributes when the same exp erimen t is describ ed in more than one section. Distinct exp eriments are retained as separate evidence units. 5 Eac h aggregated ob ject is represen ted as a structured evidence no de. The no de stores experi- men tal context, core biomedical en tities, quan titative results, source text, and extraction confidence. This formulation preserves the study-level detail needed for evidence-oriented analysis, esp ecially when similar claims differ by mo del system, disease stage, or therap eutic condition [17]. 2.4 Normalization and scoring The third stage conv erts extracted evidence in to standardized and comparable data ob jects. This stage includes biomedical entit y normalization and quality scoring. Biomedical en tities are normalized to a standardized external resource. In this release, normal- ization is aligned to T arK G [11]. Matching supp orts exact matching, synonym mapping, sym b ol standardization for genes or proteins, and fuzzy matching based on seman tic similarity . This step impro ves consistency across articles and increases inter op erabilit y with external biomedical graph resources. Eac h evidence no de is then assigned a comp osite quality score inspired by evidence-based as- sessmen t frameworks. The score com bines study design, source impact, statistical supp ort, sample size, and extraction confidence as shown in equation (5). S ( e ) = ( w 1 S type + w 2 S impact + w 3 S stat + w 4 S sample ) (1 − λ ) + λC LLM , (5) where the sample-size term is log-normalized. Con tinuous scores are additionally mapp ed to four evidence lev els (A-D) to supp ort filtering and downstream ranking. 2.5 In tegration and graph construction In the final stage, normalized evidence no des are integrated in to the p ersisten t EvidenceNet resource. This stage includes cross-do cumen t duplicate resolution, evidence–evidence relation induction, au- tomated v erification, and graph serialization. Newly extracted evidence is first compared with the existing graph to identify near-duplicate findings through fingerprin t matchin g and semantic comparison across do cumen ts. When highly o verlapping evidence is detected, the canonical no de is retained and up dated with merged pro ve- nance. This prev ents rep eated findings from inflating the graph while preserving data lineage. Evidence–evidence relations are then inferred. Candidate pairs are identified using semantic similarit y and o verlap in biomedical entities. F or related pairs, EvidenceNet assigns one of several directed relation types, including SUPPOR TS, CONTRADICTS, REFINES, EXTENDS, REPLI- CA TES, and CAUSAL_CHAIN. This la yer captures isolated findings, consistency , refinement, re- pro duction, and mechanistic ordering. Relation assignmen t uses a h ybrid strategy . Deterministic heuristics pro vide an initial prop osal based on seman tic direction, chronology , and biological ov er- lap. An LLM-based v erification step is then applied to am biguous or high-similarity pairs. This com bination balances interpretabilit y and flexibility , and improv es the reliability of relation lab els in complex biomedical contexts [31, 39]. The final resource is stored as a directed evidence-centric graph. Each evidence no de is linked to normalized biomedical entities, and evidence–evidence semantic relations are stored as typed directed edges. F or eac h evidence no de e ∈ V E and normalized entit y t ∈ V T , the graph contains edges e → t for evidence–entit y alignmen t and e i → e j for evidence–evidence relations. This top ology differs from conv entional biomedical knowledge graphs, where entities are commonly linked directly by flattened predicates. By explicitly mo delling evidence as a graph ob ject, EvidenceNet preserv es prov enance, quan titative supp ort, and contextual v alidity while remaining suitable for do wnstream graph analysis and machine learning. 6 This final stage also supp orts quality assurance. Automated verification is applied during dupli- cate resolution and relation construction. Each evidence no de also retains explicit review metadata, so manual exp ert audit can b e incorp orated without altering the schema. The resulting graph func- tions as an extracted knowledge structure and as a curated, extensible biomedical data resource. 3 Data Records W e provide tw o disease-sp ecific EvidenceNet resources, one centred on hepato cellular carcinoma (HCC) and the other on colorectal cancer (CRC). Eac h resource includes a structured evidence collection and a corresp onding graph representation. The structured collection preserves the full record-lev el description of each evidence unit, including pro venance, study context, PICO attributes, extracted en tities, quantitativ e attributes, and qualit y scores. The graph representation pro vides an in tegrated net work view in which evidence records connect to normalized biomedical entities and to other evidence records through t yp ed semantic relations. The t wo release lay ers are sync hronized through stable evidence identifiers and normalized entit y identifiers, allo wing users to mov e b et w een record-lev el insp ection and graph-lev el analysis without losing prov enance. The ov erall transforma- tion and schema are illustrated in Fig. 2, whereas the tables b elow en umerate the released fields and distributions in detail. W e deriv e evidence knowledge graphs from recent PubMed articles for eac h disease. The HCC resource con tains 7,872 evidence records and a graph with 10,328 no des, including evidence and normalized biomedical en tities, and 49,756 edges. The CR C resource con tains 6,622 evidence records and a graph with 8,795 no des and 39,361 edges. The graph no de coun ts exceed the evidence-record coun ts b ecause the graph lay er also includes normalized biomedical entit y no des that connect related evidence units across pap ers. Both resources are explicitly disease-sp ecific and retain release-lev el metadata, including disease lab el, v ersion, creator information, and up date time. In b oth datasets, the dominan t study designs are cohort, in-vivo, randomized controlled trial, in-vitro, and meta-analysis. Clinical-stage annota- tions are dominated by preclinical, clinical, phase I, phase I I, and phase I I I lab els. Evidence quality scores are concentrated in the intermediate range, with grade C as the most frequent category in b oth diseases. Small differences b et ween DOI counts and title counts reflect records that are retained through title-based pro venance recov ery when DOI metadata are incomplete or unav ailable. The net work structure reflects the evidence-centric design of the resource. In b oth graphs, the most frequent edge t yp es b et ween evidence no des are SUPPOR TS, EXTENDS, and REFINES. Less frequent but still informativ e relation t yp es include CONTRADICTS, CAUSAL_CHAIN, and REPLICA TES. Most evidence no des participate in at least one lo cal semantic neighbourho o d while the o verall graph remains interpretable and sparse. T able 2 summarizes the ov erall scale and structural c haracteristics of the released HCC and CR C EvidenceNet resources. T able 3 rep orts the distributions of evidence grades, study designs, and clinical stages. T able 4 summarizes graph relation types, and T able 5 rep orts cov erage of key pro venance, contextual, and quantitativ e fields. 7 T able 2: Expanded summary statistics of the released EvidenceNet resources. F eature HCC EvidenceNet CR C EvidenceNet Corpus and pr ovenanc e Pro cessed full-text articles 470 472 Unique DOIs 439 445 Unique source titles 457 452 Unique journals 157 179 Publication y ear range 2009–2025 2009–2025 Median publication y ear 2022 2022 Gr aph sc ale and top olo gy Evidence records 7,872 6,622 En tity no des 2,456 2,173 T otal graph no des 10,328 8,795 T otal graph edges 49,756 39,361 Graph densit y 0.000467 0.000509 Evidenc e richness A verage evidence relations 3.95 4.07 Median evidence relations 1.00 1.00 A verage linked entities 2.27 1.80 Median link ed entities 2.00 1.00 A verage core entities 5.85 6.09 Median core en tities 4.00 4.00 A verage merged records 1.13 1.16 Records with v ersion > 1 (%) 37.7 38.2 Evidenc e quality A verage comp osite_score 0.53 0.52 Median comp osite_score 0.50 0.49 Comp osite_score range 0.30–0.89 0.27–0.89 Most common evidence grade C C Field c over age (%) Records with comparison 57.1 59.7 Records with p -v alue 10.2 12.6 Records with sample size 24.8 28.0 Records with fold c hange 4.3 4.9 Records with bio-mec hanism 68.0 69.2 Records with phenot yp e 97.4 97.9 Records with source text 100.0 100.0 8 T able 3: Distribution of evidence grades, study designs, and clinical stages in the released EvidenceNet resources. V alues are shown as count (%). Category group Category HCC EvidenceNet CR C EvidenceNet Evidence grade A 60 (0.8%) 34 (0.5%) B 1,732 (22.0%) 1,251 (18.9%) C 5,886 (74.8%) 5,024 (75.9%) D 194 (2.5%) 313 (4.7%) Study design cohort 2,164 (27.5%) 1,680 (25.4%) in-viv o 1,811 (23.0%) 1,520 (23.0%) unkno wn 1,416 (18.0%) 1,412 (21.3%) in-vitro 1,115 (14.2%) 914 (13.8%) rct 703 (8.9%) 607 (9.2%) meta-analysis 426 (5.4%) 253 (3.8%) computational 192 (2.4%) 200 (3.0%) case-con trol 45 (0.6%) 36 (0.5%) Clinical stage clinical 3,087 (39.2%) 2,661 (40.2%) preclinical 2,976 (37.8%) 2,490 (37.6%) phase-i 386 (4.9%) 323 (4.9%) phase-ii 649 (8.2%) 693 (10.5%) phase-iii 756 (9.6%) 453 (6.8%) phase-iv 18 (0.2%) 2 (0.0%) T able 4: Distribution of graph edge types in the released EvidenceNet resources. V alues are sho wn as count (%). Percen tages are calculated relative to the total num b er of edges in each graph. Relation type HCC EvidenceNet CR C EvidenceNet LINKED_TO 17,849 (35.9%) 11,910 (30.3%) SUPPOR TS 13,861 (27.9%) 10,044 (25.5%) EXTENDS 9,263 (18.6%) 9,670 (24.6%) REFINES 6,115 (12.3%) 5,124 (13.0%) CONTRADICTS 1,111 (2.2%) 1,041 (2.6%) CA USAL_CHAIN 856 (1.7%) 1,101 (2.8%) REPLICA TES 701 (1.4%) 471 (1.2%) T otal edges 49,756 (100%) 39,361 (100%) 9 T able 5: Cov erage of key evidence and prov enance attributes in the released EvidenceNet resources. V alues are shown as count (%). Percen tages are calculated relative to the total num b er of evidence records in eac h resource. Field HCC EvidenceNet CR C EvidenceNet Pr ovenanc e fields Source title 7,786 (98.9%) 6,220 (93.9%) Journal name 6,820 (86.6%) 5,906 (89.2%) Impact factor 6,820 (86.6%) 5,895 (89.0%) Journal quartile 6,820 (86.6%) 5,895 (89.0%) Citation coun t 6,820 (86.6%) 5,929 (89.5%) Author list 6,820 (86.6%) 5,879 (88.8%) PICO and c ontextual fields Comparison 4,492 (57.1%) 3,951 (59.7%) Bio-mec hanism 5,355 (68.0%) 4,585 (69.2%) Phenot yp e 7,666 (97.4%) 6,480 (97.9%) Exp erimen tal con text 7,872 (100.0%) 6,622 (100.0%) Source text 7,872 (100.0%) 6,622 (100.0%) Quantitative fields p -v alue 800 (10.2%) 835 (12.6%) Sample size 1,955 (24.8%) 1,856 (28.0%) F old change 340 (4.3%) 325 (4.9%) 3.1 Record-lev el structure The