Q-BIOLAT: Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization
Protein fitness optimization is inherently a discrete combinatorial problem, yet most learning-based approaches rely on continuous representations and are primarily evaluated through predictive accuracy. We introduce Q-BIOLAT, a framework for modelin…
Authors: Truong-Son Hy
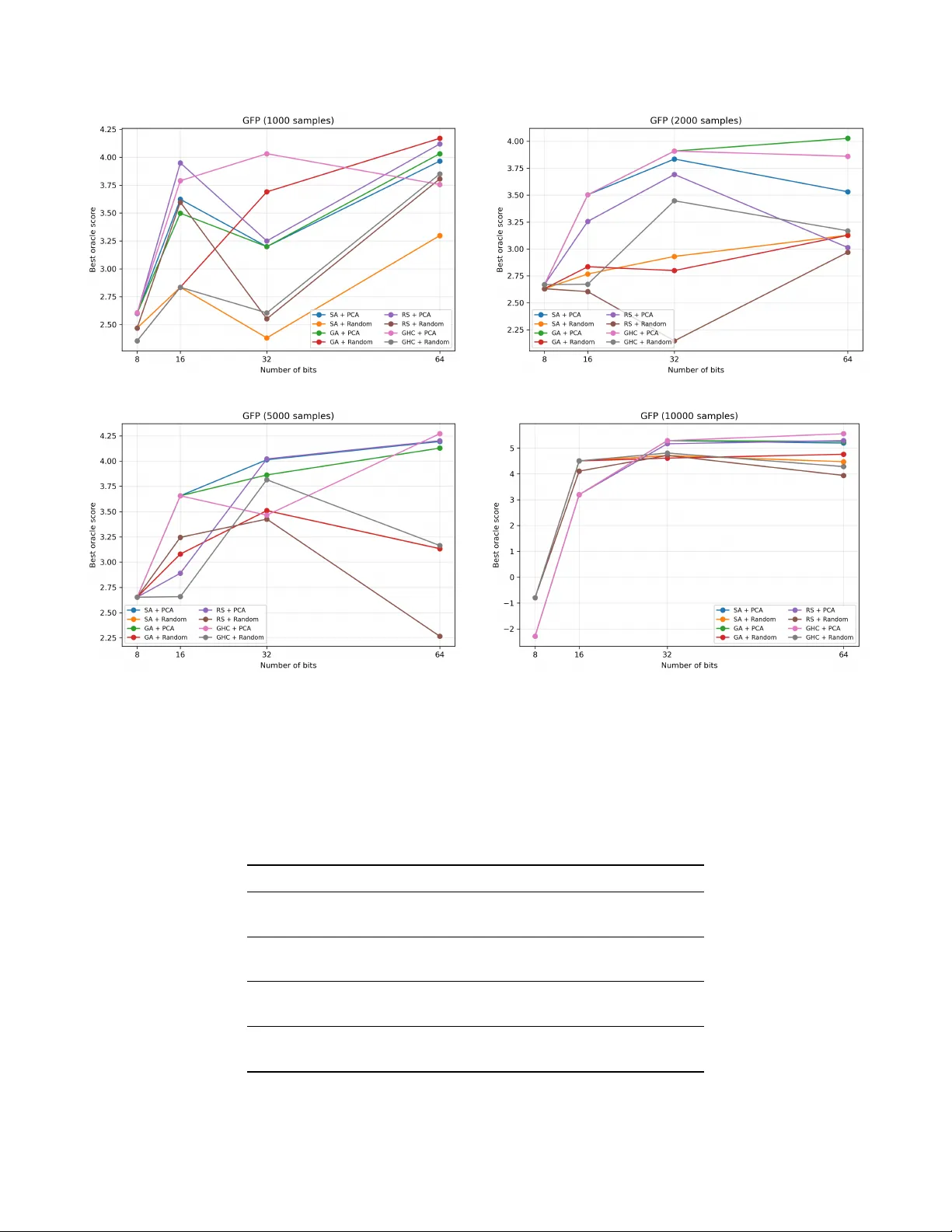
Q-BioLa t : Binar y Latent Protein Fitness Landscapes for QUBO-Based Optimization Truong-Son Hy University of Alabama at Birmingham Birmingham, Alabama, USA thy@uab.edu Abstract Protein tness optimization is inher ently a discrete combi- natorial problem, yet most learning-based appr oaches rely on continuous representations and are primarily evaluated through predictive accuracy . W e introduce Q-BioLa t , a frame- work for modeling and optimizing protein tness landscapes in compact binary latent spaces. Starting from pretrained protein language model embeddings, we construct binary latent representations and learn a quadratic unconstrained binary optimization (QUBO) surrogate that captures unary and pairwise interactions. Beyond its formulation, Q-BioLa t provides a representation- centric perspective on protein tness modeling. W e show that representations with similar predictive p erformance can induce fundamentally dierent optimization landscapes. In particular , learned autoencoder-based representations col- lapse after binarization, producing degenerate latent spaces that fail to support combinatorial search, whereas simple structured representations such as PCA yield high-entropy , decodable, and optimization-friendly latent spaces. Across multiple datasets and data regimes, we demon- strate that classical combinatorial optimization methods, in- cluding simulated annealing, genetic algorithms, and greedy hill climbing, are highly eective in structured binar y latent spaces. By expressing the objective in QUBO form, our ap- proach connects modern machine learning with discrete and quantum-inspired optimization. Our implementation and dataset are publicly available at: hps://github.com/HySonLab/Q- BIOLA T - Extended . Ke ywords: Protein Fitness Landscapes, Protein Language Models, Binary Latent Representations, QUBO Optimization, Combinatorial Optimization, Simulated Annealing, Quan- tum Annealing, Protein Engineering, Bioinformatics. 1 Introduction Understanding and optimizing protein tness landscap es is a central challenge in computational biology , with ap- plications spanning enzyme engineering, drug discovery , and synthetic biology . Protein tness landscap es are typ- ically high-dimensional, rugged, and shaped by comple x interactions between residues, making ecient exploration of sequence space dicult [ 1 – 3 ]. Recent advances in pro- tein language models (PLMs), such as ESM-2 [ 4 ] and ESM-3 [ 5 ], have enabled powerful representations of protein se- quences by learning from large-scale unlabeled data. These representations have signicantly improved performance on downstream tasks, including mutation eect prediction [6, 7] and protein design [8–10]. Despite these advances, most existing approaches rely on continuous neural predictors and gradient-based [ 9 ] or sampling-based optimization strategies [ 10 ], which are not naturally suited for discrete combinatorial search. In partic- ular , protein sequences are inherently discrete objects, and many biologically relevant optimization pr oblems are funda- mentally combinatorial in nature. Classical approaches such as evolutionary algorithms [ 10 , 11 ] and Bayesian optimiza- tion [ 12 , 13 ] have been widely applied to protein engineer- ing, but they often struggle with scalability or require large numbers of e valuations. Mor e importantly , existing methods primarily emphasize predictive accuracy , while the r ole of the underlying representation in shaping the optimization landscape remains under explored. In this work, w e introduce Q-BioLa t , a framework that maps protein sequences into binary latent spaces where t- ness landscapes can b e explicitly mo deled and optimized (see Figure 1). Starting from pr etrained protein language model embeddings, we construct compact binary representations that enable the formulation of protein tness as a quadratic unconstrained binary optimization (QUBO) [ 14 – 16 ] prob- lem. This formulation transforms protein tness modeling into a structured energy landscape o ver binar y variables, allowing us to leverage a wide range of combinatorial op- timization techniques, including simulated annealing and genetic algorithms, for ecient exploration of protein tness landscapes. Beyond its algorithmic formulation, Q-BioLa t pr ovides a representation-centric perspective on protein tness mod- eling through the lens of induced optimization landscapes. W e show that representations with similar predictive perfor- mance can exhibit markedly dierent optimization behavior , as determined by the geometry of the latent space and the structure of the corresponding QUBO interactions. In par- ticular , we obser ve a clear distinction between prediction quality and optimization eectiveness : learne d autoencoder- based representations collapse after binarization, producing degenerate latent codes that fail to support combinatorial search, whereas simple structured representations such as Truong-Son Hy Figure 1. O verview of the Q-BioLa t framework. Protein sequences are rst encoded using a pretrained protein language model (ESM) to obtain continuous embeddings. These emb eddings are transformed into binar y latent representations through projection and binarization, enabling protein tness to be mo deled as a quadratic unconstrained binar y optimization (QUBO) problem. The resulting latent tness landscape can be explored using combinatorial optimization methods and is directly compatible with quantum annealing hardware . Optimized latent codes are mapped back to high-tness protein sequences. PCA yield high-entropy and optimization-friendly latent spaces. Empirically , we demonstrate that PCA -based binary latent representations consistently outperform both random pro- jections and learne d AE/V AE representations in deco ding quality and end-to-end protein design performance across multiple datasets and data regimes. Furthermore , classical combinatorial optimization methods, including simulated annealing, genetic algorithms, and greedy hill climbing, are highly eective when applied to these structured latent land- scapes, reliably identifying high-tness regions even in data- scarce settings. W e also obser ve that optimization p erfor- mance depends on latent dimensionality , revealing a trade- o between expressivity , generalization, and searchability . Overall, Q-BioLa t bridges protein language modeling, discrete latent representation learning, and combinatorial optimization by reframing protein tness modeling as the construction of an optimization-aware energy landscape. By expressing protein tness landscapes as QUBO problems, our frame work provides a natural interface between mo dern machine learning models and discrete optimization meth- ods, while also oering a pathway toward integration with quantum and quantum-inspired optimization paradigms. 2 Related W ork Protein language models and sequence representations. Recent advances in protein language models (PLMs) have signicantly impro ved the representation of biological se- quences by leveraging large-scale unlabele d protein data. Models such as ESM-2 [ 4 ] and its successor ESM-3 [ 5 ] learn contextual embeddings that capture structural and func- tional properties of proteins directly from sequence data. More recently , multimodal protein language models have been developed to integrate sequence information with ad- ditional modalities such as structure, function, and evolu- tionary context, further enhancing representation quality and downstream performance [ 17 , 18 ]. These models have demonstrated strong p erformance across a wide range of tasks, including structure prediction, mutation eect predic- tion, and protein design [ 19 , 20 ]. In this work, we build upon PLM embeddings as a foundation, but transform them into binary latent representations to enable discrete optimization. Protein tness prediction and engineering. Predicting protein tness landscapes is a fundamental problem in com- putational biology , with applications in enzyme optimization, therapeutic design, and synthetic biology . Recent approaches combine machine learning models with experimental data, particularly from deep mutational scanning (DMS) exper- iments, to predict the ee cts of mutations [ 21 , 22 ]. Large- scale benchmarks such as Pr oteinGym hav e been introduced to systematically evaluate tness pr ediction models across diverse proteins and mutation regimes [ 3 ]. While these meth- ods achieve strong predictive performance, they typically rely on continuous repr esentations and do not directly ad- dress the combinatorial nature of sequence optimization. Furthermore, these approaches primarily focus on predic- tion rather than explicitly enabling structured optimization over discrete sequence spaces. Optimization metho ds for protein design. A variety of optimization strategies have been applied to protein engi- neering, including evolutionary algorithms [ 9 – 11 , 13 ], Bayesian optimization [ 12 , 13 ], and reinforcement learning [ 23 , 24 ]. Evolutionary methods such as genetic algorithms are widely used due to their ability to explore large discr ete search spaces, while Bayesian optimization oers sample-ecient search under expensive evaluation settings. Classical opti- mization techniques such as simulated annealing have also been applie d to combinatorial problems with rugged energy landscapes [ 25 , 26 ]. Howev er , many of these approaches op- erate directly in sequence space or continuous embedding spaces, rather than explicitly modeling pr otein tness as a structured combinatorial optimization problem. In contrast, our approach explicitly models protein tness as a structured Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization combinatorial optimization problem in a learned binary la- tent space. QUBO formulations and quantum annealing. Quadratic unconstrained binar y optimization (QUBO) [ 14 – 16 ] provides a unied framework for e xpressing a wide range of combi- natorial problems in terms of binary variables and pair wise interactions. QUBO formulations are closely r elated to the Ising model and form the basis of many optimization tech- niques, including quantum annealing [ 15 ]. Quantum anneal- ing hardware , such as that developed by D- W ave Systems, has demonstrated the ability to solve certain classes of opti- mization problems by exploiting quantum eects [ 27 ]. While applications of quantum optimization in computational bi- ology remain limited, recent work has begun to explore its potential for tasks such as molecular design and protein fold- ing. In this work, we formulate protein tness landscapes as QUBO problems in a binar y latent space, enabling di- rect compatibility with both classical combinatorial solvers and emerging quantum annealing hardware , and providing a unie d persp ective on protein design as discrete energy landscape optimization. Latent space optimization and discrete representations. Optimizing in learned latent spaces has become a common strategy in machine learning, particularly for generative modeling and design tasks [ 28 , 29 ]. Continuous latent spaces have been widely used for molecule and protein generation [ 9 , 30 , 31 ], but they often require gradient-base d optimization or sampling methods that do not naturally handle discr ete constraints. More recently , discrete and binary latent repre- sentations have been explored for enabling combinatorial search and ecient optimization [ 32 , 33 ]. Our work builds on this idea by constructing binary latent representations de- rived from protein language models and explicitly mo deling the resulting tness landscape as a QUBO problem, bridging representation learning and combinatorial optimization. Energy landscapes and combinatorial optimization. Many combinatorial optimization problems can be inter- preted through the lens of energy landscapes, where the objective function denes a surface over discrete congura- tions and optimization corresponds to navigating this land- scape toward lo w-energy (or high-tness) regions [ 34 , 35 ]. In particular , QUBO formulations are closely related to Ising models and have been widely studied in statistical physics as models of interacting binary systems [ 15 ]. The structure of the interaction