record-level comp onen t of eac h release stores the complete structured representation of in- dividual evidence units. Each record is uniquely indexed b y an evidence identifier and contains six principal information lay ers. These comprise source prov enance, PICO attributes, biomedical en tities, quantitativ e statistics, evidence quality scores, and semantic graph relations. Bibliographic co verage is high for core prov enance fields. In the HCC resource, source title is a v ailable for 98.9% of records. Journal, impact factor, journal quartile, citation count, and author information are eac h av ailable for 86.6%. In the CRC resource, the corresp onding cov erages are 93.9%, 89.2%, 89.0%, 89.0%, 89.5%, and 88.8%, resp ectiv ely . Quantitativ e fields are less uniformly rep orted in the literature. Comparator information is presen t in 57.1% of HCC records and 59.7% of CR C records. Explicit p -v alues app ear in 10.2% and 12.6% of records, sample-size v alues in 24.8% and 28.0%, and fold-change v alues in 4.3% and 4.9% of HCC and CR C records, resp ectively . The evidence-qualit y ob ject stores b oth comp onen t-level and comp osite assessmen ts. In the HCC resource, the grade distribution is C (5,886), B (1,732), D (194), and A (60). In the CR C resource, the distribution is C (5,024), B (1,251), D (313), and A (34). These v alues indicate that the released resources preserve the full sp ectrum of extracted evidence, rather than only a narro wly filtered high-confidence subset. Eac h record also stores a set of extracted biomedical entities and, when a v ailable, normalized biomedical identifiers. In HCC, the most frequent extracted semantic classes are Drug, Phenotype, Gene, and Disease. In CRC, the most frequen t classes are Drug, Disease, Gene, and Phenotype. Records may also include directed semantic relations to other evidence units, duplicate-merging pro venance, lifecycle timestamps, version information, and fields reserv ed for future man ual curation (T able 6). 10 3.2 Graph-lev el structure The graph-lev el comp onen t stores the integrated disease-specific evidence net work. It con tains t wo no de classes, namely evidence no des and normalized biomedical en tity no des. Evidence no des retain a flattened subset of the attributes present in the full record-level represen tation, including pro venance, study type, disease context, in terven tion, mec hanism, phenotype, and comp osite score. En tity no des represent normalized biomedical concepts and preserve canonical names, seman tic classes, and source-database prov enance. In the HCC resource, the normalized en tity la yer comprises 1,033 genes, 745 drugs or comp ounds, 367 diseases, 180 phenotypes, and 131 path wa ys. In the CR C resource, the corresp onding counts are 908 genes, 689 drugs or comp ounds, 352 diseases, 119 phenotypes, and 105 pathw a ys. These en tity sets form the shared concept lay er through which individual evidence units are integrated in to a coherent graph. Edges enco de b oth evidence-to-entit y alignment and evidence-to-evidence semantic relations. Evidence-to-en tity edges connect structured evidence units to normalized biomedical concepts. Evidence-to-evidence edges encode seman tic relations including support, extension, refinemen t, con tradiction, replication, and causal ordering. Because the graph is directed, edge orientation is preserv ed and should b e interpreted as part of the semantic structure of the resource (T able 7). T able 6: Core fields represen ted in the released EvidenceNet resources at the record lev el. Field Description evidence_id Unique identifier of the evidence record. source Source metadata, including DOI, title, authors, journal, publication year, cita- tion coun t, impact factor, journal quartile, and source do cument path. pico Structured study con text, including study ob ject, interv ention, comparison, and outcome metrics. core_en tities Extracted biomedical entities. Eac h entit y record includes the ra w name, se- man tic type, normalized identifier, canonical name, database source, and link- ing score when a v ailable. bio_mec hanism F ree-text description of the rep orted biological mec hanism. phenot yp e F ree-text description of the observed biological or clinical phenotype. study_design Study t yp e, suc h as cohort, in-vivo, in-vitro, or randomized trial. clinical_stage T ranslational stage of the evidence, such as preclinical, clinical, or trial phase. statistics Quantitativ e attributes, including p -v alue, fold change, confidence interv al, sample size, effect size, and statistical metho d when rep orted. score Evidence-qualit y ob ject containing impact score, statistics score, sample size score, comp osite score, llm confidence, and grade level. source_text Supp orting text span from the source article. link ed_entities List of normalized entit y identifiers used to connect the evidence record to the graph en tity lay er. evidence_relations Directed semantic relations to other evidence records, including source identi- fier, target iden tifier, relation type, similarity score, rationale, and timestamp. merged_from Iden tifiers of absorb ed duplicate evidence records retained for prov enance track- ing. review_status Review state of the evidence record. In this release, all records are mark ed as p ending review. created_at, up- dated_at, version Record v ersioning and timestamps. 11 T able 7: Core fields represen ted in the released EvidenceNet resources at the graph lev el. Field Description metadata Release-level metadata, including disease lab el, creator informa- tion, v ersion, and up date timestamp. evi_no de_attr Flattened evidence-no de fields used in the graph representation, including evidence iden tifier, bibliographic metadata, study de- sign, clinical stage, disease label, mec hanism, phenot yp e, inter- v ention, study ob ject, link ed entities, and comp osite score. en t_no de_attr Normalized biomedical entit y fields, including canonical name, se- man tic type, and source database. evi_en t_edges Directed edges connecting evidence no des to normalized entities, with en tity type and linking score. evi_evi_edges Directed semantic relations (SUPPOR TS, EXTENDS, REFINES, CONTRADICTS, CAUSAL_CHAIN, REPLICA TES) with sim- ilarit y score and rationale. The record-level and graph-level comp onen ts provide complementary views of the same biomed- ical resource. The record-level representation preserv es the full seman tic structure of individual evidence units, whereas the graph-level represen tation supports in tegration, visualization, and do wn- stream net work analysis. 4 Data Ov erview T o complemen t the structured descriptions in the Data R e c or ds section, w e further summarize the released EvidenceNet resources through three descriptive visualizations that emphasize graph top ology and dataset comp osition. Figure 3 pro vides a global view of the HCC and CRC graphs, Fig. 4 highlights represen tative lo cal motifs, and Fig. 5 summarizes their temp oral, metho dological, scoring, and seman tic-relation profiles. T ogether, these views pro vide a concise o verview of how the released evidence is organized at the disease level. A t the global level, b oth disease-specific graphs exhibit a dominan t connected bac kb one together with smaller p eripheral mo dules (Fig. 3). This structure indicates that a substantial p ortion of the literature con v erges on recurring in terven tion–mec hanism–outcome patterns that are repeatedly connected across publications. At the same time, the presence of p eripheral mo dules sho ws that the resource do es not collapse all evidence in to a single homogeneous net work. More sp ecialized themes remain preserved as localized seman tic neigh b ourho ods, allowing users to distinguish broadly connected areas of inv estigation from narrow er sub domains. The colour distribution in Fig. 3 further shows that the backbone is supp orted b y heterogeneous study designs rather than by a single exp erimen tal tier. Preclinical and clinical evidence records are in terwo v en within the same large-scale structure, indicating that EvidenceNet integrates mec hanis- tic, translational, and clinical findings into a shared disease-specific represen tation. This prop ert y is imp ortant for downstream interpretation b ecause it allows users to insp ect not only the den- sit y of evidence surrounding a topic, but also the diversit y of study t yp es that con tribute to that neigh b ourho od. Figure 4 pro vides a complementary lo cal-scale view of the same resources. Whereas Fig. 3 emphasizes global organization, the motif panels show how seman tically related evidence records form in terpretable neigh b ourho ods within the larger graphs. In b oth diseases, contradiction-ric h regions can b e iden tified in whic h closely related evidence no des are connected by mixed semant ic relations rather than b y uniformly supportive links alone. Such lo cal structures indicate that the 12 Evidence r ecor ds: 7,872 Entity nodes: 2,456 T otal edges: 49,756 Mean scor e: 0.53 Mean evidence r elations: 3.95 Displayed cr oss-paper evidence nodes: 2,724 Displayed cr oss-paper semantic links: 25,486 HCC EvidenceNet: score-filtered cross-paper semantic overview Evidence r ecor ds: 6,622 Entity nodes: 2,173 T otal edges: 39,361 Mean scor e: 0.52 Mean evidence r elations: 4.07 Displayed cr oss-paper evidence nodes: 1,903 Displayed cr oss-paper semantic links: 19,510 CRC EvidenceNet: score-filtered cross-paper semantic overview Study design (node colour) r ct meta_analysis cohort case_contr ol in_vitr o in_vivo computational unknown Semantic r elation (edge colour) SUPPOR TS EX TENDS REFINES CONTR ADICTS CAUS AL_CHAIN REPLICA TES Figure 3: Global ov erview of the released HCC and CR C EvidenceNet resources. No des represen t filtered evidence records and are coloured by study design. Edges represent cross-pap er semantic relations and are coloured b y relation type. Both diseases show a densely connected backbone together with smaller p eripheral mo dules. V ery small disconnected comp onen ts are omitted for visual clarit y . graph preserves not only evidence density but also the in ternal diversit y of how findings relate to one another across studies. The p eripheral motifs add a further la yer of in terpretability . In HCC, a compact mo dule cen tred on sirolimus and tacrolimus reflects a sp ecialized evidence cluster linked to transplant immunosup- pression, while in CR C, a p ostoperative ctDNA-cen tred mo dule illustrates how clinically fo cused themes remain preserv ed outside the main bac kb one. These examples sho w that EvidenceNet re- tains coherent subgraphs at multiple scales, allowing users to mo ve from disease-wide insp ection to finer seman tic neighbourho o ds without losing prov enance or contextual structure. Figure 5 complements the top ology-orien ted view by summarizing four key quantitativ e prop er- ties of the released resources. The annual record coun ts in Fig. 5a show that b oth disease graphs are w eighted to wards recen t literature, with a clear increase in extracted evidence in the later publication y ears. This pattern is consistent with the construction strategy of EvidenceNet, whic h prioritizes recen t full-text articles while still retaining longitudinal co verage across the disease literature. The study-design comp osition in Fig. 5b shows that the released resources remain metho dologi- cally heterogeneous. Cohort, in-vivo, in-vitro, and randomized-trial records account for m uch of the evidence volume, while meta-analysis, computational, and case-con trol studies pro vide additional complemen tary strata. The score distributions in Fig. 5c are concen trated in the mid-to-high range for b oth diseases, indicating that the release preserves broad co verage without b eing dominated by v ery lo w-confidence records. The semantic-relation distributions in Fig. 5d sho w that SUPPOR TS, EXTENDS, and REFINES constitute the ma jor cross-pap er relation classes, whereas CONTRA- DICTS, CAUSAL_CHAIN, and REPLICA TES form smaller but still informative p ortions of the graph. These summaries show that the released EvidenceNet resources are recent, metho dologically div erse, and seman tically structured for cross-pap er in terpretation and do wnstream graph analysis. 