matrix determines key properties of the land- scape, such as smoothness, ruggedness, and the distribution of local optima, which in turn str ongly inuence optimiza- tion diculty [ 36 , 37 ]. Such energy landscape perspectives have also been explored in machine learning, particularly in the analysis of non-conve x optimization and neural net- work loss surfaces [ 37 , 38 ]. While these ideas are well es- tablished in physics and optimization, their role in modern representation learning pipelines for biological sequence design remains underexplored. In this work, we adopt an energy landscape view of protein tness mo deling in binary latent spaces and study how r epresentation-induced struc- ture aects combinatorial search. Representation learning for optimization. Most e xisting representation learning approaches in protein modeling and related domains focus on improving predictive performance on downstream tasks, such as regression or classication [ 6 , 7 , 19 ]. Howev er , in design and optimization settings, the learned representation implicitly denes a search space and induces an optimization landscape. Recent work in latent space optimization has explored how learned embeddings can facilitate search in applications such as molecule design, protein engineering, and combinatorial optimization [ 39 – 42 ]. These approaches typically leverage continuous latent spaces learned by variational autoencoders or related generative models, enabling gradient-based or sampling-based search. In protein design, similar ideas have been explored through latent generative models and language-model-guided opti- mization [ 8 – 10 ], where representations are used to guide exploration of sequence space. Ho wever , these approaches are primarily evaluated through predictive metrics, likeli- hood, or sample quality , rather than explicit analysis of the induced optimization landscape. More broadly , recent work in machine learning has begun to study how representations inuence optimization and search behavior [ 43 , 44 ], but this perspective remains underdeveloped in the context of bio- logical sequence design. In contrast, our work emphasizes that representation quality should also be assessed base d on the properties of the induce d optimization landscap e, and demonstrates that dierent representations with sim- ilar predictive accuracy can lead to substantially dierent optimization outcomes. 3 Method 3.1 Overview W e propose Q-BioLa t , a framew ork for modeling and op- timizing protein tness landscapes in a binary latent space. Given a protein sequence, Q-BioLa t rst computes a pre- trained protein language model embedding, then transforms this continuous representation into a compact binary latent code. In this binary space, the tness landscape is approx- imated by a quadratic unconstrained binar y optimization (QUBO) surr ogate, which models both unary and pairwise eects among latent variables. The learned surr ogate is then optimized using combinatorial search methods such as sim- ulated annealing and genetic algorithms. Because the nal Truong-Son Hy objective is expressed in QUBO form, the framework is nat- urally compatible with both classical combinatorial solvers and quantum annealing hardware . Formally , the overall pipeline is 𝑠 → 𝑒 ( 𝑠 ) ∈ R 𝑑 → 𝑧 ( 𝑠 ) ∈ R 𝑚 → 𝑥 ( 𝑠 ) ∈ { 0 , 1 } 𝑚 → ˆ 𝑓 ( 𝑥 ) , where 𝑠 is a protein sequence, 𝑒 ( 𝑠 ) is the protein language model embe dding, 𝑧 ( 𝑠 ) is a reduced continuous latent repr e- sentation, 𝑥 ( 𝑠 ) is the binarized latent code, and ˆ 𝑓 ( 𝑥 ) is the QUBO surrogate of protein tness. 3.2 Problem Setup Let D = { ( 𝑠 𝑖 , 𝑦 𝑖 ) } 𝑁 𝑖 = 1 denote a protein tness dataset, where 𝑠 𝑖 is a pr otein sequence and 𝑦 𝑖 ∈ R is its experimentally measured tness. Our goal is to learn a surr ogate model that maps each sequence to a compact binary latent represen- tation and predicts its tness, while also enabling ecient combinatorial optimization over the latent space. Unlike conventional approaches that optimize directly in sequence space or in continuous embedding space, we aim to transform protein tness prediction into a discrete optimiza- tion problem. This is motivate d by two obser vations: rst, protein sequences are inherently discrete objects; se cond, binary latent spaces admit ecient combinatorial search and can be directly mapped to QUBO and Ising formulations. 3.3 Protein Language Model Emb eddings For each protein sequence 𝑠 𝑖 , we obtain a continuous embed- ding using a pretrained protein language model. In our exper- iments, we use ESM-based sequence embeddings [ 4 , 5 ] due to their strong empirical performance and broad adoption in protein representation learning. Given a sequence 𝑠 𝑖 of length 𝐿 𝑖 , the language mo del produces contextualized residue-level representations 𝐻 𝑖 ∈ R 𝐿 𝑖 × 𝑑 , where 𝑑 is the hidden dimension of the model. T o obtain a xed-length sequence representa- tion, we apply mean pooling across residues: 𝑒 𝑖 = 1 𝐿 𝑖 𝐿 𝑖 𝑗 = 1 𝐻 ( 𝑗 ) 𝑖 ∈ R 𝑑 . This yields one dense embedding vector per se quence. The role of the protein language model in Q-BioLa t is not to directly predict tness, but rather to provide a biologically informed continuous representation that can later be com- pressed into a discrete latent code suitable for combinatorial optimization. 3.4 Continuous-to-Binary Latent Mapping A key component of Q-BioLa t is the transformation of con- tinuous protein embeddings into compact binary latent rep- resentations. While simple projection and thresholding pro- vide a lightweight baseline, more expressiv e representations can be obtained by learning the latent mapping in a data- driven manner . In this work, we consider a unie d family of approaches for constructing binary latent codes, ranging from linear projections to learned autoencoding models. Linear projection and binarization. Given a protein embedding 𝑒 ∈ R 𝑑 , we rst consider linear dimensionality reduction methods such as random projection or principal component analysis (PCA): 𝑧 = 𝑊 𝑒, 𝑊 ∈ R 𝑚 × 𝑑 , (1) where 𝑚 ≪ 𝑑 is the latent dimension. The resulting continu- ous latent vector 𝑧 ∈ R 𝑚 is then binarized using a threshold- ing function: 𝑥 𝑘 = I ( 𝑧 𝑘 > 𝜏 𝑘 ) , (2) where 𝜏 𝑘 is typically chosen as the median of the 𝑘 -th compo- nent over the training set. This appr oach provides a simple and ecient baseline for constructing binary latent repre- sentations. Deterministic binary autoencoder . T o learn a more struc- tured latent representation, we introduce a deterministic autoencoder [ 45 , 46 ] that maps protein emb eddings into a low-dimensional latent space and reconstructs them: 𝑧 = 𝑓 𝜃 ( 𝑒 ) , ˆ 𝑒 = 𝑔 𝜙 ( 𝑧 ) , (3) where 𝑓 𝜃 and 𝑔 𝜙 denote the encoder and decoder networks, respectively . T o obtain binary latent co des, we apply a bina- rization function: 𝑥 = bin ( 𝑧 ) , (4) where bin ( ·) may be implemented using thresholding or a sign function. The model is trained to minimize reconstruc- tion loss: L AE = ∥ 𝑒 − ˆ 𝑒 ∥ 2 2 . (5) This formulation allows the latent space to adapt to the structure of protein embeddings, potentially leading to more meaningful and optimization-friendly representations. V ariational latent representations. W e further consider a probabilistic formulation based on variational auto encoders (V AEs) [ 47 ], which model a distribution over latent variables: 𝑧 ∼ 𝑞 𝜃 ( 𝑧 | 𝑒 ) , ˆ 𝑒 ∼ 𝑝 𝜙 ( 𝑒 | 𝑧 ) . (6) The model is trained by maximizing the evidence lower bound (ELBO): L V AE = E 𝑞 𝜃 ( 𝑧 | 𝑒 ) [ log 𝑝 𝜙 ( 𝑒 | 𝑧 ) ] − KL ( 𝑞 𝜃 ( 𝑧 | 𝑒 ) ∥ 𝑝 ( 𝑧 ) ) . (7) Binary latent co des can be obtaine d by applying a threshold or sampling from a Bernoulli distribution parameterized by the latent variables: 𝑥 𝑘 ∼ Bernoulli ( 𝜎 ( 𝑧 𝑘 ) ) . (8) This probabilistic formulation enables modeling uncertainty in the latent repr esentation and provides a principled way to explore the latent space. Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization Discussion. These approaches dene dierent ways of constructing binary latent spaces, ranging from simple lin- ear projections to learned deterministic and probabilistic mappings. Importantly , the choice of latent r epresentation directly aects the structure of the induce d QUBO model and, consequently , the geometry of the resulting optimiza- tion landscape. In Section 4, we empirically investigate how these dierent constructions inuence both predictive per- formance and optimization behavior . 3.5 QUBO Surrogate for Protein Fitness W e model protein tness in binary latent space with a QUBO surrogate. Given a binary latent code 𝑥 ∈ { 0 , 1 } 𝑚 , the pre- dicted tness is: ˆ 𝑓 ( 𝑥 ) = 𝑚 𝑘 = 1 ℎ 𝑘 𝑥 𝑘 + 1 ≤ 𝑘 < ℓ ≤ 𝑚 𝐽 𝑘 ℓ 𝑥 𝑘 𝑥 ℓ , where ℎ 𝑘 ∈ R captures the unar y contribution of latent bit 𝑘 , and 𝐽 𝑘 ℓ ∈ R captures the pairwise interaction between bits 𝑘 and ℓ . Equivalently , the model can be written in matrix form as: ˆ 𝑓 ( 𝑥 ) = ℎ ⊤ 𝑥 + 1 2 𝑥 ⊤ 𝐽 𝑥 , where 𝐽 ∈ R 𝑚 × 𝑚 is the symmetric Hamiltonian (i.e. pairwise interaction matrix) with zero diagonal, and ℎ ∈ R 𝑚 corre- sponds to the bias term. This representation connects the latent tness model to the classical QUBO formulation. Feature construction. For each binary code 𝑥 𝑖 , we con- struct a feature vector consisting of: • all linear terms { 𝑥 𝑖𝑘 } 𝑚 𝑘 = 1 , • all pair wise interaction terms { 𝑥 𝑖𝑘 𝑥 𝑖 ℓ } 1 ≤ 𝑘 < ℓ ≤ 𝑚 . The total number of features is therefore: 𝑚 + 𝑚 ( 𝑚 − 1 ) 2 . Parameter estimation. W e t the QUBO surrogate using ridge regression. Let Φ ( 𝑋 ) denote the design matrix formed by the linear and pairwise features of the training binar y codes, and let 𝑦 denote the corresponding tness vector . W e solve: 𝑤 ★ = arg min 𝑤 ∥ Φ ( 𝑋 ) 𝑤 − 𝑦 ∥ 2 2 + 𝜆 ∥ 𝑤 ∥ 2 2 , where 𝜆 > 0 is an ℓ 2 regularization coecient. The learned parameter vector 𝑤 ★ is then unpacked into the unary coe- cients ℎ and pairwise co ecients 𝐽 . This surrogate is attractive for three reasons. First, it is interpretable, since each term corresponds to a unar y or pairwise latent ee ct. Second, it is computationally ecient to t and evaluate for moderate latent dimensions. Third, it directly yields a QUBO objective suitable for classical and quantum combinatorial optimization. 3.6 Latent Space Optimization Once the QUBO surrogate has been tte d, we seek binary latent codes that maximize the predicted tness: 𝑥 ★ = arg max 𝑥 ∈ { 0 , 1 } 𝑚 ˆ 𝑓 ( 𝑥 ) . Because this is a discrete combinatorial problem, we employ search strategies that operate directly in the binar y latent space. Simulated annealing [ 25 ]. Simulated annealing starts from an initial latent code and iteratively proposes single-bit ips. Moves that impr ove the objective ar e accepted, while worse moves may be accepted with a temp erature-dependent probability , enabling escape from local optima. The temper- ature is gradually reduced during the search. Genetic algorithm [ 26 ]. The genetic algorithm main- tains a population of binary latent codes and evolves them through selection, crossover , and mutation. This allows broader exploration of the latent space and is particularly useful in higher-dimensional settings where multiple promising re- gions may exist. Baselines. T o contextualize performance, we also com- pare against gr eedy hill climbing, random search, and a light- weight latent Bayesian-style search baseline. Greedy hill climbing iteratively ips the single bit that most impr oves the objective until convergence. Random search samples bi- nary co des uniformly and retains the best candidate. The latent Bayesian-style metho d uses a kernel-based uncertainty heuristic over binary latent codes. 3.7 Decoding of Optimized Latent Representations A key step in Q-BioLa t is mapping optimized binar y latent codes back to valid protein sequences. Since optimization is performed in latent space , decoding pr ovides the conne ction between combinatorial search and biologically meaningful sequences. Retrieval-based decoding. As a conservative baseline, we rst use nearest-neighbor retrieval in Hamming space . Given an optimized latent code, w e identify the closest latent codes from the training set and return their associated sequences. This ensures that decoded se quences remain within the sup- port of the observed data distribution. Neural decoding. T o enable sequence generation beyond the observed dataset, we introduce a latent-conditioned mu- tation decoder . Instead of generating sequences autoregres- sively , the decoder predicts mutation patterns r elative to a wild-type sequence. For each p osition, the model predicts (i) whether a mutation occurs and (ii) the identity of the mu- tated amino acid. This formulation leverages the structur e of deep mutational scanning datasets, where sequences dier from a common backbone. Truong-Son Hy Discussion. W e nd that deco ding performance strongly depends on the structure of the latent space. In particu- lar , PCA -based binary representations produce signicantly more decodable latent codes than both random projections and AE/V AE repr esentations. These results emphasize that representation learning must be aligne d with both optimiza- tion and decoding objectives. 3.8 Quantum Annealing Compatibility An important property of Q-BioLa t is that its optimization objective is already expressed in QUBO form. This means the learned latent tness landscape can be optimized not only with classical combinatorial solvers, but also with quantum annealing hardware or other Ising/QUBO solvers without changing the mo del formulation. In this sense, Q-BioLa t provides a quantum-compatible interface between protein representation learning and discrete optimization. In the present work, we focus on classical solvers to es- tablish the feasibility of the framework and to analyze the optimization behavior in binary latent space. Nevertheless, the QUBO formulation opens a clear pathway toward future quantum-assisted protein engineering, where learned latent tness landscapes could be deploy ed on quantum annealers or related specialized optimization hardware. 3.9 Computational Complexity Let 𝑚 denote the binary latent dimension. The QUBO surro- gate uses 𝑚 + 𝑚 ( 𝑚 − 1 ) 2 = O ( 𝑚 2 ) features. Therefore , increas- ing latent dimensionality increases expressive p ower but also increases both the number of surrogate parameters and the diculty of combinatorial search. This trade-o motivates our experimental study over multiple latent dimensions. By separating the dense pr otein language model embed- ding stage from the binar y optimization stage, Q-BioLa t keeps the most expensive neural computation xed while enabling fast repeated optimization in the compact binary latent space. This makes the framew ork computationally attractive for studying representation–optimization trade- os. 4 Experiments Our experimental study is designed to answer three ques- tions: 1. First, can an external sequence-level oracle trained from protein language model embeddings provide a reliable black-box tness signal, especially in data- scarce settings? 2. Second, how do learned latent representations such as AE and V AE compare with simple projection-based binary codes for downstr eam optimization and de cod- ing? 3. Third, does the full Q-BioLa t pipeline—from sequence embedding to latent optimization and decoding—produce high-tness candidate proteins under a xed design budget? Consistent with the scope of this w ork, we place the main emphasis on end-to-end protein design performance, while using the landscape analysis as a secondary interpretive layer . 4.1 Datasets and Data-Scarce Regimes W e evaluate Q-BioLa t on protein tness landscap es from the ProteinGym benchmark [ 3 ]. Our primary experiments focus on two representative tasks: GFP and AA V . These two datasets provide complementary settings for studying both mutation-eect prediction and optimization o ver rugged protein tness landscapes. For each dataset, we construct subset sizes of { 1000 , 2000 , 5000 , 10000 } variants (see T able 1). W e treat the 1000 and 2000 settings as data-scarce regimes , and the 5000 and 10000 settings as mo derate-data regimes . Unless otherwise sp ecied, each subset is split into training, validation, and test sets using a stratie d split based on tness quantiles, with the validation split used for model sele ction and the test split reserved for nal reporting. All reported results are averaged ov er multiple random seeds. 4.2 External Sequence-Level Fitness Oracle A central component of this work is an external oracle that maps a protein sequence directly to a scalar tness esti- mate. This oracle is conceptually distinct from the internal Q-BioLa t QUBO surrogate. The external oracle serves two purposes: (i) it provides a standar dized black-box evaluator for comparing protein design methods that output sequences, and (ii) it enables evaluation of generate d sequences that do not exactly match an experimentally observed variant. For each protein sequence, we compute an ESM-based sequence embedding by mean po oling residue-lev el repre- sentations. On top of these embeddings, we train a family of classical regression models, including ridge regression, gradient-boosted decision trees (X GBo ost), and Gaussian process regression. All models are trained on the same em- bedding space to isolate the eect of the regression model rather than the upstream representation. Model selection is performed using a validation split, and the best-performing oracle for each dataset-size regime is use d in the downstream sequence-level optimization experiments. W e evaluate the external oracle using standard regression metrics, including Spearman correlation, Pearson correlation, root mean squared error (RMSE), and mean absolute err or (MAE). T able 2 summarizes the results across both GFP and AA V datasets, and Figure 2 visualizes the scaling behavior . Across both datasets, we obser ve a consistent and sys- tematic dependence of oracle performance on the amount of available data. In both low-data regimes (1000–2000 samples) and moderate-data regimes (5000–10000 samples), Gaussian Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization T able 1. ProteinGym datasets used in the experiments. W e consider two representative protein tness landscapes: GFP for uorescence-based tness and AA V capsid for viral tness. For each dataset, we construct multiple subset sizes to study data-scarce and moderate-data regimes. Dataset T ask W T length Full library size Subset sizes GFP Fluorescence / tness 238 51,714 1000, 2000, 5000, 10000 AA V Capsid / tness 735 42,328 1000, 2000, 5000, 10000 process regression achie ves the str ongest performance, re- ecting its ability to model uncertainty and adapt to limited supervision. As the dataset size increases, ridge regression becomes increasingly competitive and often achiev es near the b est performance, suggesting that the tness signal is largely captured by a linear model in the ESM embe dding space when sucient data is available. In contrast, X GBo ost underperforms in the smallest-data regime, indicating over- tting, but improves steadily with increasing dataset size. Based on these results, we use Gaussian process oracles in data-scarce settings when computationally feasible, and rely on ridge regression and X GBoost as scalable sequence-level evaluators in larger regimes. 4.3 Latent Representation Learning and Deco der Analysis W e compare four families of latent representations: (i) ran- dom projection with thresholding, (ii) PCA with threshold- ing, (iii) deterministic autoenco der (AE), and (iv ) variational autoencoder (V AE). All methods operate on the same ESM embeddings to isolate the eect of latent representation. Latent quality and collapse of learned representations. W e rst examine whether learned latent models produce usable binary codes after binarization. T able 3 reports recon- struction error , bit entropy , and the number of active latent dimensions. Although AE and V AE achieve v ery low recon- struction err or , their binarize d latent co des e xhibit near-zer o entropy and essentially no active dimensions. This indicates a collapse of the binary latent space , where most bits become constant across samples and fail to encode meaningful varia- tion. As a result, these representations do not provide a useful combinatorial search space for downstream optimization. Decoding performance of projection-base d represen- tations. W e next evaluate the deco dability of binar y la- tent codes using a mutation-conditioned deco der . T able 4 shows that pr ojection-based repr esentations, particularly PCA, signicantly outperform random projections in both mutation F1 and mutated-residue accuracy . For example, on AA V with 10000 training samples, PCA with 64 latent di- mensions achieves substantially higher decoding accuracy than random projection at the same dimensionality . Similar trends hold in the data-scarce regime on GFP . Scaling behavior with data and latent dimension. Fig- ure 3 further illustrates how decoding performance scales with the number of training samples at a xed latent dimen- sion of 64 bits. A cross both GFP and AA V datasets, PCA consistently achieves higher mutation F1 than random pro- jection, and p erformance improves as more data b ecomes available. This indicates that PCA not only preserves mean- ingful structure but also benets fr om additional sup ervision. Discussion. Taken together , these results r eveal a clear mismatch between reconstruction quality and optimization readiness. While AE and V AE reconstruct embeddings ac- curately , their binarize d latent spaces collapse and fail to support decoding or combinatorial search. In contrast, sim- ple structured representations such as PCA maintain both variability and decodability , leading to substantially better downstream performance. These ndings motivate our fo cus on PCA-based binary latent repr esentations in the end-to- end design benchmark. These observations also suggest that latent representations for protein design should b e evalu- ated not only by reconstruction or predictive accuracy , but by their ability to support b oth decoding and combinatorial optimization. 4.4 Internal QUBO Surrogate Performance Given a binar y latent representation, Q-BioLa t ts a qua- dratic surrogate over unary and pairwise latent features. This surrogate denes the QUBO objective used for combinatorial optimization. In this section, we evaluate how well dierent latent representations support the construction of such a surrogate. T able 5 reports predictive performance of the learned QUBO models using Spearman correlation on held-out test data. Importantly , this evaluation is not intended to estab- lish state-of-the-art tness prediction, but rather to assess whether the latent representation induces a meaningful op- timization landscape. Across representations, we obser ve that PCA, random pr o- jection, and even AE/V AE can achieve comparable predictive performance under the QUBO surrogate. In particular , dier- ences in Spearman correlation are often modest, indicating that multiple representations are capable of tting observed tness values to a similar degree. Howev er , when considered together with the results from Section 4.3 and the end-to-end b enchmark in Section 4.6, Truong-Son Hy T able 2. Performance of external sequence-level tness oracles built on top of ESM emb eddings across GFP and AA V datasets. Results are reported on the test set. The best Spearman correlation for each dataset and training size is highlighted in bold. Dataset Train size Oracle model Spearman ↑ Pearson ↑ RMSE ↓ MAE ↓ GFP 1000 Gaussian Process 0.657 0.682 0.759 0.612 Ridge Regression 0.637 0.665 0.859 0.652 XGBoost 0.520 0.557 0.863 0.711 2000 Gaussian Process 0.714 0.729 0.742 0.586 Ridge Regression 0.660 0.669 0.817 0.637 XGBoost 0.650 0.675 0.802 0.624 5000 Gaussian Process 0.729 0.754 0.699 0.542 Ridge Regression 0.702 0.697 0.772 0.607 XGBoost 0.635 0.685 0.772 0.596 10000 Gaussian Process 0.783 0.796 0.646 0.495 Ridge Regression 0.735 0.730 0.730 0.580 XGBoost 0.693 0.729 0.731 0.568 AA V 1000 Gaussian Process 0.671 0.650 2.390 1.906 Ridge Regression 0.706 0.656 2.618 2.058 XGBoost 0.611 0.607 2.501 2.005 2000 Gaussian Process 0.792 0.762 2.048 1.629 Ridge Regression 0.779 0.751 2.155 1.690 XGBoost 0.694 0.694 2.255 1.813 5000 Gaussian Process 0.806 0.782 1.898 1.505 Ridge Regression 0.796 0.763 1.962 1.547 XGBoost 0.729 0.724 2.094 1.659 10000 Gaussian Process 0.838 0.821 1.712 1.346 Ridge Regression 0.808 0.781 1.874 1.489 XGBoost 0.777 0.772 1.914 1.499 T able 3. Collapse of learned latent representations after binarization on GFP with 1000 training samples. Although AE and V AE achieve lo w reconstruction error , their binarized latent codes exhibit zero entrop y and no active dimensions. Representation Latent dim Bit entropy ↑ Active dims ↑ Recon. MSE ↓ AE 8 0.000 0.00 0.000125 64 0.000 0.00 0.000124 V AE 8 0.149 0.25 0.000126 64 0.002 0.00 0.000164 a clear discrepancy emerges b etween predictive accuracy and optimization eectiveness. Although AE and V AE can achieve reasonable surrogate ts, their collapsed binar y struc- ture prev ents eective decoding and downstream optimiza- tion. Similarly , random projection yields acceptable predic- tive performance but weaker deco ding and design results compared to PCA. These ndings highlight a key distinction: predictive per- formance alone does not determine the usefulness of a r epre- sentation for combinatorial optimization. Instead, the ge om- etry of the latent space and its compatibility with decoding and search play a central r ole. This observation is consistent with the theoretical analysis in Se ction A, which shows that dierent repr esentations can induce substantially dierent optimization landscapes even when predictive accuracy is similar . Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization Figure 2. T est-set performance of external sequence-level tness oracles across data regimes on GFP and AA V . Each panel shows the Spearman correlation between predicted and ground-truth tness as a function of the number of training samples. Across b oth datasets, Gaussian process regression often achieves the strongest p erformance in b oth low-data and mo derate-data regimes, reecting its ability to model uncertainty under limited supervision. As the dataset size increases, ridge r egression becomes increasingly competitive and often achieves near the best performance, indicating that the tness signal is largely captured by a linear model in the ESM embedding space. In contrast, X GBo ost underperforms in the smallest-data r egime, suggesting overtting, but impro ves steadily with additional data. These results establish a reliable sequence-lev el oracle for evaluating generated protein sequences and highlight the importance of selecting appropriate models under dierent data regimes. T able 4. Decoding quality of projection-based binar y latent representations. PCA consistently achieves higher mutation F1 and mutated-residue accuracy than random projection, and both metrics improve with increasing latent dimensionality . The better performance b etween two latent dimensions (i.e. bits) of 16 and 64 is highlighted in bold across dierent settings. The higher latent dimension (i.e. 64) consistently outperforms the lower latent dimension (i.e . 16) in decoding quality . Dataset Train size Representation Latent dim Mutation F1 ↑ AA accuracy ↑ GFP 1000 PCA 16 0.110 0.551 PCA 64 0.273 0.595 Random 16 0.102 0.510 Random 64 0.168 0.553 AA V 10000 PCA 16 0.369 0.367 PCA 64 0.603 0.602 Random 16 0.338 0.316 Random 64 0.469 0.475 Overall, these results demonstrate that while the QUBO surrogate can b e tted across a range of representations, only structured latent spaces such as PCA lead to eective end-to-end protein design. 4.5 Conservative Retrieval-Based Optimization Benchmark W e evaluate Q-BioLa t in a conservative setting where opti- mized latent codes are mapped back to obser ved sequences via nearest-neighbor retrieval in Hamming space. This bench- mark isolates the quality of the induced latent tness land- scape without confounding eects from neural decoding. Truong-Son Hy Figure 3. Scaling of decoding p erformance with dataset size for PCA -based and random-projection binary latent representations at 64 bits. The x-axis shows the numb er of training samples and the y-axis shows mutation F1. Acr oss both GFP and AA V datasets, PCA consistently achieves higher deco ding accuracy than random projection, and p erformance improves with increasing data. T able 5. Predictive performance of the internal QUBO surrogate across latent representations, training data sizes, and latent dimensions. W e report held-out test Spearman correlation. Although predictive dier ences are often modest, do wnstream optimization behavior diers substantially across representations. The better performance between the two binary-latent representations (i.e . PCA and random projection) is highlighted in bold. PCA outp erforms random projection consistently in predictive quality of the internal QUBO surrogate across dier ent settings. Dataset Train size Latent dim Representation Spearman ↑ GFP 1000 16 Random 0.291 PCA 0.147 64 Random 0.196 PCA 0.277 10000 16 Random 0.302 PCA 0.303 64 Random 0.413 PCA 0.532 AA V 1000 16 Random 0.417 PCA 0.393 64 Random 0.407 PCA 0.511 10000 16 Random 0.366 PCA 0.422 64 Random 0.534 PCA 0.564 T o ensure a fair comparison across optimization metho ds, all results are reported using PCA -base d latent repr esenta- tions with a xed latent dimension of 64 bits. T ables 6 and 7 summarize the mean retrieval-based optimization p erfor- mance across multiseed runs on GFP and AA V , respe ctively . Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization Across both datasets and all data regimes, Simulated An- nealing (SA), Genetic Algorithms (GA), and Greedy Hill Climb (GHC) consistently achieve strong performance, indi- cating that the induced QUBO landscape is well-structured for combinatorial search. In particular , these metho ds reli- ably identify high-tness regions of the observed sequence manifold even in data-scarce settings, while latent Bayesian- style optimization (LBO) is generally less competitive. These results demonstrate that PCA -base d binary latent representations produce optimization-friendly landscapes that can b e eectively explor ed by classical combinatorial solvers, providing a strong foundation for the full end-to-end design pipeline. 4.6 End-to-End Sequence Design Benchmark W e evaluate Q-BioLa t in an end-to-end protein design set- ting where optimized binary latent codes are decoded into protein sequences and scored using the external oracle. This benchmark directly measures whether optimization in bi- nary latent space leads to improved sequence-level tness. Figure 4 summarizes the best end-to-end performance achieved by PCA -based and random-projection binary latent representations across training regimes on GFP and AA V . For each dataset and train size, we report the strongest con- guration after jointly selecting the optimizer and latent dimension. Across most settings, PCA -based binar y latent repr esenta- tions achieve the strongest performance . This advantage is especially pronounced at mo derate and larger data regimes, where higher-dimensional PCA latents (32 and 64 bits) con- sistently produce the highest oracle scores. These ndings are consistent with the deco der analysis in Section 4.3, where PCA was shown to b e substantially more decodable than both random projections and learned AE/V AE latent spaces. W e also observe that larger latent dimensions generally improve end-to-end performance, particularly for PCA, in- dicating that structured latent spaces can benet from in- creased capacity without sacricing decodability . In contrast, random projections are less reliable and usually underper- form PCA, although a few low-data settings exhibit isolated exceptions. Among the optimizers, Greedy Hill Climb (GHC), Ge- netic Algorithm ( GA), and Simulated Annealing (SA) ar e consistently competitive, while latent Bayesian-style search (LBO) is generally w eaker . Overall, these results sho w that Q-BioLa t provides an eective framework for end-to-end protein design, where binary latent optimization combined with structured representations enables the discovery of high-tness protein sequences under a xed oracle evalua- tion budget. 4.7 Data-Scarcity and Scaling Analysis W e next analyze how the p erformance of Q-BioLa t evolves as the amount of available tness data increases. This exper- iment is particularly important because one of the primary motivations for using compact binar y latent representations is to enable robust optimization under limited supervision. T able 9 summarizes the best-performing oracle, latent representation, and design method acr oss data regimes on both GFP and AA V . Several consistent patterns emerge. Oracle b ehavior across data regimes. Across all datasets and training sizes, Gaussian process regression consistently provides the strongest sequence-level oracle . This highlights its ability to capture nonlinear structure in protein tness landscapes and remain robust even as the dataset size in- creases. In contrast, simpler models such as ridge regression and XGBoost r emain competitive but do not surpass Gauss- ian processes in our experiments. Latent representation. A cross all datasets and data regimes, PCA -base d binar y latent representations consistently pro- vide the strongest performance . This observation aligns with the ndings in Section 4.3, where PCA was shown to main- tain high entropy , active latent dimensions, and strong de- coding performance, while AE/V AE repr esentations collapse after binarization. Optimization methods. The best-p erforming design method varies with both dataset and data regime. In low-data set- tings, simpler metho ds such as Random Search (RS) and Greedy Hill Climbing (GHC) are often competitive, reect- ing the relatively limited structure of the learned landscape. As the dataset size increases, more sophisticated combinato- rial methods such as Genetic Algorithm (GA) and Simulated Annealing (SA) become increasingly ee ctive, suggesting that the learned QUBO landscape becomes mor e informative and structured. Scaling trends. W e obser ve a clear transition from less stable behavior in the low-data regime (1000–2000 samples) to more consistent and high-performing optimization in the moderate-data regime (5000–10000 samples). In particu- lar , PCA -based latent representations combined with higher latent dimensionality enable improved end-to-end perfor- mance, indicating that structured latent spaces can ee c- tively leverage additional data without sacricing deco dabil- ity or optimization stability . Summary . Overall, these results demonstrate that Q-BioLa t provides a robust frame work for protein design acr oss data regimes. The combination of a strong Gaussian process ora- cle, structured PCA -based latent representations, and com- binatorial optimization enables eective identication of high-tness sequences, particularly in data-scarce settings, while continuing to improve as more data becomes available . Truong-Son Hy T able 6. Retrieval-based optimization results on GFP using PCA latent representations with 64 bits. Each row reports the mean performance across multiseed runs for one optimizer . Optimizers include Simulated Annealing (SA), Genetic Algorithm (GA), Random Search (RS), Greedy Hill Climb (GHC), and Latent Bayesian Optimization (LBO). In each data size, the b est performance in each metric is highlighted in bold. Train size Optimizer Improvement ↑ NN tness ↑ Percentile ↑ 1000 SA 4.978 3.568 73.200 GA 5.044 3.440 60.925 RS 1.750 3.443 58.650 GHC 4.918 3.616 74.825 LBO -1.509 2.766 58.050 2000 SA 11.529 3.558 70.700 GA 11.103 2.908 62.025 RS 4.770 2.593 43.175 GHC 10.230 3.580 75.013 LBO -1.119 2.953 47.700 5000 SA 6.757 3.479 63.410 GA 6.628 3.689 83.330 RS 2.617 3.078 61.640 GHC 6.083 3.761 80.605 LBO -1.005 2.775 54.085 10000 SA 4.162 3.220 69.740 GA 3.938 3.607 74.595 RS 1.609 3.590 71.938 GHC 3.688 3.175 64.562 LBO -1.253 1.836 28.780 Figure 4. End-to-end sequence design performance after optimization, decoding, and oracle scoring. For each dataset and training size, we plot the best-performing PCA -based and random-projection conguration. Each point is annotate d with the corresponding latent dimension and optimizer . Across most settings, PCA -based binar y latent repr esentations achieve the strongest sequence-level performance, particularly at moderate and larger data regimes. 