13 Combined tar geted and immunotherapy -associated evidence HCC contradiction-rich neighbourhood combined tar geted drug and immunotherapy T ransplant immunosuppr ession centr ed on sir olimus/tacr olimus HCC peripheral module sir olimus (most fr equently in combination with low dosages of tacr olimus) Anti-EGFR tr eatment neighbour hood centr ed on FOLFO X plus panitumumab CRC contradiction-rich neighbourhood FOLFO X plus panitumumab P ostoperative ctDNA -associated module CRC peripheral module ctDNA -positive status af ter curative-intent sur gery r ct meta_analysis cohort case_contr ol in_vitr o in_vivo computational unknown SUPPOR TS EX TENDS REFINES CONTR ADICTS CAUS AL_CHAIN REPLICA TES Figure 4: Representativ e lo cal motifs in the HCC and CRC EvidenceNet resources. The four panels sho w example subgraphs selected from the full evidence graphs, including contradiction-ric h neigh b ourho ods and more sp ecialized p eripheral mo dules. These motifs illustrate how EvidenceNet supp orts b oth global insp ection and fine-grained exploration of disease-relev ant evidence patterns. 5 T ec hnical V alidation 5.1 Comp onen t fidelity v alidation W e first ev aluate comp onen t fidelity through a targeted man ual audit of the released HCC and CRC resources (T able 8). The audit comprises 20 evidence-extraction cases, 20 en tity-normalization cases, 12 fusion cases with merge pro venance, and 20 seman tic relations. Sampling is stratified b y disease and audit type so that the reviewed set cov ers m ultiple study designs, entit y classes, relation t yp es, and merge scenarios. All cases are sampled from released records with sufficien t textual context. En tity-linking cases are restricted to high-confidence mappings that can b e directly grounded in the source sentence, so this audit ev aluates the correctness of accepted links rather than the recall of all p ossible candidates. Evidence extraction shows high fidelity . The field-level macro accuracy reaches 98.3%, and 95.0% of review ed records are fully correct with resp ect to source supp ort, study-design assignment, and k ey-entit y capture. These results indicate that the extraction stage generally preserves the main claim and study context of the source text. En tity normalization is also robust. All review ed high-confidence links are mapp ed to the correct 14 2010 2012 2014 2016 2018 2020 2022 2024 Y ear 0 200 400 600 800 1000 1200 Evidence r ecor ds a Annual evidence records by source year HCC CR C HCC CR C 0 20 40 60 80 100 R ecor ds (%) n=7,872 n=6,622 b Study-design composition Study design cohort in vivo in vitr o r ct meta analysis computational case contr ol unknown HCC CR C 0.3 0.4 0.5 0.6 0.7 0.8 0.9 1.0 Composite scor e median 0.50 IQR 0.46--0.59 median 0.49 IQR 0.45--0.58 c Composite score distribution 0 10 20 30 40 Semantic edges (%) supports e xtends r efines contradicts causal chain r eplicates 43.4 29.0 19.2 3.5 2.7 2.2 36.6 35.2 18.7 3.8 4.0 1.7 d Semantic relation composition HCC CR C Figure 5: Quantitativ e ov erview of the released HCC and CRC EvidenceNet resources. (a) Ann ual evidence-record counts by source year. (b) Study-design comp osition of evidence records. (c) Comp osite-score distributions across records. (d) Comp osition of semantic relation types among evidence–evidence edges. T arKG [11] concept, yielding a link accuracy of 100.0%. A lo wer type-compatibility score of 80.0% is observed when assessing whether the mapp ed on tology class is the most semantically appropriate one. The remaining errors are mainly b oundary cases b et w een disease-level and phenotype-level concepts rather than failures of lexical grounding. F usion is more c hallenging than extraction and normalization, but remains reliable ov erall. The final retained record is coherent in all reviewed cases, and the fusion integrit y score reac hes 87.5%. The lo wer same-underlying-evidence rate of 83.3% suggests that a small subset of merges inv olv es closely related but not fully identical evidence statements. Seman tic linking is precise. All review ed edges connect gen uinely related evidence records, giving an edge precision of 100.0%. In addition, 90.0% of audited links are assigned the cor- rect semantic relation t yp e. Errors are concen trated in the more in terpretive CONTRADICTS and CAUSAL_CHAIN categories, whereas all reviewed SUPPOR TS, EXTENDS, R EFINES, and REPLICA TES relations are judged correct. These audits show that the released EvidenceNet resources are technically reliable at the comp o- nen t level. Easier steps such as record extraction and high-confidence entit y normalization ac hieve v ery high accuracy , whereas fusion and fine-grained relation t yping remain more difficult, as ex- p ected for literature-derived evidence graphs. 15 T able 8: Comp onen t-level fidelity audit of the released EvidenceNet resources. Primary metrics summarize the main usabilit y-oriented criterion for each comp onen t, and secondary metrics capture a stricter or more fine-grained asp ect of the same audit. Comp onen t n Primary metric V alue (%) Secondary metric V alue (%) Evidence extraction 20 Field-lev el macro ac- curacy 98.3 Strict record accuracy 95.0 En tity normalization 20 Link accuracy 100.0 T yp e compatibilit y 80.0 Evidence fusion 12 F usion integrit y score 87.5 Same-underlying- evidence rate 83.3 Seman tic relations 20 Edge precision 100.0 Relation-t yp e accuracy 90.0 5.2 In ternal consistency v alidation W e next assess in ternal consistency using question–answ ering pairs generated directly from the re- leased graph conten t. This exp erimen t tests whether evidence already enco ded in the graph can b e retriev ed and used to supp ort correct answ ers without relying on external literature. It should there- fore b e in terpreted as an internal coherence and answer-reco very test, rather than as an out-of-graph generalization b enc hmark. Separate internal QA sets are constructed for HCC and CRC, each con- taining 50 graph-derived yes/no questions. W e compare four answering settings, namely a baseline LLM (GPT-5.1) without retriev al, T arKG-only [11] retriev al-augmented generation (T arK G-RAG), lo cal literature retriev al (Article-RA G), and EvidenceNet retriev al (EvidenceNet-RAG). EvidenceNet giv es the highest accuracy in b oth diseases (T able 9). In HCC, EvidenceNet reaches 96.0% accuracy , compared with 76.0% for the baseline, 88.0% for T arK G-only retriev al, and 78.0% for lo cal literature retriev al. In CR C, EvidenceNet reaches 92.0% accuracy , compared with 76.0%, 72.0%, and 78.0% for the same baselines, resp ectiv ely . EvidenceNet also ac hieves the highest a v- erage semantic similarity b et w een predicted and reference answ ers in b oth datasets. This pattern indicates more accurate binary decisions and closer alignmen t with the reference explanation. These results show that the released graph is internally coheren t. The gain o ver the baselines suggests that the impro vemen t do es not arise simply from general biomedical kno wledge or from access to a background knowledge graph alone, but from the structured organization of literature-derived evidence records and their semantic links within EvidenceNet. T able 9: Internal consistency ev aluation using graph-derived yes/no QA pairs. Accuracy and a verage semantic similarity are computed separately for HCC and CRC using 50 internally generated questions p er disease. HCC CR C Metho d Accuracy (%) A vg. semantic similarit y A ccuracy (%) A vg. semantic similarit y Baseline LLM 76.0 0.734 76.0 0.742 T arKG-RA G 88.0 0.729 72.0 0.749 Article-RA G 78.0 0.735 78.0 0.751 EvidenceNet- RA G 96.0 0.789 92.0 0.798 16 5.3 External reasoning utility v alidation W e next ev aluate whether EvidenceNet can support question answ ering beyond facts directly in- stan tiated in the graph. F or this purpose, w e assem ble an external y es/no b enc hmark by filtering HCC-related (98 samples) and CRC-related (93 samples) question–answering instances from three public biomedical QA resources, namely PubMedQA [51], BioASQ [52], and Evidence-Inference [53]. This setting is more demanding than the internal QA task b ecause the questions are not generated from EvidenceNet itself and therefore require semantic generalization rather than direct recov ery of graph-nativ e statements. W e compare five answering settings, including a baseline LLM without retriev al, T arKG-RA G, Article-RA G, EvidenceNet-RA G, and a com bined EvidenceNet+T arK G setting. As shown in T a- ble 10, the combined setting ac hiev es the b est accuracy in b oth diseases. In HCC, accuracy increases from 56.1% for the baseline to 59.2% with EvidenceNet alone and 61.2% with EvidenceNet+T arKG. In CRC, the corresp onding v alues are 64.8%, 67.0%, and 68.1%. T arKG-RA G and Article-RAG do not consisten tly outp erform the baseline. These results indicate that EvidenceNet con tributes useful disease-specific evidence for answ ering external biomedical questions, even when the b enc hmark is not derived from the graph itself. The additional gain from combining EvidenceNet with T arKG suggests that the tw o resources contribute complemen tary information. EvidenceNet con tributes literature-grounded exp erimen tal and clinical evidence, whereas T arKG supplies broader en tity-lev el definitions and bac kground asso ciations. Absolute p erformance remains lo wer than in the internal consistency exp erimen t, whic h is exp ected b ecause the external b enc hmark spans a broader range of question formulations and knowledge requiremen ts. Nev ertheless, the consistent improv emen t across b oth diseases supp orts the utility of EvidenceNet as a retriev al substrate for downstream reasoning tasks. T able 10: External reasoning utilit y ev aluated on filtered HCC- and CR C-related yes/no questions from public biomedical QA datasets. Accuracy and av erage semantic similarity are computed separately for HCC and CR C b enc hmark subsets. HCC CR C Metho d A ccuracy (%) A vg. semantic similarit y A ccuracy (%) A vg. semantic similarit y Baseline LLM 56.1 0.601 64.8 0.647 T arKG-RA G 55.1 0.609 63.7 0.651 Article-RA G 54.1 0.600 62.6 0.668 EvidenceNet-RA G 59.2 0.612 67.0 0.664 EvidenceNet+T arKG 61.2 0.619 68.1 0.652 5.4 Structural predictive v alidation The QA-based ev aluations ab o v e mainly assess whether EvidenceNet can supp ort seman tically grounded retriev al and reasoning o ver already observ ed evidence. Predictive v alidity addresses a complemen tary question, namely whether graph structure itself con tains forward-looking informa- tion that an ticipates findings app earing only in later, previously unpro cessed literature. W e ev aluate EvidenceNet from t wo predictive persp ectiv es. The first is a general link prediction task, which mea- sures the abilit y of the graph to recov er nov el linked en tity pairs from future pap ers. The second is a scenario-based target discov ery task, which asks whether the graph can prioritize newly rep orted therap eutic targets in a more application-oriented setting. 17 F or the general link prediction task, we construct disease-sp ecific temp oral hold-out datasets using recen t papers not included in graph construction. T raining positives are linked en tity pairs already present in the released graph, whereas test p ositives are link ed en tity pairs extracted from new er pap ers but absent from the graph at training time. Random unseen pairs serve as negativ es. This yields 501 future p ositiv e pairs for HCC and 665 for CR C. W e compare four complementary predictors on the EvidenceNet graph, including a random-forest classifier using hand-crafted graph features, a no de2v ec [54] embedding mo del follo wed by logistic regression, a bipartite graph neural net work [55], and a shortest-path heuristic baseline. All four metho ds ac hieve b etter-than-random performance, indicating that the released graph con tains measurable predictiv e signal for future links (T able 11). The shortest-path baseline p er- forms best in b oth diseases, reaching an A UC of 0.831 and AP of 0.884 in HCC, and an AUC of 0.833 and AP of 0.794 in CR C. Among the learned mo dels, no de2v ec p erforms b est in HCC (AUC 0.754, AP 0.861). In CR C, no de2v ec again provides the strongest learned-mo del p erformance (AUC 0.725, AP 0.717), while the random-forest mo del remains comp etitiv e (AUC 0.703, AP 0.634). The GNN shows consisten t but more mo derate p erformance in HCC (A UC 0.699, AP 0.668) and CR C (A UC 0.658, AP 0.579). These results suggest that lo cal graph top ology is highly informative for anticipating future as- so ciations. This in terpretation is reinforced by the strong p erformance of the shortest-path baseline, whic h relies only on structural proximit y rather than complex learned parameters. The comp etitiv e p erformance of no de2v ec and random forest further indicates that b oth distributed graph represen- tations and engineered top ological features capture useful predictive signal. In the random-forest mo del, the most influential features are seman tic similarity and degree-based preferen tial attac h- men t, suggesting that future links tend to arise where seman tically related entities are already em b edded in densely connected evidence neighbourho ods. These findings sho w that the predictive utilit y of EvidenceNet is enco ded in graph organization as well as in text-level retriev al. T able 11: Multi-metho d ev aluation of future link prediction on held-out recent literature. P ositive test pairs are nov el linked entit y pairs extracted from previously unseen pap ers and absent from the released graph at training time. HCC CR C Metho d A UC AP AUC AP Random forest 0.708 0.746 0.703 0.634 no de2v ec + logistic regression 0.754 0.861 0.725 0.717 Bipartite GNN 0.699 0.668 0.658 0.579 Shortest-path baseline 0.831 0.884 0.833 0.794 W e next examine a more application-orien ted setting based on emerging therap eutic targets rep orted in unseen recen t pap ers. This task differs from generic link prediction in b oth viewp oin t and utilit y . Instead of asking whether an y missing edge can b e recov ered, it asks whether the disease- cen tred evidence neighbourho o d can prioritize biologically plausible and therap eutically relev ant targets b efore they are densely represented in the graph. W e b enc hmark four HCC targets and fiv e CR C targets curated from newly pro cessed pap ers. F or each candidate, we compute a disease– target proximit y score in EvidenceNet using graph-neighbourho o d ov erlap and compare it with an analogous score deriv ed from T arKG. Given the small num b er of b enc hmark targets, this analysis is intended as a scenario-based pro of of concept for translational prioritization rather than as a definitiv e ranking b enc hmark. As shown in T able 12, EvidenceNet assigns non-zero scores to one HCC target (SPP1) and tw o 18 CR C targets (CD47 and CTLA4), whereas T arK G assigns zero to all candidates. Across the full b enc hmark, three of nine literature-derived targets receive p ositiv e EvidenceNet scores. Among the four candidates already present as no des in EvidenceNet, three receive non-zero disease-proximit y scores. The remaining failures are mainly cold-start cases in whic h the target is absen t from the curren t graph vocabulary , making recov ery imp ossible under a purely graph-based ranking sc heme. Ev en under this conserv ativ e setup, the results suggest that EvidenceNet can supp ort weak but meaningful prioritization of emerging disease-relev ant targets when some supp orting graph con text is a v ailable. T able 12: Scenario-based prediction of therap eutic targets rep orted in unseen recent pap ers. A p ositiv e score indicates non-zero graph pro ximity b et ween the disease no de and the candidate target. Disease Benc hmark targets EvidenceNet p ositiv e-score targets T arK G p ositiv e-score targets HCC 4 1/4 (SPP1) 0/4 CR C 5 2/5 (CD47, CTLA4) 0/5 T ogether, these t wo exp erimen ts support the predictive v alidit y of EvidenceNet. The general link prediction task sho ws that the global structure of EvidenceNet contains forw ard-lo oking information ab out future entit y asso ciations. The target discov ery task shows how the same graph can b e used in a more fo cused translational setting. Alongside the QA results ab o v e, these findings indicate that EvidenceNet is useful b oth as a rep ository of structured evidence and as a graph substrate for an ticipating new relationships and prioritizing candidate therap eutic directions. 6 Usage Notes EvidenceNet can b e used at t wo complemen tary levels. The record-level release is most suitable for evidence-centric retriev al, man ual review, and in terpretation b ecause each evidence unit pre- serv es source text, study design, clinical stage, quantitativ e attributes, and pro venance. The graph- lev el release is b etter suited to visualization, semantic neigh b ourho od analysis, link prediction, and disease-sp ecific graph learning. Users in terested in conserv ative do wnstream analyses ma y further filter records b y comp osite score, evidence grade, study design, or clinical stage. More exploratory analyses ma y b enefit from retaining the full release, including low er-confidence and con tradictory evidence. This release should b e interpreted within its co verage limits. EvidenceNet is built from disease- sp ecific, recen t PubMed-indexed articles with accessible full text, rather than from the en tiret y of the biomedical literature. Absence of a no de, edge, or target in the current graph should therefore not b e in terpreted as evidence of biological absence. In some cases, it instead reflects corpus cov erage, incomplete on tology alignment, or cold-start en tities that hav e not yet entered the current graph v o cabulary . Because EvidenceNet in tentionally in tegrates heterogeneous evidence t yp es, graph proximit y should not b e interpreted as direct clinical supp ort without insp ecting no de-lev el context. Preclin- ical, translational, and clinical findings co exist in the same resource, and semantic relation edges indicate cross-pap er consistency or relev ance rather than therap eutic recommendation. W e there- fore recommend that downstream users insp ect the link ed source text, study design, and evidence score b efore drawing mec hanistic or translational conclusions. Entit y normalization is aligned to T arKG, which impro ves interoperability but may still con tain ontology-boundary cases, esp ecially b et w een disease and phenotype concepts. 19 7 Conclusion W e presen t EvidenceNet as a disease-sp ecific, evidence-centric framework for conv erting full-text biomedical literature into computable kno wledge graphs that preserv e prov enance, study con text, and quantitativ e supp ort. By treating exp erimen tally grounded findings as the primary graph unit, EvidenceNet complements conv en tional biomedical kno wledge graphs that fo cus mainly on entit y- lev el facts. The released HCC and CRC resources show that this design scales to literature-sized corp ora while retaining interpretable structure at b oth the record and graph levels. Our ev aluations demonstrate that EvidenceNet is b oth technically reliable and practically useful. Comp onen t-lev el audits supp ort the fidelity of evidence extraction, entit y normalization, duplicate handling, and semantic relation typing. Downstream v alidation further shows that the resulting graphs supp ort internal consistency c hecking, improv e external biomedical question answering when used for retriev al, and capture structural signal for future link prediction and target prioritization. These findings indicate that EvidenceNet functions b oth as a structured data resource and as a substrate for disease-sp ecific biomedical reasoning. EvidenceNet provides an evidence-a w are kno wledge resource for precision medicine and trans- lational discov ery . Because the framework is mo dular and disease-sp ecific, it can b e extended to additional disease areas, up dated with new literature, and in tegrated with external biomedical kno wledge resources and retriev al-based LLM systems. Resources of this t yp e may support more transparen t biomedical question answering, more context-sensitiv e knowledge synthesis, and more efficien t hypothesis generation from the rapidly expanding scien tific literature. App endix A LLM prompts used in EvidenceNet T o impro ve transparency and repro ducibilit y , we summarize here the principal LLM prompts used in EvidenceNet construction and ev aluation. Dynamic runtime con tent is represented by placeholders suc h as , , and . A.1 Prompts used in graph construction A.1.1 Prompt A 1. Chunk-lev el evidence extraction. This prompt is used to conv ert pap er text segmen ts into structured evidence records. A dedicated system instruction established the extraction role, JSON sc hema, and grounding constrain ts, and the user prompt supplied the disease context, text segment, and few-shot examples. F or readability , length y inserted conten t and rep eated fields were abbreviated without c hanging the prompt logic. Prompt A1. Ch unk-level evidence extraction [System instruction] You are a biomedical evidence extraction expert specializing in evidence-based medicine (EBM). Your task is to extract structured evidence from academic paper text following the PICO framework and GRADE guidelines. REQUIRED JSON FIELDS: - study_object - intervention - comparison - outcome_metrics - core_entities 20 - bio_mechanism - phenotype - study_design - clinical_stage - p_value - sample_size - fold_change - experimental_context - source_text - extraction_confidence CRITICAL RULES: 1. Only extract evidence explicitly stated in the text. 2. If a paragraph contains multiple distinct experiments, extract them separately. 3. The source_text field must be an exact verbatim quote. 4. Do not treat background statements or prior work as new evidence. 5. If a field is missing, set it to null. Output only valid JSON. Do not include markdown code blocks. [User prompt template] Extract all distinct pieces of experimental evidence from the following text segment (section: ). Target disease context: Output a JSON object with a key "evidence" containing a list of evidence items. ### FEW-SHOT EXAMPLES Example 1: Input Text: "We treated HepG2 cells with 5 Micrometre Sorafenib for 48h. The CCK-8 assay showed that cell viability was significantly reduced compared to DMSO control (p < 0.01, n=3)." Example 2: Input Text: "Hepatocellular carcinoma (HCC) is a major cause of cancer-related death. Previous studies have linked TP53 mutations to poor prognosis." Output: {"evidence": []} Example 3: Input Text: "Knockdown of GeneA decreased migration. Additionally, Western blot analysis revealed decreased phosphorylation of AKT." Output: {"evidence": [{"phenotype": "decreased migration"}, {"bio_mechanism": "decreased phosphorylation of AKT"}]} ### TEXT TO ANALYZE --- --- A.1.2 Prompt A 2. Intra-document aggregation and enric hmen t. After ch unk-lev el extraction, evidence ob jects from the same pap er were passed to a second prompt that enric hed incomplete fields and remov ed only exact duplicates, while explicitly preserving dis- tinct exp erimen ts. 21 Prompt A2. In tra-do cumen t aggregation and enrichmen t [System instruction] You are an expert in evidence synthesis for biomedical research. Your task is to enrich and complete evidence extractions from the same paper, not to reduce or merge them. Preserve every distinct piece of evidence. Only remove entries that are exact word-for-word duplicates. Output only valid JSON matching the schema. [User prompt template] The following JSON array contains evidence extracted from different sections of the same paper. Paper title: DOI: <DOI> Tasks (in order of priority): 1. Fill in missing fields using information found elsewhere in the array. 2. Add a "conflict_note" field only if two entries report directly contradictory numbers for the same metric. 3. Remove only exact word-for-word duplicate entries. 4. Keep all distinct experiments; do not merge experiments that merely study similar topics. Output a JSON object with a key "evidence" containing the list of enriched evidence items. Evidence to enrich: <EVIDENCE JSON ARRAY> A.1.3 Prompt A 3. Evidence-to-evidence relation v erification. When rule-based semantic linking b et ween evidence records was uncertain or highly similar, an LLM v erification step was used to assign a final seman tic relation lab el. Prompt A3. Evidence-to-evidence relation verification [System instruction] You are an expert in biomedical evidence synthesis and evidence-based medicine. Your task is to classify the relationship between two pieces of experimental evidence extracted from scientific papers. Output only valid JSON. No markdown and no explanation outside the JSON object. [User prompt template] Classify the relationship between Evidence A and Evidence B. Evidence A: Intervention : <A_INTERVENTION> Mechanism : <A_MECHANISM> Phenotype : <A_PHENOTYPE> Study design : <A_DESIGN> Year : <A_YEAR> Key entities : <A_ENTITIES> Evidence B: Intervention : <B_INTERVENTION> Mechanism : <B_MECHANISM> Phenotype : <B_PHENOTYPE> Study design : <B_DESIGN> Year : <B_YEAR> Key entities : <B_ENTITIES> 22 Rule-based preliminary classification: <RULE_RELATION> Choose exactly one relationship: - SUPPORTS - CONTRADICTS - REFINES - EXTENDS - REPLICATES - CAUSAL_CHAIN Respond in the following JSON format: { "relation_type": "<one of the six types>", "confidence": <float 0.0-1.0>, "rationale": "<one sentence explanation>" } A.2 Prompts used in graph ev aluation A.2.1 Prompt A 4. Graph-derived yes/no QA generation. This prompt was used to generate in ternal QA pairs from evidence records already present in the graph. A ligh tw eight JSON-only system role was used during generation. Prompt A4. Graph-deriv ed y es/no QA generation [System instruction] You are a helpful assistant that outputs JSON. [User prompt template] You are an expert in biomedical question generation. Based strictly on the following evidence snippet from a scientific paper, generate a specific scientific question and its answer. Evidence Source Text: "<SOURCE_TEXT>" Context: Intervention: <INTERVENTION> Outcome: <OUTCOME> Mechanism: <MECHANISM> Task: 1. Generate a yes/no question that can be answered by this evidence. 2. Provide the yes/no classification. 3. Provide a concise explanation justified by the evidence. Output format (JSON): { "question": "Does [Intervention] cause [Outcome]...?", "class": "yes" or "no", "answer": "Yes. [Explanation...]" } 23 A.2.2 Prompt A 5. Retriev al-augmented QA answ ering. F or QA ev aluation, EvidenceNet w as queried either alone or together with T arKG. The answering prompt required explicit filtering of retriev ed evidence, binary classification, and explanation. A fallbac k v arian t was used when no relev ant graph evidence was retrieved. Prompt A5. Retriev al-augmen ted QA answ ering [EvidenceNet-only version] You are an expert biomedical researcher. CONTEXT (EvidenceNet): <EVIDENCE CONTEXT> QUESTION: <QUESTION> TASK: 1. Filter the evidence. - Relevant: direct mentions or conceptual matches. - Irrelevant: unrelated diseases or drugs. 2. Classify the answer as YES or NO. - If relevant evidence exists, answer based on it. - If all evidence is irrelevant, answer based on general knowledge rather than defaulting to "No". 3. Explain your reasoning and cite the evidence if used. OUTPUT FORMAT: CLASSIFICATION: [YES/NO] EXPLANATION: [Detailed reasoning] [EvidenceNet + TarKG version] You are an expert biomedical researcher. SOURCES: - EvidenceNet (Clinical Trials): specific experimental evidence - TarKG (Definitions): general biological definitions CONTEXT: <COMBINED_CONTEXT> QUESTION: <QUESTION> TASK: 1. Filter the retrieved EvidenceNet evidence. 2. Use TarKG for biological definitions if needed. 3. Classify the answer as YES or NO. - If relevant evidence exists, answer based on it. - If evidence is insufficient, answer using general biomedical knowledge. - Do not answer "No" solely because direct evidence is missing. 4. Provide a brief explanation. OUTPUT FORMAT: CLASSIFICATION: [YES/NO] EXPLANATION: [Reasoning] [No-evidence fallback version] Question: <QUESTION> 24 Task: 1. Classify the answer as YES or NO based on general biomedical knowledge. 2. No specific evidence was found in the database, so rely entirely on internal knowledge. Output Format: CLASSIFICATION: [YES/NO] EXPLANATION: [Detailed explanation from general knowledge] Data a v ailability The released EvidenceNet-HCC and EvidenceNet-CR C resources, including the record-lev el ev- idence collections and graph-lev el serializations used in this study , are dep osited in figshare at https://doi.org/10.6084/m9.figshare.31888399 . The release con tains disease-sp ecific files for HCC and CR C, including structured evidence records and graph files corresp onding to the resources describ ed in the Data R e c or ds section. F unding This w ork w as supp orted by the “Pioneer” and “Leading Go ose” R&D Program of Zhejiang (Key Researc h and Developmen t Program of Zhejiang Province), China (Grant No. 2025C01115). Author con tributions Chang Zong: conceptualization, methodology , softw are, data curation, formal analysis, v alida- tion, visualization, writing—original draft. Sicheng Lv: data curation, soft ware, formal analysis, writing—review and editing. Si-tu Xue: inv estigation, v alidation, writing—review and editing. Huilin Zheng: data curation, v alidation, visualization, writing—review and editing. Jian W an: su- p ervision, metho dology , writing—review and editing. Lei Zhang: conceptualization, sup ervision, pro ject administration, funding acquisition, writing—review and editing. All authors approv ed the final man uscript. Comp eting in terests The authors declare no comp eting interests. Co de a v ailability The code used in this study is implemen ted in Python and is a v ailable on GitHub at https: //github.com/ZUST- BIT/EvidenceNet- code . The rep ository con tains scripts for literature pre- pro cessing, evidence extraction, entit y normalization, graph construction, visualization, and down- stream ev aluation, together with the dep endency sp ecifications required to repro duce the analyses rep orted here. 25 References [1] T op ol, Eric J. High-p erformance medicine: the conv ergence of h uman and artificial in telligence. Natur e me dicine 25 (1), 44–56 (2019). [2] Siso diy a, Sanjay M. Precision medicine and therapies of the future. Epilepsia 62 , S90–S105 (2021). [3] Duffy , David J. Problems, c hallenges and promises: p erspectives on precision medicine. Brief- ings in bioinformatics 17 (3), 494–504 (2016). [4] Bornmann, Lutz; Haunsc hild, Robin; Mutz, Rüdiger. Gro wth rates of mo dern science: a latent piecewise gro wth curve approac h to mo del publication n umbers from established and new literature databases. Humanities and So cial Scienc es Communic ations 8 (1), 224 (2021). [5] González-Márquez, Rita; Schmidt, Luca; Schmidt, Benjamin M; Berens, Philipp; Kobak, Dmitry . The landscap e of biomedical researc h. Patterns 5 (6) (2024). [6] Go yal, Nandita; Singh, Na vdeep. Named entit y recognition and relationship extraction for biomedical text: A comprehensive survey , recent adv ancemen ts, and future research directions. Neur o c omputing 618 , 129171 (2025). [7] Strogano v, Oleg; Schedlbauer, Am b er; Lorenzen, Emily; Kadhim, Alex; Lobanov a, Anna; Lewis, David A; Glausier, Jill R. Unpacking unstructured data: A pilot study on extracting insigh ts from neuropathological rep orts of Parkinson’s disease patients using large language mo dels. Biolo gy Metho ds and Pr oto c ols 9 (1), bpae072 (2024). [8] Seinen, T om M; F ridgeirsson, Egill A; Ioannou, Solomon; Jeannetot, Daniel; John, Luis H; K ors, Jan A; Markus, Aniek F; Pera, Victor; Rekk as, Alexandros; Williams, Ross D; et al. Use of unstructured text in prognostic clinical prediction mo dels: a systematic review. Journal of the A meric an Me dic al Informatics Asso ciation 29 (7), 1292–1302 (2022). [9] Chandak, Pa yal; Huang, Kexin; Zitnik, Marink a. Building a knowledge graph to enable preci- sion medicine. Scientific data 10 (1), 67 (2023). [10] Himmelstein, Daniel Scott; Lizee, Antoine; Hessler, Christine; Brueggeman, Leo; Chen, Sab- rina L; Hadley , Dexter; Green, Ari; Khankhanian, P ouya; Baranzini, Sergio E. Systematic in tegration of biomedical kno wledge prioritizes drugs for repurp osing. eLife 6 , e26726 (2017). https://doi.org/10.7554/eLife.26726 . [11] Zhou, Cong; Cai, Ch ui-Pu; Huang, Xiao-Tian; W u, Song; Y u, Jun-Lin; W u, Jing-W ei; F ang, Jian-Song; Li, Guo-Bo. T arK G: a comprehensiv e biomedical kno wledge graph for target dis- co very . Bioinformatics 40 (10), btae598 (2024). [12] Subbiah, Vivek. The next generation of evidence-based medicine. Natur e me dicine 29 (1), 49–58 (2023). [13] Hosseini, Mohammad-Salar; Jahanshahlou, F arid; Akbarzadeh, Mohammad Amin; Zarei, Mahdi; V aez-Gharamaleki, Y osra. F orm ulating research questions for evidence-based studies. Journal of me dicine, sur gery, and public he alth 2 , 100046 (2024). 26 [14] Armeni, P atrizio; Polat, Irem; De Rossi, Leonardo Maria; Diaferia, Lorenzo; Meregalli, Sev- erino; Gatti, Anna. Digital t wins in healthcare: is it the b eginning of a new era of evidence-based medicine? A critical review. Journal of p ersonalize d me dicine 12 (8), 1255 (2022). [15] Jain, Ritu; Subramanian, Janakiraman; Rathore, Anurag S. A review of therap eutic failures in late-stage clinical trials. Exp ert Opinion on Pharmac other apy 24 (3), 389–399 (2023). [16] Sun, Duxin; Gao, W ei; Hu, Hongxiang; Zhou, Simon. Why 90% of clinical drug developmen t fails and ho w to improv e it? A cta Pharmac eutic a Sinic a B 12 (7), 3049–3062 (2022). [17] Hanahan, Douglas. Hallmarks of cancer: new dimensions. Canc er disc overy 12 (1), 31–46 (2022). [18] K ontomanolis, Emman uel N; K outras, Antonios; Syllaios, Athanasios; Sc hizas, Dimitrios; Mas- toraki, Aik aterini; Garmpis, Nikolaos; Diakosa vv as, Michail; Angelou, Kyveli; T satsaris, Geor- gios; Pagk alos, Athanasios; et al. Role of oncogenes and tumor-suppressor genes in carcinogen- esis: a review. A ntic anc er r ese ar ch 40 (11), 6009–6015 (2020). [19] Amir-Behghadami, Mehrdad; Janati, Ali. P opulation, Interv ention, Comparison, Outcomes and Study (PICOS) design as a framework to formulate eligibility criteria in systematic reviews. Emer gency Me dicine Journal (2020). [20] Bro wn, David. A review of the PubMed PICO to ol: using evidence-based practice in health education. He alth pr omotion pr actic e 21 (4), 496–498 (2020). [21] Sutton, Reed T; Pinco c k, David; Baumgart, Daniel C; Sadowski, Daniel C; F edorak, Ric hard N; Kro eker, Karen I. An ov erview of clinical decision supp ort systems: b enefits, risks, and strategies for success. NPJ digital me dicine 3 (1), 17 (2020). [22] Musen, Mark A; Middleton, Blackford; Greenes, Rob ert A. Clinical Decision-Supp ort Sys- tems. In Biome dic al Informatics: Computer Applic ations in He alth Car e and Biome dicine , pp. 795–840 (Springer In ternational Publishing, Cham, 2021). https://doi.org/10.1007/ 978- 3- 030- 58721- 5_24 . [23] La vecc hia, Antonio. Explainable artificial intelligence in drug disco very: bridging predictive p o w er and mechanistic insigh t. Wiley Inter disciplinary R eviews: Computational Mole cular Sci- enc e 15 (5), e70049 (2025). [24] Pham, Thai-Hoang; Qiu, Y ue; Zeng, Juc heng; Xie, Lei; Zhang, Ping. A deep learning framework for high-throughput mec hanism-driven phenotype compound screening and its application to CO VID-19 drug repurp osing. Natur e machine intel ligenc e 3 (3), 247–257 (2021). [25] Mohamed, S., No v áček, V. & Noun u, A. Disco vering protein drug tar- gets using knowledge graph em b eddings. Bioinformatics . 36 , 603-610 (2019,8), h ttps://doi.org/10.1093/bioinformatics/btz600 [26] Barabási, Alb ert-László; Gulbahce, Natali; Loscalzo, Joseph. Netw ork medicine: a netw ork- based approac h to human disease. Natur e r eviews genetics 12 (1), 56–68 (2011). [27] Buphamalai, Pisan u; Kok otovic, T omislav; Nagy , V anja; Menche, Jörg. Netw ork analysis re- v eals rare disease signatures across multiple levels of biological organization. Natur e c ommuni- c ations 12 (1), 6306 (2021). 27 [28] Cha, Junha; Lee, Insuk. Single-cell netw ork biology for resolving cellular heterogeneit y in hu- man diseases. Exp erimental & mole cular me dicine 52 (11), 1798–1808 (2020). [29] Li, Qing; Geng, Shan; Luo, Hao; W ang, W ei; Mo, Y a-Qi; Luo, Qing; W ang, Lu; Song, Guan- Bin; Sheng, Jian-Peng; Xu, Bo. Signaling pathw ays inv olved in colorectal cancer: pathogenesis and targeted therap y . Signal T r ansduction and T ar gete d Ther apy 9 (1), 266 (2024). [30] Zeng, Xuezhen; W ard, Simon E; Zhou, Jingying; Cheng, Alfred SL. Liver immune micro en- vironmen t and metastasis from colorectal cancer-pathogenesis and therap eutic p ersp ectiv es. Canc ers 13 (10), 2418 (2021). [31] Thiruna vuk arasu, Arun James; Ting, Darren Shu Jeng; Elango v an, Kabilan; Gutierrez, Laura; T an, Ting F ang; Ting, Daniel Shu W ei. Large language mo dels in medicine. Natur e me dicine 29 (8), 1930–1940 (2023). [32] Clusmann, Jan; Kolbinger, Fiona R; Muti, Hannah Sophie; Carrero, Zunamys I; Eck ardt, Jan- Niklas; Laleh, Narmin Ghaffari; Löffler, Chiara Maria La vinia; Sch warzk opf, Sophie-Caroline; Unger, Michaela; V eldhuizen, Gregory P; et al. The future landscap e of large language mo dels in medicine. Communic ations me dicine 3 (1), 141 (2023). [33] Liévin, V alen tin; Hother, Christoffer Egeb erg; Motzfeldt, Andreas Geert; Winther, Ole. Can large language mo dels reason ab out medical questions? Patterns 5 (3) (2024). [34] Singhal, Karan; Azizi, Shekoofeh; T u, T ao; Mahdavi, S Sara; W ei, Jason; Chung, Hyung W on; Scales, Nathan; T anw ani, Ajay; Cole-Lewis, Heather; Pfohl, Stephen; et al. Large language mo dels enco de clinical kno wledge. Natur e 620 (7972), 172–180 (2023). [35] Song, Bosheng; Li, F en; Liu, Y uansheng; Zeng, Xiangxiang. Deep learning metho ds for biomed- ical named entit y recognition: a surv ey and qualitativ e comparison. Briefings in Bioinformatics 22 (6), bbab282 (2021). [36] Sung, Mujeen; Jeong, Min byul; Choi, Y ongh wa; Kim, Dongh yeon; Lee, Jinh yuk; Kang, Jaewoo. BERN2: an adv anced neural biomedical named en tity recognition and normalization to ol. Bioinformatics 38 (20), 4837–4839 (2022). [37] T ruhn, Daniel; Reis-Filho, Jorge; Kather, Jakob. Large language mo dels should b e used as scien tific reasoning engines, not kno wledge databases. Natur e Me dicine 29 (2023). https:// doi.org/10.1038/s41591- 023- 02594- z . [38] Dagdelen, John; Dunn, Alexander; Lee, Sangho on; W alker, Nicholas; Rosen, Andrew S; Ceder, Gerbrand; P ersson, Kristin A; Jain, Anubha v. Structured information extraction from scientific text with large language mo dels. Natur e c ommunic ations 15 (1), 1418 (2024). [39] Nori, Harsha; King, Nic holas; McKinney , Scott Ma yer; Carignan, Dean; Horvitz, Eric. Capabil- ities of GPT-4 on Medical Challenge Problems. (2023). . [40] Agra wal, Monica; Hegselmann, Stefan; Lang, Hunter; Kim, Y o on; Son tag, Da vid. Large lan- guage mo dels are few-shot clinical information extractors. In Pr o c e e dings of the 2022 Confer- enc e on Empiric al Metho ds in Natur al L anguage Pr o c essing , pp. 1998–2022 (Asso ciation for Computational Linguistics, 2022). https://doi.org/10.18653/v1/2022.emnlp- main.130 . 28 [41] Kartc hner, David; Ramalingam, Selvi; Al-Hussaini, Irfan; Kronic k, Olivia; Mitc hell, Cassie. Zero-Shot Information Extraction for Clinical Meta-Analysis using Large Language Mo dels. In Pr o c e e dings of the 22nd W orkshop on Biome dic al Natur al L anguage Pr o c essing and BioNLP Shar e d T asks , pp. 396–405 (Asso ciation for Computational Linguistics, 2023). https://doi. org/10.18653/v1/2023.bionlp- 1.37 . [42] Hu, Danqing; Liu, Bing; Zhu, Xiaofeng; Lu, Xudong; W u, Nan. Zero-shot information extrac- tion from radiological rep orts using ChatGPT. International Journal of Me dic al Informatics 183 , 105321 (2024). [43] Chen, Da vid; Alnassar, Saif A ddeen; A vison, Kate Elizab eth; Huang, Ryan S; Raman, Srini- v as. Large language mo del applications for health information extraction in oncology: scoping review. JMIR c anc er 11 , e65984 (2025). [44] W ang, Andy; Liu, Cong; Y ang, Jingye; W eng, Chunh ua. Fine-tuning large language mo dels for rare disease concept normalization. Journal of the A meric an Me dic al Informatics Asso ciation 31 (9), 2076–2083 (2024). [45] Hertling, Sv en; Paulheim, Heiko. OLaLa: Ontology Matching with Large Language Mo dels. In Pr o c e e dings of the 12th Know le dge Captur e Confer enc e 2023 , pp. 131–139 (Asso ciation for Com- puting Machinery , New Y ork, NY, USA, 2023). https://doi.org/10.1145/3587259.3627571 . [46] Shang, Y ong; Tian, Y u; Lyu, Kewei; Zhou, Tianshu; Zhang, Ping; Chen, Jiangh ua; Li, Jingsong. Electronic health record–oriented kno wledge graph system for collab orativ e clinical decision supp ort using multicen ter fragmen ted medical data: design and application study . Journal of Me dic al Internet R ese ar ch 26 , e54263 (2024). [47] Jeong, Min byul; Sohn, Jiwoong; Sung, Mujeen; Kang, Jaewoo. Impro ving medical reasoning through retriev al and self-reflection with retriev al-augmented large language mo dels. Bioinfor- matics 40 (Supplemen t_1), i119–i129 (2024). [48] Zhao, Xuejiao; Liu, Siy an; Y ang, Su-Yin; Miao, Ch un yan. MedRAG: Enhancing Retriev al- augmen ted Generation with Kno wledge Graph-Elicited Reasoning for Healthcare Copilot. In Pr o c e e dings of the ACM on W eb Confer enc e 2025 , pp. 4442–4457 (Asso ciation for Computing Mac hinery , New Y ork, NY, USA, 2025). https://doi.org/10.1145/3696410.3714782 . [49] Sohn, Jiwoong; P ark, Y ein; Y o on, Chan woong; P ark, Sihy eon; Hw ang, Hyeon; Sung, Mujeen; Kim, Hyunjae; Kang, Jaewoo. Rationale-Guided Retriev al Augmented Generation for Medi- cal Question Answering. In Pr o c e e dings of the 2025 Confer enc e of the Nations of the Ameri- c as Chapter of the Asso ciation for Computational Linguistics: Human L anguage T e chnolo gies (V olume 1: L ong Pap ers) , pp. 12739–12753 (Asso ciation for Computational Linguistics, 2025). https://doi.org/10.18653/v1/2025.naacl- long.635 . [50] Y an, Lian; Guan, Yi; W ang, Haotian; Lin, Yi; Y ang, Y ang; W ang, Boran; Jiang, Jingchi. Eirad: An evidence-based dialogue system with highly in terpretable reasoning path for automatic diagnosis. IEEE Journal of Biome dic al and He alth Informatics 28 (10), 6141–6154 (2024). [51] Jin, Qiao; Dhingra, Bh uw an; Liu, Zhengping; Cohen, William; Lu, Xinghua. PubMedQA: A Dataset for Biomedical Research Question Answ ering. In Pr o c e e dings of the 2019 Confer enc e on Empiric al Metho ds in Natur al L anguage Pr o c essing and the 9th International Joint Con- fer enc e on Natur al L anguage Pr o c essing (EMNLP-IJCNLP) , pp. 2567–2577 (Asso ciation for Computational Linguistics, 2019). https://doi.org/10.18653/v1/D19- 1259 . 