4.8 Implementation Details and Rep orting Protocol Unless otherwise specied, all metho ds use the same up- stream ESM emb edding extractor in order to isolate the ee ct Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization T able 7. Retrieval-based optimization results on AA V using PCA latent repr esentations with 64 bits. Each r ow r eports the mean performance across multiseed runs for one optimizer . Optimizers include Simulated Annealing (SA), Genetic Algorithm (GA), Random Search (RS), Greedy Hill Climb (GHC), and Latent Bayesian Optimization (LBO). In each data size, the b est performance in each metric is highlighted in bold. Train size Optimizer Improvement ↑ NN tness ↑ Percentile ↑ 1000 SA 11.364 3.303 88.225 GA 10.342 3.949 90.000 RS 1.224 1.945 77.550 GHC 9.362 3.902 91.400 LBO -9.071 -0.606 53.200 2000 SA 28.756 1.794 72.275 GA 27.108 -1.256 47.325 RS 8.523 0.343 66.050 GHC 26.511 1.856 74.388 LBO -11.177 -0.116 61.725 5000 SA 14.627 -0.654 50.165 GA 13.636 3.869 95.325 RS 4.153 1.477 77.715 GHC 13.162 6.643 94.975 LBO -5.378 -1.871 44.250 10000 SA 12.575 3.038 86.198 GA 12.321 3.720 92.948 RS 4.115 2.278 78.420 GHC 11.545 3.262 91.380 LBO -3.503 0.003 61.045 of latent representation and optimization strategy . Hyp erpa- rameters ar e selected on the validation split, and nal results are r eported on the held-out test split or on oracle-evaluated generated candidates, depending on the experiment. For de- sign metho ds, we use a xe d candidate budget and report both the best returned sequence and the average perfor- mance of the top- 𝐾 returned candidates. All reported means and standard deviations are computed o ver multiple random seeds. For this work, we publicly r elease code, exact hyper- parameter settings, and scripts for reproducing all tables and gures at: hps://github.com/HySonLab/Q- BIOLA T - Extended 5 Conclusion In this work, w e introduced Q-BioLa t , a framework for modeling and optimizing protein tness landscapes in com- pact binary latent spaces. By transforming protein sequence representations into binary codes and learning a quadratic unconstrained binary optimization (QUBO) surrogate, Q- BioLa t enables direct application of combinatorial optimiza- tion methods to protein design. Beyond its algorithmic formulation, our central contribu- tion is a representation-centric persp ective on protein tness modeling. W e show both theoretically and empirically that predictive accuracy alone is insucient for eective opti- mization. Instead, the ge ometry of the latent space—captured through its induced QUBO interactions—plays a decisive role in determining search behavior , landscape smo othness, and the ability to identify high-tness sequences. Our experiments demonstrate three key ndings. First, learned autoencoder-based representations collapse under bi- narization and fail to support optimization, despite achieving low reconstruction error . Second, simple structured represen- tations such as PCA consistently produce more informative and decodable latent spaces, leading to better optimization performance. Third, combinatorial optimization methods such as simulated annealing, genetic algorithms, and greedy hill climbing are highly eective when applied to these struc- tured binary latent landscapes, especially when combined with a strong sequence-level oracle . W e further show that Q-BioLa t performs robustly across data regimes, including data-scarce settings where tradi- tional methods often struggle. As more data becomes avail- able, both oracle quality and latent structure improve , leading to increasingly eective end-to-end protein design. Truong-Son Hy T able 8. End-to-end protein design benchmark under a xed e xternal-oracle evaluation budget. For each dataset and train size , we report the best-performing random-projection and PCA-based Q-BioLa t conguration after optimization, decoding, and oracle scoring. Optimizers include Simulated Annealing (SA), Genetic Algorithm ( GA), Random Search (RS), Greedy Hill Climb (GHC), and Latent Bayesian Optimization (LBO). The better performance between PCA and random projection is highlighted in bold. Dataset Train size Representation Optimizer Latent dim Best score ↑ T op-10 mean ↑ GFP 1000 PCA RS 64 4.119 3.903 Random GA 64 4.170 3.312 2000 PCA GA 64 4.027 3.549 Random GHC 32 3.448 2.693 5000 PCA GHC 64 4.270 3.972 Random GHC 32 3.815 2.228 10000 PCA GHC 64 5.554 5.042 Random GHC 32 4.806 3.829 AA V 1000 PCA GHC 64 5.232 2.962 Random GHC 64 4.066 -1.557 2000 PCA RS 32 2.866 0.349 Random GHC 32 4.184 -2.674 5000 PCA SA 64 3.999 2.249 Random GA 64 2.548 -1.150 10000 PCA GA 64 4.541 2.091 Random LBO 16 1.197 -4.205 T able 9. Scaling summary acr oss data regimes. W e report the b est-performing oracle, latent representation, and design method as a function of available training data. Dataset Train size Best oracle Best latent representation Best design method GFP 1000 Gaussian Process PCA Random Search (RS) 2000 Gaussian Process PCA Genetic Algorithm (GA) 5000 Gaussian Process PCA Greedy Hill Climb (GHC) 10000 Gaussian Process PCA Greedy Hill Climb (GHC) AA V 1000 Gaussian Process PCA Greedy Hill Climb (GHC) 2000 Gaussian Process PCA Random Search (RS) 5000 Gaussian Process PCA Simulated Annealing (SA) 10000 Gaussian Process PCA Genetic Algorithm (GA) Overall, Q-BioLa t bridges protein language mo deling, dis- crete repr esentation learning, and combinatorial optimiza- tion, providing a principled framework for optimization- aware protein tness modeling. By e xplicitly modeling the latent tness landscape as a QUBO pr oblem, our approach naturally conne cts modern machine learning with classi- cal and quantum optimization paradigms. This opens sev- eral promising directions for future work, including inte- gration with quantum annealing hardware , development of optimization-aware representation learning methods, and extension to multi-objective and constrained protein design. References [1] Philip A Romero and Frances H Arnold. Exploring protein tness landscapes by directed e volution. Nature reviews Molecular cell biology , 10(12):866–876, 2009. [2] Lin Chen, Zehong Zhang, Zhenghao Li, Rui Li, Ruifeng Huo, Lifan Chen, Dingyan W ang, Xiaomin Luo, Kaixian Chen, Cangsong Liao, and Mingyue Zheng. Learning protein tness landscapes with deep mutational scanning data from multiple sources. Cell Systems , 14(8): 706–721.e5, 2023. ISSN 2405-4712. doi: https://doi.org/10.1016/j.cels. 2023.07.003. URL hps://www .sciencedirect.com/science/article/pii/ S2405471223002107 . [3] Pascal Notin, Aaron Kollasch, Daniel Ritter , Lood van Niekerk, Stef- fanie Paul, Han Spinner , Nathan Rollins, Ada Shaw , Rose Orenbuch, Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization Ruben W eitzman, Jonathan Frazer , Mafalda Dias, Dinko Franceschi, Y arin Gal, and Debora Marks. Proteingym: Large-scale benchmarks for protein tness prediction and design. In A. Oh, T . Naumann, A. Globerson, K. Saenko, M. Hardt, and S. Levine, editors, Advances in Neural Information Processing Systems , volume 36, pages 64331–64379. Curran Associates, Inc., 2023. URL hps://proceedings.neurips. cc/paper_files/paper/2023/file/cac723e529f65e3fcbb0739ae91bee- Paper- Datasets_and_Benchmarks.p df . [4] Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, W ent- ing Lu, Nikita Smetanin, Robert V erkuil, Ori Kabeli, Yaniv Shmueli, Allan dos Santos Costa, Maryam Fazel-Zarandi, T om Sercu, Sal- vatore Candido, and Alexander Rives. Evolutionary-scale predic- tion of atomic-level protein structure with a language model. Sci- ence , 379(6637):1123–1130, 2023. doi: 10.1126/science.ade2574. URL hps://www.science .org/doi/abs/10.1126/science.ade2574 . [5] Thomas Hayes, Roshan Rao, Halil Akin, Nicholas J. Sofroniew , Deniz Oktay , Zeming Lin, Robert V erkuil, Vincent Q . Tran, Jonathan Deaton, Marius Wiggert, Rohil Badkundri, Irhum Shaf kat, Jun Gong, Alexander Derry , Raul S. Molina, Neil Thomas, Y ousuf A. Khan, Chetan Mishra, Carolyn Kim, Liam J. Bartie , Matthew Nemeth, Patrick D . Hsu, T om Sercu, Salvatore Candido , and Alexander Rives. Simulating 500 million years of evolution with a language model. Science , 387(6736):850–858, 2025. doi: 10.1126/science.ads0018. URL hps://www .science.org/doi/ abs/10.1126/science.ads0018 . [6] Céline Marquet, Julius Schlensok, Marina Abakarova, Burkhard Rost, and Elo die Laine. Expert-guided protein language models enable accurate and blazingly fast tness prediction. Bioinformatics , 40(11): btae621, 11 2024. ISSN 1367-4811. doi: 10.1093/bioinformatics/btae621. URL hps://doi.org/10.1093/bioinformatics/btae621 . [7] Dan Liu, Francesca Y oung, Kieran D Lamb, Adalberto Claudio Quiros, Alexandrina Pancheva, Crispin J Miller, Craig Macdonald, David L Robertson, and Ke Y uan. P lm-interact: extending protein language models to predict protein-protein interactions. Nature Communica- tions , 16(1):9012, 2025. [8] Joseph L W atson, David Juergens, Nathaniel R Bennett, Brian L T rippe, Jason Yim, Helen E Eisenach, W oody Ahern, Andrew J Borst, Robert J Ragotte, Lukas F Milles, et al. De novo design of protein structur e and function with rfdiusion. Nature , 620(7976):1089–1100, 2023. [9] Thanh V T Tran, Nhat Khang Ngo, Viet Thanh Duy Nguyen, and Truong-Son Hy . Latentde: latent-based directed evolution for protein sequence design. Machine Learning: Science and T echnology , 6(1): 015070, mar 2025. doi: 10.1088/2632- 2153/adc2e2. URL hps://doi.org/ 10.1088/2632- 2153/adc2e2 . [10] Thanh V . T . Tran and Truong Son Hy . Protein design by directed evolution guided by large language models. IEEE Transactions on Evolutionary Computation , 29(2):418–428, 2025. doi: 10.1109/TEVC. 2024.3439690. [11] Hiroki Ozawa, Ibuki Unno, Ryohei Sekine, T aichi Chisuga, Sohei Ito, and Shogo Nakano. Development of evolutionary algorithm- based protein redesign method. Cell Rep orts P hysical Science , 5(1): 101758, 2024. ISSN 2666-3864. doi: https://doi.org/10.1016/j.xcrp. 2023.101758. URL hps://www .sciencedirect.com/science/article/pii/ S2666386423006033 . [12] Timothy Atkinson, Thomas D Barrett, Scott Cameron, Bora Guloglu, Matthew Greenig, Charlie B T an, Louis Robinson, Alex Graves, Liviu Copoiu, and Alexandre Laterre. Protein se quence modelling with bayesian ow networks. Nature Communications , 16(1):3197, 2025. [13] Ruyun Hu, Lihao Fu, Y ongcan Chen, Junyu Chen, Yu Qiao, and T ong Si. Protein engineering via bayesian optimization-guided evolutionary algorithm and robotic experiments. Briengs in Bioinformatics , 24(1): bbac570, 12 2022. ISSN 1477-4054. doi: 10.1093/bib/bbac570. URL hps://doi.org/10.1093/bib/bbac570 . [14] Gary Kochenberger , Jin-Kao Hao, Fred Glover , Mark Lewis, Zhipeng Lü, Haibo W ang, and Y ang W ang. The unconstrained binary quadratic programming problem: a survey . Journal of combinatorial optimization , 28(1):58–81, 2014. [15] Andrew Lucas. Ising formulations of many np problems. Frontiers in Physics , V olume 2 - 2014, 2014. ISSN 2296-424X. doi: 10.3389/fphy .2014. 00005. URL hps://www .frontiersin.org/journals/physics/articles/10. 3389/fphy .2014.00005 . [16] Seongmin Kim, Sang- W oo Ahn, In-Saeng Suh, Alexander W Dowling, Eungkyu Lee, and T engfei Luo. Quantum annealing for combinatorial optimization: a benchmarking study . npj Quantum Information , 11(1): 77, 2025. [17] Viet Thanh Duy Nguyen and T ruong Son Hy . Multimodal pretrain- ing for unsupervised protein representation learning. Biology Meth- ods and Protocols , 9(1):bpae043, 06 2024. ISSN 2396-8923. doi: 10. 1093/biomethods/bpae043. URL hps://doi.org/10.1093/biomethods/ bpae043 . [18] Nhat Khang Ngo and Truong Son Hy . Multimodal protein repre- sentation learning and target-aware variational auto-encoders for protein-binding ligand generation. Machine Learning: Science and T echnology , 5(2):025021, apr 2024. doi: 10.1088/2632- 2153/ad3e e4. URL hps://doi.org/10.1088/2632- 2153/ad3ee4 . [19] Alexander Rives, Joshua Meier , T om Sercu, Siddharth Goyal, Zeming Lin, Jason Liu, Demi Guo, Myle Ott, C. Lawrence Zitnick, Jerry Ma, and Rob Fergus. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proceedings of the National Academy of Sciences , 118(15):e2016239118, 2021. doi: 10.1073/pnas.2016239118. URL hps://ww w .pnas.org/doi/abs/10.1073/ pnas.2016239118 . [20] Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, W ent- ing Lu, Nikita Smetanin, Robert V erkuil, Ori Kabeli, Yaniv Shmueli, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science , 379(6637):1123–1130, 2023. [21] Kevin K Y ang, Zachary Wu, and Frances H Arnold. Machine-learning- guided directed evolution for pr otein engineering. Nature methods , 16 (8):687–694, 2019. [22] Surojit Biswas, Grigory Khimulya, Ethan C Alley , K evin M Esvelt, and George M Church. Low-n protein engineering with data-ecient deep learning. Nature methods , 18(4):389–396, 2021. [23] Fanhao W ang, Tiantian Zhang, Jintao Zhu, Xiaoling Zhang, Chang- sheng Zhang, and Luhua Lai. Reinforcement learning-based target- specic de novo design of cyclic peptide binders. Journal of Medicinal Chemistry , 68(16):17287–17302, 2025. [24] Haoran Sun, Liang He, Pan Deng, Guoqing Liu, Zhiyu Zhao, Yuliang Jiang, Chuan Cao, Fusong Ju, Lijun Wu, Haiguang Liu, et al. Ac- celerating protein engineering with tness landscape modelling and reinforcement learning. Nature Machine Intelligence , 7(9):1446–1460, 2025. [25] Optimization by simulated annealing. science , 220(4598):671–680, 1983. [26] John H. Holland. Adaptation in Natural and Articial Systems: A n Introductory Analysis with A pplications to Biology , Control, and A rti- cial Intelligence . The MI T Press, 04 1992. ISBN 9780262275552. doi: 10.7551/mitpress/1090.001.0001. URL hps://doi.org/10.7551/mitpr ess/ 1090.001.0001 . [27] Mark W . Johnson, Mohammad H. S. Amin, Gildert Suzanne, T revor Lanting, Firas Hamze, Neil G. Dickson, Richard G. Harris, Andr ew J. Berkley , J. Johansson, Paul I. Bunyk, E. M. Chapple, Colin Enderud, Jeremy P. Hilton, Kamran K arimi, Eric Ladizinsky , Nicolas Ladizinsky, Travis Oh, I. G. Perminov , Chris Rich, Murray C. Thom, Elena T olka- cheva, C. J. S. Truncik, Sergey Uchaikin, J. W ang, Blake A. Wilson, and Geordie Rose. Quantum annealing with manufactured spins. Nature , 473:194–198, 2011. URL hps://api.semanticscholar .org/CorpusID: 205224761 . [28] André Hottung, Bhanu Bhandari, and Kevin Tierney . Learning a latent search space for routing problems using variational autoencoders. In International Conference on Learning Representations , 2021. URL Truong-Son Hy hps://openreview .net/forum?id=90JprV rJBO . [29] Peter J. Bentley , Soo Ling Lim, Adam Gaier , and Linh Tran. Coil: Constrained optimization in learned latent space: learning repre- sentations for valid solutions. In Proceedings of the Genetic and Evolutionary Computation Conference Companion , GECCO ’22, page 1870–1877, New Y ork, NY, USA, 2022. Association for Computing Machinery . ISBN 9781450392686. doi: 10.1145/3520304.3533993. URL hps://doi.org/10.1145/3520304.3533993 . [30] Dhruv Menon and Raghavan Ranganathan. A generative approach to materials discovery , design, and optimization. A CS Omega , 7(30): 25958–25973, 2022. doi: 10.1021/acsomega.2c03264. URL hps://doi. org/10.1021/acsomega.2c03264 . [31] Truong Son Hy and Risi Kondor . Multiresolution equivariant graph variational autoencoder . Machine Learning: Science and T echnology , 4(1):015031, mar 2023. doi: 10.1088/2632- 2153/acc0d8. URL hps: //doi.org/10.1088/2632- 2153/acc0d8 . [32] Mingrui Jiang, Keyi Shan, Chengping He, and Can Li. Ecient com- binatorial optimization by quantum-inspired parallel annealing in analogue memristor crossbar . Nature Communications , 14(1):5927, 2023. [33] T etsuro Abe, Masashi Y amashita, and Shu Tanaka. Eectiveness of binary autoenco ders for qubo-based optimization problems. arXiv preprint arXiv:2602.10037 , 2026. [34] Marc Mézard and Andrea Montanari. Information, Physics, and Com- putation . Oxford University Press, 01 2009. ISBN 9780198570837. doi: 10.1093/acprof:oso/9780198570837.001.0001. URL hps://doi.org/10. 1093/acprof:oso/9780198570837.001.0001 . [35] David W ales. Energy Landscapes: A pplications to Clusters, Biomolecules and Glasses . Cambridge Molecular Science. Cambridge University Press, 2004. [36] Antonio A unger , Gérard Ben Arous, and Jiří Černý . Random matrices and complexity of spin glasses. Communications on Pure and A pplied Mathematics , 66(2):165–201, 2013. doi: https://doi.org/10.1002/cpa. 21422. URL hps://onlinelibrary .wiley .com/doi/abs/10.1002/cpa.21422 . [37] Anna Choromanska, MIkael Hena, Michael Mathieu, Gerard Ben Arous, and Y ann LeCun. The Loss Surfaces of Multilayer Net- works. In Guy Lebanon and S. V . N. Vishwanathan, editors, Pr oceedings of the Eighteenth International Conference on A rticial Intelligence and Statistics , v olume 38 of Proceedings of Machine Learning Resear ch , pages 192–204, San Diego, California, USA, 09–12 May 2015. PMLR. URL hps://proceedings.mlr .press/v38/choromanska15.html . [38] Ian Goodfellow , Oriol Vinyals, and Andrew Saxe. Qualitatively char- acterizing neural network optimization problems. In International Conference on Learning Representations , 2015. URL hp://ar xiv .org/ abs/1412.6544 . [39] Rafael Gómez-Bombarelli, Jennifer N. W ei, David Duvenaud, José Miguel Hernández-Lobato, Benjamín Sánchez-Lengeling, Den- nis Sheberla, Jorge Aguilera-Iparraguirre, Timothy D . Hirzel, Ryan P. Adams, and Alán Aspuru-Guzik. A utomatic chemical design using a data-driven continuous representation of molecules. ACS Central Science , 4(2):268–276, 2018. doi: 10.1021/acscentsci.7b00572. URL hps://doi.org/10.1021/acscentsci.7b00572 . PMID: 29532027. [40] W engong Jin, Regina Barzilay , and T ommi Jaakkola. Junction tree variational autoencoder for molecular graph generation. In Jennifer Dy and Andreas Krause, editors, Proceedings of the 35th International Conference on Machine Learning , volume 80 of Proceedings of Machine Learning Research , pages 2323–2332. PMLR, 10–15 Jul 2018. URL hps://proceedings.mlr .press/v80/jin18a.html . [41] Ryan-Rhys Griths and José Miguel Hernández-Lobato. Constrained bayesian optimization for automatic chemical design using variational autoencoders. Chem. Sci. , 11:577–586, 2020. doi: 10.1039/C9SC04026A. URL hp://dx.doi.org/10.1039/C9SC04026A . [42] ANM Naz Abeer , Nathan M Urban, M Ryan W eil, Francis J Alexander , and Byung-Jun Y oon. Multi-obje ctive latent space optimization of generative molecular design models. Patterns , 5(10), 2024. [43] Y oshua Bengio, Aaron Courville, and Pascal Vincent. Representation learning: A review and new perspectives. IEEE Trans. Pattern Anal. Mach. Intell. , 35(8):1798–1828, August 2013. ISSN 0162-8828. doi: 10.1109/TP AMI.2013.50. URL hps://doi.org/10.1109/TP AMI.2013.50 . [44] Piotr Bojano wski, Armand Joulin, David Lopez-Pas, and Arthur Szlam. Optimizing the latent space of generative networks. In Jennifer Dy and Andreas Krause, editors, Proceedings of the 35th International Conference on Machine Learning , volume 80 of Proceedings of Machine Learning Research , pages 600–609. PMLR, 10–15 Jul 2018. URL hps: //proceedings.mlr .press/v80/b ojanowski18a.html . [45] G. E. Hinton and R. R. Salakhutdinov . Re ducing the dimensionality of data with neural networks. Science , 313(5786):504–507, 2006. doi: 10.1126/science.1127647. URL hps://w ww .science.org/doi/abs/10. 1126/science.1127647 . [46] Pascal Vincent, Hugo Larochelle, Isabelle Lajoie, Y oshua Bengio, and Pierre- Antoine Manzagol. Stacked denoising autoencoders: Learn- ing useful representations in a deep network with a local denoising criterion. J. Mach. Learn. Res. , 11:3371–3408, December 2010. ISSN 1532-4435. [47] Diederik P Kingma and Max W elling. Auto-encoding variational bayes. arXiv preprint arXiv:1312.6114 , 2013. Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization A Theoretical Insights into Binary Latent Fitness Landscapes In this section, we provide a theoretical perspective on how binary latent representations induce structured optimization landscapes, and how their geometry ae cts combinatorial search. A.1 QUBO as an Energy Landscap e Recall that the QUBO surrogate denes the predicted tness of a binary latent code 𝑥 ∈ { 0 , 1 } 𝑚 as: ˆ 𝑓 ( 𝑥 ) = ℎ ⊤ 𝑥 + 1 2 𝑥 ⊤ 𝐽 𝑥 , (9) where ℎ ∈ R 𝑚 and 𝐽 ∈ R 𝑚 × 𝑚 is a symmetric interaction matrix. This formulation can be equivalently interpreted as an energy-based model: 𝐸 ( 𝑥 ) = − ˆ 𝑓 ( 𝑥 ) , (10) where optimization corresponds to nding low-energy con- gurations. Under this view , the binary latent space { 0 , 1 } 𝑚 forms a discrete conguration space (a Boolean hypercube), and the QUBO denes an energy landscape over its vertices. The structure of this landscape is fully determined by the parameters ( ℎ, 𝐽 ) , which in turn dep end on the choice of latent representation. A.2 Local Geometr y and Bit-F lip Dynamics A fundamental operation in combinatorial optimization is a single-bit ip. Let 𝑥 ( 𝑘 ) denote the v ector obtaine d by ipping the 𝑘 -th bit of 𝑥 . The change in predicted tness is: Δ 𝑘 ( 𝑥 ) = ˆ 𝑓 ( 𝑥 ( 𝑘 ) ) − ˆ 𝑓 ( 𝑥 ) . (11) For the QUBO model, this quantity can be expr essed ex- plicitly as: Δ 𝑘 ( 𝑥 ) = ( 1 − 2 𝑥 𝑘 ) ℎ 𝑘 + ℓ ≠ 𝑘 𝐽 𝑘 ℓ 𝑥 ℓ ! . (12) This expression sho ws that local search behavior is gov- erned by the interaction structure of 𝐽 . In particular: • The term ℎ 𝑘 captures the intrinsic contribution of the 𝑘 -th latent dimension. • The sum Í ℓ 𝐽 𝑘 ℓ 𝑥 ℓ captures how other latent variables inuence the eect of ipping bit 𝑘 . Thus, optimization trajectories dep end not only on the current state 𝑥 , but also on the global interaction structure encoded in 𝐽 . A.3 Landscape Smoothness and Ruggedness The diculty of optimization is closely related to the smooth- ness of the energy landscape. W e characterize local variabil- ity using the variance of Δ 𝑘 ( 𝑥 ) over the latent space: V ar [ Δ 𝑘 ( 𝑥 ) ] ≈ ℓ 𝐽 2 𝑘 ℓ . (13) This suggests that the norm of the interaction coecients controls the ruggedness of the landscape: • Small ∥ 𝐽 𝑘 · ∥ leads to smoother landscapes with more predictable local improvements. • Large ∥ 𝐽 𝑘 · ∥ leads to highly variable local changes and a proliferation of local optima. Consequently , the structure of 𝐽 plays a central role in determining optimization diculty . A.4 Spectral Structure of the Interaction Matrix Further insight can be obtained by analyzing the spectrum of the interaction matrix 𝐽 . Let { 𝜆 𝑖 } denote its eigenvalues. Then: • Low-rank or structured 𝐽 (few dominant eigenvalues) corresponds to landscapes with coherent global direc- tions, facilitating optimization. • Random or full-rank 𝐽 with dispersed eigenvalues cor- responds to more irregular , spin-glass-like landscapes with many local minima. Since 𝐽 is learned from latent representations, its spe ctral properties are directly inuenced by the choice of represen- tation (e.g., PCA vs random projection vs learned encoders). A.5 Representation–Optimization Decoupling A key implication of this analysis is that predictive perfor- mance alone do es not determine optimization b ehavior . T wo representations may yield similar surrogate accuracy ( e.g., similar Spearman correlation), yet induce dier ent interac- tion matrices 𝐽 with distinct ge ometric properties. Observation: Representations with similar pre- dictive accuracy can induce fundamentally dif- ferent optimization landscapes. This phenomenon arises because predictive metrics eval- uate pointwise accuracy , whereas optimization depends on the global structur e of the energy landscape . As a result, rep- resentation learning for protein design should account not only for prediction quality , but also for the induced landscape geometry . A.6 Implications for Latent Representation Design The above analysis suggests several principles for designing eective latent representations: • Structured representations (e.g., PCA or learned en- coders) tend to produce smoother and more optimization- friendly landscapes. Truong-Son Hy • Overly high-dimensional latent spaces can increase model expressivity but also introduce ruggedness, making optimization more dicult. • Learned repr esentations can potentially balance ex- pressivity and smoothness by aligning latent dimen- sions with tness-relevant directions. These insights provide a theoretical foundation for under- standing the empirical observations in Section 4 and moti- vate the development of optimization-aware r epresentation learning methods. Here, we formalize ho w the learne d binary r epresentation determines the geometry , smo othness, and identiability of the induced QUBO landscape. Our goal is not to solve the global QUBO exactly , but to characterize the quantities that govern local search, global variation, and the stability of the learned surrogate. A.7 Energy Landscape and Local Fields Recall that the QUBO surrogate denes the predicted tness of a binary latent code 𝑥 ∈ { 0 , 1 } 𝑚 as: ˆ 𝑓 ( 𝑥 ) = ℎ ⊤ 𝑥 + 1 2 𝑥 ⊤ 𝐽 𝑥 , (14) where ℎ ∈ R 𝑚 and 𝐽 ∈ R 𝑚 × 𝑚 is a symmetric interaction matrix with zero diagonal. Equivalently , we may dene the energy: 𝐸 ( 𝑥 ) = − ˆ 𝑓 ( 𝑥 ) , (15) so that maximizing predicted tness is e quivalent to mini- mizing energy over the Boolean hypercube. A useful quantity for analyzing this landscap e is the local eld : 𝑔 𝑘 ( 𝑥 ) = ℎ 𝑘 + ℓ ≠ 𝑘 𝐽 𝑘 ℓ 𝑥 ℓ , (16) which measures the eective force acting on the 𝑘 -th bit given the remaining coordinates. Proposition (Bit-f lip gain and local optimality). Let 𝑥 ( 𝑘 ) denote the binary co de obtained by ipping the 𝑘 -th bit of 𝑥 . Then the change in predicted tness induced by a single-bit ip is: Δ 𝑘 ( 𝑥 ) = ˆ 𝑓 ( 𝑥 ( 𝑘 ) ) − ˆ 𝑓 ( 𝑥 ) = ( 1 − 2 𝑥 𝑘 ) 𝑔 𝑘 ( 𝑥 ) . (17) Consequently , a binary code 𝑥 ★ is a lo cal maximizer with respect to single-bit ips if and only if: Δ 𝑘 ( 𝑥 ★ ) ≤ 0 for all 𝑘 , (18) or equivalently , 𝑥 ★ 𝑘 = ( 1 , 𝑔 𝑘 ( 𝑥 ★ ) ≥ 0 , 0 , 𝑔 𝑘 ( 𝑥 ★ ) ≤ 0 , (19) with either value allowed when 𝑔 𝑘 ( 𝑥 ★ ) = 0 . Proof. W rite 𝑥 ( 𝑘 ) = 𝑥 + ( 1 − 2 𝑥 𝑘 ) 𝑒 𝑘 , where 𝑒 𝑘 is the 𝑘 -th standard basis vector . Substituting into the QUBO objective and using the symmetry of 𝐽 together with 𝐽 𝑘 𝑘 = 0 gives: ˆ 𝑓 ( 𝑥 ( 𝑘 ) ) − ˆ 𝑓 ( 𝑥 ) = ( 1 − 2 𝑥 𝑘 ) ℎ 𝑘 + ℓ ≠ 𝑘 𝐽 𝑘 ℓ 𝑥 ℓ ! . The lo cal-optimality condition follows immediately by re- quiring that no single-bit ip improves the objective . □ This proposition shows that local search in Q-BioLa t is governed by the collection of local elds { 𝑔 𝑘 ( 𝑥 ) } 𝑚 𝑘 = 1 . In particular , the interaction matrix 𝐽 determines how strongly each latent dimension depends on the current conguration of the others. A.8 Hamming Smoothness and Spectral Control W e next quantify how much the QUBO objective can change between two latent codes. Proposition (Hamming–Lipschitz continuity). For any 𝑥 , 𝑦 ∈ { 0 , 1 } 𝑚 , | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 ( 𝑦 ) | ≤ ( ∥ ℎ ∥ ∞ + ∥ 𝐽 ∥ ∞ ) 𝑑 𝐻 ( 𝑥 , 𝑦 ) , (20) where 𝑑 𝐻 ( 𝑥 , 𝑦 ) is the Hamming distance and ∥ 𝐽 ∥ ∞ = max 𝑘 Í 𝑚 ℓ = 1 | 𝐽 𝑘 ℓ | is the matrix innity norm. Proof. Let 𝛿 = 𝑥 − 𝑦 . Then ∥ 𝛿 ∥ 1 = 𝑑 𝐻 ( 𝑥 , 𝑦 ) and ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 ( 𝑦 ) = ℎ ⊤ 𝛿 + 1 2 𝛿 ⊤ 𝐽 ( 𝑥 + 𝑦 ) . (21) Therefore , | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 ( 𝑦 ) | ≤ ∥ ℎ ∥ ∞ ∥ 𝛿 ∥ 1 + 1 2 ∥ 𝛿 ∥ 1 ∥ 𝐽 ∥ ∞ ∥ 𝑥 + 𝑦 ∥ ∞ . Since 𝑥 , 𝑦 ∈ { 0 , 1 } 𝑚 , we have ∥ 𝑥 + 𝑦 ∥ ∞ ≤ 2 , which yields the result. □ This result shows that the quantity 𝐿 𝐻 : = ∥ ℎ ∥ ∞ + ∥ 𝐽 ∥ ∞ (22) acts as a global smoothness constant of the landscape in Hamming space. Corollary (Spe ctral control of landscap e variation). For any 𝑥 , 𝑦 ∈ { 0 , 1 } 𝑚 , | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 ( 𝑦 ) | ≤ 𝑑 𝐻 ( 𝑥 , 𝑦 ) ∥ ℎ ∥ 2 + √ 𝑚 ∥ 𝐽 ∥ 2 , (23) where ∥ 𝐽 ∥ 2 denotes the spectral norm of 𝐽 . Proof. Using the same decomposition with 𝛿 = 𝑥 − 𝑦 , | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 ( 𝑦 ) | ≤ ∥ ℎ ∥ 2 ∥ 𝛿 ∥ 2 + 1 2 ∥ 𝛿 ∥ 2 ∥ 𝐽 ∥ 2 ∥ 𝑥 + 𝑦 ∥ 2 . Now ∥ 𝛿 ∥ 2 ≤ 𝑑 𝐻 ( 𝑥 , 𝑦 ) and ∥ 𝑥 + 𝑦 ∥ 2 ≤ 2 √ 𝑚 , which implies the bound. □ This corollary directly motivates the spectral norm of 𝐽 as a global diagnostic of landscape roughness. In particular , smaller ∥ 𝐽 ∥ 2 implies smaller worst-case variation across the hypercube. Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization A.9 Ruggedness via Bit-Flip V ariability T o study local ruggedness, we analyze the variability of single-bit moves ov er the hypercube. Proposition (Exact second moment of bit-f lip gains). Assume 𝑥 is drawn uniformly from { 0 , 1 } 𝑚 . Then for each bit 𝑘 , E [ Δ 𝑘 ( 𝑥 ) ] = 0 , (24) and E [ Δ 𝑘 ( 𝑥 ) 2 ] = ℎ 𝑘 + 1 2 ℓ ≠ 𝑘 𝐽 𝑘 ℓ ! 2 + 1 4 ℓ ≠ 𝑘 𝐽 2 𝑘 ℓ . (25) Proof. Using Δ 𝑘 ( 𝑥 ) = ( 1 − 2 𝑥 𝑘 ) 𝑔 𝑘 ( 𝑥 ) , note that ( 1 − 2 𝑥 𝑘 ) is independent of 𝑔 𝑘 ( 𝑥 ) and has mean zero and squared value one under the uniform measure. Hence , E [ Δ 𝑘 ( 𝑥 ) ] = E [ 1 − 2 𝑥 𝑘 ] E [ 𝑔 𝑘 ( 𝑥 ) ] = 0 , and, E [ Δ 𝑘 ( 𝑥 ) 2 ] = E [ 𝑔 𝑘 ( 𝑥 ) 2 ] . Since the coordinates { 𝑥 ℓ } ℓ ≠ 𝑘 are independent Bernoulli ( 1 / 2 ) variables, E [ 𝑔 𝑘 ( 𝑥 ) ] = ℎ 𝑘 + 1 2 ℓ ≠ 𝑘 𝐽 𝑘 ℓ , V ar ( 𝑔 𝑘 ( 𝑥 ) ) = 1 4 ℓ ≠ 𝑘 𝐽 2 𝑘 ℓ . The result follows from E [ 𝑔 𝑘 ( 𝑥 ) 2 ] = V ar ( 𝑔 𝑘 ( 𝑥 ) ) + E [ 𝑔 𝑘 ( 𝑥 ) ] 2 . □ This proposition yields an exact measure of local rugged- ness. W e dene the average ruggedness: R ( 𝐽 , ℎ ) = 1 𝑚 𝑚 𝑘 = 1 E [ Δ 𝑘 ( 𝑥 ) 2 ] . (26) When the latent bits are approximately balance d and the local elds are centered, the dominant contribution is: R ( 𝐽 , ℎ ) ≈ 1 4 𝑚 ∥ 𝐽 ∥ 2 𝐹 , (27) showing that interaction energy directly drives lo cal variabil- ity . Thus, both ∥ 𝐽 ∥ 𝐹 and ∥ 𝐽 ∥ 2 are theoretically meaningful diagnostics of ruggedness. A.10 Low-Rank Structure and Eective Optimization Dimension The sp ectral structure of 𝐽 determines whether the land- scape is eectively go verned by a small numb er of collective directions. Let 𝐽 = 𝑚 𝑖 = 1 𝜆 𝑖 𝑢 𝑖 𝑢 ⊤ 𝑖 (28) be the spe ctral decomposition of 𝐽 , with eigenvalues ordered by decreasing magnitude: | 𝜆 1 | ≥ | 𝜆 2 | ≥ · · · ≥ | 𝜆 𝑚 | . For a given rank 𝑟 , we dene the truncated interaction ma- trix: 𝐽 𝑟 = 𝑟 𝑖 = 1 𝜆 𝑖 𝑢 𝑖 𝑢 ⊤ 𝑖 , (29) and the corresponding truncated objective: ˆ 𝑓 𝑟 ( 𝑥 ) = ℎ ⊤ 𝑥 + 1 2 𝑥 ⊤ 𝐽 𝑟 𝑥 . (30) Proposition (Pointwise low-rank approximation). For every 𝑥 ∈ { 0 , 1 } 𝑚 , | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 𝑟 ( 𝑥 ) | ≤ 𝑚 2 ∥ 𝐽 − 𝐽 𝑟 ∥ 2 . (31) Proof. W e have: ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 𝑟 ( 𝑥 ) = 1 2 𝑥 ⊤ ( 𝐽 − 𝐽 𝑟 ) 𝑥 . Therefore , | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 𝑟 ( 𝑥 ) | ≤ 1 2 ∥ 𝐽 − 𝐽 𝑟 ∥ 2 ∥ 𝑥 ∥ 2 2 . Since 𝑥 ∈ { 0 , 1 } 𝑚 , we have ∥ 𝑥 ∥ 2 2 ≤ 𝑚 , which pr oves the result. □ Corollary (Optimization gap under low-rank trun- cation). Let 𝑥 ★ ∈ arg max 𝑥 ˆ 𝑓 ( 𝑥 ) and 𝑥 ★ 𝑟 ∈ arg max 𝑥 ˆ 𝑓 𝑟 ( 𝑥 ) . Then, ˆ 𝑓 ( 𝑥 ★ ) − ˆ 𝑓 ( 𝑥 ★ 𝑟 ) ≤ 𝑚 ∥ 𝐽 − 𝐽 𝑟 ∥ 2 . (32) Proof. From the pr evious proposition, | ˆ 𝑓 ( 𝑥 ) − ˆ 𝑓 𝑟 ( 𝑥 ) | ≤ 𝜀 for all 𝑥 , where 𝜀 = 𝑚 2 ∥ 𝐽 − 𝐽 𝑟 ∥ 2 . Hence, ˆ 𝑓 ( 𝑥 ★ ) ≤ ˆ 𝑓 𝑟 ( 𝑥 ★ ) + 𝜀 ≤ ˆ 𝑓 𝑟 ( 𝑥 ★ 𝑟 ) + 𝜀 ≤ ˆ 𝑓 ( 𝑥 ★ 𝑟 ) + 2 𝜀 . Substituting the value of 𝜀 yields the claim. □ This corollary provides a concrete guarantee: if the spec- tral tail of 𝐽 is small, then the optimization landscape is eectively low-dimensional, and the dominant eigenmodes of 𝐽 capture most of the optimization-relevant structure. A useful scalar summary of this concentration is the eec- tive rank : 𝑟 e ( 𝐽 ) = ∥ 𝐽 ∥ 2 𝐹 ∥ 𝐽 ∥ 2 2 , (33) which is small when interaction energy is concentrated in a few dominant modes. A.11 Identiability of the Learne d QUBO Landscap e Let 𝜙 ( 𝑥 ) ∈ R 𝑝 denote the QUBO feature map containing all linear and pairwise terms, where 𝑝 = 𝑚 + 𝑚 ( 𝑚 − 1 ) 2 . (34) Let Φ ( 𝑋 ) ∈ R 𝑁 × 𝑝 be the design matrix formed from the observed latent codes in the training set. Truong-Son Hy Proposition (Non-identiability under rank deciency). If rank ( Φ ( 𝑋 ) ) < 𝑝 , then the QUBO parameters are not uniquely identiable from the observed data. In particular , there exists a nonzero v ector 𝑢 ∈ R 𝑝 such that: Φ ( 𝑋 ) 𝑢 = 0 . (35) Therefore, for any parameter vector 𝑤 , both 𝑤 and 𝑤 + 𝑢 induce identical predictions on all observed training codes, but can dier on unseen binary co des. Proof. If rank ( Φ ( 𝑋 ) ) < 𝑝 , then the null space of Φ ( 𝑋 ) is nontrivial, so there exists 𝑢 ≠ 0 with Φ ( 𝑋 ) 𝑢 = 0 . Hence, Φ ( 𝑋 ) ( 𝑤 + 𝑢 ) = Φ ( 𝑋 ) 𝑤 , so the two parameterizations agree on all observed codes. For an unseen code 𝑥 with 𝜙 ( 𝑥 ) ⊤ 𝑢 ≠ 0 , the predicted tness diers: 𝜙 ( 𝑥 ) ⊤ ( 𝑤 + 𝑢 ) ≠ 𝜙 ( 𝑥 ) ⊤ 𝑤 . □ Corollary (Dimension–sample trade-o ). Since 𝑝 = O ( 𝑚 2 ) , increasing the latent dimension rapidly enlarges the number of QUBO parameters. When the number of observed sequences is not suciently large r elative to 𝑝 , the landscape away from the observed codes is only weakly constrained. This result provides a formal explanation for the trade- o between latent dimensionality , surrogate generalization, and optimization stability . Ridge regularization selects one stable solution among many p ossibilities, but the induced landscape remains strongly dependent on the r epresentation through the feature map Φ ( 𝑋 ) . A.12 Why Prediction and Optimization Can Decouple The previous results formalize why predictive accuracy alone does not determine optimization behavior . First, predictive metrics such as Spearman correlation eval- uate surrogate quality on a nite observed set, whereas opti- mization depends on the geometry of the entire hypercube. Second, our propositions show that when the QUBO fea- ture map is rank-decient or weakly constrained, multiple interaction structures can be consistent with essentially the same obser ved t while diering away from the obser ved manifold. T ogether , these observations imply that two latent repre- sentations can achieve similar predictive accuracy while in- ducing substantially dierent optimization landscapes. This representation–optimization decoupling is not an anomaly , but rather a structural consequence of learning a surrogate over a large discrete latent space . A.13 Practical Consequences The theor y above yields several experimentally testable pre- dictions. Landscapes with smaller ∥ 𝐽 ∥ 2 , faster spectral decay , and smaller ruggedness R ( 𝐽 , ℎ ) should be easier to optimize. Likewise, increasing latent dimensionality without sucient data should increase ambiguity in the learned landscape and make optimization less stable. These further empirical analysis of the theory will be included in follow-up future works. B Additional Experimental Details B.1 Protein Sequence Emb eddings (ESM) W e represent protein sequences using pretrained protein language models from the ESM family . In particular , we use the facebook/esm2_t6_8M_UR50D checkpoint from the Hugging Face Transformers library . Given a protein sequence, we rst convert it to uppercase and remove non-alphabetic characters. The cleaned sequence is then tokenize d using the corresponding ESM tokenizer . W e apply padding and truncation with a maximum sequence length of 1024. The tokenized sequence is passe d through the pretrained transformer model in evaluation mo de without gradient com- putation. The model produces contextualized residue-level representations, from which we obtain a xed-length se- quence embedding by mean pooling over the sequence di- mension using the attention mask: 𝑒 = Í 𝐿 𝑖 = 1 𝑚 𝑖 𝐻 𝑖 Í 𝐿 𝑖 = 1 𝑚 𝑖 , (36) where 𝐻 𝑖 denotes the hidden representation of the 𝑖 -th residue and 𝑚 𝑖 is the corresponding attention mask. Embeddings are computed in mini-batches of size 8 and stored as dense vectors in float32 format. These embed- dings ar e used as inputs to downstream comp onents, in- cluding the external tness oracle and latent representation learning. B.2 External Sequence-Level Fitness Oracle For sequence-level tness evaluation, we trained an external oracle on dense protein sequence embeddings. Each dataset was stored as a NumPy ar chive containing (i) dense em- beddings, (ii) scalar tness labels, and (iii) the correspond- ing protein sequences. The emb eddings were represented in float32 , while tness targets were stored in float64 . Data spliing. W e partitioned each dataset into training, validation, and test sets using a two-stage random split. First, we split the data into a training + validation set and a held- out test set with a test ratio of 0 . 2 . Then, we further split the training + validation set into training and validation subsets using a validation ratio of 0 . 1 relative to the full dataset. This results in an eective split of 70% / 10% / 20% for training, validation, and test sets, respe ctively . All splits were performed with a xed random seed of 42 for reproducibility . Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization Evaluation metrics. W e evaluated model performance using four standard regr ession metrics: Sp earman correla- tion, Pearson correlation, root mean squared err or (RMSE), and mean absolute error (MAE). Metrics were reported on the training, validation, and test sets. Models. W e considered three regression models on top of the same embe dding representation: Ridge Regression, XGBoost, and Gaussian Process Regr ession. Ridge Regression. W e used a pipeline consisting of fea- ture standardization via StandardScaler followed by Ridge regression. The regularization strength was set to 𝛼 = 1 . 0 , and the svd solver was used. XGBoost. W e used the XGBRegressor implementation with the following hyperparameters: number of trees = 300 , maximum tree depth = 6 , learning rate = 0 . 05 , subsampling ratio = 0 . 9 , column subsampling ratio = 0 . 9 , ℓ 1 regularization coecient = 0 . 0 , and ℓ 2 regularization coecient = 1 . 0 . The objective function was squared error regression. Training was performed using 4 CP U threads, and the random seed was xed to 42. Gaussian Process Regression. W e used a pipeline con- sisting of feature standardization follow ed by Gaussian Pro- cess Regression (GPR). The kernel was dened as: 𝑘 ( 𝑥 , 𝑥 ′ ) = 𝐶 · RBF ( 𝑥 , 𝑥 ′ ) + WhiteKernel , (37) where the constant kernel 𝐶 was initialized to 1 . 0 with b ounds [ 10 − 3 , 10 3 ] , the RBF kernel had an initial length scale of 1 . 