29 [52] Krithara, Anastasia; Nen tidis, Anastasios; Bougiatiotis, Konstan tinos; P aliouras, Georgios. BioASQ-QA: A man ually curated corpus for Biomedical Question Answering. Scientific data 10 (1), 170 (2023). [53] DeY oung, Ja y; Lehman, Eric; Ny e, Benjamin; Marshall, Iain; W allace, Byron C. Evidence Inference 2.0: More Data, Better Mo dels. In Pr o c e e dings of the 19th SIGBioMe d W orkshop on Biome dic al L anguage Pr o c essing , pp. 123–132 (Association for Computational Linguistics, 2020). https://doi.org/10.18653/v1/2020.bionlp- 1.13 . [54] Gro ver, Adit ya; Lesko vec, Jure. no de2v ec: Scalable F eature Learning for Netw orks. In Pr o- c e e dings of the 22nd ACM SIGKDD International Confer enc e on Know le dge Disc overy and Data Mining , pp. 855–864 (Asso ciation for Computing Machinery , New Y ork, NY, USA, 2016). https://doi.org/10.1145/2939672.2939754 . [55] Kipf, Thomas N; W elling, Max. Semi-sup ervised classification with graph con volutional net- w orks. arXiv pr eprint arXiv:1609.02907 (2016). 30
</div>
<hr style="margin: 50px 0; border: 0; border-top: 2px solid #eee;" />
<!-- ── Original Paper Viewer ── -->
<section class="original-paper-section" id="paper-viewer-anchor">
<div style="display: flex; justify-content: space-between; align-items: center; margin-bottom: 25px;">
<h3 style="margin:0; font-size: 1.4rem; color: #222;">Original Paper</h3>
<div id="nav-top"></div>
</div>
<div id="paper-content-container" style="background: #f4f4f4; border: 1px solid #ddd; border-radius: 8px; min-height: 600px; position: relative; overflow: visible;">
<div id="loading-status" style="text-align: center; padding: 100px 20px; color: #888;">
<p>Loading high-quality paper...</p>
</div>
</div>
<div id="nav-bottom" style="margin-top: 30px; display: flex; justify-content: center;"></div>
</section>
<!-- ── Related Papers Section ── -->
<section id="related-papers-section" style="margin-top:60px; border-top:2px solid #eee; padding-top:40px;">
<h3 style="font-size:1.4rem; font-weight:800; color:#1a1a1a; margin-bottom:24px; display:flex; align-items:center; gap:10px;">
<svg width="22" height="22" viewBox="0 0 24 24" fill="none" stroke="#0366d6" stroke-width="2"><path d="M4 19.5A2.5 2.5 0 0 1 6.5 17H20"/><path d="M6.5 2H20v20H6.5A2.5 2.5 0 0 1 4 19.5v-15A2.5 2.5 0 0 1 6.5 2z"/></svg>
Related Papers
</h3>
<div id="related-papers-list" style="display:grid; grid-template-columns:repeat(auto-fill,minmax(260px,1fr)); gap:20px;">
<p style="color:#aaa; font-style:italic; font-size:0.9rem;">Loading...</p>
</div>
</section>
<!-- ── Comment Section ── -->
<section class="comments-section" style="margin-top: 80px; border-top: 3px solid #0366d6; padding-top: 50px;">
<h3 style="font-size: 1.6rem; font-weight: 800; color: #1a1a1a; margin-bottom: 35px; display: flex; align-items: center; gap: 12px;">
<svg width="28" height="28" viewBox="0 0 24 24" fill="none" stroke="#0366d6" stroke-width="2.5"><path d="M21 15a2 2 0 0 1-2 2H7l-4 4V5a2 2 0 0 1 2-2h14a2 2 0 0 1 2 2z"/></svg>
Comments & Academic Discussion
</h3>
<div id="comments-list" style="margin-bottom: 50px;">
<p style="color: #999; font-style: italic;">Loading comments...</p>
</div>
<div class="comment-form-wrap" style="background: #fdfdfd; padding: 35px; border-radius: 16px; border: 1px solid #e1e4e8; box-shadow: 0 4px 12px rgba(0,0,0,0.03);">
<h4 id="reply-title" style="margin-top: 0; margin-bottom: 20px; font-size: 1.2rem; font-weight: 800; color: #333;">Leave a Comment</h4>
<form id="comment-form" onsubmit="submitComment(event)">
<input type="hidden" id="parent-id" value="">
<div style="display: grid; grid-template-columns: 1fr 1fr; gap: 20px; margin-bottom: 20px;">
<input type="text" id="comment-author" placeholder="Your Name" required style="padding: 14px; border: 1px solid #ddd; border-radius: 8px; font-size: 1rem; outline: none; transition: border-color 0.2s;" onfocus="this.style.borderColor='#0366d6'" onblur="this.style.borderColor='#ddd'">
<input type="text" id="comment-website" placeholder="Website" style="display:none !important;" tabindex="-1" autocomplete="off">
</div>
<textarea id="comment-content" rows="5" placeholder="Share your insights or questions about this paper..." required style="width: 100%; padding: 14px; border: 1px solid #ddd; border-radius: 8px; margin-bottom: 20px; resize: vertical; box-sizing: border-box; font-size: 1rem; outline: none; transition: border-color 0.2s;" onfocus="this.style.borderColor='#0366d6'" onblur="this.style.borderColor='#ddd'"></textarea>
<div style="display: flex; justify-content: space-between; align-items: center;">
<button type="button" id="cancel-reply" onclick="resetReply()" style="display: none; background: #fff; border: 1px solid #d73a49; color: #d73a49; padding: 10px 20px; border-radius: 8px; font-weight: bold; cursor: pointer; transition: all 0.2s;">Cancel Reply </button>
<button type="submit" style="background: #0366d6; color: white; border: none; padding: 14px 40px; border-radius: 8px; font-weight: 800; cursor: pointer; font-size: 1rem; transition: background 0.2s; margin-left: auto;" onmouseover="this.style.background='#0056b3'" onmouseout="this.style.background='#0366d6'">Post Comment</button>
</div>
</form>
</div>
</section>
<script>
const arxivId = "2603.28325";
const apiUrl = "/api/comments/" + arxivId;
const pageLang = "en";
// ── Related Papers ────────────────────────────────────────
(function loadRelatedPapers() {
const container = document.getElementById('related-papers-list');
if (!container || !arxivId) return;
fetch('/api/related/' + encodeURIComponent(arxivId) + '?lang=' + pageLang)
.then(r => r.json())
.then(papers => {
if (!papers || papers.length === 0) {
document.getElementById('related-papers-section').style.display = 'none';
return;
}
container.innerHTML = papers.map(p => `
<a href="${p.url}" style="display:block; text-decoration:none; color:inherit; background:#fff; border:1px solid #e8e8e8; border-radius:10px; padding:16px; transition:box-shadow 0.2s;" onmouseover="this.style.boxShadow='0 4px 16px rgba(3,102,214,0.12)'" onmouseout="this.style.boxShadow='none'">
${p.image_url ? `<div style="aspect-ratio:16/9; overflow:hidden; border-radius:6px; margin-bottom:10px; background:#f5f5f5;"><img src="${p.image_url}" style="width:100%;height:100%;object-fit:cover;" loading="lazy" onerror="this.parentElement.style.display='none'"></div>` : ''}
<div style="font-size:0.72rem; color:#888; margin-bottom:5px;">${p.arxiv_id} · ${p.date_str}</div>
<div style="font-size:0.92rem; font-weight:700; color:#1a1a1a; line-height:1.4; display:-webkit-box; -webkit-line-clamp:2; -webkit-box-orient:vertical; overflow:hidden;">${p.title}</div>
</a>
`).join('');
})
.catch(() => {
document.getElementById('related-papers-section').style.display = 'none';
});
})();
function loadComments() {
fetch(apiUrl)
.then(res => res.json())
.then(data => {
const list = document.getElementById('comments-list');
if (!data || data.length === 0) {
list.innerHTML = `<div style="text-align:center; padding:40px; background:#fcfcfc; border-radius:12px; border:1px dashed #ddd; color:#999;">No comments yet. Be the first to share your thoughts!</div>`;
return;
}
// Group replies under parents
const top = data.filter(c => !c.parent_id);
const byParent = {};
data.filter(c => c.parent_id).forEach(c => {
byParent[c.parent_id] = byParent[c.parent_id] || [];
byParent[c.parent_id].push(c);
});
top.forEach(c => { c.replies = byParent[c.id] || []; });
list.innerHTML = top.map(renderComment).join('');
})
.catch(e => console.error('loadComments error:', e));
}
function renderComment(c) {
return `
<div class="comment-item" style="margin-bottom: 30px; border-left: 4px solid #0366d6; padding: 15px 25px; background: #fff; border-radius: 0 12px 12px 0; box-shadow: 0 2px 8px rgba(0,0,0,0.02);">
<div style="display: flex; justify-content: space-between; align-items: center; margin-bottom: 12px;">
<span style="font-weight: 800; color: #1a1a1a; font-size: 1.05rem;">${c.author}</span>
<span style="font-size: 0.85rem; color: #bbb;">${c.created_at ? c.created_at.slice(0,10) : ''}</span>
</div>
<p style="margin: 0; color: #4a4a4a; line-height: 1.7; font-size: 1.05rem; white-space: pre-wrap;">${c.content}</p>
<div style="margin-top: 15px;">
<button onclick="setReply(${c.id}, '${c.author.replace(/'/g, "\\'")}')" style="background:none; border:none; color:#0366d6; font-size:0.9rem; padding:0; cursor:pointer; font-weight:bold; display:flex; align-items:center; gap:5px;">
<svg width="14" height="14" viewBox="0 0 24 24" fill="none" stroke="currentColor" stroke-width="2.5"><path d="M15 10l-5 5 5 5"/><path d="M4 4v7a4 4 0 0 0 4 4h12"/></svg>
Reply
</button>
</div>
${c.replies && c.replies.length > 0 ? `<div style="margin-top: 25px; margin-left: 30px; border-top: 1px solid #f0f0f0; padding-top: 25px;">${c.replies.map(renderComment).join('')}</div>` : ''}
</div>
`;
}
function setReply(id, name) {
document.getElementById('parent-id').value = id;
document.getElementById('reply-title').innerText = `Reply to ${name}`;
document.getElementById('cancel-reply').style.display = 'inline-block';
document.getElementById('comment-content').focus();
document.getElementById('comment-form').scrollIntoView({ behavior: 'smooth', block: 'center' });
}
function resetReply() {
document.getElementById('parent-id').value = "";
document.getElementById('reply-title').innerText = "Leave a Comment";
document.getElementById('cancel-reply').style.display = 'none';
}
function submitComment(e) {
e.preventDefault();
const author = document.getElementById('comment-author').value;
const content = document.getElementById('comment-content').value;
const parent_id = document.getElementById('parent-id').value || null;
const website = document.getElementById('comment-website').value;
if (website) return; // honeypot
fetch(apiUrl, {
method: 'POST',
headers: { 'Content-Type': 'application/json' },
body: JSON.stringify({ author, content, parent_id })
}).then(res => {
if (res.ok) {
document.getElementById('comment-content').value = "";
resetReply();
loadComments();
alert("Comment posted successfully.");
} else {
alert("Error posting comment. Please try again.");
}
});
}
document.addEventListener('DOMContentLoaded', loadComments);
</script>
<script>
document.addEventListener("DOMContentLoaded", function() {
const arxivId = "2603.28325";
const container = document.getElementById('paper-content-container');
const loadingStatus = document.getElementById('loading-status');
let currentPage = 1;
let totalPages = 0;
const yearStr = "2026";
const monthStr = "03";
const paths = [
`/koineu_html/${yearStr}/${monthStr}/${arxivId}/index.html`
];
function updateNav() {
const navHtml = `
<div style="display: flex; align-items: center; gap: 15px; background: #fff; padding: 8px 25px; border-radius: 30px; border: 1px solid #ddd; box-shadow: 0 2px 8px rgba(0,0,0,0.05);">
<button onclick="movePage(-1)" ${currentPage === 1 ? 'disabled' : ''} style="border:0; background:none; cursor:pointer; font-weight:bold; color:${currentPage === 1 ? '#ccc' : '#0366d6'}">◀ Prev</button>
<span style="font-family: monospace; font-weight:bold; color:#333;">PAGE ${currentPage} / ${totalPages}</span>
<button onclick="movePage(1)" ${currentPage === totalPages ? 'disabled' : ''} style="border:0; background:none; cursor:pointer; font-weight:bold; color:${currentPage === totalPages ? '#ccc' : '#0366d6'}">Next ▶</button>
</div>
`;
document.getElementById('nav-top').innerHTML = navHtml;
document.getElementById('nav-bottom').innerHTML = navHtml;
}
window.movePage = function(delta) {
const next = currentPage + delta;
if (next >= 1 && next <= totalPages) {
document.getElementById('pf' + currentPage.toString(16)).style.display = 'none';
document.getElementById('pf' + next.toString(16)).style.display = 'block';
currentPage = next;
updateNav();
window.scrollTo({ top: document.getElementById('paper-viewer-anchor').offsetTop - 20, behavior: 'smooth' });
}
};
function tryLoad(idx) {
if (idx >= paths.length) {
loadingStatus.innerHTML = "<p>Original content is being processed. Available soon.</p>";
return;
}
const url = paths[idx];
const baseUrl = url.replace('index.html', '');
fetch(url).then(r => { if(!r.ok) throw new Error(); return r.text(); }).then(html => {
const parser = new DOMParser();
const doc = parser.parseFromString(html, 'text/html');
const pageContainer = doc.getElementById('page-container') || doc.body;
const styles = doc.querySelectorAll('style');
styles.forEach(s => {
const newStyle = document.createElement('style');
newStyle.textContent = s.textContent.replace(/url\((?!http|data|["']?\/)/g, `url(${baseUrl}`);
document.head.appendChild(newStyle);
});
if (pageContainer) {
const pages = pageContainer.querySelectorAll('.pf');
totalPages = pages.length || 1;
pageContainer.style.cssText = "position:relative !important; top:0 !important; left:0 !important; width:100% !important; display:block !important;";
pages.forEach((p, i) => {
p.style.display = (i === 0) ? 'block' : 'none';
p.style.cssText += "position:relative !important; margin:0 auto !important; max-width:100% !important; background:white !important; box-shadow:0 0 15px rgba(0,0,0,0.1) !important;";
p.querySelectorAll('img').forEach(img => {
const src = img.getAttribute('src');
if (src && !src.startsWith('http') && !src.startsWith('/')) img.src = baseUrl + src;
img.onerror = function() { this.style.display = 'none'; };
});
});
container.innerHTML = "";
container.appendChild(pageContainer);
updateNav();
}
}).catch(() => tryLoad(idx + 1));
}
tryLoad(0);
});
</script>
<script>
document.addEventListener("DOMContentLoaded", function() {
const arxivId = "2603.28325";
const lang = "en";
// Record View & Update Count
fetch(`/api/view/${arxivId}?lang=${lang}`, { method: 'POST' })
.then(res => res.json())
.then(data => {
if (data.status === 'success' && data.view_count !== undefined) {
const viewEl = document.getElementById('post-view-number');
if (viewEl) viewEl.innerText = Number(data.view_count).toLocaleString();
}
})
.catch(e => console.error(e));
// Load Sidebar Data
fetch(`/api/sidebar-data?lang=${lang}`)
.then(res => res.json())
.then(data => {
// Popular Posts
const popContainer = document.getElementById('dynamic-sidebar-popular');
if (data.popular_posts && data.popular_posts.length > 0) {
let popHtml = `<h4 class="sidebar-section-title">${lang === 'kr' ? '인기 게시물' : 'Popular Posts'}</h4>`;
data.popular_posts.forEach(p => {
popHtml += `
<a href="${p.url}" class="sidebar-post">
<div class="sidebar-post__img">
<img src="${p.image_url}" onerror="this.src='/images/placeholder.jpg'" alt="${p.title}" loading="lazy" />
</div>
<span class="sidebar-post__title">${p.title}</span>
</a>
`;
});
popContainer.innerHTML = popHtml;
} else {
popContainer.style.display = 'none';
}
// Recent Comments
const commentContainer = document.getElementById('dynamic-sidebar-comments');
if (data.recent_comments && data.recent_comments.length > 0) {
let cHtml = `<h4 class="sidebar-section-title">${lang === 'kr' ? '최근 댓글' : 'Recent Comments'}</h4><div style="display:flex; flex-direction:column; gap:15px;">`;
data.recent_comments.forEach(c => {
cHtml += `
<a href="${c.url}#comments-list" style="text-decoration:none; background:#f8f9fa; padding:12px; border-radius:8px; display:block; border:1px solid #eee; transition:background 0.2s;" onmouseover="this.style.background='#f0f7ff'" onmouseout="this.style.background='#f8f9fa'">
<div style="font-size:0.85rem; color:#666; margin-bottom:5px;"><strong>${c.author}</strong> on <span style="color:#0366d6;">${c.post_title}</span></div>
<div style="font-size:0.95rem; color:#333; line-height:1.4;">"${c.content}"</div>
</a>
`;
});
cHtml += `</div>`;
commentContainer.innerHTML = cHtml;
} else {
commentContainer.style.display = 'none';
}
})
.catch(e => {
document.getElementById('dynamic-sidebar-popular').style.display = 'none';
document.getElementById('dynamic-sidebar-comments').style.display = 'none';
});
});
</script>
<div class="post-share" style="margin-top: 50px;">
<a href="https://twitter.com/intent/tweet?url=http://www.koineu.com/en/posts/2026/03/2026-03-30-2603_28325&text=Building%20evidence-based%20knowledge%20graphs%20from%20full-text%20literature%20for%20disease-specific%20biomedical%20reasoning" class="share-btn" target="_blank" rel="noopener">Twitter</a>
<a href="https://www.facebook.com/sharer/sharer.php?u=http://www.koineu.com/en/posts/2026/03/2026-03-30-2603_28325" class="share-btn" target="_blank" rel="noopener">Facebook</a>
</div>
</div>
</div>
</div>
</main>
<footer class="site-footer">
<div class="container">
<div class="footer-grid">
<div>
<div class="footer-logo">KOINEU</div>
<p class="footer-desc">Global Academic Research Archive powered by AI.</p>
<div class="footer-biz-sidebar" style="margin:20px 0; font-size:0.85rem; color:var(--color-muted); line-height:1.7;">
<div style="display:flex; gap:8px;">
<span style="white-space:nowrap;"><b>Company:</b></span>
<span>미스미스터크레이지 (MissMrCrazy)</span>
</div>
<div style="display:flex; gap:8px;">
<span style="white-space:nowrap;"><b>CEO:</b></span>
<span>송호성 (Song Ho-seong)</span>
</div>
<div style="display:flex; gap:8px;">
<span style="white-space:nowrap;"><b>Biz Reg:</b></span>
<span>731-64-00881</span>
</div>
</div>
</div>
<div>
<h3 class="footer-title">Recent posts</h3>
<div class="footer-posts-list">
<a href="/en/posts/2026/03/2026-03-31-2603_29298/" class="footer-post">
<div class="footer-post__img">
<img src="/koineu_html/2026/03/2603.29298/bge.webp" onerror="this.src='/images/placeholder.jpg'" alt="thumb" loading="lazy">
</div>
<div>
<div class="footer-post__title" style="color:white;">
Machine Learning Assisted Reconstruction of Local Electronic Structure of Non-Uniformly Strained MoS2
</div>
<div class="footer-post__meta" style="font-size:0.75rem; color:#aaa; margin-top:3px;">2026-03-31</div>
</div>
</a>
<a href="/en/posts/2026/03/2026-03-31-2603_29822/" class="footer-post">
<div class="footer-post__img">
<img src="/koineu_html/2026/03/2603.29822/bg3.webp" onerror="this.src='/images/placeholder.jpg'" alt="thumb" loading="lazy">
</div>
<div>
<div class="footer-post__title" style="color:white;">
Conditional Diffusion-Based Point Cloud Imaging for UAV Position and Attitude Sensing
</div>
<div class="footer-post__meta" style="font-size:0.75rem; color:#aaa; margin-top:3px;">2026-03-31</div>
</div>
</a>
</div>
</div>
<div>
<h3 class="newsletter-title">STAY INFORMED</h3>
<p class="newsletter-desc">
Get the latest research breakthroughs delivered to your inbox.
</p>
<form class="newsletter-form" action="#" method="post">
<input type="email" name="email" class="newsletter-input" placeholder="Email Address" required />
<button type="submit" class="newsletter-btn">SUBSCRIBE</button>
</form>
</div>
</div>
<div class="footer-bottom" style="margin-top:50px; padding-top:30px; border-top:1px solid var(--color-border);">
<div style="display:flex; flex-direction:column; gap:10px;">
<div style="display:flex; flex-wrap:wrap; gap:20px; font-size:0.75rem; color:var(--color-muted);">
<span>2026 © KOINEU. All Rights Reserved.</span>
<a href="/en/about/" style="color:var(--color-muted); text-decoration:none;">About Us</a>
<a href="/en/terms/" style="color:var(--color-muted); text-decoration:none;">Terms</a>
<a href="/en/privacy/" style="color:var(--color-muted); text-decoration:none;">Privacy Policy</a>
</div>
<div style="font-size:0.7rem; color:#aaa; line-height:1.5;">
Address: 31, Hyangsoseojeong-gil, Danwol-myeon, Yangpyeong-gun, Gyeonggi-do, KR | Industry: Information & Communication
</div>
</div>
<a href="#" class="back-to-top" style="display:flex; align-items:center; gap:6px; color:var(--color-muted); font-size:0.8125rem; margin-top:auto; text-decoration:none;">
<svg width="12" height="12" viewBox="0 0 24 24" fill="none" stroke="currentColor" stroke-width="2.5"><path d="M18 15l-6-6-6 6"/></svg>
Back to top
</a>
</div>
</div>
</footer>
<script>
// ── 모바일 메뉴 토글 ──
const navToggle = document.querySelector('.nav-toggle');
const navMenu = document.querySelector('.nav-menu');
if (navToggle && navMenu) {
navToggle.addEventListener('click', () => {
const expanded = navToggle.getAttribute('aria-expanded') === 'true';
navToggle.setAttribute('aria-expanded', String(!expanded));
navMenu.classList.toggle('is-open');
});
}
// ── 검색 패널 ──
const searchBtn = document.getElementById('navSearchToggle');
const searchPanel = document.getElementById('searchPanel');
const searchClose = document.getElementById('searchClose');
if (searchBtn && searchPanel) {
searchBtn.addEventListener('click', () => {
searchPanel.style.display = 'flex';
searchPanel.querySelector('input[name="q"]')?.focus();
});
searchClose?.addEventListener('click', () => { searchPanel.style.display = 'none'; });
searchPanel.addEventListener('click', (e) => {
if (e.target === searchPanel) searchPanel.style.display = 'none';
});
document.addEventListener('keydown', (e) => {
if (e.key === 'Escape') searchPanel.style.display = 'none';
});
}
// ── Back to top ──
document.querySelector('.back-to-top')?.addEventListener('click', (e) => {
e.preventDefault();
window.scrollTo({ top: 0, behavior: 'smooth' });
});
</script>
</body>
</html>