0 with bounds [ 10 − 3 , 10 3 ] , and the white noise level was ini- tialized to 10 − 3 with b ounds [ 10 − 6 , 10 1 ] . The GPR mo del used normalize_y=True and performed 2 r estarts of the optimizer . Training protocol. All models were trained on b oth GFP and AA V datasets under four data regimes with training set sizes of { 1000 , 2000 , 5000 , 10000 } . For each dataset and data regime, the same xe d hyperparameters and random seed were used. No additional hyperparameter search was performed. Mo dels were trained on the training split and evaluated on validation and test splits. Outputs. For each trained model, we stored (i) the se- rialized mo del parameters, (ii) evaluation metrics in JSON format, and (iii) test-set predictions including sequences, ground-truth tness values, and predicted tness values. B.3 Latent Representation Learning and Deco der Implementation W e construct binar y latent representations from dense ESM embeddings using four appr oaches: PCA, random projection, deterministic autoencoder ( AE), and variational autoencoder (V AE). For all methods, we e valuate latent dimensions of 8, 16, 32, and 64. For PCA, we apply principal component analysis to the dense embeddings and retain the top components corre- sponding to the target latent dimension. The resulting con- tinuous representations ar e binarized using per-dimension median thresholding computed over the dataset. PCA -based binary representations are precomputed and reused across experiments. For random projection, we project emb eddings using a Gaussian random matrix with variance scaled by the input dimension, follow ed by the same median-based binariza- tion. This provides a lightweight baseline without learned structure. For AE and V AE, we use simple fully connected neural networks. The autoencoder consists of a two-layer encoder and decoder with a hidden dimension of 256 and ReLU ac- tivations. The encoder maps the input embedding to the latent space, and the decoder reconstructs the original em- bedding. The V AE uses a similar architecture with a shared encoder backb one and separate linear heads for the mean and log-variance. Both models ar e trained using mean squared reconstruction loss, with an additional KL divergence term for the V AE weighted by 𝛽 = 10 − 3 . Latent representations are binarized by thresholding at zero . Both AE and V AE are trained using the Adam optimizer with learning rate 10 − 3 , weight decay 10 − 5 , batch size 32, and 200 training epochs. All mo dels are trained with a xed random seed of 42 on CP U . For decoding, we train a mutation-conditioned neural de- coder that maps latent co des back to protein sequences. The decoder consists of two fully connected layers with hidden dimension 256 and ReLU activations, followed by two output heads: one for predicting mutation positions and one for predicting amino acid identities at mutate d positions. The mutation prediction head outputs a vector of length equal to the sequence length, while the amino acid head outputs logits over 20 amino acids per position. The de coder is traine d using a combination of binar y cross- entropy loss for mutation prediction and cross-entrop y loss for amino acid prediction, with a higher weight assigned to the mutation loss to account for class imbalance. T raining is performed using Adam with learning rate 10 − 3 , batch size 32, and 100 epochs. The best model is selected based on validation performance. All latent models and de coders are trained across both GFP and AA V datasets and all data regimes using consistent hyperparameters. B.4 Internal QUBO Surrogate Implementation The internal surrogate op erates directly on binary latent representations of dimension 𝑚 ∈ { 8 , 16 , 32 , 64 } and models the tness function as a combination of linear and pair wise interaction terms. Given a binary latent vector , the surrogate Truong-Son Hy includes all 𝑚 linear terms and all 𝑚 ( 𝑚 − 1 ) 2 pairwise inter- action terms. These features are constructed explicitly by concatenating the original binary variables with all pairwise products between distinct latent dimensions. The model parameters are tte d using ridge regression with ℓ 2 regularization. Specically , the surrogate is trained by solving a linear system involving the feature matrix and the target tness values, with a regularization coecient set to 10 − 3 . This closed-form solution enables ecient training even when the number of pairwise features grows quadrati- cally with the latent dimension. After training, the learned parameters are decomposed into a vector of linear coecients and a symmetric matrix of pairwise interaction coecients. The diagonal entries of the interaction matrix are set to zero, and symmetry is enforced by assigning equal weights to ( 𝑖 , 𝑗 ) and ( 𝑗 , 𝑖 ) . Predictions are computed by combining the linear contributions and pairwise interactions over the binary latent variables. W e evaluate the surrogate using standard regression met- rics including RMSE, 𝑅 2 , and Spearman correlation on both training and held-out test sets. The model is traine d using a xed random seed of 42 and a train/test split ratio of 80/20. For comparison, the implementation also includes a small multilayer p erceptron (MLP) baseline traine d using mini- batch gradient descent with ReLU activations. The MLP uses two hidden layers with dimensions 64 and 32, learning rate 10 − 3 , weight decay 10 − 5 , batch size 64, and 400 training epochs. However , this baseline is used only for reference, while all optimization experiments in the main paper rely on the QUBO surrogate. The surrogate is trained across all dataset sizes and latent dimensions and serves as the objective function for down- stream combinatorial optimization methods. B.5 Combinatorial Optimization Metho ds W e optimize the learned QUBO surrogate in the binary latent space using ve methods: Simulated Annealing (SA), Genetic Algorithm (GA), Random Search (RS), Greedy Hill Climbing (GHC), and a lightweight latent Bayesian optimization (LBO) baseline. All methods operate directly on binary vectors of dimension 𝑚 ∈ { 8 , 16 , 32 , 64 } . Simulated Annealing (SA ) starts from an initial binar y code and iteratively proposes single-bit ips. At each step, a bit index is selected uniformly at random, and the energy change induced by ipping that bit is computed eciently using the learned QUBO parameters. Moves that improve the obje ctive are always accepted, while worse mov es are accepted with a temperature-dependent probability . The tem- perature follows an exponential decay schedule with initial temperature 𝑇 0 = 1 . 0 , minimum temperature 10 − 4 , and decay factor 0 . 999 . The algorithm runs for 20,000 steps and keeps track of the best solution encountered during the traje ctory . The Genetic Algorithm (GA) maintains a population of candidate binary solutions and evolves them over multiple generations. W e use a population size of 64 and run the al- gorithm for 150 generations. At each generation, a subset of top-performing individuals (elite size 4) is preserved. New individuals are generated using tournament selection with tournament size 3, followed by single-point crossover ap- plied with probability 0.9. Ospring are further perturbed using independent bit-ip mutation with mutation rate 0.02. The population is updated at each generation, and the best solution across all generations is returned. Random Search (RS) samples binary latent codes indepen- dently and uniformly at random. W e evaluate 10,000 candi- date solutions and return the one with the highest obje ctive value. Greedy Hill Climbing (GHC) starts from an initial binary code and iteratively improves it by exhaustively evaluating all single-bit ips. At each iteration, the algorithm selects the bit ip that yields the largest improv ement in the objective and updates the current solution accordingly . The process is repeated until no improving move exists or until a maximum of 100 passes over all bits is reached. The Latent Bayesian Optimization (LBO ) baseline uses a kernel-based uncertainty heuristic over binary latent codes. Given a set of seed solutions, the method samples 5,000 candidate binary vectors uniformly at random and e valuates an acquisition function based on an upper condence b ound. The predictive mean is computed using a kernel-weighted average over seed points, where the kernel is dened using an RBF function over Hamming distance with length scale 4.0. The acquisition function is given by the sum of the predictive mean and an exploration term weighted by 𝛽 = 1 . 0 . The candidate with the highest acquisition value is selected. All optimization methods are applied consistently across datasets, latent dimensions, and data regimes using xed hyperparameters. B.6 Additional Experimental Results W e provide detailed end-to-end design results across all dataset sizes and latent dimensions in Figures 5 and 6, as well as T ables 10–17. Figure 5 presents the performance trends on the GFP dataset across dierent training regimes, while Figure 6 shows the corresponding results for the AA V dataset. These gures illustrate how performance varies with latent dimensionality and optimization method. T ables 10– 13 r eport detailed r esults for the GFP dataset at training sizes of 1000, 2000, 5000, and 10000 samples, respectively , while T ables 14–17 present the corresponding results for the AA V dataset across all data regimes. Each table includes both the best achieved score and the average performance of the top-10 candidates for each conguration, enabling a more ne-grained comparison across latent representations, optimization methods, and data regimes. Overall, these addi- tional results further corroborate the trends observed in the main text, including the consistent advantage of PCA -based Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization latent representations and the strong performance of combi- natorial optimization methods such as greedy hill climbing, genetic algorithms, and simulated annealing. Truong-Son Hy (a) GFP with 1000 training samples. (b) GFP with 2000 training samples. (c) GFP with 5000 training samples. (d) GFP with 10000 training samples. Figure 5. End-to-end design performance on GFP across dierent training sizes. The x-axis shows the number of bits and the y-axis shows the best oracle score. Each line corr esponds to an optimizer-representation pair , including simulated annealing (SA), genetic algorithm ( GA), random search (RS), and greedy hill climbing (GHC), combined with either PCA -based or random-projection binary latent representations. T able 10. Detailed end-to-end design results on GFP with 1000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA LBO 2.616 2.524 Random LBO 2.612 2.531 16 PCA RS 3.950 3.247 Random RS 3.598 2.488 32 PCA GHC 4.032 3.017 Random GA 3.690 2.525 64 PCA RS 4.119 3.903 Random GA 4.170 3.312 Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization (a) AA V with 1000 training samples. (b) AA V with 2000 training samples. (c) AA V with 5000 training samples. (d) AA V with 10000 training samples. Figure 6. End-to-end design p erformance on AA V across dierent training sizes. The x-axis shows the number of bits and the y-axis shows the best oracle score. Each line corr esponds to an optimizer-representation pair , including simulated annealing (SA), genetic algorithm ( GA), random search (RS), and greedy hill climbing (GHC), combined with either PCA -based or random-projection binary latent representations. T able 11. Detailed end-to-end design results on GFP with 2000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA GA 2.669 2.669 Random LBO 2.669 2.267 16 PCA GA 3.503 3.212 Random LBO 3.002 2.358 32 PCA GA 3.909 3.449 Random GHC 3.448 2.693 64 PCA GA 4.027 3.549 Random LBO 3.316 1.900 Truong-Son Hy T able 12. Detailed end-to-end design results on GFP with 5000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA GA 2.653 2.653 Random LBO 2.653 2.633 16 PCA GHC 3.656 3.117 Random LBO 3.568 2.356 32 PCA LBO 4.157 2.918 Random GHC 3.815 2.228 64 PCA GHC 4.270 3.972 Random LBO 3.330 2.619 T able 13. Detailed end-to-end design results on GFP with 10000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA LBO 3.162 -3.738 Random LBO 2.785 -4.556 16 PCA LBO 4.712 3.452 Random GHC 4.502 3.029 32 PCA GA 5.287 4.823 Random GHC 4.806 3.829 64 PCA GHC 5.554 5.042 Random GA 4.758 3.003 T able 14. Detailed end-to-end design results on AA V with 1000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA GA -1.687 -4.039 Random LBO -6.492 -7.534 16 PCA SA 2.484 -1.568 Random RS -0.913 -6.523 32 PCA GA 3.906 -0.680 Random SA 2.753 -3.336 64 PCA GHC 5.232 2.962 Random GHC 4.066 -1.557 Q-BioLa t : Binary Latent Protein Fitness Landscapes for QUBO-Based Optimization T able 15. Detailed end-to-end design results on AA V with 2000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA LBO -4.213 -6.054 Random LBO -3.352 -5.891 16 PCA RS 1.325 -0.440 Random LBO 1.769 -3.162 32 PCA RS 2.866 0.349 Random GHC 4.184 -2.674 64 PCA RS 1.779 -0.353 Random GHC 0.164 -1.511 T able 16. Detailed end-to-end design results on AA V with 5000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA LBO -1.727 -6.113 Random LBO -2.616 -5.168 16 PCA RS -0.245 -2.136 Random GHC 0.610 -2.630 32 PCA GA 2.929 0.666 Random RS 1.336 -5.012 64 PCA SA 3.999 2.249 Random GA 2.548 -1.150 T able 17. Detailed end-to-end design results on AA V with 10000 training samples. Bits Representation Best optimizer Best score ↑ T op-10 mean ↑ 8 PCA LBO -1.903 -3.723 Random LBO -5.172 -7.184 16 PCA RS -0.237 -2.834 Random LBO 1.197 -4.205 32 PCA SA 1.904 0.727 Random LBO -0.154 -5.185 64 PCA GA 4.541 2.091 Random GHC 0.644 -2.174
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment