Project Imaging-X: A Survey of 1000+ Open-Access Medical Imaging Datasets for Foundation Model Development
Foundation models have demonstrated remarkable success across diverse domains and tasks, primarily due to the thrive of large-scale, diverse, and high-quality datasets. However, in the field of medical imaging, the curation and assembling of such med…
Authors: Zhongying Deng, Cheng Tang, Ziyan Huang
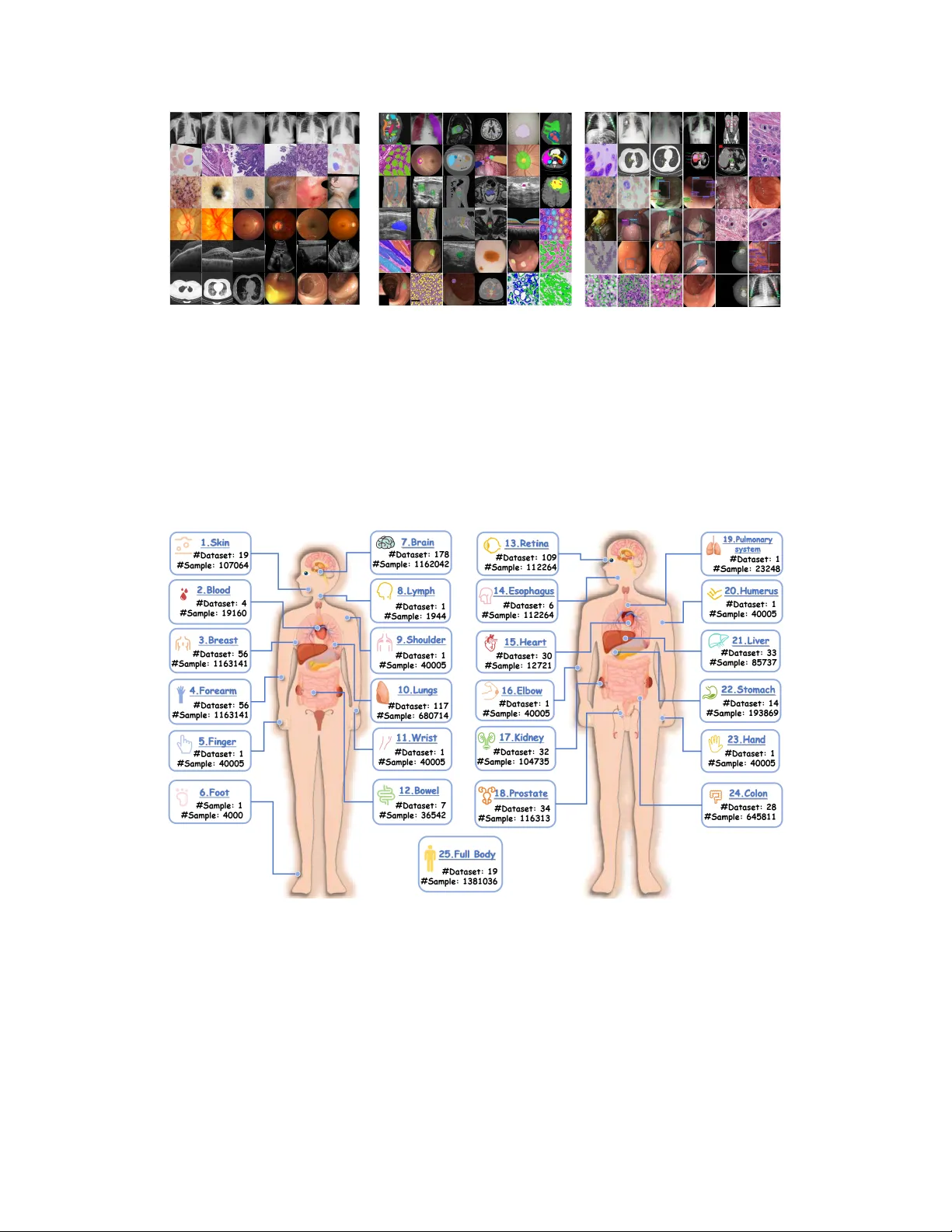
Pr oject Imaging-X: A Surv ey of 1000+ Open-Access Medical Imaging Datasets f or F oundation Model De v elopment Zhongying Deng 1 , 5 , ∗ Cheng T ang 1 , 3 , ∗ Ziyan Huang 1 , ∗ Jiashi Lin 1 , ∗ Y ing Chen 1 , ∗ Junzhi Ning 1 , ∗ Chenglong Ma 2 , 4 , ∗ Jiyao Liu 1 , 4 W ei Li 1 , 6 Y inghao Zhu 7 Shujian Gao 1 Y an yan Huang 7 Sibo Ju 8 Y anzhou Su 8 , 14 Pengcheng Chen 1 , 9 W enhao T ang 1 T ianbin Li 1 Haoyu W ang 1 , 6 Y uanfeng Ji 10 Hui Sun 1 Shaobo Min 21 Liang Peng 7 Feilong T ang 1 , 12 Haochen Xue 1 Rulin Zhou 1 Chaoyang Zhang 2 , 45 W enjie Li 2 , 6 , 13 Shaohao Rui 2 , 6 W eijie Ma 2 , 4 Xingyue Zhao 14 Y ibin W ang 2 , 4 Kun Y uan 1 Zhaohui Lu 6 Shujun W ang 15 Jinjie W ei 1 , 4 Lihao Liu 1 Dingkang Y ang 4 Lin W ang 1 Y ulong Li 1 Haolin Y ang 1 Y iqing Shen 1 Lequan Y u 7 Xiaowei Hu 16 Y un Gu 6 Y icheng Wu 12 Benyou W ang 17 Minghui Zhang 6 Angelica I. A viles-Riv ero 18 Qi Gao 4 Hongming Shan 4 Xiaoyu Ren 19 Fang Y an 1 Hongyu Zhou 20 Haodong Duan 21 Maosong Cao 1 Shanshan W ang 19 , 22 Bin Fu 1 Xiaomeng Li 23 Zhi Hou 1 Chunfeng Song 1 Lei Bai 1 Y uan Cheng 24 , 25 Y uandong Pu 1 , 6 Xiang Li 26 W enhai W ang 27 Hao Chen 23 Jiaxin Zhuang 23 Songyang Zhang 1 Huiguang He 28 , 29 Mengzhang Li 1 Bohan Zhuang 30 Zhian Bai 13 Rongshan Y u 31 Liansheng W ang 31 Y ukun Zhou 32 Xiaosong W ang 1 Xin Guo 25 Guanbin Li 33 Xiangru Lin 7 Dakai Jin 34 Mianxin Liu 1 W enlong Zhang 1 Qi Qin 1 Conghui He 1 Y uqiang Li 1 Y e Luo 35 Nanqing Dong 1 Jie Xu 1 W enqi Shao 1 Bo Zhang 1 Qiujuan Y an 1 Y ihao Liu 1 Jun Ma 36 Zhi Lu 37 Y uewen Cao 1 Zongwei Zhou 38 Jianming Liang 39 Shixiang T ang 1 Qi Duan 40 Dongzhan Zhou 1 Chen Jiang 24 , 25 Y uyin Zhou 41 Y anwu Xu 16 Jiancheng Y ang 42 , 43 Shaoting Zhang 6 Xiaohong Liu 2 , 6 Siqi Luo 1 , 6 Y i Xin 1 , 2 Chaoyu Liu 5 Haochen W en 5 , 32 Xin Chen 44 Alejandro Lozano 10 Min W oo Sun 10 Y uhui Zhang 10 Y ue Y ao 44 Xiaoxiao Sun 10 Serena Y eung-Levy 10 Xia Li 6 Jing Ke 6 Chunhui Zhang 6 Zongyuan Ge 12 Ming Hu 1 , 12 , † Jin Y e 1 , 12 , † Zhifeng Li 11 , † Y irong Chen 1 , † Y u Qiao 1 , 2 † Junjun He 1 , 2 , † 1 Shanghai Artificial Intelligence Laboratory; 2 Shanghai Innov ation Institute; 3 Shanghai Institute of Optics and Fine Mechanics; 4 Fudan Univ ersity; 5 Univ ersity of Cambridge; 6 Shanghai Jiao T ong Univ ersity; 7 The Univ ersity of Hong K ong; 8 Fuzhou Univ ersity; 9 Univ ersity of W ashington; 10 Stanford Univ ersity; 11 Incept Labs; 12 Monash Univ ersity; 13 Ruijin Hospital, Shanghai Jiao T ong Univ ersity School of Medicine; 14 Alibaba D AMO Academy; 15 The Hong K ong Polytechnic Univ ersity; 16 South China Univ ersity of T echnology; 17 The Chinese Univ ersity of Hong K ong, Shenzhen; 18 Y au Mathematical Sciences Center, Tsinghua Uni versity; 19 Chinese Academy of Sciences; 20 Tsinghua Univ ersity; 21 Independent Researcher; 22 Shenzhen Institute of Advanced T echnology , Chinese Academy of Sciences; 23 The Hong K ong Univ ersity of Science and T echnology; 24 Artificial Intelligence Innov ation and Incubation Institute, Fudan Uni versity; 25 Shanghai Academy of Artificial Intelligence for Science; 26 Nankai Univ ersity; 27 The Chinese Univ ersity of Hong K ong; 28 Institute of Automation, Chinese Academy of Sciences; 29 Univ ersity of Chinese Academy of Sciences; 30 Zhejiang Univ ersity; 31 School of Informatics, Xiamen Univ ersity; 32 Univ ersity College London; 33 Sun Y at-sen University; 34 Alibaba Group, D AMO Academy , New Y ork, NY , USA; 35 T ongji Univ ersity; 36 Univ ersity of T oronto; 37 Department of Psychological and Cognitiv e Sciences, Tsinghua Uni versity; 38 Johns Hopkins Univ ersity; 39 Arizona State Univ ersity; 40 Academy for Clinical Innov ation and T ranslation of Shanghai; 41 Univ ersity of California, Santa Cruz; 42 ELLIS Institute Finland; 43 Aalto Univ ersity; 44 Shandong Univ ersity; 45 Xi’an Jiaotong Univ ersity ∗ ∗ Equal contribution † Corresponding author Abstract Foundation models have demonstrated remarkable success across di verse domains and tasks, primarily due to the thri ve of lar ge-scale, div erse, and high-quality datasets. Howe v er , in the field of medical imaging, the curation and assem- bling of such medical datasets are highly challenging due to the reliance on clin- ical expertise and strict ethical and priv ac y constraints, resulting in a scarcity of lar ge-scale unified medical datasets and hindering the de velopment of pow- erful medical foundation models. In this work, we present the largest surv ey to date of medical image datasets, co vering ov er 1,000 open-access datasets with a systematic catalog of their modalities, tasks, anatomies, annotations, limita- tions, and potential for integration. Our analysis e xposes a landscape that is mod- est in scale, fragmented across narro wly scoped tasks, and une venly distrib uted across or gans and modalities, which in turn limits the utility of existing medical image datasets for dev eloping versatile and robust medical foundation models. T o turn fragmentation into scale, we propose a metadata-driven fusion paradigm (MDFP) that systematically inte grates public datasets with shared modalities or tasks, thereby transforming multiple small data silos into larger , more coherent resources. Building on MDFP , we release an interactive discov ery portal that en- ables end-to-end, automated medical image dataset integration, and compile all surve yed datasets into a unified, structured table that clearly summarizes their key characteristics and provides reference links, of fering the community an ac- cessible and comprehensi ve repository . By charting the current terrain and of- fering a principled path to dataset consolidation, our survey pro vides a practical roadmap for scaling medical imaging corpora, supporting faster data discov ery , more principled dataset creation, and more capable medical foundation models for the biomedical imaging research community . Our project repository can be found at https://github.com/uni- medical/Project- Imaging- X . 2 Contents 1 Introduction 5 2 An Overview of Medical Image Datasets 7 2.1 T otal Growth . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 10 2.2 Imaging Dimensionalities . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 10 2.3 Imaging Modalities . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 11 2.4 T asks . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 12 2.5 Anatomical Regions . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 14 2.6 Summary . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 15 3 2D Medical Image Datasets 15 3.1 Overvie w . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 15 3.2 CT Modality . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 16 3.3 MRI Slices . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 17 3.4 PET Slices . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 19 3.5 Ultrasound (US) Images . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 20 3.6 X-Ray Images . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 21 3.7 Optical Coherence T omography (OCT) Images . . . . . . . . . . . . . . . . . . . 22 3.8 Fundus Images . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 23 3.9 Dermoscopy Images . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 24 3.10 Histopathology . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 24 3.11 Microscopy Imaging . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 25 3.12 Infrared Imaging . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 26 3.13 Endoscopy Imaging . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 26 3.14 Other Modalities . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 27 3.15 Challenge and Opportunity . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 28 4 3D Medical Image Datasets 29 4.1 Overvie w . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 29 4.2 CT V olumes . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 30 4.3 MRI V olumes . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 32 4.4 Ultrasound V olumes . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 33 4.5 PET V olumes . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 34 4.6 Other 3D V olumes . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 36 4.7 Challenges and Opportunities . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 37 5 Medical V ideo Datasets 39 5.1 Overvie w . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 39 5.2 T ask . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 39 3 5.2.1 Classification . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 39 5.2.2 Segmentation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 40 5.2.3 Detection . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 40 5.2.4 T racking . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 41 5.2.5 Estimation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 41 5.2.6 Registration . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 41 5.3 Modalities . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 42 5.4 Anatomical Structures . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 43 5.5 Challenges and Opportunities . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 43 6 Paradigm f or Dataset Fusion 44 6.1 Dataset Collection and Processing . . . . . . . . . . . . . . . . . . . . . . . . . . 45 6.2 MDFP . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 46 6.2.1 Phase 1: Metadata Harmonization . . . . . . . . . . . . . . . . . . . . . . 47 6.2.2 Phase 2: Semantic Alignment . . . . . . . . . . . . . . . . . . . . . . . . 47 6.2.3 Phase 3: Fusion Blueprints . . . . . . . . . . . . . . . . . . . . . . . . . 48 6.2.4 Phase 4: Dataset Inde xing and Community Sharing . . . . . . . . . . . . . 49 6.2.5 Case Study: Goal-Conditioned Fusion via MDFP . . . . . . . . . . . . . 49 6.3 Interactiv e Discov ery Portal . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 50 7 Discussion 51 7.1 Limitations in T ask Definition and Evolution of Data Engineering P aradigms . . . 51 7.2 Scarcity of Multimodal Medical Datasets and Constraints in Further Dev elopment . 51 7.3 Challenges and Opportunities in Medical Foundation Models . . . . . . . . . . . . 51 8 Conclusion 52 A T ables of 2D Medical Image Datasets 131 B T ables of 3D Medical Image Datasets 131 C T ables of Medical V ideo Datasets 155 4 Figure 1: Ev olution of medical foundation models and general domain foundation models. Medical foundation models are mostly trained using millions of images, while adv anced general domain ones are trained using billions of natural images. Additionally , most medical foundation models cov er only a fe w modalities like CT and MRI, which may introduce modality-specific bias that constrains clinical applicability . 1 Introduction Medical imaging foundation models hold the promise of significantly adv ancing clinical decision- making by analyzing di verse medical imaging modalities and e xecuting multiple tasks through a single, pre-trained system. This paradigm parallels the trajectory of adv anced models in the domain of natural language processing [ 1 ] and computer vision [ 2 , 3 , 4 , 5 ], which are trained on extensiv e and div erse datasets to achie ve broad generalization across tasks and applications [ 6 , 7 , 8 , 9 , 10 ], as depicted in Figure 1 . This highlights a similar shift in medical AI from narrow , single-modality , task-specific models tow ard multi-modal, multi-functional foundation, which could better reflect the complexity of clinical workflows and enhance utility across specialties [ 11 ]. Despite this po- tential, current medical imaging foundation models, such as STUNet [ 12 ], MedSAM [ 7 ], SAM- Med3D [ 13 ], SAM-Brain3D [ 14 ] and P anDerm [ 15 ], are often tailored to well-represented settings, such as a fe w modalities like computed tomography (CT) and magnetic resonance imaging (MRI), a narrow set of tasks ( e .g. , se gmentation), or limited anatomical regions ( e .g. , brain, abdomen). Man y clinically valuable settings remain less cov ered, which introduces modality-, task-, and anatomy- specific biases that constrain generalization and clinical applicability . The root challenge lies in data a vailability and di versity [ 16 , 17 ]. Most public medical datasets con- tain only thousands of images, e .g. , BraTS series [ 18 , 19 ], which are orders of magnitude smaller than natural image datasets with billions of samples, such as Segment Anything 1 Billion (SA- 1B) [ 5 ] and LAION-5B [ 20 ]. This substantial dif ference in the number of training images between the natural image (or general) domain and the medical one is further depicted in Figure 1 . Construct- ing large, di verse medical datasets is resource-intensi ve, requiring specialized imaging equipment, expert annotations, and careful navigation of ethical and pri v acy constraints. Consequently , the cur - rent dataset landscape is highly fragmented , with data scattered across isolated, narrowly scoped collections [ 21 ]. This fragmentation not only limits pre-training scale, b ut also overlooks opportu- nities to integrate related datasets into richer , more balanced training resources. A promising direction emer ging in recent research is dataset integration [ 22 , 23 ], where multiple smaller datasets with shared modalities, anatomies, or tasks are merged into unified large-scale resources. As sho wn in Figure 2 , the merged datasets can bridge data and models, facilitating the dev elopment of foundation models [ 24 ]. While this strategy has shown potential, e xisting efforts 5 M oda l i t y : C T A T a s k : C l a s s i f i ca t i o n Ana t o m y : B r a i n , a or t a , . . . M oda l i t y : C T , M R I , P E T T a s k : S e gm e nt a t i on Ana t o m y : B r a i n , a bdom e n, . . . M oda l i t y : C T , M R I T a s k : R e gi s t r a t i on Ana t o m y : B r a i n , che s t , . . . M oda l i t y : M R I T a s k : D e t e ct i on An a t o m y : B r a i n , a bdom e n, . . . D a ta s et 1 D a ta s e t 2 D a ta s et 3 D a ta s et 4 D a ta s et N D a t a s il os W i t h i s o l a t ed m o d a l i t i es , t a s k s , a n a t o m i e s , et c . D a t a s e t int e g r a t ion O u r c o n t ri b u t i o n , b r i d gi n g d a t a a n d m o d e l s w i t h u n i f i ed d a t a t a x o n o m y Fou n d a t ion mod e ls S o l v i n g m u l t i - m o d a l i t y , m u l t i - t a s k , m u l t i - a n a t o m y c h a l l en ges Mo d a l i t y : C T , M RI , P E T , C T A , ... T a s k : C l a s s i f i c a ti o n , s e g m e n ta ti o n , d et ec ti o n , ... A n a t om y : B r a i n , c h es t, a b d o m en , p el v i s , ... T a r g et a n a t om y : B r a i n D a ta s et 1 D a ta s e t 2 D a ta s et 3 D a ta s et 4 ... CT M RI C T A ... C l a s s i f i c a ti o n Seg m en ta ti o n Reg i s tr a ti o n ... T a r g et d i s e a s e : L i v e r t u m o r . . . . . . D a ta s et 2 D a ta s et 3 D a ta s et 4 D a ta s et 5 ... CT M RI C T A ... C l a s s i f i c a ti o n Seg m en ta ti o n D et ec ti o n ... Figure 2: Conceptual o vervie w of moving from fragmented medical image data to integrated re- sources for medical foundation models. Our surve y addresses the data fragmentation issue in public medical image datasets by introducing a metadata-driv en dataset integration paradigm, which is cru- cial for the development of advanced foundation models that can tackle multi-modality , multi-task, and multi-anatomy challenges effecti v ely , ultimately enhancing clinical AI applications. typically focus on specific imaging types or organ systems [ 25 , 26 , 27 ]. Furthermore, when lacking guidance from a comprehensive o vervie w of av ailable datasets, dataset integration risks reinforcing existing biases rather than enabling balanced, general-purpose foundation model de velopment. Giv en these challenges, a comprehensi v e surv ey of medical imaging datasets is urgently needed. Such a survey can illustrate gaps in data cov erage, highlight opportunities for dataset inte gration, and establish a standardized framew ork for dataset selection and e valuation, which are crucial for the development of robust medical foundation models. A few prior surveys hav e revie wed medical imaging datasets [ 28 , 29 , 30 , 31 , 32 ], yet the y often lack subject- and image-lev el statistics, omit many recently released large-scale datasets such as T otalSegmentor [ 33 ] and AbdomenAtlas [ 34 ], and do not provide a systematic framework that links dataset characteristics to the requirements of foundation model dev elopment. T o address these limitations, we present the most comprehensi ve re vie w to date of ov er 1,000 open- access medical imaging datasets published between 2000 and 2025. W e introduce a no vel taxonomy to or ganize datasets by modality , anatomy , task, and label a v ailability . Lev eraging this taxonomy , we conduct a gap analysis to identify underrepresented modalities, tasks, and anatomies, establish- ing clear priorities for future dataset creation. Building on these insights, we further propose a metadata-driv en fusion paradigm (MDFP) for inte grating existing datasets, incorporating it into our interactiv e discov ery portal 2 that enables end-to-end process of fine-grained search, statistical anal- ysis, and dataset integration. W e conclude with a forward-looking discussion on the challenges and opportunities tow ard building truly general-purpose medical imaging foundation models. Our main contributions are summarized as follo ws: • Comprehensi ve large-scale sur vey : W e provide the most extensi v e re view to date, cov er- ing o ver 1,000 open-access medical image datasets released ov er the past 25 years, accom- panied by standardized and detailed metadata. • Integration paradigm : W e establish a structured taxonomy and present a metadata-driv en fusion paradigm (MDFP), ef fecti vely scaling-up existing medical imaging data for medical foundation model dev elopment by integrating datasets with shared characteristics. • Interactive discov ery portal : Based on the unified taxonomy and the MDFP , we build an interactiv e discovery portal that enables automated and fine-grained dataset search, inte gra- tion, and statistical analyses by modality , anatomy , task, and label type. • Gap analysis : W e identify underrepresented modalities, anatomical re gions, and tasks, highlighting critical limitations that hinder the dev elopment of future foundation models. • Accessible community resource : W e release the portal, all surveyed dataset information, related Python toolkit, and a merged large-scale dataset for public use, offering a transpar- ent and practical resource for the research community . 2 https://tchenglv520.github.io/medical- dataset- browser/ 6 The remainder of this paper is organized as demonstrated in Figure 3 . Section 2 offers a high-lev el panorama of the landscape of over 1,000 open-access medical image datasets, analyzing their dis- tribution across modalities, tasks, and anatomical regions. Section 3 zooms in on tw o-dimensional (2D) image datasets, providing a modality-specific breakdown and rev ealing e xtreme fragmentation and a long-tail distribution. Section 4 covers three-dimensional (3D) volumetric datasets, focusing on their unique clinical value and challenges of high cost and annotation complexity . Section 5 revie ws video datasets, highlighting their role in spatiotemporal analysis. T o address the perv asi ve data fragmentation, Section 6 introduces our Metadata-Dri ven Fusion P aradigm (MDFP), a system- atic workflow for integrating disparate datasets, and the corresponding interactive discovery portal for automated and ef fectiv e dataset integration. Section 7 discusses broader challenges and future directions. Finally , Section 8 concludes the surve y . Section 2 Overview of Datasets Scope 1000+ datasets Comprehensive data coverage Generation Section 3 2D Datasets Section 4 3D Datasets Section 5 V ideo Datasets T asks : • Generation • Classification • Detection • Segmentation Organs : • Retina • Breast • Brain • Colon Datasets : • EyeP ACS • MedMNIST • OCT A-500 • CheXmask Licenses Dimensions 2D 3D V ideo Detection Segmentation Registration Classification Tracking Reconstruction T asks : • Segmentation • Detection • Registration • Reconstruction Organs : • Brain • Liver • Prostat • Heart Datasets : • AutoPET • Narratives • MM-WHS • CT -RA TE T asks : • Detection • Segmentation • Classification • T racking Organs : • Colon • Esophagus • Stomach • Heart Datasets : • EndoV is • A VOS • Cataract-1K • SurgV isDom Modalities Section 6 Dataset Fusion Paradigm Section 7 Challenges & Discussion Metadata Harmonization Fusion Blueprint Semantic Alignment Dataset Indexing & Community Sharing Data Gaps & Bias Opportunities Evaluation for FMs T asks Organs Modalities : • Pathlogy • X-Ray • CT • MRI Modalities : • MRI • CT • PET • Ultrasound Modalities : • Endoscopy • Ultrasound • Microscopy • RGB Figure 3: Ov erview of the survey . W e first introduce the ov ervie w of the medical imaging datasets, followed by three sections detailing 2D, 3D, and video datasets. W e further implement inte gration strategies to merge the datasets for lar ge-scale resources, which can potentially be le veraged for the de velopment of foundation models. Finally , we discuss the challenges for foundation model dev elopment. 2 An Overview of Medical Image Datasets This section provides an overvie w of 1000+ medical image datasets released between 2000 and 2025, covering di verse anatomical structures, modalities, and tasks as illustrated in Figure 4 . These datasets are compiled from major public repositories (The Cancer Imaging Archiv e 3 , etc. ) and recent challenge sites (Grand Challenge 4 , etc. ) follo wed by deduplication, manual verification of landing pages/licences, and metadata normalization, ensuring comprehensi ve coverage. W e leave details of dataset collection process in Section 6 . T o better or ganize the landscape, we adopt the taxonomy in Figure 5 . Specifically , we begin by grouping medical imaging datasets by imaging dimensionality (2D, 3D, and video). Within each di- mensionality , we further cate gorize datasets by imaging modality . Finally , within each modality , we subcategorize datasets by task ( e .g. , segmentation, classification) and anatomical r e gion . This pro- vides a comprehensiv e basis for the analyses below , and aligns well with foundation model training needs, where dimensionality influences backbone architectural design, modality reflects acquisition physics and clinical use, task determines supervision signals and anatomical di versity shapes gener- alization in clinical practice. 3 https://www.cancerimagingarchive.net 4 https://grand- challenge.org 7 Dermoscop y MRI CT CT Pathology Endosc opy X - Ray Skin X - Ray MRI B one and S pine Chest A bdome n Other Eyes CT CT CT CT MRI MRI CT MRI Head and N eck CT Head and N eck CT Fundus photo 0 25 50 75 100 125 150 175 Number of Datasets Yearly Data Number and Dataset Count 2000 2010 2015 2020 2025 Year 10 10 10 10 10 10 10 10 Total Data Number D at a Num ber — B ar D at aset C ount D at a Num ber — Line 7 6 5 4 3 2 1 0 0 25 50 75 100 125 150 175 Number of Datasets Yearly Data Number and Dataset Count 2000 2010 2015 2020 2025 Year 10 10 10 10 10 10 10 10 Total Data Number D at a Num ber — B ar D at aset C ount D at a Num ber — Line 7 6 5 4 3 2 1 0 Ultrasound segmentation segmentation segmentation segmentation segmentation segmentation c l as sifi c ation c l as sifi c ation c l as sifi c ation de t e c ti o n de t e c ti o n de t e c ti o n segmentation segmentation det ectio n det ectio n det ectio n n ev u s mel an o ma b en i g n Dermoscop y MRI CT CT Pathology Endosc opy X - Ray Skin X - Ray MRI B one and S pine Chest A bdome n Other Eyes CT CT CT CT MRI MRI CT MRI Head and N eck CT Head and N eck CT Fundus photo 0 25 50 75 100 125 150 175 Number of Datasets Yearly Data Number and Dataset Count 2000 2010 2015 2020 2025 Year 10 10 10 10 10 10 10 10 Total Data Number D at a Num ber — B ar D at aset C ount D at a Num ber — Line 7 6 5 4 3 2 1 0 0 25 50 75 100 125 150 175 Number of Datasets Yearly Data Number and Dataset Count 2000 2010 2015 2020 2025 Year 10 10 10 10 10 10 10 10 Total Data Number D at a Num ber — B ar D at aset C ount D at a Num ber — Line 7 6 5 4 3 2 1 0 Ultrasound segmentation segmentation segmentation segmentation segmentation segmentation c l as sifi c ation c l as sifi c ation c l as sifi c ation de t e c ti o n de t e c ti o n de t e c ti o n segmentation segmentation det ectio n det ectio n det ectio n n ev u s mel an o ma b en i g n Figure 4: Overvie w of medical imaging datasets: representativ e modalities by anatomical re gion (left), dataset distrib ution across modalities, anatomical regions, and tasks (upper right), and tempo- ral trends in dataset numbers (lower right). The resulting manifest underpins all figures in this section. W e are particularly interested in the number of images in these datasets (see the right panel of Figure 4 ), as it strongly influences the effecti v eness of foundation model pre-training. Follo wing this principle, we first present the total growth ov er time and then analyze distributions by imaging dimensionality , modality , task, and organ. 2D 3D V ideo Pathlogy CT X-ray MRI Fundus ··· CT MRI Path ology Ultrasound ··· Endoscopy Microscopy Ultrasound ··· Classification Segmentation Detection Generation Brain Lung Liver Breast Retina Classification Segmentation Detection Generation T racking Brain Lung Liver Breast Retina Classification Segmentation Detection Generation Registration Brain Lung Liver Breast Retina Figure 5: T axonomy of medical imaging datasets across data dimensions, modalities, tasks, and anatomical organs. 8 (a) (b) (c) (d) (e) Figure 6: The ov ervie w of image number in medical image datasets released from 2000 to 2025. (a) T otal image number; Image number of dif ferent (b) dimensions, (c) modalities, (d) tasks, and (e) top fiv e or gans. 9 (a) (b) (c) (d) Figure 7: The distrib ution of (a) imaging dimensionalities, (b) modalities, (c) tasks, and (d) anatom- ical regions. 2.1 T otal Growth W e first examine the annual count of released imaging data to gain insight into the temporal evolution of open-access medical image datasets. Figure 6 (a) illustrates the number of imaging items publicly released per year from 2000 to 2025, with clear inflection after 2012 and another sur ge after 2023. The first phase tracks the rise of deep learning methods [ 35 ], which increased demand for extensiv e, curated training data. The recent surge be ginning in 2023 reflect the adoption of self-supervised and large-scale foundation models, which benefits from scale ev en with limited labels. These advances highlight the centrality of massiv e datasets for enabling foundation-level models, motiv ating the medical imaging community to collect substantially larger resources in pursuit of general-purpose medical AI. For example, AbdomenAtlas [ 34 ] aggregates 1.5 million 2D CT images and 5,195 3D CT v olumes. CT -RA TE [ 36 ] introduces 25,692 non-contrast 3D chest CT scans from 21,304 unique patients. These datasets rank among the largest public medical imaging resources. Nonetheless, the scale of existing medical imaging datasets, particularly in terms of 3D volumes, remains orders of magnitude smaller than data resources in natural image and language domains, where training corpora typically contain trillions of tok ens [ 37 ]. Gi ven the prohibitive cost of curating trillion- scale medical datasets, an alternati v e and more practical strategy is to integrate multiple existing datasets into lar ger , heterogeneous corpora. This observation motiv ates our dataset fusion paradigms (Section 6 ). 2.2 Imaging Dimensionalities T wo-dimensional images have height and width as their two dimensions, while three-dimensional volumes add a depth axis; videos are time-ordered 2D frames with temporal continuity . Figure 6 (b) 10 presents the total number of 2D images, 3D volumes, and videos released between 2000 and 2025, and Figure 7 (a) shows the distrib ution of them. T wo-dimensional images dominate in absolute scale, especially after 2023 (Figure 6 (b)), reflecting the wide use of 2D images for medical applications. This dominance has practical and methodologi- cal roots: 2D images are easier to store and share; patch extraction from histopathology whole-slide images (WSIs) multiplies sample counts [ 38 , 39 ]; and many long-standing benchmarks target 2D tasks [ 40 , 41 ]. In contrast, 3D and video data remain comparativ ely scarce and sho w slo wer gro wth, largely due to higher acquisition costs and storage constraints, and the complexity of curation and annotation [ 42 ]. Despite lo wer av ailability , 3D and video data are often more clinically informativ e for diagnosis and treatment planning, particularly in radiology , as they capture volumetric context and temporal dynamics that 2D cannot [ 43 ]. Increasing the av ailability of high-quality 3D and video datasets is therefore a priority for advancing clinically useful foundation models. 2.3 Imaging Modalities MRI c c c c MRI c Pathology CT X - Ray Ultrasound Fundus photo Dermoscopy Endoscopy MRI c c c c MRI c Pathology CT X - Ray Ultrasound Fundus photo Dermoscopy Endoscopy Figure 8: Representati ve modalities in medical imaging datasets. Clinical practice employs dif ferent technologies and techniques to acquire medical images; these techniques are known as imaging modalities, as depicted in Figure 8 . Each modality is designed to capture specific anatomical, functional, or molecular characteristics of the human anatomical regions/structures, and plays a critical role in clinical diagnosis and disease monitoring. Some of the most commonly used imaging modalities include: • X-ray imaging is among the oldest and most widely used techniques [ 44 ], capturing 2D projections of internal structures using X-rays. It is widely applied to detect hard tissues such as bone fractures, lung infections, or dental issues. • Computed T omography (CT) visualizes internal structure of human body with tomo- graphic acquisition of many slices to form 3D volumes. CT offers high spatial resolution and speed, making it particularly v aluable in trauma, oncology , and cardio vascular imag- ing [ 45 ]. 11 • Magnetic Resonance Imaging (MRI) generates high-contrast images of soft tissues using strong magnetic fields and radiofrequenc y pulses. It is frequently used in neuroimaging and visualizing internal organs such as the heart, liv er , and kidne ys [ 46 ], with sub-modalities including T1, T2, FLAIR, D WI, and fMRI. • Ultrasound imaging le verages high-frequency sound waves to visualize soft tissues and fluid-filled structures. It is safe, portable, and economical, and is widely used in obstetrics, cardiology , and abdominal imaging. • Positr on Emission T omography (PET) uses a radioactive tracer to detect diseased cells, featuring in functional imaging. It is commonly used for diagnosing dementia, cancers, and assessing heart conditions [ 47 ]. • Pathology imaging applies adv anced microscopy and digital slide scanning to achiev e ultra-high-resolution reconstruction and computational analysis of tissue. Beyond serving as the gold standard for histological classification, grading, and definitive cancer diagno- sis, it is increasingly central to biomarker discovery , prognostic modeling, and AI-dri ven computational pathology [ 48 ]. • Endoscopy employs a mini-camera embedded in a flexible tube, which will be inserted into the gastrointestinal tract, respiratory pathways, or other body orifices to directly vi- sualize internal organs and cavities [ 49 ]. It is widely applied in diagnostic inspection and interventional procedures. • Fundus photography captures detailed images of the retina at the back of the eye. It is essential in ophthalmology for diagnosing and monitoring diabetic retinopathy , age-related macular degeneration, glaucoma, and other retinal diseases [ 50 ]. • Dermoscopy pro vides non-in v asi ve, magnified vie ws of skin lesions, enabling observ ation of both skin surface and superficial layers. This technique rev eals fine structural details of pigmented lesions and thereby improves the diagnostic accuracy of skin lesions, particu- larly for melanoma [ 51 ]. • Other modalities include mammography for breast screening, fundus fluor escein angiog- raphy (FF A) and optical coherence tomography (OCT) for visualizing internal structures of the eyes, as well as non-imaging modalities that record electrical acti vity such as the electrocardiogram (ECG) for the heart, the electroencephalogram (EEG) for the brain, and electromy ograph y (EMG) for muscle response. Figure 6 (c) sho ws the image number of the top six modalities from 2000 to 2025. Prior to 2023, CT , pathology , and MRI account for the majority of images. Post-2023 growth is especially pro- nounced in pathology imaging, X-ray , fundus photography , and microscopy . As shown in Fig- ure 7 (b), pathology datasets contain substantially more images than other imaging modalities be- cause the gigapixel-scale WSIs are often divided into thousands of patches, each used as a separate image for analysis [ 52 ]. The inherently multi-scale nature of pathology , spanning cellular morphol- ogy to tissue-lev el architecture, further increases patch generation by requiring sampling at multi- ple magnifications. Moreover , the di v ersity of staining protocols and specimen types adds further heterogeneity and v olume [ 53 ]. These factors provide pathology with an unmatched reserv oir of fine-grained image data that underpins the training of foundation models [ 48 ]. X-ray and CT also benefit from clinical ubiquity and high throughput [ 45 ]. MRI accounts for about 10.4% of the total number of images due to its ef fecti veness in visualizing soft tissues. Despite being radiation-free, MRI imaging data grows relati vely slowly due to cost, longer acquisition, and complex multi-sequence labeling. In addition, fundus photography , microscopy , dermoscopy , and ultrasound are widely used and produce a significant number of images. Ho wev er , other modalities like PET , mammography , and endoscopy remain comparatively less av ailable in open data, which may limit the ability of foundation models trained on public corpora to fully address modality- specific clinical tasks. 2.4 T asks Medical image datasets can be collected and curated to address a wide range of tasks, each target- ing specific aspects of image analysis and interpretation in computer -aided diagnosis and clinical workflo ws. These tasks include, but are not limited to, se gmentation, classification, registration, generation, detection, and tracking. 12 Segmentation tasks in volv e assigning a class label to each individual pixel in 2D images or v ox el in 3D volumes. The goal is to delineate anatomical structures or pathological regions of interest, such as organs, tumors, and lesions, allowing for precise spatial localization and quantitati ve analysis. For example, in abdominal MRI, se gmentation can distinguish between the liv er and kidneys, facilitating downstream analysis lik e volume estimation or disease monitoring. Classification tasks aim to categorize an entire medical image or a specific region within it into predefined classes. This could in v olv e distinguishing between healthy and diseased states, grading the severity of a condition, or identifying the presence of particular disease types. For instance, in brain MRI, classification might in volv e determining whether an image corresponds to a cognitively normal subject, someone with mild cognitiv e impairment, or a patient with Alzheimer’ s disease. Registration refers to the process of aligning two or more images into a common coordinate sys- tem. This is particularly important when comparing scans from dif ferent time points (longitudinal analysis), modalities ( e.g . , MRI and PET), or subjects (for population studies). Registration tech- niques compute spatial transformations to ensure that anatomical structures in one image accurately correspond to those in another . Accurate registration is essential for tasks like image fusion, gro wth tracking, or mapping patient data to standardized anatomical atlases. Generation tasks typically use models to synthesize ne w medical images, often conditioned on specific attributes or constraints. This can help augment training datasets, simulate rare disease appearances, or recov er missing modalities. Detection focuses on ef ficient identification and localization of specific pathological findings with bounding boxes, such as lung nodules in CT , surgical instruments in endoscopic videos, or cancer cells in pathology slides. T racking monitors the movement or evolution of anatomical structures or lesions across image se- quences or time-series data, which is critical for assessing disease progression or treatment response. Reconstruction transforms incomplete or indirect ra w data obtained from sensors into a meaningful image, which often in v olves solving in v erse problems and addressing lo w-lev el vision tasks. Regression predicts continuous, quantitativ ely meaningful targets from medical images or se- quences, such as physiologic indices (ejection fraction), image-deriv ed biomarkers (arterio- lar–venular ratio), sev erity/quality scores, or v oxel-wise physical fields (dose), supporting precise monitoring, prognosis, and treatment planning. Localization seeks to identify specific anatomical points or landmarks in an image, such as corners of bones or key organ boundaries, to support diagnosis, measurement, registration, or treatment planning. Beyond these, vision-language tasks are emerging thanks to the rapid de velopment of multimodal large language models. For instance, visual question answering (VQA) aims to answer questions about giv en images in natural language, while captioning and report generation produce free-text or structured descriptions from images or image series, in different le v els of detail. In Figure 6 (d), from the task-wise statistics, classification and segmentation account for the largest share of released images o ver the past decade, while datasets tagged for generation e xhibit a marked uptick after 2023, showing the strong community interest in applying general-purpose generativ e AI for adv anced medical image analysis. In contrast, other tasks like registration, detection, and tracking, remain a relati vely small number of images over time. W e stress, ho wev er , that the ap- parent imbalance is more indicativ e of practical constraints than of community priorities, because counts are shaped by mix ed factors of label economics ( e.g. , per-image classification labels are com- parativ ely inexpensi v e) as well as acquisition and annotation burden ( e.g. , tracking requires videos with temporal labels [ 54 ]; registration often lacks easily verifiable ground truth and may depend on multi-timepoint/multimodal data [ 55 ]). Figure 7 (c) further presents the imbalanced distribution of these tasks, where generation, classifi- cation, and segmentation tasks are more extensiv ely studied compared to the remaining tasks. This imbalance may not be ideal for training a general-purpose AI e xcelling in these less-represented tasks. 13 ( a ) ( b ) ( c ) Figure 9: Representative samples of three medical image analysis tasks: (a) classification, (b) seg- mentation, and (c) detection. Each column shows images from diverse modalities and clinical ap- plications, illustrating the characteristic outputs of the respectiv e task types. V isual examples in Figure 9 intuitiv ely demonstrate the distinct outputs and clinical relev ance of each task, highlighting how the same imaging modalities can serve dif ferent analytical purposes depending on the problem at hand. 2.5 Anatomical Regions Figure 10: Anatomical structures of medical image datasets. W e also sho w the total number of datasets and images for each anatomical structure. Anatomical regions, illustrated in Figure 10 , are specific, named areas of the human anatomical re- gions used to organize and describe structures. This subsection analyzes the datasets from a compact set of anatomical regions, identified in two steps. First, we align each dataset’ s native labels with standard medical v ocab ularies. Second, we group those mapped labels into a concise set of anatom- ical regions/structures that are consistently reported across public datasets and align with major clinical workload and benchmark. This choice fa vors comparability and coverage across sources. While finer-grained systems are possible, the y are une venly annotated in the open-access datasets. Figure 6 (e) tracks medical imaging data counts for common target or gans. W e observ e that brain and lung contribute the largest volumes of images before 2023. Starting in 2023, there is an abrupt and 14 pronounced sur ge in se veral modalities, most notably brain, li ver , lung, and breast, while retina does not exhibit a comparable surge. This pattern highlights shifting research priorities toward clinically significant organs. Figure 7 (d) shows the distribution of anatomical regions. Notably , the number of fully body , retina, breast, brain, lung, and colon images significantly exceeds that of other regions, highlighting a strong research emphasis on with high clinical and societal impact, including Alzheimer’ s disease, diabetic retinopathy , and common cancers such as breast, lung, and colorectal cancer . In contrast, other anatomical re gions are less represented, such as the foot, blood, heart, bo wel, shoulder, huerus, forearm, etc. Such undercov erage often stems from practical challenges such as limited accessibility , lower disease pre v alence, or the comple xity of imaging certain anatomical sites. 2.6 Summary The public landscape of open medical imaging is distinctly long-tailed: man y small, tightly scoped datasets coexist with a smaller set of large hubs, with pronounced sk ew to ward 2D images, a subset of modalities (notably pathology , X-ray , CT , MRI), and a handful of organs (brain, lung, li ver , breast). T ask labels are like wise imbalanced where classification and segmentation dominate while other tasks are comparati vely underrepresented, lar gely reflecting practical constraints such as label economics, acquisition burden, and the scarcity of ground truth for certain tasks. These patterns imply that scale for general-purpose models is attainable, b ut not via nai ¨ ve concatenation: it requires careful normalization of counting con ventions, balanced sampling across modalities and organs, and task-aware objecti v es to av oid amplifying e xisting biases. Giv en this heterogeneity and unev en distrib ution, it is increasingly crucial to effecti v ely utilize all these datasets for training medical foundation models. Specifically , recent medical foundation mod- els across subdomains hav e been trained by integrating multiple public datasets within a modality or organ. Examples include backbone model for ophthalmology [ 56 , 57 ], histopathology [ 58 ], radio- graphy [ 59 , 60 ], segmentation-focused families in 2D [ 7 ] and 3D [ 13 , 14 ], and ev en autoencoders for generative models [ 61 , 62 ]. In short, the very imbalances mapped above become design signals for assembling data and objecti ves. W ith the proposed taxonomy and metadata-driv en fusion, this paper provides a principled path from fragmented public datasets to scalable, di verse, and clinically relev ant training distrib utions for medical foundation models. 3 2D Medical Image Datasets W e hav e collected 502 2D medical image datasets. In aggregate data count, 2D image far exceeds 3D volumes and video frames. W e partition them into 475 labeled and 45 unlabeled datasets. Labeled datasets are analyzed by modality , task, and anatomical focus; for unlabeled datasets which lack explicit task definitions, we summarize modality and anatomy . 3.1 Overview Figure 11 shows the distrib utions of dif ferent modalities, anatomical regions/structures, and tasks for 2D labeled images, which represent clear long-tail distributions. In terms of modality , pathology and X-ray dominate, followed by CT , MRI, and fundus photogra- phy . T ogether these account for the majority of images. Other modalities such as endoscopy are less representativ e. In terms of anatomy , large shares concentrate on full-body/multi-structure vie ws and a few organs with mature screening pipelines, e.g . , retina, breast, and brain, follo wed by lung and colon. By contrast, datasets targeting uterus, heart, esophagus, limb joints, and small substructures ( e.g . , nodules) remain underrepresented, indicating opportunities for tar geted curation. The dom- inating tasks include generation, classification, segmentation, regression, and detection. Howe v er , other tasks, e.g. , registration, tracking, localisation, reconstruction, and visual question answering, hav e much fe wer images. Figure 12 demonstrates representati v e e xamples of the collected 2D med- ical image datasets across different modalities and anatomical re gions. 15 (a) (b) (c) Figure 11: The distribution of different (a) modalities, (b) anatomical structures, and (c) tasks for for 2D labeled datasets. Each slice of the pie chart shows the percentage and the actual number of images. 3.2 CT Modality CT is a cornerstone of radiological imaging, providing detailed cross-sectional vie ws of the body . In 2D datasets, CT images are typically axial, sagittal, or coronal slices extracted from 3D v ol- umes. A predominant characteristic of CT datasets is their extensi ve use in se gmentation tasks. These tasks can be broadly categorized into delineating anatomical structures, such as organs for surgical planning, and identifying pathologies, such as tumors or hemorrhages for diagnosis and treatment monitoring. This makes CT datasets highly valuable for a wide range of clinical applica- tions. Among 2D labeled CT datasets, 39 provide CT slices (see T able 7 ), totaling approximately 1.4 million images. Scale v aries dramatically: from small, specialized collections such as The V isible Human Project (# 14 ) with only 2 images, to large-scale resources lik e RSN A Intracranial Hemorrhage Detection (# 6 ) with 874,000 images. CT Datasets by Anatomical Regions/Structur es. A clear trend in the distrib ution of CT datasets is the focus on specific anatomical regions. Datasets related to the brain are the most represented in terms of image v olume, primarily due to a single lar ge-scale dataset (# 6 ). Lung-related datasets are the most numerous, driv en by research in CO VID-19 and cancer screening. Con versely , data for abdominal and other structures remains relati vely scarce, highlighting potential gaps in data av ailability for de veloping models for those areas. 1) Lung (11 datasets, ∼ 60,400 images). A significant portion of the datasets is dedicated to the lungs, focusing on tasks like cancer classification in the National Lung Screening T rial (# 5 ) and CO VID-19 classification in datasets such as CO VID-19-CT SCAN IMA GES (# 8 ) and SARS-CO V -2 Ct-Scan Dataset (# 12 ). Segmentation is also a key task, as seen in CT Medical Images (# 4 ). These datasets are characterized by distinct visual patterns, such as ground-glass opacities for CO VID-19 and well- defined nodules in cancer screening, making them ideal for dev eloping specialized classifiers. 2) Brain (5 datasets, ∼ 874,400 images). Brain datasets constitute the largest collection by image count, dominated by the RSNA Intracranial Hemorrhage Detection dataset (# 6 ) for localization tasks. Other datasets lik e Brain CT Images with ICH Masks (# 17 ) focus on segmentation, while smaller sets like Cranium Image Dataset (# 11 ) are used for detection. 3) Abdomen/P elvis (7 datasets, ∼ 1,500 images). This category covers organs such as the kidney , pan- creas, colon, and prostate. Key tasks include segmentation and classification of tumors in datasets like CMB-CRC (# 18 ) for colorectal cancer and se gmentation of kidneys and pancreas in the QUBIQ challenges (# 35 ). These datasets are typically small, limiting their use for training large-scale deep learning models. They often feature multiple org ans with subtle boundaries and variable shapes, making multi-organ se gmentation a significant challenge despite limited data a vailability . 4) Full-Body/Multistructur e (5 datasets, ∼ 454,400 images). These datasets provide data from multi- ple anatomical regions or cell structures, making them suitable for pre-training generalizable models. Notable examples include RadImageNet (# 16 ), a lar ge-scale classification dataset with 34 anatomic categories, and MedMNIST (# 13 ), which contains di verse 2D slices for educational and research purposes. Their di v ersity across dif ferent anatomical regions/structures helps models learn a more generalized representation of CT imaging characteristics, reducing the risk of overfitting to a specific anatomy . 16 5) Others (11 datasets, ∼ 11,000 images). This group comprises datasets for various other anatomical regions/structures or those without a specified structure. It includes specialized collections such as 5K+ CT Images on Fractured Limbs (# 2 ) for limb fracture se gmentation and Head CT Image Data (# 25 ) for classification. Datasets with non-specific structures, like RIDER Phantom PET - CT (# 30 ) for calibration, are also in this category . CT Datasets by T asks. The distribution of datasets is heavily skewed tow ards classification, which accounts for a lar ge volume of images. Detection and localization tasks are dominated by a single lar ge dataset, while segmentation and reconstruction datasets are generally smaller in scale. 1) Classification (12 datasets, ∼ 513,900 images). Classification is the most common task, especially for pulmonary applications spurred by the CO VID-19 pandemic, with datasets like CO VID-CT (# 9 ) and SARS-CO V -2 Ct-Scan Dataset (# 12 ). Large multi-purpose datasets like RadImageNet (# 16 ) and MedMNIST (# 13 ) also contrib ute significantly to this category . Oncology is another major focus, with datasets such as the National Lung Screening T rial (# 5 ) for lung cancer . These tasks often in volv e distinguishing between dif ferent diseases or staging disease severity from a single representativ e slice. 2) Se gmentation (9 datasets, ∼ 2,100 images). Segmentation datasets are div erse but generally small. They cover or gan se gmentation, such as in the QUBIQ challenges (# 35 ), lesion se gmentation in Brain CT Images with ICH Masks (# 17 ), and quantitati v e imaging in Finding and Measuring Lungs in CT Data (# 24 ). Segmentation in CT is crucial for quantitative analysis, such as measuring tumor volume or assessing or gan health, mo ving beyond simple qualitati ve assessment. 3) Detection/Localization (2 datasets, ∼ 874,100 images). This task category is dominated by the RSN A Intracranial Hemorrhage Detection dataset (# 6 ), which contains 874,000 slices with hem- orrhage annotations. The only other dataset in this cate gory is the much smaller Cranium Image Dataset (# 11 ), also for hemorrhage detection. This task is often a precursor to se gmentation and is critical in large-scale screening programs where anomalies need to be quickly identified. 4) Reconstruction (1 dataset, 28 images). The LoDoPaB-CT dataset (# 1 ) is the sole entry dedicated to reconstruction, specifically for sparse-view reconstruction challenges. 5) Multi-task datasets (3 datasets, ∼ 500 images). A few small datasets are designed for multiple tasks. For example, CMB-CRC (# 18 ) provides data for both segmentation and classification of colorectal cancer , while CMB-PCA (# 22 ) is for classification and prediction in prostate cancer . 6) Others (12 datasets, ∼ 11,200 images). The remaining datasets are for other specific tasks or hav e no specified task. This includes AREN0534 (# 3 ) for estimation and LDCTIQA C2023 (# 26 ) for re gistration. A significant number of datasets, such as those from the TCIA archi ve like CPT A C- LSCC_CT_PET (# 23 ) and Prostate-MRI (# 32 ), hav e no explicit task listed and may be used for a variety of research purposes. 3.3 MRI Slices MRI offers superior soft-tissue contrast compared to CT and does not introduce ionizing radia- tion, making it ideal for neurological, musculoskeletal, and oncological imaging. A key feature of MRI datasets is their multi-contrast nature; a single study often includes multiple sequences ( e.g. , T1-weighted, T2-weighted, FLAIR) that highlight different tissue properties. This multi-channel information provides a rich basis for tasks like tumor segmentation and tissue characterization, though it also presents a challenge in fusing the information effecti vely . Our analysis of 24 di- verse MRI and multimodal imaging datasets (T able 8 ) reveals important trends in dataset develop- ment across modalities and clinical applications. In total, there are approximately 722,400 images , with significant variations in scale: from small, specialized collections, such as The V isible Human Project (# 42 ) with only 2 images, to large-scale resources like RadImageNet (Subset: MR) (# 56 ) with 673,000 images. MRI Datasets by Anatomical Regions/Structures. A clear trend in the distrib ution of MRI datasets is the focus on specific anatomical regions, alongside a growing number of large-scale, multi-structure collections suitable for pre-training generalizable models. Datasets related to the brain are a common focus, though typically smaller in scale. Abdominal and pelvic datasets are also 17 classification detection segmentation Landmark localizati on X- ray Pathology CT Endoscopy Mammograph X- ray Pathology X- ray CT Fundus Ultrasound Endoscopy MRI Pathology OCT X- ray Dermoscopy Pathology Fundus Ultrasound OCT CT Endoscopy Figure 12: Demonstration of the collected 2D medical datasets across dif ferent modalities and anatomical regions. present but limited in image volume. Conv ersely , data for other specific regions like the heart or spine is av ailable, highlighting di verse clinical applications. 1) Brain (2 datasets, 220 images). Datasets focused on the brain are represented by two small-scale collections for segmentation tasks: braimMRI (# 39 ) and Brain-MRI (# 40 ), each containing 110 images for analyzing brain tumors and diseases. 2) Abdomen/P elvis (4 datasets, ∼ 560 images). This category covers organs such as the colon and prostate. Ke y tasks include segmentation and classification of tumors in datasets like CMB- CRC (# 44 ) for colorectal cancer and multiple datasets for prostate cancer analysis, including CMB- PCA (# 46 ), Prostate Fused-MRI-Pathology (# 48 ), and Prostate-MRI (# 53 ). These datasets are typi- cally small, with a combined total of around 560 images. 3) Full-Body/Multistructur e (8 datasets, ∼ 704,900 images). These datasets provide data from multi- ple anatomical regions/structures, making them suitable for pre-training general izable models. This category is dominated by RadImageNet (Subset: MR) (# 56 ), a large-scale classification dataset with 673,000 images. Other notable examples include ImageCLEF 2016 (# 43 ) with 31,000 images and multi-organ challenge datasets lik e the QUBIQ series (# 57 ), (# 58 ). 4) Others (10 datasets, ∼ 16,700 images). This group comprises datasets for various other anatomical regions/structures or those without a specified structure. It includes specialized collections such as Cardiac Atrial Images (# 49 ) for heart segmentation with 8,000 images, SpinalDisease2020 (# 41 ) for spine analysis, and KNOAP2020 (# 38 ) for knee osteoarthritis. It also includes sev eral datasets from The Cancer Imaging Archive where the specific structure is not listed, such as APOLLO-5 (# 50 ) and ICDC-Glioma (GLIOMA01)_3D-MR (# 47 ). MRI Datasets by T asks. The distribution of datasets is heavily sk ewed towards classification, which accounts for the vast majority of images due to one large-scale collection. Segmentation is the next most common task, though the corresponding datasets are significantly smaller . A number of datasets are provided without a specific task, of fering resources for v arious research purposes. 1) Classification (3 datasets, 704,000 images). Classification is the most represented task by im- age volume, dominated by RadImageNet (Subset: MR) (# 56 ) (673,000 images) and ImageCLEF 18 2016 (# 43 ) (31,000 images). ImageCLEF 2015 (# 55 ) also falls into this category , although it con- tains no images. 2) Se gmentation (6 datasets, ∼ 8,900 images). Segmentation datasets are more numerous b ut contain far fewer images in total. They cov er various organs, including the heart in Cardiac Atrial Im- ages (# 49 ) (8,000 images), the brain in braimMRI (# 39 ) and Brain-MRI (# 40 ), and multiple abdom- inal organs in the QUBIQ challenges (# 57 ), (# 58 ). These datasets often require precise delineation of soft tissues with subtle intensity differences, a task for which MRI is uniquely suited. 3) Multi-task datasets (2 datasets, ∼ 500 images). A couple of small datasets are designed for mul- tiple tasks. CMB-CRC (# 44 ) provides data for both se gmentation and classification of colorectal cancer , while CMB-PCA (# 46 ) is designed for classification and prediction in prostate cancer . 4) Others (13 datasets, ∼ 9,000 images). The remaining 13 datasets cover a range of other tasks or ha ve no specified task. This includes SpinalDisease2020 (# 41 ) for detection (150 images), KNO AP2020 (# 38 ) and CMB-MML (# 45 ) for prediction, and AREN0534 (# 37 ) for estimation (239 images). A significant number of datasets (9) are provided without an explicit task, such as APOLLO-5 (# 50 ) and the ICDC-Glioma series (# 47 ), making them fle xible resources for ex- ploratory research. 3.4 PET Slices PET is a functional imaging modality that visualizes metabolic processes, often by tracking the up- take of a radioactiv e tracer . 2D PET slices are typically used in conjunction with anatomical imaging like CT or MRI for accurate localization of metabolic acti vity . Therefore, a common characteris- tic of PET datasets is their multi-modal nature (PET/CT or PET/MR). The primary tasks inv olv e detecting and quantifying regions of high metabolic activity , which are often indicati v e of cancer , inflammation, or neurological disorders. W e hav e collected 13 PET imaging datasets, a majority of which are sourced from The Cancer Imaging Archiv e (TCIA), as detailed in T able 9 . These col- lections often include multiple modalities alongside PET . Compared to CT and MRI datasets, the y span less di v erse tasks and anatomic regions, focusing primarily on brain and abdominal imaging for segmentation and classification tasks. In total, these datasets comprise approximately 41,942 images. The scale varies significantly , from small collections like CMB-GEC (# 62 ) with only 14 images to the large-scale ImageCLEF 2016 (# 60 ) dataset, which contains 31,000 images. PET Datasets by Anatomical Regions/Structures. The distribution of PET datasets shows a concentration in specific anatomical areas, with a significant number of datasets lacking explicit structural information. Datasets with multi-structure or full-body scope contribute the largest vol- ume of images, primarily due to one large collection. 1) Brain (2 datasets, ∼ 269 images). Brain-related PET datasets are represented by CMB-GEC (# 62 ) and CMB-MEL (# 63 ). These datasets focus on the detection and segmentation of cerebral microb- leeds in melanoma patients. Howe ver , their small sample sizes limit their suitability for training large-scale deep learning models. 2) Abdomen/P elvis (1 dataset, 472 images). This category contains a single dataset, CMB- CRC (# 61 ), which provides images of the colon for research on colorectal cancer . The limited size of this collection may constrain its use for dev eloping complex models. 3) Full-Body/Multistructur e (2 datasets, ∼ 31,200 images). This category is dominated by the large- scale ImageCLEF 2016 dataset (# 60 ), containing 31,000 images across skin, cell, and breast struc- tures. The other dataset, AREN0534 (# 59 ), pro vides 239 images of the kidney and lung. 4) Others (8 datasets, ∼ 10,000 images). The majority of the collected PET datasets do not specify an anatomical region. This category includes collections for various diseases, such as AREN0532 (# 69 ) for W ilms T umor research. While di verse, many of these datasets, such as CMB-MML (# 64 ) (60 im- ages), have limited numbers of images. This category also includes lar ger collections like APOLLO- 5 (# 66 ) with 6,200 images. PET Datasets by T asks. The tasks are unev enly distrib uted, with classification datasets pro viding the vast majority of images. A significant number of datasets lack explicit task labels, making them candidates for unsupervised or semi-supervised learning approaches. 19 1) Classification (1 dataset, 31,000 images). The classification task is represented by a single, lar ge- scale dataset, ImageCLEF 2016 (# 60 ), which contains 31,000 images and is designed for classifica- tion challenges. 2) Segmentation (1 dataset, 255 images). The sole dataset dedicated purely to se gmentation is CMB- MEL (# 63 ), which provides 255 images for melanoma-related cerebral microbleed se gmentation. 3) Multi-task datasets (2 datasets, 486 images). T wo small datasets are designed for multiple tasks. CMB-CRC (# 61 ) (472 images) supports both segmentation and classification for colorectal can- cer , while CMB-GEC (# 62 ) (14 images) is annotated for the same tasks in the context of cerebral microbleeds. 4) Others (9 datasets, ∼ 10,200 images). The remaining nine datasets are intended for other specific tasks or have no defined task (’N A ’). This group includes AREN0534 (# 59 ) for estimation and CMB- MML (# 64 ) for prediction. The majority , ho we ver , are general-purpose collections without specified tasks, such as APOLLO-5 (# 66 ) and AREN0532 (# 69 ), which can be v aluable for developing and testing unsupervised models or for a variety of bespok e research questions. 3.5 Ultrasound (US) Images Ultrasound imaging is a real-time, non-in vasi v e, and portable modality , making it widely used for various applications from fetal monitoring to cardiac assessment. A key characteristic of ultra- sound datasets is the inherent image noise (speckle) and operator-dependent variability , which pose significant challenges for automated analysis. Common tasks include segmentation of anatomical structures ( e.g. , cardiac chambers, fetal head) and classification of lesions ( e.g . , benign vs. malignant breast tumors). As presented in T able 10 , we have collected 19 major ultrasound imaging datasets from various sources including TCIA and Kaggle. The datasets include approximately 457,663 images in total, with RadImageNet-US (# 76 ) contributing the v ast majority (390k images). Ultrasound Datasets by Anatomical Regions/Structures. The av ailable datasets cov er a wider range of anatomical regions/structures including the skull, breast, heart, thyroid, and li v er , in addi- tion to full-body imaging, though some TCIA collections (APOLLO-5 (# 72 ) and CMB-LCA (# 608 )) lack anatomic specifications. Following the guideline that datasets containing multiple or gans are categorized separately , RadImageNet-US (# 76 ) represents the most comprehensi ve full-body cover - age with 390k images, while other datasets remain relativ ely small-scale. 1) Breast (2 dataset, ∼ 803 images). The BUSI (# 71 ) and BreastMNIST (# 77 ) datasets focus on breast ultrasound for cancer detection, providing segmented images with binary classification labels. This small-scale collection may support basic supervised learning applications. 2) Skull (1 dataset, 1,344 images). HC18 (# 70 ) targets fetal head circumference measurement through skull ultrasound imaging. As a challenge dataset with CC BY 4.0 license, it facilitates standardized benchmarking. 3) Full-Body (1 dataset, 390k images). RadImageNet-US (# 76 ) dominates the ultrasound cate gory with extensiv e coverage of 15 abdominal structures, though its commercial license may restrict accessibility . 4) Multi-structur e (2 datasets, 31,239 images). T wo datasets, including ImageCLEF 2016 (# 75 ) and AREN0534 (# 78 ), cov er multiple structures such as skin, breast, kidney , and lung. 5) Others (12 datasets, ∼ 31,200 images). The remaining datasets focus on specific organs like the heart (CAMUS (# 89 )), thyroid (TN-SCUI2020 (# 81 )), and brachial plexus (Ultrasound Nerve Seg- mentation (# 80 )), or lack detailed anatomic descriptions. These multi-modal collections currently provide 6,203 images from APOLLO-5 (# 72 ), while CMB-LCA (# 608 ) has no a v ailable images. Ultrasound Datasets by T asks. Ultrasound datasets sho w sev eral distinct task types represented, namely measurement, segmentation, and classification, along with tracking, estimation, and recon- struction. Among classification datasets, RadImageNet (US) (# 76 ) has the largest image count, while ImageCLEF 2016 (# 75 ) offers more classes (30). 1) Classification (5 dataset, ∼ 421,500 images). RadImageNet-US (# 76 ) offers large-scale multi- class classification across 15 abdominal categories. 20 2) Se gmentation (8 dataset, ∼ 26,300 images). Multiple datasets including BUSI (# 71 ), CA- MUS (# 89 ), and the Ultrasound Nerve Segmentation (# 80 ) dataset provide pixel-lev el annotations for organ and tumor segmentation, supporting computer-aided diagnosis development. The main challenge in these datasets is dealing with weak boundaries and acoustic shadowing artif acts. 3) Measur ement (1 dataset, 1,300 images). HC18 (# 70 ) specializes in biometric measurement tasks, particularly fetal head circumference calculation. 4) Unlabeled datasets (3 datasets). The TCIA datasets (APOLLO-5 (# 72 ), CMB-LCA (# 608 ), and AREN0532 (# 87 )) currently lack labels; APOLLO-5 (# 72 ) contains 6,203 images and AREN0532 (# 87 ) contains 1,021 images, while CMB-LCA (# 608 ) has none av ailable, though their multi-modal nature may enable future fusion studies. 3.6 X-Ray Images As one of the oldest and most common medical imaging techniques, 2D X-ray (radiography) pro- vides a projectional vie w of anatomical structures, excelling at visualizing bone and air -filled spaces like the lungs. X-ray datasets are characterized by their large volume, particularly for chest imaging, driv en by routine screening for diseases like pneumonia and tuberculosis. The primary tasks are classification of pathologies and segmentation or localization of abnormalities, though the ov erlap- ping of anatomical structures in the 2D projection can make these tasks challenging. T able 11 sho ws the 61 major X-ray imaging datasets from div erse sources, including TCIA, Grand Challenges, and open data platforms. These collections comprise approximately 1,657,000 images in total. The CheXmask (# 143 ) dataset dominates the quantity with 676,800 images for lung se gmentation, fol- lowed by the CheXpert (# 114 ) and VICTRE (# 139 ) datasets, while most other datasets range from hundreds to thousands of samples, presenting a long-tail distribution common in medical imaging. X-Ray Datasets by Anatomical Regions/Structures. The collected X-ray datasets cov er di verse anatomical regions, with a strong emphasis on thoracic imaging due to its clinical pre v alence in pulmonary and cardiac diagnostics. Approximately 46% of the datasets focus on the chest/lung region, reflecting the widespread use of X-rays for respiratory disease screening ( e.g. , CO VID- 19, pneumonia). Other anatomical regions/structures are less represented, with limited datasets for musculoskeletal, neurological, and abdominal applications. 1) Thorax/Lung (28 datasets, ∼ 537,900 images). This category dominates the X-ray collections, in- cluding large-scale datasets like NIH Chest X-ray 14 (# 96 ) (112,100 images) and CheXpert (# 114 ) (224,300 images). These datasets are notable for their multi-label classification tasks, where a sin- gle image can be associated with multiple pathologies. The ChestX-Det (# 121 ) series (3,600 im- ages) provides detailed annotations for lung pathologies, while MIDRC-RICORD-1c (# 128 ) (1,300 images) supports CO VID-19 research. Smaller datasets like JSR T (# 124 ) (247 images) focus on pneumonia and pulmonary nodules. Br east / Mammogr aphy (3 dataset, ∼ 248,300 images). VICTRE (# 139 ) dominates this category . VICTRE’ s (# 139 ) massiv e scale underscores breast imaging’ s importance but lacks disease annota- tions. Mammography datasets are characterized by the need to detect subtle signs of cancer , such as microcalcifications and masses, in dense breast tissue. 2) Musculoskeletal (8 datasets, ∼ 15,681 images). Musculoskeletal datasets include spine (AASCE (# 105 ), 609 images), clavicle (CRASS (# 119 ), 518 images), and pelvic bone (PENGWIN2024-T ask2 (# 148 ), 150 images) studies. The TCB-Challenge (# 118 ) (174 images) targets osteoporosis detection via bone radiographs, highlighting X-ray’ s role in orthopedic diag- nostics. A common task in these datasets is fracture detection and classification. 3) Brain/Head (2 datasets, ∼ 1,400 images). Brain datasets are limited to DENTEX (# 133 ) (1,000 images) for dental imaging and Cephalometric X-ray Image (# 126 ) (400 images) for cephalometric analysis, indicating a gap in neurological X-ray datasets compared to CT/MRI. 4) Multi-structure (5 datasets, ∼ 186,200 images). This category includes datasets spanning multi- ple distinct anatomical regions, such as MedMNIST (# 111 ) (100,000 images) and MURA (# 108 ) (40,000 images). 21 5) Others (8 datasets, ∼ 700,000 images). Includes generic collections like the CheXmask (# 143 ) (676,800 images) and X-ray Pneumonia Image Dataset (# 94 ) (5,900 images) without detailed anatomic labels. X-Ray Datasets by T asks. The datasets exhibit clear task specialization, with classification being the most pre valent application scenario. Notably , 31% of the collections (19/61) provide pixel-le vel annotations or detection labels, reflecting the clinical demand for precise localization in diagnostic imaging. 1) Classification (30 datasets, ∼ 670,100 images). This cate gory represents the largest task group, predominantly focusing on pulmonary and CO VID-19 related diagnoses. Ke y collections include CheXpert (# 114 ) (224,300 images), NIH Chest X-ray 14 (# 96 ) (112,100 images), and RANZCR CLiP (# 122 ) (30,100 images, catheter classification). The JSR T (# 124 ) dataset, though small (247 images), provides v aluable multi-class annotations for both pneumonia and pulmonary nodules. 2) Segmentation (10 datasets, ∼ 708,500 images). These datasets emphasize anatomical structure de- lineation, with CheXmask (# 143 ) (676,800 images) and Pneumothorax Masks X-Ray (# 98 ) (12,000 images) being the most substantial. The Pulmonary Chest X-Ray (# 132 ) dataset (800 images) specif- ically targets lung abnormality segmentation, while CRASS (# 119 ) (518 images) focuses on clavicle identification for orthopedic applications. 3) Detection/Localization (9 datasets, ∼ 59,700 images). Emerging needs for surgical planning are addressed by DENTEX (# 133 ) (1,005 brain images) and CL-Detection2023 (# 134 ) (555 images). The CEPHA29 (# 135 ) dataset (1,000 images) stands out for cephalometric landmark localization, despite its current data accessibility issues. 4) Others (5 datasets, ∼ 36,100 images). Unique applications include AASCE’ s (# 105 ) spinal curvature regression (609 images), CoronARe’ s (# 138 ) vascular reconstruction, and RSN A Bone Age’ s (# 146 ) bone age estimation (14,200 images). These demonstrate X-ray’ s versatility beyond con ventional diagnostic roles. 3.7 Optical Coherence T omography (OCT) Images OCT provides micrometer-resolution, cross-sectional images of biological tissues in real-time. It is analogous to "optical ultrasound," using light instead of sound. Its primary application is in ophthalmology for imaging the layers of the retina. Consequently , OCT datasets are highly spe- cialized, focusing on tasks like retinal layer segmentation for thickness mapping and classification of retinal diseases based on layer morphology . T able 12 provides 22 major optical coherence to- mography (OCT) imaging datasets from diverse sources, including Kaggle, Grand Challenges, and academic institutions. These collections demonstrate remarkable specialization in retinal imaging, comprising approximately 221k images in total. T wo large public classification benchmarks — OCT2017 (# 150 ) (about 83.5k images) and MedMNIST (# 158 ) (100k images) — account for the majority of images in the corpus. In contrast, most other datasets range from hundreds to thousands of samples, presenting a typical long-tail distribution in medical imaging resources. OCT Datasets by Anatomical Regions/Structures. Notably , almost all of the datasets focus ex- clusiv ely on retinal applications, reflecting OCT’ s primary clinical use in ophthalmology . The only exception is MedMNIST (# 158 ), which can also be applied to breast and lung. As such, we do not break down to introduce the anatomical re gions/structures. OCT Datasets by T asks. The datasets exhibit clear task specialization, with classification and segmentation being the most prev alent application scenarios. The classification task has the largest number of images, though the number of datasets for classification is less than that of the seg- mentation task. Segmentation datasets account for approximately 50% of the datasets, pro viding pixel-le vel annotations for precise anatomical analysis. 1) Classification (5 datasets, ∼ 210,200 images). This category represents the lar gest task group in terms of image number , predominantly focusing on diabetic retinopathy and glaucoma detec- tion. Ke y collections include OCT2017 (# 150 ) (83,484 images), Retinal OCT -C8 (# 151 ) (24,000 images), and MedMNIST (# 158 ) (100,000 images combining multiple modalities). The core task in these datasets is to distinguish diseases based on morphological changes in retinal layers, such as 22 the presence of drusen or intraretinal fluid. The iChallenge-A GE19 (# 152 ) dataset (1,600 images) specifically targets glaucoma classification with detailed angle closure annotations. 2) Se gmentation (11 datasets, ∼ 2,600 images). These datasets emphasize retinal layer delineation, with SinaF arsiu-009 (# 163 ) (840 images) and SinaFarsiu-018 (# 167 ) (784 images) providing the most substantial annotations. The DRA C22 (# 153 ) dataset (174 images) specializes in diabetic retinopathy lesion segmentation, while iChallenge-GO ALS (# 154 ) (300 images) offers three-layer retinal segmentation crucial for thickness measurements. 3) Prediction (3 datasets, ∼ 8,500 images). The APTOS series (APTOS-2021 (# 156 ), APTOS Cross- Country Stage 1 (# 157 ), and APT OS Cross-Country Stage 2 (# 168 )) total 8,500 images for diabetic retinopathy severity prediction, using the International Clinical Diabetic Retinopathy scale. These datasets demonstrate OCT’ s growing role in quantitativ e disease progression monitoring. 3.8 Fundus Images Fundus photography captures high-resolution color images of the retina, making it a cornerstone of ophthalmology . A key characteristic of fundus datasets is their similarity to natural RGB images in terms of data format, which allows for the direct application and transfer learning of models de- veloped for general computer vision. Howe v er , the content is highly specialized, featuring unique anatomical landmarks like the optic disc, fovea, and a complex network of blood vessels. Common tasks revolv e around detecting and grading pathologies such as diabetic retinopathy and glaucoma. The challenge lies in identifying these subtle, often minute, pathological features within a com- plex anatomical background. T able 13 sho ws 75 major fundus photography datasets from div erse sources, including Grand Challenges, Kaggle, and academic institutions. These collections demon- strate remarkable specialization in retinal imaging, comprising approximately 412,400 images in total. The AIR OGS (# 182 ) dataset dominates the quantity with 101,400 images, while most other datasets range from hundreds to thousands of samples, presenting a typical long-tail distribution in medical imaging resources. Notably , almost all of the datasets focus exclusi vely on retinal applica- tions, reflecting fundus photography’ s primary clinical use in ophthalmology diagnostics. Fundus Photography Datasets by Anatomical Regions/Structur es. The collected datasets ex- clusiv ely focus on retinal imaging, reflecting fundus photography’ s specialized application in oph- thalmology . All 75 datasets target the retina, with varying emphasis on specific anatomical structures or pathological features. This extreme specialization contrasts with other modalities like CT or MRI that cov er multiple body regions. Fundus Photography Datasets by T asks. The datasets exhibit clear task specialization, with clas- sification being the most prev alent application scenario. Approximately 30% of the collections provide pixel-le vel annotations or detection labels, enabling precise anatomical analysis crucial for diagnostic applications. 1) Classification (42 datasets, ∼ 304,200 images). This cate gory represents the largest task group, predominantly focusing on diabetic retinopathy and glaucoma detection. Ke y collections include OIA-ODIR (# 214 ) (10,000 images), APTOS 2019 (# 194 ) (5,590 images for diabetic retinopathy grading), and Y angxi (# 200 ) (20,394 images for eye axis classification). These datasets are piv otal for dev eloping automated screening systems for prev alent eye diseases, often framed as multi-class grading problems based on the number and type of lesions present. The JSIEC (# 207 ) dataset (1,000 images) stands out for its comprehensi ve coverage of 38 fundus disease categories, though sample sizes per category remain limited. 2) Se gmentation (21 datasets, ∼ 5,300 images). These datasets emphasize retinal structure delin- eation, with RIM-ONE (# 193 ) (485 images) and GAMMA CFP (# 240 ) (200 images) providing optic disc/cup annotations crucial for glaucoma assessment. The HRF Se g (# 211 ) dataset (45 im- ages) offers high-resolution vessel segmentation, while A O-SLO (# 197 ) (840 images) specializes in photoreceptor mapping. The iChallenge-GAMMA series ((# 241 ), (# 218 )) demonstrates growing interest in multi-modal retinal analysis. Segmentation tasks are critical for quantitati v e analysis, focusing on delineating blood vessels to assess vascular health, the optic disc and cup to measure glaucomatous changes, and lesions like exudates or hemorrhages to quantify disease se v erity . 23 3) Re gr ession (6 datasets, ∼ 2,300 images). The INSPIRE series ((# 208 ), (# 209 )) (70 images com- bined) focuses on arteriov enous ratio measurement, while DeepDR-T ask2 (# 219 ) (2,000 images) addresses disease progression prediction. These datasets highlight fundus photograph y’ s e xpanding role in quantitativ e disease monitoring. 3.9 Dermoscopy Images Dermoscopy inv olves imaging the skin with a specialized magnifying lens to visualize subsurface structures not visible to the naked eye. These datasets are crucial for the early detection of skin cancer , particularly melanoma. The images are typically high-resolution RGB photos of skin le- sions. Ke y tasks include the segmentation of lesion boundaries and the classification of lesions into categories ( e.g. , benign ne vus, melanoma, basal cell carcinoma). There are 17 major dermoscop y imaging datasets in our collection, as shown in T able 14 . They are collected from v arious sources, in- cluding ISIC challenges, CVPR competitions, and independent research collections. These datasets predominantly focus on skin imaging. They primarily address segmentation and classification tasks, with a strong emphasis on skin lesion analysis. These datasets include approximately 167,300 images in total, with Monk eypox (# 255 ) ha ving the largest single collection (40,200 images) and ISIC20 (# 243 ) (33,100 images), ISIC19 (# 247 ) (25,300 images), and Fitzpatrick17k (# 248 ) (16,600 images) also providing substantial sample sizes for training medical imaging models. Dermoscopy Datasets by Anatomical Regions/Structures. The vast majority of these datasets focus on skin imaging, though a fe w cover other anatomical regions. 1) Skin (13 datasets, ∼ 133,600 images). This dominant cate gory includes all ISIC challenge datasets (ISIC16-20 (# ?? )), Fitz- patrick17k (# 248 ), MED-NODE (# 249 ), PH2 (# 251 ), and others. The largest collections are Mon- keypox (# 255 ) (40,200 images), ISIC20 (# 243 ) (33,100 images), and ISIC19 (# 247 ) (25,300 im- ages). These datasets demonstrate strong clinical focus on melanoma detection and skin lesion analysis. 2) F oot (1 dataset, 2,000 images). DFUC2020 (# 252 ) specifically targets foot imaging for diabetic foot ulcer analysis. 3) Thyr oid (1 dataset, 637 images). DDTI focuses on thyroid nodule segmentation. 4) Multi-structure (1 datasets, ∼ 31,000 images). ImageCLEF2016 (# 254 ) co vers skin, cell, and breast imaging with 31,000 images. Dermoscopy Datasets by T asks. The collected datasets sho w clear task specialization, with most providing high-quality labels suitable for supervised learning. 1) Se gmentation (5 datasets, ∼ 9,400 images). K ey collections include ISIC16 (# 244 ) (1,279 images), ISIC17 (# 245 ) (2,750 images), ISIC18 (# 242 ) (2,694 images), and DDTI (637 images). These typically focus on precise lesion boundary delineation. 2) Classification (10 datasets, ∼ 157,500 images). Major collections include Monkeypox (# 255 ) (40,200 images), ISIC20 (# 243 ) (33,100 images), ImageCLEF 2016 (# 254 ) (31,000 images), and ISIC19 (# 247 ) (25,300 images). These datasets often provide multi-class categorization of skin lesions. Unlabeled dataset (1 dataset, 368 images). V itiligo (# 256 ) is the only unlabeled collection, potentially useful for unsupervised learning. 3.10 Histopathology Histopathology is the microscopic examination of tissues to study the manifestations of disease. Digital pathology datasets, particularly those based on Whole Slide Images (WSIs), possess unique characteristics. WSIs are gigapixel-resolution images, often exceeding 100,000 × 100,000 pix els, which makes it computationally infeasible to process them directly . Consequently , a standard pre- processing pipeline inv olves patch e xtraction or tiling, where the WSI is di vided into thousands of smaller , manageable patches. Common tasks include patch-level classification ( e.g. , identifying tu- morous vs. normal tissue), object-level segmentation or detection ( e.g. , delineating nuclei, glands, or mitotic figures), and WSI-le vel classification for diagnosis. The challenges in this modality stem from the massiv e image size, significant variations in staining and preparation, and the need to ag- gregate patch-level predictions into a coherent slide-level diagnosis. T ables 15 and 16 present 117 major histopathology imaging datasets from div erse sources, including grand challenges (MICCAI, ISBI), open data platforms (TCGA, TCIA, OpenDataLab), and research collections. These datasets predominantly utilize hematoxylin and eosin (H&E) staining, with some incorporating immunohis- tochemistry (IHC). They collecti vely contain approximately 2.22 million images (comprising ∼ 2.15 million patch images and ∼ 67,000 WSI), with the Quilt-1M (# 371 ) (1,000,000 images) and Patch- Camelyon (PCam) (# 345 ) (328,000 images) being the largest collections. Notably , 82% of datasets 24 provide high-quality labels suitable for supervised learning. The prohibiti ve cost of large-scale WSI annotation catalyzed a shift towards SSL, enabling the rise of Pathology Foundation Models from vast unlabeled data archives. Initial de velopment centered on algorithmic inno v ations using public datasets lik e TCGA. A subsequent "scale rev olution" utilized massive, priv ate "real-w orld" datasets, powering models like UNI (trained on over 100,000 WSIs) and Prov-Gig aPath (trained on ov er 171,000 WSIs). This addressed the "domain shift" limitations of public data, proving that dataset scale is now a primary engine of progress in the field. Histopathology Datasets by Anatomical Regions/Structures. The datasets show a strong clini- cal focus on cancer diagnosis across multiple anatomical sites. 1) Br east (25 datasets, ∼ 53,000 im- ages). Major collections include BRIGHT (# 270 ) (5,086 images), BRCA-M2C (# 304 ) (120 images), and the BreakHis series (# 299 , # 327 , # 327 , # 329 ) (combined 35,236 images across magnifications). These primarily address tumor classification and se gmentation. 2) Pr ostate (9 datasets, ∼ 42,000 images). P AND A (# 336 ) (10,616 images) and SICAPv2 (# 300 ) (18,783 images) are the largest, focusing on Gleason grading. 3) Colon/Rectum (12 datasets, ∼ 113,000 images). CRC100K (# 303 ) (100,000 images) and CoNIC2022 (# 271 ) (4,981 images) provide e xtensi ve data for colorectal can- cer analysis. 4) Multi-or gan (17 datasets, ∼ 1.18 million images). Quilt-1M (# 371 ) (1,000,000 im- ages) and MedMNIST (# 276 ) (100,000 images) cov er multiple cancer types. 5) Other s include lung (7 datasets, ∼ 38,000 images), lymph nodes (9 datasets, ∼ 537,000 images), and blood (5 datasets, ∼ 53,000 images). Histopathology Datasets by T asks. The datasets demonstrate specialized task distributions. Emerging trends include increased WSI adoption (32% of recent datasets) and multi-task collec- tions combining segmentation with classification or counting. 1) Classification (38 datasets, ∼ 709,000 images). Ke y datasets include LC25000 (# 297 ) (25,000 images, lung/colon classification) and Histopathologic Cancer Detection (# 277 ) (220,000 images). The BreakHis series (# 299 , # 327 , # 328 , # 329 ) provides multi-magnification classification (40 × - 400 × ). A ke y challenge is handling intra-class v ariation and inter-class similarity at the cellular lev el, making fine-grained classification difficult. 2) Se gmentation (31 datasets, ∼ 368,000 images). Notable collections are GlaS (# 294 ) (165 images, colorectal glands) and CRA G (# 306 ) (213 im- ages, extended from GlaS). Se gmentation targets range from macro-structures lik e tumor regions to micro-structures like individual nuclei or glands, which are essential for quantitati v e pathology . 3) Detection (6 datasets, ∼ 14,000 images). MIDOG2021 (# 281 ) (200 images) focuses on mitotic figure detection. 4) Multi-task (4 datasets, ∼ 14,000 images). PanNuke combines se gmentation and clas- sification (PanNuke (Seg) (# 298 ), 7,901 images), while CoNIC2022 (# 271 ) adds counting tasks. 5) Specialized tasks include registration (ANHIR (# 266 ), 481 images), generation (BCI (# 285 ), 4,900 images) and VQA (Quilt-1M (# 371 ), 1,000,000 images). 3.11 Microscopy Imaging T able 17 summarizes 34 major microscopy imaging datasets. These datasets predominantly utilize brightfield and fluorescence microscopy , with a strong focus on cellular and subcellular imaging. They collectiv ely contain approximately 1.8 million images , with the CellTracking2019 (# 372 ) dataset (1.44 million images), DLBCL-Morph (# 391 ) (152,200 images), and Kaggle-HP A (# 385 ) (89,460 images) being the largest collections. Unlike histopathology which focuses on tissue archi- tecture, these microscopy datasets often center on the morphology , count, and beha vior of indi vidual cells or microorganisms. Notably , most datasets provide high-quality labels suitable for supervised learning, cov ering a wide range of biological scales from single molecules to whole organisms. Microscopy Datasets by Anatomical Regions/Structur es. The datasets demonstrate specialized focus on specific anatomical structures: 1) Cellular (8 datasets, ∼ 1.51M images). Ke y collections include CellTracking2019 (# 372 ) (16,042 sequences, 1.44M frames), Kaggle-HP A (# 385 ) (89,460 images), and OCCISC ((# 381 ), (# 403 )) (945 images). These primarily address cell segmentation and tracking. 2) Ocular (5 datasets, ∼ 153,000 images). The corneal series (CornealNerv e (# 388 ), NerveT ortuosity (# 389 ), CornealEndothelial (# 387 )) and DLBCL-Morph (# 391 ) (152,200 images) focus on eye microstructure analysis. 3) Breast (1 dataset, 400 images). ICIAR2018 (# 382 ) pro- vides histopathology images for breast cancer classification. 4) Blood (3 datasets, ∼ 28,500 im- ages). Blood Cell Images (# 375 ) (12,500 images) and Leukemia Classification (# 376 ) (15,100 25 images) analyze blood cell morphology . 5) Multi-structur e (2 datasets, ∼ 31,500 images). Image- CLEF2016 (# 399 ) (31,000 images) cov ers multiple tissue types. Microscopy Datasets by T asks. There is an increased use of deep learning benchmarks (Kaggle- HP A (# 385 )) and integration of multiple tasks (CBC series (# ?? ) combining counting and detection). The datasets show clear specialization in analysis tasks: 1) Segmentation (11 datasets, ∼ 99,000 images). Kaggle-HP A (# 385 ) (89,500 images), CREMI (# 373 ), and OCCISC-Seg (# 381 ) (945 im- ages) provide precise cellular boundary delineation. A common challenge is accurately separating densely clustered or overlapping cells. 2) Classification (12 datasets, ∼ 81,400 images). ImageCLEF 2016 (# 399 ) (31,000 images), B-ALL Classification (# 378 ) (15,100 images), and ICIAR2018 (# 382 ) (400 images) enable morphological categorization. 3) Detection (3 datasets, ∼ 2,600 images). BloodCell (# 395 ) (874 images) and T uberculosis (# 396 ) (1,265 images) localize specific cellular features. 4) T rac king (1 datasets, ∼ 1.4M images). CellTracking2019 (# 372 ) dominates this cate gory with 1.4 million time-lapse frames. 5) Specialized tasks include regression (DLBCL-Morph (# 391 ), 152.2k images; CBC-Count (# 383 ), 420 images) and protein localization (Kaggle-HP A (# 385 )). 3.12 Infrared Imaging Infrared imaging in medicine captures thermal patterns or reflectance properties not visible in the normal spectrum. In the context of the collected datasets, it is primarily used in ophthalmology to image retinal structures with dif ferent light w av elengths. This modality is non-inv asi ve and can pro- vide unique contrast for features like the retinal pigment epithelium. The tasks often re v olve around image quality assessment or classification based on specific features visible in the infrared spectrum. T able 18 includes 6 major infrared reflectance imaging datasets. These collections focus exclusi vely on ocular imaging, particularly retinal analysis, using infrared reflectance technology . The datasets contain approximately 424,532 images in total, with the MRL Eye series ((# ?? )) (combined 424,490 images across 5 sub-datasets) representing the largest collection. All datasets provide high-quality labels suitable for supervised learning, with a strong emphasis on classification tasks (5/6 datasets). Infrared Datasets by Anatomical Regions/Structur es. Infrared imaging remains highly special- ized, with 100% of datasets focusing on retinal applications, and all created since 2018, suggesting growing interest in this modality . Specifically , Retina (six datasets, ∼ 424,532 images). The MRL Eye series ((# ?? )) (84,898 images per sub-dataset) provides comprehensiv e coverage of v arious retinal features. This extreme specialization in retinal imaging contrasts with other modalities that typically cov er multiple anatomical regions. Infrared Datasets by T asks. The datasets sho w clear task specialization: 1) Classification (5 datasets, ∼ 424,490 images). The MRL Eye series addresses multiple classification tasks: glasses detection (MRL-Eye-Glasses (# 407 )), eye state (MRL-Eye-State (# 408 )), reflection analysis (MRL- Eye-Reflections (# 409 )), image quality assessment (MRL-Eye-Quality (# 410 )), and sensor type identification (MRL-Eye-Sensor (# 411 )). 2) Se gmentation (one dataset, 42 images). RA VIR (# 406 ) is the only segmentation dataset, focusing on retinal blood vessel delineation with three classes (background, arteries, veins). 3.13 Endoscopy Imaging Endoscopy provides direct real-time video visualization of internal or gans and cavities through a flexible tube with a camera. Datasets are often composed of individual frames extracted from these videos. A key characteristic is the high variability in appearance due to camera motion, lighting changes, specularity , and physiological artifacts ( e.g . , bubbles, debris). Common tasks include polyp detection and segmentation for cancer screening, tool tracking for surgical navigation, and classification of tissue abnormalities. W e provide an overvie w of endoscopy imaging datasets in T able 19 , where 41 major ones are collected from div erse sources, e.g. , ISBI and MICCAI. These datasets predominantly feature endoscopic imaging (39/41), with a few incorporating multi-modal data (2/41). They cover div erse anatomical regions and tasks, totaling approximately 322,200 im- ages and videos , with EndoSlam (# 413 ) being the largest collection (76,837 images). Notably , 39% of datasets (16/41) contain over 1,000 images, making them potentially suitable for training medical vision models. 26 Endoscopy Datasets by Anatomical Regions/Structures The datasets cover several major anatomical regions, with strong emphasis on gastrointestinal tract e xamination: 1) Colon/Bowel (8 datasets, ∼ 109,400 images): This represents the most extensi vely examined region, featuring large-scale datasets like SUN_SEG (# 428 ) (49,136 images), SAR AS-ESAD (# 425 ) (33,398 images), and Ka vsir (# 412 ) (14,000 images) for polyp segmentation and detection. The CVC series (CVC-ClinicDB (# 422 ), CVC-ColonDB) provide high-quality annotations for polyps, while EndoCV2020 (# 416 ) and EndoV is15 (# 417 ) focus on artifact detection. 2) Esophagus (1 datasets, 157 images): Focused on Barrett’ s esophagus detection, with AIDA- E_2 (# 420 ) (157 images) providing a specialized benchmark. 3) Multi-structur e gastr ointestinal tr act (6 datasets, ∼ 86,000 images): Comprehensiv e collections like EndoSlam (# 413 ) (76k images) cover the entire gastrointestinal tract including esophagus, stom- ach, and colon. These are particularly valuable for de veloping generalizable endoscopic AI systems. 5) Other Regions : Includes specialized collections for uterus (FetReg (# 424 ), 2.7k images), gallblad- der (m2cai16-tool (# 418 ), 15 videos), and prostate (SARAS-MESAD (# 414 ), 50k images). While clinically important, these generally hav e smaller sample sizes. Endoscopy Datasets by T asks The datasets demonstrate a progression from single-task to multi- task benchmarks: Se gmentation (17 datasets, ∼ 20,000 images): Forms a large task category , with Kvasir-SEG (# 423 ) (8,000 images), FetReg (# 424 ) (2,718 images), and EndoV is 2018 - RSS (# 436 ) (2,840 images) providing high-quality se gmentation masks. Most focus on polyp segmentation, while specialized targets include surgical tools (EndoV is 2018-RSS (# 436 )) and placental vasculature (FetReg (# 424 )). Detection (6 datasets, ∼ 86,600 images): SARAS-MESAD (# 414 ) (50,284 images) and SARAS- ESAD (# 425 ) (33,398 images) are notable for bounding box annotations of abnormalities and in- struments. The m2cai series (# 418 ) pro vide instrument detection benchmarks. Classification (10 datasets, ∼ 77,700 images): Ranges from binary classification (MedFM2023) to fine-grained categorization (ImageCLEF (# 426 )). AIDA series (E1-E3) (# ?? ) provide histology classification benchmarks. Multi-task datasets (5 datasets, 156k images): HyperKvasir (# 429 ) (captioning, classification, local- ization), SUN_SEG (# 428 ) (segmentation, detection, classification), and Endo-FM (# 445 ) combine multiple annotation types, reflecting recent trends tow ards comprehensi ve benchmarks. Others : Includes reconstruction and depth estimation (EndoSlam (# 413 )) and registration (P2ILF (# 441 )). Some of these tasks, like in the EndoSlam (# 413 ) dataset (76,837 images), are supported by a large number of samples. 3.14 Other Modalities Finally , we introduce all the 2D datasets of other modalities that are not listed in the pre vious sub- sections. This section consolidates datasets from a variety of imaging modalities that, while less numerous than the major cate gories, represent important and often specialized clinical applications. T able 20 summarize the information of these modalities, spanning di verse modalities, including Mammography (4 datasets), X-Ray (3), Fundus (2), Colposcop y (2), and others. These datasets col- lectiv ely contain approximately 858,000 images , with the Digital Mammography (# 451 ) dataset be- ing the largest (640,000 images), followed by MRL Eye Gender (# 462 ) (84,898 images) and ADDI ALZHEIMER’S DETECTION CHALLENGE (# 450 ) (34,614 images). The datasets demonstrate a strong emphasis on classification tasks (75%) and co ver all major anatomical re gions, though with unev en distrib ution across modalities. Datasets by Anatomical Regions/Structures. The datasets cover comprehensiv e anatomical structures with a particular concentration on thoracic and retinal imaging. 1) Thoracic/Lung (2 datasets, ∼ 27,000 images). This category includes collections like V inDr-CXR (# 467 ) (18,000 images) and V inDr-PCXR (# 466 ) (9,125 images) for lung abnormalities. 2) Retina (3 datasets, ∼ 88,000 images). Retinal imaging features collections like MRL Eye Gender (# 462 ) (84,898 im- ages) and specialized datasets for various ophthalmic diseases. 3) Br east (4 datasets, ∼ 663,000 27 images). The Digital Mammography (# 451 ) dataset dominates this category with 640,000 images, supplemented by specialized collections like CMMD (# 457 ) (1,775) and V inDr -Mammo (# 464 ) (19,992). 4) Brain/Head (2 datasets, ∼ 5,000 images). While smaller in quantity , these include important collections like Br35H (# 458 ) (3,060) for brain tumors. 5) Whole-body/Multi-structur e collections like OralCancer (# 460 ) (131 images) provide cross-anatomical co verage. Datasets by T asks. The datasets demonstrate clear task specialization across modalities. 1) Clas- sification (15 datasets, ∼ 798,000 images): Mammography datasets like The Digital Mammography DREAM Challenge (# 451 ) and retinal collections (MRL Eye Gender (# 462 )) dominate this cate- gory . 2) Se gmentation (4 datasets, ∼ 2,300 images): Notable collections include CDD-CESM (# 459 ) (2,006 images). 3) Multi-task datasets like CDD-CESM (# 459 ) (segmentation+classification) pro- vide versatile training opportunities. 4) Emer ging tasks like reconstruction (BigNeuron (# 452 )) demonstrate expanding research frontiers. 3.15 Challenge and Opportunity The landscape of 2D medical imaging datasets presents a distinct duality . On one hand, its sheer volume, particularly in modalities like histopathology and radiography , offers a scale for model pre- training that is unparalleled in the medical domain. On the other hand, this abundance is coupled with significant fragmentation, heterogeneity , and the inherent limitations of two-dimensional rep- resentations, posing unique challenges for the de velopment of rob ust and generalizable foundation models. Key Challenges in 2D Medical Imaging Datasets. The primary obstacles stem from the div ersity and nature of 2D data acquisition and annotation practices. Extr eme fragmentation and heter og eneity represent a major barrier . The vast number of 2D datasets are scattered across numerous indepen- dent repositories and challenges, often with inconsistent imaging protocols, varying resolutions, and non-standardized metadata. This leads to significant domain shifts between datasets of the same modality , complicating large-scale integration efforts. For instance, histopathology slides exhibit wide v ariations in staining and preparation, while chest X-rays dif fer in projection and e xposure settings. P ervasive data imbalance and long-tail distributions introduce substantial biases. As our analy- sis reveals, modalities like pathology , X-ray , and fundus photography dominate the data landscape, while clinically vital modalities such as endoscopy and ultrasound remain underrepresented. This imbalance extends to anatomical re gions and tasks; for example, over 80% of images in our col- lection come from just thoracic and breast datasets, lea ving other regions ( e.g . , abdominal organs) critically underserved. This also creates modality-specific limitations; for instance, X-Ray datasets in this collection average only ∼ 12.5K images per dataset. Foundation models pre-trained on such ske wed data may fail to generalize to less common modalities or pathologies, limiting their clinical utility . Furthermore, annotation quality and scalability present a persistent challenge. The creation of lar ge- scale 2D datasets often relies on weak supervision, such as labels extracted from radiology reports, which can be noisy and imprecise. While pix el-le vel annotations are the gold standard, they are labor-intensi v e and scarce at scale. The lack of a unified annotation ontology across datasets makes it difficult to harmonize labels for multi-dataset training, hindering the creation of truly comprehensive benchmarks. Finally , the inher ent limitation of 2D r epr esentation is a fundamental constraint. A single 2D im- age, whether a projection like an X-ray or a slice from a volume, provides only a partial view of the underlying three-dimensional anatomy . This loss of spatial conte xt can be a critical handicap for diagnosing complex diseases that require volumetric understanding, such as assessing tumor morphology or subtle structural changes. Opportunities for Advancement. Despite these challenges, the 2D medical imaging domain of- fers exceptional opportunities to advance foundation models. The unprecedented scale for self- supervised pr e-training is the most significant advantage. W ith millions of av ailable images, thoracic imaging (pathology and chest radiography) has achiev ed a critical mass for large-scale AI training. This scale, alongside exceptionally standardized lar ge collections (such as the >80K retinal image 28 (a) (b) (c) Figure 13: The distribution of different (a) modalities, (b) anatomical structures, and (c) tasks for 3D datasets. Each slice of the pie chart sho ws the percentage and the actual number of images. datasets), enables the effecti ve application of self-supervised learning paradigms, such as masked auto-encoding and contrasti v e learning, to build foundational backbones that can be fine-tuned for a multitude of downstream tasks. The rich diversity of modalities enables powerful multi-modal learning . The breadth of 2D imaging, spanning from macroscopic radiographic images to microscopic pathology slides, provides a fertile ground for dev eloping models that can reason across dif ferent biological scales and data sources. A particularly promising a venue is the integration of imaging data with unstructured clinical te xt. Large datasets paired with radiology reports, such as MIMIC-CXR [ 63 ] and CheXpert [ 64 ], unlock the potential for vision-language pre-training, allowing models to learn semantically rich represen- tations that align visual features with clinical narrativ es. Moreov er , the widespread clinical use and lower cost of 2D imaging modalities create opportunities for high-impact, scalable clinical applications . Foundation models trained on common 2D data like X-rays, fundus, or dermoscopy images can be deployed for large-scale screening programs in resource-constrained settings. This can democratize access to expert-lev el diagnostics for conditions like tuberculosis, diabetic retinopathy , and skin cancer , addressing critical global health challenges. In summary , while the path to building generalist 2D medical foundation models is fraught with challenges of data heterogeneity and annotation quality , the opportunities are immense. Strategic dataset consolidation, prioritization of balanced anatomical coverage, and the dev elopment of stan- dardized multi-task annotations, coupled with adv anced self-supervised and multi-modal learning techniques, can harness the v ast scale of 2D data to create transformati ve AI tools for global health- care. 4 3D Medical Image Datasets W e hav e collected 591 3D medical image datasets, comprising 1,242,022+ volumes in total. Al- though the total number of volumes is considerably smaller than that of 2D datasets, 3D datasets provide richer spatial information that is essential for volumetric analysis and clinical decision- making. W e categorize these 3D datasets according to their modalities, tasks, and body parts. The labeled datasets dominate the collection, while unlabeled datasets provide additional opportunities for self-supervised learning approaches. 4.1 Overview W e first provide an overvie w of 3D medical image datasets. Figure 13 shows the distrib utions of different modalities, anatomical structures, and tasks for 3D datasets, which represent clear long-tail distributions. In terms of modality , MRI and CT are the most popular, while other modalities, like PET , ultrasound, and OCT , are less representati ve. From the perspectiv e of anatomical structures, the brain, abdomen, and lung have the largest number of datasets, while the prostate, teeth, and other structures are still limited in their dataset numbers. The dominating tasks include classification, segmentation, and other tasks. Howe ver , other tasks, e .g. , registration, localization, and detection, hav e much fewer datasets. Figure 14 demonstrates representativ e examples of the collected 3D medical image datasets across different modalities and anatomical re gions. 29 Figure 14: 3D visualization examples of medical imaging datasets across dif ferent modalities and anatomical structures. 4.2 CT V olumes CT is a widely used imaging modality that employs X-rays to visualize internal structures in three dimensions. W e identify 252 3D CT datasets comprising approximately 516,087 v olumes in total, as summarized in T able 21 . These datasets diver ge considerably in scale and annotation quality , from small, domain-specific collections (for example, 3D-IRCADb [ 65 ] with 20 li ver volumes) to large, multi-center compilations such as CT -RA TE [ 36 ] (50,188 v olumes). Large collections like CT -RA TE and M3D [ 66 ] aim to cover a wide range of acquisition protocols b ut often depend on semi-automated or weak supervision for annotations, while curated challenge datasets lik e T otalSeg- mentator [ 67 ] (1,204 volumes) deli ver expert-verified labels across 104 anatomical structures. Anno- tation consistency remains a persistent challenge: manual lesion delineation is laborious, operator - dependent, and subject to inter-observ er v ariability , as illustrated by se gmentation benchmarks such as LiTS [ 68 ]. Re garding clinical representati veness, CT datasets range from broad population-based cohorts like NLST [ 69 ] to small, specific single-center collections (e.g. 3D-IRCADb), whereas multi-institution benchmarks like AMOS [ 70 ] (500 CT + 100 MRI scans, collected across multiple centers and vendors) better reflect real-world di v ersity in scanner types and imaging protocols [ 70 ]. CT Datasets by anatomical structures. CT datasets sho w strong concentration in lung/chest ap- plications, dri ven by lar ge-scale screening programs and CO VID-19 research. Whole-body datasets represent an emerging trend for foundation model de velopment, while traditional abdominal and bone imaging remain important clinical applications. 1) Lung/Chest (96 datasets, 279,285 volumes). This dominant category reflects CT’ s primary clinical role in thoracic imaging. Major applications include CO VID-19 analysis (STOIC2021 (# 519 ) with 10,735 volumes, CO V19-CT -DB (# 521 ) with 7,750 v olumes), lung cancer screening (NLST (# 641 ) with 26,254 volumes), chest abnormalities detection (CT -RA TE (# 480 ) with 50,188 volumes), and nodule detection (LUNA16 (# 507 ) with 888 volumes, LIDC-IDRI (# 638 ) with 1,018 volumes). The category benefits from e xtensiv e public health initiati ves and automated screening demands. 30 2) Whole-body (7 datasets, 123,557 volumes). An emerging category driven by foundation model dev elopment needs. Ke y datasets include M3D (# 481 ) (120,000 volumes), T otalSegmentator (# 472 ) (1,204 volumes), and AutoPET series (# 473 ) (2,233 v olumes combined). These comprehensi ve collections enable multi-organ se gmentation and cross-anatomical learning. 3) Abdomen (55 datasets, 46,305 volumes). T raditional CT application focusing on multi-organ segmentation and tumor analysis. Notable collections include AbdomenAtlas (# 479 ) (20,460 v ol- umes), FLARE series (# 484 ) (7,311 volumes combined), AbdomenCT -1K (# 498 ) (1,062 volumes), and specialized or gan datasets like KiTS series (# 494 ) for kidney analysis (1,329 volumes com- bined). These datasets support both or gan-specific and comprehensi ve abdominal analysis. 4) Bone/Spine (10 datasets, 41,641 v olumes). Specialized orthopedic applications including CT - Pelvic1K (# 548 ) (1,184 volumes), CTSpine1K (# 547 ) (1,005 volumes), V erSe series (# 545 ) (460 volumes combined), and RibFrac2020 (# 549 ) (660 volumes). T asks focus on bone segmentation, fracture detection, and spinal analysis. 5) Head and Neck (21 datasets, 8,969 v olumes). Applications in radiation therapy planning and head/neck cancer treatment. Key datasets include HECKTOR series (# 542 ) (1,462 volumes com- bined), SegRap2023 (# 529 ) (400 v olumes), and v arious structural segmentation challenges. 6) Br ain (19 datasets, 4,887 v olumes). CT brain imaging focuses on emer gency applications includ- ing strok e detection (ISLES 2024 (# 527 ) with 250 v olumes), hemorrhage analysis (InST ANCE2022 (# 526 ) with 200 v olumes), and trauma assessment. Most brain imaging utilizes MRI, with CT serv- ing specialized acute care roles. CT Datasets by T asks. CT datasets demonstrate strong task di versity , with segmentation dom- inating due to CT’ s excellent structural contrast. Classification applications lev erage large-scale screening datasets, while specialized tasks like reconstruction and registration support adv anced imaging workflo ws. 1) Se gmentation (150 datasets, 266,862 volumes). Segmentation represents the dominant task cate- gory , reflecting CT’ s strength in structural imaging. Applications include multi-organ segmentation (T otalSegmentator (# 472 ) with 1,204 v olumes, AbdomenAtlas (# 479 ) with 20,460 volumes, M3D (# 481 ) with 120,000 volumes), organ-specific se gmentation (KiTS series (# 494 ) with 1,329 v ol- umes for kidneys, LiTS (# 490 ) with 201 volumes for liv er), and specialized targets like airway segmentation (AIIB23 (# 509 ) with 312 volumes) and fracture detection (RibFrac2020 (# 549 ) with 660 volumes). 2) Classification (93 datasets, 206,483 volumes). Classification tasks focus on disease screening and diagnostic applications. Major datasets include chest abnormalities detection (CT -RA TE (# 480 ) with 50,188 volumes), CO VID-19 severity assessment (ST OIC2021 (# 519 ) with 10,735 volumes), lung cancer screening (NLST (# 641 ) with 26,254 volumes), and v arious cancer staging applications across TCGA collections. These datasets enable automated diagnosis and population-level screen- ing. 3) Reconstruction (5 datasets, 130,668 volumes). Emer ging task category driv en by dose reduction and image enhancement needs. Key datasets include M3D (# 481 ) (120,000 v olumes) for multi- modal reconstruction, LDCT -and-Projection-data (# 637 ) (299 volumes) for lo w-dose reconstruction, and specialized synthesis applications. 4) Localization (6 datasets, 124,107 volumes). Localization tasks primarily support workflo w au- tomation and anatomical reference. The M3D dataset (# 481 ) (120,000 volumes) provides compre- hensiv e localization annotations across multiple or gans and structures. 5) Re gistr ation (20 datasets, 123,382 volumes). Registration applications focus on longitudinal analysis and multi-modal fusion. K ey datasets include Learn2Reg series (# 535 ) for lung CT (450 volumes) and abdomen CT -CT/MR-CT registration (# 537 ) (172 volumes), supporting motion cor - rection and atlas construction. 6) Detection (20 datasets, 52,542 volumes). Detection tasks tar get specific anatomical structures and pathological findings. Notable applications include pulmonary nodule detection (LUN A16 (# 507 ) with 888 v olumes, LIDC-IDRI (# 638 ) with 1,018 v olumes), pulmonary embolism detection (RSNA STR (# 657 ) with 12,195 volumes), and lesion detection across v arious or gans. 31 4.3 MRI V olumes Magnetic Resonance Imaging (MRI) provides rich soft-tissue contrast and div erse sequence types for v olumetric analysis. W e identified 231 3D MRI datasets comprising approximately 523,847 vol- umes in total, as summarized in T able 22 . These datasets span a wide range of sequences (T1, T2, FLAIR, D WI) and specialized protocols, varying greatly in scale and focus, from small studies such as MRBrainS13 (# 899 ) for brain tissue segmentation[ 71 ] to large-scale resources like OpenMind (# 963 )[ 72 ]. Dataset quality is shaped by sequence heterogeneity and scanner variability: standard- ized collections such as BraTS [ 73 ] enforce uniform preprocessing across four canonical sequences, whereas multi-site datasets lik e O ASIS-3 (# 756 )[ 74 ] include heterogeneous acquisition protocols and magnetic field strengths. Annotation consistency remains challenging; peritumoral or edema- related boundaries are kno wn to be more ambiguous than enhancing or core regions in brain tu- mor tasks, contrib uting to inter-observ er v ariability . In terms of clinical representativeness, MRI datasets range from healthy young adult cohorts in HCP (# 839 )[ 75 ] to elderly dementia popula- tions in ADNI (# 816 )[ 76 ], while multi-v endor datasets such as M&Ms (# 735 ) (Siemens, Philips, GE, Canon) capture broader scanner and protocol di versity and highlight persistent cross-vendor generalization gaps. MRI Datasets by anatomical structures. MRI is predominantly used in neuroimaging, with brain datasets dominating the 3D MRI landscape. Cardiac and abdominal applications show more limited representation, though they pro vide valuable specialized resources. 1) Brain/Neur o (155 datasets, 356,751 volumes). The brain represents the most studied anatomy in 3D MRI, featuring major collections including BraTS series for tumor segmentation (BraTS 2023 (# 802 ) with 5,880 volumes, UPENN-GBM (# 942 ) with 3,680 volumes), Alzheimer’ s research datasets (O ASIS-3 (# 756 ) with 5,699 v olumes, ADNI (# 816 ) with 2,500 volumes, T ADPOLE (# 779 ) with 1,667 volumes), stroke studies (ISLES 2022 (# 788 ) with 400 volumes), and multiple sclerosis research (MSSEG-2 (# 780 ) with 100 volumes). Brain datasets dominate the 3D MRI landscape in both dataset count and total volumes. 2) Head and Neck (3 datasets, 114,643 v olumes). Dominated by the OpenMind collection (# 963 ) (114,570 volumes), which represents a breakthrough in large-scale MR data collection. Other datasets include specialized head and neck cancer applications (AAPM-R T -MA C (# 897 ) with 55 volumes). 3) Pr ostate (15 datasets, 3,704 volumes). Prostate MRI represents a well-established clinical appli- cation, with notable collections including PI-CAI (# 776 ) (1,500 volumes), Prostate-MR-US-Biopsy (# 772 ) (1,151 volumes for fusion imaging), PROST A TEx (# 764 ) (204 volumes for classification), Prostate-MR-Segmentation (# 773 ) (116 v olumes), and PROMISE12 (# 761 ) (50 v olumes for seg- mentation). These datasets support cancer diagnosis, treatment planning, and MR-US fusion work- flows. 4) Breast (11 datasets, 3,262 v olumes). Breast MRI applications include Duke-Breast-Cancer - MR (# 834 ) (922 volumes), I-SPY1 (# 813 ) (847 volumes), I-SPY2 (# 944 ) (719 v olumes), A CRIN- Contralateral-Breast-MR (# 814 ) (984 v olumes), and specialized collections. These datasets support cancer diagnosis, treatment response assessment, and radiomics research. 5) Car diac (13 datasets, 2,991 volumes). Cardiac MRI datasets focus on ventricular/myocardial segmentation and functional quantification. K ey collections include M&Ms (# 735 ) (375 volumes), M&Ms-2 (# 736 ) (360 volumes), LAScarQS++ 2024 (# 738 ) (200+ volumes), MyoPS++ 2024 (# 740 ) (200+ v olumes), A CDC (# 734 ) (150 volumes), and EMIDEC (# 881 ) (150 v olumes). These datasets support automated cardiac analysis and multi-center validation studies. 6) Knee (2 datasets, 1,823 v olumes). Include MRNet (# 880 ) (1,370 v olumes for knee abnormali- ties detection) and SKI10 (# 778 ) (150 v olumes for cartilage se gmentation), supporting orthopedic applications and sports medicine research. 7) Others (17 datasets, 1,539 volumes). Include li ver applications (LLD-MMR2023 (# 935 ) with 498 volumes), spine imaging, gastrointestinal tract studies, and v arious specialized anatomical re gions. 8) Whole-body (2 datasets, 1,016 v olumes). Include T otalSegmentator MRI (# 733 ) (616 volumes) and UW -Madison GI Tract (# 883 ) (467 v olumes), providing comprehensi ve anatomical coverage for foundation model dev elopment. 32 MRI Datasets by T asks. 3D MRI datasets are predominantly designed for segmentation and clas- sification tasks, reflecting MRI’ s strength in soft-tissue contrast and anatomical delineation. The task distribution aligns with MRI’ s clinical applications in detailed tissue analysis and disease char - acterization. 1) Classification (80 datasets, 322,508 volumes). Classification represents the largest category by to- tal v olumes, dominated by the OpenMind collection (# 963 ) (114,570 v olumes) and lar ge-scale neu- roimaging studies. Major applications include Alzheimer’ s disease classification (O ASIS-3 (# 756 ) with 5,699 volumes, ADNI (# 816 ) with 2,500 volumes, T ADPOLE (# 779 ) with 1,667 volumes), population studies (Human Connectome Project (# 839 ) with 1,206 volumes, Brain Genomics Su- perstruct Project (# 838 ) with 1,570 volumes), and cancer staging (PROST A TEx (# 764 ) with 204 volumes for prostate cancer). These datasets enable automated diagnosis, disease staging, and population-lev el brain research. 2) Segmentation (114 datasets, 151,433 volumes). Segmentation represents the largest category by dataset count, lev eraging MRI’ s excellent soft-tissue contrast. Major applications include brain tumor delineation (BraTS 2023 (# 802 ) with 5,880 v olumes, UPENN-GBM (# 942 ) with 3,680 vol- umes, MSD01_BrainT umor (# 803 ) with 750 volumes), cardiac segmentation (M&Ms (# 735 ) with 375 volumes, ACDC (# 734 ) with 150 volumes), prostate segmentation (PI-CAI (# 776 ) with 1,500 volumes, Prostate-MR-US-Biopsy (# 772 ) with 1,151 volumes), and whole-body segmentation (T o- talSegmentator MRI (# 733 ) with 616 v olumes). The div ersity in anatomical targets reflects MRI’ s versatility in tissue delineation. 3) Reconstruction (15 datasets, 127,464 volumes). MR reconstruction focuses on acceleration tech- niques and image enhancement. Key datasets include fastMR (# 809 ) (1,594 volumes), CMRxRecon (# 930 ) (300 volumes for cardiac reconstruction), and OpenMind (# 963 ) which also supports re- construction tasks. This category addresses critical clinical needs for faster MR acquisition and improv ed image quality . 4) Re gistration (31 datasets, 17,808 volumes). Registration applications include multi-timepoint studies, atlas construction, and multi-modal fusion. Notable datasets include Learn2Re g series (# 757 ) (O ASIS, Hippocampus, LUMIR), CuRIOUS series (# 743 ) for MR-US registration, and vari- ous longitudinal studies for disease progression monitoring. These datasets enable temporal analysis and cross-modal alignment. 5) T rac king (5 datasets, 1,855 v olumes). Motion tracking applications primarily in cardiac MRI, including ST A COM 2011 (# 907 ) (1,158 v olumes) for cardiac motion analysis and v arious dif fusion tractography studies. These datasets support dynamic analysis and fiber tracking applications. 6) Detection (4 datasets, 1,245 v olumes). Detection tasks focus on automated identification of anatomical landmarks and pathological structures, including aneurysm detection (ADAM2020 (# 956 ) with 255 volumes) and v arious brain pathology identification tasks. 4.4 Ultrasound V olumes 3D ultrasound provides real-time volumetric imaging widely used for interv entional guidance and multi-modal fusion. W e identify 27 ultrasound-related 3D datasets containing approximately 56,609 volumes, as summarized in T able 23 . Most of these datasets appear within multi-modal collections (e.g., US/MR or US/CT), reflecting ultrasound’ s predominant role in image-guided and fusion-based clinical workflows rather than as a standalone modality . Data quality is strongly operator-dependent, with clinical acquisitions sho wing higher variability compared to controlled research settings (e.g., the CuRIOUS series (# 979 )[ 77 ]). In terms of representativ eness, e xisting 3D ultrasound datasets are primarily deriv ed from high-end interventional systems, underrepresenting handheld or point- of-care imaging scenarios common in real-world clinical practice. Ultrasound Datasets by anatomical structures. The av ailable 3D ultrasound datasets span di- verse anatomical regions, with multi-modal combinations being particularly common for re gistra- tion and fusion applications. 1) Brain (9 datasets, ∼ 500 volumes). Brain ultrasound datasets focus primarily on US-MR registra- tion for neurosurgical guidance. The CuRIOUS series (# 979 ) (2018, 2019, 2022) provides datasets for brain tumor applications, while Learn2Re g LUMIR (# 987 ) (269 v olumes) supports multi-modal 33 registration research. These datasets enable US-guided brain interventions and intraoperative navi- gation. 2) Car diac (3 datasets, ∼ 1,400 volumes). Cardiac ultrasound datasets include ST ACOM 2011 (# 907 ) (1,158 volumes for motion tracking), CETUS2014 (# 965 ) (45 volumes), and MVSeg- 3DTEE2023 (# 966 ) (175 volumes for mitral valve segmentation). These datasets support automated echocardiography , cardiac function quantification, and structural heart analysis. 3) Pr ostate (2 datasets, ∼ 1,300 v olumes). Prostate datasets focus on US-MR fusion for biopsy guid- ance and treatment planning. Prostate-MR-US-Biopsy (# 988 ) (1,151 volumes) and µ -Re gPro2023 (# 989 ) (108 volumes) support fusion imaging applications critical for prostate cancer diagnosis and intervention. 4) Kidney (4 datasets, ∼ 1,400 volumes). Pediatric kidney datasets from the AREN series (# 557 ) (AREN0532, AREN0533, AREN0534) provide multi-modal collections including ultrasound for W ilms tumor research, supporting both classification and segmentation tasks in pediatric oncology . 5) Br east (1 dataset, 200 volumes). TDSC-ABUS2023 (# 964 ) provides automated breast ultrasound data for breast cancer detection, supporting segmentation, classification, and detection tasks in breast imaging screening workflo ws. 6) Other Abdominal Or gans (8 datasets, ∼ 1,100 v olumes). Include pancreas (CPT A C-PD A (# 598 )), liv er (AHEP0731 (# 625 )), uterus (CPT A C-UCEC (# 600 )), and other organs from multi-modal can- cer imaging collections, primarily supporting classification tasks for oncological applications. Ultrasound Datasets by T asks. 3D ultrasound datasets are dominated by registration applications, reflecting the modality’ s role in multi-modal image fusion and guidance systems. 1) Registr ation (15 datasets, ∼ 2,000 volumes). Registration represents the dominant task category , reflecting ultrasound’ s critical role in real-time guidance and multi-modal fusion. Major applications include US-MR brain registration (CuRIOUS series (# 979 )), prostate fusion imaging (Prostate-MR- US-Biopsy (# 988 ), µ -RegPro2023 (# 989 )), cardiac motion tracking (ST A COM 2011 (# 907 )), and multi-modal brain re gistration (Learn2Reg LUMIR (# 987 )). This dominance reflects ultrasound’ s primary clinical v alue in providing real-time guidance for interv entions and fusion with other imag- ing modalities. 2) Classification (10 datasets, ∼ 1,800 volumes). Classification tasks focus primarily on cancer stag- ing and diagnosis across multiple organs, including kidney tumors (AREN series (# 557 )), pancreatic cancer (CPT AC-PD A (# 598 )), liv er cancer (AHEP0731 (# 625 )), and other malignancies. These ap- plications lev erage ultrasound’ s accessibility for screening and staging workflows. 3) Se gmentation (8 datasets, ∼ 800 volumes). Se gmentation applications target organ and struc- ture delineation for cardiac analysis (CETUS2014 (# 965 ), MVSeg-3DTEE2023 (# 966 )), tumor seg- mentation (AREN0533-Tumor -Annotations (# 559 ), AREN0534 (# 561 )), and breast lesion detection (TDSC-ABUS2023 (# 964 )). These datasets support automated measurement and volumetric analy- sis critical for clinical assessment. 4) Detection (1 dataset, 200 volumes). Detection tasks focus on automated lesion identification, ex emplified by TDSC-AB US2023 (# 964 ) for breast cancer screening, supporting computer -aided detection workflo ws in clinical practice. 4.5 PET V olumes Positron Emission T omography (PET) pro vides functional information complementary to anatom- ical imaging. Public PET volumes are scarce and often appear in multi-modality collections ( e .g. , PET/CT , PET/MR). W e identify 65 PET -related 3D datasets with 95,456 v olumes in total, as pre- sented in T able 24 . These collections span di verse anatomic regions with a strong focus on oncol- ogy applications, particularly in lung/chest (15 datasets), head and neck (11 datasets), and brain (8 datasets) regions. This significant expansion largely comes from comprehensive cancer imaging archiv es, multi-center studies, and lar ge-scale neuroimaging initiativ es. PET Datasets by anatomical structur es. These datasets use PET primarily for oncology applica- tions across various anatomical regions, though multi-modal combinations are the norm rather than 34 the exception. The distrib ution sho ws clear preferences for certain anatomical regions where PET imaging provides the most clinical v alue. 1) Lung/Chest (15 datasets, ∼ 55,000+ v olumes). This represents the largest category by dataset count, reflecting PET’ s critical role in lung cancer diagnosis and staging. K ey collections include QID W (# 646 ) (52,000 volumes for quality assurance), Lung-PET -CT -Dx (# 639 ) (355 volumes), CPT AC-LU AD (# 597 ) (244 volumes), ACRIN-NSCLC-FDG-PET (# 1001 ) (242 v olumes), CPT A C- LSCC (# 596 ) (212 volumes), and NSCLC-Radiogenomics (# 603 ) (211 v olumes). The dominance of lung-related datasets demonstrates PET’ s established clinical utility in pulmonary oncology . 2) Head and Neck (11 datasets, ∼ 4,200 volumes). Head and neck cancers represent a major ap- plication area for PET imaging, with notable collections including HECKTOR 2022 (# 544 ) (883 volumes), HNSCC (# 565 ) (627 v olumes), TCGA-HNSC (# 576 ) (479 v olumes), HECKTOR 2021 (# 543 ) (325 volumes), Head-Neck-PET -CT (# 634 ) (298 volumes), QIN-HEADNECK (# 615 ) (279 volumes), and A CRIN-HNSCC-FDG-PET -CT (# 998 ) (260 volumes). These datasets support both tumor segmentation and treatment response assessment. 3) Brain (8 datasets, ∼ 13,300 volumes). Brain PET datasets focus primarily on neurodegenerativ e diseases and pro vide the lar gest indi vidual dataset v olumes. Major collections include OASIS-3 (# 756 ) (5,699 volumes), ADNI (# 816 ) (2,500 volumes), T ADPOLE (# 779 ) (1,667 v olumes), and PPMI (# 1037 ) (683 volumes) for Alzheimer’ s and Parkinson’ s disease research, alongside smaller oncology-focused datasets like A CRIN-FMISO-Brain (# 1000 ) (45 v olumes). 4) Abdominal Organs (7 datasets, ∼ 1,400 v olumes). Include specialized datasets for li v er , pancreas, and kidney imaging. Notable collections include AREN0532 (# 557 ) (544 volumes) and AREN0534 (# 561 ) (239 volumes) for pediatric kidney tumors, AHEP0731 (# 625 ) (190 volumes) for liver cancer , and CPT AC-PD A (# 598 ) (168 volumes) for pancreatic cancer . 5) Whole-body/Multi-or gan (3 datasets, ∼ 2,300 volumes). Comprehensiv e whole-body PET datasets include AutoPET II (# 993 ) (1,219 volumes), AutoPET (# 992 ) (1,014 volumes), and fastPET -LD (# 1042 ) (68 volumes), providing valuable resources for pan-cancer detection and seg- mentation tasks. 6) Br east (3 datasets, ∼ 240 volumes). Specialized breast cancer datasets include BREAST - DIA GNOSIS (# 628 ) (88 volumes), A CRIN-FL T -Breast (# 999 ) (83 volumes), and QIN-Breast (# 614 ) (68 volumes), supporting breast cancer diagnosis and treatment monitoring. PET Datasets by T asks. PET datasets reflect the modality’ s primary clinical applications in on- cology and neurology , with task distribution strongly aligned with PET’ s role in functional and metabolic imaging for disease diagnosis, staging, and treatment monitoring. 1) Classification (45 datasets, ∼ 60,000+ volumes). Classification represents the dominant task cat- egory , reflecting PET’ s core clinical utility in disease staging, treatment response assessment, and diagnostic classification. Oncology applications span multiple cancer types, including lung cancer datasets (CPT A C-LU AD (# 597 ) with 244 volumes, ACRIN-NSCLC-FDG-PET (# 1001 ) with 242 volumes, TCGA-LUSC (# 1012 ) with 37 v olumes), head and neck cancer studies (TCGA-HNSC (# 576 ) with 479 v olumes, A CRIN-HNSCC-FDG-PET -CT (# 998 ) with 260 volumes), and v arious other malignancies across different anatomical sites. Neurological applications focus on neurode- generativ e diseases, particularly Alzheimer’ s disease classification (OASIS-3 (# 756 ) with 5,699 vol- umes, ADNI (# 816 ) with 2,500 v olumes, T ADPOLE (# 779 ) with 1,667 volumes) and Parkinson’ s disease research (PPMI (# 1037 ) with 683 v olumes). The dominance of classification tasks aligns with PET’ s clinical role in providing metabolic information for staging and prognosis. 2) Se gmentation (20 datasets, ∼ 25,000 volumes). Segmentation tasks focus primarily on tumor delineation and organ-at-risk identification for radiation therapy planning. Major collections include AutoPET II (# 993 ) (1,219 volumes), AutoPET (# 992 ) (1,014 volumes), HECKT OR 2022 (# 544 ) (883 volumes), and HNSCC (# 565 ) (627 volumes). These datasets support automated tumor v olume definition, which is critical for radiotherapy planning and treatment monitoring. The emphasis on head and neck, lung, and whole-body segmentation reflects PET’ s established role in oncology workflo w integration. 3) Multi-task datasets (8 datasets, ∼ 5,000 volumes). Several datasets provide annotations for mul- tiple tasks, enabling comprehensiv e analysis approaches. Examples include Head-Neck-PET -CT 35 (# 634 ) (298 v olumes for both se gmentation and classification), NSCLC-Radiogenomics (# 603 ) (211 volumes for segmentation and classification), and A CRIN-FMISO-Brain (# 1000 ) (45 volumes for segmentation and classification). This multi-task approach reflects the clinical reality where PET images are used for multiple diagnostic and therapeutic purposes simultaneously . 4) Detection (3 datasets, ∼ 400 volumes). Detection tasks focus on lesion identification and local- ization, ex emplified by Lung-PET -CT -Dx (# 639 ) (355 volumes for classification and detection) and fastPET -LD (# 1042 ) (68 volumes for detection). While less common than classification, detection tasks are important for automated screening and lesion characterization in clinical workflo ws. 5) Registr ation (3 datasets, ∼ 1,200 volumes). Registration applications appear primarily in the HECKTOR series (# 543 ) (2021 and 2022), supporting multi-timepoint analysis for treatment re- sponse assessment. This reflects PET’ s gro wing role in longitudinal monitoring of therapy ef fects and disease progression. 4.6 Other 3D V olumes Beyond the major modalities, we collect 26 3D datasets from specialized imaging techniques with 5,381+ volumes in total, as presented in T able 25 . These modalities serv e specific clinical niches and emer ging applications, with OCT dominating the collection due to large-scale ophthalmology datasets. The di versity reflects the ev olution of medical imaging technology and specialized clinical needs. Other Modalities According to Imaging T echnology . Each modality addresses specific clini- cal applications and anatomical targets, with OCT leading in volume due to comprehensi ve retinal imaging datasets. 1) Optical Coher ence T omography (OCT) (14 datasets, 4,288+ volumes). OCT dominates this cat- egory , primarily targeting retinal and ophthalmologic applications. The OLIVES dataset (# 1071 ) alone contributes 1,268 volumes for diabetic condition analysis, while the newly added OCT A-500 dataset (# 1080 ) provides 500 volumes for comprehensi v e retinal OCT A analysis. Specialized col- lections include GAMMA (# 1072 ) (300 volumes for glaucoma analysis), RETOUCH (# 1068 ) (112 volumes for retinal disease segmentation), and various Duke Univ ersity datasets for age-related macular degeneration and diabetic macular edema. The OCT A2024 dataset (# 1081 ) supports ad- vanced OCT to OCT A translation research. T asks primarily focus on classification, se gmentation, and reconstruction of retinal pathologies, supporting automated screening for eye diseases. 2) Digital Subtraction Angiogr aphy (3D DSA) (4 datasets, 454 volumes). DSA applications fo- cus on cerebrov ascular imaging, particularly aneurysm detection and analysis. Ke y datasets in- clude CADA series (# 1059 ) for cerebral aneurysm detection (372 volumes combined) and SHINY - ICAR US (# 1060 ) for internal carotid artery aneurysm se gmentation (82 volumes). These datasets support critical neurov ascular intervention planning and risk assessment. 3) Cone-beam CT (CBCT) (4 datasets, 581 volumes). CBCT serves specialized applications in dental imaging and treatment planning. Notable collections include T oothFairy2023 (# 1067 ) for dental surgery planning (443 v olumes), pancreatic CT -CBCT registration datasets (40 volumes), and pelvic reference data for prostate cancer treatment (58 volumes). These datasets bridge diagnostic and interventional imaging workflo ws. 4) 3D Micr oscopy (3 datasets, 54 volumes). Microscopy datasets target cellular and subcellular anal- ysis, including MitoEM (# 1056 ) for mitochondrial ultrastructure (2 volumes), platelet ultrastructure analysis (2 volumes), and prostate cancer pathology (50 v olumes). Though small in v olume, these datasets enable high-resolution structural analysis at the cellular lev el. Other Modalities According to T asks. T ask distrib ution reflects the specialized nature of these modalities, with classification dominating due to large-scale OCT screening applications. 1) Classification (11 datasets, 3,914 volumes). Classification tasks predominantly target disease screening and diagnosis, especially in ophthalmology . Major applications include diabetic condition screening (OLIVES (# 1071 ) with 1,268 volumes), glaucoma detection (OCT Glaucoma Detection (# 1070 ) with 1,110 volumes), and various retinal disease classification tasks. These datasets enable automated screening systems for population health initiativ es. 36 2) Se gmentation (17 datasets, 1,987 volumes). Segmentation applications span multiple modalities and anatomical targets, from retinal layer segmentation in OCT to aneurysm delineation in DSA and dental structure se gmentation in CBCT . The div ersity of targets reflects the specialized nature of each modality’ s clinical applications. 3) Registr ation (3 datasets, 498 volumes). Registration tasks primarily support treatment planning and longitudinal analysis, including CBCT -CT registration for radiation therapy and structural- functional alignment in ophthalmology . 4) Reconstruction/T r anslation (1 dataset, TBD volumes). Advanced reconstruction and translation tasks include OCT to OCT A image translation, enabling cross-modal analysis and synthetic data generation for retinal imaging applications. These specialized modalities complement major imaging modalities by addressing specific clini- cal needs and emerging applications, contrib uting to the comprehensiv e landscape of 3D medical imaging datasets. 4.7 Challenges and Opportunities The 3D medical imaging landscape presents unique challenges and opportunities that distinguish it from 2D medical imaging. Despite providing richer spatial information essential for volumetric analysis and clinical decision-making, 3D datasets remain significantly constrained by fundamental limitations in data acquisition, annotation complexity , and resource allocation. Key Challenges in 3D Medical Imaging Datasets. The primary challenges stem from the inher- ent complexity and cost of 3D data acquisition and processing. High acquisition and annotation costs represent the most significant barrier , as 3D imaging requires expensi ve specialized equipment (CT , MRI, and PET scanners) and e xpert radiologists for v olumetric annotation, resulting in the modest growth observed compared to 2D datasets. This economic constraint directly impacts data av ailability and di versity . Dataset overlap and duplication presents another critical challenge that researchers must be aware of when conducting external v alidation studies. Some datasets in our collection contain ov erlapping or identical data under different names, particularly when larger datasets consolidate multiple smaller collections. F or instance, the OCT2017 dataset and MedMNIST OCT dataset contain identical reti- nal OCT images, as MedMNIST integrates multiple publicly av ailable datasets including OCT2017. Similar ov erlaps exist across other modalities where comprehensi v e datasets mer ge smaller special- ized collections. Researchers should exercise caution when selecting datasets for external validation to avoid inadvertently using overlapping data that could lead to overly optimistic performance esti- mates and compromised generalizability assessments. Complexity and cost. On the data side, challenges are multifaceted. Acquisition costs remain pro- hibitiv ely high due to the expense of imaging hardware, long scanning times, and patient compliance issues. Storage costs escalate rapidly as each volumetric scan can range from hundreds of megabytes to sev eral gigabytes, requiring rob ust archiving infrastructure. Annotation costs are substantial be- cause volumetric segmentation demands time-consuming, slice-by-slice delineation by expert radi- ologists. On the model side, these data characteristics translate into significant computational chal- lenges. The high dimensionality of 3D medical images substantially increases memory consumption and processing time during training and inference, often necessitating specialized hardware and op- timization strategies. Moreover , the lo w signal-to-noise ratio of many volumetric acquisitions and the small size of pathological re gions further complicate feature e xtraction and model generaliza- tion. T ogether , these f actors underscore the intricate interplay between data and model complexity in 3D medical imaging research. Modality and anatomical imbalances create substantial gaps in representation. While CT (261 datasets, 753,421 volumes) and MRI (231 datasets, 523,847 volumes) dominate the landscape, crit- ical modalities like ultrasound (27 datasets, 56,609 v olumes), PET (65 datasets, 95,456 volumes), and emerging v olumetric techniques remain underrepresented relativ e to their clinical importance. Anatomically , while the concentration on brain and abdomen/liv er regions has expanded signifi- cantly , cardiac, musculoskeletal, and certain specialized applications still hav e relativ ely limited resources, though recent large-scale initiati v es are beginning to address these gaps. 37 T ask-specific limitations further constrain the utility of existing 3D datasets. The ov erwhelming dominance of classification and segmentation tasks, while clinically important, reflects the field’ s incomplete transition from task-oriented to foundation-oriented data engineering paradigms. Reg- istration and reconstruction tasks remain underrepresented despite their critical importance for lon- gitudinal studies and treatment monitoring. Additionally , the scarcity of multi-task datasets limits the dev elopment of versatile clinical AI systems capable of handling comple x, real-world diagnostic workflo ws. Opportunities for Advancement. Despite these challenges, the 3D medical imaging domain presents remarkable opportunities for transformativ e adv ancement. The substantial collection of unlabeled 3D volumes (219 datasets with hundreds of thousands of volumes) of fers unprecedented potential for self-supervised learning and contrastiv e pretraining. Large repositories like TCIA for CT and HCP for MRI provide the scale necessary for foundation model pretraining, while multi- sequence MR data enables sophisticated cross-modal consistency training and modality dropout techniques. F oundation model-driven data augmentation emerges as a particularly promising direction. W ell- trained generative foundation models can participate in semi-supervised learning frameworks, gen- erating synthetic 3D v olumes that reflect real clinical presentations while addressing priv acy con- straints. This approach is especially valuable for rare diseases and underrepresented anatomical regions where data acquisition remains challenging. Multimodal inte grati on presents opportunities to lev erage complementary information across imag- ing modalities. PET/CT and PET/MR combinations demonstrate the clinical v alue of multimodal approaches, while the emer gence of vision-language datasets that combine 3D v olumes with clini- cal reports and radiology texts opens ne w possibilities for cross-modal reasoning and clinical con- text understanding. Be yond traditional imaging modality combinations, innov ati ve cross-domain multimodal approaches are emerging, such as integrating macroscopic imaging (CT/MRI) with mi- croscopic pathology data. These pathology-imaging combinations offer unique opportunities to bridge the gap between radiological findings and histological ground truth, enabling AI systems to learn from both macroscopic anatomical structures and microscopic tissue characteristics. Such approaches can significantly enhance diagnostic accuracy by combining CT’ s ability to detect and localize lesions with pathology’ s role as the diagnostic gold standard, creating more robust and clinically-relev ant AI systems. Adv ances in cross-modal alignment techniques enable more sophis- ticated fusion strategies that can fav orably enhance diagnostic capabilities across these div erse data types. Multi-task learning paradigms offer transformati ve potential for 3D medical imaging, analogous to the "one-for -all" paradigm ex emplified by ChatGPT in natural language processing. Rather than training separate models for individual tasks, integrated frameworks can simultaneously address multiple tasks ( e.g . , segmentation, classification, and detection) within unified architectures. This approach not only improves computational ef ficiency but also enables knowledge transfer across related tasks, particularly v aluable giv en the limited scale of individual 3D datasets. Multi-task datasets that provide di verse annotation types for the same volumetric data can unlock syner gistic learning ef fects, where performance on individual tasks benefits from joint optimization across mul- tiple objecti ves. The vision of a unified diagnostic and generati ve model that can handle multiple clinical tasks simultaneously represents a paradigm shift to ward more versatile and ef ficient clinical AI systems, similar to ho w foundation models have revolutionized natural language understanding and generation. Looking forward, the transition toward foundation-oriented data engineering paradigms demands fundamental changes in how 3D medical datasets are conceptualized and structured. Future dataset designs should prioritize adaptability and extensibility , enabling researchers to deri ve ne w tasks and applications from existing resources. Strategic dataset consolidation through systematic metadata harmonization, combined with adv ances in self-supervised learning and cross-modal reasoning, po- sitions the 3D medical imaging domain for significant breakthroughs in clinical AI applications. 38 (a) (b) (c) Figure 15: The distribution of different (a) modalities, (b) anatomical structures, and (c) tasks for video datasets 5 Medical V ideo Datasets Medical video datasets are crucial resources for de veloping algorithms that le verage spatiotemporal information in dynamic clinical scenarios, such as minimally inv asi ve surgery , medical education, and video-based diagnosis. In contrast to static image datasets, video data facilitates the model- ing of motion patterns, procedural workflo ws, and temporal consistencies, which are essential for tasks such as surgical instrument tracking, cross-frame anatomical structure segmentation, or phys- iological motion estimation. This survey identifies 77 medical video datasets, comprising a total of 166,691 samples. These datasets span a diverse range of tasks, imaging modalities, and anatomical structures. All video datasets are illustrated in the T ab. 26 . 5.1 Overview Fig. 15 illustrates the distribution of video datasets across different anatomical structures, imaging modalities, and tasks. The most pre v alent anatomical structures are the stomach, colon, and esopha- gus, with each category indi vidually accounting for about 30% of the total videos. In contrast, other anatomical structures, such as the retina, heart, pupil, and iris, are significantly underrepresented, each constituting less than 2% of the total collection. Similarly , the distribution across imaging modalities is highly ske wed, with endoscopy alone accounting for a substantial 85.9% of the videos. Consequently , modalities such as ultrasound microscopy , and RGB remain scarce, which highlights a critical need for larger-scale datasets to mitigate potential modality bias, particularly in the devel- opment of foundation models. In contrast to the severe imbalances observed across modalities, the task distribution is more moderate, though still demonstrably long-tailed. Classification, detection, and segmentation represent the most common tasks, followed by estimation, generation, and VQA, whereas tracking, retrie v al, and registration are notably underrepresented, warranting further in vesti- gation. Given the se v ere modality imbalance, task-le vel information helps distinguish the properties of the collected video datasets. Therefore, the organizational structure of this section deviates from that of Sections 3 and 4 . Specifically , the video datasets are introduced primarily based on their as- sociated tasks rather than modality , with a supplementary analysis of the corresponding modalities and anatomical structures. 5.2 T ask Below we introduce the major tasks in the collected video datasets. Figure 16 demonstrates common tasks in medical video datasets, including video classification, video segmentation, video detection, video tracking, video estimation, and video registration. 5.2.1 Classification Classification in medical videos in volves assigning cate gorical labels to entire sequences or specific temporal segments by le veraging spatial-temporal features. This task is fundamental to a wide range of clinical and surgical applications, such as sur gical phase recognition, skill assessment, and disease diagnosis. T o date, 40 datasets comprising 81,701 samples hav e been identified for this purpose, with endoscopy dominating as the primary modality . Performance is commonly ev aluated using metrics 39 such as accurac y , F1-score, and the Area Under the R OC Curv e (A UC), while temporal metrics like the segmental edit score are also employed to assess sequence-le v el consistency . Representativ e datasets span di verse surgical domains. Cholec80 (# 1098 ) and its deri vati v es, in- cluding CholecT50 (# 1082 ) and the CholecTriplet challenges (# 1083 ),(# 1107 ), provide laparoscopic cholecystectomy videos annotated with surgical phases, instrument presence, and fine-grained triplets. These resources serve as benchmarks for workflow analysis and activity recognition. SurgV isDom (# 1084 ) contains 488 bo wel surgery videos with phase anno- tations, enabling cross-domain generalization studies. As the largest public dataset for gastroin- testinal endoscopy , HyperKvasir (# 1152 ) is the largest publicly a v ailable gastrointestinal endoscopy dataset, comprising 373 videos and ov er 110,000 video frames annotated for anatomy and pathol- ogy , supporting classification, localization, and captioning. In the field of ophthalmic surgery , the CA T ARACTS (# 1085 ) and Cataract-1K datasets (# 1130 ) provide microscopy videos annotated for surgical phases, instruments, and pix el-lev el segmentation, facilitating multi-task modeling. Other influential resources include the EndoV is W orkflow and Skill Assessment (SWSA) series (# 1087 ) for phase and skill classification, and SAR-RARP50 (# 1127 ), the first public robot-assisted radical prostatectomy dataset with synchronized action and instrument annotations. 5.2.2 Segmentation Medical video segmentation in v olves the frame-by-frame delineation of anatomical structures, pathological re gions, or surgical instruments to enable precise spatio-temporal analysis. This task is crucial for applications including real-time surgical guidance, quantitati ve organ motion track- ing, and automated assessment of lesion dynamics. Our revie w encompasses 32 datasets tailored for segmentation, containing a total of 18,739 video instances. Performance is typically ev aluated using metrics such as the Dice similarity coefficient, Intersection-over -Union (IoU), and pixel-le vel accuracy . Representativ e datasets highlight both surgical and microscopic domains. The Robotic Instru- ment Segmentation (RIS) (# 1116 ) and Kidney Boundary Detection (KBD) (# 1114 ) datasets in- troduced pixel-le v el annotations for robotic surgical tools and anatomical boundaries, establish- ing early benchmarks for intraoperati ve vision. In ophthalmology , Cataract-1K (# 1130 ) combines phase annotations with 2,256 manually se gmented frames for cataract sur gery , enabling joint analy- sis of workflo w and fine-grained structures. The HyperKvasir dataset (# 1152 ), while primarily used for gastrointestinal classification, also includes segmentation masks for anatomical landmarks and pathological findings across 373 endoscopic videos. More recent challenges extend segmentation to complex multi-modal and 3D contexts, For instance, P2ILF (# 1120 ) combines laparoscopic video and CT for li ver landmark delineation, while SAR-RARP50 (# 1127 ) is the first public dataset of robot-assisted radical prostatectomy videos with synchronized instrument se gmentation and action recognition. 5.2.3 Detection V ideo detection aims to identify and localize tar get objects, such as lesions, instruments, or anatom- ical landmarks, within individual frames of a video while le veraging temporal continuity to improv e robustness. This capability is crucial for early disease screening, intraoperative navigation, and au- tomated procedural quality assessment. W e identified 27 datasets for the detection task. Commonly used eval uation metrics include precision, recall, mean A verage Precision (mAP), and frame-le vel F1-core. In our surve y , these 28 datasets comprise 49,507 samples emphasize detection. Evaluations typically report precision, recall, mean A verage Precision (mAP) at bounding-box or mask-le vel IoU thresholds, and frame-lev el F1. For temporally aggregated predictions, some studies additionally re- port video-mAP or track-aware scores to penalize fragmented detections. Representativ e resources span lesion, artifact, and instrument detection across multiple surgical do- mains. GIANA (# 1110 ) and EndoCV (# 1105 ) provide endoscopic polyp detection benchmarks with bounding-box or mask annotations, stressing generalization across centers and devices. Instrument- centric datasets include the m2cai16-tool-locations (# 1108 ) set and the large-scale SurgT oolLoc challenges (2022–2023) (# 1091 ),(# 1125 ) with tens of thousands of annotated frames for robotic and laparoscopic tools, enabling strong baselines for real-time instrument awareness and do wn- stream workflow understanding. Beyond the abdomen, ophthalmic datasets such as CA T ARA CTS or Cataract-1K and LensID (# 1131 ) support tool and structure detection in cataract surgery , while 40 PitV is (# 1124 ) focuses on transsphenoidal neurosurgery with step- and instrument-lev el labels. Broader clinical cov erage is of fered by A V OS (# 1112 ), a multi-procedure open-surgery corpus with dense annotations that enables cross-procedure detection, tracking, and localization. Recent multi-domain collections such as SARAS-MESAD (# 1155 ) further test robustness by mixing real and phantom data under shared action or instrument vocab ularies. Across these datasets, anno- tation granularity ranges from sparsely sampled frames to densely labeled clips, with boxes, in- stance masks, or k eypoints. Emerging trends include spatiotemporal tube proposals, joint detection- tracking protocols, and rob ustness benchmarks under realistic corruptions, which together mov e detection from frame-wise recognition tow ard reliable, clinically usable video understanding. 5.2.4 T racking T racking in medical videos entails following the spatiotemporal trajectories of objects of interest, such as surgical tools or anatomical landmarks, across consecuti ve frames. This task is fundamental to applications such as workflow analysis, motion quantification, and dynamic process monitoring. Our surv ey identified 8 datasets with 2,420 samples dedicated to tracking. The tracking task usually employs metrics such as Multiple Object T racking Accurac y (MO T A), Multiple Object T racking Precision (MO TP), identity switches (IDSW), and track purity . Representativ e datasets focus on the surgical domain. For example, the m2cai16-tool-locations dataset (# 1108 ) pro vided laparoscopic tool-tip trajectories, while the EndoV is tracking challenges expanded to encompass tracking, tissue motion estimation, and joint detection–tracking tasks. SurgT (# 1141 ) and SARAS-MESAD (# 1155 ) further incorporated stereoscopic vie ws, soft-tissue tracking, and phantom–real domain variations. Beyond endoscop y , STIR (# 1142 ) provided infrared–visible paired videos for surgical tissue tracking, and the large-scale dataset A V OS deliver ed dense per - frame annotations across 47 hours of open surgery from 23 procedure types. Specialized datasets such as HiSW A-RLLS (# 1143 ) for robotic li ver resection and EgoSurgery (# 1144 ) with egocen- tric video plus gaze data highlight emerging subfields, reflecting a recent trend tow ard multi-task benchmarks that unify detection, se gmentation, and temporal association for comprehensiv e spa- tiotemporal understanding. 5.2.5 Estimation Estimation tasks in medical video analysis aim to deri ve quantitativ e variables from temporal se- quences, such as depth maps, motion fields, or physiological parameters. Applications include 3D reconstruction from monocular endoscopic videos, camera pose estimation for na vigation, respira- tory motion estimation, and sur gical skill scoring. Our surv ey identified two datasets dedicated to this task. The SimCol-to-3D (# 1113 ) dataset contains simulated colonoscopy videos for depth pre- diction and camera pose estimation, with 15 sequences annotated for both simulated and real pro- cedures, enabling ev aluation under controlled and clinical conditions. The challenge also includes a Colposcopy subset with 30 videos for depth estimation in gynecological imaging. The Endovis 2019-SCRE (# 1151 ) dataset contains videos from 9 medical sites for the task of dense depth estima- tion. The corresponding depth maps were obtained from structured light data captured using porcine cadav ers. Evaluation metrics are task-specific, including mean absolute error (MAE), endpoint error (EPE), and correlation coefficients. Moreover , recent works ha ve increasingly adopted multi-task formulations that jointly estimate depth, pose, and motion to improv e downstream surgical naviga- tion and workflo w understanding. 5.2.6 Registration Registration in medical video analysis in volv es aligning multimodal data, such as 2D video en- doscopy with 3D computed tomography (CT), to establish a consistent spatial correspondence across imaging modalities. This process is crucial for intraoperativ e guidance, anatomical structure map- ping, and enhanced visualization of surgical fields. In our survey , one datasets with a total of 167 samples were identified for registration tasks. The P2ILF dataset (# 1120 ) provides paired endoscopy videos and CT scans, and is designed for ev aluating multimodal re gistration methods. The dataset included 25 cases (10 for training, 10 for validation, and 5 for testing) with both 3D model and video-endoscopic data, supporting cross- modality alignment and benchmarking re gistration accurac y . The registration is performed between the landmarks of the 3D model and those extracted from the videos. 41 Lower GI tr act Upper GI tract √ × Video Classification Video Segmentation Video Detection Video Estimation Video T racking Bagging Prost ate: 1~4 s Video Registration Figure 16: Demonstration of the collected video datasets from different tasks. The figure for the video estimation task is from the EndoV is 2022-SimCol-to-3D dataset (# 1113 ), and the figure for the video registration task is from the EndoV is 2022-P2ILF dataset (# 1120 ). Endoscopy Ultrasound RGB Microscopy Figure 17: Illustration of four major modalities in the collected video datasets. Evaluation metrics for registration commonly include T arget Registration Error (TRE), the Dice Similarity Coefficient (DSC) for segmented structures, and success rates within clinically acceptable error thresholds. T ogether , the P2ILF dataset form a comprehensiv e benchmark for de veloping and validating multimodal re gistration approaches in minimally in v asi ve li v er surgery . 5.3 Modalities Among the 77 medical video datasets identified in our survey , endoscopy constitutes the v ast major - ity with 56 datasets, underscoring its central role in documenting dynamic intraoperati ve and diag- nostic procedures. This prev alence is attributable to sev eral factors: the routine integration of video recording systems in surgical suites, the ease of acquiring high-resolution footage during standard procedures, and the relati v ely straightforward annotation of visible anatomical structures or sur gical tools without requiring complex multi-view reconstruction. Longstanding community initiativ es, such as the EndoV is challenges, hav e further accelerated dataset generation and standardization, fostering a virtuous cycle between benchmark a v ailability and method dev elopment. In contrast, other imaging modalities are notably underrepresented. Microscopy videos (10 datasets) are often recorded during ophthalmic surgery to demonstrate detailed anatomical structures of the eye, such as the retina, iris, and pupil. Ultrasound (7 dataset) are infrequently acquired as continuous cine sequences due to clinical workflow constraints and the need for specialized protocols, such as dynamic perfusion studies or echocardiography loops. The RGB refers to videos captured with a camera in open en vironments and is most often associated with non-surgical scenarios, such as instructional recordings for emer gency care, nursing, or simulated sur gical procedures. Figure 17 illustrates four major modalities in the collected video datasets. 42 5.4 Anatomical Structures Most of the video datasets (40 datasets) focus on abdominal anatomical structures, with a lar ge pro- portion related to the gallbladder (13 datasets) and the colon (6 datasets). This is because many of these datasets were collected during procedures such as cholecystectomy and endoscopy . Se ven datasets focus on eye-related anatomical structures, including the iris and pupil, with most of the videos collected during cataract surgery . The remaining video datasets cov er a wide range of anatomical structures across the body , including the thyroid (1 dataset), pituitary (2 dataset) and placenta (1 dataset). A portion of the videos were also collected from non-human structures, such as artificial blood vessels (1 dataset) and porcine cada vers (1 dataset). 5.5 Challenges and Opportunities The de velopment of medical video datasets has enabled substantial progress across segmentation, detection, tracking, and registration, yet the field continues to face enduring challenges . Annotation quality . On one hand, generating pixel- or frame-le vel ground truth requires e xtensi ve expert labor , particularly in domains such as surgical tool segmentation and landmark tracking, where precision and temporal consistenc y are essential [ 78 , 79 , 80 , 81 , 82 ]. Few datasets e xcept the CaDIS dataset provide fine-grained, frame-lev el annotations covering full scenes in videos. On the other hand, there may be v ariations in annotation quality across different videos, even within the same dataset, due to dif ferences in sur geon skill when multiple annotators are inv olved. Sparse or weak labels hav e been proposed as a compromise, b ut the y often limit the reliability of do wnstream ev aluation. Semi-supervised and synthetic data augmentation approaches show promise, though their acceptance in clinical research requires rigorous validation. Data privacy . Unlike natural video, medical recordings inherently encode sensitiv e patient informa- tion. De-identification is particularly challenging in endoscopy and surgery , where anatomical con- text itself can serve as a patient identifier . Consequently , dataset releases are frequently restricted in scale or geographic scope, hampering the establishment of broadly generalizable benchmarks [ 83 , 84 , 85 ]. Addressing this requires technical advances in anonymization as well as standardized regulatory and ethical frame works that enable secure multi-center data sharing. Domain shift represents a further persistent issue. Substantial v ariability arises from dif ferences in imaging de vices, acquisition protocols, and surgical practices, often causing models trained on one dataset to fail when applied to another . This problem has been observed across lesion detec- tion, artifact remov al, and instrument recognition benchmarks [ 86 , 87 , 88 , 89 ]. While phantom or synthetic data help isolate algorithmic behavior , bridging these controlled conditions with the com- plexity of real clinical en vironments remains an open research frontier . Rob ust domain adaptation, self-supervised pretraining, and benchmark designs that explicitly incorporate cross-institutional variation are therefore pressing needs. Computational bur den. From a computational standpoint, the scale of medical video poses formidable demands. High-resolution intraoperativ e recordings can span hours, making storage, annotation, and real-time analysis resource-intensiv e. Real-time deployment, for instance in robotic surgery or intraoperati ve na vigation, requires methods that balance accuracy with computational ef- ficiency [ 90 ]. Furthermore, emerging benchmarks increasingly combine multiple tasks—detection, segmentation, and tracking—placing pressure on algorithm design to unify spatiotemporal reasoning under constrained latency . These challenges, ho we ver , also motiv ate transformati ve opportunities . The rise of multimodal datasets such as P2ILF and SAR-RARP50 opens pathways to ward comprehensiv e scene understand- ing, aligning 2D video streams with 3D imaging modalities and enabling clinically relev ant multi- modal registration [ 91 , 92 ]. The integration of large pre-trained models and foundation architectures has the potential to mitigate annotation bottlenecks and impro ve generalization across institutions, provided that interpretability and domain alignment are addressed. Longstanding community ini- tiativ es, such as the EndoV is series challenges, further underscore the importance of standardized ev aluation protocols for reproducibility and clinical translation. Clinically , the opportunities are pro- found. Accurate lesion detection and temporal localization can support early diagnosis in screening procedures, while reliable instrument tracking and w orkflo w analysis enable intraoperati v e decision support and skill assessment [ 93 , 94 ]. More broadly , the con v ergence of diverse datasets, robust 43 ID Field Brief description 1 dataset_name Official name or commonly used short name of the dataset. 2 release_date First public release date (YYYY -MM or YYYY -MM-DD; use NA if unknown). 3 homepage_url Stable URL or DOI for the dataset homepage, paper , or repository . 4 organization Institution(s) releasing or hosting the dataset; multiple entries allowed, separated by commas. 5 challenge_series Name of the associated challenge or benchmark series; NA if not challenge-based. 6 license Data usage license or access polic y as specified by the download agreement. 7 dataset_description Short free-text summary of source, modality , tasks, scale, and key characteristics. 8 modality_primary Primary imaging modality or modalities (e.g., CT , MR, X-ray , Fundus). 9 modality_secondary Subtype or sequence within the primary modality (e.g., MR:T1, CT:CT A; NA if unspecified). 10 anatomical_structure T arget org an, region, or lesion; multiple structures allo wed. 11 disease Disease or clinical condition(s) represented; NA for non disease-specific datasets. 12 data_volume T otal size and split, preferably as JSON (e.g., {"total":..., "train":...} ). 13 valid_image_n Usable sample count after cleaning, optionally in the same JSON format as data_volume . 14 label_presence Annotation av ailability: labeled , unlabeled , or mixed . 15 task_type Supported computational tasks (e.g., segmentation, detection, classification, VQA). 16 num_classes_per_task JSON describing, per task, the number of classes/targets and relevant settings. T able 2: Definition of data_meta fields for dataset-lev el metadata. D a t a se t So u rce Po o l T C I A T C G A I SI C ··· A D N I F i l t e r Mo d e 1: JSO N R u l e s " d i m e n s i o n " : " 2 D " , " m o d a l i t i e s " : [ " C T " , " M R " ] , " t a s k _ t y p e s ": [ . . . ] , " a n a t o m i e s " : [ . . . ] , " l i c e n s e " : [ . . . ] , " m i n _ v a l i d _ n u m ": 1 0 0 0 , ... F i l t e r Mo d e 2: D i re ct Se a rch Ke yw o rd , d a t a se t n a me s, o rg a n i za t i o n , . . . D a t a se t F u si o n 2. A l i g n m e n t : a l i g n d a t a s e t t a s k s e m a n t i c s a n d l a b e l s 3. B l u e p r i n t : us e m e t a d a t a to c l u s t e r d a t a s e t s a n d a s s e s s p o t e n t i a l f o r f u s i o n 4. I n d e x i n g : b u i l d p u b l i c i n d i c e s a n d v i s u a l s u m m a r i e s 1. H a r m o n i z a t i o n : s t a n d a r d i z e m e t a d a t a a c r o s s d a t a s e t s D a t a se t St a t i st i cs I n t e ra ct i ve D i sco ve ry Po rt a l M e t a d a t a S u m m a r y S t a t i s t i c s V i s u a l i z a t i o n T a b l e f o r M a t c h i n g D a t a s e t s Figure 18: Pipeline of our dataset collection, processing, fusion, and summarization system based on the proposed dataset fusion paradigm, implemented in our interactiv e discov ery portal. benchmarking, and advanced learning paradigms is steering medical video analysis from narro w research prototypes tow ard clinically indispensable technologies. In summary , medical video datasets face inherent challenges in annotation, priv acy , domain robust- ness, and scalability , but these limitations are also drivers of innov ation. W ith sustained progress in dataset div ersity , federated ev aluation, and integration with lar ge-scale learning systems, the field is positioned to deliv er clinically impactful solutions in the coming decade. 6 Paradigm f or Dataset Fusion Despite the ab undance of public medical imaging datasets, their fragmentation significantly hinders their effecti ve use in lar ge-scale model training. T o address this, we propose the Metadata-Driven Fusion P aradigm (MDFP), grounded in our comprehensi ve collection and curation of medical imag- ing datasets, of fering an efficient, scalable, and metadata-centric strategy to systematize discovery , 44 Block Role Representativ e fields and examples record Sample identifiers and cross-references dataset_name , image_path , optional sample IDs used to locate the un- derlying media file and join annotations across tasks. context Clinical and textual conte xt Subject- and acquisition-le vel metadata (e.g., subject ID, age, sex, site, modality , anatomy), extensible extra dictionary , free-text descriptions or reports. media_geometry Media attributes and geometry Media-lev el task configuration and spatio-temporal metadata, including task_type , leaf_task , annotation_type , dimension , pixel spacing, orientation, slice/frame indices, timestamps, camera parameters. tasks T ask-specific annotation payloads Structured labels for the predefined tasks, such as segmentation masks, de- tection boxes, class labels, polygons, or keypoints, grouped by dimension and schema_variant . T able 4: Four logical blocks— record , context , media_geometry , and tasks —structuring annotations_{task}.jsonl . The blocks separate sample identity , context, media-le vel geom- etry , and task-specific annotation payloads. auditing, and composition of multiple datasets. By operating primarily on metadata rather than raw pixels, MDFP reduces handling overhead and priv acy risk, impro ves reproducibility and auditability , and enables rapid goal-conditioned dataset assembly . Based on MDFP , we build an interactiv e dis- cov ery portal that supports fine-grained dataset search, inte gration, and statistical analysis. Figure 18 ov erviews the full system, and Figure 19 details MDFP . The remainder of this section proceeds in the follo wing order: we describe dataset collection and processing (Section 6.1 ); introduce MDFP and its four phases (Section 6.2 ); incorporate the afore- mentioned components to form the interactiv e discov ery protal (Section 6.3 ). 6.1 Dataset Collection and Processing All datasets included in this study were obtained from publicly accessible web-based repositories, such as The Cancer Imaging Archi ve (TCIA) 5 , Grand Challenge 6 , OpenNeuro 7 , Kaggle 8 , Neu- roImaging T ools and Resources Collaboratory (NITRC) 9 , Synapse 10 , CodaLab 11 , GitHub 12 , etc. After collecting the medical imaging datasets from these sources, we org anize them into a multi- dimensional database that serves as a comprehensive overvie w table. This database categorizes each dataset by multiple attributes, including dimension, modality , anatomical structure, number of cases, label av ailability , and task type, along with other essential metadata. Such an organization enables flexible querying and filtering, allowing researchers to quickly retrie v e datasets that match specific research needs, i.e. , training a 3D foundation model for CT , MRI, and PET . For each indi vidual dataset, we preserve the original directory layout as much as possible and en- rich it with two JSONL files. The file data-meta.jsonl records dataset-level information such as release date, imaging modality , homepage URL, and license, using 16 well-defined fields (summa- rized in T able 2 ). The file annotations-{task}.jsonl stores per-media, task-specific annotations (for example, mask file paths for segmentation tasks or bounding boxes for detection tasks). Each JSON object describes a single annotated media item for a particular task and is decomposed into four logical information blocks: record , context , media_geometry , and tasks . The record block contains stable identifiers such as dataset_name (dataset identifier) and image_path (path or ke y of the underlying image, volume, or video file within that dataset), which link the annota- tion back to the original media file and allow annotations for the same sample to be joined across tasks. The context block collects optional subject- and acquisition-level metadata and free-text descriptions (e.g., subject ID, age, sex, site, modality , anatomy , and an e xtensible extra dictio- nary for dataset-specific fields). The media_geometry block captures media-lev el attributes and 5 https://www.cancerimagingarchive.net 6 https://grand- challenge.org 7 https://openneuro.org 8 https://www.kaggle.com/ 9 https://www.nitrc.org 10 https://www.synapse.org 11 https://codalab.lisn.upsaclay.fr 12 https://github.com 45 T arge t task ty pe s : [ c la ssif ic a t io n , se gm e nta tion, de te c tion , r e gr e ssion] ; L ic e n se allow li st : none . in c lu de u n labe le d : y e s; pr e f e r la be le d in se le c tion: y e s; m i n v a l i d_i m a ge _n : pe r da ta se t: 100. S e le c ti on e n a b le d : F a lse ( m in_ da ta se t s_ p e r _m oda l i ty = 2, m in_or gs_p e r _ m oda li t y = 2) ... G oal : P r et ra in a 2 D m edica l f o unda t io n m o del o v er m o da lit ies [ CT , M R, F undus ] . R ef i n em en t M D F P Pro c e ss P ha s e 1: H a rm oni s a t i on P ha s e 2: A l i gnm e nt P ha s e 3: Bl ue pri nt P ha s e 4: Inde xi ng I np ut Cu r at e d D at abas e O ut p ut m o d a lit y n _d at as e t s s u m _i m age n _or gs la b e le d CT 10 1173965 4 1 F und us 42 280311 17 0. 95238 M R 5 681025 2 1 Figure 19: Detailed process of the proposed metadata-dri ven fusion paradigm (MDFP). spatial/temporal geometry that are shared by all annotations on the same media item, including the high-lev el task_type (one of 12 predefined task categories such as segmentation, detection, or clas- sification), the dataset-specific leaf_task , the annotation_type (e.g., binary masks, bounding boxes, polygons), the dimension (2D, 3D, or video), and imaging metadata such as pixel spacing, orientation, slice or frame indices, timestamps, and camera parameters. Finally , the tasks block contains the structured task-specific annotation payloads themselv es (e.g., mask references, box co- ordinates, class labels, keypoints), organized in a schema that is consistent across datasets for the same task type. A compact ov erview of these four blocks and their representative fields is giv en in T able 4 . Building on the standardized directory structure, we further unify file formats with a focus on pre- serving quantitative information and metadata. Specifically , all v olumetric imaging data ( e.g. , CT , MRI, PET) are conv erted to NIfTI (.nii.gz), with vox el spacing, orientation (qform/sform), and in- tensity scaling (slope/intercept or equiv alent) preserved; dynamic PET is stored as 4D NIfTI with companion JSON/TSV files for frame timing and calibration ( e.g. , SUV factors). When appropriate, de-identified source DICOMs are retained as an optional raw layer . For 2D modalities ( e.g. , radio- graphs, ultrasound frames), we preserv e full dynamic range using lossless 16-bit formats (TIFF or 16-bit PNG) together with sidecar JSON for essential metadata (pixel spacing, orientation, window settings, modality-specific tags). For video data ( e.g. , endoscopy , ultrasound cine, sur gical record- ings), we store sequences in compressed formats (MP4 or A VI with H.264/H.265 encoding) while maintaining 8-bit color depth for most clinical videos; specialized applications requiring higher dynamic range ( e.g . , fluorescence microscopy , high-speed recordings) are preserv ed as 16-bit se- quences when av ailable. V ideo metadata, including frame rate, resolution, acquisition timestamps, and equipment parameters, are recorded in companion JSON files to ensure reproducibility and temporal consistenc y . Optional 8-bit PNG thumbnails may be generated solely for visualization and documentation; these are never used as training inputs when quantitati ve intensity matters. This preprocessing and standardization workflo w maintains dataset fidelity and compatibility , facilitates seamless integration into model-training pipelines, and enables reproducible and comparable bench- marking across studies. 6.2 MDFP MDFP systematizes discov ery , auditing, and composition through four sequential phases that oper- ate primarily on structured metadata, strengthening pri v acy , auditability , and reproducibility while av oiding raw-pix el handling. T able 5 presents these phases of MDFP , outlining their core objectives and associated metadata fields. These phases are tightly aligned with our systematic metadata collection framew ork, ensuring 46 T able 5: MDFP W orkflow Overvie w with K e y Metadata Fields Phase Objective Metadata Fields 1. Harmoniza- tion Standardize modalities, tasks, and anatomy . modality_primary , dimension , anatomical_structure , organization , challenge_series 2. Alignment Align semantic labels and tasks across datasets. task_type , modality_secondary , label_presence , notes 3. Blueprint Cluster datasets; assess integrati ve po- tential and data scale. data_volume , valid_image_n , storage_size_gb 4. Indexing Create public metadata indices and vi- sualization tools for easy access. dataset_name , release_date , homepage_url , license consistency , completeness, and interoperability across heterogeneous datasets. Below , we detail each phase. 6.2.1 Phase 1: Metadata Harmonization Phase 1 resolves semantic heterogeneity by enforcing a rigorously defined metadata schema. Rather than creating a new v ocab ulary , we ground our schema in authoritative medical terminologies, such as the Unified Medical Language System (UMLS) and Medical Subject Headings (MeSH) [ 95 ]. This process is semi-automated, lev eraging API-driv en searches against these ontologies, followed by LLM-based refinement to programmatically align disparate dataset descriptors into a consistent, machine-readable form. Concretely , we: • Standardize primary modality ( modality_primary ): Mapped to an enumerated set in- cluding CT , MR, PET , US, X-ray , etc. , with niche modalities deterministically aligned to this taxonomy . • Normalize data dimensionality ( dimension ): Parsed directly from dataset metadata to determine whether the data is 2D, 3D, or video (2D + time). • Establish hierar chical classification : Instead of a simple anatomical tag, we implement a multi-level classification system based on standard medical ontologies ( e.g. , UMLS, MeSH). This provides rich, hierarchical context. For e xample, a dataset on cataracts w ould be classified under Eye Diseases → Lens Diseases → Cataract . • Record prov enance and context ( organization , challenge_series ): Identifying the originating institution and any associated benchmark or competition series. • Document annotation granularity ( annotation_type ): Explicitly cataloging the type and granularity of labels provided, including landmark coordinates, pixel-le vel segmenta- tion masks, region-lev el bounding boxes, image-level classification labels, or multi-modal annotations. This metadata enables researchers to identify datasets compatible with their task requirements and facilitates appropriate fusion strategies during retrie v al. This harmonization step yields a uniform, richly annotated metadata table that transforms a frag- mented corpus into an interoperable resource, thereby enabling reliable cross-dataset comparison, reproducible filtering, and seamless integration. 6.2.2 Phase 2: Semantic Alignment Phase 2 mitig ates inconsistencies by mapping abstract machine learning tasks to their concrete clin- ical significance. This crucial step in v olves a systematic re vie w of dataset documentation, source publications, and of ficial guidelines to understand the intended real-w orld application. By doing so, we align heterogeneous labeling con ventions and ev aluation objectiv es with tangible clinical goals. Specifically , we: • Define downstr eam tasks ( downstream_task ): W e standardize ML tasks and explicitly map them to their clinical applications. F or example: 47 – A classification task might correspond to clinical diagnosis ( e.g. , malignant vs. benign tumor), severity grading ( e.g. , staging diabetic retinopathy), or tr eatment r e- sponse pr ediction . – A segmentation task may be used for lesion delineation ( e.g. , outlining a tumor boundary), volumetric quantification ( e.g . , measuring organ volume to track disease progression), or radiother apy/sur gical planning . – A detection task is often used for disease scr eening ( e.g . , identifying candidate pulmonary nodules in a chest CT). – A regression task can quantify clinical biomarkers ( e .g. , predicting bone mineral density from a CT scan or cardiac ejection fraction from an ultrasound video). • Indicate label a vailability ( label_presence ): Denoting whether ground-truth annota- tions are provided ( labeled ) or not ( unlabeled ). • Specify secondary imaging modalities ( modality_secondary ): Capturing finer -grained protocol-lev el distinctions under each primary modality , such as T1 or T2 sequences for MR. • Document special considerations ( notes ): Capturing dataset-specific nuances, assump- tions, or known limitations in free-te xt form. This alignment phase yields a clinically-grounded task vocab ulary that supports meaningful inter - pretation, goal-oriented filtering, and enhances the reliability of cross-dataset benchmarking. 6.2.3 Phase 3: Fusion Blueprints This phase le verages harmonized metadata to design strategic dataset integration plans. Specifically , we perform grouping and categorization based on combinations of primary and secondary imag- ing modalities ( modality_primary , modality_secondary ), clinical tasks ( task_type ), and anatomical cov erage ( anatomical_structure ). This grouping process consolidates datasets with similar or identical attributes into unified groups, pro viding a structured foundation for designing fu- sion blueprints that guide principled dataset integration. Quantitativ e ev aluations are systematically deriv ed from metadata, encompassing the follo wing aspects: • Data V olume ( data_volume ) : Assess total images available, along with explicit training, validation, and testing splits. • V alid Image Counts ( valid_image_n ) : Determine precisely ho w many images hav e re- liable and validated annotations, critical for training supervised models. These quantitativ e statistics are obtained directly from official dataset documentation, README files, pub- lished papers, and other authoritativ e public resources provided by dataset curators. • Storage Estimation ( storage_size_gb ) : Ev aluate practical storage requirements, essen- tial for infrastructure planning. • Anatomical and T ask Diversity ( anatomical_structure , task_type ) : Quantify anatomical breadth and task variety within each fusion cluster, ensuring coverage div er - sity crucial for generalization. This structured assessment produces a principled basis for scalable dataset mer ging, balancing quan- tity , annotation quality , and content div ersity to support robust foundation model training. Dur - ing fusion blueprint design, we explicitly account for data heterogeneity captured in Phase 1, in- cluding v ariations in imaging protocols ( e .g. , dif ferences in CT reconstruction parameters or MRI field strengths), image resolutions, and annotation granularities. Our tool systematically identi- fies and flags datasets with incompatible annotation types ( e.g. , mixing pixel-lev el segmentation with bounding-box detection) and imaging protocol dif ferences, alerting researchers to potential in- tegration challenges. These metadata annotations help researchers mak e informed decisions about whether harmonization preprocessing, protocol-aware sampling strategies, or domain-adaptive train- ing approaches are needed to ensure cross-dataset compatibility and model robustness. 48 T able 6: MDFP-derived composition for the 2D CT/MR/Fundus goal. sum_image is sum(valid_image_n) ; labeled_ratio is the fraction of datasets that are labeled. modality n_datasets sum_image n_orgs labeled_ratio CT 10 1,173,965 4 1.000 MR 5 681,025 2 1.000 Fundus 42 280,311 17 0.952 6.2.4 Phase 4: Dataset Indexing and Community Sharing Phase 4 transforms the harmonized metadata into a structured, publicly accessible dataset index to support community-scale discovery and reuse. W e consolidate key metadata elements for each dataset, including: • Dataset name ( dataset_name ): the canonical name of the dataset for standardized refer- encing; • Release date ( release_date ): official publication or release timestamp, enabling tempo- ral filtering; • Homepage URL ( homepage_url ): direct access link to dataset documentation or hosting platform; • License ( license ): clearly defined usage permissions, ensuring legal compliance and re- producibility . This index ed representation f acilitates rapid dataset disco very , promotes responsible reuse, and pro- vides the infrastructure foundation for large-scale model pretraining, benchmarking, and open col- laboration. 6.2.5 Case Study: Goal-Conditioned Fusion via MDFP As shown in Figure 19 , to demonstrate how MDFP supports foundation-model pretraining with reproducible, goal-aligned data composition, we instantiate a concrete target: a 2D model over modalities {CT , MR, Fundus} and tasks {classification, segmentation, detection, regression}. In Phases 1–2 (Harmonization and Alignment), we apply the following filtering and standardization procedures: First, we standardize all datasets by mapping primary modalities to our controlled vo- cabulary (CT , MR, Fundus) and normalize data dimensionality to 2D only , explicitly excluding 3D volumetric data and video sequences to maintain dimensional consistenc y . Second, we establish hi- erarchical anatomical classifications using UMLS/MeSH ontologies for each dataset, enabling con- sistent cross-dataset anatomical mapping. Third, we perform semantic alignment by mapping ma- chine learning task types to their clinical applications, ensuring that selected datasets cov er the four target task families: classification (diagnosis, se verity grading), segmentation (lesion delineation, volumetric quantification), detection (disease screening), and regression (clinical biomarker quan- tification). Fourth, we apply quality filters including minimum sample size ( valid_image_n ≥ 100) to ensure statistical reliability . Both labeled and unlabeled datasets are retained, with preference giv en to labeled ones when multiple alternati ves exist. No license restrictions are enforced in this demonstration, though license-aware filtering is supported by the frame w ork. In Phases 3–4 (Blueprint and Indexing), we perform grouping and categorization based on the har- monized metadata from Phases 1–2. Specifically , we group datasets by modality-task combinations, assess the integrati ve potential of each cluster by quantifying data volume, annotation av ailability , and anatomical co verage, and generate a fusion blueprint that summarizes the composition strate gy . Finally , we create a structured metadata index with all essential fields ( dataset_name , modality , task , valid_image_n , license , homepage_url ) to enable reproducible access and community sharing. The resulting integrated dataset composition is summarized in T able 6 . The curated pool com- prises 57 datasets and 2,135,301 v alidated images across three imaging modalities: CT (10 datasets, 1,173,965 images, 4 organisations), MR (5 datasets, 681,025 images, 2 organisations), and Fundus (42 datasets, 280,311 images, 17 organisations). These numbers correspond to the summary fields 49 n_datasets , sum_image (defined as P valid_image_n ), n_orgs , and labeled_ratio in T a- ble 6 . All CT and MR datasets are fully annotated ( labeled_ratio =1.000), and Fundus datasets achiev e a high annotation rate ( labeled_ratio =0.952), yielding strong supervision across modal- ities while satisfying the case-study constraints (2D only; CT/MR/Fundus) and covering the four target task families defined in Phase 2 (classification, segmentation, detection, regression). This configuration constitutes a concrete instantiation of goal-conditioned dataset integration via MDFP . These aggre gate statistics have direct implications for foundation-model pretraining. The high la- beled fractions support multi-task supervised objectiv es, while the remaining unlabeled images can be exploited with auxiliary self-supervised losses. At the same time, the ske w in sum_image to- ward CT and MR (CT+MR ≈ 1.85 M images) suggests employing modality-a ware sampling strate- gies (e.g., temperature-based sampling or per-dataset caps) and task-stratified batching to prev ent ov er-representation of these modalities. Finally , although this case study restricts sources to 2D CT/MR/Fundus datasets for clarity , the same MDFP pipeline can be rerun with relaxed configura- tion (e.g., enabling allow_3d_as_2d_sources=true ) to augment the 2D pool with projected 3D or video data when broader cov erage is desired. 6.3 Interactive Disco very P ortal Combining the aforementioned components together, we b uild a lightweight interactive discovery portal, namely the Medical Dataset Br owser , to triage and refine candidate datasets before schema- lev el alignment. The portal is deployed as a single page static application on GitHub Pages 13 , exe- cutes entirely client-side, and consumes at runtime the standardized JSON artifact produced in 6.1 (for example, the cleaned and mer ged manifest). This design eliminates serv er-side dependencies, simplifies reproducibility , and enables pri vac y-preserving exploration. Below , we detail the pipeline. Dataset Filtering. Starting from the dataset source pool prepared in Section 6.1 , the portal exposes two complementary modes for dataset filtering. First, Rule-based filtering (“Filter Mode 1”). This mode implements the MDFP , accepting an editable JSON specification that encodes deterministic selection criteria, e.g. , image dimension (2D/3D), modality sets (CT/MRI/US/Pathology , etc. ), task types (segmentation, detection, classification, report generation), organ/anatomy whitelists, license constraints, minimum sample sizes, and year ranges. This recipe-lik e abstraction makes selections auditable and perfectly reproducible. Specifically , the controls for Phase 1&2 in MDFP (harmonization and alignment) and Phase 3&4 (blueprint and indexing) are integrated into the page to pre vie w downstream ef fects before commit- ting a batch run. During execution, the interface highlights in-progress elements and then surfaces consolidated outputs for inspection. In parallel, dir ect faceted sear ch (“Filter Mode 2”) pro vides dropdown facets and a free text query for fast exploratory narro wing. Both modes drive li ve visual summaries, complete bar and dough- nut charts of dimension-modality-task distrib utions, so users can immediately assess co verage and balance of the current subset. Statistics and summaries. After the dataset filtering, the portal renders liv e bar/pie summaries of modality , dimension, task, and anatomy distributions; a MDFP Phase-4 audit table exposes fields essential for screening and compliance: name , dimension , modality , task , organ , images (counts), year , organization , license , and link . These statistics and summaries can be ex- ported to CSV/JSON for benchmarking pipelines. T ogether , these views close the loop from search to fusion to shareable artifacts. Implementation details. The index.html bootstraps by loading the preprocessed JSON mani- fest, initializes an in-memory filter store, and applies deterministic, order -independent rule ev alua- tion entirely on the client. V isual analytics and tables are rendered with lightweight, dependency- minimal components; results are paginated to maintain interactivity on medium-to-lar ge corpora. Because the application is a self-contained static bundle, any user can fork, reconfigure the selection recipe, and redeploy an identical retrie v al en vironment without additional infrastructure. 13 https://tchenglv520.github.io/medical- dataset- browser/ 50 7 Discussion 7.1 Limitations in T ask Definition and Evolution of Data Engineering Paradigms Current open-access medical imaging datasets exhibit limitations in task definition, reflecting the task-oriented nature of early deep learning practices [ 96 , 11 ]. Most datasets target indirect down- stream tasks ( e.g. segmentation, classification, or detection), which served as proxies for clinical goals but remain distant from real-world applications. W ith the advancement of foundation models and LLMs, AI systems are shifting to ward direct, clinically rele v ant tasks such as disease diagno- sis, patient condition assessment, and treatment recommendation [ 97 ]. This paradigm shift creates a critical mismatch: existing datasets, designed for classical computer vision tasks, cannot be di- rectly utilized without substantial transformation. Ho we ver , re-annotation for clinically oriented tasks incurs prohibiti vely high costs, as these tasks demand e xpert-lev el medical kno wledge and high-quality annotations from domain specialists. For instance, a lung nodule segmentation mask does not indicate whether the nodule is benign, malignant, or requires biopsy—information essential for clinical decision support but absent in e xisting annotations. Bridging this gap requires resource-intensiv e re-annotation by radiologists, a process that scales poorly across large datasets. Consequently , the medical AI community faces a dual challenge: legac y datasets are misaligned with contemporary needs, yet creating new foundation-model-ready datasets remains economically and logistically prohibitive. Future data engineering must priori- tize fle xible annotation frame w orks that capture clinically meaningful information upfront, enabling adaptation to ev olving AI paradigms without complete re-annotation. 7.2 Scarcity of Multimodal Medical Datasets and Constraints in Further De velopment Multimodal medical data that integrates imaging modalities (CT , MRI, pathology) with clinical re- ports, genomics, and temporal records holds exceptional v alue for clinical diagnosis, yet remains exceedingly rare in the public domain [ 11 ]. Most open-access datasets are unimodal and lack stan- dardized frame works for multimodal collection and annotation [ 98 , 99 ], significantly restricting research in cross-modal reasoning and joint representation learning essential for next-generation medical AI. The challenge extends be yond data av ailability to fundamental issues of modal align- ment and semantic consistency . Dif ferent modalities operate on disparate scales: pathology captures microscopic cellular details, radiology visualizes organ-le v el structures, and clinical notes document temporal disease progression. Harmonizing these heterogeneous streams requires sophisticated alignment protocols and cross- modal v alidation standards that current datasets rarely pro vide. F or example, aligning a radiologist’ s report timestamp with the corresponding imaging study , or synchronizing pathology findings with longitudinal treatment records, demands metadata infrastructure lar gely absent in existing resources. Moreov er , the absence of standardized multimodal benchmarks impedes systematic ev aluation of cross-modal architectures. Researchers lack unified frameworks to assess whether models effec- tiv ely integrate complementary information across modalities or leverage modal-specific strengths to compensate for indi vidual limitations. This e v aluation gap slows dev elopment of clinically viable systems capable of synthesizing di verse diagnostic information as human clinicians do. The techni- cal complexity of multimodal data management compounds these challenges. Institutions struggle with storage, versioning, and synchronization of large-scale heterogeneous datasets, while priv ac y regulations complicate cross-institutional data sharing. W ithout robust infrastructure and standard- ized curation protocols, the field remains fragmented, with isolated efforts failing to achie ve the critical mass needed for breakthrough advances in multimodal medical AI. 7.3 Challenges and Opportunities in Medical Foundation Models Medical foundation models demand unprecedented scale and diversity in training data, yet current resources remain insufficient for de v eloping truly generalizable systems [ 100 , 101 , 102 , 103 ]. The gap between a vailable data and foundation model requirements is particularly e vident in specialized domains such as pediatric imaging, rare diseases, and longitudinal treatment monitoring. Three interconnected challenges fundamentally constrain progress in this field. Scale and Representational Di versity . Be yond sheer quantity , foundation models require compre- hensiv e coverage across disease presentations, imaging protocols, clinical specialties, and patient 51 demographics to dev elop rob ust internal representations. Current medical datasets typically capture narrow slices of clinical reality , missing the long-tail distribution of rare conditions and atypical presentations that characterize real medical practice. This limitation is especially acute in underrep- resented populations and emerging disease v ariants. Licensing and Privacy Constraints. Unlike general-domain AI where datasets can be freely shared, medical data faces dual constraints from patient pri v acy regulations (e.g., HIP AA, GDPR) and institutional intellectual property policies. Even when foundation models can generate high- quality synthetic data for training augmentation [ 104 , 81 ], restricti ve licensing pre vents these en- hanced datasets from benefiting the broader research community . This regulatory landscape frag- ments the field, forcing redundant efforts across institutions and limiting collaborati ve progress [ 105 ]. Contextual and T emporal Intelligence. Effecti ve medical AI must transcend pattern recognition to understand clinical workflows, resource constraints, and patient-specific contexts [ 106 ]. For in- stance, models must distinguish between emergency protocols and routine screening, interpret how prior treatments influence current presentations, and track disease progression over time. Current training paradigms inadequately address these temporal reasoning and workflow integration capa- bilities essential for real-world deployment. Addressing these challenges requires coordinated ef forts to establish data gov ernance frame works that balance priv ac y protection with research adv ancement. W ithout systemic solutions—including federated learning infrastructures, standardized licensing models, and clinically-grounded e v alua- tion benchmarks—medical foundation models will remain confined to narro w applications rather than achieving the general intelligence needed for transformati v e clinical impact. 8 Conclusion This comprehensiv e survey of ov er 1,000 open-access medical image datasets rev eals a fragmented and imbalanced landscape that fundamentally constrains the de velopment of medical foundation models. Existing datasets remain predominantly small-scale, task-specific, and modality-restricted, with pronounced disparities across anatomical re gions and imaging modalities. These limitations reflect the field’ s incomplete transition from task-oriented to foundation-oriented data engineer- ing paradigms. T o address these challenges, we formulate the Metadata-Dri ven Fusion Paradigm (MDFP), a systematic framew ork for dataset integration that enables the construction of larger , more div erse training resources essential for foundation model de v elopment. Our analysis identifies three critical gaps: the scarcity of multimodal datasets that limits cross-modal reasoning capabilities, re- strictiv e licensing and priv acy regulations that fragment collaborative efforts, and the absence of contextual intelligence necessary for real-world clinical deployment. The dominance of segmenta- tion and classification tasks, alongside the underrepresentation of emerging applications like visual question answering and multimodal reasoning, underscores the urgent need for comprehensive data engineering strategies. Looking forward, advancing the development of medical foundation models requires a collectiv e shift to ward openness, ef ficiency , and inclusivity in data engineering. A key priority should be to encourage broader public release of medical imaging datasets, thereby enhancing transparency , re- producibility , and equitable access across institutions and regions. In parallel, research on synthetic data generation holds promise for mitigating pri v acy and data scarcity challenges, while annotation- efficient learning approaches can enable effecti ve use of partially labeled or weakly supervised data. Moreov er , the public release of foundation models trained on priv ate or institution-specific data, ev en when raw datasets cannot be shared, represents a practical pathway to democratize access to advanced medical AI capabilities. T ogether , these strategies constitute a sustainable and collabora- tiv e frame w ork for b uilding truly generalizable and clinically impactful medical foundation models. Acknowledgment W e sincerely thank all researchers, clinicians, institutions, and organizations who hav e contributed to the de velopment and public release of medical imaging datasets. Their dedicated ef forts in data col- lection, annotation, curation, and sharing ha ve laid the foundation for significant progress in medical AI. The open a v ailability of these resources has not only accelerated methodological innovation and 52 benchmark creation b ut also fostered collaboration across disciplines, enabling the broader commu- nity to explore new directions in multimodal learning, foundation model dev elopment, and clinical translation. W ithout their commitment to advancing science through openness and collaboration, this surve y and many of the achie vements in the field w ould not hav e been possible. References [1] Josh Achiam, Steven Adler , Sandhini Agarwal, Lama Ahmad, Ilge Akkaya, Florencia Leoni Aleman, Diogo Almeida, Jank o Altenschmidt, Sam Altman, Shyamal Anadkat, et al. Gpt-4 technical report. arXiv pr eprint arXiv:2303.08774 , 2023. 5 [2] Alec Radford, Jong W ook Kim, Chris Hallac y , Aditya Ramesh, Gabriel Goh, Sandhini Agar- wal, Girish Sastry , Amanda Askell, Pamela Mishkin, Jack Clark, Gretchen Krueger , and Ilya Sutske ver . Learning transferable visual models from natural language supervision, 2021. 5 [3] Y uhao Zhang, Hang Jiang, Y asuhide Miura, Christopher D Manning, and Curtis P Langlotz. Contrastiv e learning of medical visual representations from paired images and text. arXiv pr eprint arXiv:2010.00747 , 2020. 5 [4] Oriane Siméoni, Huy V . V o, Maximilian Seitzer , Federico Baldassarre, Maxime Oquab, Cijo Jose, V asil Khalidov , Marc Szafraniec, Seungeun Y i, Michaël Ramamonjisoa, Francisco Massa, Daniel Haziza, Luca W ehrstedt, Jian yuan W ang, T imothée Darcet, Théo Moutakanni, Leonel Sentana, Claire Roberts, Andrea V edaldi, Jamie T olan, John Brandt, Camille Couprie, Julien Mairal, Hervé Jégou, Patrick Labatut, and Piotr Bojanowski. DINOv3. arXiv pr eprint arXiv:2508.10104 , 2025. 5 [5] Alexander Kirillo v , Eric Mintun, Nikhila Ra vi, Hanzi Mao, Chloe Rolland, Laura Gustafson, T ete Xiao, Spencer Whitehead, Alexander C Berg, W an-Y en Lo, et al. Segment anything. arXiv pr eprint arXiv:2304.02643 , 2023. 5 [6] Andre w Sellergren, Sahar Kazemzadeh, T iam Jaroensri, Atilla Kiraly , Madeleine T ra verse, T imo K ohlber ger , Shawn Xu, Fayaz Jamil, Cían Hughes, Charles Lau, et al. Medgemma technical report. arXiv pr eprint arXiv:2507.05201 , 2025. 5 [7] Jun Ma, Y uting He, Feifei Li, Lin Han, Chenyu Y ou, and Bo W ang. Segment anything in medical images. Natur e Communications , 15(1):654, 2024. 5 , 15 [8] Richard J Chen, T ong Ding, Ming Y Lu, Drew FK W illiamson, Guillaume Jaume, Bowen Chen, Andrew Zhang, Daniel Shao, Andrew H Song, Muhammad Shaban, et al. T owards a general-purpose foundation model for computational pathology . Natur e Medicine , 2024. 5 [9] Ming Hu, Chenglong Ma, W ei Li, W anghan Xu, Jiamin W u, Jucheng Hu, T ianbin Li, Guo- hang Zhuang, Jiaqi Liu, Y ingzhou Lu, et al. A surve y of scientific lar ge language models: From data foundations to agent frontiers. arXiv pr eprint arXiv:2508.21148 , 2025. 5 [10] Zheyuan Zhang, T ianyi Ma, Zehong W ang, Y iyang Li, Shifu Hou, W eixiang Sun, Kaiwen Shi, Y ijun Ma, W ei Song, Ahmed Abbasi, et al. Llms4all: A re vie w on lar ge language models for research and applications in academic disciplines. arXiv pr eprint arXiv:2509.19580 , 2025. 5 [11] Michael Moor, Oishi Banerjee, Zahra Shakeri Hossein Abad, Harlan M Krumholz, Jure Leskov ec, Eric J T opol, and Pranav Rajpurkar . Foundation models for generalist medical artificial intelligence. Natur e , 616(7956):259–265, 2023. 5 , 51 [12] Ziyan Huang, Haoyu W ang, Zhongying Deng, Jin Y e, Y anzhou Su, Hui Sun, Junjun He, Y un Gu, Lixu Gu, Shaoting Zhang, et al. Stu-net: Scalable and transferable medical im- age segmentation models empowered by lar ge-scale supervised pre-training. arXiv pr eprint arXiv:2304.06716 , 2023. 5 [13] Haoyu W ang, Sizheng Guo, Jin Y e, Zhongying Deng, Junlong Cheng, T ianbin Li, Jianpin Chen, Y anzhou Su, Ziyan Huang, Y iqing Shen, et al. Sam-med3d: A vision foundation model for general-purpose se gmentation on volumetric medical images. IEEE T ransactions on Neural Networks and Learning Systems , 2025. 5 , 15 53 [14] Zhongying Deng, Haoyu W ang, Ziyan Huang, Lipei Zhang, Angelica I A viles-Ri v ero, Chaoyu Liu, Junjun He, Zoe K ourtzi, and Carola-Bibiane Schönlieb. Brain founda- tion models with hypergraph dynamic adapter for brain disease analysis. arXiv pr eprint arXiv:2505.00627 , 2025. 5 , 15 [15] Siyuan Y an, Zhen Y u, Clare Primiero, Cristina V ico-Alonso, Zhonghua W ang, Litao Y ang, Philipp Tschandl, Ming Hu, Lie Ju, Gin T an, et al. A multimodal vision foundation model for clinical dermatology . Natur e Medicine , pages 1–12, 2025. 5 [16] Nahid Ul Islam, DongAo Ma, Jiaxuan Pang, Shiv asakthi Senthil V elan, Michael Gotway , and Jianming Liang. Foundation x: integrating classification, localization, and segmentation through lock-release pretraining strategy for chest x-ray analysis. In 2025 IEEE/CVF W inter Confer ence on Applications of Computer V ision (W ACV) , pages 3647–3656. IEEE, 2025. 5 [17] Lei Bai, Zhongrui Cai, Y uhang Cao, Maosong Cao, W eihan Cao, Chiyu Chen, Haojiong Chen, Kai Chen, Pengcheng Chen, Y ing Chen, et al. Intern-s1: A scientific multimodal foundation model. arXiv pr eprint arXiv:2508.15763 , 2025. 5 [18] Alessandro Crimi and Spyridon Bakas. Brainlesion: Glioma, Multiple Scler osis, Str ok e and T raumatic Brain Injuries: 6th International W orkshop, BrainLes 2020, Held in Conjunction with MICCAI 2020, Lima, P eru, October 4, 2020, Revised Selected P apers, P art I , volume 12658. Springer Nature, 2021. 5 [19] Bjoern H Menze, Andras Jakab, Stef an Bauer , Jayashree Kalpathy-Cramer, Keyv an Farahani, Justin Kirby , Y uliya Burren, Nicole Porz, Johannes Slotboom, Roland Wiest, et al. The mul- timodal brain tumor image segmentation benchmark (brats). IEEE transactions on medical imaging , 34(10):1993–2024, 2014. 5 [20] Christoph Schuhmann, Romain Beaumont, Richard V encu, Cade Gordon, Ross Wightman, Mehdi Cherti, Theo Coombes, Aarush Katta, Clayton Mullis, Mitchell W ortsman, et al. Laion-5b: An open large-scale dataset for training next generation image-text models. Ad- vances in neural information pr ocessing systems , 35:25278–25294, 2022. 5 [21] Martin J W illemink, W ojciech A Koszek, Cailin Hardell, Jie W u, Dominik Fleischmann, Hugh Harv ey , Les R Folio, Ronald M Summers, Daniel L Rubin, and Matthe w P Lungren. Preparing medical imaging data for machine learning. Radiology , 295(1):4–15, 2020. 5 [22] Jin Y e, Junlong Cheng, Jianpin Chen, Zhongying Deng, T ianbin Li, Haoyu W ang, Y anzhou Su, Ziyan Huang, Jilong Chen, Lei Jiang, et al. Sa-med2d-20m dataset: Segment an ything in 2d medical imaging with 20 million masks. arXiv pr eprint arXiv:2311.11969 , 2023. 5 [23] Jin Y e, Guoan W ang, Y anjun Li, Zhongying Deng, W ei Li, T ianbin Li, Haodong Duan, Ziyan Huang, Y anzhou Su, Benyou W ang, et al. Gmai-mmbench: A comprehensive multimodal ev aluation benchmark to wards general medical ai. Advances in Neural Information Pr ocess- ing Systems , 37:94327–94427, 2024. 5 [24] Fatemeh Haghighi, Michael B Gotway , and Jianming Liang. Learning anatomy-disease en- tangled representation. In 2025 IEEE/CVF W inter Confer ence on Applications of Computer V ision (W A CV) , pages 4129–4141. IEEE, 2025. 5 [25] Marc Combalia, Noel Codella, V eronica Rotemberg, Cristina Carrera, Stephen Dusza, David Gutman, Brian Helba, Harald Kittler , Nicholas R K urtansky , Konstantinos Liopyris, et al. V alidation of artificial intelligence prediction models for skin cancer diagnosis using der- moscopy images: the 2019 international skin imaging collaboration grand challenge. The Lancet Digital Health , 4(5):e330–e339, 2022. 6 , 138 [26] Ziyan Huang, Zhongying Deng, Jin Y e, Haoyu W ang, Y anzhou Su, T ianbin Li, Hui Sun, Junlong Cheng, Jianpin Chen, Junjun He, Y un Gu, Shaoting Zhang, Lixu Gu, and Y u Qiao. A-ev al: A benchmark for cross-dataset and cross-modality ev aluation of abdominal multi- organ segmentation. Medical Image Analysis , 101:103499, 2025. ISSN 1361-8415. doi: https://doi.org/10.1016/j.media.2025.103499. 6 54 [27] Zhaodong W u, Qiaochu Zhao, Ming Hu, Y ulong Li, Haochen Xue, Zhengyong Jiang, An- gelos Stefanidis, Qiufeng W ang, Imran Razzak, Zongyuan Ge, et al. Mswal: 3d multi-class segmentation of whole abdominal lesions dataset. In International Confer ence on Medical Image Computing and Computer -Assisted Intervention , pages 378–388. Springer , 2025. 6 [28] Johann Li, Guangming Zhu, Cong Hua, Mingtao Feng, Basheer Bennamoun, Ping Li, Xi- aoyuan Lu, Juan Song, Peiyi Shen, Xu Xu, et al. A systematic collection of medical image datasets for deep learning. A CM Computing Surve ys , 2021. 6 [29] Saad M Khan, Xiaoxuan Liu, Siddharth Nath, Edward K orot, Li via Faes, Siegfried K W agner, Pearse A Keane, Neil J Sebire, Matthe w J Burton, and Alastair K Denniston. A global revie w of publicly av ailable datasets for ophthalmological imaging: barriers to access, usability , and generalisability . The Lancet Digital Health , 3(1):e51–e66, 2021. 6 [30] David W en, Saad M Khan, Antonio Ji Xu, Hussein Ibrahim, Luk e Smith, Jose Caballero, Luis Zepeda, Carlos de Blas Perez, Alastair K Denniston, Xiaoxuan Liu, et al. Characteristics of publicly av ailable skin cancer image datasets: a systematic revie w . The Lancet Digital Health , 4(1):e64–e74, 2022. 6 [31] Masoud T afa vvoghi, Lars Ailo Bongo, Nikita Shvetso v , Lill-T ov e Rasmussen Busund, and Kajsa Møllersen. Publicly av ailable datasets of breast histopathology h&e whole-slide im- ages: a scoping re vie w . Journal of P athology Informatics , 15:100363, 2024. 6 [32] Katharine A Dishner, Bala McRae-Posani, Arka Bho wmik, Maxine S Jochelson, Andrei Holodny , Katja Pinker , Sarah Eskreis-W inkler , and Joseph N Stember . A survey of publicly av ailable mri datasets for potential use in artificial intelligence research. J ournal of Ma gnetic Resonance Imaging , 59(2):450–480, 2024. 6 [33] Jakob W asserthal, Hanns-Christian Breit, Manfred T . Me yer , Maurice Pradella, Daniel Hinck, Alexander W . Sauter, T obias Heye, Daniel T . Boll, Joshy Cyriac, Shan Y ang, Michael Bach, and Martin Segeroth. T otalsegmentator: Robust segmentation of 104 anatomic structures in ct images. Radiology: Artificial Intelligence , 5(5):e230024, 2023. 6 [34] Chongyu Qu, Tiezheng Zhang, Hualin Qiao, Y ucheng T ang, Alan L Y uille, Zongwei Zhou, et al. Abdomenatlas-8k: Annotating 8,000 ct volumes for multi-organ segmentation in three weeks. Advances in Neural Information Pr ocessing Systems , 36:36620–36636, 2023. 6 , 10 [35] Kaiming He, Xiangyu Zhang, Shaoqing Ren, and Jian Sun. Deep residual learning for image recognition. In Pr oceedings of the IEEE confer ence on computer vision and pattern r eco gni- tion , pages 770–778, 2016. 10 [36] Ibrahim Ethem Hamamci, Sezgin Er , Furkan Almas, A yse Gulnihan Simsek, Sevv al Nil Esir - gun, Irem Dogan, Muhammed Furkan Dasdelen, Omer Faruk Durugol, Bastian Wittmann, T amaz Amiranashvili, Enis Simsar , Mehmet Simsar, Emine Bensu Erdemir , Abdullah Alan- bay , Anjany Sekubo yina, Berkan Lafci, Christian Bluethgen, Mehmet Kemal Ozdemir , and Bjoern Menze. De veloping generalist foundation models from a multimodal dataset for 3d computed tomography , 2024. URL . 10 , 30 , 143 [37] Aaron Grattafiori, Abhimanyu Dubey , Abhina v Jauhri, Abhinav Pande y , Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Ale x V aughan, et al. The llama 3 herd of models. arXiv pr eprint arXiv:2407.21783 , 2024. 10 [38] Andre w A Bork owski, Marilyn M Bui, L Brannon Thomas, Catherine P Wilson, Lauren A DeLand, and Stephen M Mastorides. Lung and colon cancer histopathological image dataset (lc25000). arXiv pr eprint arXiv:1912.12142 , 2019. 11 , 139 , 140 [39] Fabio A Spanhol, Luiz S Oliv eira, Caroline Petitjean, and Laurent Heutte. A dataset for breast cancer histopathological image classification. Ieee transactions on biomedical engineering , 63(7):1455–1462, 2015. 11 , 139 , 140 [40] Bastiaan S V eeling, Jasper Linmans, Jim W ink ens, T aco Cohen, and Max W elling. Rota- tion equiv ariant cnns for digital pathology . In International Conference on Medical image computing and computer-assisted intervention , pages 210–218. Springer , 2018. 11 , 139 55 [41] Jiancheng et al. Y ang. Medmnist v2-a large-scale lightweight benchmark for 2d and 3d biomedical image classification. Scientific Data , 10(1):41, 2023. 11 , 136 , 137 [42] Nima T ajbakhsh, Laura Jeyaseelan, Qian Li, Jeffre y N. Chiang, Zhihao W u, and Xiao wei Ding. Embracing imperfect datasets: A revie w of deep learning solutions for medical image segmentation. Medical Imag e Analysis , 63:101693, 2020. ISSN 1361-8415. doi: https: //doi.org/10.1016/j.media.2020.101693. 11 [43] Lauren H W illiams and Trafton Dre w . What do we kno w about volumetric medical image interpretation?: a revie w of the basic science and medical image perception literatures. Cog- nitive Resear ch: Principles and Implications , 4(1):21, 2019. 11 [44] Madan M. Rehani and Da vid Nacouzi. Higher patient doses through x-ray imaging proce- dures. Physica Medica , 79:80–86, Nov ember 2020. doi: 10.1016/j.ejmp.2020.10.017. 11 [45] Carlo Liguori, Giulia Frauenfelder , Carlo Massaroni, Paola Saccomandi, Francesco Giurazza, Francesca Pitocco, Riccardo Marano, and Emiliano Schena. Emerging clinical applications of computed tomography . Medical De vices: Evidence and Resear ch , 8:265–78, 06 2015. doi: 10.2147/MDER.S70630. 11 , 12 [46] Jessica Lohrke, Thomas Frenzel, Jan Endrikat, Filipe Caseiro Alves, Thomas M Grist, Meng Law , Jeong Min Lee, Tim Leiner , K un-Cheng Li, Konstantin Nikolaou, et al. 25 years of contrast-enhanced mri: dev elopments, current challenges and future perspectiv es. Advances in Therapy , 33(1):1–28, 2016. 12 [47] Shiying W ang, John A Hossack, and Ale xander L Klibanov . From anatomy to functional and molecular biomarker imaging and therap y: ultrasound is safe, ultrafast, portable, and inexpensi ve. In vestigative Radiology , 55(9):559–572, 2020. 12 [48] Gregory V erghese, Jochen K Lennerz, Danny Ruta, W en Ng, Selvam Thav araj, Kalliopi P Siziopikou, Threnesan Naidoo, Swapnil Rane, Roberto Salgado, Sarah E Pinder, et al. Computational pathology in cancer diagnosis, prognosis, and prediction–present day and prospects. The J ournal of P athology , 260(5):551–563, 2023. 12 [49] Niehls Kurnia wan and Martin Keuchel. Flexible gastro-intestinal endoscopy—clinical chal- lenges and technical achiev ements. Computational and Structural Biotec hnology Journal , 15:168–179, 2017. 12 [50] Nishtha P anwar , Philemon Huang, Jiaying Lee, Pearse A Keane, Tjin Swee Chuan, Ashutosh Richhariya, Stephen T eoh, T ock Han Lim, and Rupesh Agrawal. Fundus photography in the 21st century—a re vie w of recent technological adv ances and their implications for worldwide healthcare. T elemedicine and e-Health , 22(3):198–208, 2016. 12 [51] Harold Kittler , H Pehamberger , K W olff, and MJTIO Binder . Diagnostic accuracy of der- moscopy . The Lancet Oncology , 3(3):159–165, 2002. 12 [52] Chetan L. Srinidhi, Ozan Ciga, and Anne L. Martel. Deep neural network models for com- putational histopathology: A surve y . Medical Image Analysis , 67:101813, 2021. ISSN 1361- 8415. doi: https://doi.org/10.1016/j.media.2020.101813. 12 [53] David T ellez, Geert Litjens, Péter Bándi, W outer Bulten, John-Melle Bokhorst, Francesco Ciompi, and Jeroen van der Laak. Quantifying the ef fects of data augmentation and stain color normalization in conv olutional neural networks for computational pathology . Medical Image Analysis , 58:101544, 2019. ISSN 1361-8415. doi: https://doi.org/10.1016/j.media. 2019.101544. 12 [54] Hassan Al Hajj, Mathieu Lamard, Pierre-Henri Conze, Soumali Roycho wdhury , Xiao wei Hu, Gabija Maršalkait ˙ e, Odysseas Zisimopoulos, Muneer Ahmad Dedmari, Fenqiang Zhao, Jonas Prellber g, et al. Cataracts: Challenge on automatic tool annotation for cataract surgery . Medical image analysis , 52:24–41, 2019. 13 , 155 [55] Kristy K Brock, Sasa Mutic, T odd R McNutt, Hua Li, and Marc L Kessler . Use of image re g- istration and fusion algorithms and techniques in radiotherapy: Report of the aapm radiation therapy committee task group no. 132. Medical Physics , 44(7):e43–e76, 2017. 13 56 [56] Y ukun Zhou, Mark A Chia, Siegfried K W agner , Murat S A yhan, Dominic J W illiamson, Robbert R Struyv en, Timing Liu, Moucheng Xu, Mateo G Lozano, Peter W oodward-Court, et al. A foundation model for generalizable disease detection from retinal images. Nature , 622(7981):156–163, 2023. 15 [57] Y ilan W u, Bo Qian, Tingyao Li, Y iming Qin, Zhouyu Guan, T ingli Chen, Y ali Jia, Ping Zhang, Dian Zeng, Sayok o Moroi, et al. An eyecare foundation model for clinical assistance: a randomized controlled trial. Natur e Medicine , pages 1–10, 2025. 15 [58] Jinxi Xiang, Xiyue W ang, Xiaoming Zhang, Y inghua Xi, Feyisope Eweje, Y ijiang Chen, Y uchen Li, Colin Ber gstrom, Matthe w Gopaulchan, T ed Kim, et al. A vision–language foun- dation model for precision oncology . Natur e , 638(8051):769–778, 2025. 15 [59] Fernando Perez-Garcia, Harshita Sharma, Sam Bond-T aylor , Kenza Bouzid, V alentina Sal- vatelli, Maximilian Ilse, Shruthi Bannur , Daniel C Castro, Anton Schwaighofer , Matthew P Lungren, et al. Exploring scalable medical image encoders beyond text supervision. Nature Machine Intelligence , 7(1):119–130, 2025. 15 [60] DongAo Ma, Jiaxuan P ang, Michael B Gotway , and Jianming Liang. A fully open ai founda- tion model applied to chest radiography . Natur e , pages 1–11, 2025. 15 [61] Chenglong Ma, Y uanfeng Ji, Jin Y e, Zilong Li, Chenhui W ang, Junzhi Ning, W ei Li, Lihao Liu, Qiushan Guo, T ianbin Li, Junjun He, and Hongming Shan. Meditok: A unified tok enizer for medical image synthesis and interpretation. arXiv pr eprint arXiv:2505.19225 , 2025. 15 [62] Maya V arma, Ashwin K umar , Rogier van der Sluijs, Sophie Ostmeier, Louis Blankemeier , Pierre Chambon, Christian Bluethgen, Jip Prince, Curtis Langlotz, and Akshay Chaudhari. Medvae: Efficient automated interpretation of medical images with large-scale generalizable autoencoders. arXiv pr eprint arXiv:2502.14753 , 2025. 15 [63] Alistair EW Johnson, T om J Pollard, Seth J Berkowitz, Nathaniel R Greenbaum, Matthe w P Lungren, Chih-ying Deng, Roger G Mark, and Stev en Horng. Mimic-cxr, a de-identified publicly av ailable database of chest radiographs with free-text reports. Scientific data , 6(1): 317, 2019. 29 [64] Jeremy Irvin, Prana v Rajpurkar , Michael K o, Y ifan Y u, Silviana Ciurea-Ilcus, Chris Chute, Henrik Marklund, Behzad Haghgoo, Robyn Ball, Katie Shpanskaya, et al. Chexpert: A lar ge chest radiograph dataset with uncertainty labels and e xpert comparison. In Pr oceedings of the AAAI Confer ence on Artificial Intelligence , v olume 33, pages 590–597, 2019. 29 , 135 [65] Luc Soler , Ale xandre Hostettler , V incent Agnus, Arnaud Charnoz, Julien Fasquel, Johan Moreau, Anne Osswald, Mourad Bouhadjar , and Jacques Marescaux. 3D image reconstruc- tion for comparison of algorithm database: A patient specific anatomical and medical image database. IRCAD, Strasbour g, F r ance, T ech. Rep , 1(1), 2010. 30 , 148 [66] Fan Bai, Y uxin Du, Tiejun Huang, Max Q.-H. Meng, and Bo Zhao. M3d: Advanc- ing 3d medical image analysis with multi-modal large language models. arXiv pr eprint arXiv:2404.00578 , 2024. doi: 10.48550/arXiv .2404.00578. 30 , 143 [67] Jakob W asserthal, Hanns-Christian Breit, Manfred T . Me yer , Maurice Pradella, Daniel Hinck, Alexander W . Sauter, T obias Heye, Daniel T . Boll, Joshy Cyriac, Shan Y ang, Michael Bach, and Martin Segeroth. T otalsegmentator: Robust segmentation of 104 anatomic structures in ct images. Radiology: Artificial Intelligence , 5(5):e230024, 2023. doi: 10.1148/ryai.230024. 30 , 143 [68] Patrick Bilic, Patrick F Christ, Hongwei Li, Eugene V orontso v , A vi Ben-Cohen, Georgios Kaissis, Adi Szeskin, Colin Jacobs, Gabriel Efrain Humpire Mamani, Gabriel Chartrand, et al. The liv er tumor segmentation benchmark (lits). arXiv pr eprint arXiv:1901.04056 , 2019. 30 , 143 [69] National Lung Screening T rial Research T eam. Data from the national lung screen- ing trial (nlst) [data set]. The Cancer Imaging Archive, 2013. URL https://www. cancerimagingarchive.net/collection/nlst/ . 30 , 144 , 146 , 148 57 [70] Y uanfeng Ji, Haotian Bai, Chongjian Ge, Jie Y ang, Y e Zhu, Ruimao Zhang, Zhen Li, Lingyan Zhanng, W anling Ma, Xiang W an, et al. Amos: A large-scale abdominal multi-organ bench- mark for versatile medical image segmentation. Advances in neural information pr ocessing systems , 35:36722–36732, 2022. 30 , 143 [71] Adrienne M. Mendrik, Koen L. V incken, Hugo J. Kuijf, Marcel Breeuwer , W illem H. Bouvy , Jeroen de Bresser , Amir Alansary , Marleen de Bruijne, Aaron Carass, A yman El-Baz, Amod Jog, Ran veer Katyal, Ali R. Khan, Fedde v an der Lijn, Qaiser Mahmood, Ryan Mukher - jee, Annegreet v an Opbroek, Sahil P aneri, Sergio Pereira, Mikael Persson, Martin Rajchl, Duygu Sarikaya, Orjan Smedby , Carlos A. Silv a, Henri A. Vrooman, Saurabh Vyas, Chun- liang W ang, Liang Zhao, Geert Jan Biessels, and Max A. V ier ge ver . Mrbrains challenge: Online ev aluation frame work for brain image se gmentation in 3t mri scans. Computational Intelligence and Neur oscience , 2015:813696, 2015. doi: 10.1155/2015/813696. 32 , 151 [72] T assilo W ald, Constantin Ulrich, Jonathan Suprijadi, Sebastian Zie gler , Michal Nohel, Robin Peretzke, Gregor Köhler , and Klaus H. Maier-Hein. An openmind for 3d medical vision self-supervised learning. arXiv pr eprint arXiv:2412.17041 , 2024. doi: 10.48550/arXi v .2412. 17041. 32 , 152 [73] Bjoern H Menze, Andras Jakab, Stef an Bauer , Jayashree Kalpathy-Cramer, Keyv an Farahani, Justin Kirby , Y uliya Burren, Nicole Porz, Johannes Slotboom, Roland W iest, Le vente Lanczi, Elizabeth Gerstner , Marc-Andre W eber , T al Arbel, Brian B A vants, Nicholas A yache, Pa- tricia Buendia, D. Louis Collins, Nicolas Cordier , Jason J Corso, Antonio Criminisi, Tilak Das, Hervé Delingette, Cagatay Demiralp, Christopher R Durst, Michel Dojat, Senan Doyle, Joana Festa, Florence F orbes, Ezequiel Geremia, Ben Glocker , Polina Golland, Xiaotao Guo, Andac Hamamci, Khan M Iftekharuddin, Raj Jena, Nigel M John, Ender K onukoglu, Danial Lashkari, José Antonio Mariz, Raphael Meier, Sergio Pereira, Doina Precup, Stephen J Price, T ammy Riklin Ravi v , Syed M. S Reza, Michael Ryan, Duygu Sarikaya, Lawrence Schwartz, Hoo-Chang Shin, Jamie Shotton, Carlos A Silva, Nuno Sousa, Nagesh K Subbanna, Gábor Szekely , Thomas J T aylor, Owen M Thomas, Nicholas J T ustison, Gözde Unal, Flor V asseur , Max W intermark, Dong Hye Y e, Liang Zhao, Binsheng Zhao, Darko Zikic, Marcel Prasta wa, Mauricio Reyes, and K oen V an Leemput. The multimodal brain tumor image segmentation benchmark (brats). IEEE T ransactions on Medical Imaging , 34(10):1993–2024, 2015. doi: 10.1109/TMI.2014.2377694. 32 , 149 [74] Pamela J LaMontagne, T ammie L S Benzinger , John C Morris, Sarah K eefe, Russ Hornbeck, Chengjie Xiong, Elizabeth Grant, Jason Hassenstab, Krista Moulder , Andrei G Vlassenko, Marcus E Raichle, Carlos Cruchaga, and Daniel Marcus. Oasis-3: Longitudinal neuroimag- ing, clinical, and cognitiv e dataset for normal aging and Alzheimer disease. medRxiv , 2019. doi: 10.1101/2019.12.13.19014902. Preprint under CC-BY -ND 4.0 International. 32 , 148 , 154 [75] David C V an Essen, Stephen M Smith, Deanna M Barch, T imothy E J Behrens, Essa Y acoub, Kamil Ugurbil, and WU-Minn HCP Consortium. The wu-minn human connectome project: an o vervie w . Neur oImage , 80:62–79, 2013. doi: 10.1016/j.neuroimage.2013.05.041. 32 , 150 [76] Susanne G Mueller, Michael W W einer, Leon J Thal, Ronald C Petersen, Clifford R Jack, W illiam Jagust, John Q Trojano wski, Arthur W T oga, and Laurel Beckett. W ays to ward an early diagnosis in alzheimer’ s disease: The alzheimer’ s disease neuroimaging initiati ve (adni). Alzheimer’ s & Dementia , 1(1):55–66, 2005. doi: 10.1016/j.jalz.2005.06.003. 32 , 149 , 154 [77] Y iming Xiao, Hassan Ri v az, Matthieu Chabanas, Maryse Fortin, Ines Machado, Y angming Ou, Mattias P Heinrich, Julia A Schnabel, Xia Zhong, Andreas Maier , W olfgang W ein, Roozbeh Shams, Samuel Kadoury , Da vid Drobn y , Marc Modat, and Ingerid Reinertsen. Evaluation of mri to ultrasound registration methods for brain shift correction: The curi- ous2018 challenge. IEEE T ransactions on Medical Imaging , 39(3):777–786, 2019. doi: 10.1109/TMI.2019.2935060. 33 , 148 , 153 [78] Jonas Bernal et al. Gastrointestinal image analysis (giana) challenge: Endovis sub- challenge on polyp detection, localization, and segmentation. https://endovissub2017- roboticinstrumentsegmentation.grand-challenge.or g/Data/, 2017. 43 58 [79] G. Hattab et al. Kidney edge detection in laparoscopic image data for computer-assisted surgery . BMC Medical Imaging , 21(1):119, 2021. doi: 10.1186/s12880- 021- 00650- z. 43 , 156 [80] M. Maška et al. The cell tracking challenge: 10 years of objectiv e benchmarking. Nature Methods , 20:1010–1020, 2023. doi: 10.1038/s41592- 023- 01879- y. 43 [81] W ei Li, Ming Hu, Guoan W ang, Lihao Liu, Kaijing Zhou, Junzhi Ning, Xin Guo, Zongyuan Ge, Lixu Gu, and Junjun He. Ophora: A large-scale data-driven text-guided ophthalmic surgical video generation model. arXiv preprint , 2025. 43 , 52 , 157 [82] K yungmo Kim, Kyoungb un Lee, Sungduk Cho, Dong Un Kang, Seongkeun P ark, Y unsook Kang, Hyunjeong Kim, Gheeyoung Choe, K yung Chul Moon, Kyu Sang Lee, et al. P aip 2020: Microsatellite instability prediction in colorectal cancer . Medical Image Analysis , 89: 102886, 2023. [83] Hanna Borgli, V ajira Thambawita, Pia H Smedsrud, Stev en Hicks, Debesh Jha, Sigrun L Eskeland, Kristin Ranheim Randel, Konstantin Pogorelov , Mathias Lux, Duc T ien Dang Nguyen, et al. Hyperkvasir , a comprehensiv e multi-class image and video dataset for gas- trointestinal endoscopy . Scientific data , 7(1):283, 2020. 43 , 142 , 157 [84] Emmett D Goodman, Krishna K P atel, Y ilun Zhang, William Locke, Chris J K ennedy , Rohan Mehrotra, Stephen Ren, Melody Guan, Orr Zohar , Maren Do wning, et al. Analyzing sur gical technique in di verse open surgical videos with multitask machine learning. J AMA sur g ery , 159(2):185–192, 2024. 43 , 156 [85] Ming Hu, Zhengdi Y u, Feilong T ang, Kaiwen Chen, Y ulong Li, Imran Razzak, Junjun He, T olga Birdal, Kaijing Zhou, and Zongyuan Ge. T owards dynamic 3d reconstruction of hand- instrument interaction in ophthalmic surgery . arXiv pr eprint arXiv:2505.17677 , 2025. 43 [86] Jordi Bernal et al. Gastrointestinal image analysis (giana) challenge dataset. MICCAI En- doV is Challenge, 2018. A vailable at https://endovissub2017-giana.grand-challenge.or g/. 43 [87] Sharib Ali, Felix Zhou, Christian Daul, Barbara Braden, Adam Bailey , Stefano Realdon, James East, Georges W agnieres, V ictor Loschenov , Enrico Grisan, et al. Endoscopy artifact detection (ead 2019) challenge dataset. arXiv pr eprint arXiv:1905.03209 , 2019. 43 , 142 [88] V ivek Singh Bawa, Gurkirt Singh, Francis KapingA, Inna Skarga-Banduro va, Elettra Oleari, Alice Leporini, Carmela Landolfo, Pengfei Zhao, Xi Xiang, Gongning Luo, et al. The saras endoscopic surgeon action detection (esad) dataset: Challenges and methods. arXiv preprint arXiv:2104.03178 , 2021. 43 , 142 , 157 [89] Siyuan Y an, Xieji Li, Ming Hu, Y iwen Jiang, Zhen Y u, and Zongyuan Ge. Make: Multi- aspect kno wledge-enhanced vision-language pretraining for zero-shot dermatological assess- ment. In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , pages 369–379. Springer , 2025. 43 [90] Aneeq Zia, Max Berniker , Rogerio Nespolo, Conor Perreault, Ziheng W ang, Ben- jamin Mueller , et al. Surgical visual understanding (surgvu) dataset. arXiv pr eprint arXiv:2501.09209 , 2025. Dataset of hundreds of hours of robotic surgical training videos with tool presence and task labels. 43 , 156 [91] Safa Ali et al. Preoperativ e-to-intraoperati ve laparoscopy fusion (p2ilf) challenge: Dataset and tasks. https://p2ilf.grand-challenge.or g/, 2022. Accessed 2024-05-13. 43 , 156 [92] Dimitrios Psychogyios, Emanuele Colleoni, Beatrice V an Amsterdam, Chih-Y ang Li, Shu- Y u Huang, Y uchong Li, Fucang Jia, Baosheng Zou, Guotai W ang, Y ang Liu, et al. Sar- rarp50: Segmentation of sur gical instrumentation and action recognition on robot-assisted radical prostatectomy challenge. arXiv pr eprint arXiv:2401.00496 , 2023. 43 [93] Negin Ghamsarian, Y osuf El-Shabrawi, Sahar Nasirihaghighi, Doris Putzgruber -Adamitsch, Martin Zinkernagel, Sebastian W olf, Klaus Schoeffmann, and Raphael Sznitman. Cataract- 1k: Cataract sur gery dataset for scene segmentation, phase recognition, and irre gularity de- tection. arXiv pr eprint arXiv:2312.06295 , 2023. 43 , 156 59 [94] Haav ard Bor gli, V ajira Thamba wita, Pia H. Smedsrud, Stev en Hicks, et al. Hyperkvasir , a comprehensiv e multi-class image and video dataset for gastrointestinal endoscopy . Scientific Data , 7(1):283, 2020. doi: 10.1038/s41597- 020- 00622- y. 43 [95] Oli vier Bodenreider . The unified medical language system (umls): integrating biomedical terminology . Nucleic acids r esear ch , 32(suppl_1):D267–D270, 2004. 47 [96] Kai Zhang, Rong Zhou, Eashan Adhikarla, Zhiling Y an, Y ixin Liu, Jun Y u, Zhengliang Liu, Xun Chen, Brian D Davison, Hui Ren, et al. A generalist vision–language foundation model for div erse biomedical tasks. Natur e Medicine , 30(11):3129–3141, 2024. 51 [97] Xun Zhu, Fanbin Mo, Zheng Zhang, Jiaxi W ang, Y iming Shi, Ming W u, Chuang Zhang, Miao Li, and Ji W u. Enhancing multi-task learning capability of medical generalist foundation model via image-centric multi-annotation data. arXiv pr eprint arXiv:2504.09967 , 2025. 51 [98] Shih-Cheng Huang, Malte Jensen, Serena Y eung-Le vy , Matthe w P Lungren, Hoifung Poon, and Akshay S Chaudhari. Multimodal foundation models for medical imaging-a systematic revie w and implementation guidelines. medRxiv , pages 2024–10, 2024. 51 [99] Shujian Gao, Y uan W ang, and Zekuan Y u. Barl: Bilateral alignment in representation and label spaces for semi-supervised volumetric medical image segmentation, 2025. URL https: //arxiv.org/abs/2510.16863 . 51 [100] Raphael Schäfer, Till Nicke, Henning Höfener , Annkristin Lange, Dorit Merhof, Friedrich Feuerhake, V olkmar Schulz, Johannes Lotz, and Fabian Kiessling. Overcoming data scarcity in biomedical imaging with a foundational multi-task model. Natur e Computational Science , 4(7):495–509, 2024. 51 [101] T ianbin Li, Y anzhou Su, W ei Li, Bin Fu, Zhe Chen, Ziyan Huang, Guoan W ang, Cheng- long Ma, Y ing Chen, Ming Hu, et al. Gmai-vl & gmai-vl-5.5 m: A large vision-language model and a comprehensi ve multimodal dataset to wards general medical ai. arXiv pr eprint arXiv:2411.14522 , 2024. 51 [102] Y anzhou Su, T ianbin Li, Jiyao Liu, Chenglong Ma, Junzhi Ning, Cheng T ang, Sibo Ju, Jin Y e, Pengcheng Chen, Ming Hu, et al. Gmai-vl-r1: Harnessing reinforcement learning for multimodal medical reasoning. arXiv pr eprint arXiv:2504.01886 , 2025. 51 [103] Y uan W ang, Jiaxiang Liu, Shujian Gao, Bin Feng, Zhihang T ang, Xiaotang Gai, Jian Wu, and Zuozhu Liu. V2t-cot: From vision to te xt chain-of-thought for medical reasoning and diagnosis. In International Confer ence on Medical Image Computing and Computer -Assisted Intervention , pages 658–668. Springer , 2025. 51 [104] Ming Hu, Siyuan Y an, Peng Xia, Feilong T ang, W enxue Li, Peibo Duan, Lin Zhang, and Zongyuan Ge. Dif fusion model dri ven test-time image adaptation for robust skin lesion clas- sification. arXiv pr eprint arXiv:2405.11289 , 2024. 52 [105] Richard Sidebottom, Iain L yburn, Michael Brady , and Sarah V innicombe. Fair shares: b uild- ing and benefiting from healthcare ai with mutually beneficial structures and de v elopment partnerships. British J ournal of Cancer , 125(9):1181–1184, 2021. 52 [106] Michelle M Li, Ben Y Reis, Adam Rodman, Tianxi Cai, Noa Dagan, Ran D Balicer , Joseph Loscalzo, Isaac S K ohane, and Marinka Zitnik. One patient, many contexts: Scaling medical ai through contextual intelligence. arXiv preprint , 2025. 52 [107] Johannes Leuschner , Maximilian Schmidt, Daniel Otero Baguer , and Peter Maass. LoDoP aB- CT, a benchmark dataset for low-dose computed tomography reconstruction. Scientific Data , 8:109, 2021. doi: 10.1038/s41597- 021- 00893- z. URL https://www.nature.com/ articles/s41597- 021- 00893- z . 132 [108] Darshan D. Ruikar , K. C. Santosh, Ra vindra S. He gadi, Lakhan Rupnar, and V i vek A. Choud- hary . 5k+ ct images on fractured limbs: A dataset for medical imaging research. Journal of Medical Systems , 45(4):51, 2021. doi: 10.1007/s10916- 021- 01724- 9. 132 60 [109] P . Ehrlich, Y . Y . Chi, M. M. Chintagumpala, F . A. Hoffer , E. J. Perlman, J. A. Kalapurakal, A. W arwick, R. C. Shamberger , G. Khanna, T . E. Hamilton, K. W . Gow , A. C. Paulino, E. J. Gratias, E. A. Mullen, J. I. Geller, P . E. Grundy , C. V . Fernandez, M. L. Ritchey , and J. S. Dome. Combination chemotherapy and surgery in treating young patients with wilms tumor (aren0534) [data set]. https://doi.org/10.7937/TCIA.5M9S- 6Y97 , 2021. The Cancer Imaging Archiv e (TCIA); DOI: 10.7937/TCIA.5M9S-6Y97. Accessed 2025-08-21. 132 , 133 , 134 [110] Marc K ohli, James J. Morrison, Judy W awira, Matthew B. Morgan, Jason Hostetter , Brad Genereaux, Mohannad Hussain, and Ste ve G. Langer . Creation and curation of the society of imaging informatics in medicine hackathon dataset. Journal of Digital Imaging , 31(1):9–12, 2018. doi: 10.1007/s10278- 017- 0003- 5. 132 [111] National Lung Screening Trial Research T eam. Data from the national lung screening trial (nlst). The Cancer Imaging Archive (TCIA), https://doi.org/10.7937/TCIA.HMQ8-J677, 2013. Accessed: 2025-08-22. 132 [112] Adam E. Flanders, Luciano M. Prev edello, George Shih, Safwan S. Halabi, Jayashree Kalpathy-Cramer , Robyn Ball, John T . Mongan, Anouk Stein, Felipe C. Kitamura, Matthew P . Lungren, Gagandeep Choudhary , Lesley Cala, Luiz Coelho, Monique Mogensen, Fann y Morón, Elka Miller , Ichiro Ikuta, V ahe Zohrabian, Olivia McDonnell, Christie Lincoln, Lub- dha Shah, David Joyner , Amit Agarw al, Ryan K. Lee, Jaya Nath, and RSNA-ASNR 2019 Brain Hemorrhage CT Annotators. Construction of a machine learning dataset through col- laboration: The rsna 2019 brain ct hemorrhage challenge. Radiology: Artificial Intelligence , 2(3):e190211, 2020. doi: 10.1148/ryai.2020190211. URL https://doi.org/10.1148/ ryai.2020190211 . 132 [113] Xingyi Y ang, Xuehai He, Jinyu Zhao, Y ichen Zhang, Shanghang Zhang, and Pengtao Xie. Covid-ct-dataset: A ct scan dataset about covid-19. arXiv preprint , 2020. doi: 10.48550/arXi v .2003.13865. URL . 132 [114] wjXiaoChuangw. Co vid-19-ct scan images. https://tianchi.aliyun.com/dataset/dataDetail?dataId=93666, 2021. Dataset; accessed 2025-08-22. 132 [115] SunneY i. Chest ct-scan images dataset. https://tianchi.aliyun.com/dataset/93929, 2025. V er- sion v1. 132 [116] Murtadha D. Hssayeni, Muayad S. Croock, A ymen D. Salman, Hassan Falah Al-khafaji, Zakaria A. Y ahya, and Behnaz Ghoraani. Intracranial hemorrhage se gmentation using a deep con v olutional model. Data , 5(1):14, 2020. doi: 10.3390/data5010014. URL https://doi. org/10.3390/data5010014 . 132 [117] Eduardo Soares, Plamen Angelov , Sarah Biaso, Michele Higa Froes, and Daniel Kanda Abe. Sars-cov-2 ct-scan dataset: A lar ge dataset of real patients ct scans for sars-cov-2 identifi- cation. medRxiv , 2020. doi: 10.1101/2020.04.24.20078584. URL https://doi.org/10. 1101/2020.04.24.20078584 . 132 [118] Jiancheng Y ang, Rui Shi, and Bingbing Ni. Medmnist classification decathlon: A lightweight automl benchmark for medical image analysis. In IEEE 18th International Symposium on Biomedical Ima ging (ISBI) , pages 191–195, 2021. doi: 10.1109/ISBI48211.2021.9433967. 132 , 135 , 136 [119] National Library of Medicine. The visible human project. https://www.nlm.nih.gov/ research/visible/visible_human.html , 1994. The creation of complete, anatomically detailed, three-dimensional representations of the normal male and female human bodies. Accessed 2025-08-22. 132 , 133 [120] Alba García Seco de Herrera, Roger Schaer , Stefano Bromuri, and Henning Müller . Overvie w of the imageclef 2016 medical task. In CLEF 2016 W orking Notes , v olume 1609, pages 219– 232. CEUR-WS.org, 2016. URL http://ceur- ws.org/Vol- 1609/16090219.pdf . 132 , 133 , 134 , 135 , 138 , 142 61 [121] Xueyan Mei, Zelong Liu, Philip M Robson, Brett Marinelli, Mingqian Huang, Amish Doshi, Adam Jacobi, Chendi Cao, Katherine E Link, Thomas Y ang, et al. Radimagenet: an open radiologic deep learning research dataset for ef fecti ve transfer learning. Radiology: Artificial Intelligence , 4(5):e210315, 2022. 132 , 133 , 134 [122] Cancer Moonshot Biobank. Cancer moonshot biobank – colorectal cancer collection (cmb- crc). The Cancer Imaging Archiv e, https://doi.or g/10.7937/DJG7-GZ87, 2022. URL https: //doi.org/10.7937/DJG7- GZ87 . Dataset. 132 , 133 , 139 [123] Cancer Moonshot Biobank. Cancer moonshot biobank – gastroesophageal cancer collection (cmb-gec). The Cancer Imaging Archive (TCIA), https://doi.or g/10.7937/E7KH-R486, 2022. V ersion 6 [dataset]. 132 , 133 , 139 [124] Cancer Moonshot Biobank. Cancer moonshot biobank - melanoma collection (cmb-mel). https://doi.org/10.7937/GWSP-WH72, 2022. Data set. 132 , 133 , 139 [125] Cancer Moonshot Biobank. Cancer moonshot biobank – multiple myeloma collection (cmb- mml). https://www .cancerimagingarchi ve.net/collection/cmb-mml/, 2022. Dataset. 132 , 133 , 139 [126] Cancer Moonshot Biobank. Cancer moonshot biobank – prostate cancer collection (cmb- pca). https://www .cancerimagingarchive.net/collection/cmb-pca/, 2022. V ersion 9. 132 , 133 , 139 [127] National Cancer Institute Clinical Proteomic T umor Analysis Consortium (CPT A C). The clinical proteomic tumor analysis consortium lung squamous cell carcinoma collection (cptac-lscc) (version 15). The Cancer Imaging Archiv e, 2018. URL https://www. cancerimagingarchive.net/collection/cptac- lscc/ . [Data set]. 132 , 133 , 145 , 154 [128] K evin S. Mader . Finding and measuring lungs in ct data. Kaggle, URL: https://www .kaggle.com/datasets/kmader/finding-lungs-in-ct-data, 2017. Accessed: 2025- 08-22. 132 [129] Felipe Campos Kitamura. Head ct - hemorrhage. https://www.kaggle.com/datasets/ felipekitamura/head- ct- hemorrhage , 2019. Accessed: 2025-08-22. 132 [130] W onkyeong Lee, Fabian W agner, Adrian Galdran, Y ongyi Shi, W enjun Xia, Ge W ang, Xu- anqin Mou, Md Atik Ahamed, Abdullah Al Zubaer Imran, Ji Eun Oh, Kyungsang Kim, Jong T ak Baek, Dongheon Lee, Boohwi Hong, Philip T empelman, Donghang L yu, Adrian Kuiper , Lars van Blokland, Maria Baldeon Calisto, Scott Hsieh, Minah Han, Jongduk Baek, Andreas Maier , Adam W ang, Garry Evan Gold, and Jang-Hwan Choi. Low-dose computed tomography perceptual image quality assessment. Medical Image Analysis , 99:103343, 2025. doi: 10.1016/j.media.2024.103343. 132 [131] Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Research Network. Applied proteogenomics organizational learning and outcomes (apollo-5). https://wiki.cancerimagingarchi ve.net/display/Public/APOLLO-5, 2023. Limited access; ac- cessed 2025-08-22. 132 , 133 [132] Mirabela Rusu, Prabhakar Rajiah, Robert Gilk eson, Ming Y ang, Christopher Donatelli, Rahul Thawani, Frank J. Jacono, Patrick Linden, and Anant Madabhushi. Co-registration of pre- operativ e ct with e x vi v o sur gically excised ground glass nodules to define spatial extent of in v asi ve adenocarcinoma on in viv o imaging: a proof-of-concept study . Eur opean Radiology , 27(10):4209–4217, 2017. doi: 10.1007/s00330- 017- 4813- 0. 132 , 140 [133] Cancer Moonshot Biobank. Cancer moonshot biobank – lung cancer collection (cmb- lca). The Cancer Imaging Archive (TCIA), https://doi.org/10.7937/3CX3-S132, 2022. URL https://doi.org/10.7937/3CX3- S132 . Dataset. 132 [134] P . Muzi, M. W anner , and P . Kinahan. Data from rider lung pet-ct. The Cancer Imaging Archiv e, 2015. URL https://doi.org/10.7937/k9/tcia.2015.ofip7tvm . 132 , 133 , 146 , 154 62 [135] K. M. Kelly , P . D. Cole, Q. Pei, R. Bush, K. B. Roberts, D. C. Hodgson, K. M. McCarten, S. Y . Cho, and C. Schw artz. Combination chemotherap y and radiation therapy in treating young patients with newly diagnosed hodgkin lymphoma (ahod0831) (v ersion 1) [data set]. https://www.cancerimagingarchive.net/collection/ahod0831/ , 2022. The Can- cer Imaging Archiv e (TCIA); V ersion 1; Accessed 2025-08-21. 132 , 133 , 135 [136] Peter Choyke, Baris Turkbe y , Peter Pinto, Maria Merino, and Bradford W ood. Data from prostate-mri. The Cancer Imaging Archive, 2016. URL https://doi.org/10.7937/K9/ TCIA.2016.6046GUDv . 132 [137] C. V . Fernandez, E. A. Mullen, Y .-Y . Chi, P . F . Ehrlich, E. J. Perlman, J. A. Kalapurakal, G. Khanna, A. C. Paulino, T . E. Hamilton, K. W . Gow , Z. T ochner, F . A. Hof fer , J. S. W ithycombe, R. C. Shamberger , Y . Kim, J. I. Geller , J. R. Anderson, P . E. Grundy , and J. S. Dome. V incristine, dactinomycin, and doxorubicin with or without radiation ther- apy or observation only in treating younger patients who are undergoing surgery for ne wly diagnosed stage i, stage ii, or stage iii wilms’ tumor (aren0532) (version 1) [data set]. https://doi.org/10.7937/6PJ1- M859 , 2022. The Cancer Imaging Archiv e (TCIA); V ersion 1; DOI: 10.7937/6PJ1-M859; Accessed 2025-08-21. 132 , 133 , 134 [138] Carole H Müller , Carole Gonzalez, Katharina Breininger , Shadi Albarqouni, Emma W achter, Palla vi Agraw al, Dominik Auer, Marius Erdt, Hongwei Chen, Doreen Miranda, et al. The qubiq challenge: quantifying uncertainty in biomedical image segmentation. In Uncertainty for Safe Utilization of Machine Learning in Medical Imaging and Clinical Image-Based Pr o- cedur es , pages 59–70. Springer , 2020. 132 , 133 [139] Hongwei Bran Li, Fernando Na varro, Ivan Ezhov , Amirhossein Bayat, Dhritiman Das, Flo- rian K ofler , Suprosanna Shit, Diana W aldmannstetter, Johannes C P aetzold, Xiaobin Hu, et al. Qubiq: Uncertainty quantification for biomedical image segmentation challenge. arXiv pr eprint arXiv:2405.18435 , 2024. 132 , 133 [140] Jukka Hirv asniemi, Jos Runhaar , Rianne A van der Heijden, Maryam Zokaeinikoo, Mingrui Y ang, Xiaojuan Li, Jimin T an, Haresh Rengaraj Rajamohan, Y uyue Zhou, Cem M Deniz, et al. The knee osteoarthritis prediction (knoap2020) challenge: An image analysis challenge to predict incident symptomatic radiographic knee osteoarthritis from mri and x-ray images. Osteoarthritis and Cartilage , 31(1):115–125, 2023. 133 , 135 [141] Alibaba Tianchi. braimmri - brain mri se gmentation dataset. https://tianchi.aliyun. com/dataset/dataDetail?dataId=127459 , 2022. Brain tumor MRI segmentation dataset with 110 images. License: CC BY -NC-SA. Accessed 2025-08-22. 133 [142] Alibaba T ianchi. Brain-mri - brain disease mri segmentation dataset. https://tianchi. aliyun.com/dataset/127583 , 2020. Brain disease MRI segmentation dataset using FLAIR sequences with 110 images. License: CC BY -NC-SA. Accessed 2025-08-22. 133 [143] Alibaba Tianchi. Spinaldisease2020 - spinal disease mri detection dataset. https:// tianchi.aliyun.com/competition/entrance/531796/information , 2020. Spinal disease detection dataset using T1 and T2 MRI sequences with 150 images. License: CC BY -NC-SA. Accessed 2025-08-22. 133 [144] Alba G Seco De Herrera, Stefano Bromuri, Roger Schaer , and Henning Müller . Ov erview of the medical tasks in imageclef 2016. CLEF working notes. Evora, P ortugal , 2016. 133 , 138 , 139 [145] Cancer Moonshot Biobank. Cancer moonshot biobank – colorectal cancer collection (cmb- crc) (version 8). https://www.cancerimagingarchive.net/collection/cmb- crc/ , 2022. DOI:10.7937/djg7-gz87; accessed 2025-08-21. 133 , 134 , 135 [146] Cancer Moonshot Biobank. Cancer moonshot biobank – lung cancer collection (cmb-lca) (version 9). https://www.cancerimagingarchive.net/collection/cmb- lca/ , 2025. DOI:10.7937/3CX3-S132; CC BY 4.0; accessed 2025-08-21. 133 , 134 , 135 63 [147] Anant Madabhushi and Michael D. Feldman. Fused Radiology-Pathology Prostate Dataset (Prostate Fused-MRI-Pathology). The Cancer Imaging Archi ve, 2016. URL https://www. cancerimagingarchive.net/collection/prostate- fused- mri- pathology/ . 133 , 139 [148] HeyWhale. Cardiac atrial images - cardiac mri segmentation dataset. https://www. heywhale.com/mw/dataset/5e4de9618ee624002d4c4117 , 2020. Cardiac atrial MRI segmentation dataset with 8,000 images for cardiac disease analysis. License: CC BY 4.0. Accessed 2025-08-22. 133 [149] The Cancer Imaging Archiv e (TCIA). Apollo-5-da-rad. https://www. cancerimagingarchive.net/tcia- downloads/apollo- 5- da- rad/ , 2025. Ac- cessed 2025-08-21; ISSN: 2474-4638; TCIA Site License (CC BY -NC-ND). 133 , 134 , 135 [150] Prostate-mri, 2011. URL https://wiki.cancerimagingarchive.net/display/ Public/PROSTATE- MRI . 133 , 140 [151] Alba Garcia Seco De Herrera, Henning Müller , and Stefano Bromuri. Overview of the im- ageclef 2015 medical classification task. In W orking Notes of CLEF 2015–Cr oss Language Evaluation F orum, CEUR , volume 1391. CEUR W orkshop Proceedings, 2015. 133 , 134 [152] Thomas LA van den Heuvel, Dagmar de Bruijn, Chris L de Korte, and Bram van Ginneken. Automated measurement of fetal head circumference using 2d ultrasound images. PloS one , 13(8):e0200412, 2018. 134 [153] W alid Al-Dhabyani, Mohammed Gomaa, Hussien Khaled, and Aly Fahmy . Dataset of breast ultrasound images. Data in Brief , 28:104863, 2020. 134 [154] V aleria De Luca, T obias Benz, Satoshi K ondo, Lars König, D Lübke, Sven Rothlübbers, Oudom Somphone, Stéphane Allaire, MA Lediju Bell, D YF Chung, et al. The 2014 liver ultrasound tracking benchmark. Physics in Medicine & Biology , 60(14):5571, 2015. 134 , 157 [155] Anna Montoya, Hasnin, kaggle446, shirzad, W ill Cukierski, and yffud. Ultrasound nerv e seg- mentation. https://kaggle.com/competitions/ultrasound- nerve- segmentation , 2016. Kaggle. 134 [156] Jianqiao Zhou, Xiaohong Jia, Dong Ni, Alison Noble, Ruobing Huang, T ao T an, and Manh The V an. Thyroid nodule segmentation and classification in ultrasound i mages, March 2020. URL https://doi.org/10.5281/zenodo.3715942 . 134 [157] Alba García Seco de Herrera, Roger Schaer , Stefano Bromuri, and Henning Müller . Overvie w of the ImageCLEF 2016 medical task. In W orking Notes of CLEF 2016 (Cr oss Language Evaluation F orum) , September 2016. 134 , 141 [158] Cancer Moonshot Biobank. Cancer moonshot biobank – melanoma collection (cmb-mel) (version 9). https://www.cancerimagingarchive.net/collection/cmb- mel/ , 2022. DOI:10.7937/gwsp-wh72; accessed 2025-08-21. 134 [159] Y aosheng Lu, Mengqiang Zhou, Dengjiang Zhi, Minghong Zhou, Xiaosong Jiang, Ruiyu Qiu, Zhanhong Ou, Huijin W ang, Di Qiu, Mei Zhong, Xiaoxing Lu, Gaowen Chen, and Jieyun Bai. The jnu-ifm dataset for segmenting pubic symphysis-fetal head. Data in Brief , 41:107904, 2022. ISSN 2352-3409. doi: https://doi.org/10.1016/j.dib .2022.107904. URL https:// www.sciencedirect.com/science/article/pii/S2352340922001160 . 134 [160] Y i Guo, Shichong Zhou, Jun Shi, and Y uanyuan W ang. Ultrasound image enhancement challenge 2023, April 2023. URL https://doi.org/10.5281/zenodo.7841250 . 134 [161] Haifan Gong, Guanqi Chen, Ranran W ang, Xiang Xie, Mingzhi Mao, Y izhou Y u, Fei Chen, and Guanbin Li. Multi-task learning for thyroid nodule segmentation with thyroid region prior . In 2021 IEEE 18th International Symposium on Biomedical Imaging (ISBI) , pages 257–261, 2021. doi: 10.1109/ISBI48211.2021.9434087. 134 64 [162] Sarah Leclerc, Erik Smistad, João Pedrosa, Andreas Østvik, Frederic Cervenansky , Florian Espinosa, T orvald Espeland, Erik Andreas Rye Berg, Pierre-Marc Jodoin, Thomas Grenier, Carole Lartizien, Jan D’hooge, Lasse Lovstakken, and Olivier Bernard. Deep learning for segmentation using an open large-scale dataset in 2d echocardiography . IEEE T r ansactions on Medical Imaging , 38(9):2198–2210, 2019. doi: 10.1109/TMI.2019.2900516. 134 [163] Haifan Gong, Guanqi Chen, Ranran W ang, Xiang Xie, Mingzhi Mao, Y izhou Y u, Fei Chen, and Guanbin Li. Multi-task learning for thyroid nodule segmentation with thyroid region prior . In 2021 IEEE 18th International Symposium on Biomedical Imaging (ISBI) , pages 257–261, 2021. doi: 10.1109/ISBI48211.2021.9434087. 134 [164] Moi Hoon Y ap, Gerard Pons, Joan Marti, Sergi Ganau, Melcior Sentis, Re yer Zwiggelaar , Adrian K Da vison, and Robert Marti. Automated breast ultrasound lesions detection using con v olutional neural networks. IEEE journal of biomedical and health informatics , 22(4): 1218–1226, 2017. 134 [165] Hanna Piotrzk owska-Wróble wska, Katarzyna Dobruch-Sobczak, Michał Byra, and Andrzej Nowicki. Open access database of raw ultrasonic signals acquired from malignant and benign breast lesions. Medical physics , 44(11):6105–6109, 2017. 134 [166] Anna Pa wło wska, Anna ´ Cwierz-Pie ´ nko wska, Agnieszka Domalik, Dominika Jagu ´ s, Piotr Kasprzak, Rafał Matko wski, Łukasz Fura, Andrzej No wicki, and Norbert ˙ Zołek. Curated benchmark dataset for ultrasound based breast lesion analysis. Scientific Data , 11(1):148, 2024. 134 [167] Daniel S Kerman y , Michael Goldbaum, W enjia Cai, Carolina CS V alentim, Huiying Liang, Sally L Baxter , Ale x McK eown, Ge Y ang, Xiaokang W u, Fangbing Y an, et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell , 172(5):1122– 1131, 2018. 135 [168] Prav een Govi. Coronahack - chest x-ray-dataset. https://www.kaggle.com/datasets/ praveengovi/coronahack- chest- xraydataset , 2019. Kaggle dataset (uploader: prav eengovi). Accessed 2025-08-21. 135 [169] Xiaosong W ang, Y ifan Peng, Le Lu, Zhiyong Lu, Mohammadhadi Bagheri, and Ronald M Summers. Chestx-ray8: Hospital-scale chest x-ray database and benchmarks on weakly- supervised classification and localization of common thorax diseases. In Pr oceedings of the IEEE Confer ence on Computer V ision and P attern Reco gnition , pages 2097–2106, 2017. 135 [170] Linda W ang, Zhong Qiu Lin, and Ale xander W ong. Co vid-net: a tailored deep con volutional neural netw ork design for detection of covid-19 cases from chest x-ray images. Scientific Reports , 10(1):19549, Nov 2020. ISSN 2045-2322. doi: 10.1038/s41598- 020- 76550- z. URL https://doi.org/10.1038/s41598- 020- 76550- z . 135 [171] Anna Zawacki, Carol W u, George Shih, Julia Elliott, Mikhail Fomitche v , Mohannad Hus- sain, ParasLakhani, Phil Culliton, and Shunxing Bao. Siim-acr pneumothorax segmenta- tion. https://kaggle.com/competitions/siim- acr- pneumothorax- segmentation , 2019. Kaggle. 135 [172] Raddar. IRMA x-ray dataset. https://www.kaggle.com/datasets/raddar/ irma- xray- dataset , 2020. Kaggle dataset (uploader: raddar); contains 14,000 X-ray images; used in ImageCLEF medical annotation tasks. Accessed 2025-08-21. 135 [173] Moulay A. Akhloufi and Mohamed Chetoui. Chest XR CO VID-19 detection. https:// cxr- covid19.grand- challenge.org/ , August 2021. Online; accessed September 2021. 135 [174] Joseph Paul Cohen, Paul Morrison, and Lan Dao. Covid-19 image data collec- tion. arXiv pr eprint arXiv:2003.11597 , 2020. URL https://github.com/ieee8023/ covid- chestxray- dataset . 135 [175] Amanullah Asraf and Zabirul Islam. CO VID19, pneumonia and normal chest x-ray pa dataset. https://data.mendeley.com/datasets/jctsfj2sfn/1 , April 2021. Mendeley Data (V1); doi:10.17632/jctsfj2sfn.1; CC BY 4.0; Accessed 2025-08-21. 135 65 [176] L Rodney Long, Sameer Antani, Dah-Jye Lee, Daniel M Krainak, and George R Thoma. Biomedical information from a national collection of spine x-rays: film to content-based retriev al. In Medical Imaging 2003: P ACS and Inte grated Medical Information Systems: Design and Evaluation , volume 5033, pages 70–84. SPIE, 2003. 135 [177] Liansheng W ang, Cong Xie, Y i Lin, Hong-Y u Zhou, Kailin Chen, Dalong Cheng, Florian Du- bost, Benjamin Collery , Bidur Khanal, Bishesh Khanal, Rong T ao, Shangliang Xu, Upasana Upadhyay Bharadw aj, Zhusi Zhong, Jie Li, Shuxin W ang, and Shuo Li. Ev aluation and com- parison of accurate automated spinal curvature estimation algorithms with spinal anterior- posterior x-ray images: The AASCE2019 challenge. Medical Image Analysis , 72:102115, 2021. ISSN 1361-8415. doi: https://doi.org/10.1016/j.media.2021.102115. URL https: //www.sciencedirect.com/science/article/pii/S1361841521001614 . 135 [178] Pranav Raikote (pranavraikokte). Covid-19 image dataset: 3 way classification - covid- 19, viral pneumonia, normal. https://www.kaggle.com/datasets/pranavraikokte/ covid19- image- dataset , 2020. Kaggle dataset (uploader: pranavraikokte); contains CO VID-19, viral pneumonia, and normal chest X-ray images. Accessed 2025-08-21. 135 [179] Stefan Jaeger , Sema Candemir, Sameer Antani, Yì-Xiáng J Wáng, Pu-Xuan Lu, and George Thoma. T wo public chest x-ray datasets for computer -aided screening of pulmonary diseases. Quantitative imaging in medicine and sur g ery , 4(6):475, 2014. 135 [180] Pranav Rajpurkar , Jeremy Irvin, Aarti Bagul, Daisy Ding, T ony Duan, Hershel Mehta, Bran- don Y ang, Kaylie Zhu, Dillon Laird, Robyn L Ball, et al. MURA: Large dataset for ab- normality detection in musculoskeletal radiographs. arXiv pr eprint arXiv:1712.06957 , 2017. 135 [181] John Suckling. The mammographic images analysis society digital mammogram database. In Exerpta Medica. International Congress Series, 1994 , volume 1069, pages 375–378, 1994. 135 [182] MD Anouk Stein, Carol W u, Chris Carr, Geor ge Shih, Jamie Dulkowski, kalpathy , Leon Chen, Luciano Prevedello, MD Marc K ohli, Mark McDonald, Peter, Phil Culliton, Safwan Halabi MD, and T ian Xia. Rsna pneumonia detection challenge. https://kaggle. com/competitions/rsna- pneumonia- detection- challenge , 2018. Kaggle. 135 [183] Duc Nguyen, DungNB, Ha Q. Nguyen, Julia Elliott, NguyenThanhNhan, and Phil Culliton. V inbigdata chest x-ray abnormalities detection. https://kaggle.com/competitions/ vinbigdata- chest- xray- abnormalities- detection , 2020. Kaggle. 135 [184] Paras Lakhani, John Mongan, Chinmay Singhal, Quan Zhou, Katherine P Andriole, W illiam F Auf fermann, PM Prasanna, Theresa X Pham, Michael Peterson, Peter J Bergquist, et al. The 2021 siim-fisabio-rsna machine learning co vid-19 challenge: Annotation and stan- dard exam classification of covid-19 chest radiographs. J ournal of Digital Imaging , 36(1): 365–372, 2023. 135 [185] Ecem Sogancioglu, Bram V an Ginnek en, Finn Behrendt, Marcel Bengs, Alexander Schlaefer , Miron Radu, Di Xu, K e Sheng, Fabien Scalzo, Eric Marcus, et al. Nodule detection and generation on chest x-rays: Node21 challenge. IEEE T r ansactions on Medical Imaging , 2024. 135 [186] K eni Zheng and Sokratis Makrogiannis. Bone texture characterization for osteoporosis diag- nosis using digital radiography . In 2016 38th Annual International Confer ence of the IEEE Engineering in Medicine and Biology Society (EMBC) , pages 1034–1037. IEEE, 2016. 135 [187] Laurens Hogeweg, Clara I. Sánchez, Pim A. de Jong, Pragnya Maduskar , and Bram van Ginneken. Clavicle segmentation in chest radiographs. Medical Ima ge Analysis , 16(8):1490– 1502, 2012. 135 [188] Siham T abik, Anabel Gómez-Ríos, José Luis Martín-Rodríguez, Iván Sevillano-García, Manuel Rey-Area, David Charte, Emilio Guirado, Juan-Luis Suárez, Julián Luengo, MA V alero-González, et al. CO VIDGR dataset and CO VID-SDNet methodology for pre- dicting co vid-19 based on chest x-ray images. IEEE J ournal of Biomedical and Health Infor- matics , 24(12):3595–3605, 2020. 135 66 [189] Jie Lian, Jingyu Liu, Shu Zhang, Kai Gao, Xiaoqing Liu, Dingwen Zhang, and Y izhou Y u. A structure-a ware relation network for thoracic diseases detection and segmentation. IEEE T ransactions on Medical Imaging , 40(8):2042–2052, 2021. 135 [190] Jarrel Seah, Jen, Maggie, Meng Law , Phil Culliton, and Sarah Dowd. RANZCR CLiP - catheter and line position challenge. https://kaggle.com/competitions/ ranzcr- clip- catheter- line- classification , 2020. Kaggle. 135 [191] Narinder Singh Punn and Sonali Agarwal. Covid-19 posteroanterior chest x-ray fused (cpcxr) dataset, 2020. URL https://dx.doi.org/10.21227/x2r3- xk48 . 135 [192] Junji Shiraishi, Shigehiko Katsuragawa, Junpei Ikezoe, Tsuneo Matsumoto, T akeshi K obayashi, K en-ichi Komatsu, Mitate Matsui, Hiroshi Fujita, Y oshie K odera, and Kunio Doi. Dev elopment of a digital image database for chest radiographs with and without a lung nod- ule: receiv er operating characteristic analysis of radiologists’ detection of pulmonary nodules. American Journal of Roentg enology , 174(1):71–74, 2000. 135 [193] Hasib Zunair and A Ben Hamza. Synthesis of covid-19 chest x-rays using unpaired image- to-image translation. Social network analysis and mining , 11(1):1–12, 2021. 135 [194] Ching-W ei W ang, Cheng-T a Huang, Meng-Che Hsieh, Chung-Hsing Li, Sheng-W ei Chang, W ei-Cheng Li, Rémy V andaele, Raphaël Marée, Sébastien Jodogne, Pierre Geurts, Cheng Chen, Guoyan Zheng, Chengwen Chu, Hengameh Mirzaalian, Ghassan Hamarneh, T omaž Vrtov ec, and Bulat Ibragimov . Ev aluation and comparison of anatomical landmark detection methods for cephalometric x-ray images: A grand challenge. IEEE T ransactions on Medical Imaging , 34(9):1890–1900, 2015. doi: 10.1109/TMI.2015.2412951. 135 [195] E. B. Tsai, S. Simpson, M. P . Lungren, M. Hershman, L. Roshkov an, E. Colak, B. J. Er- ickson, G. Shih, A. Stein, J. Kalpathy-Cramer , J. Shen, M. A. F . Hafez, S. John, P . Rajiah, B. P . Pogatchnik, J. T . Mongan, E. Altinmakas, E. Ranschaert, F . C. Kitamura, L. T opf f, L. Moy , J. P . Kanne, and C. C. W u. Data from medical imaging data resource cen- ter (midrc) - rsna international covid radiology database (ricord) release 1c - chest x-ray , covid+ (midrc-ricord-1c). https://www.cancerimagingarchive.net/collection/ midrc- ricord- 1c/ , 2021. V ersion 1 (updated 2021-01-15); DOI: 10.7937/91ah-v663; The Cancer Imaging Archiv e (TCIA); License: CC BY -NC 4.0; Accessed 2025-08-21. 135 [196] Daniel S Kerman y , Michael Goldbaum, W enjia Cai, Carolina CS V alentim, Huiying Liang, Sally L Baxter , Ale x McK eown, Ge Y ang, Xiaokang W u, Fangbing Y an, et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. cell , 172(5):1122– 1131, 2018. 135 , 136 [197] Muhammad EH Chowdhury , T awsifur Rahman, Amith Khandakar , Rashid Mazhar , Muham- mad Abdul Kadir , Zaid Bin Mahbub, Khandakar Reajul Islam, Muhammad Salman Khan, Atif Iqbal, Nasser Al Emadi, et al. Can ai help in screening viral and co vid-19 pneumonia? IEEE Access , 8:132665–132676, 2020. 135 [198] Sergii Stirenko, Y uriy Kochura, Ole g Alienin, Oleksandr Rokovyi, Y uri Gordienko, Peng Gang, and W ei Zeng. Chest x-ray analysis of tuberculosis by deep learning with segmen- tation and augmentation. In 2018 IEEE 38th International Confer ence on Electr onics and Nanotechnology (ELN ANO) , pages 422–428. IEEE, 2018. 135 [199] Ibrahim Ethem Hamamci, Sezgin Er , Enis Simsar, Atif Emre Y uksel, Sadullah Gultekin, Serife Damla Ozdemir , Kaiyuan Y ang, Hongwei Bran Li, Sarthak Pati, Bernd Stadlinger , et al. Dentex: An abnormal tooth detection with dental enumeration and diagnosis benchmark for panoramic x-rays. arXiv pr eprint arXiv:2305.19112 , 2023. 135 [200] Jun Cao, Juan Dai, Xuguang Li, Bingsheng Huang, Ching-W ei W ang, and Hongyuan Zhang. Cephalometric landmark detection in lateral x-ray images, April 2023. URL https://doi. org/10.5281/zenodo.7835592 . 135 [201] Muhammad Anwaar Khalid, Kanwal Zulfiqar , Ulfat Bashir , Areeba Shaheen, Rida Iqbal, Zarnab Rizwan, Ghina Rizwan, and Muhammad Moazam Fraz. Cepha29: Automatic cephalometric landmark detection challenge 2023. arXiv pr eprint arXiv:2212.04808 , 2022. 135 67 [202] Maxim Popov , Akmaral Amanturdiev a, Nuren Zhaksylyk, Alsabir Alkanov , Adilbek Saniyazbekov , T emirgali Aimyshev , Eldar Ismailov , Ablay Bulegenov , Arystan Kuzhukeye v , Aizhan Kulanbaye v a, et al. Dataset for automatic region-based coronary artery disease diag- nostics using x-ray angiography images. Scientific data , 11(1):20, 2024. 135 [203] Dequan W ang, Xiaosong W ang, Lilong W ang, Mengzhang Li, Qian Da, Xiaoqiang Liu, Xi- angyu Gao, Jun Shen, Junjun He, T ian Shen, et al. A real-world dataset and benchmark for foundation model adaptation in medical image classification. Scientific Data , 10(1):574, 2023. 135 [204] Serkan Çimen, Mathias Unberath, Alejandro Frangi, and Andreas Maier . CoronARe: A coro- nary artery reconstruction challenge. In International W orkshop on Computational Methods for Molecular Imaging , pages 96–104. Springer , 2017. 135 [205] A. Badano, C. G. Graf f, A. Badal, D. Sharma, R. Zeng, F . W . Samuelson, S. Glick, and K. J. Myers. The victre trial: Open-source, in-silico clinical trial for ev aluating digital breast tomosynthesis [data set]. https://www.cancerimagingarchive.net/ collection/victre/ , 2019. The Cancer Imaging Archive (TCIA); CC BY 3.0; Accessed 2025-08-21. 135 [206] Nicolas Gaggion, Candelaria Mosquera, Martina Aineseder, Lucas Mansilla, Die go Milone, and Enzo Ferrante. Che xmask database: a large-scale dataset of anatomical segmentation masks for chest x-ray images (v ersion 0.1) [data set]. https://physionet.org/content/ chexmask- cxr- segmentation- data/0.1/ , June 2023. PhysioNet; License: CC BY -NC- SA 4.0; Accessed 2025-08-21. 135 [207] Pingjun Chen. Knee osteoarthritis sev erity grading dataset. https://data. mendeley.com/datasets/56rmx5bjcr/1 , September 2018. Mendeley Data (V1); doi:10.17632/56rmx5bjcr .1; CC BY 4.0. 135 [208] Baidu AI Studio. X-ray hand joint classification dataset [data set], 2021. URL https: //aistudio.baidu.com/datasetdetail/69582/0 . Accessed: 2025-05-22. 135 [209] Safwan S Halabi, Luciano M Prev edello, Jayashree Kalpathy-Cramer , Artem B Mamonov , Alexander Bilbily , Mark Cicero, Ian Pan, Lucas Araújo Pereira, Rafael T eix eira Sousa, Nita- mar Abdala, et al. The RSN A pediatric bone age machine learning challenge. Radiology , 290 (2):498–503, 2019. 135 [210] Mingquan Lin, Gregory Holste, Song W ang, Y iliang Zhou, Y ishu W ei, Imon Banerjee, Pengyi Chen, T ianjie Dai, Y uexi Du, Nicha C Dvornek, et al. Cxr-lt 2024: A miccai challenge on long-tailed, multi-label, and zero-shot disease classification from chest x- ray . Medical Imag e Analysis , page 103739, 2025. 135 [211] Y anzhen Liu, Sutuk e Y ibulayimu, Gang Zhu, Chao Shi, Chendi Liang, Chunpeng Zhao, Xin- bao W u, Y udi Sang, and Y u W ang. Automatic pelvic fracture segmentation: a deep learning approach and benchmark dataset. F r ontiers in Medicine , 12:1511487, 2025. 135 [212] Y anzhen Liu, Sutuke Y ib ulayimu, Y udi Sang, Gang Zhu, Chao Shi, Chendi Liang, Qiyong Cao, Chunpeng Zhao, Xinbao W u, and Y u W ang. Preoperativ e fracture reduction planning for image-guided pelvic trauma surgery: A comprehensive pipeline with learning. Medi- cal Image Analysis , 102:103506, 2025. ISSN 1361-8415. doi: https://doi.org/10.1016/j. media.2025.103506. URL https://www.sciencedirect.com/science/article/pii/ S1361841525000544 . 135 [213] Chenglong Ma, Y uanfeng Ji, Jin Y e, Lu Zhang, Y ing Chen, Tianbin Li, Mingjie Li, Junjun He, and Hongming Shan. T owards interpretable counterfactual generation via multimodal autoregression. arXiv preprint , 2025. 135 [214] Huazhu Fu, Fei Li, Xu Sun, Xingxing Cao, Jingan Liao, Jose Ignacio Orlando, Xing T ao, Y uexiang Li, Shihao Zhang, Mingkui T an, et al. Age challenge: angle closure glaucoma ev aluation in anterior segment optical coherence tomography . Medical Imag e Analysis , 66: 101798, 2020. 136 68 [215] Bo Qian, Hao Chen, Xiangning W ang, Zhouyu Guan, Tingyao Li, Y ixiao Jin, Y ilan W u, Y ang W en, Haoxuan Che, Gitaek Kwon, et al. Drac 2022: A public benchmark for dia- betic retinopathy analysis on ultra-wide optical coherence tomography angiography images. P atterns , 5(3), 2024. 136 [216] Huihui Fang, Fei Li, Huazhu Fu, Junde W u, Xiulan Zhang, and Y anwu Xu. Dataset and ev aluation algorithm design for goals challenge. In International W orkshop on Ophthalmic Medical Image Analysis , pages 135–142. Springer , 2022. 136 [217] Mahdi Kazemian Jahromi, Raheleh Kafieh, Hossein Rabbani, Alireza Mehri Dehnavi, Alireza Peyman, Fedra Hajizadeh, and Mohammadreza Ommani. An automatic algorithm for se g- mentation of the boundaries of corneal layers in optical coherence tomography images using gaussian mixture model. J ournal of Medical Signals & Sensors , 4(3):171–180, 2014. 136 [218] T ahereh Mahmudi, Rahele Kafieh, Hossein Rabbani, Alireza Mehri Dehnavi, and Moham- madreza Akhlagi. Comparison of macular octs in right and left eyes of normal people, 2025. URL https://durham- repository.worktribe.com/output/1136638 . 136 [219] W eiyi Zhang, Peranut Chotcomwongse, Y inwen Li, Pusheng Xu, Ruijie Y ao, Lianhao Zhou, Y uxuan Zhou, Hui Feng, Qiping Zhou, Xinyue W ang, et al. Predicting diabetic macular edema treatment responses using oct: Dataset and methods of aptos competition. arXiv pr eprint arXiv:2505.05768 , 2025. 136 , 137 [220] Peyman Gholami, Priyanka Roy , Mohana Kuppuswamy Parthasarathy , and V asudev an Laksh- minarayanan. Octid: Optical coherence tomography image database. Computers & Electrical Engineering , 81:106532, 2020. 136 [221] Leyuan Fang, Shutao Li, Qing Nie, Joseph A Izatt, Cynthia A T oth, and Sina Farsiu. Sparsity based denoising of spectral domain optical coherence tomography images. Biomedical optics expr ess , 3(5):927–942, 2012. 136 [222] Hossein Rabbani, Michael J Allingham, Priyatham S Mettu, Scott W Cousins, and Sina Far - siu. Fully automatic segmentation of fluorescein leakage in subjects with diabetic macular edema. In vestigative ophthalmolo gy & visual science , 56(3):1482–1492, 2015. 136 [223] Rolando Estrada, Michael J Allingham, Priyatham S Mettu, Scott W Cousins, Carlo T omasi, and Sina Farsiu. Retinal artery-vein classification via topology estimation. IEEE transactions on medical imaging , 34(12):2518–2534, 2015. 136 [224] Rolando Estrada, Carlo T omasi, Scott C Schmidler, and Sina Farsiu. Tree topology esti- mation. IEEE tr ansactions on pattern analysis and machine intelligence , 37(8):1688–1701, 2014. 136 [225] Ziyun Y ang, Somayyeh Soltanian-Zadeh, Kengyeh K Chu, Haoran Zhang, Lama Moussa, Ariel E W atts, Nicholas J Shaheen, Adam W ax, and Sina Farsiu. Connectivity-based deep learning approach for segmentation of the epithelium in in viv o human esophageal oct images. Biomedical optics expr ess , 12(10):6326–6340, 2021. 136 [226] Mingchao Li, Songtao Y uan, and Qiang Chen. Octa-500, 2019. URL https://dx.doi. org/10.1016/j.media.2024.103092 . 136 [227] Zhuangzhuang Chen, Hualiang W ang, Chubin Ou, and Xiaomeng Li. Mutri: Multi-vie w tri- alignment for oct to octa 3d image translation. In IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , 2025. 136 , 155 [228] Jayanthi Siv aswamy , SR Krishnadas, Gopal Datt Joshi, Madhulika Jain, and A Ujjwaft Syed T abish. Drishti-gs: Retinal image dataset for optic nerve head (onh) segmentation. In 2014 IEEE 11th international symposium on biomedical imaging (ISBI) , pages 53–56. IEEE, 2014. 137 [229] Muhammad Moazam Fraz, Paolo Remagnino, Andreas Hoppe, Bunyarit Uyyanon v ara, Al- icja R. Rudnicka, Christopher G. Owen, and Sarah A. Barman. Chase db1: Retinal vessel reference dataset, 2012. URL https://researchdata.kingston.ac.uk/96/ . 137 69 [230] AD Hoover , V alentina Kouznetso v a, and Michael Goldbaum. Locating blood vessels in reti- nal images by piece wise threshold probing of a matched filter response. IEEE T r ansactions on Medical Imaging , 19(3):203–210, 2000. 137 [231] Joes Staal, Michael D Abràmoff, Meindert Niemeijer , Max A V ier gev er , and Bram V an Gin- neken. Ridge-based vessel segmentation in color images of the retina. IEEE transactions on medical imaging , 23(4):501–509, 2004. 137 [232] Prasanna Porwal, Samiksha Pachade, Ravi Kamble, Manesh Kokare, Girish Deshmukh, V ivek Sahasrabuddhe, and Fabrice Meriaudeau. Indian diabetic retinopathy image dataset (idrid): a database for diabetic retinopathy screening research. Data , 3(3):25, 2018. 137 [233] Jorge Cuadros and Geor ge Bresnick. Eyepacs: an adaptable telemedicine system for diabetic retinopathy screening. J ournal of diabetes science and technolo gy , 3(3):509–516, 2009. 137 [234] Sven Holm, Greg Russell, V incent Nourrit, and Niall McLoughlin. Dr hagis—a fundus image database for the automatic extraction of retinal surface vessels from diabetic patients. J ournal of Medical Imaging , 4(1):014503–014503, 2017. 137 [235] Ning Li, T ao Li, Chunyu Hu, Kai W ang, and Hong Kang. A benchmark of ocular disease intelligent recognition: One shot for multi-disease detection. In International symposium on benchmarking, measuring and optimization , pages 177–193. Springer , 2020. 137 [236] Samiksha Pachade, Prasanna Porw al, Dhanshree Thulkar , Manesh K okare, Girish Deshmukh, V ivek Sahasrabuddhe, Luca Giancardo, Gwenolé Quellec, and Fabrice Mériaudeau. Retinal fundus multi-disease image dataset (rfmid): A dataset for multi-disease detection research. Data , 6(2):14, 2021. 137 [237] Michael D Abràmof f, James C F olk, Dennis P Han, Jonathan D W alker , Da vid F W illiams, Stephen R Russell, Pascale Massin, Beatrice Cochener , Philippe Gain, Li T ang, et al. Au- tomated analysis of retinal images for detection of referable diabetic retinopathy . J AMA ophthalmology , 131(3):351–357, 2013. 137 [238] Huihui et al. Fang. Adam challenge: Detecting age-related macular degeneration from fundus images. IEEE transactions on medical ima ging , 41(10):2828–2847, 2022. 137 [239] Coen et al. De V ente. Airogs: Artificial intelligence for rob ust glaucoma screening challenge. IEEE transactions on medical imaging , 43(1):542–557, 2023. 137 [240] T omi Kauppi, V alentina Kalesn ykiene, Joni-Kristian Kamarainen, Lasse Lensu, Iiris Sorri, Asta Raninen, Raija V outilainen, and Hannu Uusitalo. The diaretdb1 diabetic retinopathy database and ev aluation protocol. In BMVC , volume 1, page 10, 2007. 137 [241] Attila Budai, Rüdiger Bock, Andreas Maier , Joachim Hornegger , and Georg Michelson. Ro- bust v essel segmentation in fundus images. International journal of biomedical imaging , 2013(1):154860, 2013. 137 [242] Huazhu Fu, Fei Li, José Ignacio Orlando, Hrv oje Boguno vi ´ c, Xu Sun, Jingan Liao, Y anwu Xu, Shaochong Zhang, and Xiulan Zhang. Palm: P athologic myopia challenge, 2019. URL https://dx.doi.org/10.21227/55pk- 8z03 . 137 [243] Carlos Hernandez-Matas, Xenophon Zabulis, Areti T riantafyllou, Panagiota Anyfanti, Stella Douma, and Antonis A Ar gyros. Fire: fundus image registration dataset. Artificial Intelli- gence in V ision and Ophthalmology , 1(4):16–28, 2017. 137 [244] T ianchi. Retina fundus image registration, 2021. URL https://tianchi.aliyun.com/ dataset/dataDetail?dataId=90112 . 137 [245] Ruhan Liu, Xiangning W ang, Qiang W u, Ling Dai, Xi F ang, T ao Y an, Jaemin Son, Shiqi T ang, Jiang Li, Zijian Gao, et al. Deepdrid: Diabetic retinopathy—grading and image quality estimation challenge. P atterns , 3(6), 2022. 137 70 [246] Johannes Rückert, Louise Bloch, Raphael Brüngel, Ahmad Idrissi-Y aghir , Henning Schäfer , Cynthia S Schmidt, Sven Koitka, Obioma Pelka, Asma Ben Abacha, Alba G. Seco de Herrera, et al. Rocov2: Radiology objects in context version 2, an updated multimodal image dataset. Scientific Data , 11(1):688, 2024. 137 [247] Qiao et al. Hu. Automated separation of binary o verlapping trees in lo w-contrast color retinal images. In International conference on medical image computing and computer-assisted intervention , pages 436–443, 2013. 137 [248] Junde W u, Huihui Fang, Fei Li, Huazhu Fu, Fengbin Lin, Jiongcheng Li, Y ue Huang, Qinji Y u, Sifan Song, Xinxing Xu, Y anyu Xu, W ensai W ang, Lingxiao W ang, Shuai Lu, Huiqi Li, Shihua Huang, Zhichao Lu, Chubin Ou, Xifei W ei, Bingyuan Liu, Riadh Kobbi, Xiaoy- ing T ang, Li Lin, Qiang Zhou, Qiang Hu, Hrvoje Bogunovic, José Ignacio Orlando, Xiulan Zhang, and Y anwu Xu. Gamma challenge: Glaucoma grading from multi-modality images. Medical Image Analysis , 90:102938, 2023. doi: 10.1016/j.media.2023.102938. 137 [249] Francisco José Fumero et al. Batista. Rim-one dl: A unified retinal image database for as- sessing glaucoma using deep learning. Image Analysis and Ster eology , 39(3):161–167, 2020. 137 [250] sshikamaru. Glaucoma detection, 2022. URL https://www.kaggle.com/datasets/ sshikamaru/glaucoma- detection . 137 [251] Andres et al. Diaz-Pinto. Cnns for automatic glaucoma assessment using fundus images: an extensi ve v alidation. Biomedical engineering online , 18(1):29, 2019. 137 [252] Stephanie J et al. Chiu. Automatic cone photoreceptor segmentation using graph theory and dynamic programming. Biomedical optics e xpr ess , 4(6):924–937, 2013. 137 [253] Uyen TV Nguyen, Alauddin Bhuiyan, Laurence AF Park, Ryo Ka wasaki, Tien Y W ong, Jie Jin W ang, P aul Mitchell, and K otagiri Ramamohanarao. An automated method for reti- nal arterio venous nicking quantification from color fundus images. IEEE T ransactions on Biomedical Engineering , 60(11):3194–3203, 2013. 137 [254] jr2ngb . Cataract dataset, 2019. URL https://www.kaggle.com/datasets/jr2ngb/ cataractdataset . 137 [255] Chi Liu, Xiaotong Han, Zhixi Li, Jason Ha, Guankai Peng, W ei Meng, and Mingguang He. A self-adapti ve deep learning method for automated eye laterality detection based on color fundus photography . Plos one , 14(9):e0222025, 2019. 137 [256] Adria Perez-Rovira, T MacGillivray , Emanuele T rucco, KS Chin, K Zutis, C Lupascu, Domenico T egolo, Andrea Giachetti, Peter J W ilson, A Doney , et al. V AMPIRE: V essel assessment and measurement platform for images of the retina. In 2011 Annual International Confer ence of the IEEE Engineering in Medicine and Biology Society , pages 3391–3394. IEEE, 2011. 137 [257] Ahmed Almazroa, Sami Alodhayb, Essameldin Osman, Eslam Ramadan, Mohammed Hum- madi, Mohammed Dlaim, Muhannad Alkatee, Kaamran Raahemifar , and V asudev an Laksh- minarayanan. Retinal fundus images for glaucoma analysis: the riga dataset. In Medical Imaging 2018: Imaging Informatics for Healthcare, Resear ch, and Applications , volume 10579, pages 55–62. SPIE, 2018. 137 [258] Samaneh Abbasi-Sureshjani, Iris Smit-Ockeloen, Jiong Zhang, and Bart T er Haar Romeny . Biologically-inspired supervised vasculature segmentation in slo retinal fundus images. In International Confer ence Image Analysis and Recognition , pages 325–334. Springer , 2015. 137 [259] Jie wei Jiang, Xiyang Liu, Lin Liu, Shuai W ang, Erping Long, Haoqing Y ang, Fuqiang Y uan, Deying Y u, Kai Zhang, Liming W ang, et al. Predicting the progression of ophthalmic dis- ease based on slit-lamp images using a deep temporal sequence network. PloS one , 13(7): e0201142, 2018. 137 71 [260] Ling-Ping Cen, Jie Ji, Jian-W ei Lin, Si-T ong Ju, Hong-Jie Lin, T ai-Ping Li, Y un W ang, Jian- Feng Y ang, Y u-Fen Liu, Shaoying T an, et al. Automatic detection of 39 fundus diseases and conditions in retinal photographs using deep neural networks. Natur e communications , 12 (1):4828, 2021. 137 [261] Jan Odstrcilik, Radim Kolar , Attila Budai, Joachim Hornegger , Jiri Jan, Jiri Gazarek, T omas Kubena, Pa vel Cernosek, Ondrej Svoboda, and Elli Angelopoulou. Retinal vessel se gmen- tation by improv ed matched filtering: ev aluation on a new high-resolution fundus image database. IET Imag e Pr ocessing , 7(4):373–383, 2013. 137 [262] Huihui Fang, Fei Li, Junde W u, Huazhu Fu, Xu Sun, Jaemin Son, Shuang Y u, Menglu Zhang, Chenglang Y uan, Cheng Bian, et al. Refuge2 challenge: A treasure tro ve for multi-dimension analysis and ev aluation in glaucoma screening. arXiv preprint , 2022. 137 [263] V icavr , 2011. URL http://www.varpa.es/research/ophtalmology.html#vicavr . 137 [264] Abdullah Sarhan, Jon Rokne, Reda Alhajj, and Andrew Crichton. T ransfer learning through weighted loss function and group normalization for vessel se gmentation from retinal images. In 2020 25th International Confer ence on P attern Recognition (ICPR) , pages 9211–9218. IEEE, 2021. 137 [265] Subhadeep Chakraborty . Drimdb (diabetic retinopathy images database), 2024. URL https: //www.kaggle.com/ds/4523071 . 137 [266] Y ihao Li, Philippe Zhang, Y ubo T an, Jing Zhang, Zhihan W ang, W eili Jiang, Pierre-Henri Conze, Mathieu Lamard, Gwenolé Quellec, and Mostafa El Habib Daho. Automated detec- tion of myopic maculopathy in mmac 2023: achiev ements in classification, segmentation, and spherical equi valent prediction. In International Conference on Medical Image Computing and Computer-Assisted Intervention , pages 1–17. Springer , 2023. 137 [267] M. A. Rodriguez, H. AlMarzouqi, and P . Liatsis. Multi-label retinal disease classification using transformers, 2022. URL . 137 [268] V eronica Elisa Castillo Benítez, Ingrid Castro Matto, Julio César Mello Román, José Luis Vázquez Noguera, Miguel García-T orres, Jordan A yala, Diego P Pinto-Roa, Pedro E Gardel-Sotomayor , Jacques Facon, and Sebastian Alberto Grillo. Dataset from fundus im- ages for the study of diabetic retinopathy . Data in brief , 36:107068, 2021. 137 [269] Mir T anvir Islam, Shafin T Mashfu, Abrar Faisal, Sadman Chowdhury Siam, Intisar T ahmid Naheen, and Riasat Khan. Deep learning-based glaucoma detection with cropped optic cup and disc and blood vessel se gmentation. Ieee Access , 10:2828–2841, 2021. 137 [270] Liu Li, Mai Xu, Xiaofei W ang, Lai Jiang, and Hanruo Liu. Attention based glaucoma detec- tion: A large-scale database and cnn model. In The IEEE Confer ence on Computer V ision and P attern Recognition (CVPR) , June 2019. 137 [271] Kai Jin, Xingru Huang, Jingxing Zhou, Y unxiang Li, Y an Y an, Y ibao Sun, Qianni Zhang, Y aqi W ang, and Juan Y e. Fives: A fundus image dataset for artificial intelligence based vessel se gmentation. Scientific Data , 9(1):475, 2022. 137 [272] Isha Kansal, V ikas Khullar , Preeti Sharma, Supreet Singh, Junainah Abd Hamid, and A John- son Santhosh. Multiple model visual feature embedding and selection method for an ef ficient ocular disease classification. Scientific Reports , 15(1):5157, 2025. 137 [273] Oleksandr K ov alyk, Juan Morales-Sánchez, Rafael V erdú-Monedero, Inmaculada Sellés- Nav arro, Ana Palazón-Cabanes, and José-Luis Sancho-Gómez. Papila: Dataset with fundus images and clinical data of both e yes of the same patient for glaucoma assessment. Scientific Data , 9(1):291, 2022. 137 [274] Ungsoo Kim. Machine learning for pseudopapilledema, Aug 2018. URL osf.io/2w5ce . 137 72 [275] Oli via Cardozo, V erena Ojeda, Rodrigo Parra, Julio César Mello-Román, José Luis Vázquez Noguera, Miguel García-T orres, Federico Divina, Sebastian A. Grillo, Cyn- thia V illalba, Jacques Facon, V eronica Elisa Castillo Benítez, Ingrid Castro Matto, and Diego Aquino-Brítez. Dataset of fundus images for the diagnosis of ocular toxoplasmo- sis. Data in Brief , 48:109056, 2023. ISSN 2352-3409. doi: https://doi.org/10.1016/ j.dib .2023.109056. URL https://www.sciencedirect.com/science/article/pii/ S2352340923001749 . 137 [276] Noel Codella, V eronica Rotemberg, Philipp Tschandl, M Emre Celebi, Stephen Dusza, David Gutman, Brian Helba, Aadi Kalloo, K onstantinos Liopyris, Michael Marchetti, et al. Skin lesion analysis to ward melanoma detection 2018: A challenge hosted by the international skin imaging collaboration (isic). arXiv pr eprint arXiv:1902.03368 , 2019. 138 [277] V eronica Rotemberg, Nicholas Kurtansky , Brigid Betz-Stablein, Liam Caf fery , Emmanouil Chousakos, Noel Codella, Marc Combalia, Stephen Dusza, Pascale Guitera, David Gutman, et al. A patient-centric dataset of images and metadata for identifying melanomas using clinical context. Scientific data , 8(1):34, 2021. 138 [278] David Gutman, Noel CF Codella, Emre Celebi, Brian Helba, Michael Marchetti, Nabin Mishra, and Allan Halpern. Skin lesion analysis tow ard melanoma detection: A challenge at the international symposium on biomedical imaging (isbi) 2016, hosted by the international skin imaging collaboration (isic). arXiv pr eprint arXiv:1605.01397 , 2016. 138 [279] Noel CF Codella, David Gutman, M Emre Celebi, Brian Helba, Michael A Marchetti, Stephen W Dusza, Aadi Kalloo, Konstantinos Liopyris, Nabin Mishra, Harald Kittler , et al. Skin lesion analysis toward melanoma detection: A challenge at the 2017 international sym- posium on biomedical imaging (isbi), hosted by the international skin imaging collaboration (isic). In 2018 IEEE 15th international symposium on biomedical ima ging (ISBI 2018) , pages 168–172. IEEE, 2018. 138 [280] Jeremy Ka wahara, Sara Daneshvar , Giuseppe Argenziano, and Ghassan Hamarneh. Seven- point checklist and skin lesion classification using multitask multimodal neural nets. IEEE journal of biomedical and health informatics , 23(2):538–546, 2018. 138 [281] Matthe w Groh, Caleb Harris, Luis Soenksen, Felix Lau, Rachel Han, Aerin Kim, Arash K oochek, and Omar Badri. Ev aluating deep neural networks trained on clinical images in dermatology with the fitzpatrick 17k dataset. In Pr oceedings of the IEEE/CVF confer ence on computer vision and pattern r ecognition , pages 1820–1828, 2021. 138 [282] Ioannis Giotis, Nynke Molders, Sander Land, Michael Biehl, Marcel F Jonkman, and Nicolai Petkov . Med-node: A computer-assisted melanoma diagnosis system using non-dermoscopic images. Expert systems with applications , 42(19):6578–6585, 2015. 138 [283] Andre GC P acheco, Gusta vo R Lima, Amanda S Salomao, Breno Krohling, Igor P Biral, Gabriel G De Angelo, Fábio CR Alv es Jr , José GM Esgario, Alana C Simora, Pedro BC Castro, et al. Pad-ufes-20: A skin lesion dataset composed of patient data and clinical images collected from smartphones. Data in brief , 32:106221, 2020. 138 [284] T eresa Mendonça, Pedro M Ferreira, Jor ge S Marques, André RS Marcal, and Jor ge Rozeira. Ph 2-a dermoscopic image database for research and benchmarking. In 2013 35th annual international conference of the IEEE engineering in medicine and biology society (EMBC) , pages 5437–5440. IEEE, 2013. 138 [285] Moi Hoon Y ap, Ryo Hachiuma, Azadeh Alavi, Raphael Brüngel, Bill Cassidy , Manu Goyal, Hongtao Zhu, Johannes Rückert, Moshe Olshansk y , Xiao Huang, et al. Deep learning in dia- betic foot ulcers detection: A comprehensi ve ev aluation. Computers in biology and medicine , 135:104596, 2021. 138 [286] Xiaoxiao Sun, Jufeng Y ang, Ming Sun, and Kai W ang. A benchmark for automatic visual classification of clinical skin disease images. In Eur opean confer ence on computer vision , pages 206–222. Springer , 2016. 138 73 [287] Jufeng Y ang, Xiaoping W u, Jie Liang, Xiaoxiao Sun, Ming-Ming Cheng, Paul L Rosin, and Liang W ang. Self-paced balance learning for clinical skin disease recognition. IEEE trans- actions on neural networks and learning systems , 31(8):2832–2846, 2019. 138 [288] Shams Nafisa Ali, Md. T azuddin Ahmed, Joydip P aul, T asnim Jahan, S. M. Sakeef Sani, Nawshaba Noor , and T aufiq Hasan. Monkeypox skin lesion detection using deep learning models: A preliminary feasibility study . arXiv pr eprint arXiv:2207.03342 , 2022. 138 [289] Md. T azuddin Ahmed, Joydip Paul, T asnim Jahan, Shams Nafisa Ali, S. M. Sakeef Sani, Nawshaba Noor, Anzirun Nahar Asma, and T aufiq Hasan. A web-based mpox skin lesion de- tection system using state-of-the-art deep learning models considering racial div ersity . arXiv pr eprint arXiv:2306.14169 , 2023. 138 [290] V itiligo images, 2019. URL https://www.kaggle.com/datasets/shaikhshahid/ vitiligo- images . 138 [291] W outer Bulten, Kimmo Kartasalo, Po-Hsuan Cameron Chen, Peter Ström, Hans Pinckaers, Kunal Nagpal, Y uannan Cai, David F Steiner , Hester V an Bov en, Robert V ink, et al. Artificial intelligence for diagnosis and gleason grading of prostate cancer: the panda challenge. Natur e medicine , 28(1):154–163, 2022. 139 , 140 [292] Guy Nir , Soheil Hor , Davood Karimi, Ladan Fazli, Brian F Skinnider , Peyman T a vassoli, Dmitry T urbin, Carlos F V illamil, Gang W ang, R Storey W ilson, et al. Automatic grading of prostate cancer in digitized histopathology images: Learning from multiple experts. Medical image analysis , 50:167–180, 2018. 139 [293] Xuehai He, Y ichen Zhang, Luntian Mou, Eric Xing, and Pengtao Xie. Pathvqa: 30000+ questions for medical visual question answering. arXiv pr eprint arXiv:2003.10286 , 2020. 139 [294] Gabriele Campanella, Matthe w G Hanna, Edi Brogi, and Thomas J Fuchs. Breast metastases to axillary lymph nodes. (No T itle) , 2019. 139 [295] Neeraj Kumar , Ruchika V erma, Deepak Anand, Y anning Zhou, Omer Fahri Onder , Efstratios Tsougenis, Hao Chen, Pheng-Ann Heng, Jiahui Li, Zhiqiang Hu, et al. A multi-or gan nucleus segmentation challenge. IEEE transactions on medical imaging , 39(5):1380–1391, 2019. 139 [296] Ruchika V erma, Neeraj Kumar , Abhijeet Patil, Nikhil Cherian Kurian, Swapnil Rane, Si- mon Graham, Quoc Dang V u, Mieke Zwager , Shan E Ahmed Raza, Nasir Rajpoot, et al. Monusac2020: A multi-or gan nuclei segmentation and classification challenge. IEEE T rans- actions on Medical Imaging , 40(12):3413–3423, 2021. 139 [297] Qian Da, Xiaodi Huang, Zhongyu Li, Y anfei Zuo, Chenbin Zhang, Jingxin Liu, W en Chen, Jiahui Li, Dou Xu, Zhiqiang Hu, et al. Digestpath: A benchmark dataset with challenge revie w for the pathological detection and segmentation of digesti v e-system. Medical imag e analysis , 80:102485, 2022. 139 [298] Geert Litjens, Peter Bandi, Babak Ehteshami Bejnordi, Oscar Geessink, Maschenka Balken- hol, Peter Bult, Altuna Halilovic, Meyk e Hermsen, Rob V an de Loo, Rob V ogels, et al. 1399 h&e-stained sentinel lymph node sections of breast cancer patients: the camelyon dataset. GigaScience , 7(6):giy065, 2018. 139 [299] Ji ˇ rí Borov ec, Jan Kybic, Ignacio Arganda-Carreras, Dmitry V Sorokin, Gloria Bueno, Alexan- der V Khvostiko v , Spyridon Bakas, Eric I-Chao Chang, Stefan Heldmann, Kimmo Kartasalo, et al. Anhir: automatic non-rigid histological image registration challenge. IEEE tr ansactions on medical imaging , 39(10):3042–3052, 2020. 139 [300] Zhi Lu, Gustav o Carneiro, Andrew P Bradley , Daniela Ushizima, Masoud S Nosrati, An- drea GC Bianchi, Claudia M Carneiro, and Ghassan Hamarneh. Evaluation of three algo- rithms for the segmentation of o verlapping cervical cells. IEEE journal of biomedical and health informatics , 21(2):441–450, 2016. 139 , 141 74 [301] Marc Aubreville, Nikolas Stathonikos, Christof A Bertram, Robert Klopfleisch, Natalie T er Hoeve, Francesco Ciompi, Frauke W ilm, Christian Marzahl, T aryn A Donov an, Andreas Maier , et al. Mitosis domain generalization in histopathology images—the midog challenge. Medical Image Analysis , 84:102699, 2023. 139 [302] W outer Bulten, Péter Bándi, Jef frey Hoven, Rob v an de Loo, Johannes Lotz, Nick W eiss, Jeroen van der Laak, Bram van Ginneken, Christina Hulsbergen-van de Kaa, and Geert Lit- jens. Epithelium segmentation using deep learning in h&e-stained prostate specimens with immunohistochemistry as reference standard. Scientific r eports , 9(1):864, 2019. 139 [303] Kimberly H Allison, Lisa M Reisch, Patricia A Carne y , Donald L W eaver , Stuart J Schnitt, Frances P O’Malley , Berta M Geller, and Joann G Elmore. Understanding diagnostic v ariabil- ity in breast pathology: lessons learned from an expert consensus re vie w panel. Histopathol- ogy , 65(2):240–251, 2014. 139 [304] Simon Graham, Mostafa Jahanifar , Quoc Dang V u, Gior gos Hadjigeorghiou, Thomas Leech, David Snead, Shan E Ahmed Raza, Fayyaz Minhas, and Nasir Rajpoot. Conic: Colon nuclei identification and counting challenge 2022. arXiv pr eprint arXiv:2111.14485 , 2021. 139 [305] Je vgenij Gamper , Navid Alemi Koohbanani, Ksenija Benet, Ali Khuram, and Nasir Rajpoot. Pannuke: an open pan-cancer histology dataset for nuclei instance segmentation and classifi- cation. In Eur opean congr ess on digital pathology , pages 11–19. Springer , 2019. 139 [306] Nikita V Orlov , W ayne W Chen, David Mark Eckley , T omasz J Macura, Lior Shamir, Elaine S Jaffe, and Ilya G Goldberg. Automatic classification of lymphoma images with transform- based global features. IEEE T r ansactions on Information T echnology in Biomedicine , 14(4): 1003–1013, 2010. 139 , 140 [307] Paip 2021 challenge: Perineural in v asion in multiple org an cancer . https://paip2021. grand- challenge.org/ , 2021. Accessed: 2025-08-21. 139 [308] Elisa Drelie Gelasca, Jiyun Byun, Bogusla w Obara, and BS Manjunath. Evaluation and benchmark for biological image se gmentation. In 2008 15th IEEE international conference on image pr ocessing , pages 1816–1819. IEEE, 2008. 139 [309] Jiancheng Y ang, Rui Shi, and Bingbing Ni. Medmnist classification decathlon: A lightweight automl benchmark for medical image analysis. In IEEE 18th International Symposium on Biomedical Imaging (ISBI) , pages 191–195, 2021. 139 [310] Histopathologic cancer detection dataset. https://www.kaggle.com/competitions/ histopathologic- cancer- detection , 2018. 139 [311] Addison Howard, Andy Lawrence, Bud Sims, Eddie Tinsle y , Jarek Kazmierczak, Katy Borner , Leah Godwin, Marcos Nov aes, Phil Culliton, Richard Holland, Rick W atson, and Y ingnan Ju. Hubmap - hacking the kidney . https://kaggle.com/competitions/ hubmap- kidney- segmentation , 2020. Kaggle. 139 [312] Zhang Li, Jiehua Zhang, T ao T an, Xichao T eng, Xiaoliang Sun, Hong Zhao, Lihong Liu, Y ang Xiao, Byungjae Lee, Y ilong Li, et al. Deep learning methods for lung cancer segmentation in whole-slide histopathology images—the acdc@ lunghp challenge 2019. IEEE Journal of Biomedical and Health Informatics , 25(2):429–440, 2020. 139 [313] Anubha Gupta, Shiv Gehlot, Shubham Goswami, Sachin Motwani, Ritu Gupta, Álvaro García Faura, Dejan Štepec, T omaž Martin ˇ ci ˇ c, Reza Azad, Dorit Merhof, et al. Segpc-2021: A challenge & dataset on segmentation of multiple myeloma plasma cells from microscopic images. Medical Imag e Analysis , 83:102677, 2023. 139 [314] Lucia Ballerini, Robert B Fisher, Ben Aldridge, and Jonathan Rees. A color and texture based hierarchical k-nn approach to the classification of non-melanoma skin lesions. In Color medical image analysis , pages 63–86. Springer , 2013. 139 [315] Jun Ma, Ronald Xie, Shamini A yyadhury , Cheng Ge, Anubha Gupta, Ritu Gupta, Song Gu, Y ao Zhang, Gihun Lee, Joonkee Kim, et al. The multimodality cell segmentation challenge: tow ard uni versal solutions. Nature methods , 21(6):1103–1113, 2024. 139 75 [316] Mart v an Rijthov en, W itali Aswolinskiy , Leslie T essier, Maschenka Balkenhol, Joep Bo- gaerts, Jeroen van der Laak, Roberto Salgado, and Francesco Ciompi. T iger training dataset (wsirois subset) with roi-level annotations, 2022. URL https://zenodo.org/records/ 6014422 . Includes WSIR OIS, WSIBULK, WSITILS subsets; WSIBULK contains whole- slide images with tumor bulk annotations compatible with se gmentation pipelines. 139 [317] Shengjie Liu, Chuang Zhu, Feng Xu, Xinyu Jia, Zhongyue Shi, and Mulan Jin. Bci: Breast cancer immunohistochemical image generation through pyramid pix2pix. In Pr oceedings of the IEEE/CVF confer ence on computer vision and pattern r ecognition , pages 1815–1824, 2022. 139 [318] Chu Han, Xipeng P an, Lixu Y an, Huan Lin, Bingbing Li, Su Y ao, Shanshan Lv , Zhenwei Shi, Jinhai Mai, Jiatai Lin, et al. Wsss4luad: Grand challenge on weakly-supervised tissue semantic se gmentation for lung adenocarcinoma. arXiv preprint , 2022. 139 [319] Mohamed Amgad, Habiba Elfandy , Hagar Hussein, Lamees A Atteya, Mai A T Elsebaie, Lamia S Abo Elnasr , Rokia A Sakr, Hazem SE Salem, Ahmed F Ismail, Anas M Saad, et al. Structured cro wdsourcing enables con volutional se gmentation of histology images. Bioinfor - matics , 35(18):3461–3467, 2019. 139 [320] Mohamed Amgad, Lamees A Atteya, Hagar Hussein, Kareem Hosny Mohammed, Ehab Hafiz, Maha A T Elsebaie, Ahmed M Alhussein y , Mohamed Atef AlMoslemany , Abdel- magid M Elmatboly , Philip A Pappalardo, et al. Nucls: A scalable cro wdsourcing approach and dataset for nucleus classification and segmentation in breast cancer . GigaScience , 11: giac037, 2022. 139 [321] K yungmo Kim, Kyoungb un Lee, Sungduk Cho, Dong Un Kang, Seongkeun P ark, Y unsook Kang, Hyunjeong Kim, Gheeyoung Choe, K yung Chul Moon, Kyu Sang Lee, et al. P aip 2020: Microsatellite instability prediction in colorectal cancer . Medical Image Analysis , 89: 102886, 2023. 139 [322] Eduardo Conde-Sousa, João V ale, Ming Feng, K ele Xu, Y in W ang, V incenzo Della Mea, David La Barbera, Ehsan Montahaei, Mahdieh Baghshah, Andreas T urzynski, et al. Herohe challenge: predicting her2 status in breast cancer from hematoxylin–eosin whole-slide imag- ing. J ournal of Imaging , 8(8):213, 2022. 139 [323] Y iping Jiao, Jeroen V an Der Laak, Shadi Albarqouni, Zhang Li, T ao T an, Abhir Bhalerao, Shenghua Cheng, Jiabo Ma, Johnathan Pocock, Josien PW Pluim, et al. L ysto: The lympho- cyte assessment hackathon and benchmark dataset. IEEE journal of biomedical and health informatics , 28(3):1161–1172, 2023. 139 [324] Zaneta Swiderska-Chadaj, Hans Pinckaers, Mart V an Rijthov en, Maschenka Balkenhol, Mar - garita Melnikov a, Oscar Geessink, Quirine Manson, Mark Sherman, Antonio Polonia, Jeremy Parry , et al. Learning to detect lymphocytes in immunohistochemistry with deep learning. Medical image analysis , 58:101547, 2019. 139 [325] K orsuk Sirinukunwattana, Josien PW Pluim, Hao Chen, Xiaojuan Qi, Pheng-Ann Heng, Y un Bo Guo, Li Y ang W ang, Bogdan J Matuszewski, Elia Bruni, Urko Sanchez, et al. Gland segmentation in colon histology images: The glas challenge contest. Medical image analysis , 35:489–502, 2017. 139 [326] Simon Graham, Quoc Dang V u, Shan E Ahmed Raza, A yesha Azam, Y ee W ah Tsang, Jin T ae Kwak, and Nasir Rajpoot. Hov er-net: Simultaneous se gmentation and classification of nuclei in multi-tissue histology images. Medical imag e analysis , 58:101563, 2019. 139 [327] Julio Silva-Rodríguez, Adrián Colomer, María A Sales, Rafael Molina, and V alery Naranjo. Going deeper through the gleason scoring scale: An automatic end-to-end system for histol- ogy prostate grading and cribriform pattern detection. Computer methods and pr ograms in biomedicine , 195:105637, 2020. 139 [328] Jan Jantzen, Jonas Norup, Geor gios Dounias, and Beth Bjerre gaard. Pap-smear benchmark data for pattern classification. Natur e inspired smart information systems (NiSIS 2005) , pages 1–9, 2005. 139 76 [329] Richard J Chen and Rahul G Krishnan. Self-supervised vision transformers learn visual concepts in histopathology . arXiv pr eprint arXiv:2203.00585 , 2022. 139 [330] Shahira Abousamra, David Belinsky , John V an Arnam, Felicia Allard, Eric Y ee, Rajarsi Gupta, T ahsin Kurc, Dimitris Samaras, Joel Saltz, and Chao Chen. Multi-class cell detection using spatial context representation. In Pr oceedings of the IEEE/CVF International Confer- ence on Computer V ision , pages 4005–4014, 2021. 139 [331] Simon Graham, Hao Chen, Jevgenij Gamper, Qi Dou, Pheng-Ann Heng, David Snead, Y ee W ah Tsang, and Nasir Rajpoot. Mild-net: Minimal information loss dilated netw ork for gland instance se gmentation in colon histology images. Medical image analysis , 52:199– 211, 2019. 139 [332] Chuang Zhu, W enkai Chen, T ing Peng, Y ing W ang, and Mulan Jin. Hard sample aware noise robust learning for histopathology image classification. IEEE transactions on medical imaging , 41(4):881–894, 2021. 139 [333] Xinmi Huo, K ok Haur Ong, Kah W eng Lau, Laurent Gole, David M Y oung, Char Loo T an, Xiaohui Zhu, Chongchong Zhang, Y onghui Zhang, Longjie Li, et al. A comprehensive ai model de velopment frame work for consistent gleason grading. Communications Medicine , 4 (1):84, 2024. 139 [334] Mitko V eta, Y ujing J Heng, Nikolas Stathonikos, Babak Ehteshami Bejnordi, Francisco Beca, Thomas W ollmann, Karl Rohr, Manan A Shah, Dayong W ang, Mikael Rousson, et al. Pre- dicting breast tumor proliferation from whole-slide images: the tupac16 challenge. Medical image analysis , 54:111–121, 2019. 139 [335] Feng Y ang, Mahdieh Poostchi, Hang Y u, Zhou Zhou, Kamolrat Silamut, Jian Y u, Richard J Maude, Stefan Jaeger , and Sameer Antani. Deep learning for smartphone-based malaria parasite detection in thick blood smears. IEEE journal of biomedical and health informatics , 24(5):1427–1438, 2019. 139 [336] Anders Boesen Lindbo Larsen, Jacob Schack V estergaard, and Rasmus Larsen. Hep-2 cell classification using shape index histograms with donut-shaped spatial pooling. IEEE trans- actions on medical imaging , 33(7):1573–1580, 2014. 139 [337] Elisa Drelie Gelasca, Jiyun Byun, Boguslaw Obara, and B.S. Manjunath. Evaluation and benchmark for biological image segmentation. In IEEE International Confer ence on Im- age Pr ocessing , Oct 2008. URL http://vision.ece.ucsb.edu/publications/elisa_ ICIP08.pdf . 139 , 140 [338] Adam Shephard, Mostafa Jahanifar , Ruoyu W ang, Muhammad Dawood, Simon Graham, Kastytis Sidlauskas, Syed Ali Khurram, Nasir Rajpoot, and Shan E Ahmed Raza. T iager: T umor -infiltrating lymphoc yte scoring in breast cancer for the tiger challenge. arXiv pr eprint arXiv:2206.11943 , 2022. 140 [339] Mahdi S. Hosseini, Y ueyang Zhang, and Konstantinos N. Plataniotis. Encoding visual sensi- tivity by maxpol conv olution filters for image sharpness assessment. IEEE T ransactions on Image Pr ocessing , 28(9):4510–4525, September 2019. ISSN 1941-0042. doi: 10.1109/tip. 2019.2906582. URL http://dx.doi.org/10.1109/TIP.2019.2906582 . 140 [340] Blood cell images, 2018. URL https://www.kaggle.com/datasets/ paultimothymooney/blood- cells . 140 , 141 [341] J. N. Kather , F . G. Zöllner, F . Bianconi, S. M. Melchers, L. R. Schad, T . Gaiser , A. Marx, and C.-A. W eis. Collection of textures in colorectal cancer histology , 2016. URL https: //doi.org/10.5281/zenodo.53169 . 140 [342] Feng Xu, Chuang Zhu, W enqi T ang, Y ing W ang, Y u Zhang, Jie Li, Hongchuan Jiang, Zhongyue Shi, Jun Liu, and Mulan Jin. Predicting axillary lymph node metastasis in early breast cancer using deep learning on primary tumor biopsy slides. F r ontiers in oncology , 11: 759007, 2021. 140 77 [343] Shazia Akbar, Mohammad Peikari, Sherine Salama, Azadeh Y azdan Panah, Sharon Nofech- Mozes, and Anne L Martel. Automated and manual quantification of tumour cellularity in digital slides for tumour burden assessment. Scientific reports , 9(1):14099, 2019. 140 [344] Ching-W ei W ang, Nabila Puspita Firdi, Tzu-Chiao Chu, Mohammad Faiz Iqbal Faiz, Moham- mad Zafar Iqbal, Y ifan Li, Bo Y ang, Mayur Mallya, Ali Bashashati, Fei Li, et al. Atec23 chal- lenge: automated prediction of treatment ef fectiv eness in o v arian cancer using histopatholog- ical images. Medical Imag e Analysis , 99:103342, 2025. 140 [345] Philippe W eitz, Masi V alkonen, Leslie Solorzano, Circe Carr, Kimmo Kartasalo, Constance Boissin, Sonja K oi vukoski, Aino Kuusela, Dusan Rasic, Y anbo Feng, et al. The acrobat 2022 challenge: automatic registra tion of breast cancer tissue. Medical image analysis , 97:103257, 2024. 140 [346] JaeW oong Shin, Jeongun Ryu, Aaron V alero Puche, Jinhee Lee, Biagio Brattoli, W onkyung Jung, Soo Ick Cho, Kyunghyun Paeng, Chan-Y oung Ock, Donggeun Y oo, et al. Ocelot 2023: Cell detection from cell–tissue interaction challenge. Medical Image Analysis , 106:103751, 2025. 140 [347] Maryam Asadi-Aghbolaghi, Hossein Farahani, Allen Zhang, Ardalan Akbari, Sirim Kim, Ashley Chow , Sohier Dane, OCEAN Challenge Consortium, OTT A Consortium, Da vid G Huntsman, et al. Machine learning-driv en histotype diagnosis of o v arian carcinoma: insights from the ocean ai challenge. medRxiv , pages 2024–04, 2024. 140 [348] Sanne V ermorgen, Thijs Gelton, Peter Bult, Heidi VN Kusters-V andevelde, Jitka Hausnerová, K oen V an de V ijv er , Ben Davidson, Ingunn Marie Stefansson, Loes FS K ooreman, Adelina Qerimi, et al. Endometrial pipelle biopsy computer-aided diagnosis: a feasibility study . Mod- ern P athology , 37(2):100417, 2024. 140 [349] Patchcamelyon (pcam). https://patchcamelyon.grand- challenge.org/ , 2018. Grand Challenge dataset page. 140 [350] Bone marrow cytomorphology , 2021. URL https://wiki.cancerimagingarchive. net/pages/viewpage.action?pageId=101941770 . 140 [351] Parsa Ghahremani, Joseph Marino, Juan Hernandez-Prera, Jor ge V . de la Iglesia, Robbert J. Slebos, Christine H. Chung, and Saad Nadeem. An ai-ready multiplex staining dataset for reproducible and accurate characterization of tumor immune microen vironment. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2023 , volume 14225 of Lectur e Notes in Computer Science , pages 1–10. Springer , Cham, 2023. doi: 10.1007/978- 3- 031- 43987- 2_68. 140 [352] Anubha Gupta, Rahul Duggal, Shi vam Gehlot, Ritu Gupta, Ankit Mangal, Lalit K umar , Niraj Thakkar , and Debdoot Satpathy . Gcti-sn: Geometry-inspired chemical and tissue in variant stain normalization of microscopic medical images. Medical Image Analysis , 65:101788, 2020. doi: 10.1016/j.media.2020.101788. 140 [353] Ov arian bev acizumab response, 2023. URL https://wiki.cancerimagingarchive. net/pages/viewpage.action?pageId=83593077 . 140 [354] Cmb-lca: Combined multimodal biomarkers – lung cancer atlas, 2022. URL https://wiki. cancerimagingarchive.net/pages/viewpage.action?pageId=93258420 . 140 [355] Cptac-coad, 2021. URL https://wiki.cancerimagingarchive.net/pages/ viewpage.action?pageId=70227852 . 140 [356] Hungarian colorectal screening, 2022. URL https://wiki.cancerimagingarchive. net/pages/viewpage.action?pageId=91357370 . 140 [357] Dlbcl morphology , 2022. URL https://wiki.cancerimagingarchive.net/pages/ viewpage.action?pageId=119702520 . 140 [358] Cptac-ov , 2021. URL https://wiki.cancerimagingarchive.net/pages/viewpage. action?pageId=70227856 . 140 78 [359] Codex imaging of hepatocellular carcinoma (hcc), 2023. URL https://wiki. cancerimagingarchive.net/pages/viewpage.action?pageId=140313174 . 140 [360] Cptac-brca, 2021. URL https://wiki.cancerimagingarchive.net/pages/ viewpage.action?pageId=70227748 . 140 [361] Christian Matek, Sascha Schw arz, Karl Spiekermann, and Carsten Marr . Human-level recog- nition of blast cells in acute myeloid leukaemia with con volutional neural networks. Nature Machine Intelligence , 1:538–544, 2019. doi: 10.1038/s42256- 019- 0101- 9. 140 [362] Ritu Gupta and Anubha Gupta. Mimm_sbilab dataset: Microscopic images of multiple myeloma. (No T itle) , 2019. 140 [363] Le Hou, Rajarsi Gupta, John S V an Arnam, Y uwei Zhang, Kaustubh Siv alenka, Dimitris Samaras, T ahsin M Kurc, and Joel H Saltz. Dataset of segmented nuclei in hematoxylin and eosin stained histopathology images of ten cancer types. Scientific data , 7(1):185, 2020. 140 [364] Joel Saltz, Rajarsi Gupta, Le Hou, T ahsin Kurc, Pankaj Singh, V incent Nguyen, Dimitris Samaras, Kenneth R Shroyer , Tingting Zhao, Ryan Batiste, et al. Spatial organization and molecular correlates of tumor-infiltrating lymphocytes using deep learning on pathology im- ages. Cell Reports , 23(1):181–193.e7, 2018. doi: 10.1016/j.celrep.2018.10.077. URL https://pmc.ncbi.nlm.nih.gov/articles/PMC6250765/ . 140 [365] Ritu Gupta, Shubham Gehlot, and Anubha Gupta. C-nmc: B-lineage acute lymphoblastic leukaemia: A blood cancer dataset. Medical Engineering & Physics , 103:103793, 2022. doi: 10.1016/j.medengphy .2022.103793. URL https://www.sciencedirect.com/science/ article/pii/S1350453322001609 . 140 [366] National Cancer Institute Clinical Proteomic T umor Analysis Consortium (CPT A C). The clinical proteomic tumor analysis consortium acute myeloid leukemia collection (cptac-aml) (version 5), 2019. URL https://doi.org/10.7937/tcia.2019.b6foe619 . [dataset]. 140 [367] Frauke W ilm, Marco Fragoso, Christian Marzahl, Jingna Qiu, Chloé Puget, Laura Diehl, Christof A Bertram, Robert Klopfleisch, Andreas Maier, Katharina Breininger , and Marc Aubreville. Pan-tumor canine cutaneous cancer histology (catch) dataset. arXiv preprint arXiv:2201.11446 , 2022. URL . 140 [368] Lawrence W ilkinson, Benjamin W ang, Charles Johnson, ..., and Marco Gerlinger . Machine learning guided prognosis stratification using spatiotemporal patterns of tumor -infiltrating lymphocytes in neoadjuvant-treated prostate cancer . Eur opean Ur ology , 80:653–663, 2021. doi: 10.1016/j.eururo.2021.06.028. URL https://www.sciencedirect.com/science/ article/pii/S030228382101020X . 140 [369] Saman Farahmand, Aileen I Fernandez, Fahad Shabbir Ahmed, Da vid L Rimm, Jef frey H Chuang, Emily Reisenbichler , and K ourosh Zarringhalam. Deep learning trained on hema- toxylin and eosin tumor region of interest predicts her2 status and trastuzumab treatment response in her2+ breast cancer . Modern P athology , 35(1):44–51, 2022. 140 [370] Christian M Schürch, Sharmila S Bhate, Greg L Barlo w , David J Phillips, Lukas Noti, Inti Zlobec, Philip Chu, Sierra Black, Joy Demeter , Alejandra Méndez-Mancilla, et al. Coordi- nated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer in- vasi v e front. Cell , 183(4):1341–1359.e19, 2020. doi: 10.1016/j.cell.2020.10.033. URL https://www.cell.com/cell/fulltext/S0092- 8674(20)30870- 9 . 140 [371] AL Martel, Sharon Nofech-Mozes, Sherine Salama, Shazia Akbar , and Mohammad Peikari. Assessment of residual breast cancer cellularity after neoadjuv ant chemotherapy using digital pathology [data set]. The Cancer Imaging Ar chive , 2019. 140 [372] Harini B Arunachalam, Rituparna Mishra, Ovidiu Daescu, Karen Cederberg, Dinesh Rakheja, Annapurna Sengupta, David Leonard, and Patrick Lea v ey . V iable and necrotic tumor assess- ment from whole slide images of osteosarcoma using machine-learning and deep-learning models. PLOS ONE , 14(4):e0210706, 2019. doi: 10.1371/journal.pone.0210706. URL https://pubmed.ncbi.nlm.nih.gov/30995247/ . 140 79 [373] W isdom Oluchi Ikezogw o, Mehmet Saygin Seyfioglu, F atemeh Ghezloo, Dylan Stefan Chan Gev a, Fatwir Sheikh Mohammed, Pav an Kumar Anand, Ranjay Krishna, and Linda Shapiro. Quilt-1m: One million image-te xt pairs for histopathology , 2023. 140 [374] Martin Maška, Vladimír Ulman, Pablo Delgado-Rodriguez, Estibaliz Gómez-de Mariscal, T ereza Ne ˇ casová, Fidel A Guerrero Peña, Tsang Ing Ren, Elliot M Meyero witz, Tim Scherr , Katharina Löffler , et al. The cell tracking challenge: 10 years of objectiv e benchmarking. Natur e Methods , 20(7):1010–1020, 2023. 141 [375] Cremi: Miccai challenge on circuit reconstruction from electron microscopy images, 2016. URL https://cremi.org/ . 141 [376] Michał W ieczorek, Jakub Siłka, Katarzyna W iltos, and Marcin W o ´ zniak. Transformer based semantic segmentation network for medical imaging application. In International Confer ence on Artificial Intelligence and Soft Computing , pages 380–389. Springer , 2024. 141 [377] A. Santos and colleagues. Deep learning for leukemia classification: Performance analysis and challenges across multiple architectures. Diagnostics , 10:1014, 2020. doi: 10.3390/ diagnostics10121014. 141 [378] Anubha Gupta and Ritu Gupta. Isbi 2019 c-nmc challenge: Classification in cancer cell imaging. Select Pr oceedings , 2:27, 2019. 141 [379] Juan C Caicedo, Allen Goodman, Kyle W Karhohs, Beth A Cimini, Jeanelle Ackerman, Marzieh Haghighi, CherK eng Heng, T im Becker , Minh Doan, Claire McQuin, et al. Nucleus segmentation across imaging experiments: the 2018 data science bowl. Natur e methods , 16 (12):1247–1253, 2019. 141 [380] Guilherme Aresta, T eresa Araújo, Scotty Kwok, Sai Saketh Chennamsetty , Mohammed Safwan, V arghese Alex, Bahram Marami, Marcel Prasta wa, Monica Chan, Michael Dono van, et al. Bach: Grand challenge on breast cancer histology images. Medical image analysis , 56: 122–139, 2019. 141 [381] Mohammad Mahmudul Alam and Mohammad T ariqul Islam. Machine learning approach of automatic identification and counting of blood cells. Healthcar e T echnology Letters , 6(4): 103–108, 2019. 141 [382] Fariba Shaker , S Amirhassan Monadjemi, Jav ad Alirezaie, and Ahmad Reza Naghsh-Nilchi. A dictionary learning approach for human sperm heads classification. Computers in biology and medicine , 91:181–190, 2017. 141 [383] T rang Le, Casper F W insnes, Ulrika Ax elsson, Hao Xu, Jayasankar Mohanakrishnan Kaimal, Diana Mahdessian, Shubin Dai, Ilya S Makarov , Vladislav Ostankovich, Y ang Xu, et al. Analysis of the human protein atlas weakly supervised single-cell classification competition. Natur e methods , 19(10):1221–1229, 2022. 141 [384] Loris Nanni, Michelangelo Paci, Florentino Luciano Caetano dos Santos, Heli Skottman, Kati Juuti-Uusitalo, and Jari Hyttinen. T exture descriptors ensembles enable image-based classification of maturation of human stem cell-deri v ed retinal pigmented epithelium. PLoS One , 11(2):e0149399, 2016. 141 [385] Bikash R. Alam and colleagues. Automated segmentation of corneal endothelial cell images. Scientific Reports , 9:2284, 2019. doi: 10.1038/s41598- 019- 38859- w. 141 [386] A. de Bonnay , P . Thévenaz, J. Y un, et al. Automated analysis of in vi v o confocal microscop y images of the corneal sub-basal nerv e plexus and dendritic cells. T r anslational V ision Science & T echnology , 11:35, 2022. doi: 10.1167/tvst.11.2.35. 141 [387] Fabio Scarpa, Xiaodong Zheng, Y uichi Ohashi, and Alfredo Ruggeri. Automatic ev aluation of corneal nerve tortuosity in images from in vi vo confocal microscopy . In vestigative oph- thalmology & visual science , 52(9):6404–6408, 2011. 141 80 [388] Hady Ahmady Phoulady and Peter R Mouton. A new cervical cytology dataset for nucleus de- tection and image classification (cervix93) and methods for cervical nucleus detection. arXiv pr eprint arXiv:1811.09651 , 2018. 141 [389] Damir Vrabac, Akshay Smit, Rebecca Rojansky , Y asodha Natkunam, Ranjana H. Advani, Andrew Y . Ng, Sebastian Fernandez-Pol, and Pranav Rajpurkar . Dlbcl-morph: Morphological features computed using deep learning for an annotated digital dlbcl image set, 2020. 141 [390] Petteri T eikari, Marc Santos, Charissa Poon, and Kullerv o Hynynen. Deep learning con- volutional networks for multiphoton microscopy v asculature segmentation. arXiv pr eprint arXiv:1606.02382 , 2016. 141 [391] Sen Li, Zeyu Du, Xiangjie Meng, and Y ang Zhang. Multi-stage malaria parasite recognition by deep learning. GigaScience , 10(6):giab040, 2021. 141 [392] Y ide Zhang, Y inhao Zhu, Evan Nichols, Qingfei W ang, Siyuan Zhang, Cody Smith, and Scott How ard. A poisson-gaussian denoising dataset with real fluorescence microscopy images. In CVPR , 2019. 141 [393] Heywhale. Blood cell detection dataset. https://www.heywhale.com/mw/dataset/ 62c2af90913a54a66038165a , 2022. Dataset for blood cell object detection. 141 [394] T awsifur Rahman, Amith Khandakar, Muhammad Abdul Kadir , Khandaker Rejaul Islam, Khandakar F Islam, Rashid Mazhar , T ahir Hamid, Mohammad T ariqul Islam, Saad Kashem, Zaid Bin Mahb ub, et al. Reliable tuberculosis detection using chest x-ray with deep learning, segmentation and visualization. Ieee Access , 8:191586–191601, 2020. 141 [395] Soroush Jav adi and Seyed Abolghasem Mirroshandel. A novel deep learning method for automatic assessment of human sperm images. Computers in Biology and Medicine , 109: 182–194, 2019. doi: 10.1016/j.compbiomed.2019.04.030. 141 [396] Ali Hatamizadeh, Y ufan Xu, Demetri T erzopoulos, et al. Ravir: A dataset and methodology for the semantic segmentation and quantitative analysis of retinal arteries and v eins in infrared reflectance imaging. arXiv pr eprint arXiv:2203.04041 , 2022. 141 [397] Eduard Sojka et al. Mrl e ye dataset. http://mrl.cs.vsb.cz/eyedataset , 2018. 141 [398] K onstantin Pogorelov , Kristin Ranheim Randel, Carsten Griwodz, Sigrun Losada Eskeland, Thomas de Lange, Dag Johansen, Concetto Spampinato, Duc-T ien Dang-Nguyen, Mathias Lux, Peter Thelin Schmidt, et al. Kvasir: A multi-class image dataset for computer aided gastrointestinal disease detection. In Pr oceedings of the 8th ACM on Multimedia Systems Confer ence , pages 164–169, 2017. 142 [399] Kutse v Bengisu Ozyoruk, Guliz Irem Gokceler, Gulfize Coskun, Kagan Incetan, Y asin Al- malioglu, F aisal Mahmood, Ev a Curto, Luis Perdigoto, Marina Oli veira, Hasan Sahin, Helder Araujo, Henrique Alexandrino, Nicholas J. Durr, Hunter B. Gilbert, and Mehmet T uran. En- doslam dataset and an unsupervised monocular visual odometry and depth estimation ap- proach for endoscopic videos: Endo-sfmlearner , 2020. 142 [400] Sharib Ali and Noha M Ghatwary . Endoscopic computer vision challenges 2.0. In EndoCV@ ISBI , pages 5–8, 2022. 142 [401] Jorge Bernal, Nima T ajkbaksh, Francisco Javier Sanchez, Bogdan J Matusze wski, Hao Chen, Lequan Y u, Quentin Angermann, Olivier Romain, Bjørn Rustad, Ilangko Balasingham, et al. Comparativ e v alidation of polyp detection methods in video colonoscopy: results from the miccai 2015 endoscopic vision challenge. IEEE transactions on medical imaging , 36(6): 1231–1249, 2017. 142 [402] Andru P T winanda, Didier Mutter , Jacques Marescaux, Michel de Mathelin, and Nicolas Padoy . Single-and multi-task architectures for tool presence detection challenge at m2cai 2016. arXiv pr eprint arXiv:1610.08851 , 2016. 142 , 155 , 156 [403] Part of aida-e: Confocal endoscopy in celiac imaging, 2016. URL https:// aidasub- cleceliachy.grand- challenge.org/ . 142 81 [404] Part of aida-e: Esophagus microendoscopy images in barrett’ s surveillance, 2016. URL https://aidasub- clebarrett.grand- challenge.org/ . 142 [405] Part of aida-e: Gastric chromoendoscopy images in cancer surveillance, 2016. URL https: //aidasub- chromogastro.grand- challenge.org/ . 142 [406] David Vázquez, Jorge Bernal, F Javier Sánchez, Gloria Fernández-Esparrach, Antonio M López, Adriana Romero, Michal Drozdzal, and Aaron Courville. A benchmark for endolu- minal scene se gmentation of colonoscopy images. Journal of healthcare engineering , 2017 (1):4037190, 2017. 142 [407] Debesh Jha, Pia H Smedsrud, Michael A Riegler , Pål Halvorsen, Thomas De Lange, Dag Johansen, and Håvard D Johansen. Kvasir -seg: A segmented polyp dataset. In International confer ence on multimedia modeling , pages 451–462. Springer , 2019. 142 [408] Sophia Bano, Alessandro Casella, Francisco V asconcelos, Sara Moccia, George Attilakos, Ruwan Wimalasundera, Anna L David, Dario Paladini, Jan Deprest, Elena De Momi, et al. Fetreg: Placental vessel segmentation and registration in fetoscopy challenge dataset. arXiv pr eprint arXiv:2106.05923 , 2021. 142 [409] Ge-Peng Ji, Guobao Xiao, Y u-Cheng Chou, Deng-Ping Fan, Kai Zhao, Geng Chen, and Luc V an Gool. V ideo polyp segmentation: A deep learning perspective. Machine Intelligence Resear ch , 19(6):531–549, 2022. 142 , 157 [410] Gastrointestinal image analysis challenge, 2016. URL https://giana. grand- challenge.org/Home/ . 142 [411] Detection of abnormalities in gastroscopic images, 2015. URL https:// endovissub- abnormal.grand- challenge.org/EndoVisSub- Abnormal/ . 142 [412] Early barrett’ s cancer detection, 2015. URL https://endovissub- barrett. grand- challenge.org/ . 142 [413] Sharib Ali, Noha Ghatwary , Barbara Braden, Dominique Lamarque, Adam Baile y , Stefano Realdon, Renato Cannizzaro, Jens Rittscher, Christian Daul, and James East. Endoscopy disease detection challenge 2020. arXiv pr eprint arXiv:2003.03376 , 2020. 142 [414] Instrument segmentation and tracking, 2015. URL https://endovissub- instrument. grand- challenge.org/EndoVisSub- Instrument/ . 142 [415] Max Allan, Satoshi Kondo, Sebastian Bodenstedt, Stefan Le ger , Rahim Kadkhodamoham- madi, Imanol Luengo, Felix Fuentes, Evangello Flouty , Ahmed Mohammed, Marius Peder - sen, A vinash K ori, V arghese Ale x, Ganapathy Krishnamurthi, Da vid Rauber , Robert Mendel, Christoph Palm, Sophia Bano, Guinther Saibro, Chi-Sheng Shih, Hsun-An Chiang, Juntang Zhuang, Junlin Y ang, Vladimir Iglovikov , Anton Dobrenkii, Madhu Reddiboina, Anubhav Reddy , Xingtong Liu, Cong Gao, Mathias Unberath, Myeonghyeon Kim, Chanho Kim, Chae- won Kim, Hyejin Kim, Gyeongmin Lee, Ihsan Ullah, Miguel Luna, Sang Hyun Park, Mahdi Azizian, Danail Stoyanov , Lena Maier-Hein, and Stefanie Speidel. 2018 robotic scene seg- mentation challenge, 2020. URL . 142 [416] Surgical instrument multi-domain segmentation challenge, 2023. URL https://www. synapse.org/Synapse:syn47193563/wiki/ . 142 [417] Synthetic data for instrument se gmentation in surgery , 2023. URL https://www.synapse. org/Synapse:syn50908388/wiki/620516 . 142 [418] Preoperati ve to intraoperativ e laparoscopy fusion, 2022. URL https://p2ilf. grand- challenge.org/P2ILF/ . 142 [419] Haozheng Xu, Alistair W eld, Chi Xu, Alfie Roddan, João Cartucho, Mert Asim Karaoglu, Alexander Ladikos, Y angke Li, Y iping Li, Daiyun Shen, et al. Surgripe challenge: Bench- mark of surgical robot instrument pose estimation. Medical Image Analysis , page 103674, 2025. 142 82 [420] Salman Maqbool, Aqsa Riaz, Hasan Sajid, and Osman Hasan. m2caiseg: Semantic seg- mentation of laparoscopic images using con v olutional neural networks. arXiv preprint arXiv:2008.10134 , 2020. 142 [421] Zhao W ang, Chang Liu, Shaoting Zhang, and Qi Dou. Foundation model for endoscop y video analysis via lar ge-scale self-supervised pre-train. In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , pages 101–111. Springer, 2023. 142 , 155 [422] Hao Ding, Y uqian Zhang, T uxun Lu, Ruixing Liang, Hongchao Shu, Lalithkumar Seeni- vasan, Y onghao Long, Qi Dou, Cong Gao, Y icheng Leng, et al. Segstrong-c: Segmenting surgical tools rob ustly on non-adversarial generated corruptions–an endovis’ 24 challenge. arXiv pr eprint arXiv:2407.11906 , 2024. 142 [423] Xinwei Ju, Rema Daher , Razvan Caramalau, Baoru Huang, Danail Stoyano v , and Fran- cisco V asconcelos. Segcol challenge: Semantic segmentation for tools and fold edges in colonoscopy data. arXiv pr eprint arXiv:2412.16078 , 2024. 142 , 156 [424] Federated learning for sur gical vision, 2024. URL https://www.synapse.org/Synapse: syn53137385/wiki/ . 142 [425] Kaggle. Intel & mobileodt cervical cancer screening. https://www.kaggle.com/ competitions/intel- mobileodt- cervical- cancer- screening , 2017. Kaggle com- petition dataset. 142 [426] AIcro wd and Alzheimer’ s Disease Data Initiati ve. ADDI alzheimer’ s detection challenge. https://www.aicrowd.com/challenges/ addi- alzheimers- detection- challenge , 2021. Challenge and dataset page. 142 [427] Thomas Schaffter et al. Evaluation of combined artificial intelligence and radiologist assess- ment to interpret screening mammograms. J AMA Network Open , 3(9):e2011872, 2020. URL https://pubmed.ncbi.nlm.nih.gov/32997146/ . 142 [428] Hanchuan Peng et al. Bigneuron: Large-scale 3d neuron reconstruction from optical mi- croscopy images. Neur on , 87(2):252–256, 2015. URL https://www.ncbi.nlm.nih.gov/ pmc/articles/PMC4683726/ . 142 [429] R. H. Khan et al. Grand challenge on human activity classification with radar: Datasets, methods, and results. IEEE J ournal of Biomedical and Health Informatics , 2023. URL https://ieeexplore.ieee.org/document/10134905 . 142 [430] Debesh Jha et al. Kvasircapsule-se g. https://www.kaggle.com/datasets/ debeshjha1/kvasircapsuleseg , 2021. Segmentation subset from Kvasir -Capsule; dataset page suggests citing related work if used. 142 [431] SCUT V ision and Learning Lab . Scdb: Simple concept database (synthetic concept dataset). https://github.com/SCUT- VLlab/SCDB , 2020. Synthetic concept dataset; not a medical imaging dataset. 142 [432] Y uhui Ma, Huaying Hao, Huazhu Fu, Jiong Zhang, Jianlong Y ang, Jiang Liu, Y alin Zheng, and Y itian Zhao. R OSE: A retinal OCT -Angiography v essel se gmentation dataset and new model. arXiv pr eprint arXiv:2007.05201 , 2020. URL 05201 . 142 [433] Hongmin Cai, Jinhua W ang, T ingting Dan, Jiao Li, Zhihao Fan, W eiting Y i, Chunyan Cui, Xinhua Jiang, Li Li, et al. An online mammograph y database with biopsy confirmed types. Scientific Data , 10(123), 2023. doi: 10.1038/s41597- 023- 02025- 1. URL https://www. nature.com/articles/s41597- 023- 02025- 1 . 142 [434] Ahmed Hamada. Br35H: Brain tumor detection 2020. https://www.kaggle.com/ datasets/ahmedhamada0/brain- tumor- detection , 2020. Community dataset; no peer- revie wed dataset paper . 142 83 [435] Rana Khaled, Maha Helal, Omar Alfarghaly , Omnia Mokhtar , Abeer Elkorany , Hebatalla El Kassas, and Aly F ahmy . Categorized contrast enhanced mammography dataset for diag- nostic and artificial intelligence research. Scientific data , 9(1):122, 2022. 142 [436] Shi vam Barot and Parth Patel. Oral cancer (lips and tongue) images. https://www.kaggle. com/datasets/shivam17299/oral- cancer- lips- and- tongue- images , 2020. Kaggle dataset. 142 [437] Salman Sajid. Oral diseases (panoramic x-ray) dataset. https://www.kaggle.com/ datasets/salmansajid05/oral- diseases , 2023. Kaggle dataset. 142 [438] Roman Fusek. Pupil localization using geodesic distance. In Advances in V isual Computing (ISVC 2018) , v olume 11241 of Lectur e Notes in Computer Science , pages 433–444. Springer , 2018. URL https://mrl.cs.vsb.cz/eyedataset.html . 142 [439] Ludovic Roux and colleagues. Mitos & atypia 2014 grand challenge: Detection of mitosis and ev aluation of nuclear atypia in breast cancer h&e images. https://mitos- atypia- 14. grand- challenge.org/ , 2014. Grand Challenge dataset page. 142 [440] H. T . Nguyen, H. Q. Nguyen, H. H. Pham, K. Lam, L. T . Le, M. Dao, and V . V u. V indr - mammo: A large-scale benchmark dataset for computer-aided diagnosis in full-field digital mammography . Scientific Data , 10:277, 2023. doi: 10.1038/s41597- 023- 02100- 7. URL https://www.nature.com/articles/s41597- 023- 02100- 7 . 142 [441] H. T . Nguyen, H. H. Pham, N. T . Nguyen, H. Q. Nguyen, T . Q. Huynh, M. Dao, and V . V u. V indr-spinexr: A deep learning framework for spinal lesions detection and classification from radiographs. arXiv preprint , 2021. URL 2106.12930 . 142 [442] H. H. Pham, T . T . Tran, and H. Q. Nguyen. V indr -pcxr: An open, large-scale pediatric chest x-ray dataset for interpretation of common thoracic diseases. PhysioNet (version 1.0.0), 2022. URL https://physionet.org/content/vindr- pcxr/1.0.0/ . 142 [443] H. Q. Nguyen, K. Lam, L. T . Le, H. H. Pham, et al. V indr -cxr: An open dataset of chest x-rays with radiologist’ s annotations. Scientific Data , 9:429, 2022. doi: 10.1038/s41597- 022- 01498- w. URL https://www.nature.com/articles/ s41597- 022- 01498- w . 142 [444] Philipp Tschandl, Cliff Rosendahl, and Harald Kittler . The ham10000 dataset, a lar ge col- lection of multi-source dermatoscopic images of common pigmented skin lesions. Scientific Data , 5:180161, 2018. doi: 10.1038/sdata.2018.161. URL https://www.nature.com/ articles/sdata2018161 . 142 [445] Samiksha P achade, Prasanna Porwal, Dhanshree Thulkar , Manesh Kokare, Girish Desh- mukh, V iv ek Sahasrabuddhe, Luca Giancardo, Gwenolé Quellec, and Fabrice Mériaudeau. Retinal fundus multi-disease image dataset (rfmid): A dataset for multi-disease detection re- search. Data , 6(2):14, 2021. doi: 10.3390/data6020014. URL https://www.mdpi.com/ 2306- 5729/6/2/14 . 142 [446] J. R. Harish K umar , Chandra Sekhar Seelamantula, J. H. Gagan, Y ogish S. Kamath, Neetha I. R. Kuzhuppilly , U. V i vekanand, Preeti Gupta, and Shilpa Patil. Chák¸ su: A glaucoma spe- cific fundus image database. Scientific Data , 10:70, 2023. doi: 10.1038/s41597- 023- 01943- 4. URL https://www.nature.com/articles/s41597- 023- 01943- 4 . 142 [447] Andrea Acevedo, Anna Merino, Santiago Alférez, Ángel Molina, Laura Boldú, and José Rodellar . A dataset for microscopic peripheral blood cell images for dev elopment of automatic recognition systems, 2020. URL https://data.mendeley.com/datasets/ snkd93bnjr/1 . 142 [448] Sergios Gatidis, T obias Hepp, Marcel Früh, Christian La Fougère, Konstantin Nikolaou, Christina Pfannenberg, Bernhard Schölkopf, Thomas Küstner , Clemens Cyran, and Daniel Rubin. A whole-body fdg-pet/ct dataset with manually annotated tumor lesions. Scientific Data , 9:601, 2022. doi: 10.1038/s41597- 022- 01718- 3. 143 , 153 84 [449] Sergios Gatidis, T obias Hepp, Marcel Früh, Christian La Fougère, Konstantin Nikolaou, Christina Pfannenberg, Bernhard Schölkopf, Thomas Küstner , Clemens Cyran, and Daniel Rubin. The autopet ii dataset: a large-scale whole-body fdg-pet/ct dataset for lesion se gmen- tation. Scientific Data , 10:1–11, 2023. doi: 10.1038/s41597- 023- 02416- y. 143 , 153 [450] M. J. J. de Grauw , E. T . Scholten, E. J. Smit, M. J. C. M. Rutten, M. Prokop, B. v an Ginneken, and A. Hering. The uls23 challenge: A baseline model and benchmark dataset for 3d univ ersal lesion segmentation in computed tomography . Medical Image Analysis , 102:103525, 2025. doi: 10.1016/j.media.2025.103525. 143 [451] K e Y an, Xiaosong W ang, Le Lu, and Ronald M. Summers. Deeplesion: Automated mining of large-scale lesion annotations and univ ersal lesion detection with deep learning. Journal of Medical Imaging , 5(3):036501, 2018. doi: 10.1117/1.JMI.5.3.036501. 143 [452] Blaine Rister , Kaushik Shiv akumar , T omomi Nobashi, and Daniel L. Rubin. Ct-org: Ct volumes with multiple or gan segmentations. Dataset. The Cancer Imaging Archiv e, 2019. URL https://www.cancerimagingarchive.net/collection/ct- org/ . 143 [453] Zoé Lambert, Caroline Petitjean, Bernard Dubray , and Su Ruan. Segthor: Segmentation of thoracic organs at risk in ct images. In 2020 T enth International Confer ence on Image Pr o- cessing Theory , T ools and Applications (IPT A) , pages 1–6, 2020. doi: 10.1109/IPT A50016. 2020.9286453. 143 [454] W enxuan Li, Chongyu Qu, Xiaoxi Chen, Pedro RAS Bassi, Y ijia Shi, Y uxiang Lai, Qian Y u, Huimin Xue, Y ixiong Chen, Xiaorui Lin, et al. Abdomenatlas: A large-scale, detailed- annotated, & multi-center dataset for efficient transfer learning and open algorithmic bench- marking. Medical Image Analysis , page 103285, 2024. URL https://github.com/ MrGiovanni/AbdomenAtlas . 143 [455] Y uanfeng Ji, Chongjian Ge, Ruijiang Li, and Ping Luo. Amos-mm: Abdominal multimodal analysis challenge: Structured description of the challenge design, 2024. URL https:// doi.org/10.5281/zenodo.10992155 . 27th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2024). 143 [456] Jun Ma, Y ao Zhang, Song Gu, Xingle An, Zhihe W ang, Cheng Ge, Congcong W ang, Fan Zhang, Y u W ang, Y inan Xu, Shuiping Gou, Franz Thaler , Christian Payer , Darko Štern, Ed- ward G.A. Henderson, Dónal M. McSweeney , Andrew Green, Price Jackson, Lachlan McIn- tosh, Quoc-Cuong Nguyen, Abdul Qayyum, Pierre-Henri Conze, Ziyan Huang, Ziqi Zhou, Deng-Ping F an, Huan Xiong, Guoqiang Dong, Qiongjie Zhu, Jian He, and Xiaoping Y ang. Fast and low-gpu-memory abdomen ct or gan se gmentation: The flare challenge. Medical Image Analysis , 82:102616, 2022. doi: 10.1016/j.media.2022.102616. 143 [457] Jun Ma, Y ao Zhang, Song Gu, Cheng Ge, Shihao Ma, Adamo Y oung, Cheng Zhu, Kangkang Meng, Xin Y ang, and Ziyan Huang. Unleashing the strengths of unlabeled data in pan- cancer abdominal organ quantification: the flare22 challenge. In International Confer ence on Medical Image Computing and Computer -Assisted Intervention , pages 1–10. Springer , 2023. 143 [458] Jun Ma, Y ao Zhang, Song Gu, Cheng Ge, Ershuai W ang, Qin Zhou, Ziyan Huang, Pengju L yu, Jian He, and Bo W ang. Automatic or gan and pan-cancer se gmentation in abdomen ct: the flare 2023 challenge. arXiv pr eprint arXiv:2408.12534 , 2024. doi: 10.48550/arXi v .2408. 12534. 143 [459] Xiangde Luo, W enjun Liao, Jianghong Xiao, Jieneng Chen, T ao Song, Xiaofan Zhang, Kang Li, Dimitris N. Metaxas, Guotai W ang, and Shaoting Zhang. WORD: A large scale dataset, benchmark and clinical applicable study for abdominal organ segmentation from ct image. Medical Image Analysis , 82:102642, 2022. doi: 10.1016/j.media.2022.102642. 143 [460] Xiangde Luo, Zihan Li, Shaoting Zhang, W enjun Liao, and Guotai W ang. Rethinking abdom- inal org an segmentation (raos) in the clinical scenario: A robustness e v aluation benchmark with challenging cases. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2024 , pages 531–541. Springer , 2024. doi: 10.1007/978- 3- 031- 72114- 4_51. 143 85 [461] T obias Heimann, Bram van Ginneken, Martin A. Styner , Y ulia Arzhae va, V olker Aurich, Christian Bauer , Axel Beck, Christian Beck er , Reinhard Beichel, Gyor gy Bekes, Fabio Bello, Georg Binnig, Horst Bischof, Andrej Bornik, Patrick Cashman, Y uan Chi, Alberto Cor - dov a, Benoit Daw ant, Marta Fidrich, Daniel Furukawa, Laurent Grenacher , Joachim Horne g- ger , Dagmar Kainmueller, Richard Kitne y , Hide Kobatake, Hans Lameck er , Thomas Lange, Joohwi Lee, Brian Lennon, Rong Li, Sen Li, Hans-Peter Meinzer , Gabor Nemeth, Dorin Raicu, Axel Rau, Erik M. v an Rikxoort, Mikael Rousson, Laszlo Rusko, Kemal Saddi, Guenter Schmidt, Denis Seghers, Atsushi Shimizu, Pieter Slagmolen, Eric Sorantin, Gabriela Soza, Rosanee Susomboon, John W aite, Andreas W immer , and Iris W olf. Comparison and ev aluation of methods for li ver segmentation from ct datasets. IEEE T ransactions on Medical Imaging , 28(8):1251–1265, 2009. doi: 10.1109/TMI.2009.2013851. 143 [462] Holger R. Roth, Le Lu, Amal Farag, H.-C. Shin, Jiamin Liu, Evrim B. T urkbey , and Ronald M. Summers. Deeporgan: Multi-le vel deep con v olutional networks for automated pancreas segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2015, P art I , volume 9349 of Lectur e Notes in Computer Science , pages 556–564, 2015. doi: 10.1007/978- 3- 319- 24553- 9_68. 143 [463] A. Emre Ka vur , N. Sinem Gezer , Mustafa Barı ¸ s, Sinem Aslan, Pierre-Henri Conze, Vladimir Groza, Duc Duy Pham, Soumick Chatterjee, Philipp Ernst, Sav a¸ s Özkan, Bora Baydar , Dmitry Lachinov , Shuo Han, Josef Pauli, Fabian Isensee, Matthias Perkonigg, Rachana Sathish, Ronnie Rajan, Debdoot Sheet, Gurbandurdy Dovleto v , Oliv er Speck, Andreas Nürn- berger , Klaus H. Maier-Hein, Gözde Bozda ˘ gı Akar, Gözde Ünal, O ˘ guz Dicle, and M. Alper Selver . Chaos challenge - combined (ct-mr) health y abdominal organ segmentation. Medi- cal Image Analysis , 69:101950, Apr 2021. ISSN 1361-8415. doi: https://doi.org/10.1016/ j.media.2020.101950. URL http://www.sciencedirect.com/science/article/pii/ S1361841520303145 . 143 [464] Nicholas Heller , Niranjan Sathianathen, Arveen Kalapara, Edward W alczak, Keenan Moore, Heather Kaluzniak, Joel Rosenberg, Paul Blake, Zachary Rengel, Makinna Oestreich, Joshua Dean, Michael Trade well, Aneri Shah, Resha T ejpaul, Zachary Edgerton, Matthew Peterson, Shaneabbas Raza, Subodh Regmi, Nikolaos P apanikolopoulos, and Christopher W eight. The kits19 challenge data: 300 kidney tumor cases with clinical context, ct semantic se gmenta- tions, and surgical outcomes. arXiv preprint , 2019. 143 [465] Nicholas Heller, F abian Isensee, Dasha Trofimo v a, Resha T ejpaul, Zhongchen Zhao, Huai Chen, Lisheng W ang, Alex Golts, Daniel Khapun, Daniel Shats, Y oel Shoshan, Flora Gilboa- Solomon, Y asmeen George, Xi Y ang, Jianpeng Zhang, Jing Zhang, Y ong Xia, Mengran W u, Zhiyang Liu, Ed W alczak, Sean McSweeney , Ran v eer V asdev , Chris Hornung, Rafat So- laiman, Jamee Schoephoerster , Bailey Abernathy , David W u, Safa Abdulkadir, Ben Byun, Justice Spriggs, Grif fin Struyk, Alexandra Austin, Ben Simpson, Michael Hagstrom, Sierra V irnig, John French, Nitin V enkatesh, Sarah Chan, Keenan Moore, Anna Jacobsen, Susan Austin, Mark Austin, Subodh Regmi, Nikolaos P apanikolopoulos, and Christopher W eight. The kits21 challenge: Automatic segmentation of kidne ys, renal tumors, and renal cysts in corticomedullary-phase ct. arXiv pr eprint arXiv:2307.01984 , 2023. 143 [466] Nicholas Heller , Andrew W ood, Fabian Isensee, T im Rädsch, Resha T eipaul, Nikolaos Pa- panikolopoulos, and Christopher W eight. Kidney and Kidney T umor Se gmentation: MIC- CAI 2023 Challenge , KiTS 2023, Held in Conjunction with MICCAI 2023, V ancouver , BC, Canada, October 8, 2023, Pr oceedings , v olume 14540. Springer Nature, 2024. 143 [467] Shishuai Hu, Zehui Liao, Y iwen Y e, and Y ong Xia. Boundary-aware netw ork for kidney parsing. In Lesion Se gmentation in Surgical and Diagnostic Applications: MICCAI 2022 Challenges, CuRIOUS 2022, KiP A 2022 and MELA 2022, Held in Conjunction with MICCAI 2022, Singapor e, September 18–22, 2022, Pr oceedings , pages 9–17. Springer, 2023. 143 [468] Jun Ma, Y ao Zhang, Song Gu, Cheng Zhu, Cheng Ge, Y ichi Zhang, Xingle An, Congcong W ang, Qiyuan W ang, Xin Liu, Shucheng Cao, Qi Zhang, Shangqing Liu, Y unpeng W ang, Y uhui Li, Jian He, and Xiaoping Y ang. Abdomenct-1k: Is abdominal organ segmentation a solved problem? IEEE T ransactions on P attern Analysis and Machine Intelligence , 44(10): 6695–6714, 2022. doi: 10.1109/TP AMI.2021.3100536. 143 86 [469] Michela Antonelli, Annika Reinke, Spyridon Bakas, Keyv an Farahani, Annette K opp- Schneider , Bennett A Landman, Geert Litjens, Bjoern Menze, Olaf Ronneberger , Ronald M Summers, Bram v an Ginneken, Michel Bilello, Patrick Bilic, P atrick F Christ, Richard K G Do, Marc Gollub, Jennifer Golia-Pernicka, Stephan H Heckers, William R Jarnagin, Mau- reen K McHugo, Sandy Napel, Eugene V orontsov , Lena Maier-Hein, and M Jorge Car- doso. The medical segmentation decathlon. Natur e Communications , 13(1):4128, 2022. doi: 10.1038/s41467- 022- 30695- 9. 143 , 148 , 149 [470] Pechin Lo, Bram van Ginneken, Joseph M Reinhardt, T arunashree Y av arna, Pim A de Jong, Benjamin Irving, Catalin Fetita, Margarete Ortner , Rômulo Pinho, Jan Sijbers, Marco Feuer- stein, Anna Fabija ´ nska, Christian Bauer, Reinhard Beichel, Carlos S Mendoza, Rafael W iemker , Jaesung Lee, Anthony P Reev es, Silvia Born, Oli ver W einheimer , Ev a M v an Rikx- oort, Juerg Tschirren, K en Mori, Benjamin Odry , David P Naidich, Ieneke Hartmann, Eric A Hoffman, Mathias Prok op, Jesper H Pedersen, and Marleen de Bruijne. Extraction of airways from ct (exact’09). IEEE T ransactions on Medical Imaging , 31(11):2093–2107, 2012. doi: 10.1109/TMI.2012.2209674. 143 [471] Ev a v an Rikxoort, Bram v an Ginneken, and Sjoerd K erkstra. Lobe and lung analysis 2011 (lola11) dataset. Zenodo, 2011. 143 [472] Arnaud Arindra Adiyoso Setio, Alberto T rav erso, Thomas de Bel, Moira S.N. Berens, Cas van den Bogaard, Piergiorgio Cerello, Hao Chen, Qi Dou, Maria Evelina Fantacci, Bram Geurts, Robbert van der Gugten, Pheng Ann Heng, Bart Jansen, Michael M.J. de Kaste, V alentin Koto v , Jack Y u-Hung Lin, Jeroen T .M.C. Manders, Alexander Sónora-Mengana, Juan Carlos García-Naranjo, Evgenia Papav asileiou, Mathias Prokop, Marco Saletta, Cor- nelia M. Schaefer-Prokop, Ernst T . Scholten, Luuk Scholten, Miranda M. Snoeren, Ernesto Lopez T orres, Jef V andemeulebroucke, Nicole W alasek, Guido C.A. Zuidhof, Bram van Gin- neken, and Colin Jacobs. V alidation, comparison, and combination of algorithms for auto- matic detection of pulmonary nodules in computed tomography images: the luna16 challenge. Medical Image Analysis , 42:1–13, 2017. doi: 10.1016/j.media.2017.06.015. 143 [473] Minghui Zhang, Y angqian W u, Hanxiao Zhang, Y ulei Qin, Hao Zheng, W en T ang, Corey Arnold, Chenhao Pei, Pengxin Y u, Y ang Nan, et al. Multi-site, multi-domain airway tree modeling. Medical Image Analysis , 90:102957, 2023. doi: 10.1016/j.media.2023.102957. 143 [474] Y ang Nan, Xiaodan Xing, Shiyi W ang, Zeyu T ang, Federico N. Felder , Sheng Zhang, Roberta Eufrasia Ledda, Xiaoliu Ding, Ruiqi Y u, W eiping Liu, Feng Shi, T ian yang Sun, Zehong Cao, Minghui Zhang, Y un Gu, Hanxiao Zhang, Jian Gao, Pingyu W ang, W en T ang, Pengxin Y u, Han Kang, Junqiang Chen, Xing Lu, Boyu Zhang, Michail Mamalakis, Francesco Prinzi, Gianluca Carlini, Lisa Cuneo, Abhirup Banerjee, Zhaohu Xing, Lei Zhu, Zacharia Mesbah, Dhruv Jain, Tsiry Mayet, Hongyu Y uan, Qing L yu, Abdul Qayyum, Moona Mazher , Athol W ells, Simon L. F . W alsh, and Guang Y ang. Hunting imaging biomarkers in pulmonary fibrosis: Benchmarks of the aiib23 challenge. Medical Image Analysis , 97: 103253, 2024. doi: 10.1016/j.media.2024.103253. 143 [475] Maria de la Iglesia V aya, Jose Manuel Saborit, Joaquim Angel Montell, Antonio Per- tusa, Aurelia Bustos, Miguel Cazorla, Joaquin Galant, Xavier Barber, Domingo Orozco- Beltran, Francisco Garcia-Garcia, Marisa Caparros, German Gonzalez, and Jose Maria Salinas. Bimcv covid-19+: a large annotated dataset of rx and ct images from covid- 19 patients. arXiv pr eprint arXiv:2006.01174 , 2020. doi: 10.21227/w3a w- rv39. URL https://arxiv.org/abs/2006.01174 . 143 [476] S. Desai, A. Baghal, T . W ongsurawat, P . Jenjaroenpun, T . Powell, S. Al-Shukri, K. Gates, P . Farmer , M. Rutherford, G. Blake, T . Nolan, K. Se xton, W . Bennett, K. Smith, S. Syed, and F . Prior . Chest imaging representing a covid-19 positive rural u.s. population. Scientific Data , 7(1):414, 2020. doi: 10.1038/s41597- 020- 00741- 6. 143 [477] Peng An, Sheng Xu, Stephanie A. Harmon, Evrim B. T urkbey , Thomas H. Sanford, Amel Amalou, Michael Kassin, Nicole V arble, Maxime Blain, V ictoria Anderson, Francesca 87 Patella, Gianpaolo Carrafiello, Baris T . Turkbe y , and Bradford J. W ood. Ct images in covid- 19. The Cancer Imaging Archi ve, 2020. URL https://www.cancerimagingarchive. net/collection/ct- images- in- covid- 19/ . 143 [478] S. P . Morozov , A. E. Andreychenk o, N. A. P avlo v , A. V . Vladzymyrskyy , N. V . Ledikhova, V . A. Gombolevskiy , I. A. Blokhin, P . B. Gelezhe, A. V . Gonchar , and V . Y u. Chernina. Mosmeddata: Chest ct scans with covid-19 related findings dataset. Digital Diagnostics , 1 (1):49–59, 2020. doi: 10.17816/DD46826. 143 [479] Emily B. Tsai, Scott Simpson, Matthew P . Lungren, Michelle Hershman, Leonid Roshko van, Errol Colak, Bradley J. Erickson, George Shih, Anouk Stein, Jayashree Kalpathy-Cramer , Jody Shen, Mona Hafez, Susan John, Prabhakar Rajiah, Brian P . Pogatchnik, John Mongan, Emre Altinmakas, Erik R. Ranschaert, Felipe C. Kitamura, Laurens T opff, Linda Moy , Jef- frey P . Kanne, and Carol C. W u. The rsna international covid-19 open annotated radiology database (ricord). Radiology , 299(1):E204–E213, 2021. doi: 10.1148/radiol.2021203957. 143 [480] Jun Ma, Chongjian Ge, Y ixin W ang, Xingle An, Jian Gao, Zhaowei Y u, Y efeng Zhang, Dong Nie, Bingbing Li, Xiaoyan Meng, Jie Zhuo, and Qianqian Zhao. Covid-19-20 lung ct lesion segmentation challenge. Medical Image Analysis , 70:102193, 2021. 143 [481] Joel Saltz, Michael Saltz, Prateek Prasanna, Robert Moffitt, Janos Hajagos, Eric Bremer , Joseph Balsamo, and T ahsin K urc. Stony brook uni versity covid-19 positi ve cases (covid- 19-ny-sb u). The Cancer Imaging Archi ve, 2021. URL https://doi.org/10.7937/TCIA. BBAG- 2923 . 143 , 153 [482] Shokouh Shakouri, Mohammad Amin Bakhshali, Parv aneh Layegh, Behzad Kiani, Farid Ma- soumi, Saeedeh Ataei Nakhaei, and Sayyed Mostafa Mostafa vi. Covid19-ct-dataset: an open- access chest ct image repository of 1000+ patients with confirmed covid-19 diagnosis. BMC Resear ch Notes , 14:178, 2021. doi: 10.1186/s13104- 021- 05592- x. 143 [483] Eric B Tsai, Sam Simpson, Matthe w P Lungren, Meredith Hershman, Larissa Roshko- van, Ege Colak, Bradley J Erickson, Gordon Shih, Adam Stein, Jagadish Kalpathy-Cramer , Jun Shen, Marwan AF Hafez, Sarah John, Prabhakar Rajiah, Brett P Pogatchnik, Jen- nifer T Mongan, Emel Altinmakas, Erik Ranschaert, Fumi Cli ve Kitamura, Landon T opff, Lawrence Moy , Jef frey P Kanne, and Chin-Chang W u. Medical imaging data resource center (midrc) – rsna international covid-19 open radiology database (ricord) release 1b – chest ct covid- (midrc-ricord-1b) [data set]. The Cancer Imaging Archiv e, 2021. URL https://www.cancerimagingarchive.net/collection/midrc- ricord- 1b/ . 143 [484] Marie-Pierre Rev el, Soufiane Boussouar , Celine de Margerie-Mellon, Iryna Saab, Thibault Lapotre, Didier Mompoint, Amira Salhi, Anne-Lise Agdamdoua, K eyv an Razazi, and Guil- laume Chassagnon. Study of thoracic ct in covid-19: The stoic project. Radiology , 301: E361–E370, 2021. doi: 10.1148/radiol.2021210384. URL https://doi.org/10.1148/ radiol.2021210384 . Published online June 29, 2021. 143 [485] Ma Jun, Ge Cheng, W ang Y ixin, An Xingle, Gao Jiantao, Y u Ziqi, Zhang Minqing, Liu Xin, Deng Xue yuan, Cao Shucheng, et al. Covid-19 ct lung and infection segmentation dataset. (No T itle) , 2020. 143 [486] Dimitrios K ollias, Anastasios Arsenos, and Stefanos Kollias. A deep neural architecture for harmonizing 3-d input data analysis and decision making in medical imaging. Neur ocomput- ing , 542:126244, 2023. doi: 10.1016/j.neucom.2023.126244. 143 [487] Reuben Dorent, Ro ya Khajavi, T agwa Idris, Erik Ziegler , Bhanusupriya Somarouthu, Heather Jacene, Ann LaCasce, Jonathan Deissler, Jan Ehrhardt, Sofija Engelson, Stefan M. Fischer , Y un Gu, Heinz Handels, Satoshi Kasai, Satoshi K ondo, Klaus Maier-Hein, Julia A. Schn- abel, Guotai W ang, Litingyu W ang, T assilo W ald, Guang-Zhong Y ang, Hanxiao Zhang, Minghui Zhang, Steve Pieper , Gordon Harris, Ron Kikinis, and T ina Kapur . Lnq 2023 chal- lenge: Benchmark of weakly-supervised techniques for mediastinal lymph node quantifica- tion. MELB A , 2025. doi: 10.59275/j.melba.2025- d482. 143 88 [488] Gongning Luo, Kuanquan W ang, Jun Liu, Shuo Li, Xinjie Liang, Xiangyu Li, Shaowei Gan, W ei W ang, Suyu Dong, W enyi W ang, Pengxin Y u, Enyou Liu, Hongrong W ei, Na W ang, Jia Guo, Huiqi Li, Zhao Zhang, Ziwei Zhao, Na Gao, Nan An, Ashkan Pakzad, Bojidar Rangelov , Jiaqi Dou, Song Tian, Zeyu Liu, Y i W ang, Ampatishan Siv alingam, Kumarade v an Punithakumar , Zhaowen Qiu, and Xin Gao. Ef ficient automatic segmentation for multi-lev el pulmonary arteries: The parse challenge. arXiv pr eprint arXiv:2304.03708 , 2023. doi: 10. 48550/arXiv .2304.03708. 143 [489] Joao Pedrosa, Guilherme Aresta, Carlos Ferreira, Marcio Rodrigues, Patricia Leitao, An- dre Silv a Carv alho, Joao Rebelo, Eduardo Ne grao, Isabel Ramos, Antonio Cunha, and Au- relio Campilho. Lndb: A lung nodule database on computed tomography . arXiv pr eprint arXiv:1911.08434 , 2019. doi: 10.48550/arXiv .1911.08434. 143 [490] Mojtaba Masoudi, Hamid-Reza Pourreza, Mahdi Saadatmand-T arzjan, Noushin Eftekhari, Fateme Shafiee Zargar , and Masoud Pezeshki Rad. A new dataset of computed-tomography angiography images for computer-aided detection of pulmonary embolism. Scientific Data , 5:180180, 2018. doi: 10.1038/sdata.2018.180. 143 [491] Xiangyu Li, Gongning Luo, Kuanquan W ang, Hongyu W ang, Jun Liu, Xinjie Liang, Jie Jiang, Zhenghao Song, Chunyue Zheng, Haokai Chi, et al. The state-of-the-art 3d anisotropic intracranial hemorrhage segmentation on non-contrast head ct: The instance challenge. arXiv pr eprint arXiv:2301.03281 , 2023. 143 [492] ISLES Challenge Or ganizers. ISLES 2024: Ischemic Stroke Lesion Se gmentation Challenge. ISLES Challenge, 2024. URL https://www.isles- challenge.org/ . Longitudinal mul- timodal multicenter real-world data for acute to subacute ischemic stroke. 144 , 149 [493] Gašper Podobnik, Primož Strojan, Primož Peterlin, Bulat Ibragimov , and T omaž Vrtov ec. Han-seg: The head and neck or gan-at-risk ct & mr segmentation dataset. Medical Physics , 2023. doi: 10.1002/mp.16197. 144 [494] Xiangde Luo, Jia Fu, Y unxin Zhong, Shuolin Liu, Bing Han, Mehdi Astaraki, Simone Bendazzoli, Iuliana T oma-Dasu, Y iwen Y e, Ziyang Chen, Y ong Xia, Y anzhou Su, Jin Y e, Junjun He, Zhaohu Xing, Hongqiu W ang, Lei Zhu, Kaixiang Y ang, Xin Fang, Zhi- wei W ang, Chan W oong Lee, Sang Joon P ark, Jaehee Chun, Constantin Ulrich, Klaus H. Maier-Hein, Nchongmaje Ndipenoch, Alina Miron, Y ongmin Li, Y imeng Zhang, Y u Chen, Lu Bai, Jinlong Huang, Chengyang An, Lisheng W ang, Kaiwen Huang, Y unqi Gu, T ao Zhou, Mu Zhou, Shichuan Zhang, W enjun Liao, Guotai W ang, and Shaoting Zhang. Seg- rap2023: A benchmark of or gans-at-risk and gross tumor volume segmentation for radiother - apy planning of nasopharyngeal carcinoma. arXiv pr eprint arXiv:2312.09576 , 2023. URL https://arxiv.org/abs/2312.09576 . 144 [495] Patrik F . Raudaschl, Paolo Zaffino, Gregory C. Sharp, Maria Francesca Spadea, Antong Chen, Benoit M. Dawant, Thomas Albrecht, T obias Gass, Christoph Langguth, Marcel Lüthi, Flo- rian Jung, Oliver Knapp, Stefan W esarg, Richard Mannion-Haworth, Mik e Bo wes, Annaliese Ashman, Gwenael Guillard, Alan Brett, Graham V incent, Mauricio Orbes-Arteaga, David Cárdenas-Peña, German Castellanos-Dominguez, Nav a Aghdasi, Y angming Li, Angelique Berens, Kris Moe, Blake Hannaford, Rainer Schubert, and Karl D. Fritscher . Evaluation of segmentation methods on head and neck ct: Auto-segmentation challenge 2015. Medical Physics , 44(5):2020–2036, 2017. doi: 10.1002/mp.12197. 144 , 146 [496] Jun Shi. Structseg2019 gtv segmentation. https://structseg2019.grand-challenge.org, 2019. URL https://dx.doi.org/10.21227/h75x- gt46 . 144 [497] Alessa Hering, Keelin Murphy , and Bram van Ginneken. Learn2reg: comprehensiv e multi- task medical image registration challenge, dataset and ev aluation in the era of deep learn- ing. IEEE T ransactions on Medical Imaging , 42(3):697–712, 2021. doi: 10.1109/TMI.2022. 3213983. 144 , 148 , 152 , 153 [498] Kasper Marstal, Floris Berendsen, Niels Dekker , Marius Staring, and Stefan Klein. The continuous registration challenge: Ev aluation-as-a-service for medical image registration al- gorithms. In 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019) , pages 1399–1402, V enice, Italy , April 2019. doi: 10.1109/ISBI.2019.8759559. 144 89 [499] Richard Castillo, Edward Castillo, Rudy Guerra, V alen E. Johnson, Tra vis McPhail, Amit K. Garg, and Thomas Guerrero. A framew ork for ev aluation of deformable image registration spatial accuracy using large landmark point sets. Physics in Medicine and Biology , 54(7): 1849–1870, 2009. doi: 10.1088/0031- 9155/54/7/001. 144 [500] K eelin Murphy , Bram van Ginneken, Joseph M. Reinhardt, Sven Kabus, Kai Ding, Xiang Deng, Kunlin Cao, Kaifang Du, Gary E. Christensen, V incent Garcia, T om V ercauteren, Nicholas A yache, Oli vier Commo wick, Grégoire Malandain, Ben Glocker , Nikos P aragios, Nassir Nav ab, Vladlena Gorbuno va, Jon Sporring, Marleen De Bruijne, Xiao Han, Mat- tias P . Heinrich, Julia A. Schnabel, Mark Jenkinson, Cristian Lorenz, Marc Modat, Jamie R. McClelland, Sebastien Ourselin, Sascha E.A. Muenzing, Max A. V ier ge ver , Dante De Ni- gris, D. Louis Collins, T al Arbel, Marta Peroni, Rui Li, Gregory C. Sharp, Alexander Schmidt-Richberg, Jan Ehrhardt, Rene W erner, Dirk Smeets, Dirk Loeckx, Gang Song, Nicholas T ustison, Brian A vants, James C. Gee, Marius Staring, Stefan Klein, Berend C. Stoel, Martin Urschler , Manuel W erlberger , Jef V andemeulebroucke, Simon Rit, Da vid Sar- rut, and Josien P .W . Pluim. Evaluation of registration methods on thoracic ct: The em- pire10 challenge. IEEE T ransactions on Medical Imaging , 30(11):1901–1920, 2011. doi: 10.1109/TMI.2011.2158349. 144 [501] V incent Andrearczyk, V alentin Oreiller , Mario Jreige, Martin V allieres, Joel Castelli, Hesham Elhalawani, Sarah Boughdad, John O Prior, and Adrien Depeursinge. Overvie w of the hecktor challenge at miccai 2020: Automatic head and neck tumor segmentation in pet/ct. In Head and Neck T umor Se gmentation , pages 1–21. Springer , 2020. 144 , 153 [502] V incent Andrearczyk, V alentin Oreiller , Sarah Boughdad, Catherine Chez Le Rest, Hesham Elhalawani, Mario Jreige, John O. Prior, Martin V allières, Dimitris V isvikis, Mathieu Hatt, and Adrien Depeursinge. Overview of the hecktor challenge at miccai 2021: Automatic head and neck tumor segmentation and outcome prediction in pet/ct images. arXiv preprint arXiv:2201.04138 , 2022. URL . 144 , 153 [503] V incent Andrearczyk, V alentin Oreiller , Sarah Boughdad, Catherine Chez Le Rest, Hesham Elhalawani, Mario Jreige, John O. Prior, Martin V allières, Dimitris V isvikis, Mathieu Hatt, and Adrien Depeursinge. Overview of the hecktor challenge at miccai 2022: Automatic head and neck tumor segmentation and outcome prediction in pet/ct images. Medical Image Analysis , 84:102375, 2023. doi: 10.1016/j.media.2022.102375. URL https://doi.org/ 10.1016/j.media.2022.102375 . 144 , 153 [504] Anjany Sekuboyina, Malek E Husseini, Amirhossein Bayat, Maximilian Loffler , Hans Liebl, Hongwei Li, Giles T etteh, Jan Kukacka, Christian Payer , Darko Stern, Martin Urschler, Maodong Chen, Dalong Cheng, Nikolas Lessmann, Y ujin Hu, T ianfu W ang, Dong Y ang, Daguang Xu, Felix Ambellan, T amaz Amiranashvili, Moritz Ehlke, Hans Lamecker , Sebas- tian Lehnert, Marilia Lirio, Nicolas Perez de Olaguer , Heiko Ramm, Manish Sahu, Ale xan- der T ack, Stefan Zacho w , T ao Jiang, Xinjun Ma, Christoph Angerman, Xin W ang, Ke vin Brown, Alexandre Kirszenber g, Elodie Puybareau, Di Chen, Y iwei Bai, Brandon H Rapazzo, T imyoas Y eah, Amber Zhang, Shangliang Xu, Feng Hou, Zhiqiang He, Chan Zeng, Zheng Xiangshang, Xu Liming, T ucker J Netherton, Raymond P Mumme, Laurence E Court, Zixun Huang, Chenhang He, Li-W en W ang, Sai Ho Ling, Le Duy Huynh, Nicolas Boutry , Ro- man Jakubicek, Jiri Chmelik, Supriti Mulay , Mohanasankar Siv aprakasam, Johannes C Paet- zold, Suprosanna Shit, Iv an Ezho v , Benedikt W iestler , Ben Glocker , Alexander V alentinitsch, Markus Rempfler, Bjorn H Menze, and Jan S Kirschk e. V erse: A vertebrae labelling and segmentation benchmark for multi-detector ct images. Medical Image Analysis , 73:102166, 2021. doi: 10.1016/j.media.2021.102166. 144 [505] Hans Liebl, David Schinz, Anjany Sekuboyina, Luca Malagutti, Maximilian T Löffler , Amirhossein Bayat, Malek El Husseini, Giles T etteh, Katharina Grau, Eva Niederreiter , Thomas Baum, Benedikt Wiestler , Bjoern Menze, Rickmer Braren, Claus Zimmer , and Jan S Kirschke. A computed tomography vertebral segmentation dataset with anatom- ical variations and multi-vendor scanner data. Scientific Data , 8(1):284, 2021. doi: 10.1038/s41597- 021- 01060- 0. 144 [506] Y ang Deng, Ce W ang, Y uan Hui, Qian Li, Jun Li, Shiwei Luo, Mengke Sun, Quan Quan, Shuxin Y ang, Y ou Hao, Pengbo Liu, Honghu Xiao, Chunpeng Zhao, Xinbao W u, and 90 S. Ke vin Zhou. Ctspine1k: A large-scale dataset for spinal v ertebrae se gmentation in com- puted tomography . Machine Learning for Biomedical Ima ging (MELB A) , 2:1–23, 2025. doi: 10.59275/j.melba.2025- b3f2. 144 [507] Pengbo Liu, Hu Han, Y uanqi Du, Heqin Zhu, Y inhao Li, Feng Gu, Honghu Xiao, Jun Li, Chunpeng Zhao, Li Xiao, Xinbao W u, and S. Ke vin Zhou. Deep learning to segment pelvic bones: large-scale ct datasets and baseline models. International Journal of Computer As- sisted Radiology and Sur gery , 16(5):749–756, 2021. doi: 10.1007/s11548- 021- 02363- 8. 144 [508] Liang Jin, Jiancheng Y ang, Kaiming Kuang, Bingbing Ni, Y iyi Gao, Y ingli Sun, Pan Gao, W eiling Ma, Mingyu T an, Hui Kang, Jiajun Chen, and Ming Li. Deep-learning-assisted detection and se gmentation of rib fractures from ct scans: De velopment and validation of fracnet. EBioMedicine , 62:103106, 2020. doi: 10.1016/j.ebiom.2020.103106. 144 [509] Bennett A. Landman, Zhoubing Xu, Juan Eugenio Iglesias, Martin Styner, Thomas Robin Langerak, and Arno Klein. Miccai multi-atlas labeling beyond the cranial vault–w orkshop and challenge. In Pr oc. MICCAI Multi-Atlas Labeling Be yond the Cranial V ault W orkshop Challenge , v olume 5, page 12, 2015. doi: 10.7303/syn3193805. 144 [510] V al J Lo we, Fenghai Duan, Rathan M Subramaniam, JoRean D Sicks, Justin Romanof f, T wyla Bartel, Jian Q Michael Y u, Brian Nussenbaum, Jeremy Richmon, Charles D Arnold, David Cognetti, and Brendan C Stack Jr . Multicenter trial of [18f]fluorodeoxyglucose positron emission tomography/computed tomography staging of head and neck cancer and negati v e predictive v alue and surgical impact in the n0 neck: Results from acrin 6685. Jour - nal of Clinical Oncology , 37(20):1704–1712, 2019. doi: 10.1200/JCO.18.01182. 144 , 153 [511] P . Kinahan, M. Muzi, B. Bialecki, B. Herman, and L. Coombs. Data from the acrin 6668 trial nsclc-fdg-pet (version 2). The Cancer Imaging Archive, 2019. [Data set]. 144 , 153 [512] Lale Kostak oglu, Fenghai Duan, Michael O. Idowu, Paul R. Jolles, Harry D. Bear, Mark Muzi, Jean Cormack, John P . Muzi, Daniel A. Pryma, Jennifer M. Specht, Linda Hov anessian-Larsen, John Miliziano, Sharon Mallett, Anthony F . Shields, and David A. Mankof f. A phase ii study of 3’-deoxy-3’-18f-fluorothymidine pet in the assessment of early response of breast cancer to neoadjuvant chemotherap y: Results from acrin 6688. J ournal of Nuclear Medicine , 56(11):1681–1689, 2015. doi: 10.2967/jnumed.115.160663. 144 , 153 [513] P . Kinahan, M. Muzi, B. Bialecki, and L. Coombs. Data from acrin-fmiso-brain (version 2). The Cancer Imaging Archiv e, 2018. 144 , 153 [514] Children’ s Oncology Group. AREN0532. The Cancer Imaging Archive, 2022. URL https: //www.cancerimagingarchive.net/collection/aren0532/ . Clinical trial data for W ilms tumor treatment study . 144 , 152 , 153 [515] Mark Rozenfeld and Patricia Jordan. AREN0532-T umor-Annotations. The Cancer Imaging Archiv e, 2023. URL https://www.cancerimagingarchive.net/analysis- result/ aren0532- tumor- annotations/ . T umor annotations for AREN0532 W ilms tumor dataset. 144 [516] Children’ s Oncology Group. AREN0533. The Cancer Imaging Archive, 2022. URL https: //www.cancerimagingarchive.net/collection/aren0533/ . Clinical trial data for stage III/IV W ilms tumor treatment study . 144 , 152 [517] Mark Rozenfeld and Patricia Jordan. AREN0533-T umor-Annotations. The Cancer Imaging Archiv e, 2023. URL https://www.cancerimagingarchive.net/analysis- result/ aren0533- tumor- annotations/ . T umor annotations for AREN0533 W ilms tumor dataset. 144 , 153 [518] Children’ s Oncology Group. AREN0534. The Cancer Imaging Archive, 2021. URL https: //www.cancerimagingarchive.net/collection/aren0534/ . Clinical trial data for bi- lateral W ilms tumor treatment study . 144 , 153 91 [519] Kara M. Kelly , Paul D. Cole, Q. Pei, Rachel Bush, Keith B. Roberts, David C. Hodgson, Kristina M. McCarten, Si Y oung Cho, and Christopher Schwartz. Combination chemotherapy and radiation therapy in treating young patients with newly diagnosed hodgkin lymphoma (ahod0831) (version 1). Data set. The Cancer Imaging Archive, 2022. URL https://doi. org/10.7937/CV5M- 1H59 . 144 , 153 [520] M. Rozenfeld and P . Jordan. Annotations for vincristine, dactinomycin, and doxorubicin with or without radiation therap y or observ ation only in treating younger patients who are undergoing sur gery for newly diagnosed stage i, ii, or iii wilms’ tumor (aren0532-tumor- annotations) [data set]. The Cancer Imaging Archive, 2023. URL https://doi.org/10. 7937/KJA4- 1Z76 . 144 , 147 , 153 , 154 [521] T atiana Bejarano, Mariluz De Ornelas-Couto, and Iv aylo B. Mihaylov . Head-and-neck squa- mous cell carcinoma patients with ct taken during pre-treatment, mid-treatment, and post- treatment (hnscc-3dct-rt). Data set, 2018. The Cancer Imaging Archi v e. 144 [522] A. Grossberg, A. Mohamed, H. Elhalawani, W . Bennett, K. Smith, T . Nolan, B. W illiams, S. Chamchod, J. Heukelom, A. Kanwar , T . Browne, K. Hutcheson, G. Gunn, A. Garden, W . Morrison, S. Frank, D. Rosenthal, J. Freymann, and C. Fuller . Imaging and clinical data archiv e for head and neck squamous cell carcinoma patients treated with radiotherap y . Sci- entific Data , 5:180173, 2018. doi: 10.1038/sdata.2018.173. 144 , 153 [523] K yle Smith, K e vin Clark, W illiam Bennett, Thomas Nolan, John Kirby , Michael W olfsberger , James Moulton, Brian V endt, and John Freymann. Data from ct colonography . The Cancer Imaging Archiv e, 2015. URL https://www.cancerimagingarchive.net/collection/ ct- colonography/ . 144 [524] Holger R. Roth, Le Lu, Ari Seff, Ke vin M. Cherry , Joanne Hoffman, Shijun W ang, Jiamin Liu, Evrim T urkbey , and Ronald M. Summers. A new 2.5d representation for lymph node detection using random sets of deep conv olutional neural network observations. In Medical Image Computing and Computer-Assisted Intervention – MICCAI 2014 , volume 8673, pages 520–527, 2014. doi: 10.1007/978- 3- 319- 10404- 1_65. 144 [525] Dennis Mackin, Xenia Ray , Lifei Zhang, Da vid Fried, Jinzhong Y ang, Brian T aylor , Edgardo Rodriguez-Riv era, Cristina Dodge, Aaron K yle Jones, and Laurence Court. Data from cre- dence cartridge radiomics phantom ct scans (cc-radiomics-phantom). The Cancer Imaging Archiv e, 2017. 144 [526] Muhammad Shafiq ul Hassan, Geoffre y Zhang, Kujtim Latifi, Ghanim Ullah, Robert Gillies, and Eduardo G. Moros. Computed tomography texture phantom dataset for ev aluating the impact of ct imaging parameters on radiomic features. Scientific Reports , 9:1–10, 2019. doi: 10.1038/s41598- 019- 45349- 9. 144 [527] Rachel Ger , Shouhao Zhou, Pai-Chun Chi, Hannah Lee, Rick Layman, K yle Jones, David Goff, Clifton D. Fuller, Rebecca M. Howell, Heng Li, R. Jason Stafford, Laurence E. Court, and Dennis Mackin. Data from ct phantom scans for head, chest, and con- trolled protocols on 100 scanners (cc-radiomics-phantom-3). Data set. The Cancer Imag- ing Archiv e, 2019. URL https://www.cancerimagingarchive.net/collection/ cc- radiomics- phantom- 3/ . 144 [528] N. Mayr, W . T . C. Y uh, S. Bowen, M. Harkenrider , M. V . Knopp, E. Y .-P . Lee, E. Leung, S. S. Lo, W . Small Jr ., and A. H. W olfson. Cervical cancer – tumor heterogeneity: Serial functional and molecular imaging across the radiation therapy course in advanced cervical cancer (version 1). Data set. The Cancer Imaging Archive, 2023. URL https://www. cancerimagingarchive.net/collection/cc- tumor- heterogeneity/ . 145 , 153 [529] S. Kirk, Y . Lee, F . R. Lucchesi, N. D. Aredes, N. Gruszauskas, J. Catto, K. Garcia, R. Jarosz, V . Duddalwar , B. V arghese, K. Rieger-Christ, and J. Lemmerman. The cancer genome atlas urothelial bladder carcinoma collection (tcga-blca) (version 8) [data set]. The Cancer Imaging Archiv e, 2016. URL https://doi.org/10.7937/K9/TCIA.2016.8LNG8XDR . 145 92 [530] S. Kirk, Y . Lee, F . R. Lucchesi, N. D. Aredes, N. Gruszauskas, J. Catto, K. Garcia, R. Jarosz, V . Duddalwar , B. V arghese, K. Rieger-Christ, and J. Lemmerman. The cancer genome atlas colon adenocarcinoma collection (tcga-coad) (version 8) [data set]. The Cancer Imaging Archiv e, 2016. URL https://www.cancerimagingarchive.net/collection/ tcga- coad/ . 145 [531] The cancer genome atlas esophageal carcinoma collection (tcga-esca). The Cancer Imaging Archiv e, 2016. V ersion 3. 145 , 150 [532] Lisa Scarpace, T om Mikkelsen, Soonmee Cha, Sudhir Rao, Sandeep T ekchandani, David Gutman, Joel H. Saltz, Bradley J. Erickson, Nick Pedano, Adam E. Flanders, Jill Barnholtz- Sloan, Quinn Ostrom, Daniel Barboriak, and Lori J. Pierce. The cancer genome atlas glioblas- toma multiforme collection (tcga-gbm) (version 5) [data set]. The Cancer Imaging Archive, 2016. URL https://www.cancerimagingarchive.net/collection/tcga- gbm/ . 145 [533] M. L. Zuley , R. Jarosz, S. Kirk, Y . Lee, R. Colen, K. Garcia, D. Delbeke, M. Pham, P . Nagy , G. Sevinc, M. Goldsmith, S. Khan, J. M. Net, F . R. Lucchesi, and N. D. Aredes. The cancer genome atlas head-neck squamous cell carcinoma collection (tcga-hnsc) (version 6) [data set]. The Cancer Imaging Archive, 2016. URL https://www.cancerimagingarchive. net/collection/tcga- hnsc/ . 145 , 153 [534] M. Linehan, R. Gautam, S. Kirk, Y . Lee, C. Roche, E. Bonaccio, J. Filippini, K. Rieger-Christ, J. Lemmerman, and R. Jarosz. The cancer genome atlas cervical kidne y renal papillary cell carcinoma collection (tcga-kirp) (version 4) [data set]. The Cancer Imaging Archiv e, 2016. URL https://www.cancerimagingarchive.net/collection/tcga- kirp/ . 145 , 153 [535] O. Akin, P . Elnajjar, M. Heller , R. Jarosz, B. J. Erickson, S. Kirk, Y . Lee, M. W . Linehan, R. Gautam, R. V ikram, K. M. Garcia, C. Roche, E. Bonaccio, and J. Filippini. The can- cer genome atlas kidney renal clear cell carcinoma collection (tcga-kirc) (version 3). Data set. The Cancer Imaging Archi ve. https://doi.org/10.7937/K9/TCIA.2016.V6PBVTDR, 2016. Accessed 2020-05-29. 145 [536] A. B. Shinagare, R. V ikram, C. Jaf fe, O. Akin, J. Kirby , E. Huang, J. Freymann, N. I. Sainani, C. A. Sado w , T . K. Bathala, D. L. Rubin, A. Oto, M. T . Heller , V . R. Surabhi, V . Katabathina, and S. G. Silverman. Radiogenomics of clear cell renal cell carcinoma: preliminary findings of the cancer genome atlas–renal cell carcinoma (tcga–rcc) imaging research group. Abdom- inal Imaging , 40(6):1684–1692, 2015. doi: 10.1007/s00261- 015- 0386- z. 145 [537] Spyridon Bakas, Hamed Akbari, Aristeidis Sotiras, Michel Bilello, Martin Rozycki, Justin S. Kirby , John B. Freymann, K eyv an Farahani, Christos Dav atzikos, Justin Kirby , Y uliya Bur - ren, Nicole Porz, Johannes Slotboom, Roland W iest, Lev ente Lanczi, Elisabeth Gerstner , Marc-Andre W eber , T al Arbel, Brian B. A vants, Nicholas A yache, Patricia Buendia, D. Louis Collins, Nicolas Cordier , Jason J. Corso, Antonio Criminisi, Tilak Das, Herve Delingette, Cagatay Demiralp, Christopher R. Durst, Michel Dojat, Senan Doyle, Joana Festa, Florence Forbes, Ezequiel Geremia, Ben Glocker , Polina Golland, Xiaotao Guo, Andac Hamamci, Khan Iftekharuddin, Raj Jena, Nigel John, Ender K onukoglu, Danial Lashkari, Jose A. Mariz, Raphael Meier, Sergio Pereira, Doina Precup, S. J. Price, T ammy Riklin-Ra viv , Syed M. S. Reza, Michael Ryan, Lawrence Schwartz, Hoo-Chang Shin, Jamie Shotton, Carlos Silv a, Nuno Sousa, Nagesh Subbanna, Gabor Szek ely , Thomas T aylor , Owen Thomas, Nicholas T ustison, Gozde Unal, Flor V asseur , Max Wintermark, Dong Hye Y e, Liang Zhao, Binsheng Zhao, Darko Zikic, Marcel Prastawa, Mauricio Reyes, and Koen van Leemput. Advancing the cancer genome atlas glioma mri collections with expert se gmentation labels and radiomic features. Scientific Data , 4:170117, 2017. doi: 10.1038/sdata.2017.117. 145 , 149 , 150 [538] B. J. Erickson, S. Kirk, Y . Lee, O. Bathe, M. Kearns, C. Gerdes, K. Rieger-Christ, and J. Lem- merman. The cancer genome atlas li ver hepatocellular carcinoma collection (tcg a-lihc). Data set. The Cancer Imaging Archi ve, V ersion 5, 2016. URL https://doi.org/10.7937/K9/ TCIA.2016.IMMQW8UQ . 145 , 153 , 154 [539] B. Albertina, M. W atson, C. Holback, R. Jarosz, S. Kirk, Y . Lee, K. Rieger -Christ, and J. Lem- merman. The cancer genome atlas lung adenocarcinoma collection (tcga-luad) (version 4). Data set. The Cancer Imaging Archiv e. https://doi.or g/10.7937/K9/TCIA.2016.JGNIHEP5, 2016. 145 93 [540] S. Kirk, Y . Lee, F . R. Lucchesi, N. D. Aredes, N. Gruszauskas, J. Catto, K. Garcia, R. Jarosz, V . Duddalwar , B. V arghese, K. Rie ger-Christ, and J. Lemmerman. The cancer genome at- las lung squamous cell carcinoma collection (tcga-lusc) (version 8) [data set]. The Cancer Imaging Archiv e, 2016. URL https://www.cancerimagingarchive.net/collection/ tcga- lusc/ . 145 [541] C. Holback, R. Jarosz, F . Prior , D. G. Mutch, P . Bhosale, K. Garcia, Y . Lee, S. Kirk, C. A. Sadow , S. Le vine, E. Sala, P . Elnajjar, T . Morg an, and B. J. Erickson. The cancer genome atlas ov arian cancer collection (tcga-ov) (version 4). The Cancer Imaging Archi v e, 2016. URL https://www.cancerimagingarchive.net/collection/tcga- ov/ . [Data set]. 145 [542] Hebert Alberto V arg as, Erich P Huang, Y ulia Lakhman, Joseph E Ippolito, Priya Bhosale, V incent Mellnick, Atul B Shinagare, Maria Anello, Justin Kirby , Brenda Fevrier -Sulli v an, John Freymann, C Carl Jaffe, and Evis Sala. Radiogenomics of high-grade serous o varian cancer: Multireader multi-institutional study from the cancer genome atlas ovarian cancer imaging research group. Radiology , 285(2):482–492, 2017. doi: 10.1148/radiol.2017161870. 145 [543] Lucian Beer , Huseyin Sahin, Ivana Blazic, Hebert A. V argas, Harini V eeraraghav an, Justin Kirby , Brigitte Fevrier -Sulli van, John B. Freymann, Carl C. Jaffe, Thomas P . Conrads, George L. Maxwell, Kristine M. Darcy , Eric Huang, and Evis Sala. Data from integration of ct-based qualitativ e and radiomic features with proteomic variables in patients with high- grade serous ov arian cancer: An exploratory analysis (tcga-ov-proteogenomics). The Cancer Imaging Archiv e, 2019. 145 [544] S. Kirk, Y . Lee, F . R. Lucchesi, N. D. Aredes, N. Gruszauskas, J. Catto, K. Garcia, R. Jarosz, V . Duddalwar , B. V arghese, K. Rieger -Christ, and J. Lemmerman. The cancer genome atlas rectum adenocarcinoma collection (tcga-read) (version 8) [data set]. The Cancer Imaging Archiv e, 2016. URL https://www.cancerimagingarchive.net/collection/ tcga- read/ . 145 [545] C. Roche, E. Bonaccio, and J. Filippini. The cancer genome atlas sarcoma collection (tcga-sarc) (version 3). Data set, The Cancer Imaging Archive. https://doi.org/10.7937/K9/TCIA.2016.CX6YLSUX, 2016. 145 [546] National Cancer Institute Clinical Proteomic Tumor Analysis Consortium (CP- T AC). The clinical proteomic tumor analysis consortium lung adenocarcinoma collection (cptac-luad) (version 15). The Cancer Imaging Archive. A v ailable: https://www .cancerimagingarchiv e.net/collection/cptac-luad/, 2018. [dataset]. 145 , 154 [547] National Cancer Institute Clinical Proteomic T umor Analysis Consortium (CPT AC). The clinical proteomic tumor analysis consortium pancreatic ductal adenocarcinoma collection (cptac-pda) (version 15). The Cancer Imaging Archive. A v ailable: https://www .cancerimagingarchiv e.net/collection/cptac-pda/, 2018. [dataset]. 145 , 153 , 154 [548] National Cancer Institute Clinical Proteomic T umor Analysis Consortium (CPT A C). The clinical proteomic tumor analysis consortium uterine corpus endometrial carcinoma col- lection (cptac-ucec) (version 13). The Cancer Imaging Archi ve, 2019. URL https: //doi.org/10.7937/k9/tcia.2018.3r3juisw . [dataset]. 145 , 153 , 154 [549] Hugo J.W .L. Aerts, Emmanuel R. V elazquez, Robbert T .H. Leijenaar , Chintan Parmar , Patri- cia Grossmann, Sara Carvalho, Jonas Bussink, Renske Monshouwer, Benjamin Haibe-Kains, Daniël Rietveld, Frank Hoebers, Marieke M. Rietbergen, Chris R. Leemans, Andre Dekker , John Quackenb ush, E.R. Gillies, and Philippe Lambin. Decoding tumour phenotype by non- in v asi ve imaging using a quantitati ve radiomics approach. Natur e Communications , 5(4006): 4006, 2014. doi: 10.1038/ncomms5006. 145 , 146 , 154 [550] Sameh Bakr , Olivier Ge vaert, Santiago Echegaray , Ke vin A yers, Min Zhou, Mahshid Shafiq, Hong Zheng, Jonathan A. Benson, W eidong Zhang, Ann Leung, Michael Kadoch, Christo- pher D. Hoang, Joseph B. Shrager , Andre w Quon, Daniel L. Rubin, Sylvia K. Ple vritis, and 94 Sandy Napel. A radiogenomic dataset of non-small cell lung cancer . Scientific Data , 5: 180202, 2018. doi: 10.1038/sdata.2018.202. 145 , 154 [551] Jef frey Bradley and Ken Forster . Data from nsclc-cetuximab. The Cancer Imag- ing Archiv e, 2018. URL https://www.cancerimagingarchive.net/collection/ nsclc- cetuximab/ . 145 [552] Leonard W ee, Hugo J.L. Aerts, Petros Kalendralis, and Andre Dekker . Data from nsclc- radiomics-interobserver1. The Cancer Imaging Archive, 2019. URL https://www. cancerimagingarchive.net/collection/nsclc- radiomics- interobserver1/ . 145 [553] Cancer Moonshot Biobank. Cancer moonshot biobank – gastroesophageal cancer collection (cmb-gec). V ersion 6, The Cancer Imaging Archiv e, dataset, 2022. URL https://www. cancerimagingarchive.net/collection/cmb- gec/ . [dataset]. 145 , 153 , 154 [554] J. Kalpathy-Cramer , A. Beers, A. Mamonov , E. Ziegler , R. Lewis, A. B. Almeida, G. Harris, S. Pieper , A. Sharma, L. T arbox, J. T obler , F . Prior , A. Flanders, J. Dulk owski, B. Fe vrier- Sulliv an, C. Jaf fe, J. Freymann, and J. Kirby . Crowds cure cancer: Crowdsourced data collected at the rsna 2017 annual meeting. The Cancer Imaging Archi ve, 2019. URL https://doi.org/10.7937/K9/TCIA.2018.OW73VLO2 . [Data set]. 145 [555] T . Urban, E. Ziegler , S. Pieper , J. Kirby , D. Rukas, B. Beardmore, B. Somarouthu, E. Ozkan, G. Lelis, B. Fe vrier -Sulli van, S. Nandekar , A. Beers, C. Jaf fe, J. Freymann, D. Clunie, G. J. Harris, and J. Kalpathy-Cramer . Cro wds cure cancer: Crowdsourced data collected at the rsna 2018 annual meeting. Data set, The Cancer Imaging Archiv e, 2019. URL https: //www.cancerimagingarchive.net/analysis- result/crowds- cure- 2018/ . 145 [556] Xia Li, Richard G. Abramson, Lori R. Arlinghaus, Hakmook Kang, Anuradha B. Chakrav arthy , V andana G. Abramson, Jaime Farle y , Ingrid A. Mayer , Mark C. K elley , In- grid M. Meszoely , Julie Means-Po well, Ana M. Grau, Melinda Sanders, and Thomas E. Y an- keelov . Multiparametric magnetic resonance imaging for predicting pathological response after the first cycle of neoadjuvant chemotherapy in breast cancer . In vestigative Radiology , 50(4):195–204, 2015. doi: 10.1097/RLI.0000000000000100. 145 , 154 [557] Andriy Fedorov , David Clunie, Erik J. Ulrich, Christian Bauer , Andreas W ahle, Bradley Brown, Mark Onken, Joerg Riesmeier, Stefan Pieper , Ron Kikinis, Joseph Buatti, and Rein- hard R. Beichel. Dicom for quantitative imaging biomarker dev elopment: a standards based approach to sharing clinical data and structured pet/ct analysis results in head and neck cancer research. P eerJ , 4:e2057, 2016. doi: 10.7717/peerj.2057. 146 , 154 [558] D. Goldgof, L. Hall, S. Hawkins, M. Schabath, O. Stringfield, A. Garcia, Y . Balagurunathan, J. Kim, S. Eschrich, A. Ber glund, R. Gatenby , and R. Gillies. Data from qin lung ct (ver - sion 2). The Cancer Imaging Archi ve, 2015. URL https://doi.org/10.7937/K9/TCIA. 2015.NPGZYZBZ . Data set. 146 [559] Jayashree Kalpathy-Cramer , Sandy Napel, Dmitry Goldgof, and Binsheng Zhao. Multi-site collection of lung ct data with nodule se gmentations (v ersion 3) [data set]. The Cancer Imag- ing Archiv e. DOI:10.7937/k9/tcia.2015.1b uvfjr7, 2015. 146 [560] Charles Fenimore, Michael F . McNitt-Gray , Da vid Clunie, Marios A. Ga vrielides, Nicholas Petrick, Ehsan Samei, B. Chen, G. Saiprasad, K. Jen-Sho Chen, K. Boedeker , H. Chen-Mayer , J. Barudin, B. Beute, K. Byrne, G. Edeb urn, S. Kaplan, J. Sherman, and K. Slazak. Data from qiba ct-1c (version 1). The Cancer Imaging Archive [Data set], 2016. 146 [561] Marilyn F . McNitt-Gray , Hyun G. Kim, Bin Zhao, Lawrence H. Schwartz, Da vid A. Clu- nie, K enneth Cohen, Nicholas Petrick, Charles Fenimore, Zheng Q. J. Lu, and Anthony J. Buckler . Qiba volct group 1b round 2 no change size measurements (qiba-volct-1b) [data set]. The Cancer Imaging Archi ve. https://www.cancerimagingarchive.net/ analysis- result/qiba- volct- 1b/ , 2020. 146 [562] B. Zhao, Q. Li, Y . Liang, H. Y ang, M. A. Ga vrielides, L. H. Schwartz, D. C. Sulli van, and N. A. Petrick. Qiba anthropomorphic abdominal phantom ct scans. Data set, The Cancer Imaging Archiv e, 2021. DOI:10.7937/TCIA.RMV0-9Y95. 146 95 [563] Y unliang Cai, Said Osman, Manas Sharma, Mark Landis, and Shuo Li. Multi-modality verte- bra recognition in arbitrary views using 3d deformable hierarchical model. IEEE T ransactions on Medical Imaging , 34(8):1676–1693, 2015. doi: 10.1109/TMI.2015.2392054. 146 [564] Robert Korez, Bulat Ibragimov , Boštjan Likar, Franjo Pernuš, and T omaž Vrtovec. A framew ork for automated spine and vertebrae interpolation-based detection and model-based segmentation. IEEE T r ansactions on Medical Imaging , 34(8):1649–1662, 2015. doi: 10.1109/TMI.2015.2389334. 146 [565] T omaž Vrtovec, Jianhua Y ao, Ben Glocker , T obias Klinder , Alejandro Frangi, Guoyan Zheng, and Shuo Li. Computational methods and clinical applications for spine imaging: Third in- ternational w orkshop and challenge, csi 2015, held in conjunction with miccai 2015, munich, germany , october 5, 2015, proceedings. In Lectur e Notes in Computer Science , v olume 9402. Springer , 2016. doi: 10.1007/978- 3- 319- 41827- 8. 146 [566] E. I. Hwang, M. Kool, P . C. Burger , D. Capper , L. Chavez, S. Brabetz, C. Williams-Hughes, C. Billups, L. Heier , A. Jaju, J. Michalski, Y . Li, S. Leary , T . Zhou, A. v on Deimling, D. T . W . Jones, M. Fouladi, I. F . Pollack, A. Gajjar , and J. M. Olson. Extensi ve molecular and clinical heterogeneity in patients with histologically diagnosed cns-pnet treated as a single entity: A report from the children’ s oncology group randomized acns0332 trial. J ournal of Clinical Oncology , 36(34):3388–3395, 2018. doi: 10.1200/jco.2017.76.4720. 146 [567] Henri M. Katzenstein, Mary R. Langham, Mark H. Malogolo wkin, Mark D. Krailo, An- drew J. T owbin, Mark B. McCarville, Milton J. Finegold, Sw ati Rang anathan, Stephan Dunn, Emily D. McGahren, George M. Tiao, Ann F . O’Neill, Mohammed Qayed, W illiam L. Fur- man, Caroline Xia, Carlos Rodriguez-Galindo, and Rebecca L. Meyers. Risk-based ther- apy in treating younger patients with ne wly diagnosed li ver cancer (ahep0731) (version 2). The Cancer Imaging Archiv e, 2021. URL https://www.cancerimagingarchive.net/ collection/ahep0731/ . 146 , 153 , 154 [568] Madhavi P atnana, Sapna Patel, and Anne S. Tsao. Data from anti-pd-1 immunotherapy lung [data set]. The Cancer Imaging Archi ve, 2019. 146 , 154 [569] M. P atnana, S. Patel, and A. Tsao. Anti-pd-1 immunotherap y melanoma dataset. Data set. The Cancer Imaging Archiv e. DOI:10.7937/tcia.2019.1ae0qtcu, 2019. 146 , 154 [570] B. Nicolas Bloch, Ashali Jain, and C. Carl Jaf fe. Breast-diagnosis [data set]. The Cancer Imaging Archiv e, 2015. URL https://www.cancerimagingarchive.net/collection/ breast- diagnosis/ . 146 , 154 [571] Nancy L. Bartlett, W yndham H. W ilson, Sin-Ho Jung, Eric D. Hsi, Matthew J. Maurer , Lisa D. Pederson, Marie-Yv onne C. Polley , Brian N. Pitcher , Bruce D. Cheson, Brad S. Kahl, Jonathan W . Friedberg, Louis M. Staudt, Nina D. W agner-Johnston, Kristie A. Blum, Jeremy S. Abramson, Nishitha M. Reddy , Jane N. W inter , Joanne E. Chang, Ajay K. Gopal, Amy Chadburn, Songya Mathew , Richard I. Fisher , Kristie Richards, Heiko Schloder , An- drew D. Zelenetz, and John P . Leonard. Rituximab and combination chemotherapy in treat- ing patients with dif fuse large b-cell non-hodgkin’ s lymphoma (calgb50303) (version 2). The Cancer Imaging Archiv e, 2020. URL https://www.cancerimagingarchive.net/ collection/calgb50303/ . 146 , 154 [572] Anthony P . Reev es, Anna M. Biancardi, David Y ankele vitz, Serge y Fotin, Brian M. Keller , Ajit Jirapatnakul, and Jae Lee. A public image database to support research in computer aided diagnosis. In 31st Annual International Confer ence of the IEEE Engineering in Medicine and Biology Society , pages 3715–3718. IEEE, 2009. doi: 10.1109/IEMBS.2009.5334807. 146 [573] Nadya Shusharina, Thomas Bortfeld, Carlos Cardenas, Bastien De, Ke vin Diao, Sandra Her- nandez, Y iwen Liu, Sarin Maroongroge, Jonas Söderberg, and Mohamed Soliman. Cross- modality brain structures image se gmentation for the radiotherapy target definition and plan optimization. In Se gmentation, Classification, and Re gistration of Multi-modality Medical Imaging Data , volume 12587 of Lectur e Notes in Computer Science , pages 3–15, 2021. doi: 10.1007/978- 3- 030- 71827- 5_1. 146 96 [574] Nicolas Gerber , Mauricio Reyes, Livia Barazzetti, Hans Martin Kjer , Sergio V era, Martin Stauber , Pav el Mistrik, Mario Ceresa, Nerea Mangado, W ilhelm W immer, Thomas Stark, Rasmus R. P aulsen, Stefan W eber, Marco Ca versaccio, and Miguel A. González Ballester . A multiscale imaging and modelling dataset of the human inner ear . Scientific Data , 4(1): 170132, 2017. doi: 10.1038/sdata.2017.132. URL https://doi.org/10.1038/sdata. 2017.132 . 146 [575] Ki Ang, Qiang Zhang, David I Rosenthal, P atricia F Nguyen-T an, Elliot J Sherman, Robert S W eber , Jef fre y M Galvin, John A Bonner , James Harris, Adel K El-Naggar , Maura L Gilli- son, Robert C Jordan, Alexander A K onski, W aldemar L Thorstad, Anthony Trotti, Jeremy J Beitler , Adam S Garden, W illiam J Spanos, Stanley S Y om, and Randall S Axelrod. Random- ized phase iii trial of concurrent accelerated radiation plus cisplatin with or without cetuximab for stage iii to i v head and neck carcinoma: Rtog 0522. J ournal of Clinical Oncology , 32(27): 2940–2950, 2014. doi: 10.1200/jco.2013.53.5633. 146 , 154 [576] Martin V allières, Emily Kay-Riv est, Léo Jean Perrin, Xavier Liem, Christophe Furstoss, Hugo J. W . L. Aerts, Nader Khaouam, Phuc Félix Nguyen-T an, Chang-Shu W ang, Khalil Sultanem, Johan Seuntjens, and Imad El Naqa. Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer . Scientific Reports , 7(1):10117, 2017. doi: 10.1038/s41598- 017- 10371- 5. 146 , 154 [577] Jinzhong Y ang, Harini V eeraraghav an, Samuel G. III Armato, K eyv an Farahani, Justin S. Kirby , Jayashree Kalpathy-Kramer , W outer v an Elmpt, Andre Dekker , Xiao Han, Xue Feng, Paul Aljabar , Bruno Oliv eira, Brent van der Heyden, Leonid Zamdborg, Dao Lam, Mark Gooding, and Gregory C. Sharp. Autosegmentation for thoracic radiation treatment planning: A grand challenge at aapm 2017. Medical Physics , 45(10):4568–4581, 2018. doi: 10.1002/ mp.13141. 146 [578] T aylor R. Moen, Baiyu Chen, Da vid R. III Holmes, Xinhui Duan, Zhicong Y u, Lifeng Y u, Shuai Leng, Joel G. Fletcher , and Cynthia H. McCollough. Low-dose ct image and projection dataset. Medical Physics , 48(2):902–911, 2021. doi: 10.1002/mp.14594. 146 [579] Samuel G. Armato III, Geof frey McLennan, Luc Bidaut, Michael F . McNitt-Gray , Charles R. Meyer , Anthony P . Reeves, Binsheng Zhao, Denise R. Aberle, Claudia I. Henschk e, Eric A. Hoffman, Ella A. Kazerooni, Heber MacMahon, Edwin J. R. V an Beeke, David Y ankelevitz, Alberto M. Biancardi, Peyton H. Bland, Matthew S. Bro wn, Roger M. Engelmann, Gary E. Laderach, Daniel Max, Richard C. Pais, David P . Y . Qing, Rachael Y . Roberts, Amanda R. Smith, Adam Starke y , Poonam Batrah, Philip Caligiuri, Ali Farooqi, Gregory W . Gladish, C. Matilda Jude, Reginald F . Munden, Iva Petkovska, Leslie E. Quint, La wrence H. Schwartz, Baskaran Sundaram, Lori E. Dodd, Charles Fenimore, David Gur , Nicholas Petrick, John Freymann, Justin Kirby , Brian Hughes, Alessi V ande Casteele, Sangeeta Gupte, Maha Sal- lamm, Michael D. Heath, Michael H. Kuhn, Ekta Dharaiya, Richard Burns, Da vid S. Fryd, Marcos Salganicof f, V ikram Anand, Uri Shreter , Stephen V astagh, and Barbara Y . Croft. The lung image database consortium (lidc) and image database resource initiati ve (idri): a com- pleted reference database of lung nodules on ct scans. Medical Physics , 38(2):915–931, 2011. doi: 10.1118/1.3528204. 146 [580] Peijun Li, Sheng W ang, T ao Li, Jichao Lu, Y uhan Huangfu, and Dong W ang. A lar ge-scale ct and pet/ct dataset for lung cancer diagnosis (lung-pet-ct-dx). The Cancer Imaging Archi v e, 2020. [Data set]. 146 , 154 [581] Olya Grov e, Anders E Berglund, Matthew B Schabath, Hugo JWL Aerts, Andre Dekker , Hua W ang, Emmanuel R V elazquez, Philippe Lambin, Y uhua Gu, Y oganand Balagurunathan, Edward Eikman, Robert A Gatenby , Ste ven Eschrich, and Robert J Gillies. Quantitativ e computed tomographic descriptors associate tumor shape complexity and intratumor hetero- geneity with prognosis in lung adenocarcinoma. PLOS One , 10(3):e0118261, 2015. doi: 10.1371/journal.pone.0118261. 146 [582] Jie Su, Jennifer Y in Y ee Kwan, Shao Hui Huang, Laleh S. Ghoraie, W ei Xu, Biu Chan, Kwok W ong Y ip, Meredith Giuliani, Andre w Bayley , John Kim, Andrew John Hope, Jolie Ringash, John Cho, Andrea McNiv en, Aaron Hansen, Da vid Goldstein, Jose Rui de Almeida, Hugo J. W . L. Aerts, John N. W aldron, Benjamin Haibe-Kains, Brian O’Sulli van, Scott V . 97 Bratman, and Fei-Fei Liu. Radiomic biomarkers to refine risk models for distant metastasis in hpv-related oropharyngeal carcinoma. International Journal of Radiation Oncology Biology Physics , 102:1107–1116, 2018. doi: 10.1016/j.ijrobp.2018.01.057. 146 [583] K enneth Marek, Sohini Cho wdhury , Andre w Siderowf, Shirley Lasch, Christopher S Coffe y , Chelsea Caspell-Garcia, T anya Simuni, Danna Jennings, Caroline M T anner, John Q T ro- janowski, Leslie M Shaw , John Seibyl, Norbert Schuff, Andre w Singleton, Karl Kieb urtz, Arthur W T oga, Brit Mollenhauer , Doug Galask o, Lana M Chahine, Daniel W eintraub, T a- tiana Foroud, Duygu T osun-T urgut, Kathleen Poston, V anessa Arnedo, Mark Frasier, and T odd Sherer . The parkinson’ s progression markers initiativ e (ppmi) – establishing a pd biomarker cohort. Annals of Clinical and T r anslational Neurolo gy , 5(12):1460–1477, 2018. doi: 10.1002/acn3.644. 146 , 154 [584] Petr Jordan, Philip M. Adamson, Vrunda Bhattbhatt, Surabhi Beriwal, Sangyu Shen, Oskar Radermecker , Supratik Bose, Linda S. Strain, Michael Offe, David Fraley , Sara Principi, Dong Hye Y e, Adam S. W ang, John V an Heteren, Nghia-Jack V o, and T aly Gilat Schmidt. Pediatric chest-abdomen-pelvis and abdomen-pelvis ct images with expert organ contours. Medical Physics , 49(5):3523–3528, 2022. doi: 10.1002/mp.15485. 146 [585] Paul LaT our . Quantitativ e imaging data warehouse supports research needs. RSN A News, Radiological Society of North America, 2015. A v ailable at: https://www .rsna.org/ne ws/2015/july/quantitativ e-imaging-data-warehouse. 146 , 153 , 154 [586] Michael Kistler , Serena Bonaretti, Marcel Pfahrer , Roman Niklaus, and Philippe Büchler . The virtual sk eleton database: An open access repository for biomedical research and collab- oration. Journal of Medical Internet Researc h , 15(11):e245, 2013. doi: 10.2196/jmir .2930. 146 [587] Samuel G. Armato III, Lubomir Hadjiiski, Georgia D. T ourassi, Karen Drukker , Maryellen L. Giger , Feng Li, Geor ge Redmond, K e yvan Farahani, Justin S. Kirby , and Laurence P . Clarke. Spie-aapm-nci lung nodule classification challenge dataset, 2015. URL https://www. cancerimagingarchive.net/collection/spie- aapm- lung- ct- challenge/ . 146 [588] Jonathan Shapey , Aaron Kujaw a, Reuben Dorent, Guotai W ang, Andreas Dimitriadis, Dmitrii Grishchuk, Ian P addick, Neil Kitchen, Robert Bradford, Shakeel R. Saeed, Sotirios Bisdas, Sebastien Ourselin, and T om V ercauteren. Segmentation of vestib ular schwannoma from mri, an open annotated dataset and baseline algorithm. Scientific Data , 8(1), 2021. doi: 10.1038/s41597- 021- 01064- w. 146 [589] Xiahai Zhuang, Lei Li, Christian Payer , Darko Štern, Martin Urschler, Mattias P . Heinrich, Julien Oster , Chunliang W ang, Örjan Smedby , Cheng Bian, Xin Y ang, Pheng-Ann Heng, Aliasghar Mortazi, Ulas Bagci, Guanyu Y ang, Chenchen Sun, Gaetan Galisot, Jean Yves Ramel, Thierry Brouard, Qianqian T ong, W eixin Si, Xiangyun Liao, Guodong Zeng, Zenglin Shi, Guo yan Zheng, Chengjia W ang, T om MacGillivray , Da vid Newby , Ka wal Rhode, Se- bastien Ourselin, Raad Mohiaddin, Jennifer Keegan, Da vid Firmin, and Guang Y ang. Ev alu- ation of algorithms for multi-modality whole heart segmentation: an open-access grand chal- lenge. Medical Image Analysis , 58:101537, 2019. doi: 10.1016/j.media.2019.101537. 146 , 148 [590] K yle Smith, K e vin Clark, W illiam Bennett, Thomas Nolan, John Kirby , Michael W olfsberger , James Moulton, Brian V endt, and John Freymann. Data from soft-tissue-sarcoma. The Cancer Imaging Archiv e, 2015. URL https://www.cancerimagingarchive.net/collection/ soft- tissue- sarcoma/ . 146 , 154 [591] T ianchi Platform. Segmenting soft tissue sarcomas. Alibaba Tianchi Competition Platform, 2021. URL https://tianchi.aliyun.com/dataset/dataDetail?dataId=89694 . Preprocessed subset of TCIA Soft-tissue-Sarcoma data con verted to 3D HDF5 arrays. 146 , 154 [592] Rashed Karim, Lauren E. Blake, Jun Inoue, Qian T ao, Siyuan Jia, Robert J. Housden, Prasanna Bhagirath, Jeffre y L. Duval, Máximo V arela, Javier M. Behar , Laurent Cadour , 98 Rob J. v an der Geest, Helene Cochet, Maria Drangov a, Martine Sermesant, Reza Razavi, Oleg Aslanidi, Rajiv Rajani, and Karl Rhode. Algorithms for left atrial wall segmen- tation and thickness – evaluation on an open-source ct and mri image database. Med- ical Image Analysis , 50:36–53, 2018. doi: 10.1016/j.media.2018.08.004. URL http: //stacom.cardiacatlas.org . Creati ve Commons Attrib ution 4.0 License. 146 [593] Shuang Song, Rui Xu, Y ong Luo, Bo Du, Zhijian Y ang, Jiancheng Y ang, Kaiming Kuang, Bingbing Ni, Chang Chen, Deping Zhao, Dong Xie, Xiwen Sun, Jingyun Shi, Y unlang She, Mengmeng Zhao, Jiajun Deng, Junqi W u, and Tingting W ang. Mediastinal lesion analysis. Zenodo: https://doi.or g/10.5281/zenodo.6361949, 2022. 146 [594] Errol Colak, Felipe C. Kitamura, Stephen B. Hobbs, Carol C. Wu, Matthew P . Lungren, Lu- ciano M. Pre vedello, Jayashree Kalpath y-Cramer , Robyn L. Ball, Geor ge Shih, Anouk Stein, Safwan S. Halabi, Emre Altinmakas, Meng Law , P arveen Kumar , Karam A. Manzalawi, Den- nis Charles Nelson Rubio, Jacob W . Sechrist, Pauline Germaine, Eva Castro Lopez, T omas Amerio, Pushpender Gupta, Manoj Jain, Fernando U. Kay , Cheng T ing Lin, Saugata Sen, Jonathan W esley Rev els, Carola C. Brussaard, John Mongan, For the RSNA-STR Annota- tors, and Dataset Curation Contributors. The rsna pulmonary embolism ct dataset. Radi- ology: Artificial Intelligence , 3(2):e200254, 2021. doi: 10.1148/ryai.2021200254. URL https://doi.org/10.1148/ryai.2021200254 . 146 [595] Nicholas Heller, F abian Isensee, Dasha Trofimo v a, Resha T ejpaul, Zhongchen Zhao, Huai Chen, Lisheng W ang, Alex Golts, Daniel Khapun, Daniel Shats, Y oel Shoshan, Flora Gilboa- Solomon, Y asmeen George, Xi Y ang, Jianpeng Zhang, Jing Zhang, Y ong Xia, Mengran W u, Zhiyang Liu, Ed W alczak, Sean McSweeney , Ran v eer V asdev , Chris Hornung, Rafat So- laiman, Jamee Schoephoerster , Bailey Abernathy , David W u, Safa Abdulkadir, Ben Byun, Justice Spriggs, Grif fin Struyk, Alexandra Austin, Ben Simpson, Michael Hagstrom, Sierra V irnig, John French, Nitin V enkatesh, Sarah Chan, Keenan Moore, Anna Jacobsen, Susan Austin, Mark Austin, Subodh Regmi, Nikolaos P apanikolopoulos, and Christopher W eight. neheller/knight: The of ficial repository of the isbi 2022 knight challenge. GitHub repository: https://github.com/neheller/KNIGHT , 2021. MIT License. 146 [596] Kingsley Kuan, Mathieu Ra v aut, Gaura v Manek, Huiling Chen, Jie Lin, Babar Nazir, Cen Chen, Tse Chiang Howe, Zeng Zeng, and V ijay Chandrasekhar . Deep learning for lung cancer detection: T ackling the kaggle data science bowl 2017 challenge. arXiv preprint arXiv:1705.09435, 2017. Conference on Information and Knowledge Management, Nov em- ber 2017, Singapore. 146 [597] Open Source Imaging Consortium. Osic pulmonary fibrosis progression. Kaggle competition, 2020. https://www .kaggle.com/competitions/osic-pulmonary-fibrosis-progression. 146 [598] Richard A. Ban v ard. The visible human project® image data set from inception to completion and be yond. In Pr oceedings of COD A T A 2002: F r ontiers of Scientific and T echnical Data, T rack I-D-2: Medical and Health Data , Montréal, Canada, October 2002. 147 [599] Nadya Shusharina, Thomas Bortfeld, Carlos Cardenas, and Jinzhong Y ang. Anatomical brain barriers to cancer spread: Segmentation from ct and mr images challenge design document. https://doi.org/10.5281/zenodo.3746561, 2020. 23rd International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2020), Lima, Peru, October 4–8, 2020. 147 [600] Marco Mueller, Per Poulsen, Rune Hansen, W ilk o V erbakel, Ross Berbeco, Dianne Ferguson, Shinichiro Mori, Lei Ren, John C. Roeske, Lei W ang, Pengpeng Zhang, and Paul Keall. The markerless lung target tracking aapm grand challenge (match) results. Medical Physics , 49 (2):1161–1180, 2021. doi: 10.1002/mp.15418. 147 [601] John Kipritidis, Bilal T ahir , Guillaume Cazoulat, Shankar Siv a, P aul Keall, Michael Hof- man, Jason Callahan, Peter Greer, Thomas Eade, Nicholas Hardcastle, W ei Chen, Jack Peter- son, Robert O’Brien, Michael Liston, and Ann Haw orth. The v ampire challenge: A multi- institutional v alidation study of ct v entilation imaging. Medical Physics , 46(3):1198–1217, 2019. 147 99 [602] Y ashin Dicente Cid, V itali Liauchuk, V assili Ko vale v , and Henning Müller . Overvie w of ImageCLEFtuberculosis 2018 - detecting multi-drug resistance, classifying tuberculosis type, and assessing severity score. In CLEF2018 W orking Notes , CEUR W orkshop Proceedings, A vignon, France, September 10-14 2018. CEUR-WS.org. 147 [603] Y ashin Dicente Cid, Alexander Kalino vsky , V itali Liauchuk, V assili K o v ale v , and Henning Muller . Overvie w of ImageCLEFtuberculosis 2017 - predicting tuberculosis type and drug resistances. In CLEF2017 W orking Notes , CEUR W orkshop Proceedings, Dublin, Ireland, September 11-14 2017. CEUR-WS.org. 147 [604] Michael Green and Arnaldo Mayer . fastpet-ld. Zenodo, https://doi.org/10.5281/zenodo.4781986, 2021. 147 , 154 [605] T aylor R. Moen, Baiyu Chen, Da vid R. III Holmes, Xinhui Duan, Zhicong Y u, Lifeng Y u, Shuai Leng, Joel G. Fletcher , and Cynthia H. McCollough. Low-dose ct image and projection dataset. Medical Physics , 48(2):902–911, 2020. doi: 10.1002/mp.14594. 147 [606] Dagmar Grob, Luuk Oostveen, Jan Rühaak, Stefan Heldmann, Brian Mohr , Koen Michielsen, Sabrina Dorn, Mathias Prokop, Marc Kachelrieß, Monique Brink, and Ioannis Sechopoulos. Accuracy of re gistration algorithms in subtraction ct of the lungs: A digital phantom study . Medical Physics , 46(5):2264–2274, 2019. doi: 10.1002/mp.13496. 147 [607] Author Name. Cad-pe: A computed tomography pulmonary embolism dataset. arXiv pr eprint arXiv:2003.13440 , 2020. 147 [608] Bram v an Ginneken, Samuel G. Armato, Bartjan de Hoop, Saskia v an de V orst, Tim Duin- dam, Marcel Niemeijer , K evin Murphy , A M R Schilham, Alessandro Retico, Maria E Fan- tacci, Niccolo Camarlinghi, Federico Bagagli, Isabella Gori, Tsuyoshi Hara, Hiroyuki Fu- jita, Guido Gargano, Riccardo Belloti, F D Carlo, Rosella Megna, Savino T angaro, Luis Bolanos, Paolo Cerello, S C Cheran, E L T orres, and Mathias Prokop. Comparing and combining algorithms for computer -aided detection of pulmonary nodules in computed to- mography scans: the anode09 study . Medical Image Analysis , 14(6):707–722, 2010. doi: 10.1016/j.media.2010.05.005. 147 [609] Rina D Rudyanto, Sjoerd Kerkstra, Ev a M van Rikxoort, Catalin Fetita, Pierre-Yves Bril- let, Christophe Lefe vre, W enzhe Xue, Xiangjun Zhu, Jianming Liang, Ilkay Öksüz, De vrim Ünay , Kamuran Kadipa ¸ sao ˘ glu, Raúl San José Estépar , James C Ross, George R W ashko, Juan-Carlos Prieto, Marcela Hernández Hoyos, Maciej Orkisz, Hans Meine, Markus Hülle- brand, Christina Stöcker , Fernando Lopez Mir , V alery Naranjo, Eliseo V illanue v a, Mar- ius Staring, Changyan Xiao, Berend C Stoel, Anna Fabijanska, Erik Smistad, Anne C El- ster , Frank Lindseth, Amir Hossein Foruzan, Ryan Kiros, Karteek Popuri, Dana Cobzas, Daniel Jimenez-Carretero, Andres Santos, Maria J Ledesma-Carbayo, Michael Helmberger , Martin Urschler, Michael Pienn, Dennis G H Bosboom, Arantza Campo, Mathias Prokop, Pim A de Jong, Carlos Ortiz-de Solorzano, Arrate Muñoz-Barrutia, and Bram v an Ginneken. Comparing algorithms for automated vessel segmentation in computed tomography scans of the lung: the vessel12 study . Medical Image Analysis , 18(7):1217–1232, 2014. doi: 10.1016/j.media.2014.07.003. 147 [610] Binsheng Zhao, La wrence H. Schw artz, Mark G. Kris, and Gregory J. Riely . Coffee-break lung ct collection with scan images reconstructed at multiple imaging parameters (version 3). The Cancer Imaging Archiv e, 2015. 147 [611] Aaron Babier, Binghao Zhang, Rafid Mahmood, K e vin L. Moore, Thomas G. Purdie, An- drea L. McNi ven, and Timoth y C.Y . Chan. Openkbp: The open-access knowledge-based planning grand challenge and dataset. Medical Physics , 48(9):5549–5561, 2021. doi: 10.1002/mp.14845. URL https://doi.org/10.1002/mp.14845 . 147 [612] E. M. Eslick, J. Kipritidis, D. Gradinscak, M. J. Stevens, D. L. Bailey , B. Harris, J. T . Booth, and P . J. Keall. Ct ventilation as a functional imaging modality for lung cancer radiotherapy (ct-vs-pet-ventilation-imaging) v ersion 1. The Cancer Imaging Archi ve, 2022. URL https: //doi.org/10.7937/3ppx- 7s22 . Data set. 147 , 154 100 [613] A. W . Moa wad, D. Fuentes, A. Morshid, A. M. Khalaf, M. M. Elmohr , A. Abusaif, J. D. Hazle, A. O. Kaseb, M. Hassan, A. Mahvash, J. Szklaruk, A. Qayyom, and K. M. Elsayes. Multimodality annotated hcc cases with and without advanced imaging se gmentation. Data set, The Cancer Imaging Archi ve, 2021. URL https://www.cancerimagingarchive. net/collection/hcc- tace- seg/ . 147 [614] David Zimmerer , Jens Petersen, Gregor Köhler , Paul Jäger , Peter Full, Klaus Maier-Hein, T obias Roß, T im Adler, Annika Reinke, and Lena Maier -Hein. Medical out-of-distribution analysis challenge 2022. Zenodo, https://doi.org/10.5281/zenodo.6362313, 2022. 25th In- ternational Conference on Medical Image Computing and Computer Assisted Interv ention (MICCAI 2022). 147 [615] T . T ong and M. Li. Abdominal or pelvic enhanced ct images within 10 days before surgery of 230 patients with stage ii colorectal cancer (stageii-colorectal-ct). The Cancer Imaging Archiv e, 2022. URL https://www.cancerimagingarchive.net/collection/ stageii- colorectal- ct/ . 147 [616] Michael Rutherford, Seong K. Mun, Betty Le vine, William Bennett, Kirk Smith, Phil Farmer , Quasar Jarosz, Ulrike W agner, John Freyman, Geri Blake, La wrence T arbox, Keyv an Fara- hani, and Fred Prior . A dicom dataset for ev aluation of medical image de-identification. Scientific Data , 8(1):183, 2021. doi: 10.1038/s41597- 021- 00967- y. 147 , 154 [617] Geof frey D. Hugo, Elisabeth W eiss, William C. Sleeman, Salim Balik, Paul J. K eall, Jun Lu, and Jeffrey F . W illiamson. A longitudinal four -dimensional computed tomography and cone beam computed tomography dataset for image-guided radiation therapy research in lung cancer . Medical Physics , 44(2):762–771, 2017. doi: 10.1002/mp.12059. 147 , 155 [618] Binsheng Zhao. Lung phantom (version 2). The Cancer Imaging Archive [Data set], 2015. URL https://doi.org/10.7937/k9/tcia.2015.08a1ixoo . 147 [619] Karen A Kurdziel, Andrea B Apolo, Liza Lindenber g, Esther Mena, Y olanda Y McKin- ney , Stephen S Adler, and Peter L Choyke. Data from naf prostate [dataset]. The Cancer Imaging Archiv e, 2015. URL https://www.cancerimagingarchive.net/collection/ naf- prostate/ . 147 , 154 [620] Sharib Ali, Y ueming Jin, Y amid Espinel López, Emmanuel Buc, Bertrand Le Roy , Patrick T eoule, Christoph Reissfelder , Adam Bailey , Zahir Soonawalla, Ale x Gordon-W eeks, Michael Silv a, Lena Maier-Hein, and Adrien Bartoli. Preoperativ e to intraoperati ve la- paroscopy fusion. Zenodo, 25th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2022), 2022. URL https://doi.org/10.5281/ zenodo.6362162 . 147 [621] K evin Mader . Finding and measuring lungs in ct data, 2019. URL https://www.kaggle. com/datasets/kmader/finding- lungs- in- ct- data . 147 [622] Adrian Thummerer , Erik van der Bijl, Arthur Jr . Galapon, Joost J. C. V erhoeff, Johannes A. Langendijk, Stefan Both, Cornelis A. T . v an den Berg, and Matteo Maspero. Synthrad2023 grand challenge dataset: Generating synthetic ct for radiotherapy . Medical Physics , 50(7): 4664–4674, 2023. doi: 10.1002/mp.16529. 147 [623] Luca Giancardo et al. The image analysis for cta endo v ascular stroke therapy (iacta-est) data challenge. In Pr oceedings of the 2023 IEEE International Symposium on Biomedical Imaging (ISBI) . IEEE, 2023. URL https://lgiancauth.github.io/iacta- est- 2023/ . 147 [624] ISBI Challenge Organizers. Isbi 2023 challenge - apis (advancements in pulmonary imaging segmentation), 2023. URL https://bivl2ab.uis.edu.co/challenges/apis . Brain imaging segmentation challenge dataset. 147 [625] Mark J. Gooding, Shafak Aluwini, T eresa Guerrero Urbano, Y asmin McQuinlan, Deborah Om, Floor H. E. Staal, T anguy Perennec, Sana Azzarouali, Carlos E. Cardenas, Antony Carver , Stine Sofia K orreman, and Jean-Emmanuel Bibault. Fully automated radiotherapy treatment planning: A scan to plan challenge. Radiotherapy and Oncology , 200:110513, 2024. doi: 10.1016/j.radonc.2024.110513. 147 101 [626] Birger C Lassen, Colin Jacobs, Jan-Martin K uhnigk, Bram v an Ginneken, and Eric M van Rikxoort. Robust semi-automatic segmentation of pulmonary subsolid nodules in chest com- puted tomography scans. Physics in Medicine and Biology , 60(3):1307–1323, 2015. doi: 10.1088/0031- 9155/60/3/1307. 147 [627] Andrey Fedorov , Matthe w Hancock, David Clunie, Mathias Brochhausen, Jonathan Bona, Justin Kirby , John Freymann, Ste ve Pieper , Hugo J. W . L. Aerts, Ron Kikinis, and Fred Prior . Dicom re-encoding of volumetrically annotated lung imaging database consortium (lidc) nodules. Medical Physics , 47(11):5953–5965, 2020. doi: 10.1002/mp.14445. 147 [628] S. V . Zolotov a, A. V . Golanov , I. N. Pronin, A. V . Dalechina, A. A. Nikolaev a, A. S. Belyashov a, D. Y . Usache v , E. A. Kondrate v a, P . V . Druzhinina, B. N. Shirokikh, T . N. Saparov , M. G. Belyaev , and A. I. Kurmuko v . Burdenko’ s glioblastoma progression dataset (burdenk o-gbm-progression) (version 1). Data set, 2023. URL https://www. cancerimagingarchive.net/collection/burdenko- gbm- progression/ . 147 [629] J. L. T atum, J. D. Kalen, L. V . Ile va, L. A. Rif fle, S. K eita, N. P atel, P . M. Jacobs, C. Sanders, A. James, S. Difilippantonio, L. Thang, M. G. Hollingshead, J. Phillips, Y . Evrard, D. A. Clunie, Y . Liu, C. Suloway , K. E. Smith, U. W agner , and J. H. Doroshow . Imaging characterization of a metastatic patient deriv ed model of adenocarcinoma colon: (pdmr- 997537-175-t). Data set, The Cancer Imaging Archi ve, 2020. URL https://www. cancerimagingarchive.net/collection/pdmr- 997537- 175- t/ . 147 , 151 , 154 [630] Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Research Net- work. Data from the applied proteogenomics organizational learning and outcomes lung squa- mous cell carcinoma [apollo-5-lscc] collection. The Cancer Imaging Archive, 2021. URL https://wiki.cancerimagingarchive.net/display/Public/APOLLO- 5- LSCC . 147 , 154 [631] Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Research Network. Data from the applied proteogenomics organizational learning and outcomes lung adenocarcinoma cohort [apollo-5-luad] collection. The Cancer Imaging Archiv e, 2021. URL https://wiki.cancerimagingarchive.net/pages/viewpage.action? pageId=96337930 . 147 [632] Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Research Network. Data from the applied proteogenomics organizational learning and outcomes esophageal squamous cell carcinoma [apollo-5-esca] collection. The Cancer Imaging Archiv e, 2021. URL https://wiki.cancerimagingarchive.net/display/Public/ APOLLO- 5- ESCA . 147 [633] Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Research Net- work. Data from the applied proteogenomics organizational learning and outcomes pancre- atic adenocarcinoma [apollo-5-paad] collection. The Cancer Imaging Archive, 2021. URL https://wiki.cancerimagingarchive.net/display/Public/APOLLO- 5- PAAD . 147 [634] Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Research Net- work. Data from the applied proteogenomics organizational learning and outcomes thy- moma [apollo-5-thym] collection. The Cancer Imaging Archiv e, 2021. URL https: //wiki.cancerimagingarchive.net/display/Public/APOLLO- 5- THYM . 147 [635] Anant Madabhushi and Mirabela Rusu. Fused radiology-pathology lung (lung-fused-ct- pathology). Data set. The Cancer Imaging Archi ve, V ersion 1, 2018. URL https: //www.cancerimagingarchive.net/collection/lung- fused- ct- pathology/ . 147 [636] H. Lee Moffitt Cancer Center & Research Institute. Long and short survi v al in adenocarci- noma lung cts (luad-ct-survi v al). The Cancer Imaging Ar chive , 2017. URL https://wiki. cancerimagingarchive.net/pages/viewpage.action?pageId=24284406 . 147 [637] Carlos Rodriguez-Galindo, Mark D. Krailo, Matthew J. Krasin, Lillian Huang, M. Beth McCarville, John Hicks, Farzana Pashankar , and Alberto S. Pappo. Radiation therapy , amifostine, and chemotherapy in treating young patients with newly diagnosed nasopha- ryngeal cancer (arar0331). The Cancer Imaging Archive, 2022. URL https://www. cancerimagingarchive.net/collection/arar0331/ . V ersion 1 [Data set]. 147 , 154 102 [638] Ev an Porter , P atrick Fuentes, Irene Sala, Zohaib Siddiqui, Ross Levitin, Nicholas Myziuk, Blake Squires, Thomas Gonzalez, Pierre Chen, T im Guerrero, and Ing a Grills. Gamma knife mr/ct/rtstruct sets with hippocampal contours (gammaknife-hippocampal). V ersion 1. The Cancer Imaging Archive [Data set], 2022. URL https://doi.org/10.7937/Q967- X166 . 147 [639] M. M. Gounder, M. R. Mahoney , B. A. V an T ine, V . Ravi, S. Attia, H. A. Deshpande, A. A. Gupta, M. M. Milhem, R. M. Conry , S. Movv a, M. J. Pishvaian, R. F . Riedel, T . Sabagh, W . D. T ap, N. Horvat, E. Basch, L. H. Schwartz, R. G. Maki, N. P . Agaram, and G. K. Schwartz. Sorafenib tosylate in treating patients with desmoid tumors or aggressi ve fi- bromatosis (a091105) (version 1) [data set]. The Cancer Imaging Archiv e, 2023. URL https://www.cancerimagingarchive.net/collection/a091105/ . 147 [640] Amber L Simpson, Jacob Peoples, John M Creasy , Gabor Fichtinger , Natalie Gangai, Kr - ishna N Kesha v amurthy , Andras Lasso, Jinru Shia, Michael I D’Angelica, and Richard K G Do. Preoperati ve ct and survi v al data for patients under going resection of colorectal li ver metastases. Scientific Data , 11:172, 2024. doi: 10.1038/s41597- 024- 02981- 2. 147 [641] A Grossberg, A Mohamed, H Elhalaw ani, W Bennett, K Smith, T Nolan, S Chamchod, M Kantor , T Bro wne, K Hutcheson, G Gunn, AS Garden, SJ Frank, DI Rosenthal, J Frey- mann, and C Fuller . Data from head and neck cancer ct atlas (version 2). Dataset. The Cancer Imaging Archiv e, 2017. 147 [642] T awfik Giaddui, W ensha Chen, James Y u, Lili Lin, Charles B. Simone II, Lin Y uan, Y . U. T . Gong, Q. Jackie W u, Radhe Mohan, Xiaodong Zhang, John B. Bluett, Michael Gillin, Ke vin Moore, Ellen O’Meara, John Presley , Jef fre y D. Bradley , Zhongxing Liao, James Galvin, and Y ing Xiao. Data from nrg-1308 (version 1). Data set, 2016. 147 [643] Michael A. Ga vrielides, Laura M. Kinnard, K enneth J. Myers, Jennifer Peregoy , W illiam F . Pritchard, Rui Zeng, Jose Esparza, John Karanian, and Nicholas Petrick. Data from phantom fda. Data set. The Cancer Imaging Archi ve, 2015. 147 [644] Luohai Chen, W ei W ang, Kaizhou Jin, Bing Y uan, Huangying T an, Jian Sun, Y u Guo, Y anji Luo, Shi-Ting Feng, Xianjun Y u, Min-Hu Chen, and Jie Chen. Prediction of sunitinib efficac y using computed tomography in patients with pancreatic neuroendocrine tumors. International Journal of Cancer , 152(1):90–99, 2023. doi: 10.1002/ijc.34294. 147 [645] L. W ee, H. Aerts, P . Kalendralis, and A. Dekker . Rider lung ct segmentation labels from: Decoding tumour phenotype by nonin v asi ve imaging using a quantitativ e radiomics approach. Data set. The Cancer Imaging Archive, 2020. URL https://doi.org/10.7937/tcia. 2020.jit9grk8 . 147 [646] M. McNitt-Gray , S. Napel, A. Jaggi, S.A. Mattonen, L. Hadjiiski, M. Muzi, D. Goldgof, Y . Balagurunathan, L.A. Pierce, P .E. Kinahan, E.F . Jones, A. Nguyen, A. V irkud, H.-P . Chan, N. Emaminejad, M. W ahi-Anwar , M. Daly , M. Abdalah, H. Y ang, L. Lu, W . Lv , A. Rahmim, A. Gastounioti, S. Pati, S. Bakas, D. K ontos, B. Zhao, J. Kalpathy-Cramer , and K. Farahani. Standardization in quantitative imaging: A multicenter comparison of radiomic features from different softw are packages on digital reference objects and patient data sets. T omogr aphy , 6 (2):118–128, 2020. doi: 10.18383/j.tom.2019.00031. 147 [647] K endall J Kiser , Sara Ahmed, Sonja Stieb, Abdallah S R Mohamed, Hesham Elhalaw ani, Peter Y S Park, Nathan S Doyle, Brandon J W ang, Arko Barman, Zhao Li, W Jim Zheng, Clifton D Fuller, and Luca Giancardo. Plethora: Pleural ef fusion and thoracic cavity segmen- tations in diseased lungs for benchmarking chest ct processing pipelines. Medical Physics , 47(11):5941–5952, 2020. doi: 10.1002/mp.14424. 147 [648] R. B. Ger , J. Y ang, Y . Ding, M. C. Jacobsen, C. E. Cardenas, C. D. Fuller, R. M. Howell, H. Li, R. J. Stafford, S. Zhou, and L. E. Court. Synthetic head and neck and phantom images for determining deformable image registration accuracy in magnetic resonance imaging. Medical Physics , 45(9):4315–4321, 2018. doi: 10.1002/mp.13090. 147 103 [649] R B Puchalski, N Shah, J Miller , R Dalle y , S R Nomura, J-G Y oon, K A Smith, M Lan- kerovich, D Bertagnolli, K Bickley , A F Boe, K Brouner , S Butler , S Caldejon, M Chapin, S Datta, N Dee, T Desta, T Dolbeare, N Dotson, A Ebbert, D Feng, X Feng, M Fisher , G Gee, J Goldy , L Gourley , B W Gregor , G Gu, N Hejazinia, J Hohmann, P Hothi, R Ho ward, K Joines, A Kriedberg, L Kuan, C Lau, F Lee, H Lee, T Lemon, F Long, N Mastan, E Mott, C Murthy , K Ngo, E Olson, M Reding, Z Riley , D Rosen, D Sandman, N Shapovalo v a, C R Slaughterbeck, A Sodt, G Stockdale, A Szafer , W W akeman, P E W ohnoutka, S J White, D Marsh, R C Rostomily , L Ng, C Dang, A Jones, B Keogh, H R Gittleman, J S Barnholtz- Sloan, P J Cimino, M S Uppin, C D K eene, F R Farrokhi, J D Lathia, M E Berens, A Iav arone, A Bernard, E Lein, J W Phillips, S W Rostad, C Cobbs, M J Hawrylycz, and G D Foltz. An anatomic transcriptional atlas of human glioblastoma. Science , 360(6389):660–663, 2018. doi: 10.1126/science.aaf2666. 148 [650] Lung Image Database Consortium (LIDC). RIDER Pilot [data set]. The Cancer Imaging Archiv e (TCIA), 2023. URL https://doi.org/10.7937/m87f- mz83 . 148 [651] Mattea L. W elch, Sejin Kim, Andrew J. Hope, Shao Hui Huang, Zhibin Lu, Joseph Mar- silla, Michal Kazmierski, Katrina Rey-McIntyre, Tirth Patel, Brian O’Sulliv an, John W aldron, Scott Bratman, Benjamin Haibe-Kains, and T ony T adic. Radcure: An open-source head and neck cancer ct dataset for clinical radiation therapy insights. Medical Physics , 2024. doi: 10.1002/mp.16972. 148 [652] A. W . Moawad, A. A. Ahmed, M. ElMohr , M. Eltaher , M. A. Habra, S. Fisher , N. Perrier , M. Zhang, D. Fuentes, and K. Elsayes. V oxel-lev el segmentation of pathologically-prov en adrenocortical carcinoma with ki-67 expression (adrenal-acc-ki67-se g). Data set, The Cancer Imaging Archiv e, 2023. URL https://www.cancerimagingarchive.net/collection/ adrenal- acc- ki67- seg/ . 148 [653] Aasheesh Kanwar , Brandon Merz, Cheryl Claunch, Shushan Rana, Arthur Hung, and Reid F . Thompson. Stress-testing pelvic autosegmentation algorithms using anatomical edge cases. Physics and Imaging in Radiation Oncology , 25:100413, 2023. doi: 10.1016/j.phro.2023. 100413. 148 [654] Antonio Pepe, Gian Marco Melito, and Jan Egger, editors. Segmentation of the Aorta: T owards the Automatic Segmentation, Modeling, and Meshing of the Aortic V essel T r ee fr om Multicenter Acquisition , volume 14539 of Lecture Notes in Computer Science , Febru- ary 2024. Springer . ISBN 978-3-031-53241-2. doi: 10.1007/978- 3- 031- 53241- 2. URL https://link.springer.com/book/10.1007/978- 3- 031- 53241- 2 . 148 [655] Sharib Ali, Y amid Espinel, Y ueming Jin, Peng Liu, Bianca Güttner, Xukun Zhang, Lihua Zhang, T om Dowrick, Matthew J Clarkson, Shiting Xiao, Y if an W u, Y ijun Y ang, Lei Zhu, Dai Sun, Lan Li, Micha Pfeiffer , Shahid F arid, Lena Maier-Hein, Emmanuel Buc, and Adrien Bartoli. An objective comparison of methods for augmented reality in laparoscopic liver re- section by preoperativ e-to-intraoperati ve image fusion from the miccai2022 challenge. Med- ical Image Analysis , 99:103371, 2025. doi: 10.1016/j.media.2024.103371. 148 [656] Andre w Hoopes, Jocelyn S. Mora, Adrian V . Dalca, Bruce Fischl, and Malte Hof fmann. Synthstrip: Skull-stripping for any brain image. Neur oImage , 260:119474, 2022. doi: https: //doi.org/10.1016/j.neuroimage.2022.119474. 148 [657] David Zimmerer , Peter M Full, Fabian Isensee, Paul Jäger , Tim Adler , Jens Petersen, Gregor Köhler, T obias Ross, Annika Reinke, Antanas Kascenas, Bjørn Sand Jensen, Al- ison Q O’Neil, Jeremy T an, Benjamin Hou, James Batten, Huaqi Qiu, Bernhard Kainz, Nina Shvetsov a, Irina Fedulova, Dmitry V Dylov , Baolun Y u, Jianyang Zhai, Jingtao Hu, Runxuan Si, Sihang Zhou, Siqi W ang, Xinyang Li, Xuerun Chen, Y ang Zhao, Ser- gio Na v al Marimont, Giacomo T arroni, V ictor Saase, Lena Maier -Hein, and Klaus Maier - Hein. Mood 2020: A public benchmark for out-of-distribution detection and localization on medical images. IEEE T ransactions on Medical Imaging , 41(10):2728–2738, 2022. doi: 10.1109/TMI.2022.3170077. 148 , 151 [658] Y udi Sang, Y anzhen Liu, Sutuke Y ib ulayimu, Y unning W ang, Benjamin D. Killeen, Mingxu Liu, Ping-Cheng Ku, Ole Johannsen, Karol Gotkowski, Maximilian Zenk, Klaus Maier-Hein, 104 Fabian Isensee, Peiyan Y ue, Y i W ang, Haidong Y u, Zhaohong Pan, Y utong He, Xiaokun Liang, Daiqi Liu, Fuxin F an, Artur Jurg as, Andrzej Skalski, Y uxi Ma, Jing Y ang, Szymon Płotka, Rafał Litka, Gang Zhu, Y ingchun Song, Mathias Unberath, Mehran Armand, Dan Ruan, S. Ke vin Zhou, Qiyong Cao, Chunpeng Zhao, Xinbao W u, and Y u W ang. Benchmark of se gmentation techniques for pelvic fracture in ct and x-ray: Summary of the pengwin 2024 challenge. arXiv pr eprint arXiv:2504.02382 , 2025. doi: 10.48550/arXiv .2504.02382. 148 [659] W ei Huang, W ei Liu, Xiaoming Zhang, Xiaoli Y in, Xu Han, Chunli Li, Y uan Gao, Y u Shi, Le Lu, Ling Zhang, Lei Zhang, and Ke Y an. Triphasic-aided liv er lesion se gmentation in non- contrast ct (trials) challenge. In Pr oceedings of the 27th International Confer ence on Medical Image Computing and Computer-Assisted Intervention (MICCAI) , page TBD. Springer , 2024. doi: 10.1007/978- 3- 031- 72114- 4_38. 148 [660] Maraw an Elbatel, Xiaomeng Li, Mohamed Ghonim, Mohanad Ghonim, Amr Muham- mad Abdo Salem, Nouran Elghitany , Noha Elghitany , Amira Adel, Susan Adil Ali, and A ya Y assin. T riphasic-aided liver lesion segmentation in non-contrast ct. Zenodo dataset for the 27th International Conference on Medical Image Computing and Computer Assisted In- tervention (MICCAI 2024) challenge, 2024. URL https://doi.org/10.5281/zenodo. 10992127 . 148 [661] Hongwei Bran Li, Fernando Na varro, Ivan Ezhov , Amirhossein Bayat, Dhritiman Das, Flo- rian Kofler , Suprosanna Shit, Diana W aldmannstetter , Johannes C. P aetzold, Xiaobin Hu, Benedikt Wiestler , Lucas Zimmer, T amaz Amiranashvili, Chinmay Prabhakar , Christoph Berger , Jonas W eidner , Michelle Alonso-Basant, Arif Rashid, Ujjwal Baid, W esam Adel, Deniz Ali, Bhakti Baheti, Y ingbin Bai, Ishaan Bhatt, Sabri Can Cetindag, W enting Chen, Li Cheng, Prasad Dutand, Lara Dular, Mustafa A. Elattar , Ming Feng, Shengbo Gao, Henk- jan Huisman, W eifeng Hu, Shubham Innani, W ei Jiat, Da v ood Karimi, Hugo J. K uijf, Jin T ae Kwak, Hoang Long Le, Xiang Lia, Huiyan Lin, T ongliang Liu, Jun Ma, Kai Ma, T ing Ma, Ilkay Oksuz, Robbie Holland, Arlindo L. Oli veira, Jimut Bahan Pal, Xuan Pei, Maoying Qiao, Anindo Saha, Raghavendra Selvan, Linlin Shen, Joao Lourenco Silva, Ziga Spiclin, Sanjay T albar , Dadong W ang, W ei W ang, Xiong W ang, Y in W ang, Ruiling Xia, Kele Xu, Y anwu Y an, Mert Y ergin, Shuang Y u, Lingxi Zeng, Y ingLin Zhang, Jiachen Zhao, Y efeng Zheng, Martin Zukov ec, Richard Do, Anton Becker , Amber Simpson, Ender K onukoglu, Andras Jakab, Spyridon Bakas, Leo Josko wicz, and Bjoern Menze. Qubiq: Uncertainty quantifica- tion for biomedical image segmentation challenge. arXiv preprint , 2024. doi: 10.48550/arXi v .2405.18435. 148 [662] Jelmer M W olterink, T im Leiner , Bob D de V os, Rogier W van Hamersvelt, Max A V ierge v er , and Ivana Išgum. An evaluation of automatic coronary artery calcium scoring methods with cardiac ct using the orcascore frame work. Medical Physics , 43(5):2361–2373, 2016. doi: 10.1118/1.4945696. 148 [663] Shih-Cheng Huang, Zepeng Huo, Ethan Steinberg, Chia-Chun Chiang, Curtis Langlotz, Matthew P Lungren, Serena Y eung, Nigam Shah, and Jason Alan Fries. Inspect: A multimodal dataset for pulmonary embolism diagnosis and prognosis. arXiv pr eprint arXiv:2311.10798 , 2023. 148 [664] Y inda Chen, Che Liu, Xiaoyu Liu, Rossella Arcucci, and Zhiwei Xiong. Bimcv-r: A land- mark dataset for 3d ct text-image retrie val. In Medical Imag e Computing and Computer - Assisted Intervention – MICCAI 2024 , volume 15011 of Lectur e Notes in Computer Science , pages 124–134. Springer , 2024. doi: 10.1007/978- 3- 031- 72120- 5_12. 148 [665] Jürgen W allner , Irene Mischak, and Jan Egger . Computed tomography data collection of the complete human mandible and v alid clinical ground truth models. Scientific Data , 6:190003, 2019. doi: 10.1038/sdata.2019.3. 148 [666] Y aqi W ang, Shuai W ang, F an Y e, W eiwei Cui, Y ifan Zhang, Liaoyuan Zeng, and Xingru Huang. Semi-supervised teeth segmentation. Zenodo, dataset, International Conference on Medical Image Computing and Computer Assisted Interv ention (MICCAI) 2023, 2023. URL https://zenodo.org/record/7840021 . 148 105 [667] T ugba Akinci D’Antonoli, Lucas K. Ber ger , Ashraya K. Indrakanti, Nathan V ishw anathan, Jakob W eiß, Matthias Jung, Zeynep Berkarda, Alexander Rau, Marco Reisert, Thomas Küst- ner , Ale xandra W alter , Elmar M. Merkle, Daniel Boll, Hanns-Christian Breit, Andrew Phillip Nicoli, Martin Segeroth, Joshy Cyriac, Shan Y ang, and Jakob W asserthal. T otalsegmenta- tor mri: Rob ust sequence-independent segmentation of multiple anatomic structures in mri. Radiology , 314(2):e241613, 2025. doi: 10.1148/radiol.241613. 148 [668] Oli vier Bernard, Alain Lalande, Clement Zotti, Frederick Cervenansky , Xin Y ang, Pheng- Ann Heng, Irem Cetin, Karim Lekadir , Oscar Camara, Miguel Angel Gonzalez Ballester, Gerard Sanroma, Sandy Napel, Steffen Petersen, Georgios Tziritas, Elias Grinias, Mahendra Khened, V arghese Alex K ollerathu, Ganapathy Krishnamurthi, Marc-Michel Rohé, Xavier Pennec, Maxime Sermesant, Fabian Isensee, Paul Jäger , Klaus H. Maier-Hein, Peter M. Full, Ivo W olf, Sandy Engelhardt, Christian F . Baumgartner , Lisa M. K och, Jelmer M. W olterink, Ivana Išgum, Y eonggul Jang, Y oonmi Hong, Jay Patrav ali, Shubham Jain, Olivier Humbert, and Pierre-Marc Jodoin. Deep learning techniques for automatic mri cardiac multi-structures segmentation and diagnosis: Is the problem solved? IEEE T ransactions on Medical Ima ging , 37(11):2514–2525, 2018. doi: 10.1109/TMI.2018.2837502. 148 [669] V ictor M Campello, Polyxeni Gk ontra, Cristian Izquierdo, Carlos Martin-Isla, Alireza So- joudi, Peter M Full, Klaus Maier-Hein, Y ao Zhang, Zhiqiang He, Jun Ma, Mario Parreno, Alberto Albiol, Fanwei K ong, Shawn C Shadden, Jorge Corral Acero, V aanathi Sundaresan, Mina Saber, Mustafa Elattar, Hongwei Li, Bjoern Menze, Firas Khader , Christoph Haar- bur ger , Cian M Scannell, Mitko V eta, Adam Carscadden, Kumaradev an Punithakumar , Xiao Liu, Sotirios A Tsaftaris, Xiaoqiong Huang, Xin Y ang, Lei Li, Xiahai Zhuang, David V ilades, Martin L Descalzo, Andrea Guala, Lucia La Mura, Matthias G Friedrich, Ria Garg, Julie Lebel, Filipe Henriques, Mahir Karakas, Ersin Ca vus, Stef fen E Petersen, Sergio Escalera, Santi Se gui, Jose F Rodriguez-Palomares, and Karim Lekadir . Multi-centre, multi-vendor and multi-disease cardiac segmentation: The m&ms challenge. IEEE T ransactions on Medi- cal Imaging , 40(12):3543–3554, 2021. doi: 10.1109/TMI.2021.3090082. 148 [670] O. Camara, E. Konuk oglu, M. Pop, K. M. Moeller , M. Sermesant, and A. Y oung. Statistical atlases and computational models of the heart. multi-disease, multi-vie w , and multi-center right ventricular se gmentation in cardiac mri challenge: 12th international workshop, stacom 2021, held in conjunction with miccai 2021, strasbourg, france, september 27, 2021, revised selected papers. In Lectur e Notes in Computer Science , volume 13025. Springer , 2021. doi: 10.1007/978- 3- 030- 93722- 5. 148 [671] Lei Li, V eronika A. Zimmer, Julia A. Schnabel, and Xiahai Zhuang. Left atrial and scar quantification and segmentation: First challenge, lascarqs 2022, held in conjunction with miccai 2022, singapore, september 18, 2022, proceedings. In Lectur e Notes in Computer Science , volume 13586, pages 1–10. Springer , 2023. 148 [672] Lei Li, V eronika A. Zimmer , Julia A. Schnabel, and Xiahai Zhuang. Lascarqs++: Multi- center left atrial and scar quantification and segmentation challenge. In CARE 2024: Compr ehensive Analysis and Retrieval of Medical Images . Springer , 2024. URL https: //www.zmic.org.cn/care_2024/track2/ . 148 [673] Xiahai Zhuang. Multiv ariate mixture model for myocardial segmentation combining multi- source images. IEEE T ransactions on P attern Analysis and Machine Intelligence , 41(12): 2933–2946, 2019. 148 [674] Lei Li, V eronika A. Zimmer , Julia A. Schnabel, and Xiahai Zhuang. Myops++: Multi-center myocardial pathology segmentation challenge. In CARE 2024: Compr ehensive Analysis and Retrieval of Medical Images . Springer , 2024. URL https://www.zmic.org.cn/care_ 2024/track4/ . Multi-center myocardial pathology segmentation from multi-sequence CMR data. 148 [675] Lei Li, V eronika A. Zimmer , Julia A. Schnabel, and Xiahai Zhuang. Whs++: Multi-center whole heart segmentation challenge. In CARE 2024: Compr ehensive Analysis and Re- trieval of Medical Images . Springer, 2024. URL https://www.zmic.org.cn/care_2024/ track5/ . Multi-center whole heart segmentation for se v en cardiac substructures. 148 106 [676] Y iming Xiao, Hassan Ri v az, Matthieu Chabanas, Maryse Fortin, Ines Machado, Y angming Ou, Mattias P . Heinrich, Julia A. Schnabel, Xia Zhong, Andreas Maier, W olfgang W ein, Roozbeh Shams, Samuel Kadoury , Da vid Drobn y , Marc Modat, and Ingerid Reinertsen. Evaluation of mri to ultrasound registration methods for brain shift correction: The curi- ous2018 challenge. IEEE T ransactions on Medical Imaging , 39(3):777–786, 2020. doi: 10.1109/TMI.2019.2935060. 148 , 153 [677] Y iming Xiao, Maryse Fortin, Geirmund Unsgård, Hassan Ri v az, and Ingerid Reinertsen. Ret- rospectiv e e v aluation of cerebral tumors (resect): a clinical database of pre-operati ve mri and intra-operativ e ultrasound in low-grade glioma sur geries. Medical Physics , 44(7):3875–3882, 2017. doi: 10.1002/mp.12268. 148 , 153 [678] Bahareh Behboodi, Francois-Xa vier Carton, Matthieu Chabanas, Sandrine De Ribaupierre, Ole Solheim, Bodil K. R. Munkvold, Hassan Ri v az, Y iming Xiao, and Ingerid Reinertsen. Open access segmentations of intraoperati ve brain tumor ultrasound images. Medical Physics , 51(9):6525–6532, 2024. doi: 10.1002/mp.17317. 148 , 153 [679] Reuben Dorent, Aaron Kujawa, Marina Ivory , Spyridon Bakas, Nicola Rieke, Samuel Joutard, Ben Glocker , Jor ge Cardoso, Marc Modat, Kayhan Batmanghelich, Arseniy Belkov , Maria Baldeon Calisto, Jae W on Choi, Benoit M Da want, Hexin Dong, Sergio Escalera, Y ubo Fan, Lasse Hansen, Mattias P Heinrich, Smriti Joshi, V ictoriya Kashtano v a, Hyeon Gyu Kim, Satoshi K ondo, Christian N Kruse, Susana K Lai-Y uen, Hao Li, Han Liu, Buntheng L y , Ipek Oguz, Hyungseob Shin, Boris Shirokikh, Zixian Su, Guotai W ang, Jianghao W u, Y anwu Xu, Kai Y ao, Li Zhang, Sébastien Ourselin, Jonathan Shapey , and T om V ercauteren. Crossmoda 2021 challenge: Benchmark of cross-modality domain adaptation techniques for v estibular schwannoma and cochlea segmentation. Medical Image Analysis , 83:102628, 2023. doi: 10.1016/j.media.2022.102628. 148 [680] Daniel S Marcus, T racy H W ang, Jamie Parker , John G Csernansky , John C Morris, and Randy L Buckner . Open access series of imaging studies (oasis): Cross-sectional mri data in young, middle aged, nondemented, and demented older adults. Journal of Cognitive Neuro- science , 19(9):1498–1507, 2007. doi: 10.1162/jocn.2007.19.9.1498. 148 [681] Daniel S. Marcus, Anthony F . Fotenos, John G. Csernansky , John C. Morris, and Randy L. Buckner . Open access series of imaging studies (oasis): Longitudinal mri data in nonde- mented and demented older adults. J ournal of Cognitive Neur oscience , 22(12):2677–2684, 2010. doi: 10.1162/jocn.2009.21407. 148 [682] N AMIC W iki contributors. 2009 prostate segmentation challenge miccai, 2017. URL https://www.na- mic.org/w/index.php?title=2009_prostate_segmentation_ challenge_MICCAI&oldid=97190 . [Online; accessed 19-August-2025]. 148 , 151 [683] Geert Litjens, Robert T oth, W endy van de V en, Caroline Hoeks, Sjoerd Kerkstra, Bram v an Ginneken, Graham V incent, Gwenael Guillard, Neil Birbeck, Jindang Zhang, Robin Strand, Filip Malmberg, Y angming Ou, Christos Dav atzikos, Matthias Kirschner, Florian Jung, Jing Y uan, W u Qiu, Qinquan Gao, Philip Eddie Edwards, Bianca Maan, Ferdinand van der Hei- jden, Soumya Ghose, Jhimli Mitra, Jason Dowling, Dean Barratt, Henkjan Huisman, and Anant Madabhushi. Evaluation of prostate segmentation algorithms for mri: the promise12 challenge. Medical Image Analysis , 18(2):359–373, 2014. doi: 10.1016/j.media.2013.12.002. URL https://doi.org/10.1016/j.media.2013.12.002 . 148 [684] Geert Litjens, Jurgen Futterer , and Henkjan Huisman. Data From Prostate-3T. The Cancer Imaging Archiv e, 2015. 148 [685] Brian N. Bloch, Anant Jain, and Cliv e C. Jaffe. Data from prostate-diagnosis [dataset]. The Cancer Imaging Archiv e, 2015. URL https://www.cancerimagingarchive.net/ collection/prostate- diagnosis/ . 149 [686] Samuel G. Armato, Henkjan Huisman, Karen Drukker , Lubomir Hadjiiski, Justin S. Kirby , Nicholas Petrick, George Redmond, Maryellen L. Giger , K enny Cha, Artem Mamonov , Jayashree Kalpathy-Cramer , and K eyv an Farahani. Prostatex challenges for computerized classification of prostate lesions from multiparametric magnetic resonance images. Journal of Medical Imaging , 5(4):044501, 2018. doi: 10.1117/1.JMI.5.4.044501. 149 107 [687] Geert Litjens, Oscar Debats, Jelle Barentsz, Nico Karssemeijer , and Henkjan Huisman. Spie- aapm prostatex challenge data (v ersion 2). The Cancer Imaging Archi ve, 2017. 149 [688] W ei Huang, Xi Li, Y Chen, Xi Li, M-C Chang, M.J. Oborski, D.I. Malyarenko, M Muzi, G.H. Jajamovich, A. Fedorov , A Tudorica, S.N. Gupta, C.M. Laymon, K.I. Marro, H.A. Dyvorne, J.V . Miller , D.P . Barbodiak, T .L. Chenev ert, T .E. Y ankeelov , J.M. Mountz, P .E. Kinahan, R. Kikinis, B. T aouli, F . Fennessy , and J. Kalpathy-Cramer . V ariations of dynamic contrast- enhanced magnetic resonance imaging in ev aluation of breast cancer therapy response: A multicenter data analysis challenge. T r anslational Oncology , 7(1):153–166, 2014. doi: 10. 1593/tlo.13838. 149 [689] W . Huang, C. Ryan, B. Beckett, A. T udorica, A. Mansoor , A. Afzal, M. Holtorf, and T . Aston. Qin-sarcoma, 2016. 149 [690] A. B. Mamonov and J. Kalpathy-Cramer . Data from qin gbm treatment response. The Cancer Imaging Archiv e. DOI:10.7937/k9/tcia.2016.nQF4gpn2, 2016. 149 [691] Kathleen M Schmainda, Melissa A Prah, Jennifer M Connelly , and Scott D Rand. Glioma dsc-mri perfusion data with standard imaging and rois. The Cancer Imaging Archiv e (TCIA), 2016. URL https://www.cancerimagingarchive.net/collection/ qin- brain- dsc- mri/ . 149 [692] Andrey Fedorov , Michael Schwier , David Clunie, Carsten Herz, Stev e Pieper, Ron Kikinis, Clare T empany , and Fiona Fennessy . An annotated test-retest collection of prostate multi- parametric mri. Scientific Data , 5:180281, 2018. doi: 10.1038/sdata.2018.281. 149 [693] Thomas E. Y ankeelov , Gregory S. Karczmar , and Richard G. Abramson. Data from qin- breast-02. The Cancer Imaging Archi ve, 2019. 149 [694] Shyam Natarajan, Ana Priester , Daniel Mar golis, Jiaoti Huang, and Leonard Marks. Prostate mri and ultrasound with pathology and coordinates of tracked biopsy (prostate-mri-us- biopsy). Data set, The Cancer Imaging Archiv e, 2020. URL https://doi.org/10.7937/ TCIA.2020.A61IOC1A . 149 , 153 [695] Quande Liu, Qi Dou, and Pheng-Ann Heng. Shape-aware meta-learning for generalizing prostate mri segmentation to unseen domains. In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , pages 329–338. Springer , 2020. 149 [696] A. Meyer , D. Schindele, D. von Reibnitz, M. Rak, M. Schostak, and C. Hansen. Prostatex zone segmentations, 2020. URL https://www.cancerimagingarchive.net/ analysis- result/prostatex- seg- zones/ . 149 [697] Anneke Meyer , Grzegorz Chlebus, Marko Rak, Daniel Schindele, Martin Schostak, Bram v an Ginneken, Andrea Schenk, Hans Meine, Horst K. Hahn, Andreas Schreiber , and Christian Hansen. Anisotropic 3d multi-stream cnn for accurate prostate segmentation from multi- planar mri. Computer Methods and Pr ogr ams in Biomedicine , 200:105821, 2020. doi: 10. 1016/j.cmpb .2020.105821. 149 [698] Anindo Saha, Joeran Bosma, Jasper T wilt, Bram van Ginneken, Derya Y akar , Mattijs Elschot, Jeroen V eltman, Jurgen Fütterer , Maarten de Rooij, et al. Artificial intelligence and radiol- ogists at prostate cancer detection in MRI — the PI-CAI challenge. In Medical Imaging with Deep Learning, short paper tr ack , 2023. URL https://openreview.net/forum? id=XfXcA9- 0XxR . 149 [699] Zachary M. C. Baum, Shaheer U. Saeed, Zhe Min, Y ipeng Hu, and Dean C. Barratt. Mr to ultrasound registration for prostate challenge - dataset. Zenodo, https://doi.org/10.5281/zenodo.8004388, 2023. V ersion 1.1.0. 149 , 153 [700] T obias Heimann, Bryan J. Morrison, Martin A. Styner, Marc Niethammer, and Simon K. W arfield. Se gmentation of knee images: A grand challenge. In MICCAI W orkshop on Medical Image Analysis for the Clinic – A Grand Challenge , pages 207–214, 2010. URL https: //www.ski10.org/ . 149 108 [701] Razv an V . Marinescu, Neil P . Oxtoby , Alexandra L. Y oung, Esther E. Bron, Arthur W . T oga, Michael W . W einer, Frederik Barkhof, Nick C. Fox, Polina Golland, Stefan Klein, and Daniel C. Alexander . Predicting alzheimer’ s disease progression: Results from the tad- pole challenge. Alzheimer’s & Dementia , 16:e039538, 2020. doi: 10.1002/alz.039538. 149 , 154 [702] Oli vier Commo wick, Frédéric Cerv enansky , François Cotton, and Michel Dojat, editors. MSSEG-2 Challenge Pr oceedings: Multiple Scler osis Ne w Lesions Se gmentation Challenge Using a Data Manag ement and Pr ocessing Infrastructur e , 2021. URL https://portal. fli- iam.irisa.fr/msseg- 2/data/ . 149 [703] Oli vier Commowick, Michaël Kain, Romain Casey , Roxana Ameli, Jean-Christophe Ferré, Anne Kerbrat, Thomas T ourdias, Frédéric Cervenansk y , Sorina Camarasu-Pop, T ristan Glatard, Sandra V ukusic, Gilles Edan, Christian Barillot, Michel Dojat, and François Cotton. Multiple sclerosis lesions segmentation from multiple e xperts: The miccai 2016 challenge dataset. Neur oIma ge , 244:118589, 2021. doi: 10.1016/j.neuroimage.2021.118589. 149 [704] Martin Styner, Joohwi Lee, Brian Chin, Matthew S Chin, Olivier Commo wick, Hoai-Huong T ran, V alerie Jewells, and Simon W arfield. 3d segmentation in the clinic: A grand challenge ii: Ms lesion se gmentation. MID AS J ournal , 2008:1–6, 2008. doi: 10.54294/lmkqvm. 149 [705] Jason R. T aylor, Nitin W illiams, Rhodri Cusack, T ibor Auer , Meredith A. Shafto, Marie Dixon, Lorraine K. T yler , Cam-CAN, and Richard N. Henson. The cambridge centre for ageing and neuroscience (cam-can) data repository: Structural and functional mri, meg, and cognitiv e data from a cross-sectional adult lifespan sample. Neur oImage , 144:262–269, 2017. doi: 10.1016/j.neuroimage.2015.09.018. 149 [706] Oskar Maier , Bjoern H. Menze, Janina von der Gablentz, Levin Häni, Mattias P . Heinrich, Matthias Liebrand, Stefan W inzeck, Abdul Basit, Paul Bentley , Liang Chen, Daan Chris- tiaens, Francis Dutil, Karl Egger , Chaolu Feng, Ben Glocker , Michael Götz, T om Haeck, Hanna-Leena Halme, Mohammad Havaei, Khan M. Iftekharuddin, Pierre-Marc Jodoin, K on- stantinos Kamnitsas, Elias Kellner , Antti K orvenoja, Hugo Larochelle, Christian Ledig, Jia- Hong Lee, Frederik Maes, Qaiser Mahmood, Klaus H. Maier-Hein, Richard McKinley , John Muschelli, Chris Pal, Linmin Pei, Janaki Raman Rangarajan, Syed M. S. Reza, Da vid Robben, Daniel Rueckert, Eero Salli, Paul Suetens, Ching-W ei W ang, Matthias Wilms, Jan S. Kirschke, Ulrike M. Krämer, Thomas F . Münte, Peter Schramm, Roland W iest, Heinz Han- dels, and Mauricio Reyes. Isles 2015 - a public e valuation benchmark for ischemic stroke lesion segmentation from multispectral mri. Medical Image Analysis , 35:250–269, 2016. doi: 10.1016/j.media.2016.07.009. 149 [707] Stefan W inzeck, Arsany Hakim, Richard McKinley , Jose AASR Pinto, V ictor Alv es, Car- los Silva, Maxim Pisov , Egor Kri v ov , Mikhail Belyaev , Miguel Monteiro, Arlindo Oli veira, Y oungwon Choi, Myunghee Cho Paik, Y ongchan Kwon, Hanbyul Lee, Beom Joon Kim, Joong-Ho W on, Mobarakol Islam, Hongliang Ren, David Robben, Paul Suetens, Enhao Gong, Y ilin Niu, Junshen Xu, John M Pauly , Christian Lucas, Mattias P Heinrich, Luis C Rivera, Laura S Castillo, Laura A Daza, Andrew L Beers, Pablo Arbelaezs, Oskar Maier , Ken Chang, James M Brown, Jayashree Kalpathy-Cramer , Greg Zaharchuk, Roland Wiest, and Mauricio Reyes. Isles 2016 and 2017—benchmarking ischemic stroke lesion outcome prediction based on multispectral mri. F r ontier s in Neur olo gy , 9:679, 2018. doi: 10.3389/fneur .2018.00679. 149 [708] Arsany Hakim, Soren Christensen, Stefan W inzeck, Maarten G Lansberg, Mark W Par- sons, Christian Lucas, David Robben, Roland W iest, Mauricio Reyes, and Gre g Zaharchuk. Predicting infarct core from computed tomography perfusion in acute ischemia with ma- chine learning: Lessons from the isles challenge. Str ok e , 52(7):2328–2337, 2021. doi: 10.1161/STR OKEAHA.120.030696. 149 [709] Moritz R. Hernandez Petzsche, Ezequiel de la Rosa, Uta Hanning, Roland Wiest, W aldo V alenzuela, Mauricio Re yes, Maria Ines Meyer , Sook-Lei Liew , Florian Kofler , Iv an Ezhov , David Robben, Ale xander Hutton, T assilo Friedrich, T eresa Zarth, Johannes Bürkle, The Anh Baran, Björn Menze, Gabriel Broocks, Lukas Meyer , Claus Zimmer , T obias Boeckh-Behrens, 109 Maria Berndt, Benno Ikenberg, Benedikt W iestler, and Jan S. Kirschke. Isles 2022: A multi- center magnetic resonance imaging stroke lesion segmentation dataset. Scientific Data , 9(1): 762, 2022. doi: 10.1038/s41597- 022- 01875- 5. 149 [710] Hugo J. Kuijf, J. Matthijs Biesbroek, Jeroen de Bresser, Rutger Heinen, Simon Ander- matt, Mariana Bento, Matt Berseth, Mikhail Belyaev , M. Jorge Cardoso, Adria Casamitjana, D. Louis Collins, Mahsa Dadar, Achilleas Geor giou, Mohsen Ghafoorian, Dakai Jin, April Khademi, Jesse Knight, Hongwei Li, Xa vier Llado, Miguel Luna, Qaiser Mahmood, Richard McKinley , Alireza Mehrtash, Sebastien Ourselin, Bo yong Park, Hyunjin Park, Sang Hyun Park, Simon Pezold, Elodie Puybareau, Leticia Rittner , Carole H. Sudre, Sergi V alverde, V eronica V ilaplana, Roland W iest, Y ongchao Xu, Ziyue Xu, Guodong Zeng, Jianguo Zhang, Guoyan Zheng, Christopher Chen, W iesje v an der Flier , Frederik Barkhof, Max A. V ierge v er , and Geert Jan Biessels. Standardized assessment of automatic segmentation of white matter hyperintensities; results of the wmh segmentation challenge. IEEE T ransactions on Medical Imaging , 38(11):2556–2568, 2019. doi: 10.1109/TMI.2019.2905770. 149 [711] Bjoern H. Menze, Andras Jakab, Stefan Bauer , Jayashree Kalpathy-Cramer , Ke yvan Fara- hani, Justin Kirby , Y uliya Burren, Nicole Porz, Johannes Slotboom, Roland W iest, Levente Lanczi, Elizabeth Gerstner , Marc-André W eber, T al Arbel, Brian B. A vants, Nicholas A yache, Patricia Buendia, D. Louis Collins, Nicolas Cordier , Jason J. Corso, Antonio Criminisi, Tilak Das, Hervé Delingette, Ça ˘ gatay Demiralp, Christopher R. Durst, Michel Dojat, Senan Do yle, Joana Festa, Florence F orbes, Ezequiel Geremia, Ben Glocker , Polina Golland, Xiaotao Guo, Andac Hamamci, Khan M. Iftekharuddin, Raj Jena, Nigel M. John, Ender K onukoglu, Danial Lashkari, José António Mariz, Raphael Meier , Sérgio Pereira, Doina Precup, Stephen J. Price, T ammy Riklin Ravi v , Syed M. S. Reza, Michael Ryan, Duygu Sarikaya, La wrence Schwartz, Hoo-Chang Shin, Jamie Shotton, Carlos A. Silv a, Nuno Sousa, Nagesh K. Subbanna, Ga- bor Szekely , Thomas J. T aylor, Owen M. Thomas, Nicholas J. T ustison, Gozde Unal, Flor V asseur , Max W intermark, Dong Hye Y e, Liang Zhao, Binsheng Zhao, Darko Zikic, Marcel Prastawa, Mauricio Reyes, and K oen V an Leemput. The multimodal brain tumor image seg- mentation benchmark (BRA TS). IEEE T ransactions on Medical Ima ging , 34(10):1993–2024, 2015. doi: 10.1109/TMI.2014.2377694. 149 [712] Spyridon Bakas, Bjoern Menze, Christos Davatzik os, Jayashree Kalpathy-Cramer , K ey- van Farahani, Michel Bilello, Suyash Mohan, John B. Freymann, Justin S. Kirby , Man- meet Ahluw alia, V olodymyr Statsevych, Raymond Huang, Hassan Fathallah-Shaykh, Roland W iest, Andras Jakab, Rivka R. Colen, Aikaterini Kotrotsou, Daniel Marcus, Mikhail Milchenko, Arash Nazeri, Marc-Andre W eber , Abhishek Mahajan, and Ujjwal Baid. Mic- cai brain tumor se gmentation (brats) 2020 benchmark: Prediction of surviv al and pseudopro- gression. Zenodo, 2020. 23rd International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2020), Lima, Peru, 4-8 October 2020. 149 [713] Ujjwal Baid, Satyam Ghodasara, Suyash Mohan, Michel Bilello, Evan Calabrese, Errol Colak, Ke yv an Farahani, Jayashree Kalpathy-Cramer , Felipe C. Kitamura, Sarthak Pati, Luciano M. Prevedello, Jeffre y D. Rudie, Chiharu Sako, Russell T . Shinohara, T imothy Bergquist, Rong Chai, James Eddy , Julia Elliott, W alter Reade, Thomas Schaffter , Thomas Y u, Jiaxin Zheng, Ahmed W . Moawad, Luiz Otavio Coelho, Olivia McDonnell, Elka Miller , Fann y E. Moron, Mark C. Oswood, Robert Y . Shih, Loizos Siakallis, Y ulia Bronstein, James R. Mason, Anthony F . Miller , Gagandeep Choudhary , Aanchal Agarwal, Cristina H. Besada, Jamal J. Derakhshan, Mariana C. Diogo, Daniel D. Do-Dai, Luciano Farage, John L. Go, Mohiuddin Hadi, V irginia B. Hill, Michael Iv , David Joyner , Christie Lincoln, Eyal Lotan, Asako Miyakoshi, Mariana Sanchez-Montano, Jaya Nath, Xuan V . Nguyen, Manal Nicolas-Jilwan, Johanna Ortiz Jimenez, Kerem Ozturk, Bojan D. Petrovic, Chintan Shah, Lubdha M. Shah, Manas Sharma, Onur Simsek, Achint K. Singh, Salil Soman, V olodymyr Statsevych, Brent D. W einberg, Robert J. Y oung, Ichiro Ikuta, Amit K. Agarwal, Sword C. Cambron, Richard Silbergleit, Alexandru Dusoi, Alida A. Postma, Laurent Letourneau- Guillon, Gloria J. Guzman Perez-Carrillo, Atin Saha, Neetu Soni, Greg Zaharchuk, V ahe M. Zohrabian, Y ingming Chen, Milos M. Cekic, Akm Rahman, Juan E. Small, V arun Sethi, Christos Da vatzik os, John Mongan, Christopher Hess, Soonmee Cha, Javier V illanue v a- Meyer , John B. Freymann, Justin S. Kirby , Benedikt W iestler , Priscila Cri vellaro, Ri vka R. Colen, Aikaterini Kotrotsou, Daniel Marcus, Mikhail Milchenko, Arash Nazeri, Hassan 110 Fathallah-Shaykh, Roland W iest, Andras Jakab, Marc-Andre W eber , Abhishek Mahajan, Bjoern Menze, Adam E. Flanders, and Spyridon Bakas. The rsna-asnr-miccai brats 2021 benchmark on brain tumor segmentation and radiogenomic classification. arXiv pr eprint arXiv:2107.02314 , 2021. doi: 10.48550/arXiv .2107.02314. 149 [714] Spyridon Bakas, Bjoern Menze, Jan Kirschke, Benedikt W iestler , Juan Eugenio Iglesias, Mar - ius George Linguraru, Adam Flanders, Michel Bilello, John Freymann, K eyv an Farahani, and Christos Dav atzikos. The international brain tumor segmentation (brats) cluster of challenges. In Pr oceedings of the MICCAI 2023 Challenges , pages 1–10. Springer , 2023. 149 [715] K elly Payette, Priscille de Dumast, Hamza Kebiri, Hui Ji, Md Mahfuzur Rahman Siddiquee, Daguang Xu, and Roxane Licandro. An automatic multi-tissue human fetal brain segmenta- tion benchmark using the fetal tissue annotation dataset. Scientific Data , 8:167, 2021. doi: 10. 1038/s41597- 021- 00946- 3. URL https://doi.org/10.1038/s41597- 021- 00946- 3 . 149 [716] K elly Payette, Celine Steger , Priscille de Dumast, Andras Jakab, Meritxell Bach Cuadra, Lana V asung, Roxane Licandro, Matthe w Barkovich, and Hongwei Li. Fetal tissue annotation challenge 2022 dataset. Zenodo, https://doi.org/10.5281/zenodo.6683366 , 2022. V ersion v2; associated with MICCAI 2022. 149 [717] Jure Zbontar, Florian Knoll, Anuroop Sriram, T ullie Murrell, Zhengnan Huang, Matthe w J. Muckley , Aaron Defazio, Ruben Stern, Patricia Johnson, Mary Bruno, Marc Parente, Krzysztof J. Geras, Joe Katsnelson, Hersh Chandarana, Zizhao Zhang, Michal Drozdzal, Adriana Romero, Michael Rabbat, Pascal V incent, Nafissa Y akubov a, James Pinkerton, Duo W ang, Erich Owens, C. Lawrence Zitnick, Michael P . Recht, Daniel K. Sodickson, and Yvonne W . Lui. fastmri: An open dataset and benchmarks for accelerated mri. ArXiv e- prints , 2018. 149 [718] Ale Neubert, Jurgen Fripp, Craig Engstrom, Duncan W alker , Raphael Schwarz, and Stuart Crozier . Automatic quantification of 3d morphology and appearance of intervertebral discs in high resolution mri. In Pr oceedings of the 21st Annual Meeting of the International Society for Magnetic Resonance in Medicine (ISMRM) , page 1612, 2013. 149 [719] Bharat B. Biswal, Maarten Mennes, Xi-Nian Zuo, Sanjay Gohel, Christina Kelly , Stephen M. Smith, Christian F . Beckmann, Justin S. Adelstein, Randy L. Buckner , Stanley Colcombe, Alexander -M Dogono wski, Monika Ernst, Damien A. Fair , Michelle Hampson, Matthew J. Hoptman, James S. Hyde, V esa J. Kiviniemi, Rolf Kötter , Shi-Jen Li, Ching-Po Lin, Michael J. Lowe, Charles E. Mackay , David J. Madden, Kristof fer H. Madsen, Daniel S. Margulies, Helen S. Mayberg, Ke vin McMahon, Catherine S. Monk, Ste wart H. Mostofsky , Bonnie J. Nagel, James J. Pekar , Scott J. Peltier , Ste ven E. Petersen, V iktor Riedl, Serge A. R. B. Rombouts, Barbara Rypma, Bradley L. Schlaggar , Sabine Schmidt, Rachael D. Seidler , Gre g J. Siegle, Christian Sorg, Guo-Jun T eng, Juha V eijola, Arno V illringer , Mar- tin W alter , Lei W ang, Xin-Chen W eng, Susan Whitfield-Gabrieli, Peter W illiamson, Chris- tian W indischberger , Y u-Feng Zang, Hong-Y u Zhang, F . Xavier Castellanos, and Michael P . Milham. T ow ard discovery science of human brain function. Pr oceedings of the National Academy of Sciences of the United States of America , 107(10):4734–4739, 2010. doi: 10.1073/pnas.0911855107. 149 [720] Jin W ang, Marisa N. L ytle, Y ael W eiss, Brianna L. Y amasaki, and James R. Booth. A longi- tudinal neuroimaging dataset on language processing in children ages 5, 7, and 9 years old. Scientific Data , 9:4, 2022. doi: 10.1038/s41597- 021- 01106- 3. 149 [721] David Newitt, Nola Hylton, on behalf of the I-SPY 1 Network, and A CRIN 6657 T rial T eam. Multi-center breast dce-mri data and segmentations from patients in the i-spy 1/acrin 6657 trials. The Cancer Imaging Archiv e, 2016. URL https://doi.org/10.7937/K9/TCIA. 2016.HdHpgJLK . 149 [722] Paul Kinahan, Mark Muzi, Brent Bialecki, Barbara Herman, and Lauren Coombs. Acrin- contralateral-breast-mr (acrin 6667). Data set. The Cancer Imaging Archiv e, 2021. URL https://doi.org/10.7937/Q1EE- J082 . 149 111 [723] David C. Ne witt, Sav annah C. P artridge, Zheng Zhang, Jessica E. Gibbs, Thomas Chene vert, Mark Rosen, Patrick J. Bolan, Helga S. Marques, Justin Romanoff, Lara Cimino, Bonnie N. Joe, Heidi Umphrey , Haydee Ojeda-Fournier , Basak Dogan, Karen Y . Oh, Hiroyuki Abe, Jennifer Drukteinis, Laura J. Esserman, and Nola M. Hylton. Acrin 6698/i-spy2 breast dwi. Data set. The Cancer Imaging Archive, 2021. URL https://doi.org/10.7937/tcia. kk02- 6d95 . 149 [724] Michael W . W einer, Danielle Harve y , Jacqueline Hayes, Susan M. Landau, Paul S. Aisen, Ronald C. Petersen, Duygu T osun, Dallas P . V eitch, Clifford R. Jr Jack, Charles Decarli, An- drew J. Saykin, Jordan Grafman, Thomas C. Ne ylan, and Department of Defense Alzheimer’ s Disease Neuroimaging Initiati v e. Effects of traumatic brain injury and posttraumatic stress disorder on de v elopment of alzheimer’ s disease in vietnam veterans using the alzheimer’ s dis- ease neuroimaging initiati ve: Preliminary report. Alzheimers Dement (N Y) , 3(2):177–188, 2017. doi: 10.1016/j.trci.2017.02.005. 149 , 154 [725] Francisca S. Rodriguez, Ling Zheng, and Helena C. Chui. Psychometric characteristics of cognitiv e reserve: how high education might improv e certain cognitive abilities in aging. De- mentia and Geriatric Cognitive Disorders , 47(4-6):335–344, 2019. doi: 10.1159/000501150. 149 [726] Kathryn A Ellis, Ashley I Bush, David Darby , Daniela De Fazio, Jonathan Foster , Peter Hudson, Nicola T Lautenschlager , Nat Lenzo, Ralph N Martins, Paul Maruff, Colin L Mas- ters, Andre w Milner , K erryn E Pike, Christopher Rowe, Greg Sa v age, Cassandra Szoeke, Ke vin T addei, V ictor L V illemagne, Michael W oodward, David Ames, and AIBL Re- search Group. The australian imaging, biomarkers and lifestyle (aibl) study of aging: methodology and baseline characteristics of 1112 individuals recruited for a longitudinal study of alzheimer’ s disease. International Psychog eriatrics , 21(4):672–687, 2009. doi: 10.1017/S1041610209009405. 149 , 154 [727] Lukas Snoek, Maite M. van der Miesen, T inka Beemsterboer , Andries van der Leij, An- nemarie Eigenhuis, and H. Stev en Scholte. The amsterdam open mri collection, a set of multimodal mri datasets for indi vidual difference analyses. Scientific Data , 8(1):85, 2021. doi: 10.1038/s41597- 021- 00870- 6. 149 [728] Saloni Krishnan, Salomi S Asaridou, Gabriel J Cler, Harriet J Smith, Hannah E W illis, Máiréad P Healy , P aul A Thompson, Dorothy VM Bishop, and Kate E W atkins. Functional organisation for verb generation in children with de v elopmental language disorder . Neur oIm- age , 226:117599, 2021. doi: 10.1016/j.neuroimage.2020.117599. 149 [729] Macarena Suarez-Pellicioni, Marisa N. L ytle, Jessica W . Y ounger , and James R. Booth. A longitudinal neuroimaging dataset on arithmetic processing in school children. Scientific Data , 6(1):190040, 2019. doi: 10.1038/sdata.2019.40. 150 [730] Matthe w J Kempton, T oby SA Underw ood, Sarah Brunton, Fotios Stylios, Anke Schmechtig, Ulrich Ettinger , Matthew S Smith, Simon Lovestone, W illiam R Crum, Sophia Frangou, Stev e CR W illiams, and Andre w Simmons. A comprehensi ve testing protocol for mri neu- roanatomical se gmentation techniques: Evaluation of a nov el lateral ventricle se gmentation method. Neur oImage , 58(4):1051–1059, 2011. doi: 10.1016/j.neuroimage.2011.06.080. 150 [731] Endre Grøvik, Darvin Y i, Elizabeth T ong, Michael Iv , Daniel Rubin, Greg Zaharchuk, and Ghiam Y amin. Deep learning enables automatic detection and segmentation of brain metas- tases on multisequence mri. J ournal of Magnetic Resonance Imaging , 51(1):175–182, 2020. doi: 10.1002/jmri.26766. 150 [732] David Newitt and Nola M. Hylton. Single site breast dce-mri data and se gmentations from patients undergoing neoadjuvant chemotherapy . Data set, The Cancer Imaging Archi ve, 2016. URL https://doi.org/10.7937/K9/TCIA.2016.QHsyhJKy . 150 [733] Li-Ming Hsu, W oomi Ban, Tzu-Hao Chao, Shihui Song, Da vid H. Cerri, Laura W alton, Mar- garet Broadwater , Seung-Hwan Lee, and Y en-Y u Ian Shih. Camri rat brain mri data. Open- Neuro, 2021. URL https://openneuro.org/datasets/ds002870/versions/1.0.1 . 150 112 [734] Dorit Kliemann, Ralph Adolphs, T im Armstrong, Paola Galdi, David A. Kahn, T essa Rusch, A. Zeynep Enkavi, Deuhua Liang, Ste ven Lograsso, W enying Zhu, Rona Y u, Remya Nair , L ynn K. Paul, and J. Michael T yszka. Caltech conte center , a multimodal data resource for e xploring social cognition and decision-making. Scientific Data , 9(1):138, 2022. doi: 10.1038/s41597- 022- 01171- 2. 150 [735] Sangil Lee and Joseph Kable. Cognitive training dataset. OpenNeur o , 2020. URL https: //openneuro.org/datasets/ds002843/versions/1.0.1 . 150 [736] Ali Khan, Maged Goubran, Da vid A. Rudk o, Joseph Gati, T rev or Szekeres, Colin Holmes, and T erry Peters. High-resolution 3t and 7t extension of the colin27 atlas for deep-brain targeting. In Pr oceedings of the Or ganization for Human Brain Mapping Annual Meeting , Hambur g, Germany , June 2014. 150 [737] T akuya Ito, Kaustubh R. Kulkarni, Douglas H. Schultz, Ravi D. Mill, Richard H. Chen, Levi I. Solomyak, and Michael W . Cole. Cogniti ve task information is transferred between brain regions via resting-state netw ork topology . Nature Communications , 8(1):1027, 2017. doi: 10.1038/s41467- 017- 01000- w. 150 [738] David A.A. Baranger , Y aroslav O. Halchenko, Skye Satz, Rachel Ragozzino, Satish Iyengar , Holly A. Swartz, and Anna Manelis. Aberrant le vels of cortical myelin distinguish indi viduals with depressive disorders from healthy controls. Neur oIma ge: Clinical , 32:102790, 2021. doi: 10.1016/j.nicl.2021.102790. 150 [739] Marisa N. L ytle, T ali Bitan, and James R. Booth. A neuroimaging dataset on orthographic, phonological and semantic word processing in school-aged children. Data in Brief , 28: 105091, 2020. doi: 10.1016/j.dib.2019.105091. 150 [740] Kate Nussenbaum and Catherine A Hartley . Dev elopmental change in prefrontal cortex re- cruitment supports the emergence of v alue-guided memory . eLife , 10:e69796, 2021. doi: 10.7554/eLife.69796. 150 [741] Ashirbani Saha, Michael R Harowicz, Lars J Grimm, Connie E Kim, Sujata V Ghate, Ruth W alsh, and Maciej A Mazurowski. A machine learning approach to radiogenomics of breast cancer: a study of 922 subjects and 529 dce-mri features. British Journal of Cancer , 119 (4):508–516, 2018. doi: 10.1038/s41416- 018- 0185- 8. URL https://doi.org/10.1038/ s41416- 018- 0185- 8 . 150 [742] Christian Meyer , Srikanth Padmala, and Luiz Pessoa. Dynamic threat processing. J ournal of Cognitive Neur oscience , 31(4):522–542, 2018. doi: 10.1162/jocn_a_01363. 150 [743] W illiam K. Lloyd, Jayne Morriss, Birthe Macdonald, Karin Joanknecht, Julie Nihouarn, and Carien M. v an Reekum. Longitudinal change in ex ecutiv e function is associated with im- paired top-do wn frontolimbic regulation during reappraisal in older adults. Neur oImage , 225: 117488, 2021. doi: 10.1016/j.neuroimage.2020.117488. 150 [744] Carrie Elizabeth Gold. Exploring the resting state neural activity of monolinguals and late and early bilinguals. Master’ s thesis, Brigham Y oung University , 2018. URL: http://hdl.lib .byu.edu/1877/etd9704. 150 [745] A vram J. Holmes, Marisa O. Hollinshead, T imothy M. O’Keefe, V ictor I. Petrov , Gabriele R. Fariello, Lawrence L. W ald, Bruce Fischl, Bruce R. Rosen, Ross W . Mair , Joshua L. Roffman, Jordan W . Smoller, and Randy L. Buckner . Brain genomics superstruct project initial data release with structural, functional, and behavioral measures. Scientific Data , 2:150031, 2015. doi: 10.1038/sdata.2015.31. URL http://dx.doi.org/10.1038/sdata.2015.31 . 150 [746] Biomedical Image Analysis Group, Imperial College London. Ixi dataset. http://brain- dev elopment.or g/ixi-dataset/. Accessed: 2025-08-19. 150 [747] J Hirvasniemi, J Runhaar , RA v an der Heijden, M Zokaeinikoo, M Y ang, X Li, J T an, HR Ra- jamohan, Y Zhou, CM Deniz, F Cali v a, C Iriondo, JJ Lee, F Liu, AM Martinez, N Namiri, V Pedoia, E P anfilov , N Bayramoglu, HH Nguyen, MT Nieminen, S Saarakkala, A Tiulpin, E Lin, A Li, V Li, EB Dam, AS Chaudhari, R Kijo wski, S Bierma-Zeinstra, EHG Oei, and 113 S Klein. The knee osteoarthritis prediction (knoap2020) challenge: An image analysis chal- lenge to predict incident symptomatic radiographic knee osteoarthritis from mri and x-ray im- ages. Osteoarthritis and Cartilage , 31(1):115–125, 2023. doi: 10.1016/j.joca.2022.10.001. 150 [748] Bradley Erickson, Zeynettin Akkus, Jesse Sedlar , and Panagiotis K orfiatis. Data from lgg- 1p19qdeletion (version 2). Data set, 2017. URL https://doi.org/10.7937/K9/TCIA. 2017.DWEHTZ9V . 150 [749] T ommaso Di Noto, Guillaume Marie, Sebastien T ourbier , Y asser Alemán-Gómez, Oscar Es- teban, Guillaume Saliou, Meritxell Bach Cuadra, Patric Hagmann, and Jonas Richiardi. T o- wards automated brain aneurysm detection in tof-mra: Open data, weak labels, and anatomi- cal knowledge. Neur oinformatics , 2022. doi: 10.1007/s12021- 022- 09597- 0. 150 [750] Leon Y Cai, Qi Y ang, Praitayini Kanakaraj, V ishwesh Nath, Allen T Ne wton, Heidi A Ed- monson, Jef frey Luci, Benjamin N Conrad, Ga vin R Price, Colin B Hansen, Cailey I Ker - ley , Karthik Ramadass, Fang-Cheng Y eh, Hakmook Kang, Eleftherios Garyfallidis, Maxime Descoteaux, Francois Rheault, K urt G Schilling, and Bennett A Landman. Masiv ar: Multi- site, multiscanner , and multisubject acquisitions for studying v ariability in diffusion weighted magnetic resonance imaging. Magnetic Resonance in Medicine , 86(6):3304–3320, 2021. doi: 10.1002/mrm.28926. 150 [751] Ian B Malone, David Cash, Gerard R Ridgway , David G MacManus, Sebastien Ourselin, Nick C Fox, and Jonathan M Schott. Miriad—public release of a multiple time point alzheimer’ s mr imaging dataset. Neur oImag e , 70:33–36, 2013. doi: 10.1016/j.neuroimage. 2012.12.044. 150 [752] Anahit Babayan, Miray Erbey , Deniz Kumral, Janis D Reinelt, Andrea M F Reiter , Josefin Röbbig, H Lina Schaare, Marie Uhlig, Alfred Anwander , Pierre-Louis Bazin, Annette Horstmann, Leonie Lampe, V adim V Nikulin, Hadas Okon-Singer , Sven Preusser , André Pampel, Christiane S Rohr , Julia Sacher, Angelika Thöne-Otto, Sabrina T rapp, Till Nier- haus, Denise Altmann, Katrin Arelin, Maria Blöchl, Edith Bongartz, P atric Breig, Elena Ces- naite, Sufang Chen, Roberto Cozatl, Saskia Czerwonatis, Gabriele Dambrauskaite, Maria Dreyer , Jessica Enders, Melina Engelhardt, Marie Michele Fischer, Norman Forschack, Jo- hannes Golchert, Laura Golz, C Alexandrina Guran, Susanna Hedrich, Nicole Hentschel, Daria I Hoffmann, Julia M Huntenburg, Rebecca Jost, Anna K osatschek, Stella Kunzen- dorf, Hannah Lammers, Mark E Lauckner, K eyv an Mahjoory , Ahmad S Kanaan, Natacha Mendes, Ramona Menger , Enzo Morino, Karina Näthe, Jennifer Neubauer , Handan Noyan, Sabine Oligschläger , Patricia Panczyszyn-T rze wik, Dorothee Poehlchen, Nadine Putzke, Sab- rina Roski, Marie-Catherine Schaller , Anja Schieferbein, Benito Schlaak, Robert Schmidt, Krzysztof J Gorgole wski, Hanna Maria Schmidt, Anne Schrimpf, Sylvia Stasch, Maria V oss, Annett W iedemann, Daniel S Margulies, Michael Gaebler , and Arno V illringer . A mind- brain-body dataset of mri, eeg, cognition, emotion, and peripheral physiology in young and old adults. Scientific Data , 6:180308, 2019. doi: 10.1038/sdata.2018.308. 150 [753] Li via T omov a, Kimberly L. W ang, T odd Thompson, Gillian A. Matthe ws, Akiko T aka- hashi, Kay M. T ye, and Rebecca Saxe. Acute social isolation ev okes midbrain crav- ing responses similar to hunger . Natur e Neur oscience , 23(12):1597–1605, 2020. doi: 10.1038/s41593- 020- 00742- z. 150 [754] Hilary Richardson, Grace Lisandrelli, Alexa Riobueno-Naylor , and Rebecca Saxe. Develop- ment of the social brain from age three to twelve years. Natur e Communications , 9(1):1027, 2018. doi: 10.1038/s41467- 018- 03399- 2. 150 [755] Joanes Grandjean, Claudio Canella, Charlotte Anckaerts, Gökberk A yrancı, Sana Bougacha, Thomas Bienert, David Buehlmann, Laura Coletta, Domenico Gallino, Nicola Gass, Coralie M. Garin, Romain E. Jacob, Deniz Kirik, Jun Li, Émilien Macé, Dan Madularu, Anne E. Mechling, Stephen J. Sawiak, Pekka Stenroos, T omokazu Tsurugizawa, Annemie van der Linden, V alerio Zerbi, Markus W enk, T obias K ober , Christa Baltes, Markus Rudin, Sophie Achard, Thomas Knöpfel, N. Jon Shah, Karl Deisseroth, Jukka K. Huttunen, and Alessandro Gozzi. Common functional netw orks in the mouse brain re v ealed by multi-centre 114 resting-state fmri analysis. Neur oImage , 205:116278, 2020. doi: 10.1016/j.neuroimage.2019. 116278. 150 [756] Jonathan D. Po wer , Mark Plitt, Stephen J. Gotts, Prantik Kundu, V alerie V oon, Peter A. Bandettini, and Alex Martin. Ridding fmri data of motion-related influences: Remov al of signals with distinct spatial and physical bases in multiecho data. Pr oceedings of the Na- tional Academy of Sciences of the United States of America , 115(9):E2105–E2114, 2018. doi: 10.1073/pnas.1720985115. 150 [757] Rotem Botvinik-Nezer , Roni Iwanir , Felix Holzmeister , Jür gen Huber , Magnus Johannesson, Michael Kirchler , Anna Dreber, Colin F Camerer, Russell A Poldrack, and T om Schonberg. fmri data of mixed g ambles from the neuroimaging analysis replication and prediction study . Scientific Data , 6:106, 2019. doi: 10.1038/s41597- 019- 0113- 7. 150 [758] Samuel A. Nastase, Y un Fei Liu, Hanna Hillman, Asieh Zadbood, Liat Hasenfratz, Neggin Kesha v arzian, Janice Chen, Christopher J. Honey , Y aara Y eshurun, Mor Rege v , Mai Nguyen, Claire H. C. Chang, Christopher Baldassano, Olg a Lositsky , Erez Simony , Michael A. Cho w , Y uan Chang Leong, Paula P . Brooks, Emily Micciche, Gina Choe, Ariel Goldstein, T amara V anderwal, Y aroslav O. Halchenko, Kenneth A. Norman, and Uri Hasson. The “narrativ es” fmri dataset for e valuating models of naturalistic language comprehension. Scientific Data , 8 (1):250, 2021. doi: 10.1038/s41597- 021- 01033- 3. 150 [759] Michael Hanke, Felix Baumgartner , V ittorio Iacovella, Patricia Broderick, Uri Hasson, Thorsten Kahnt, Thomas Y ates, Benedikt A. Poser , Rainer Goebel, and Arno V illringer . A naturalistic neuroimaging database for understanding the brain using ecological stimuli. Sci- entific Data , 8(1):1–18, 2020. doi: 10.1038/s41597- 020- 00680- 2. 150 [760] R Nathan Spreng, Roni Setton, Udi Alter , Benjamin N Cassidy , Bri Darboh, Elizabeth DuPre, Karin Kantarovich, Amber W Lockro w , Laetitia Mwilambwe-Tshilobo, W en-Ming Luh, Prantik Kundu, and Gary R T urner . Neurocognitiv e aging data release with behav- ioral, structural and multi-echo functional mri measures. Scientific Data , 9:119, 2022. doi: 10.1038/s41597- 022- 01231- 7. 150 [761] Bart Larsen, V alur Olafsson, Finnegan Calabro, Charles Laymon, Brenden T erv o-Clemmens, Elizabeth Campbell, Davneet Minhas, David Montez, Julie Price, and Beatriz Luna. Matu- ration of the human striatal dopamine system re vealed by pet and quantitativ e mri. Natur e Communications , 11(1):846, 2020. doi: 10.1038/s41467- 020- 14693- 3. 150 , 154 [762] Jalil Rasgado-T oledo, Fernando Lizcano-Cortes, V ictor Enrique Olalde-Mathieu, Giov anna Licea-Haquet, Miguel Angel Zamora-Ursulo, Magda Giordano, and Azalea Reyes-Aguilar . A dataset to study pragmatic language and its underlying cognitiv e processes. F r ontier s in Human Neur oscience , 15, 2021. doi: 10.3389/fnhum.2021.666210. 150 [763] Subha Madhavan, Jean-Claude Zenklusen, Y uri Kotliaro v , Himanso Sahni, Howard A. Fine, and Kenneth Buetow . Rembrandt: helping personalized medicine become a reality through integrati ve translational research. Molecular Cancer Researc h , 7(2):157–167, 2009. doi: 10.1158/1541- 7786.MCR- 08- 0435. 150 [764] Simon Duchesne, Y assine Benhajali, Francois Carbonell, Christian Dansereau, Genevie ve Albouy , Melanie Pelland, Pierre Orban, Jean St-Aubin, Maxime Descoteaux, Emmanuel Stip, and Pierre Bellec. Structural and functional multi-platform mri series of a single human volunteer ov er more than fifteen years. Scientific Data , 6(1):1–18, 2019. doi: 10.1038/ s41597- 019- 0262- 8. 150 [765] Diego Angeles-V aldez, Jalil Rasgado-T oledo, V ictor Issa-Garcia, Thania Balducci, V iviana V illicaña, Alely V alencia, Jorge Julio Gonzalez-Olvera, Ernesto Re yes-Zamorano, and Ed- uardo A. Garza-V illarreal. The mexican magnetic resonance imaging dataset of patients with cocaine use disorder: Sudmex conn. Scientific Data , 9:133, 2022. doi: 10.1038/ s41597- 022- 01251- 3. 150 [766] Rajendra A Morey , Sarah L Da vis, Courtney C Haswell, Jennifer C Naylor , Jason D Kilts, Stev en T Szabo, Larry J Shampine, Gillian J P arke, Delin Sun, Chelsea A Sw anson, Henry R 115 W agner , Mid-Atlantic MIRECC W orkgroup, and Christine E Marx. Widespread cortical thickness is associated with neuroactive steroid le v els. F r ontiers in Neur oscience , 13:1118, 2019. doi: 10.3389/fnins.2019.01118. 150 [767] Ekaterina V . Pechenkov a, Y ana R. Panikrato v a, Maria A. F omina, Alena D. Rumshiskaya, Darya A. Bazhenova, Liudmila A. Mako vskaya, Irina S. Lebedev a, and V alentin E. Sinitsyn. Speech disfluencies: Neurophysiological aspect in normal population. OpenNeuro, 2021. URL https://openneuro.org/datasets/ds003469/versions/1.0.0 . 150 [768] Michal Raf al Zareba, Magdalena Fafro wicz, T adeusz Marek, Ewa Beldzik, Halszka Oginska, Anna Beres, Piotr Faba, Justyna Janik, Koryna Le wando wska, Monika Ostrogorska, Barbara Sikora-W achowicz, Aleksandra Zyrko wska, and Aleksandra Domagalik. Neuroimaging of chronotype, sleep quality and daytime sleepiness: Structural t1-weighted magnetic resonance brain imaging data from 136 young adults. Data in Brief , 41:107956, 2022. doi: 10.1016/j. dib .2022.107956. 150 [769] Oli vier Gev aert, Leslie A. Mitchell, Achal S. Achrol, Jing Xu, Sergio Echegaray , Gary K. Steinberg, Stephen H. Cheshier, Sandy Napel, Greg Zaharchuk, and Sylvia K. Plevritis. Glioblastoma multiforme: Exploratory radiogenomic analysis by using quantitativ e image features. Radiology , 273(1):168–174, 2014. doi: 10.1148/radiol.14131731. 150 [770] W . Lingle, B. J. Erickson, M. L. Zule y , R. Jarosz, E. Bonaccio, J. Filippini, J. M. Net, L. Levi, E. A. Morris, G. G. Figler , P . Elnajjar , S. Kirk, Y . Lee, M. Giger , and N. Gruszauskas. The cancer genome atlas breast in vasi ve carcinoma collection (tcga-brca) (version 3). Data set. The Cancer Imaging Archiv e, 2016. URL https://www.cancerimagingarchive.net/ collection/tcga- brca/ . 150 [771] W eidong Guo, Hui Li, Y uhua Zhu, Lianlian Lan, Shaohua Y ang, Karen Drukker , Emily A Morris, Elizabeth S Burnside, Gretchen J Whitman, Maryellen L Giger , Y onggang Ji, and TCGA Breast Phenotype Research Group. Prediction of clinical phenotypes in in vasi v e breast carcinomas from the integration of radiomics and genomics data. J ournal of Medical Imaging , 2(4):041007, 2015. doi: 10.1117/1.JMI.2.4.041007. 150 [772] Gustav Nilsonne, Sandra T amm, Paolo d’Onofrio, Hanna Å Thuné, Johanna Schw arz, Catha- rina La vebratt, Jia Jia Liu, Kristof fer N T Månsson, T ina Sundelin, John Axelsson, Claus Lamm, Predrag Petrovic, Peter Fransson, Göran Kecklund, Håkan Fischer , Mats Lekander , and T orbjörn Åkerstedt. A multimodal brain imaging dataset on sleep depri vation in young and old humans: The sleepy brain study 1, version 3. Karolinska Institutet ResearchData, 2021. URL https://doi.org/10.5878/87y5- kh22 . 150 [773] Peter V an Schuerbeek, Chris Baeken, and Johan De Mey . The heterogeneity in retrieved relations between the personality trait ‘harm av oidance’ and gray matter v olumes due to v ari- ations in the vbm and roi labeling processing settings. PLOS ONE , 11(4):e0153865, 2016. doi: 10.1371/journal.pone.0153865. 150 [774] Cyril R. Pernet, Phil McAleer , Marianne Latinus, Krzysztof J. Gorgole wski, Ian Charest, Patricia E. G. Bestelmeyer , Rebecca H. W atson, David Fleming, Frances Crabbe, Mitchell V aldes-Sosa, and Pascal Belin. The human voice areas: Spatial org anization and inter- individual variability in temporal and e xtra-temporal cortices. Neur oIma ge , 119:164–174, 2015. doi: 10.1016/j.neuroimage.2015.06.050. 150 [775] Russell A Poldrack, Emily Congdon, W illiam T riplett, Krzysztof J Gorgole wski, Karla H Karlsgodt, Jeanette A Mumford, Fred W Sabb, Nelson B Freimer , Edith D London, T yrone D Cannon, and Robert M Bilder . A phenome-wide examination of neural and cognitiv e function. Scientific Data , 3:160110, 2016. doi: 10.1038/sdata.2016.110. 151 [776] Jonathan D. Power , Mark Plitt, Prantik Kundu, Peter A. Bandettini, and Alex Martin. T em- poral interpolation alters motion in fmri scans: Magnitudes and consequences for artifact detection. PLoS One , 12(9):e0182939, 2017. doi: 10.1371/journal.pone.0182939. 151 [777] Lucca Pizzato T ondo, Thiago W endt V iola, Gabriel R Fries, Bruno Kluwe-Schiav on, Leonardo Mello Rothmann, Renata Cupertino, Pedro Ferreira, Alexandre Rosa Franco, 116 Scott D Lane, Laura Stertz, Zhongming Zhao, Ruifeng Hu, Thomas Meyer , Joy M Schmitz, Consuelo W alss-Bass, and Rodrigo Grassi-Oli veira. White matter deficits in cocaine use disorder: con ver gent evidence from in viv o diffusion tensor imaging and e x vivo proteomic analysis. T ranslational Psychiatry , 11(1):252, 2021. doi: 10.1038/s41398- 021- 01367- x. 151 [778] Emma K. T owlson, Petra E. Vértes, Ulrich Müller , and Sebastian E. Ahnert. Brain networks rev eal the effects of antipsychotic drugs on schizophrenia patients and controls. F r ontiers in Psychiatry , 10:611, 2019. doi: 10.3389/fpsyt.2019.00611. 151 [779] W ei Liao, Y un-Shuang Fan, Siqi Y ang, Jiao Li, Xujun Duan, Qian Cui, and Huafu Chen. Preservation effect: Cigarette smoking acts on the dynamic of influences among unifying neuropsychiatric triple netw orks in schizophrenia. Schizophr enia Bulletin , 45(6):1242–1250, 2019. doi: 10.1093/schbul/sby184. 151 [780] G. Zeng, D. Belavy , S. Li, and G. Zheng. Ev aluation and comparison of automatic interverte- bral disc localization and segmentation methods with 3d multi-modality mr images: A grand challenge. In Pr oceedings of the 5th International W orkshop and Challenge on Computa- tional Methods and Clinical Applications for Spine Imaging (CSI 2018) , volume 11397 of Medical Image Computing and Computer Assisted Intervention – W orkshop and Challeng e , pages 163–171, Cham, Switzerland, 2018. Springer . doi: 10.1007/978- 3- 030- 13736- 6_14. 151 [781] Nicholas Bien, Pranav Rajpurkar , Robyn L. Ball, Jeremy Irvin, Allison Park, Erik Jones, Michael Bereket, Bhavik N. Patel, Kristen W . Y eom, Katie Shpanskaya, Safwan Halabi, Evan Zucker , Gary Fanton, Derek F . Amanatullah, Christopher F . Beaulieu, Geof frey M. Riley , Russell J. Stew art, Francis G. Blankenberg, Da vid B. Larson, Ricky H. Jones, Cur - tis P . Langlotz, Andrew Y . Ng, and Matthe w P . Lungren. Deep-learning-assisted diagno- sis for knee magnetic resonance imaging: Dev elopment and retrospecti v e v alidation of mr - net. PLOS Medicine , 15(11):e1002699, 2018. doi: 10.1371/journal.pmed.1002699. URL https://doi.org/10.1371/journal.pmed.1002699 . 151 [782] Alain Lalande, Zhihao Chen, Thomas Decourselle, Abdul Qayyum, Thibaut Pommier , Luc Lorgis, Ezequiel de la Rosa, Alexandre Cochet, Yves Cottin, Dominique Ginhac, Michel Salomon, Raphaël Couturier , and Fabrice Meriaudeau. Emidec: A database usable for the automatic ev aluation of myocardial infarction from delayed-enhancement cardiac mri. Data , 5(4):89, 2020. doi: 10.3390/data5040089. 151 [783] Li W ang, Dong Nie, Guannan Li, Elodie Puybareau, Jose Dolz, Qian Zhang, Fan W ang, Jing Xia, Zhengwang W u, Jiawei Chen, Kim-Han Thung, T oan Duc Bui, Jitae Shin, Guodong Zeng, Guoyan Zheng, Vladimir S. Fonov , Andrew Doyle, Y ongchao Xu, Pim Moeskops, Josien PW Pluim, Christian Desrosiers, Ismail Ben A yed, Gerard Sanroma, Oualid M. Benkarim, Adria Casamitjana, V eronica V ilaplana, W eili Lin, Gang Li, and Dinggang Shen. Benchmark on automatic 6-month-old infant brain segmentation algorithms: The iseg-2017 challenge. IEEE T r ansactions on Medical Imaging , 38(9):2219–2230, 2019. doi: 10.1109/TMI.2019.2901712. 151 [784] Sangjune L. Lee, Poonam Y adav , Y in Li, Jason J. Meudt, Jessica Strang, Dustin Hebel, Alyx Alfson, Stephanie J. Olson, T era R. Kruser , Jennifer B. Smilowitz, Kailee Borchert, Brianne Loritz, Laila Gharzai, Shervin Karimpour , John Bayouth, and Michael F . Bassetti. Dataset for gastrointestinal tract segmentation on serial mris for abdominal tumor radiotherapy . Data in Brief , 57:111159, 2024. doi: 10.1016/j.dib.2024.111159. 151 [785] Marco Pizzolato, Marco P alombo, Elisenda Bonet-Carne, C. M. T ax, Francesco Grussu, Andrada Ianus, Fabian Bogusz, T omasz Pieciak, Lipeng Ning, Hugo Larochelle, Maxime Descoteaux, Micah Chamberland, Stefano B. Blumberg, Thomy Mertzanidou, Daniel C. Alexander , Maryam Afzali, Santiago Aja-Fernandez, Derek K. Jones, Carl-Fredrik W estin, Y ogesh Rathi, Steven H. Baete, Lucilio Cordero-Grande, Thilo Ladner , Paddy J. Slator , Josef V . Hajnal, Jean-Philippe Thiran, Anthony N. Price, F arshid Sepehrband, F an Zhang, and Jana Hutter . Acquiring and predicting multidimensional diffusion (mudi) data: An open challenge. In Mathematics and V isualization , pages 195–208. Springer , 2020. doi: 10.1007/978- 3- 030- 52893- 5_17. 151 117 [786] Y ue Sun, Kun Gao, Zhengwang W u, Zhihao Lei, Y ing W ei, Jun Ma, Xiaoping Y ang, Xue Feng, Li Zhao, T rung Le Phan, Jitae Shin, T ao Zhong, Y u Zhang, Lequan Y u, Caizi Li, Ramesh Basnet, M. Omair Ahmad, M. N. S. Swamy , W enao Ma, Qi Dou, T oan Duc Bui, Camilo Bermudez Noguera, Bennett Landman, Ian H. Gotlib, Kathryn L. Humphreys, Sarah Shultz, Longchuan Li, Sijie Niu, W eili Lin, V alerie Jewells, Gang Li, Dinggang Shen, and Li W ang. Multi-site infant brain segmentation algorithms: The iseg-2019 challenge. IEEE T r ansactions on Medical Imaging , 40(5):1363–1376, 2021. doi: 10.1109/TMI.2021. 3055428. 151 [787] Aaron Carass, Snehashis Roy , Amrit Jog, Jennifer L. Cuzzocreo, Emily Magrath, Ale xan- dru Gherman, Jon Button, Jennifer Nguyen, Ferran Prados, Carole H. Sudre, M. Jorge Car- doso, Natalia Cawle y , Olga Ciccarelli, Claudia A M Wheeler-Kingshott, Sebastien Ourselin, Luisa Catanese, Hemant Deshpande, Pierre Maurel, Olivier Commowick, Christian Barillot, Xavier T omas-Fernandez, Simon K W arfield, Subodh V aidya, Akhil Chunduru, Ramanathan Muthuganapathy , Ganapathy Krishnamurthi, Andre w Jesson, T al Arbel, Olaf Maier , Heinz Handels, Lovina Ohemeng Iheme, De vrim Unay , Subhasis Jain, Diana M Sima, Dries Smeets, Mohsen Ghafoorian, Bram Platel, Alex Birenbaum, Hayit Greenspan, Pierre-Louis Bazin, Pe- ter A Calabresi, Ciprian M Crainiceanu, Lina M Ellingsen, Daniel S Reich, Jerry L Prince, and Dzung L Pham. Longitudinal multiple sclerosis lesion segmentation: Resource and chal- lenge. Neur oIma ge , 148:77–102, 2017. doi: 10.1016/j.neuroimage.2016.12.064. 151 [788] Huijun Chen et al. Carotid v essel wall segmentation and atherosclerotic lesion detection chal- lenge. In Pr oceedings of the MICCAI 2022 Challenge on Car otid V essel W all Se gmentation and Ather oscler otic Lesion Detection . MICCAI, 2022. 151 [789] Y ue Sun, Limei W ang, Kun Gao, Shihui Y ing, W eili Lin, Kathryn L. Humphreys, Gang Li, Sijie Niu, Mingxia Liu, and Li W ang. Self-supervised learning with application for infant cerebellum se gmentation and analysis. Natur e Communications , 14:4717, 2023. doi: 10. 1038/s41467- 023- 40446- z. 151 [790] Kathryn M. Schmainda and Marjan Prah. Data from brain-tumor -progression. The Cancer Imaging Archiv e, 2018. URL https://doi.org/10.7937/K9/TCIA.2018.15quzvnb . 151 [791] Chun Y uan, Li Chen, Niranjan Balu, Mahmud Mossa-Basha, Jenq-Neng Hwang, David Saloner , and Peter Douglas. Carotid vessel wall segmentation challenge. Zenodo: https://doi.org/10.5281/zenodo.4575301, 2021. International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) endorsed e vent. 151 [792] Zhaohan Xiong, Qing Xia, Zhiqiang Hu, Ning Huang, Cheng Bian, Y efeng Zheng, Sulaiman V esal, Nishant Ravikumar , Andreas Maier, Xin Y ang, Pheng-Ann Heng, Dong Ni, Caizi Li, Qianqian T ong, W eixin Si, Elodie Puybareau, Y ounes Khoudli, Thierry Géraud, Chen Chen, W enjia Bai, Daniel Rueckert, Lingchao Xu, Xiahai Zhuang, Xinzhe Luo, Shuman Jia, Maxime Sermesant, Y ashu Liu, Kuanquan W ang, Davide Borra, Alessandro Masci, Cris- tiana Corsi, Coen de V ente, Mitko V eta, Rashed Karim, Chandrakanth Jayachandran Preetha, Sandy Engelhardt, Menyun Qiao, Y uanyuan W ang, Qian T ao, Marta Nu nez Garcia, Oscar Camara, Nicolo Savioli, Pablo Lamata, and Jichao Zhao. A global benchmark of algorithms for se gmenting the left atrium from late gadolinium-enhanced cardiac magnetic resonance imaging. Medical Imag e Analysis , 67:101832, 2021. doi: 10.1016/j.media.2020.101832. 151 [793] Chiara Maffei, Gabriel Girard, Kurt G Schilling, Dogu Baran A ydogan, Nagesh Adluru, An- drey Zhylka, Y e W u, Matteo Mancini, Andac Hamamci, Alessia Sarica, Dav ood Karimi, Fang-Cheng Y eh, Mert E Y ildiz, Ali Gholipour , Andrea Quattrone, Aldo Quattrone, Pe w- Thian Y ap, Alberto de Luca, Josien Pluim, Alexander Leemans, V iv ek Prabhakaran, Bar - bara B Bendlin, Andrew L Alexander , Bennett A Landman, Erick J Canales-Rodríguez, Muhamed Barakovic, Jonathan Rafael-Patino, Thomas Y u, Gaëtan Rensonnet, Simona Schi- avi, Alessandro Daducci, Marco Pizzolato, Elda Fischi-Gomez, Jean-Philippe Thiran, Geor ge Dai, Giorgia Grisot, Santi Puch, Marc Ramos, Nikola Lazovski, Paulo Rodrigues, V esna Prchkovska, Robert Jones, Julia Lehman, Suzanne Haber , and Anastasia Y endiki. Insights from the irontract challenge: Optimal methods for mapping brain pathways from multi-shell diffusion mri. Neur oImag e , 257:119327, 2022. doi: 10.1016/j.neuroimage.2022.119327. 151 118 [794] Jonathan Rafael-Patino, Gabriel Girard, Raphaël T ruffet, Marco Pizzolato, Emmanuel Caruyer , and Jean-Philippe Thiran. The diffusion-simulated connectivity (disco) dataset. Data Brief , 38:107429, 2021. doi: 10.1016/j.dib.2021.107429. 151 [795] Melanie Ganz et al. Realnoisemri: A realistic mri reconstruction challenge with motion degraded scans. PsyArXiv , 2021. URL https://psyarxiv.com/vzh4g . 151 [796] C. E. Cardenas, A. S. R. Mohamed, G. Sharp, M. Gooding, H. V eeraraghav an, and J. Y ang. Data from aapm rt-mac grand challenge 2019. The Cancer Imaging Archi ve, 2019. URL https://www.cancerimagingarchive.net/collection/aapm- rt- mac/ . 151 [797] Danielle F Pace, Adrian V Dalca, T al Gev a, Andrew J Po well, Mehdi H Moghari, and Polina Golland. Interactive whole-heart se gmentation in congenital heart disease. In Medical Imag e Computing and Computer Assisted Interventions–MICCAI 2015 , v olume 9351 of Lectur e Notes in Computer Science , pages 80–88, 2015. doi: 10.1007/978- 3- 319- 24574- 4_10. 151 [798] Alessandro Daducci, Emmanuel Caruyer , Maxime Descoteaux, and Jean-Philippe Thiran. Hardi reconstruction challenge. In Pr oceedings of the 2013 IEEE 10th International Sympo- sium on Biomedical Imaging , pages 834–837. IEEE, 2013. 151 [799] Bram v an Ginnek en, T obias Heimann, and Martin A. Styner . 3d segmentation in the clinic: A grand challenge. In W orkshop on 3D Segmentation in the Clinic: A Grand Challenge , pages 7–15, Brisbane, Australia, October 2007. Medical Image Computing and Computer Assisted Intervention (MICCAI). 151 [800] Klaus H. Maier-Hein, Peter F . Neher, Jean-Christophe Houde, Emmanuel Caruyer , Alessan- dro Daducci, T im Dyrby , Bram Stieltjes, and Maxime Descoteaux. T ractography challenge ismrm 2015 data. Zenodo, 2015. URL https://zenodo.org/records/572345 . 151 [801] Carole H. Sudre, Kimberlin V an W ijnen, Florian Dubost, Hieab Adams, David Atkinson, Frederik Barkhof, Mahlet A. Birhanu, Esther E. Bron, Robin Camarasa, Nish Chaturv edi, Y uan Chen, Zihao Chen, Shuai Chen, Qi Dou, T avia Evans, Ivan Ezhov , Haojun Gao, Marta Girones Sanguesa, Juan Domingo Gispert, Beatriz Gomez Anson, Alun D. Hughes, M. Arfan Ikram, Silvia Ingala, H. Rolf Jae ger , Florian K ofler , Hugo J. K uijf, Denis Kutnar , Minho Lee, Bo Li, Luigi Lorenzini, Bjoern Menze, Jose Luis Molinuev o, Y iwei Pan, Elodie Puybareau, Rafael Rehwald, Ruisheng Su, Pengcheng Shi, Lorna Smith, Therese Tillin, Guil- laume T ochon, Helene Urien, Bas H. M. v an der V elden, Isabelle F . v an der V elpen, Benedikt W iestler , Frank J. W olters, Pinar Y ilmaz, Marius de Groot, Meike W . V ernooij, and Marleen de Bruijne. Where is v aldo? v ascular lesions detection and segmentation challenge at miccai 2021. Medical Imag e Analysis , 91:103029, 2024. doi: 10.1016/j.media.2023.103029. 151 [802] Hugo J. Kuijf, Edwin Bennink, K oen L. V incken, Nick W eav er , Geert Jan Biessels, and Max A. V ierge v er . Mr brain segmentation challenge 2018 data (mrbrains18). DataverseNL, V ersion 1.0, 2024. URL https://doi.org/10.34894/E0U32Q . 151 [803] C. T obon-Gomez, M. De Craene, K. McLeod, L. T autz, W . Shi, A. Hennemuth, A. Prakosa, H. W ang, G. Carr-White, S. Kapetanakis, A. Lutz, V . Rasche, T . Schaeffter , C. Butakoff, O. Friman, T . Mansi, M. Sermesant, X. Zhuang, S. Ourselin, H.-O. Peitgen, X. Pennec, R. Razavi, D. Rueck ert, A. F . Frangi, and K. S. Rhode. Benchmarking framew ork for myocar- dial tracking and deformation algorithms: an open access database. Medical Imag e Analysis , 17(6):632–648, 2013. doi: 10.1016/j.media.2013.03.008. 151 , 153 [804] W outer Boekel, Eric-Jan W agenmakers, Luam Belay , Josine V erhagen, Scott D Bro wn, and Birte U Forstmann. A purely confirmatory replication study of structural brain-behavior cor - relations. Corte x , 66:115–133, 2015. doi: 10.1016/j.cortex.2014.11.019. 151 [805] Qixiang Lin, Zhengjia Dai, Mingrui Xia, Zaizhu Han, Ruiwang Huang, Gaolang Gong, Chao Liu, Y anchao Bi, and Y ong He. A connecti vity-based test-retest dataset of multi-modal mag- netic resonance imaging in young healthy adults. Scientific Data , 2:150056, 2015. doi: 10.1038/sdata.2015.56. 151 [806] Arno Klein and Jason T ourville. 101 labeled brain images and a consistent human cortical labeling protocol. F r ontiers in Neur oscience , 6:171, 2012. doi: 10.3389/fnins.2012.00171. URL https://dx.doi.org/10.3389/fnins.2012.00171 . 151 119 [807] Ana Luísa Pinho, Alexis Amadon, Baptiste Gauthier , Nicolas Clairis, André Knops, Sarah Genon, Elvis Dohmatob, Juan Jesús T orre, Chantal Ginisty , Séverine Becuwe-Desmidt, Séverine Roger , Y ann Lecomte, V alérie Berland, Laurence Laurier , Véronique Joly-T estault, Gaëlle Médiouni-Cloarec, Christine Doublé, Bernadette Martins, Eric Salmon, Manuela Pi- azza, David Melcher, Mathias Pessiglione, V irginie V an W assenhove, Evelyn Eger , Gaël V aroquaux, Stanislas Dehaene, Lucie Hertz-Pannier , and Bertrand Thirion. Individual brain charting dataset extension, second release of high-resolution fmri data for cogniti ve mapping. Sci Data , 7(1), 2020. URL https://doi.org/10.1038/s41597- 020- 00670- 4 . 151 [808] James V Haxby , J. Swaroop Guntupalli, Andrew C Connolly , Y aroslav O Halchenko, Bryan R Conroy , M. Ida Gobbini, Michael Hanke, and Peter J. Ramadge. A common, high- dimensional model of the representational space in human v entral temporal cortex. Neur on , 72(2):404–416, 2011. doi: 10.1016/j.neuron.2011.08.026. 151 [809] Chantal MW T ax, Francesco Grussu, Enrico Kaden, Lipeng Ning, Umesh Rudrapatna, John Evans, Samuel St-Jean, Alexander Leemans, Santi Puch, Matt Ro we, and Francesco Gal- busera. Cross-scanner and cross-protocol diffusion mri data harmonisation: a benchmark database and ev aluation of algorithms. Neur oImage , 195:285–299, 2019. 151 [810] Alberto De Luca, Andrada Ianus, Alexander Leemans, Marco Palombo, Noam Shemesh, Hui Zhang, Daniel C. Alexander , Markus Nilsson, Martijn Froeling, Geert-Jan Biessels, Mauro Zucchelli, Matteo Frigo, Enes Albay , Sara Sedlar, Abib Alimi, Samuel Deslauriers-Gauthier , Rachid Deriche, Rutger Fick, Maryam Afzali, T omasz Pieciak, Fabian Bogusz, Santiago Aja- Fernández, Evren Özarslan, Derek K. Jones, Haoze Chen, Mingwu Jin, Zhijie Zhang, Fengx- iang W ang, V ishwesh Nath, Prasanna P arvathaneni, Jan Morez, Jan Sijbers, Ben Jeurissen, Shreyas Fadna vis, Stefan Endres, Ariel Rokem, Eleftherios Garyfallidis, Irina Sanchez, V esna Prchkovska, Paulo Rodrigues, Bennet A. Landman, and Kurt G. Schilling. On the general- izability of diffusion mri signal representations across acquisition parameters, sequences and tissue types: Chronicles of the memento challenge. Neur oImag e , 240:118367, 2021. doi: 10.1016/j.neuroimage.2021.118367. 151 [811] Shuo W ang, Chen Qin, Chengyan W ang, Kang W ang, Haoran W ang, Chen Chen, Cheng Ouyang, Xutong Kuang, Chengliang Dai, Y uanhan Mo, Zhang Shi, Chenchen Dai, Xin- rong Chen, He W ang, and W enjia Bai. The extreme cardiac mri analysis challenge un- der respiratory motion (cmrxmotion). In Statistical Atlases and Computational Models of the Heart. Re gular and CMRxMotion Challenge P apers , pages 3–12. Springer, 2022. doi: 10.1007/978- 3- 031- 23443- 9_1. 151 [812] James L. T atum, Joseph D. Kalen, P aula M. Jacobs, Lilia V . Ilev a, Lisa A. Rif fle, Melinda G. Hollingshead, and James H. Dorosho w . A spontaneously metastatic model of bladder cancer: imaging characterization. Journal of T ranslational Medicine , 17(1):425, 2019. doi: 10.1186/ s12967- 019- 02177- y. 151 [813] J. L. T atum, J. D. Kalen, P . M. Jacobs, L. V . Ilev a, L. A. Riffle, S. Keita, N. P atel, C. Sanders, A. James, S. Difilippantonio, L. Thang, M. G. Hollingshead, Y . Evrard, E. Edmondson, D. A. Clunie, Y . Liu, C. Sulow ay , K. E. Smith, U. W agner , J. B. Freymann, J. Kirby , and J. H. Doroshow . Imaging characterization of a metastatic patient derived model of adenocarcinoma pancreas: (pdmr-521955-158-r4). Data set, The Cancer Imaging Archiv e, V ersion 1. URL: https://www .cancerimagingarchiv e.net/collection/pdmr-521955-158-r4/, 2022. 152 [814] SB Amin, KJ Anderson, CE Boudreau, E Martinez-Ledesma, E Kocaka vuk, KC John- son, FP Barthel, FS V arn, C Kassab, X Ling, H Kim, M Barter , CC Lau, C Y ee Ngan, M Chapman, JW Koehler , AD Miller, JP Long, CR Miller , BF Porter , DR Rissi, C Mazcko, AK LeBlanc, PJ Dickinson, RA Packer , AR T aylor , Jr . JH Rossmeisl, KD W oolard, AB He- imberger , JM Le vine, and RGW V erhaak. Canine glioma characterization project for icdc (icdc-glioma) 01. Dataset. The Cancer Imaging Archive, 2020. URL https://www. cancerimagingarchive.net/collection/icdc- glioma/ . V ersion 01. 152 [815] Ryan L Muetzel, Lotte M E Blanken, Sandra Thijssen, Aad v an der Lugt, V incent W V Jaddoe, Frank C V erhulst, Henning Tiemeier , and T onya White. Resting-state networks in 6-to-10 year old children. Human Br ain Mapping , 37(12):4286–4300, 2016. doi: 10.1002/ hbm.23309. 152 120 [816] David E. V aillancourt, Mark B. Spraker , Janey Prodoehl, Ian Abraham, Daniel M. Corcos, and Xiaobo J. Zhou. High-resolution diffusion tensor imaging in the substantia nigra of de nov o parkinson disease. Neur ology , 72(16):1378–1384, 2009. doi: 10.1212/01.wnl.0000340982. 01727.6e. 152 [817] Y angming Ou, Lilla Zöllei, Kallirroi Retzepi, V ictor Castro, Sara V Bates, Ste ve Pieper, Katherine P Andriole, Shawn N Murphy , Randy L Gollub, and Patricia Ellen Grant. Using clinically acquired mri to construct age-specific adc atlases: Quantifying spatiotemporal adc changes from birth to 6-year old. Human Brain Mapping , 38(6):3052–3068, 2017. doi: 10.1002/hbm.23573. 152 [818] Edward F . Jackson. Rider phantom mri. Data set, The Cancer Imaging Archiv e, 2015. URL https://www.cancerimagingarchive.net/collection/rider- phantom- mri/ . 152 [819] Charles R. Meyer , Thomas L. Chene vert, Craig J. Galbán, T imothy D. Johnson, David A. Hamstra, Alna waz Rehemtulla, and Brian D. Ross. Rider breast mri. Data set, The Cancer Imaging Archiv e, 2015. 152 [820] Félix Quinton, Romain Popof f, Benoît Presles, Sarah Leclerc, F abrice Meriaudeau, Guil- laume Nodari, Olivier Lopez, Julie Pellegrinelli, Olivier Chev allier , Dominique Ginhac, Jean-Marc Vrigneaud, and Jean-Louis Alberini. A tumour and liv er automatic segmenta- tion (atlas) dataset on contrast-enhanced magnetic resonance imaging for hepatocellular car- cinoma. Data , 8(5):79, 2023. doi: 10.3390/data8050079. URL https://doi.org/10. 3390/data8050079 . 152 [821] Soumick Chatterjee, Hendrik Mattern, Marc Dörner , Alessandro Sciarra, Florian Dubost, Hannes Schnurre, Rupali Khatun, Chun-Chih Y u, Tsung-Lin Hsieh, Y i-Shan Tsai, Y i-Zeng Fang, Y ung-Ching Y ang, Juinn-Dar Huang, Marshall Xu, Siyu Liu, Fernanda L. Ribeiro, Saskia Bollmann, Karthikesh V arma Chintalapati, Chethan Mysuru Radhakrishna, Sri Chan- dana Hudukula Ram Kumar , Raviteja Sutrave, Abdul Qayyum, Moona Mazher , Imran Raz- zak, Cristobal Rodero, Steven Niederen, Fengming Lin, Y an Xia, Jiacheng W ang, Riyu Qiu, Liansheng W ang, Arya Y azdan Panah, Rosana El Jurdi, Guanghui Fu, Janan Arslan, Ghislain V aillant, Romain V alabregue, Didier Dormont, Bruno Stankof f, Olivier Colliot, Luisa V argas, Isai Daniel Chacon, Ioannis Pitsiorlas, Pablo Arbelaez, Maria A. Zuluaga, Stefanie Schreiber , Oliv er Speck, and Andreas Nürnber ger . Smile-uhura challenge – small vessel segmentation at mesoscopic scale from ultra-high resolution 7t magnetic resonance angiograms. arXiv pr eprint arXiv:2411.09593 , 2024. URL . 152 [822] Chengyan W ang, Jun L yu, Shuo W ang, Chen Qin, K unyuan Guo, Xinyu Zhang, Xiaotong Y u, Y an Li, F anwen W ang, Jianhua Jin, Zhang Shi, Ziqiang Xu, Y apeng Tian, Sha Hua, Zhensen Chen, Meng Liu, Mengting Sun, Xutong Kuang, Kang W ang, Haoran W ang, Hao Li, Y inghua Chu, Guang Y ang, W enjia Bai, Xiahai Zhuang, He W ang, Jing Qin, and Xiaobo Qu. Cmrxrecon: A publicly a v ailable k-space dataset and benchmark to adv ance deep learning for cardiac mri. Scientific Data , 11(1):687, 2024. doi: 10.1038/s41597- 024- 03525- 4. 152 [823] Iris V os, Ynte Ruigrok, Edwin Bennink, Myrthe Buser , Bir gitta V elthuis, and Hugo Kuijf. Data of the Circle of Willis Intracranial Artery Classification and Quantification (CR O WN) Challenge. Datav erseNL, 2023. URL https://doi.org/10.34894/R05G1L . V ersion 2.3. 152 [824] Nancy R. Ne wlin, Kurt Schilling, Serge K oudoro, Bramsh Qamar Chandio, Praitayini Kanakaraj, Daniel Mo yer , Claire E. Kelly , Sila Genc, Jian Chen, Joseph Y uan-Mou Y ang, Y e W u, Y ifei He, Jiawei Zhang, Qingrun Zeng, Fan Zhang, Nagesh Adluru, V ishwesh Nath, Sudhir Pathak, W alter Schneider , Anurag Gade, Y ogesh Rathi, T om Hendriks, Anna V i- lanov a, Maxime Chamberland, T omasz Pieciak, Dominika Ciupek, Antonio T ristán-V ega, Santiago Aja-Fernández, Maciej Malawski, Gani Ouedraogo, Julia Machnio, Christian Ew- ert, Paul M. Thompson, Neda Jahanshad, Eleftherios Garyfallidis, and Bennett A. Land- man. Miccai-cdmri 2023 quantconn challenge findings on achieving rob ust quantitati ve connectivity through harmonized preprocessing of dif fusion mri. MELBA , 2024. doi: 10.59275/j.melba.2024- 9c68. 152 121 [825] Rina Bao, Y a’nan Song, Sara V Bates, Rebecca J W eiss, Anna N Foster , Camilo Jaimes, Susan Sotardi, Y ue Zhang, Randy L Hirschtick, P . Ellen Grant, and Y angming Ou. Boston neonatal brain injury data for hypoxic ischemic encephalopathy (bonbid-hie): I. mri and le- sion labeling. Scientific Data , 12(1):53, 2025. doi: 10.1038/s41597- 024- 03986- 7. 152 [826] Meng Lou, Xiaoqing Liu, Y uqing Zhang, Y izhou Y u, and Hong-Y u Zhou. Liver lesion di- agnosis challenge on multi-phase mri (lld-mmri2023). Zenodo, International Conference on Medical Image Computing and Computer Assisted Interv ention (MICCAI) 2023, 2023. URL https://doi.org/10.5281/zenodo.7852363 . 152 [827] Massachusetts General Hospital, Harv ard Medical School, National Institutes of Health/National Cancer Institute, Sage Bionetworks, Uni versity of W isconsin-Madison, and Intel Corporation. Neurofibromatosis tumor segmentation on whole-body mri, 2023. URL https://doi.org/10.5281/zenodo.7989646 . 152 [828] Logan W illiams, Abdulah Fa waz, Simon Dahan, Emma Robinson, Jonathan O’Muircheartaigh, Andre Marquand, and Seyed Mostafa Kia. Surface learning for clinical neuroimaging: regressing clinical phenotypes for cortical surf ace metrics. Zenodo, 2023. International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) 2023. 152 [829] Myrthe A. D. Buser , Dominique C. Simons, Matthijs Fitski, Marc H. W . A. Wijnen, Anne- mieke S. Littooij, A. H. ter Brugge, I. N. V os, M. H. A. Janse, M. de Boer , R. ter Maat, J. Sato, S. Kido, S. Kondo, S. Kasai, M. W odzinski, H. Muller, J. Y e, J. He, Y . Kirch- hoff, M. R. Rokkus, G. Haokai, S. Zitong, M. Fernández-Patón, D. V eiga-Canuto, D. G. Ellis, M. R. Aizenberg, B. H. M. van der V elden, H. Kuijf, A. De Luca, and A. F . W . van der Steeg. Automated segmentation of pediatric neuroblastoma on multi-modal mri: Results of the sppin challenge at miccai 2023. arXiv preprint , 2025. doi: 10.48550/arXiv .2505.00369. URL . 152 [830] Andrey Malinin, Andreas Athanasopoulos, Muhamed Barakovic, Meritxell Bach Cuadra, Mark J. F . Gales, Cristina Granziera, Mara Graziani, Nik olay Kartashe v , K onstantinos Kyr - iakopoulos, Po-Jui Lu, Nataliia Molchanov a, Antonis Nikitakis, V atsal Raina, Francesco La Rosa, Eli Si vena, V asileios Tsarsitalidis, Efi Tsompopoulou, and Elena V olf. Shifts 2.0: Extending the dataset of real distributional shifts. arXiv pr eprint arXiv:2206.15407 , 2022. doi: 10.48550/ARXIV .2206.15407. 152 [831] Sunny Jansen and T erry V an Dyke. Tcia mouse-astrocytoma collection (mouse-astrocytoma) [data set]. The Cancer Imaging Archiv e, 2015. URL https://doi.org/10.7937/K9TCIA. 2017.SGW7CAQW . 152 [832] R. Jain, L. M. Poisson, D. Gutman, L. Scarpace, S. N. Hwang, C. A. Holder , M. W intermark, A. Rao, R. R. Colen, J. Kirby , J. Freymann, C. C. Jaf fe, T . Mikkelsen, and A. Flanders. Outcome prediction in patients with glioblastoma by using imaging, clinical, and genomic biomarkers: Focus on the nonenhancing component of the tumor . Radiology , 272(2):484– 493, 2014. doi: 10.1148/radiol.14131691. 152 [833] S. Bakas, C. Sako, H. Akbari, M. Bilello, A. Sotiras, G. Shukla, J. D. Rudie, N. F . Santamaria, A. F . Kazerooni, S. Pati, S. Rathore, E. Mamourian, S. M. Ha, W . Parker , J. Doshi, U. Baid, M. Bergman, Z. A. Binder, R. V erma, R. Lustig, A. S. Desai, S. J. Bagley , Z. Mourelatos, J. Morrissette, C. D. W att, S. Brem, R. L. W olf, E. R. Melhem, M. P . Nasrallah, S. Mohan, D. M. O’Rourke, and C. Da v atzikos. The univ ersity of pennsylv ania glioblastoma (upenn- gbm) cohort: advanced mri, clinical, genomics, & radiomics. Scientific Data , 9(1):453, 2022. doi: 10.1038/s41597- 022- 01560- 7. 152 [834] R. Chitalia, S. Pati, M. Bhalerao, S. P . Thakur, N. Jahani, V . Belenky , E. S. McDonald, J. Gibbs, D. C. Newitt, N. M. Hylton, D. K ontos, and S. Bakas. Expert tumor annotations and radiomics for locally advanced breast cancer in dce-mri for acrin 6657/i-spy1. Scientific Data , 9(1), 2022. doi: 10.1038/s41597- 022- 01555- 4. 152 [835] W en Li, Da vid C. Newitt, Jessica Gibbs, Lisa J. W ilmes, Ella F . Jones, V ignesh A. Arasu, Fredrik Strand, Natsuko Onishi, Alex Anh-Tu Nguyen, John K ornak, Bonnie N. Joe, Ellen R. 122 Price, Heloisa Ojeda-Fournier , Mana Eghtedari, Kimberly W . Zamora, Shauna A. W oodard, Heather Umphre y , W endy Bernreuter , Melissa Nelson, and Nola M. Hylton. I-spy 2 breast dynamic contrast enhanced mri trial (ispy2) (version 1) [data set], 2022. URL https:// www.cancerimagingarchive.net/collection/ispy2/ . 152 [836] Sarthak Pati, Ruchika V erma, Hamed Akbari, Michel Bilello, V ir ginia B Hill, Chiharu Sako, Ramon Correa, Niha Beig, Ludovic V enet, Siddhesh Thakur , Prashant Serai, Sung Min Ha, Geri D Blake, Russell T Shinohara, Pallavi T iw ari, and Spyridon Bakas. Reproducibility analysis of multi-institutional paired e xpert annotations and radiomic features of the i vy glioblastoma atlas project (ivy GAP) dataset. Medical Physics , 47(12):6039–6052, 2020. doi: 10.1002/mp.14556. 152 [837] Y ibin W ang, W illiam Neil Duggar , Da vid Michael Caballero, T oms V engaloor Thomas, Neha Adari, Eswara Kumar Mundra, and Haifeng W ang. A brain mri dataset and baseline e v al- uations for tumor recurrence prediction after gamma knife radiotherapy . Scientific Data , 10(1):785, 2023. doi: 10.1038/s41597- 023- 02683- 1. URL https://doi.org/10.1038/ s41597- 023- 02683- 1 . 152 [838] K. Owczarczyk, D. Prezzi, D. Boisfwr , R. Adams, and V . Goh. Expert anal cancer consensus staging (exact). The Cancer Imaging Archiv e, 2023. URL https://www. cancerimagingarchive.net/collection/exact/ . 152 [839] M. Rozenfeld and P . Jordan. Annotations for chemotherapy and radiation therapy in treating young patients with newly diagnosed, previously untreated, high-risk medulloblastoma/pnet (acns0332-tumor-annotations). Data set, The Cancer Imaging Archi v e, 2022. URL https: //doi.org/10.7937/D8A8- 6252 . 152 [840] Ev an Calabrese, Ja vier E. V illanue v a-Meyer , Jeffre y D. Rudie, Andreas M. Rauschecker , Ujjwal Baid, Spyridon Bakas, Soonmee Cha, John T . Mongan, and Christopher P . Hess. The univ ersity of california san francisco preoperative dif fuse glioma mri (ucsf-pdgm) (version 5). The Cancer Imaging Archiv e, a v ailable at https://doi.org/10.7937/tcia.bdgf- 8v37 , 2022. 152 [841] Santiago Cepeda, Sergio García-García, Ignacio Arrese, Francisco Herrero, Trinidad Escud- ero, T omás Zamora, and Rosario Sarabia. The río hortega university hospital glioblastoma dataset: a comprehensi ve collection of preoperati v e, early postoperati v e and recurrence mri scans (rhuh-gbm). Data in Brief , 50:109617, 2023. doi: 10.1016/j.dib .2023.109617. 152 [842] Daniel Barboriak. Data from rider_neuro_mri. The Cancer Imaging Archiv e, 2015. URL http://doi.org/10.7937/K9/TCIA.2015.VOSN3HN1 . 152 [843] Andre w V assantachart, Y anhui Cao, Zhenwei Shen, Ke vin Cheng, Mark Gribble, Jeremy C. Y e, Gelareh Zada, K evin Hurth, Arun Mathew , Sandra Guzman, and W ei Y ang. Segmentation and classification of grade i and ii meningiomas from magnetic resonance imaging: An open annotated dataset (meningioma-se g-class). The Cancer Imaging Archi ve (TCIA), 2023. URL https://doi.org/10.7937/0TKV- 1A36 . V ersion 1; Data set. 152 [844] Amir Reza Sadri, Andre w Janowczyk, Ruchika V erma, Jacob Antunes, Anant Madabhushi, Palla vi Tiw ari, and Satish V iswanath. Mrqy quality measures for tcia mri datasets. The Cancer Imaging Archiv e, 2020. 152 [845] A. Beers, E. Gerstner, B. Rosen, D. Clunie, S. Pieper, A. Fedorov , and J. Kalpathy-Cramer . Dicom-seg con v ersions for tcga-lgg and tcga-gbm segmentation datasets. [Data set]. The Cancer Imaging Archiv e, 2018. 152 [846] Kimberley M. Timmins, Irene C. van der Schaaf, Edwin Bennink, Ynte M. Ruigrok, Xin- gle An, Michael Baumgartner , Pascal Bourdon, Riccardo De Feo, T ommaso Di Noto, Florian Dubost, Augusto Fa v a-Sanches, Xue Feng, Corentin Giroud, Inteneural Group, Minghui Hu, Paul F . Jaeger , Juhana Kaiponen, Michał Klimont, Y uexiang Li, Hongwei Li, Y i Lin, Timo Loehr, Jun Ma, Klaus H. Maier-Hein, Guillaume Marie, Bjoern Menze, Jonas Richiardi, Saifeddine Rjiba, Dhav al Shah, Suprosanna Shit, Jussi T ohka, Thierry Urruty , Urszula W ali ´ nska, Xiaoping Y ang, Y unqiao Y ang, Y in Y in, Birgitta K. V elthuis, 123 and Hugo J. Kuijf. Comparing methods of detecting and se gmenting unruptured intracra- nial aneurysms on tof-mras: The adam challenge. Neur oImag e , 238:118216, 2021. doi: 10.1016/j.neuroimage.2021.118216. 152 [847] T omaž Vrtovec, Jianhua Y ao, Ben Glocker , T obias Klinder , Alejandro Frangi, Guoyan Zheng, and Shuo Li. Computational Methods and Clinical Applications for Spine Imaging: Thir d International W orkshop and Challenge, CSI 2015, Held in Conjunction with MICCAI 2015, Munich, Germany , October 5, 2015, Pr oceedings , volume 9402. Springer , 2016. 152 [848] David W Shattuck, Mubeena Mirza, V itria Adisetiyo, Cornelius Hojatkashani, Georges Sala- mon, Katherine L Narr , Russell A Poldrack, Robert M Bilder , and Arthur W T oga. Construc- tion of a 3d probabilistic atlas of human cortical structures. Neur oIma ge , 39(3):1064–1080, 2007. doi: 10.1016/j.neuroimage.2007.09.031. 152 [849] Continuous Registration Challenge Organizers. Continuous registration challenge. https://continuousregistration.grand-challenge.or g/home/, 2018. Grand Challenge. 152 [850] Columbia Uni versity Medical Center . Cumc12 dataset, 2018. URL https:// continuousregistration.grand- challenge.org/data/ . Accessed: 2025-08-16. 152 [851] Ilya Nelk enbaum, Galia Tsarf aty , Nahum Kiryati, Eli K onen, and Arnaldo Mayer . Automatic segmentation of white matter tracts using multiple brain mri sequences. In 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI) , pages 368–371. IEEE, 2020. 152 [852] Gongning Luo, Mingwang Xu, Hongyu Chen, Xinjie Liang, Xing T ao, Dong Ni, Hyunsu Jeong, Chulhong Kim, Raphael Stock, Michael Baumgartner , Y annick Kirchhoff, Maximilian Rokuss, Klaus Maier-Hein, Zhikai Y ang, T ianyu F an, Nicolas Boutry , Dmitry T ereshchenko, Arthur Moine, Maximilien Charmetant, Jan Sauer , Hao Du, Xiang-Hui Bai, V ipul Pai Raikar, Ricardo Montoya-del Angel, Robert Marti, Miguel Luna, Dongmin Lee, Abdul Qayyum, Moona Mazher , Qihui Guo, Changyan W ang, Navchetan A wasthi, Qiaochu Zhao, W ei W ang, Kuanquan W ang, Qiucheng W ang, and Suyu Dong. T umor detection, segmenta- tion and classification challenge on automated 3d breast ultrasound: The tdsc-abus chal- lenge. arXiv pr eprint arXiv:2501.15588 , 2025. doi: 10.48550/arXiv .2501.15588. URL https://arxiv.org/abs/2501.15588 . 152 [853] Oli vier Bernard, Brecht Heyde, Martino Alessandrini, and Daniel Barbosa. Challenge on en- docardial three-dimensional ultrasound se gmentation (cetus). The MIDAS J ournal - MICCAI 2014 W orkshop: Challenge on Endocardial Three-dimensional Ultrasound Se gmentation , 2014. doi: 10.54294/j78w0v. CC BY 4.0. 152 [854] Patrick Carnahan. T owards P atient Specific Mitral V alve Modelling via Dynamic 3D T rans- esophageal Ec hocar diogr aphy . PhD thesis, The Univ ersity of W estern Ontario (Canada), 2023. 152 [855] Reinhard Beichel, Eric J. Ulrich, Christian Bauer , Daniel W . Byrd, James P . Muzi, Mark Muzi, Paul E. Kinahan, John J. Sunderland, Michael M. Graham, and John M. Buatti. Qin pet phantom. Data set. The Cancer Imaging Archiv e, 2015. URL https://doi.org/10. 7937/k9/tcia.2015.zpukhckb . 153 [856] Donglai W ei, Zudi Lin, Daniel Franco-Barranco, Nils W endt, Xingyu Liu, W enjie Y in, Xin Huang, Aarush Gupta, W on-Dong Jang, Xueying W ang, Ignacio Ar ganda-Carreras, Jef f W . Lichtman, and Hanspeter Pfister . Mitoem dataset: Large-scale 3d mitochondria instance segmentation from em images. In Medical Image Computing and Computer Assisted Inter - vention – MICCAI 2020 , v olume 12265 of Lectur e Notes in Computer Science , pages 66–76. Springer , 2020. doi: 10.1007/978- 3- 030- 59722- 1_7. 154 [857] Matthe w D. Guay , Zeyad A. S. Emam, Adam B. Anderson, Maria A. Aronova, Irina D. Pokrovskaya, Brian Storrie, and Richard D. Leapman. Dense cellular se gmentation for em using 2d–3d neural netw ork ensembles. Scientific Reports , 11:2561, 2021. doi: 10.1038/ s41598- 021- 81590- 0. 154 124 [858] W eisi Xie, Nicholas P . Reder, C. F . K oyuncu, Paul Leo, Scott Hawley , H. Huang, C. Mao, Nadia Postupna, Soyoung Kang, R. Serafin, G. Gao, Q. Han, K. Bishop, L. Barner, P . Fu, J. Wright, C. Keene, J. V aughan, A. Janowczyk, A. Madabhushi, and Jonathan T . C. Liu. 3d pathology of prostate biopsies with biochemical recurrence outcomes: raw h&e-analog datasets and image translation-assisted segmentation in 3d (itas3d) datasets (pca_bx_3dpathology), 2023. 155 [859] Matthias Iv antsits, Leonid Gouber grits, Jan-Martin Kuhnigk, Markus Huellebrand, Jan Brün- ing, T abea K ossen, Boris Pfahringer , Jens Schaller , Andreas Spuler, Titus Kuehne, and Anja Hennemuth. Cerebral aneurysm detection and analysis challenge 2020 (cada). In F irst Chal- lenge, CAD A 2020, Held in Conjunction with the 23rd International Confer ence on Medical Image Computing and Computer -Assisted Intervention (MICCAI 2020) , volume 12643 of Lectur e Notes in Computer Science , pages 3–17, Cham, Switzerland, 2021. Springer . doi: 10.1007/978- 3- 030- 72862- 5_1. 155 [860] Nicolas Dazeo and Ignacio Larrabide. Shiny-icarus: Segmentation over three dimensional rotational angiography of internal carotid artery with aneurysm. ISBI 2023 Challenge Dataset, 2023. URL https://www.synapse.org/#!Synapse:syn45774070 . 155 [861] Matthias Ivantsits, Leonid Goubergrits, Jan-Martin Kuhnigk, Matthias Huellebrand, Juliane Bruening, Thomas Kossen, Bojan Pfahringer , Jürgen Schaller, Andreas Spuler , Tilo Kühne, Y anfei Jia, Xiaoxin Li, Saenc Shit, Bjoern Menze, Zheng Su, Ji Ma, Zhen Nie, Kunal Jain, Y an Liu, Y ichao Lin, and Anja Hennemuth. Detection and analysis of cerebral aneurysms based on x-ray rotational angiography - the cada 2020 challenge. Medical Image Analysis , 77:102333, 2022. doi: 10.1016/j.media.2021.102333. 155 [862] T ri Nguyen, Mukul Narwani, Mark Larson, Y icong Li, Shuhan Xie, Hanspeter Pfister, Donglai W ei, Nir Shavit, Lu Mi, Alexandra Pacureanu, W ei-Chung Lee, and Aaron T Kuan. The xpress challenge: Xray projectomic reconstruction–extracting segmentation with sk ele- tons. arXiv pr eprint arXiv:2302.03819 , 2023. 155 [863] J. Hong, M. Reyngold, C. Crane, J. Cuaron, C. Hajj, J. Mann, M. Zinov oy , E. Y orke, E. Lo- Castro, A. P . Apte, and G. Mageras. Breath-hold ct and cone-beam ct images with expert manual organ-at-risk segmentations from radiation treatments of locally advanced pancre- atic cancer . The Cancer Imaging Archiv e, 2021. URL https://doi.org/10.7937/TCIA. ESHQ- 4D90 . Data set. 155 [864] Afua A. Y orke, Gary C. McDonald, Da vid Solis, and Thomas Guerrero. Pelvic reference data (version 1). Data set, The Cancer Imaging Archi ve, 2019. URL https://doi.org/10. 7937/TCIA.2019.WOSKQ5OO . 155 [865] Federico Bolelli, Luca Lumetti, Shankeeth V inayahalingam, Mattia Di Bartolomeo, Arrigo Pellacani, K evin Marchesini, Niels v an Nistelrooij, Pieter v an Lierop, T ong Xi, Y usheng Liu, Rui Xin, T ao Y ang, Lisheng W ang, Haoshen W ang, Chenfan Xu, Zhiming Cui, Marek Michal W odzinski, Henning Müller, Y annick Kirchhof f, Maximilian Rokuss, Klaus H. Maier-Hein, Jaehwan Han, W an Kim, Hong-Gi Ahn, T omasz Szczepa ´ nski, Michal Grzeszczyk, Przemys- law K orzenio wski, V icent Caselles Ballester, Xavier Bur gos-Artizzu, Ferran Prados Carrasco, Stefaan Berge, Bram van Ginneken, Ale x Anesi, and Costantino Grana. Segmenting the infe- rior alv eolar canal in cbcts volumes: the toothfairy challenge. IEEE T ransactions on Medical Imaging , 44(4):1890–1906, 2024. doi: 10.1109/TMI.2024.3523096. 155 [866] Hrvoje Bogunovic, Freerk V enhuizen, Sophie Klimscha, Stefanos Apostolopoulos, Alireza Bab-Hadiashar , Ulas Bagci, Mirza Faisal Beg, Loza Bekalo, Qiang Chen, Carlos Ciller , Karthik Gopinath, Amirali K. Gostar , Kiwan Jeon, Zexuan Ji, Sung Ho Kang, Dara D. K oozekanani, Donghuan Lu, Dustin Morley , Keshab K. Parhi, Hyoung Suk Park, Abdolreza Rashno, Marinko Sarunic, Saad Shaikh, Jayanthi Siv aswamy , Ruwan T ennakoon, Shivin Y a- dav , Sandro De Zanet, Sebastian M. W aldstein, Bianca S. Gerendas, Caroline Klav er , Clara I. Sanchez, and Ursula Schmidt-Erfurth. Retouch – the retinal oct fluid detection and segmen- tation benchmark and challenge. IEEE T r ansactions on Medical Imaging , 38(8):1858–1874, 2019. doi: 10.1109/TMI.2019.2901398. 155 125 [867] R OCC Organizers. Retinal oct classification challenge. In Pr oceedings of the 2017 Medical Image Understanding and Analysis Conference , 2017. URL https://rocc. grand- challenge.org/ . 155 [868] Stefan Maetschke, Bhavna Antony , Hiroshi Ishikawa, Gadi W ollstein, Joel S. Schuman, and Rahil Garnavi. A feature agnostic approach for glaucoma detection in oct v olumes. PLoS ONE , 14(7):e0219126, 2019. doi: 10.1371/journal.pone.0219126. 155 [869] Mohit Prabhushankar , Kiran K okilepersaud, Y ash-yee Logan, Stephanie T rejo Corona, Ghas- san AlRegib, and Charles W ykoff. Oliv es dataset: Ophthalmic labels for in vestig ating visual eye semantics. In Advances in Neural Information Pr ocessing Systems 35 (NeurIPS 2022) T rack on Datasets and Benchmarks , 2022. doi: 10.48550/arXi v .2209.11195. 155 [870] Junde W u, Huihui Fang, Fei Li, Huazhu Fu, Fengbin Lin, Jiongcheng Li, Lexing Huang, Qinji Y u, Sifan Song, Xinxing Xu, Y anyu Xu, W ensai W ang, Lingxiao W ang, Shuai Lu, Huiqi Li, Shihua Huang, Zhichao Lu, Chubin Ou, Xifei W ei, Bingyuan Liu, Riadh K obbi, Xiaoying T ang, Li Lin, Qiang Zhou, Qiang Hu, Hrv oje Bogunovic, José Ignacio Orlando, Xiulan Zhang, and Y anwu Xu. Gamma challenge: Glaucoma grading from multi-modality images. arXiv pr eprint arXiv:2202.06511 , 2022. doi: 10.48550/arXiv .2202.06511. URL https://arxiv.org/abs/2202.06511 . 155 [871] Sina Farsiu, Stephanie J. Chiu, Rachelle V . O’Connell, Francisco A. Folg ar , Eric Y uan, Joseph A. Izatt, Cynthia A. T oth, and Age-Related Eye Disease Study 2 Ancillary Spec- tral Domain Optical Coherence T omography Study Group. Quantitativ e classification of eyes with and without intermediate age-related macular degeneration using optical coherence to- mography . Ophthalmology , 121(1):162–172, 2014. doi: 10.1016/j.ophtha.2013.07.013. 155 [872] Stephanie J. Chiu, Joseph A. Izatt, Rachelle V . O’Connell, Katrina P . W inter , Cynthia A. T oth, and Sina Farsiu. V alidated automatic segmentation of amd pathology including drusen and geographic atrophy in sd-oct images. Investigative Ophthalmology & V isual Science , 53(1): 53–61, 2012. doi: 10.1167/iovs.11- 7640. 155 [873] Stephanie J. Chiu, Michael J. Allingham, Priyatham S. Mettu, Scott W . Cousins, Joseph A. Izatt, and Sina Farsiu. K ernel regression based segmentation of optical coherence tomography images with diabetic macular edema. Biomedical Optics Expr ess , 6(4):1172–1194, 2015. doi: 10.1364/BOE.6.001172. 155 [874] Pratul P Srini vasan, Leo A Kim, Priyatham S Mettu, Scott W Cousins, Grant M Comer, Joseph A Izatt, and Sina Farsiu. Fully automated detection of diabetic macular edema and dry age-related macular degeneration from optical coherence tomography images. Biomedical Optics Expr ess , 5(10):3568–3577, 2014. doi: 10.1364/BOE.5.003568. 155 [875] Somayyeh Soltanian-Zadeh, Kazuhiro K urokawa, Zhuolin Liu, Furu Zhang, Osamah Saeedi, Daniel X. Hammer, Donald T . Miller , and Sina Farsiu. W eakly supervised individual ganglion cell segmentation from adapti ve optics oct images for glaucomatous damage as- sessment. Optica , 8(5):642–651, 2021. doi: 10.1364/OPTICA.418274. URL https: //doi.org/10.1364/OPTICA.418274 . 155 [876] Huazhu Fu, Y anwu Xu, Xiulan Zhang, Fei Li, José Ignacio Orlando, and Hrvoje Bogunovic. Structural-functional transition in glaucoma assessment. https://zenodo.org/record/7835341, 2023. 155 [877] T ianchi Platform. Eye OCT Datasets. Tianchi Dataset Platform, 2021. URL https:// tianchi.aliyun.com/dataset/dataDetail?dataId=90672 . Retinal OCT images for disease classification and segmentation. 155 [878] Mingchao Li et al. Octa-500: a retinal dataset for optical coherence tomography angiography study . Medical Image Analysis , 93:103092, 2024. URL https://ieee- dataport.org/ open- access/octa- 500 . Large-scale OCT A dataset with 500 subjects for retinal disease analysis. 155 126 [879] Chinedu Innocent Nwoye, T ong Y u, Saurav Sharma, Aditya Murali, Deepak Alapatt, Armine V ardazaryan, Kun Y uan, Jonas Hajek, W olfgang Reiter , Amine Y amlahi, et al. Cholec- triplet2022: Show me a tool and tell me the triplet—an endoscopic vision challenge for surgical action triplet detection. Medical Image Analysis , 89:102888, 2023. 155 , 156 [880] Chinedu Innocent Nwoye, Deepak Alapatt, T ong Y u, Armine V ardazaryan, Fangf ang Xia, Zixuan Zhao, T ong Xia, Fucang Jia, Y uxuan Y ang, Hao W ang, et al. Cholectriplet2021: A benchmark challenge for surgical action triplet recognition. Medical Imag e Analysis , 86: 102803, 2023. 155 [881] Aneeq Zia, Kiran Bhattacharyya, Xi Liu, Ziheng W ang, Satoshi Kondo, Emanuele Colleoni, Beatrice V an Amsterdam, Razeen Hussain, Raabid Hussain, Lena Maier -Hein, et al. Surgical visual domain adaptation: Results from the miccai 2020 surgvisdom challenge. arXiv preprint arXiv:2102.13644 , 2021. 155 [882] Surgical workflo w analysis in the sensoror, 2018. https://endovissub2017- workflow. grand- challenge.org/ . 155 [883] Martin W agner , Beat-Peter Müller-Stich, Anna Kisilenko, Duc Tran, Patrick Heger , Lars Mündermann, David M Lubotsky , Benjamin Müller , T ornike Davitashvili, Manuela Capek, Annika Reinke, Carissa Reid, T ong Y u, Armine V ardazaryan, Chinedu Innocent Nwoye, Nicolas Padoy , Xinyang Liu, Eung-Joo Lee, Constantin Disch, Hans Meine, T ong Xia, Fucang Jia, Satoshi Kondo, W olfgang Reiter, Y ueming Jin, Y onghao Long, Meirui Jiang, Qi Dou, Pheng Ann Heng, Isabell T wick, Kadir Kirtac, Enes Hosgor , Jon Lindström Bolmgren, Michael Stenzel, Björn von Siemens, Long Zhao, Zhenxiao Ge, Haiming Sun, Di Xie, Mengqi Guo, Daochang Liu, Hannes G. K enngott, Felix Nickel, Moritz von Frankenber g, Franziska Mathis-Ullrich, Annette K opp-Schneider , Lena Maier-Hein, Ste- fanie Speidel, and Sebastian Bodenstedt. Comparativ e validation of machine learning al- gorithms for surgical workflow and skill analysis with the heichole benchmark. Medi- cal Image Analysis , 86:102770, 2023. ISSN 1361-8415. doi: https://doi.org/10.1016/j. media.2023.102770. URL https://www.sciencedirect.com/science/article/pii/ S1361841523000312 . 155 [884] Arnaud Huaulmé, Duygu Sarikaya, Kévin Le Mut, Fabien Despinoy , Y onghao Long, Qi Dou, Chin-Boon Chng, W enjun Lin, Satoshi K ondo, Laura Brav o-Sánchez, et al. Micro-surgical anastomose workflow recognition challenge report. Computer Methods and Pr ogr ams in Biomedicine , 212:106452, 2021. 155 [885] Arnaud Huaulmé, Kanako Harada, Quang-Minh Nguyen, Bogyu Park, Seungb um Hong, Min-K ook Choi, Michael Pev en, Y unshuang Li, Y onghao Long, Qi Dou, et al. Peg transfer workflo w recognition challenge report: Does multi-modal data improve recognition? arXiv pr eprint arXiv:2202.05821 , 2022. 155 [886] Surgical tool localization in endoscopic videos. Endoscopic V ision Challenge (MICCAI 2022) website, 2022. https://surgtoolloc.grand- challenge.org/ . 155 [887] Y upeng Zhuo, Andrew W . Kirkpatrick, Kyle Couperus, Oanh Tran, and Juan W achs. The trauma thompson challenge report miccai 2023. In T rauma Thompson Challenge , pages 61– 71. Springer , 2023. 155 [888] Open suturing skills challenge. Endoscopic V ision Challenge (MICCAI 2025) website, 2025. https://opencas.dkfz.de/endovis/challenges/2025/ . 155 [889] Fedsurg: Federated learning for surgical vision. Endoscopic V ision Challenge (MIC- CAI 2024) website, 2022. https://www.synapse.org/Synapse:syn53137385/wiki/ 625370 . 155 [890] Jie wen Y ang, Xinpeng Ding, Ziyang Zheng, Xiaowei Xu, and Xiaomeng Li. Graphecho: Graph-driv en unsupervised domain adaptation for echocardiogram video segmentation. In Pr oceedings of the IEEE/CVF International Confer ence on Computer V ision (ICCV) , pages 11878–11887, October 2023. 155 127 [891] Andru P T winanda, Sherif Shehata, Didier Mutter, Jacques Marescaux, Michel De Mathelin, and Nicolas Padoy . Endonet: a deep architecture for recognition tasks on laparoscopic videos. IEEE transactions on medical imaging , 36(1):86–97, 2016. 155 [892] The critical view of safety challenge. Endoscopic V ision Challenge (MICCAI 2024) website, 2024. https://www.cvschallenge.org/ . 156 [893] Negin Ghamsarian, Mario T aschwer , Doris Putzgruber-Adamitsch, Stephanie Sarny , and Klaus Schoeffmann. Relev ance detection in cataract surgery videos by spatio- tempo- ral action localization. In 25th International Conference on P attern Recognition, ICPR 2020, V irtual Event / Milan, Italy , J anuary 10-15, 2021 , pages 10720–10727. IEEE, 2020. doi: 10.1109/ICPR48806.2021.9412525. URL https://doi.org/10.1109/ICPR48806. 2021.9412525 . 156 [894] Klaus Schoeffmann, Heinrich Husslein, Sabrina Kletz, Stefan Petscharnig, Bernd Münzer , and Christian Beecks. V ideo retriev al in laparoscopic video recordings with dynamic content descriptors. Multim. T ools Appl. , 77(13):16813–16832, 2018. doi: 10.1007/ s11042- 017- 5252- 2. URL https://doi.org/10.1007/s11042- 017- 5252- 2 . 156 [895] Ming Hu, Peng Xia, Lin W ang, Siyuan Y an, Feilong T ang, Zhongxing Xu, Y imin Luo, Kaimin Song, Jurgen Leitner , Xuelian Cheng, et al. Ophnet: A large-scale video benchmark for ophthalmic sur gical workflo w understanding. arXiv pr eprint arXiv:2406.07471 , 2024. 156 [896] Ming Hu, Lin W ang, Siyuan Y an, Don Ma, Qingli Ren, Peng Xia, W ei Feng, Peibo Duan, Lie Ju, and Zongyuan Ge. Nurvid: A lar ge expert-le vel video database for nursing procedure ac- tivity understanding. Advances in Neur al Information Pr ocessing Systems , 36:18146–18164, 2023. 156 [897] Ralf Stauder , Daniel Ostler, Michael Kranzfelder , Sebastian Koller , Hubertus Feußner , and Nassir Navab . The tum lapchole dataset for the m2cai 2016 workflow challenge. arXiv pr eprint arXiv:1610.09278 , 2016. 156 [898] Sharib Ali, Mariia Dmitriev a, Noha Ghatwary , Sophia Bano, Gorkem Polat, Alptekin T em- izel, Adrian Krenzer , Amar Hekalo, Y un Bo Guo, Bogdan Matusze wski, et al. Deep learning for detection and segmentation of artefact and disease instances in gastrointestinal endoscopy . Medical image analysis , 70:102002, 2021. 156 [899] Lalith Sharan, Gabriele Romano, Sven K oehler , Halvar Kelm, Matthias Karck, Raf faele De Simone, and Sandy Engelhardt. Mutually improved endoscopic image synthesis and landmark detection in unpaired image-to-image translation. IEEE J ournal of Biomedical and Health Informatics , 26(1):127–138, 2021. 156 [900] Institute of Biomedical Engineering, Univ ersity of Oxford. A-afma ultrasound challenge dataset (1.0), 2020. URL https://doi.org/10.5281/zenodo.4305956 . 156 [901] Gastrointestinal image analysis, 2021. https://giana.grand- challenge.org/ . 156 [902] Stefanie Speidel, Lena Maier-Hein, Danail Stoyanov , Sebastian Bodenstedt, Martin W agner , Beat Müller , Jonathan Chen, Benjamin Müller, Franziska Mathis-Ullrich, P aul Scheikl, Jorge Bernal, A ymeric Histache, Gloria Fernandes-Esparrach, Xavier Dray , Sophia Bano, Alessan- dro Casella, Francisco V asconcelos, Sara Moccia, Chinedu Nwoye, Deepak Alapatt, Armine V ardazaryan, Nicolas Padoy , Arnaud Huaulme, Kanako Harada, Pierre Jannin, Aneeq Zia, Kiran Bhattacharyya, Xi Liu, Ziheng W ang, and Anthony Jarc. Endoscopic vision challenge 2021. 24th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2021), 2021. URL https://doi.org/10.5281/zenodo.4572973 . 156 [903] Anita Rau, Sophia Bano, Y ueming Jin, P ablo Azagra, Ja vier Morlana, Ra wen Kader , Edward Sanderson, Bogdan J Matuszewski, Jae Y oung Lee, Dong-Jae Lee, et al. Simcol3d—3d reconstruction during colonoscopy challenge. Medical Image Analysis , 96:103195, 2024. 156 128 [904] Sebastian Bodenstedt, Max Allan, Anthony Agustinos, Xiaofei Du, Luis Garcia-Peraza- Herrera, Hannes Kenngott, Thomas Kurmann, Beat Müller-Stich, Sebastien Ourselin, Daniil Pakhomo v , et al. Comparative e valuation of instrument segmentation and tracking methods in minimally in v asi ve sur gery . arXiv pr eprint arXiv:1805.02475 , 2018. 156 [905] Max Allan, Ale x Shvets, Thomas K urmann, Zichen Zhang, Rahul Duggal, Y un-Hsuan Su, Nicola Riek e, Iro Laina, Ni v editha Kala v akonda, Sebastian Bodenstedt, et al. 2017 robotic instrument segmentation challenge. arXiv preprint , 2019. 156 [906] T obias Ross et al. Robust medical instrument segmentation challenge 2019. arXiv preprint arXiv:2003.10299 , 2020. 156 [907] Imanol Luengo, Maria Grammatikopoulou, Rahim Mohammadi, Chris W alsh, Chinedu Inno- cent Nwo ye, Deepak Alapatt, Nicolas Pado y , Zhen-Liang Ni, Chen-Chen F an, Gui-Bin Bian, et al. 2020 cataracts semantic segmentation challenge. arXiv pr eprint arXiv:2110.10965 , 2021. 156 [908] Markus W agner, Stefan Bodenstedt, et al. Endoscopic vision challenge (endovis) 2021: Heichole surgical workflo w analysis and full scene segmentation. arXiv pr eprint arXiv:2109.14956 , 2021. 156 [909] Hao Ding et al. Segstrong-c: Segmenting surgical tools robustly on non-adversarial generated corruptions – an endovis’24 challenge. arXiv preprint arXi v:2407.11906, 2024. 156 [910] Sophia Bano, Alessandro Casella, Francisco V asconcelos, Abdul Qayyum, Abdesslam Ben- zinou, Moona Mazher , Fabrice Meriaudeau, Chiara Lena, Ilaria Anita Cintorrino, Gaia Ro- mana De Paolis, Jessica Biagioli, Daria Grechishnikov a, Jing Jiao, Bizhe Bai, Y anyan Qiao, Binod Bhattarai, Rebati Raman Gaire, Ronast Subedi, Eduard V azquez, Szymon Płotka, Aneta Lisowska, Arkadiusz Sitek, George Attilakos, Ruw an W imalasundera, Anna L. David, Dario P aladini, Jan Deprest, Elena De Momi, Leonardo S. Mattos, Sara Moccia, and Danail Stoyanov . Placental vessel segmentation and registration in fetoscop y: Litera- ture revie w and miccai fetreg2021 challenge findings. Medical Image Analysis , 92:103066, 2024. ISSN 1361-8415. doi: https://doi.org/10.1016/j.media.2023.103066. URL https: //www.sciencedirect.com/science/article/pii/S1361841523003262 . 156 [911] Adrito Das, Danyal Z. Khan, Dimitrios Psychogyios, Y itong Zhang, John G. Hanrahan, Francisco V asconcelos, Y ou Pang, Zhen Chen, Jinlin W u, Xiaoyang Zou, Guoyan Zheng, Abdul Qayyum, Moona Mazher, Imran Razzak, T ianbin Li, Jin Y e, Junjun He, Szymon Płotka, Joanna Kaleta, Amine Y amlahi, Antoine Jund, Patrick Godau, Satoshi Kondo, Satoshi Kasai, K ousuke Hirasawa, Dominik Rivoir , Stefanie Speidel, Alejandra Pérez, Santiago Rodriguez, P ablo Arbeláez, Danail Stoyanov , Hani J. Marcus, and Sophia Bano. Pitvis- 2023 challenge: W orkflow recognition in videos of endoscopic pituitary surgery . Medi- cal Image Analysis , 106:103716, 2025. ISSN 1361-8415. doi: https://doi.org/10.1016/j. media.2025.103716. URL https://www.sciencedirect.com/science/article/pii/ S1361841525002634 . 156 [912] Aneeq Zia, Kiran Bhattacharyya, Xi Liu, Max Berniker , Ziheng W ang, Rogerio Nespolo, Satoshi K ondo, Satoshi Kasai, K ousuke Hirasawa, Bo Liu, et al. Surgical tool classification and localization: results and methods from the miccai 2022 surgtoolloc challenge. arXiv pr eprint arXiv:2305.07152 , 2023. 156 [913] M. Allan et al. Robust scene se gmentation in robotic endoscop y: Endovis 2018 robotics scene segmentation challenge. arXiv pr eprint arXiv:2001.11190 , 2020. Training dataset comprises frames from 16 robotic nephrectomy procedures recorded with da V inci Xi systems; an- notated classes include surgical instruments, suturing materials, anatomical structures, and background[935884295898146†L49-L72]. 156 [914] Dimitrios Psychogyios, Emanuele Colleoni, Beatrice V an Amsterdam, et al. Sar -rarp50: Seg- mentation of surgical instrumentation and action recognition on robot-assisted radical prosta- tectomy challenge. arXiv pr eprint arXiv:2401.00496 , 2024. Releases the first multimodal dataset of 50 suturing video se gments of robot-assisted radical prostatectomy , providing both instrument segmentation and action recognition labels. 156 129 [915] T obias Rückert, Da vid Rauber , Raphaela Mäerkl, Leonard Klausmann, Suemeyye R. Y ildiran, et al. Comparative validation of surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation in endoscopy: Results of the phakir 2024 challenge. arXiv pr eprint arXiv:2507.16559 , 2025. Dataset of 13 full-length laparoscopic videos with annotations for phase recognition, ke ypoint estimation, and instrument instance segmentation. 156 [916] Negin Ghamsarian, Mario T aschwer , Doris Putzgruber-Adamitsch, Stephanie Sarny , Y osuf El-Shabrawi, and Klaus Schoef fmann. Lensid: A cnn-rnn-based framework to wards lens irregularity detection in cataract surgery videos. In Medical Imag e Computing and Computer Assisted Intervention (MICCAI) , v olume 12908 of Lectur e Notes in Computer Science , pages 76–86. Springer , 2021. doi: 10.1007/978- 3- 030- 87237- 3_8. 156 [917] Ziyi W ang, Bo Lu, Y onghao Long, Fangxun Zhong, T ak-Hong Cheung, Qi Dou, and Y unhui Liu. Autolaparo: A new dataset of integrated multi-tasks for image-guided surgical automa- tion in laparoscopic hysterectomy . In International Confer ence on Medical Image Computing and Computer-Assisted Intervention , pages 486–496. Springer , 2022. 156 [918] W -Y Hong, C-L Kao, Y -H Kuo, J-R W ang, W -L Chang, and C-S Shih. Cholecseg8k: a semantic segmentation dataset for laparoscopic cholecystectomy based on cholec80. arXiv pr eprint arXiv:2012.12453 , 2020. 156 [919] Oluwatosin Alabi, K o K o Zayar T oe, Zijian Zhou, Charlie Budd, Nicholas Raison, Miao- jing Shi, and T om V ercauteren. Cholecinstanceseg: A tool instance segmentation dataset for laparoscopic surgery . Scientific Data , 12(1):825, 2025. 156 [920] Maria Grammatikopoulou, Evangello Flouty , Abdolrahim Kadkhodamohammadi, Gwenolé Quellec, Andre Chow , Jean Nehme, Imanol Luengo, and Danail Stoyanov . Cadis: Cataract dataset for sur gical rgb-image segmentation. Medical Image Analysis , 71:102053, 2021. 156 [921] Pietro Mascagni, Deepak Alapatt, Aditya Murali, Armine V ardazaryan, Alain Garcia, Nariaki Okamoto, Guido Costamagna, Didier Mutter, Jacques Marescaux, Bernard Dallemagne, et al. Endoscapes, a critical view of safety and surgical scene segmentation dataset for laparoscopic cholecystectomy . Scientific Data , 12(1):331, 2025. 156 [922] Matthias Carstens, Franziska M Rinner , Sebastian Bodenstedt, Alexander C Jenke, Jür gen W eitz, Marius Distler , Stefanie Speidel, and Fiona R Kolbinger . The dresden surgical anatomy dataset for abdominal organ segmentation in surgical data science. Scientific Data , 10(1):1–8, 2023. 156 [923] Sharib Ali, Debesh Jha, Noha Ghatwary , Stefano Realdon, Renato Cannizzaro, Osama E Salem, Dominique Lamarque, Christian Daul, Michael A Riegler , Kim V Anonsen, et al. A multi-centre polyp detection and segmentation dataset for generalisability assessment. Scien- tific Data , 10(1):75, 2023. 156 [924] Hemin Ali Qadir, Y ounghak Shin, Jacob Bergsland, and Ilangko Balasingham. Accurate real-time polyp detection in videos from concatenation of latent features extracted from con- secutiv e frames. In 2022 IEEE International Confer ence on Bioinformatics and Biomedicine (BIBM) , pages 2461–2466, 2022. doi: 10.1109/BIBM55620.2022.9995323. 157 [925] Sahar Nasirihaghighi, Negin Ghamsarian, Leonie Peschek, Matteo Munari, Heinrich Hus- slein, Raphael Sznitman, and Klaus Schoeffmann. Gynsurg: A comprehensiv e gynecology laparoscopic surgery dataset. arXiv preprint , 2025. 157 [926] João Cartucho, Alistair W eld, Samyakh Tukra, Haozheng Xu, Hiroki Matsuzaki, T aiyo Ishikawa, Minjun Kwon, Y ong Eun Jang, Kwang-Ju Kim, Gwang Lee, et al. Surgt chal- lenge: Benchmark of soft-tissue trackers for robotic surgery . Medical image analysis , 91: 102985, 2024. 157 [927] Adam Schmidt, Mert Asim Karaoglu, Soham Sinha, Mingang Jang, Ho-Gun Ha, Kyungmin Jung, K yeongmo Gu, Ihsan Ullah, Hyunki Lee, Jonáš Šer ` ych, et al. Point tracking in surgery– the 2024 sur gical tattoos in infrared (stir) challenge. arXiv pr eprint arXiv:2503.24306 , 2025. 157 130 [928] Stefanie Speidel, Lena Maier-Hein, Danail Sto yanov , Max Kirchner , Ale xander Jenke, Se- bastian Bodenstedt, Fiona K olbinger , Oliver Lester Saldanha, Jakob Nikolas Kather , S. K evin Zhou, Shang Zhao, Qiyuan W ang, Dai Sun, T obias Rueckert, Christoph P alm, Dirk W ilhelm, Hubertus Feußner, Daniel Rueckert, Hao Ding, Mathias Unberath, Adam Schmidt, Tim Sal- cudean, Omid Mohareri, Simon DiMaio, Hanna Hoffmann, Jan Egger , Setareh Bady , Frank Hölzle, Rainer Röhrig, Behrus Puladi, Rema Daher , Xinwei Ju, Razvan Caramalau, Baoru Huang, Francisco V asconcelos, Aneeq Zia, Max Berniker , Conor Perreault, Rogerio Nespolo, Ziheng W ang, Anthony Jarc, Annika Reinke, and Sophia Bano. Endoscopic vision challenge 2024 (endovis-classification-tracking + endovis-se gmentation). 27th International Confer - ence on Medical Image Computing and Computer Assisted Intervention (MICCAI 2024), 2024. URL https://doi.org/10.5281/zenodo.11119034 . 157 [929] Ryo Fujii, Masashi Hatano, Hideo Saito, and Hiroki Kajita. Egosur gery-phase: A dataset of surgical phase recognition from e gocentric open surgery videos. In MICCAI , 2024. 157 [930] Thyroid nodule segmentation and classification, 2020. https://tn- scui2020. grand- challenge.org/ . 157 [931] David Ouyang, Bryan He, Amirata Ghorbani, Neal Y uan, Joseph Ebinger, Curtis P Langlotz, Paul A Heidenreich, Robert A Harrington, David H Liang, Euan A Ashle y , et al. V ideo-based ai for beat-to-beat assessment of cardiac function. Natur e , 580(7802):252–256, 2020. 157 [932] The gastrointestinal atlas, 2000. http://www.gastrointestinalatlas.com/english/ english.html . 157 [933] Klaus Schoeffmann, Mario T aschwer, Stephanie Sarny , Bernd Münzer, Manfred Jürgen Primus, and Doris Putzgruber . Cataract-101: video dataset of 101 cataract surgeries. In Pr oceedings of the 9th A CM multimedia systems confer ence , pages 421–425, 2018. 157 [934] Deepak Gupta, Kush Attal, and Dina Demner-Fushman. A dataset for medical instructional video classification and question answering. Scientific Data , 10(1):158, 2023. 157 [935] A ysen Degerli, Morteza Zabihi, Serkan Kiranyaz, T ahir Hamid, Rashid Mazhar, Ridha Hamila, and Moncef Gabbouj. Early detection of myocardial infarction in low-quality echocardiography . IEEE Access , 9:34442–34453, 2021. 157 [936] Max Allan, Jonathan McLeod, Congcong W ang, Jean Claude Rosenthal, Zhenglei Hu, Niklas Gard, Peter Eisert, K e Xue Fu, Tre v or Zef firo, W enyao Xia, Zhanshi Zhu, Huoling Luo, Fucang Jia, Xiran Zhang, Xiaohong Li, Lalith Sharan, T om Kurmann, Sebastian Schmid, Raphael Sznitman, Dimitris Psychogyios, Mahdi Azizian, Danail Stoyano v , Lena Maier- Hein, and Stefanie Speidel. Stereo correspondence and reconstruction of endoscopic data challenge. arXiv pr eprint arXiv:2101.01133 , 2021. doi: 10.48550/arXiv .2101.01133. 157 [937] Jan Cychnerski, T omasz Dziubich, and Adam Brzeski. Ers: a novel comprehensive en- doscopy image dataset for machine learning, compliant with the mst 3.0 specification. arXiv pr eprint arXiv:2201.08746 , 2022. 157 [938] Jannis Born, Nina W iedemann, Manuel Cossio, Charlotte Buhre, Gabriel Brändle, K onstantin Leidermann, A vinash Aujayeb, Michael Moor , Bastian Rieck, and Karsten Bor gwardt. Ac- celerating detection of lung pathologies with explainable ultrasound image analysis. Ap- plied Sciences , 11(2):672, Jan 2021. ISSN 2076-3417. doi: 10.3390/app11020672. URL http://dx.doi.org/10.3390/app11020672 . 157 A T ables of 2D Medical Image Datasets B T ables of 3D Medical Image Datasets 131 T able 7: 2D CT datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 1 LoDoPaB-CT [ 107 ] 2020 2D CT Lung 28 Y es Recon NA 2 5K+ CT Images on Fractured Limbs [ 108 ] 2021 2D CT Limbs 24 Y es Seg Bone Fracture 3 AREN0534 [ 109 ] 2021 3D, 2D Multi a Kidney , Lung 239 No Est Kidney T umor 4 CT Medical Images [ 110 ] 2017 2D CT Lung 475 Y es Seg NA 5 National Lung Screening Trial [ 111 ] 2013 3D, 2D CT , Pathology Lung 26.7k No Cls Lung Cancer 6 RSNA Intracranial Hemorrhage Detection [ 112 ] 2019 2D CT Brain 874k Y es Loc Intracranial Hemorrhage 7 CT diagnosis of CO VID-19 [ 113 ] 2021 2D CT Lung 275 Y es Cls Lung COVID-19 8 CO VID-19-CT SCAN IMAGES [ 114 ] 2021 2D CT Lung 1.4k Y es Cls Lung COVID-19 9 CO VID_CT_COVID-CT [ 113 ] 2021 2D CT Lung 746 Y es Cls Lung COVID-19 10 Chest CT -Scan images Dataset [ 115 ] 2021 2D CT Lung 1k Y es Cls Lung Cancer 11 Cranium Image Dataset [ 116 ] 2020 2D CT Brain 50 Y es Det Intracranial Hemorrhage 12 SARS-CO V -2 Ct-Scan Dataset [ 117 ] 2021 2D CT Lung 2.5k Y es Cls Lung Disease 13 MedMNIST [ 118 ] 2020 2D Multi b Retina, Breast, Lung 100k Y es Cls Multi-disease 14 The V isible Human Project [ 119 ] 1994 3D, 2D CT, MR, etc. Full Body 2 No NA Skin Lesion 15 ImageCLEF 2016 [ 120 ] 2015 2D Multi c Skin, Cell, Breast 31k Y es Cls Head & Neck Tumor 16 RadImageNet (Subset: CT) [ 121 ] 2022 2D CT Full Body 292.4k Y es Cls Abdomen, lung, etc. d 17 Brain CT Images with ICH Masks [ 116 ] 2019 2D CT Brain 82 Y es Seg Intracranial Hemorrhage 18 CMB-CRC [ 122 ] 2022 3D, 2D Multi e Colon 472 No Seg, Cls Colorectal Cancer 19 CMB-GEC [ 123 ] 2022 3D, 2D CT, WSI, PET Brain 14 No Seg, Cls Melanoma 20 CMB-MEL [ 124 ] 2022 3D, 2D Multi f Brain 255 No Seg Melanoma 21 CMB-MML [ 125 ] 2021 2D, 3D Multi g NA 60 No NA Multiple Myeloma 22 CMB-PCA [ 126 ] 2022 2D, 3D CT, MR, WSI Prostate 31 No Cls, Pred Prostate Cancer 23 CPT AC-LSCC_CT_PET [ 127 ] 2018 2D, 3D CT , PET , Histopathology NA 238 No NA NA 24 Finding and Measuring Lungs in CT Data [ 128 ] 2019 2D, 3D CT Lung 534 Y es Seg NA 25 Head CT Image Data [ 129 ] 2019 2D CT Head 200 Y es Cls NA 26 LDCTIQA C2023 [ 130 ] 2023 2D CT NA 1k Y es Reg NA 27 APOLLO-5 [ 131 ] 2022 2D, 3D Multi h NA 6.2k No NA NA 28 Lung-Fused-CT -Pathology [ 132 ] 2018 2D, 3D CT , Histopathology Lung 36 Y es Seg Lung Disease 29 CMB-LCA [ 133 ] 2022 2D, 3D Multi i NA 0 No NA NA 30 RIDER Phantom PET -CT [ 134 ] 2011 2D CT , PET NA 2.2k No NA NA 31 AHOD0831 [ 135 ] 2022 3D, 2D Multi j NA 0 No NA Hodgkin Lymphoma 32 Prostate-MRI [ 136 ] 2011 3D, 2D Multi k Prostate 26 No NA Prostate Cancer 33 AREN0532 [ 137 ] 2022 3D, 2D Multi l NA 1k No NA Wilms T umor 34 ImageCLEF 2016 (Duplicate) [ 120 ] 2015 2D Multi c Skin, Cell, Breast 31k Y es Cls Head & Neck Tumor 35 QUBIQ2020 [ 138 ] 2020 2D CT , MR Kidney , Pancreas, etc. 150 Y es Seg NA 36 QUBIQ2021_2D_CT [ 139 ] 2021 2D CT , MR Kidney , Pancreas, etc. 268 Y es Seg NA Overall 1994 ∼ 2022 2D Multi Full Body 1.4m NA Multi Multi a Multi-modalities of AREN0534: CT , MR, PET , Ultrasound. b Multi-modalities of MedMNIST : OCT , X-Ray, CT , Pathology , Fundus Photography . c Multi-modalities of ImageCLEF: MR, US, Histopathology , X-Ray , CT , PET , Endoscop y , Dermoscopy , EEG, ECG, EMG, Microscopy , Fundus. d The complete list of diseases for RadImageNet includes: prostate lesion, adrenal pathology , gallstone, arterial pathology , urolithiasis, pancreatic lesion, etc. e Multi-modalities of CMB-CRC: CT , MR, US, X-ray , PET , WSI. f Multi-modalities of CMB-MEL: CT , US, WSI, PET . g Multi-modalities of CMB-MML: CT , MR, PET , WSI. h Multi-modalities of APOLLO-5: CT , MR, US, PET , X-Ray . i Multi-modalities of CMB-LCA: CT , MR, US, Histopathology , X-ray . j Multi-modalities of AHOD0831: CT , MR, PET , X-Ray . k Multi-modalities of Prostate-MRI: MR, CT , PET , Pathology . l Multi-modalities of AREN0532: CT , MR, Ultrasound, PET . Abbre viations: Cls=Classification, Det=Detection, Est=Estimation, Histo=Histopathology , Loc=Localization, Pred=Prediction, Recon=Reconstruction, Reg=Registration, Se g=Segmentation, US=Ultrasound, WSI=Whole-slide images. 132 T able 8: 2D MRI datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 37 AREN0534 [ 109 ] 2021 2D, 3D Multi a Kidney , Lung 239 No Est Kidney Tumor 38 KNOAP2020 [ 140 ] 2020 2D, 3D MR, X-Ray Knee 30 Y es Pred Osteoarthritis 39 braimMRI [ 141 ] 2022 2D MR Brain 110 Y es Seg Brain T umor 40 Brain-MRI [ 142 ] 2020 2D MR Brain 110 Y es Seg Brain Disease 41 SpinalDisease2020 [ 143 ] 2020 2D MR Spine 150 Y es Det Spinal Disease 42 The Visible Human Project [ 119 ] 1994 2D, 3D CT , MR, Others Full Body 2 No NA Skin Lesion 43 ImageCLEF 2016 [ 144 ] 2015 2D Multi b Skin, Cell, Breast 31k Y es Cls H&N Tumor 44 CMB-CRC [ 145 ] 2022 2D, 3D Multi c Colon 472 No Seg, Cls Colorectal Cancer 45 CMB-MML [ 146 ] 2021 2D, 3D Multi d NA 60 No Pred Multiple Myeloma 46 CMB-PCA [ 126 ] 2022 2D, 3D CT , MR, Histo Prostate 31 No Cls, Pred Prostate Cancer 47 ICDC-Glioma (GLIOMA01)_3D-MR 2021 2D, 3D MR, Histo NA 650 No NA Glioma 48 Prostate Fused-MRI-Pathology [ 147 ] 2016 2D, 3D MR, Histo Prostate (Pelvis) 29 No NA Prostate Cancer 49 Cardiac Atrial Images [ 148 ] 2020 2D MR Atrium 8k Y es Se g Cardiac Disease 50 APOLLO-5 [ 149 ] 2022 2D, 3D Multi e NA 6.2k No NA NA 51 CMB-LCA [ 146 ] 2022 2D, 3D Multi f NA 0 No NA Lung Cancer 52 AHOD0831 [ 135 ] 2022 2D, 3D Multi g NA 0 No NA Hodgkin Lymphoma 53 Prostate-MRI [ 150 ] 2011 2D, 3D Multi h Prostate 26 No NA Prostate Cancer 54 AREN0532 [ 137 ] 2022 2D, 3D Multi i NA 1k No NA Wilms T umor 55 ImageCLEF 2015 [ 151 ] NA 2D, 3D Multi j Skin, Cell, Breast 0 Y es Cls NA 56 RadImageNet (Subset: MR) [ 121 ] 2022 2D MR Full Body 673k Y es Cls Whole Body Abnorm. 57 QUBIQ2020 [ 138 ] 2020 2D CT , MR Kidney , etc. k 150 Y es Se g Pathologies 58 QUBIQ2021_2D_MR [ 139 ] 2021 2D CT , MR Kidney , etc. k 268 Y es Se g Pathologies Overall 1994 ∼ 2022 2D Multi Full Body 721.5k NA Multi Multi a Multi-modalities of AREN0534: CT , MR, PET , US. b Multi-modalities of ImageCLEF 2016: MR, US, Histo, X-Ray , CT , PET , Endo, Dermo, EEG, ECG, EMG, Micro, Fundus. c Multi-modalities of CMB-CRC: CT , MR, US, X-Ray , PET , Histo. d Multi-modalities of CMB-MML: CT , MR, PET , Histo. e Multi-modalities of APOLLO-5: CT , MR, US, PET , X-Ray . f Multi-modalities of CMB-LCA: CT , MR, US, Histo, X-Ray . g Multi-modalities of AHOD0831: CT , MR, PET , X-Ray . h Multi-modalities of Prostate-MRI: MR, CT , PET , Histo. i Multi-modalities of AREN0532: CT , MR, US, PET . j Multi-modalities of ImageCLEF 2015: MR, US, Histo, X-Ray . k etc. in QUBIQ Structures: Pancreas, Brain, Prostate. Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Pred=Prediction, Est=Estimation; Histo=Histopathology , US=Ultrasound, Endo=Endoscopy , Dermo=Dermoscop y , Micro=Microscopy , Abnorm.=Abnormalities, H&N=Head & Neck. T able 9: 2D PET datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 59 AREN0534 [ 109 ] 2021 3D, 2D Multi a Kidney , Lung 239 No Est Kidney 60 ImageCLEF 2016 [ 120 ] 2015 2D Multi b Skin, Cell, Breast 31k Y es Cls H&N Tumor 61 CMB-CRC [ 122 ] 2022 3D, 2D Multi c Colon 472 No Seg, Cls Colorectal Cancer (H&E stained tissue) 62 CMB-GEC [ 123 ] 2022 3D, 2D CT , WSI, PET Brain 14 No Seg, Cls Melanoma (Cerebral microbleeds) 63 CMB-MEL [ 124 ] 2022 3D, 2D Multi d Brain 255 No Seg Melanoma (Cerebral microbleeds) 64 CMB-MML [ 125 ] 2021 2D, 3D Multi e NA 60 No Pred Multiple Myeloma 65 CPT A C-LSCC_CT_PET [ 127 ] 2018 2D, 3D CT, PET , Histo N A 238 No NA NA 66 APOLLO-5 [ 131 ] 2022 2D, 3D Multi f NA 6.2k No NA NA 67 RIDER Phantom PET -CT [ 134 ] 2011 2D CT , PET N A 2.2k No NA NA 68 AHOD0831 [ 135 ] 2022 3D, 2D Multi g NA 0 No NA Hodgkin Lymphoma 69 AREN0532 [ 137 ] 2022 3D, 2D Multi i NA 1k No N A W ilms Tumor Overall 2011 ∼ 2022 2D Multi Full Body 41.7k NA Multi Multi a Multi-modalities of AREN0534: CT , MR, PET , Ultrasound. b Multi-modalities of ImageCLEF 2016: MR, US, Histo, X-Ray , CT , PET , Endo, Derm, EEG, ECG, EMG, Microscopy , Fundus. c Multi-modalities of CMB-CRC: CT , MR, US, DX, PET , WSI. d Multi-modalities of CMB-MEL: CT , US, WSI, PET (SWI). e Multi-modalities of CMB-MML: CT , MR, PET , WSI. f Multi-modalities of APOLLO-5: CT , MR, US, PET , X-Ray . g Multi-modalities of AHOD0831: CT , MR, PET , X-Ray . h Multi-modalities of Prostate-MRI: MR, CT , PET , Patho. i Multi-modalities of AREN0532: CT , MR, US, PET . Abbre viations: Seg=Se gmentation, Cls=Classification, Est=Estimation, Pred=Prediction, H&N=Head & Neck, US=Ultrasound, Histo=Histopathology , Patho=P athology , Endo=Endoscop y , Derm=Dermoscopy , WSI=Whole-slide Images, DX=Digital Radiography . 133 T able 10: 2D Ultrasound datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 70 HC18 [ 152 ] 2018 2D US Skull 1.3k Y es Meas NA 71 BUSI [ 153 ] 2019 2D US Breast 647 Y es Seg Breast Cancer 72 APOLLO-5 [ 149 ] 2022 2D, 3D Multi a NA 6.2k No NA NA 73 CMB-LCA [ 146 ] 2022 2D, 3D Multi b NA 0 No NA NA 74 ImageCLEF 2015 [ 151 ] 2015 2D, 3D Multi c Skin, Cell, Breast 0 Y es Cls NA 75 ImageCLEF 2016 [ 120 ] 2016 2D Multi d Skin, Cell, Breast 31k Y es Cls Head & Neck T umor 76 RadImageNet (Subset: US) [ 121 ] 2022 2D US Full Body 390k Y es Cls Abdominal Structures 77 BreastMNIST [ 153 ] 2021 2D US Breast 156 Y es Cls Breast Cancer 78 AREN0534 [ 109 ] 2021 2D, 3D Multi e Kidney , Lung 239 No Est Kidney Tumor 79 CLUST15 [ 154 ] 2015 2D US Liv er 34 Y es Track NA 80 Ultrasound Nerve Segmentation [ 155 ] 2016 2D US Brachial Plexus 11.3k Y es Seg NA 81 TN-SCUI2020 [ 156 ] 2020 2D US Thyroid 3.6k Y es Seg Leukemia 82 ImageCLEF 2016 [ 157 ] 2015 2D Multi f Skin, Cell, Breast 31k Y es Cls Head & Neck T umor 83 CMB-CRC [ 145 ] 2022 2D, 3D Multi g Colon 472 No Seg, Cls Colorectal Cancer 84 CMB-MEL [ 158 ] 2022 2D, 3D Multi h Brain 255 No Seg Melanoma, Cerebral microbleed 85 PSFHS [ 159 ] 2023 2D US NA 4.7k Y es Seg NA 86 USenhance2023 [ 160 ] 2023 2D US NA 1.5k Y es Recon NA 87 AREN0532 [ 137 ] 2022 2D, 3D Multi i NA 1k No NA Wilms T umor 88 TN3K [ 161 ] 2021 2D US Head and Neck 3.5k Y es Seg Thyroid Nodules 89 CAMUS [ 162 ] 2019 2D US Heart 1.8k Y es Seg Cardiac Disease 90 DDTI [ 163 ] 2020 2D US Thyroid 637 Y es Seg Thyroid Nodule 91 UDIA T -B [ 164 ] 2017 2D US Breast 163 Y es Det Breast Lesion 92 OASB UD [ 165 ] 2017 2D US Breast 200 Y es Se g Breast Cancer 93 BrEaST [ 166 ] 2024 2D US Breast 256 Y es Cls Breast Cancer Overall 2015 ∼ 2024 2D Multi Full Body 490.0k NA Multi Multi a Multi-modalities of APOLLO-5: CT , MR, US, PET , X-Ray . b Multi-modalities of CMB-LCA: CT , MR, US, Histo, DX (WSI). c Multi-modalities of ImageCLEF 2015: MR, US, Histo, X-Ray . d Multi-modalities of ImageCLEF 2016 [ 120 ]: MR, US, Histo, X-Ray , CT , PET , Endo, Dermo, EEG, ECG, EMG, Microscopy , Fundus (Electron Microscopy). e Multi-modalities of AREN0534: CT , MR, PET , US. f Multi-modalities of ImageCLEF 2016: MR, US, Histo, X-Ray , CT , PET , Endo, Dermo, Others, EEG, ECG, EMG, Electron Microscopy , Fundus Photography . g Multi-modalities of CMB-CRC: CT , MR, US, DX, PET , WSI. h Multi-modalities of CMB-MEL: CT , US, WSI, PET (SWI). i Multi-modalities of AREN0532: CT , MR, US, PET . Abbre viations: Seg=Se gmentation, Cls=Classification, Est=Estimation, Recon=Reconstruction, Meas=Measurement, Track=T racking; US=Ultrasound, Histo=Histopathology , WSI=Whole-slide image, Endo=Endoscopy , Dermo=Dermoscopy . 134 T able 11: 2D X-Ray datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 94 Chest X-ray [ 167 ] 2018 2D X-Ray Lung 5.9k Y es Cls Pneumonia 95 CoronaHack [ 168 ] 2020 2D X-Ray Lung 5.9k Y es Cls CO VID-19, Pneumonia 96 NIH Chest X-ray 14 [ 169 ] 2017 2D X-Ray Lung 112.1k Y es Cls Thorax diseases 97 COVIDx CXR-2 [ 170 ] 2020 2D X-Ray Lung 30.9k Y es Cls CO VID-19 98 Pneumothorax Masks X-Ray [ 171 ] 2020 2D X-Ray Lung 12.0k Y es Seg Pneumothorax 99 IRMA X-ray [ 172 ] 2020 2D X-Ray Brain, Lung 14.7k Y es Cls NA 100 Chest XR COVID-19 [ 173 ] 2021 2D X-Ray Lung 21.4k Y es Cls CO VID-19 101 COVID-19-Image [ 174 ] 2020 2D X-Ray Lung 93 Y es Cls COVID-19 102 Chest X-ray P A Dataset [ 175 ] 2021 2D X-Ray Lung 4.6k No Cls COVID-19, Pneumonia 103 NHANES II X-ray [ 176 ] 2021 2D X-Ray Lung 17.1k No NA N A 104 KNOAP2020 [ 140 ] 2020 2D, 3D MR, X-Ray Knee 30 Y es Pred Osteoarthritis 105 AASCE [ 177 ] 2019 2D X-Ray Spine 609 Y es Reg NA 106 Covid-19 Image Dataset [ 178 ] 2021 2D X-Ray Lung 345 Y es Cls Lung diseases 107 Pulmonary Chest X-Ray (ChinaSet) [ 179 ] 2021 2D X-Ray Lung 800 Y es Cls Lung diseases 108 MURA [ 180 ] 2021 2D X-Ray Multi-bone a 40.0k Y es Cls Musculoskeletal 109 SIIM-ACR Pneumothorax Se g [ 171 ] 2020 2D X-Ray Lung 12.1k Y es Seg Pneumothorax 110 MIAS Mammography [ 181 ] 2021 2D X-Ray Breast 322 Y es Cls Breast cancer 111 MedMNIST [ 118 ] 2020 2D Multi b Retina, Breast, Lung 100k Y es Cls Multi-diseases 112 RSNA Pneumonia Detection [ 182 ] 2018 2D X-Ray Lung 26.7k Y es Det Lung diseases 113 V inBigData Chest X-ray [ 183 ] 2020 2D X-Ray Lung 15.0k Y es Det Heart atrium 114 CheXpert [ 64 ] 2021 2D X-Ray Lung 224.3k Y es Cls Diabetic retinopathy 115 SIIM-FISABIO-RSNA CO VID-19 [ 184 ] 2021 2D X-Ray Lung 6.1k Y es Det Tuberculosis 116 NODE21 [ 185 ] 2021 2D X-Ray Lung 5.5k Y es Det Breast cancer 117 ImageCLEF 2016 [ 120 ] 2016 2D Multi c Skin, Cell, Breast 31.0k Y es Cls Head & Neck tumor 118 TCB-Challenge [ 186 ] 2016 2D X-Ray Bone 174 Y es Cls Osteoporotic bone 119 CRASS [ 187 ] 2012 2D X-Ray Clavicle 518 Y es Seg Clavicles 120 COVIDGR [ 188 ] 2020 2D X-Ray Lung 852 Y es Cls COVID-19 121 ChestX-Det [ 189 ] 2021 2D X-Ray Lung 3.6k Y es Seg Lung diseases 122 RANZCR CLiP [ 190 ] 2020 2D X-Ray Breast 30.1k Y es Cls NA 123 CPCXR [ 191 ] 2020 2D X-Ray Lung 1.2k Y es NA Pneumonia, CO VID-19 124 JSR T [ 192 ] 2000 2D X-Ray Lung 247 Y es Cls Lung nodule 125 Synthetic COVID-19 CXR [ 193 ] 2020 2D X-Ray Lung 21.3k Y es Cls, Gen CO VID-19 126 Cephalometric X-ray Image [ 194 ] 2014 2D X-Ray Skull 400 Y es Loc NA 127 CMB-CRC [ 145 ] 2022 2D, 3D Multi d Colon 472 No Seg, Cls Colorectal cancer 128 MIDRC-RICORD-1c [ 195 ] 2021 2D X-Ray Lung 1.3k Y es Cls NA 129 Chest X-ray Imaging [ 196 ] 2017 2D X-Ray Lung 5.9k Y es Cls NA 130 COVID-19 Chest X-ray DB [ 197 ] 2021 2D X-Ray NA 3.9k Y es Cls COVID-19 131 SZ-CXR [ 198 ] 2018 2D X-Ray Lung 566 Y es Seg NA 132 Pulmonary Chest X-Ray Seg [ 179 ] 2021 2D X-Ray Lung 800 Y es Seg Lung diseases 133 DENTEX [ 199 ] 2023 2D X-Ray Brain 1.0k Y es Det NA 134 CL-Detection2023 [ 200 ] 2023 2D X-Ray NA 555 Y es Det N A 135 ISBI2023 CEPHA29 [ 201 ] NA 2D X-Ray NA 1.0k Y es Loc NA 136 ARCADE [ 202 ] 2023 2D X-Ray NA 1.5k Y es Seg NA 137 MedFM2023 [ 203 ] 2023 2D X-Ray NA 4.8k Y es Cls NA 138 CoronARe [ 204 ] NA 2D X-Ray N A 0 Y es Recon Coronary artery diseases 139 VICTRE [ 205 ] 2019 2D X-Ray Breast 217.9k No NA N A 140 APOLLO-5 [ 149 ] 2022 2D, 3D Multi e NA 6.2k No NA N A 141 CMB-LCA [ 146 ] 2022 2D, 3D Multi f NA 0 No NA N A 142 AHOD0831 [ 135 ] 2022 2D, 3D Multi g NA 0 No NA Hodgkin L ymphoma 143 CheXmask [ 206 ] 2023 2D X-Ray NA 676.8k Y es Seg Lung diseases 144 Knee Osteoarthritis Dataset [ 207 ] 2020 2D X-Ray Knee 0 Y es Cls Knee osteoarthritis 145 RUS_CHN [ 208 ] 2021 2D X-Ray Hand 0 Y es Cls Hand joints 146 RSNA Bone Age [ 209 ] 2017 2D X-Ray Hand 14.2k Y es Est Hand bone 147 CXR-L T [ 210 ] 2023 2D X-Ray Breast, Lung 377.1k Y es Cls Multi-diseases 148 PENGWIN2024-T ask2 [ 211 , 212 ] 2025 2D X-Ray Pelvic Bone 150 Y es Seg Pelvic bone fragments 149 ICG-CXR [ 213 ] 2025 2D X-Ray Lung 11.4k Y es Gen Lung diseases Overall 2014 ∼ 2025 2D Multi Full Body 2.1m NA Multi Multi a Structures of MURA: Elbow , Finger, Forearm, Hand, Humerus, Shoulder , Wrist. b Multi-modalities of MedMNIST : OCT , X-Ray, CT , Pathology , Fundus. c Multi-modalities of ImageCLEF 2016: MR, US, Histo, X-Ray , CT , PET , Endo, Derm, EEG, ECG, EMG, Microscopy , Fundus. d Multi-modalities of CMB-CRC: CT , MR, US, DX, PET , WSI. e Multi-modalities of APOLLO-5: CT , MR, US, PET , X-Ray . f Multi-modalities of CMB-LCA: CT , MR, US, Histo, DX. g Multi-modalities of AHOD0831: CT , MR, PET , X-Ray . Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Recon=Reconstruction, Reg=Registration, Loc=Localization, Est=Estimation, Pred=Prediction, Gen=Generation. 135 T able 12: 2D OCT datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 150 OCT2017 [ 196 ] 2018 2D OCT Retina 83.5k Y es Cls NA 151 Retinal OCT - C8 [ 41 ] 2021 2D OCT Retina 24k Y es Cls NA 152 iChallenge - AGE19 [ 214 ] 2019 2D OCT Retina 1.6k Y es Cls NA 153 DRAC22 [ 215 ] 2022 2D OCT Retina 174 Y es Seg Diabetic Retinopathy Lesions 154 iChallenge - GOALS [ 216 ] 2022 2D OCT Retina 300 Y es Seg NA 155 Eye OCT Datasets [ 217 , 218 ] 2021 2D OCT Retina 148 Y es Cls NA 156 APTOS-2021 [ 219 ] 2022 2D OCT Retina 2.6k Y es Pred Diabetic Retinopathy 157 APTOS Cross-Country Datasets_s tage1 [ 219 ] 2022 2D OCT Retina 2.6k Y es Pred NA 158 MedMNIST [ 118 ] 2020 2D Multi a Retina, Breast, Lung 100k Y es Cls NA 159 Canada OCT Retinal Images (Subset) [ 220 ] 2018 2D OCT Retina 25 Y es Seg Retinal Structures 160 SinaFarsiu-002-Fang_TMI_2013 [ 221 ] 2013 2D OCT Retina 195 Y es Seg NA 161 SinaFarsiu-003-Fang_BOE_2012 [ 221 ] 2012 2D OCT Retina 51 Y es Seg NA 162 SinaFarsiu-008-Chiu_BOE_2012 [ 221 ] 2012 2D OCT Retina 23 Y es Seg NA 163 SinaFarsiu-009-Chiu_BOE_2013 [ 221 ] 2013 2D OCT Retina 840 Y es Seg NA 164 SinaFarsiu-010-Rabbani_IO VS_2014 [ 222 ] 2015 2D OCT Retina 24 Y es Seg NA 165 SinaFarsiu-012-Estrada_TMI_2015 [ 223 ] 2015 2D OCT Retina 60 Y es Seg NA 166 SinaFarsiu-013-Estrada_P AMI_2015 [ 224 ] 2015 2D OCT Retina 90 Y es Seg NA 167 SinaFarsiu-018-Y ang_BOE_2021 [ 225 ] 2021 2D OCT Retina 784 Y es Seg NA 168 APTOS Cross-Country Datasets_s tage2 [ 219 ] 2022 2D OCT Retina 3.3k Y es Pred Diabetic Retinopathy 169 OCT A-500_2D-Fundus [ 226 ] 2020 2D OCT Retina 500 Y es Seg N/A 170 OCT A2024 (MuT ri)_2D-Fundus [ 227 ] 2024 2D OCT Retina 848 Y es Seg NA Overall 2012 ∼ 2022 2D Multi Retina, Breast, Lung 221.7k Y es Multi Multi a Multi-modalities of MedMNIST : OCT , X-Ray, CT , Pathology , Fundus Photography . Abbre viations: Seg=Se gmentation, Cls=Classification, Pred=Prediction, Det=Detection, Recon=Reconstruction, Reg=Registration, Loc=Localization, Est=Estimation. 136 T able 13: 2D fundus datasets. # Dataset Y ear Dim Modality Structur e Images Label T ask Diseases 171 DRISHTI-GS [ 228 ] 2014 2D Fundus Photo Retina 101 Y es Seg Optic Disc 172 CHASE [ 229 ] 2009 2D Fundus Photo Retina 28 Y es Seg NA 173 ST ARE [ 230 ] 2004 2D Fundus Photo Retina 40 Y es Seg NA 174 DRIVE [ 231 ] 2003 2D Fundus Photo Retina 40 Y es Seg NA 175 IDRID2018 [ 232 ] 2018 2D Fundus Photo Retina 81 Y es Seg, Cls Diabetic Retinopathy 176 EyeP A CS [ 233 ] 2015 2D Fundus Photo Retina 88.7k Y es Cls Diabetic Retinopathy 177 DRHAGIS [ 234 ] 2017 2D Fundus Photo Retina 40 Y es Seg DR Lesions 178 ODIR [ 235 ] 2019 2D Fundus Photo Retina 8k Y es Cls Ocular Diseases (DR screening) 179 RIADD (RFMiD) [ 236 ] 2020 2D Fundus Photo Retina 3.2k Y es Cls Retinal Diseases 180 MESSIDOR-2 [ 237 ] 2013 2D Fundus Photo Retina 1.7k Y es Cls Diabetic Retinopathy 181 iChallenge-ADAM [ 238 ] 2020 2D Fundus Photo Retina 400 Y es Cls Diabetic Retinopathy 182 AIROGS [ 239 ] 2021 2D Fundus Photo Retina 101.4k No Cls Diabetic Retinopathy 183 DiaRetDB [ 240 ] 2009 2D Fundus Photo Retina 89 No Det DR Lesions 184 HRF [ 241 ] NA 2D Fundus Retina 45 No Seg NA 185 iChallenge-P ALM19 [ 242 ] 2019 2D Fundus Retina 800 Y es Seg NA 186 Retina Fundus Image Reg. [ 243 ] 2021 2D Fundus Photo Retina 129 Y es Reg NA 187 APTOS-2019 [ 244 ] 2021 2D Fundus Photo Retina 3.7k Y es Cls Diabetic Retinopathy 188 MedMNIST [ 41 ] 2020 2D Multi a Retina, Breast, Lung 100k Y es Cls NA 189 DeepDR-T ask1 [ 245 ] 2020 2D Fundus Photo Eye V essel 2k Y es Cls Breast Cancer 190 ImageCLEF 2016 [ 246 ] 2015 2D Multi b Skin, Cell, Breast 31k Y es Cls Head & Neck Tumor 191 RITE [ 247 ] 2013 2D Fundus Retina 40 Y es Seg Retinal V essel 192 GAMMA (T ask1, CFP) [ 248 ] 2021 2D Fundus (CFP) Retina 200 Y es Cls Grading 193 RIM-ONE [ 249 ] 2020 2D Fundus Retina 485 Y es Seg Optic Disc and Cup 194 APTOS 2019 Blindness Det. [ 219 ] 2019 2D Fundus Retina 5.6k Y es Cls Grading 195 Glaucoma Detection [ 250 ] 2020 2D Fundus Retina 650 Y es Cls Glaucoma 196 ACRIMA [ 251 ] 2019 2D Fundus Retina 705 Y es Cls Glaucoma 197 AO-SLO Photoreceptor Seg. [ 252 ] 2013 2D Fundus Retina 840 Y es Seg A O-SLO Cone Photoreceptor 198 Arteriovenous Nicking [ 253 ] NA 2D Fundus Retina 90 Y es Cls Retinal Artery-V ein Nicking 199 Retina [ 254 ] 2019 2D Fundus Retina 601 Y es Cls Fundus Diseases 200 Y angxi [ 255 ] 2019 2D Fundus Retina 20.4k Y es Cls Eye Axis 201 William Hoyt [ 256 ] 2004 2D Fundus Retina 856 Y es Cls Fundus Diseases 202 V ampire [ 256 ] 2011 2D Fundus Retina 8 Y es Seg V essel 203 Retinal Fundus Imgs for Glaucoma [ 257 ] 2018 2D Fundus Retina 2.9k Y es Cls NA 204 RetinaCheck (IOST AR) [ 258 ] 2016 2D Fundus Retina 30 Y es Seg V essel 205 Ophthalmic Slit Lamp [ 259 ] 2018 2D Fundus Retina 60 No N A NA 206 Miles Iris [ 254 ] 2013 2D Fundus (Iris) Retina 833 No Cls Retinal Structures 207 JSIEC [ 260 ] 2019 2D Fundus Retina 1k Y es Cls Fundus Diseases 208 INSPIRE (Stereo) [ 254 ] 2011 2D Fundus Retina 30 Y es Reg NA 209 INSPIRE (A VR) [ 254 ] 2011 2D Fundus Retina 40 Y es Reg NA 210 HRF Quality Assessment [ 261 ] 2013 2D Fundus Retina 36 Y es Reg NA 211 HRF Segmentation [ 241 ] 2013 2D Fundus Retina 45 Y es Seg V essel 212 iChallenge-REFUGE2 [ 262 ] 2020 2D Fundus Photo (CFP) Retina 1.6k Y es Cls Glaucoma 213 GAMMA [ 248 ] 2021 2D, 3D Fundus Retina 200 Y es Cls NA 214 OIA-ODIR [ 235 ] 2019 2D Fundus NA 10k Y es Cls NA 215 V ARP A [ 263 ] 2019 2D Fundus Retina 58 Y es Cls NA 216 OR VS [ 264 ] 2020 2D Fundus Retina 49 Y es Seg NA 217 Retinal Img Quality Assess [ 265 ] 2020 2D Fundus Retina 216 Y es Cls NA 218 iChallenge-GAMMA_3D-OCT [ 248 ] 2021 2D Fundus Retina 300 Y es Seg Glaucoma 219 DeepDR-T ask2 [ 245 ] 2020 2D Fundus NA 2k Y es Reg NA 220 DeepDR-T ask3 [ 245 ] 2020 2D Fundus NA 246 Y es Cls NA 221 MMAC2023 [ 266 ] 2023 2D Fundus NA 0 Y es Cls NA 222 RFMiD 2.0 [ 236 ] 2023 2D Fundus Photo NA 860 Y es Cls Retinal Fundus Multi-Disease 223 MuReD [ 267 ] 2022 2D Fundus Photo NA 2.2k Y es Cls Retinal Diseases 224 Retinal V essel T ortuosity 2008 2D Fundus Photo Retina 60 Y es Reg NA 225 ImageCLEF 2016 NA 2D Multi c Skin, Cell, Breast 31k Y es Cls NA 226 P ARA GUA Y [ 268 ] NA 2D Fundus Photo NA 0 Y es Cls Diabetic Retinopathy 227 BEH [ 269 ] NA 2D Fundus Photo NA 0 Y es NA Glaucoma 228 BiDR NA 2D Fundus Photo NA 0 Y es NA Diabetic Retinopathy 229 HarvardGlaucoma NA 2D Fundus Photo NA 0 Y es NA Glaucoma 230 FUND NA 2D Fundus Photo NA 0 Y es NA NA 231 LAG [ 270 ] NA 2D Fundus Photo NA 0 Y es NA Glaucoma 232 DHRF NA 2D Fundus Photo Retina 6.2k Y es Cls Diabetic Retinopathy 233 E-ophta NA 2D Fundus Photo Retina 926 Y es Seg N A 234 FIVES [ 271 ] NA 2D Fundus Photo Retina 800 Y es Seg V essel 235 OcularD [ 272 ] NA 2D Fundus Photo Retina 6.4k Y es Cls NA 236 P APILA [ 273 ] NA 2D Fundus Photo Retina 488 Y es Seg N A 237 Papilledema [ 274 ] 2018 2D Fundus Photo Retina 1.4k Y es Cls Papilledema 238 ROD 2023 2D Fundus Photo Retina 281 Y es Cls Retinal Occlusion 239 T oxoFundus [ 275 ] 2023 2D Fundus Photo Retina 411 Y es Cls Ocular T oxoplasmosis 240 GAMMA (T ask3, CFP) [ 248 ] 2021 2D Fundus (CFP) Retina 200 Y es Seg Optic Disc and Cup 241 iChallenge-GAMMA_2D- Fundus [ 248 ] 2021 2D Fundus Retina 300 Y es Seg Glaucoma Overall 2003 ∼ 2023 2D Multi Multi 443.1k NA Multi Multi a Multi-modalities of MedMNIST: OCT , X-Ray , CT , P athology , Fundus Photography . b Multi-modalities of ImageCLEF 2016: MR, US, Histopathology, X-Ray , CT , PET , Endoscopy , Dermoscopy , EEG, ECG, EMG, Electron Microscop y, Fundus Photograph y. c Multi-modalities of ImageCLEF 2016: MR, US, Histopathology, X-Ray , CT , PET , Endoscopy , Dermoscopy , EEG, ECG, EMG, Microscop y, Fundus Photograph y. Abbreviations: Seg=Segmentation, Det=Detection, Cls=Classification, Reg=Registration, US=Ultrasound, DR=Diabetic Retinopathy . 137 T able 14: 2D dermoscopy datasets. # Dataset Y ear Dim Modality Structur e Images Label T ask Diseases 242 ISIC18 [ 276 ] 2018 2D Dermoscopy Skin 2.7k Y es Seg Skin lesion 243 ISIC20 [ 277 ] 2020 2D Dermoscopy Skin 33.1k Y es Cls Benign melanoma, malignant melanoma 244 ISIC16 [ 278 ] 2016 2D Dermoscopy Skin 1.3k Y es Seg Skin lesion 245 ISIC17 [ 279 ] 2016 2D Dermoscopy Skin 2.8k Y es Seg Skin lesion 246 Derm7pt [ 280 ] 2021 2D Dermoscopy Skin 2.0k Y es Cls Skin lesion 247 ISIC19 [ 25 ] 2019 2D Dermoscopy Skin 25.3k Y es Cls Cells 248 Fizpatrick 17k [ 281 ] 2021 2D Dermoscopy Skin 16.6k Y es Cls NA 249 MED-NODE [ 282 ] 2015 2D Dermoscopy Skin 170 Y es Cls Brain 250 P AD-UFES-20 [ 283 ] 2020 2D Dermoscopy Skin 2.3k Y es Cls Thoracic diseases 251 PH2 [ 284 ] 2014 2D Dermoscopy Skin 200 Y es Cls Cells 252 DFUC 2020 [ 285 ] 2020 2D Dermoscopy Foot 2.0k Y es Seg Breast cancer 253 SD-128 / SD-198 / SD-260 [ 286 , 287 ] 2021 2D Dermoscopy Skin 6.6k Y es Cls Fetal structure 254 ImageCLEF 2016 [ 144 ] 2015 2D Multi a Skin, Cell, Breast 31k Y es Cls Head & neck tumor 255 Monkeypox Skin Image Dataset [ 288 , 289 ] 2022 2D Dermoscopy Skin 40.2k Y es Cls Monkeypox 256 Vitiligo Images [ 290 ] 2019 2D Dermoscopy Skin 368 No NA Vitiligo 257 ImageCLEF 2016 [ 120 ] NA 2D Multi a Skin, Cell, Breast 31k Y es Cls NA Overall 2014 ∼ 2022 2D Multi Skin, Cell, Breast 197.6k NA Multi Multi a Multi-modalities of ImageCLEF 2016: MR, US, Histopathology , X-Ray , CT , PET , Endoscopy , Dermoscopy , EEG, ECG, EMG, Microscop y , Fundus Photography . Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Recon=Reconstruction, Reg=Registration, Loc=Localization, Est=Estimation, US=Ultrasound, EM=Electron Microscopy . 138 T able 15: 2D histopathology datasets. (part 1/2) # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 258 P ANDA_radboud [ 291 ] 2020 2D Histopathology (P atch) Prostate 5.1k Y es Seg Prostate Cancer 259 Gleason [ 292 ] 2019 2D Histopathology (Patch) Prostate 331 Y es Seg Prostate Cancer 260 PathologyVQA [ 293 ] 2020 2D Histopathology (P atch) Full Body 5.0k Y es VQA NA 261 SLN-Breast [ 294 ] 2019 2D Histopathology (WSI) L ymph 166 Y es Cls Breast Lymph Node 262 MoNuSeg [ 295 ] 2018 2D Histopathology (Patch) Nuclei 51 Y es Seg NA 263 MoNuSA C2020 [ 296 ] 2019 2D Histopathology (Patch) Lung, Prostate, etc. a 914 Y es Seg NA 264 DigestPath19 [ 297 ] 2019 2D Histopathology (WSI) Colon 212 Y es Det Signet Ring Cell 265 CAMEL YON17 [ 298 ] 2016 2D Histopathology (WSI) Breast 500 Y es Cls Breast Cancer 266 ANHIR [ 299 ] 2018 2D Histopathology (WSI) Kidney , Breast, etc. b 481 Y es Reg NA 267 Overlapping Cervical Cells [ 300 ] 2015 2D Histopathology (Patch) Cervix 17 Y es Seg Cervical Cells 268 MIDOG2022 [ 301 ] 2022 2D Histopathology (Patch) Lung, Breast, Skin 405 Y es Det Mitotic Figure 269 A CR OBA T [ 302 ] 2023 2D Histopathology (WSI) Breast 750 Y es Reg NA 270 BRIGHT [ 303 ] 2021 2D Histopathology (Patch) Breast 5.1k Y es Cls Pathological Benign 271 CoNIC2022 [ 304 ] 2022 2D Histopathology (Patch) Colon 5.0k Y es Seg Colon Nuclei 272 PanNuk e [ 305 ] 2021 2D Histopathology (WSI) Multi-organ 481 Y es Seg, Cls Multiple Cancers 273 Malignant L ymphoma Cls [ 306 ] 2021 2D Histopathology (Patch) Lymph 374 Y es Cls Lymphoma 274 P AIP2021 [ 307 ] 2021 2D Histopathology (WSI) Colon, Prostate 150 Y es Det Colon/Prostate Cancer 275 Breast Cancer Cell Seg [ 308 ] 2021 2D Histopathology (Patch) Breast 58 Y es Seg Breast Cancer 276 MedMNIST [ 309 ] 2020 2D Multi c Retina, Breast, Lung 100k Y es Cls Multi-disease 277 Histopathologic Cancer Det [ 310 ] 2018 2D Histopathology (Patch) L ymph 220k Y es Cls Breast Cancer 278 HuBMAP [ 311 ] 2020 2D Histopathology (Patch) Kidney 15 Y es Seg Kidney Tissue 279 A CDC-LungHP [ 312 ] 2019 2D Histopathology (WSI) Lung 200 Y es Seg Lung Cancer 280 SegPC 2021 [ 313 ] 2021 2D Histopathology (Patch) Blood 498 Y es Seg Plasma Cells 281 MIDOG2021 [ 301 ] 2021 2D Histopathology (Patch) Full Body 200 Y es Det Prostate Cancer 282 Dermofit Image Library [ 314 ] 2021 2D Histopathology (Patch) Skin 1.3k Y es Cls Lung Adenocarcinoma 283 W eakly Supervised Cell Seg [ 315 ] 2022 2D Histopathology (P atch) Full Body 30 Y es Seg Prostate Cancer 284 TIGER-wsibulk [ 316 ] 2022 2D Histopathology (WSI) Breast 93 Y es Seg Pneumothorax 285 BCI [ 317 ] 2022 2D Histopathology (Patch) Breast 4.9k Y es Gen Lesion 286 WSSS4LU AD [ 318 ] 2021 2D Histopathology (P atch) Lung 10.2k Y es Seg Coronary Artery 287 Breast Cancer Seg [ 319 ] 2019 2D Histopathology (Patch) Breast 151 Y es Seg Neurons 288 NuCLS [ 320 ] 2021 2D Histopathology (Patch) Nuclei 3.1k Y es Seg Kidney 289 ImageCLEF 2016 [ 144 ] 2015 2D Multi d Skin, Cell, Breast 31k Y es Cls Head & Neck Tumor 290 P AIP2020 [ 321 ] 2020 2D Histopathology (WSI) Liver 118 Y es Cls Colorectal Cancer 291 HER OHE [ 322 ] 2019 2D Histopathology (WSI) Lung 510 Y es Cls GI diseases 292 L ymphocyte Assessment [ 323 ] 2019 2D Histopathology (P atch) L ymphocyte 20k Y es Cls Lymphocyte Number 293 L YON19 [ 324 ] 2019 2D Histopathology (Patch) Lymphoc yte 441 Y es Cls L ymphocytes 294 GlaS [ 325 ] 2015 2D Histopathology (Patch) Cell 165 Y es Seg Colorectal Adenocarcinoma 295 CoNSeP [ 326 ] 2018 2D Histopathology (P atch) Colon 41 Y es Seg Colorectal Nuclei 296 PCam [ 40 ] 2018 2D Histopathology (Patch) Breast 328k Y es Seg Metastatic Tissue 297 LC25000 [ 38 ] 2019 2D Histopathology (Patch) Colon 25k Y es Cls Lung and Colon Tissue 298 PanNuk e (Seg) [ 305 ] 2021 2D Histopathology (Patch) Full Body 7.9k Y es Seg Nucleus 299 BreakHis (40x) [ 39 ] 2016 2D Histopathology (Patch) Breast 2.0k Y es Cls Breast Tumors 300 SICAPv2 [ 327 ] 2020 2D Histopathology (Patch) Prostate 18.8k Y es Cls Prostate Cancer 301 Kumar [ 295 ] 2018 2D Histopathology (Patch) Cell 54 Y es Seg Multi-organ Nuclei 302 HErle v [ 328 ] 2008 2D Histopathology (Patch) Cervix 5.6k Y es Cls Cervical Cancer 303 CRC100K [ 329 ] 2018 2D Histopathology (Patch) Colon 100k Y es Cls Colorectal Cancer 304 BRCA-M2C [ 330 ] 2021 2D Histopathology (Patch) Breast 120 Y es Seg Breast Cancer 305 warwick [ 325 ] 2015 2D Histopathology (Patch) Colon 330 Y es Seg Colorectal Gland 306 CRA G [ 331 ] 2019 2D Histopathology (Patch) Colon 213 Y es Seg Colorectal Cancer 307 Chaoyang [ 332 ] 2021 2D Histopathology (Patch) Blood 6.2k Y es Cls Red Blood Cell 308 CMB-CRC [ 122 ] 2022 3D, 2D Multi e Colon 472 No Seg, Cls Colorectal Cancer 309 CMB-GEC [ 123 ] 2022 3D, 2D CT , Histopathology (WSI), PET Brain 14 No Seg, Cls Melanoma 310 CMB-MEL [ 124 ] 2022 3D, 2D Multi f Brain 255 No Seg Melanoma 311 CMB-MML [ 125 ] 2021 2D, 3D Multi g NA 60 No Pred Multiple Myeloma 312 CMB-PCA [ 126 ] 2022 2D, 3D CT , MR, Histopathology (WSI) Prostate 31 No Cls, Pred Prostate Cancer 313 A GGC22 [ 333 ] 2022 2D Histopathology (Patch) Gland 150 Y es Seg Gland Segmentation 314 TUP AC [ 334 ] 2015 2D Histopathology (P atch) Brain 573 Y es Reg Breast Cancer 315 Prostate Fused-MRI-Pathology [ 147 ] 2016 2D, 3D MR, Histopathology (WSI) Prostate 29 No NA Prostate Cancer 316 Malaria Cell Image Dataset [ 335 ] 2021 2D Histopathology (Patch) Cell 27.6k Y es Cls Malaria 317 HEp-2 Cell Classification [ 336 ] 2020 2D Histopathology (P atch) Cell 13.6k Y es Cls HEp-2 Cells 318 Breast Cancer Cell Seg Dataset [ 337 ] 2020 2D Histopathology (Patch) Breast, Cell 58 Y es Seg Breast Cancer a Full structure of MoNuSA C2020: Lung (Thorax), Prostate (Pelvis), Kidney (Abdomen), Breast (Thorax). b Full structure of ANHIR: Kidney (Abdomen), Breast (Thorax), Colon (Abdomen), Spleen, Lung (Thorax). c Multi-modalities of MedMNIST : OCT , X-Ray, CT , Histopathology (Patch), Fundus Photography . d Multi-modalities of ImageCLEF 2016: MR, US, Histopathology (Patch), X-Ray , CT , PET , Endoscop y , Dermoscopy , Others. e Multi-modalities of CMB-CRC: CT , MR, US, DX, PET , Histopathology (WSI). f Multi-modalities of CMB-MEL: CT , US, Histopathology (WSI), PET . g Multi-modalities of CMB-MML: CT , MR, PET , Histopathology (WSI). Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Reg=Registration, VQA=V isual Question Answering, Gen=Generation, Pred=Prediction. 139 T able 16: 2D histopathology datasets. (part 2/2) # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 319 TIGER-wsirois [ 338 ] 2022 2D Histopathology (P atch) Breast 2.0k Y es Seg Breast Cancer 320 TIGER-wsitils [ 338 ] 2022 2D Histopathology (Patch) Breast 82 Y es Reg Breast Cancer 321 Breast Cancer Cell Seg 2 [ 337 ] 2020 2D Histopathology (Patch) Breast 58 Y es Seg Breast cancer 322 Malignant L ymphoma Cls Dataset [ 306 ] 2020 2D Histopathology (Patch) L ymph 374 Y es Cls Lymphoma 323 Lung and Colon Histopathology [ 38 ] 2020 2D Histopathology (Patch) Lung, Colon 25k Y es Cls Lung and Colon Cancer 324 FocusPath [ 339 ] 2020 2D Histopathology (Patch) N A 864 Y es IQA Histopathology Image 325 Blood Cell Images [ 340 ] 2019 2D Histopathology (Patch) Blood 12.5k Y es Det Blood Cell 326 Colorectal Histology MNIST [ 341 ] 2016 2D Histopathology (Patch) Colon 5.0k Y es Cls Colorectal Tissue 327 BreakHis 100x [ 39 ] 2016 2D Histopathology (Patch) Breast 9.1k Y es Cls Breast Cancer 328 BreakHis 200x [ 39 ] 2016 2D Histopathology (Patch) Breast 9.1k Y es Cls Breast Cancer 329 BreakHis 400x [ 39 ] 2016 2D Histopathology (Patch) Breast 9.1k Y es Cls Breast Cancer 330 BCNB T ask-1 [ 342 ] 2021 2D Histopathology (WSI) Breast 1.1k Y es Cls Leukemia 331 BCNB T ask-2 [ 342 ] 2021 2D Histopathology (WSI) Breast 1.1k Y es Cls Breast Cancer 332 BCNB T ask-3 [ 342 ] 2021 2D Histopathology (WSI) Breast 1.1k Y es Cls Breast Cancer 333 BCNB T ask-4 [ 342 ] 2021 2D Histopathology (WSI) Breast 1.1k Y es Cls Breast Cancer 334 BCNB T ask-5 [ 342 ] 2021 2D Histopathology (WSI) Breast 1.1k Y es Cls Breast Cancer 335 BCNB T ask-6 [ 342 ] 2021 2D Histopathology (WSI) Breast 1.1k Y es Cls Breast Cancer 336 P AND A [ 291 ] 2020 2D Histopathology (P atch) Prostate 10.6k Y es Cls Prostate Cancer 337 P AND A_karolinska [ 291 ] 2020 2D Histopathology (Patch) Prostate 5.5k Y es Seg Prostate Cancer 338 P AIP 2023 [ 343 ] 2022 2D Histopathology (Patch) Pancreas 103 Y es Seg Liver Cancer 339 A TEC23 [ 344 ] 2023 2D Histopathology (WSI) Ovary 468 Y es Cls Ovarian Cancer 340 ACR OBA T2023 [ 345 ] 2023 2D Histopathology (WSI) Breast 1.2k Y es Reg Breast Cancer 341 OCELOT2023 [ 346 ] 2023 2D Histopathology (WSI) Colon 667 Y es Det Colon Cancer 342 OCEAN [ 347 ] 2023 2D Histopathology (WSI) Ovary 1.6k Y es Cls Ovarian Cancer 343 Endo-Aid [ 348 ] 2022 2D Histopathology (WSI) GI Tract 91 No Cls GI Polyps 344 P AIP2023 [ 343 ] 2022 2D Histopathology (P atch) Pancreas 103 Y es Seg Pancreatic Cancer 345 PatchCamelyon[ 349 ] 2018 2D Histopathology (Patch) L ymph Node 295k Y es Cls Metastatic Tissue 346 Bone Marrow Cytomorphology[ 350 ] 2021 2D Histopathology (Patch) Bone Marrow 171k Y es Cls Blood Cells 347 Lung-Fused-CT -Pathology[ 132 ] 2018 2D, 3D CT , Histopathology (WSI) Lung 36 Y es Seg Lung Cancer 348 HNSCC-mIF-mIHC[ 351 ] 2020 2D Histopathology (P atch) Head & Neck 3.2k No NA HNSCC 349 SN-AM[ 352 ] 2019 2D Histopathology (Patch) L ymph Node 190 Y es Seg Melanoma 350 Ovarian Bev acizumab Response[ 353 ] 2023 2D Histopathology (WSI) Ovary 285 No NA Ovarian Cancer 351 CMB-LCA[ 354 ] 2022 2D, 3D Multi a Lung 0 No NA Lung Cancer 352 CPT A C-COAD[ 355 ] 2021 2D Histopathology (WSI) Colon 373 Y es Cls Colon Adenocarcinoma 353 Hungarian-Colorectal- Screening[ 356 ] 2022 2D Histopathology (WSI) Colorectal 200 No NA Colorectal Polyps 354 DLBCL-Morphology[ 357 ] 2022 2D Histopathology (P atch) Lymph Node 246 Y es Seg DLBCL 355 CPT A C-O V[ 358 ] 2021 2D Histopathology (WSI) Ovary 222 No NA Ovarian Cancer 356 CODEX imaging of HCC[ 359 ] 2023 2D Histopathology (WSI) Liver 646 No NA Liver HCC 357 Prostate-MRI[ 150 ] 2011 3D, 2D Multi b Prostate 26 No NA Prostate Cancer 358 CPT A C-BRCA[ 360 ] 2021 2D Histopathology (WSI) Breast 642 No NA Breast Cancer 359 AML-Cytomorphology_LMU[ 361 ] 2019 2D Histopathology (WSI) Blood 18.4k Y es Cls Acute Myeloid Leukemia 360 MiMM_SBILab [ 362 ] 2019 2D Histopathology (WSI) Bone Marrow 85 Y es Loc Multiple Myeloma 361 Pan-Cancer-Nuclei-Seg [ 363 ] 2020 2D Histopathology (WSI) Multi-organ 5.1k Y es Seg Pan-Cancer 362 TIL-WSI-TCGA [ 364 ] 2018 2D Histopathology (WSI) Multi-organ 5.2k Y es Cls Pan-Cancer 363 C-NMC 2019 [ 365 ] 2019 2D Histopathology (WSI) Blood 15.1k Y es Cls Leukemia 364 CPT A C-AML [ 366 ] 2019 2D Histopathology (WSI) Bone Marrow 122 No NA Acute Myeloid Leukemia 365 CA TCH [ 367 ] 2022 2D Histopathology (WSI) Skin 350 Y es Seg Skin Cancer 366 NADT -Prostate [ 368 ] 2021 2D Histopathology (WSI) Prostate 1.4k No NA Prostate Cancer 367 HER2 tumor ROIs [ 369 ] 2022 2D Histopathology (WSI) Breast 273 Y es Seg HER2+ Breast Cancer 368 CRC_FFPE- CODEX_CellNeighs [ 370 ] 2020 2D Histopathology (WSI) Colorectal 200 No NA Colorectal Cancer 369 Post-NA T -BRCA [ 371 ] 2019 2D Histopathology (WSI) Breast 96 Y es Cls Breast Cancer 370 Osteosarcoma T umor Assessment [ 372 ] 2019 2D Histopathology (WSI) Bone 1.1k Y es Cls Osteosarcoma 371 Quilt-1M [ 373 ] 2023 2D Histopathology (Patch) Multi-or gan 1m Y es VQA Multi-organ Pathology Overall 2008 ∼ 2023 2D Multi Full Body 2.6m NA Multi Multi a Multi-modalities of CMB-LCA: CT , MR, US, Histopathology (WSI), DX. b Multi-modalities of Prostate-MRI: MR, CT , PET , Histopathology (WSI). Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Reg=Registration, Loc=Localization, IQA=Image Quality Assessment, VQA=V isual Question Answering. 140 T able 17: 2D microscopy datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 372 CellT racking2019 [ 374 ] 2019 2D Microscopy Cell 1.4M Y es Tracking N A 373 CREMI [ 375 ] 2016 2D Microscopy Brain 375 Y es Seg N A 374 Bacteria Detection [ 376 ] 2021 2D Microscopy NA 366 Y es Seg N A 375 Blood Cell Images [ 340 ] 2021 2D Microscopy Blood 12.5k Y es Cls Blood 376 Leukemia Classification [ 377 ] 2021 2D Microscopy N A 15.1k Y es Cls Leukemia 377 CellT racking2021 [ 374 ] 2021 2D+3D+Video Microscopy Cell 0 Y es Tracking, Seg Lung Disease 378 B-ALL Classification [ 378 ] 2018 2D Microscopy Cell 15.1k Y es Cls Brain Tumor 379 2018 Data Science Bowl [ 379 ] 2018 2D Microscopy Nuclei 670 Y es Seg Skin Lesions 380 GSB2016 [ 157 ] 2015 2D Multi a Skin, Cell, Breast 31k Y es Cls Head & Neck T umor 381 OCCISC (SemSeg) [ 300 ] 2014 2D Microscopy Cell 945 Y es Seg Cervical Cytology 382 ICIAR 2018 (Microscopy) [ 380 ] 2017 2D Microscopy Breast 400 Y es Cls Breast Cancer 383 CBC (Counting) [ 381 ] 2019 2D Microscopy Full Body 420 Y es Reg NA 384 HuSHeM [ 382 ] 2017 2D Microscopy Pelvic 216 Y es Cls Sperm Head Morphology 385 Kaggle-HP A [ 383 ] 2021 2D Microscopy N A 89.5k Y es Seg Protein Localization 386 nanni2016texture [ 384 ] 2016 2D Microscopy Retina 195 Y es Cls Cell Shape 387 Corneal Endothelial Cell [ 385 ] 2019 2D Microscopy Retina 385 Y es Seg NA 388 Corneal Nerve [ 386 ] 2008 2D Microscopy Retina 90 Y es Cls Corneal Abnormalities 389 Corneal Nerve T ortuosity [ 387 ] 2011 2D Microscopy Retina 30 Y es Cls Nerve T ortuosity 390 Cervix93 Cytology [ 388 ] 2018 2D Microscopy Cervix 93 Y es Cls Cervical Cancer 391 DLBCL-Morph [ 389 ] 2020 2D Microscopy Retina 152.2k Y es Reg DLBCL Lymphoma 392 2-PM V essel Dataset [ 390 ] 2016 2D Microscopy V essel 12 Y es Seg NA 393 BBBC041 [ 391 ] 2012 2D Microscopy Cell 1.3k Y es Seg Malaria 394 FMD [ 392 ] 2019 2D Microscopy Surface 5.1k Y es Cls, Seg Surface Defect 395 Blood Cell Detection [ 393 ] 2022 2D Microscopy NA 874 Y es Det NA 396 T uberculosis Image [ 394 ] 2020 2D Microscopy NA 1.3k Y es Det T uberculosis 397 MHSMA [ 395 ] 2019 2D Microscopy N A 1.5k Y es Cls NA 398 ICIAR 2018 (Microscopy) [ 380 ] 2017 2D Microscopy , WSI NA 400 Y es Seg Breast Cancer 399 ImageCLEF 2016 [ 157 ] 2016 2D Multi a Skin, Cell, Breast 31k Y es Cls NA 400 CellT racking2024 [ 374 ] 2024 2D+3D+Video Microscopy Cell 0 Y es Tracking, Seg N A 401 CellT racking2022 [ 374 ] 2022 2D+3D+Video Microscopy Cell 0 Y es Tracking, Seg N A 402 CellT racking2023 [ 374 ] 2023 2D+3D+Video Microscopy Cell 0 Y es Tracking, Seg N A 403 OCCISC (InstSeg) [ 381 ] 2014 2D Microscopy Cell 945 Y es Seg N A 404 CBC (Detection) [ 394 ] 2019 2D Microscopy Full Body 420 Y es Det NA 405 ICIAR 2018 (WSI) [ 380 ] 2018 2D Microscopy , WSI NA 400 Y es Seg Breast Cancer Overall 2008 ∼ 2024 2D Multi Full Body 1.8m Y es Multi Multi a Multi-modalities of GSB2016 and ImageCLEF 2016: MR, US, Histopathology , X-Ray , CT , PET , Endoscopy , Dermoscopy , EEG, ECG, EMG, Microscopy , Electron Microscop y , Fundus Photography . Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Reg=Registration, T racking=T racking, WSI=Whole-Slide Images. T able 18: 2D infrared datasets. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 406 RA VIR [ 396 ] 2022 2D Infrared Retina 42 Y es Seg Blood vessel 407 MRL Eye Glasses cls [ 397 ] 2018 2D Infrared Retina 84.9k Y es Cls NA 408 MRL Eye Eye state cls [ 397 ] 2018 2D Infrared Retina 84.9k Y es Cls NA 409 MRL Eye Reflections cls [ 397 ] 2018 2D Infrared Retina 84.9k Y es Cls NA 410 MRL Eye Image quality cls [ 397 ] 2018 2D Infrared Retina 84.9k Y es Cls NA 411 MRL Eye Sensor type cls [ 397 ] 2018 2D Infrared Retina 84.9k Y es Cls NA Overall 2018 ∼ 2022 2D Infrared Retina 424.5k Y es Cls, Seg Blood vessel Abbre viations: Seg=Se gmentation, Cls=Classification. 141 T able 19: 2D endoscopy datasets. # Dataset Y ear Dim Modality Structure Images Label Task Diseases 412 Kavsir[ 398 ] 2017 2D Endoscopy Colon 14k Y es Cls NA 413 EndoSlam[ 399 ] 2021 2D Endoscopy Colon, Liver , Stomach, Kidney 76.8k Y es Recon, Est NA 414 SARAS-MESAD[ 88 ] 2021 2D Endoscopy Prostate 50.3k No Det GI disease 415 EAD19[ 87 ] 2018 2D Endoscopy Stomach, Bladder, Colon 2.1k Y es Det Endo Artifact 416 EndoCV2020-Sub Challenge1[ 400 ] 2019 2D Endoscopy Colon 2.3k Y es Det, Seg Polyp 417 EndoVis15[ 401 ] 2015 2D Endoscopy Colon 612 Y es Seg Polyp 418 Surgical tool detection challenge (m2cai16-tool)[ 402 ] 2016 2D Endoscopy Gallbladder 15 Y es Det NA 419 AIDA-E_1 [ 403 ] 2015 2D Endoscopy Stomach, Liver 181 Y es Cls Celiac Disease 420 AIDA-E_2 [ 404 ] 2015 2D Endoscopy Esophagus 157 Y es Cls Barrett’s Esophagus 421 AIDA-E_3 [ 405 ] 2015 2D Endoscopy Stomach, Colon 88 Y es Cls Metaplasia, Dysplasia 422 CVC-ClinicDB[ 406 ] 2021 2D Endoscopy Bowel 1.4k Y es Seg Polyp 423 Kvasir-SEG[ 407 ] 2020 2D Endoscopy Bowel 8k Y es Seg N A 424 FetReg[ 408 ] 2022 2D Endoscopy Uterus 2.7k Y es Seg Placental V asculature 425 SARAS-ESAD[ 88 ] 2020 2D Endoscopy Bowel 33.4k Y es Det Skin lesion 426 ImageCLEF 2016[ 120 ] 2015 2D Multi a Skin, Cell, Breast 31k Y es Cls H&N tumor 427 ISBI-AIDA-CECI 2015 2D Endoscopy Liv er, Stomach 181 Y es Cls Celiac diseases 428 SUN_SEG[ 409 ] 2022 2D+V ideo Endoscopy Colon 49.1k Y es Seg, Det, Cls Polyp 429 HyperKvasir[ 83 ] 2020 2D+Video Endoscopy Esophagus, Stomach, Colon 6.5k Y es Cls, Caption, Loc GI disease 430 Gastrointestinal Image ANAlysis (GIANA) [ 410 ] 2016 2D Endoscopy Colon 600 Y es Cls V ascular Malformation 431 EndoVis 2015 - D A GI [ 411 ] 2015 2D Endoscopy NA 389 Y es Det Cholecystectomy 432 EndoVis 2015 - EBCD [ 412 ] 2015 2D Endoscopy NA 150 Y es Seg Barrett’s Epithelium 433 EndoCV2020-Sub Challenge2[ 413 ] 2019 2D Endoscopy NA 386 Y es Det NA 434 EndoVis 2015 - APDCV[ 401 ] 2015 2D Endoscopy NA 612 Y es Seg Colonic Polyp 435 EndoVis 2015 - IST_2D-Endoscop y [ 414 ] 2015 2D+V ideo Endoscopy N A 100 Y es Seg Surgical Instruments 436 EndoVis 2018 - RSS[ 415 ] 2018 2D Endoscopy N A 2.8k Y es Seg Surgical Instruments 437 ISBI-AIDA-EMIBS 2015 2D Endoscopy NA 262 Y es Cls Gastric 438 ISBI-AIDA-GCICS 2015 2D Endoscopy NA 176 Y es Cls Gastric 439 EndoVis2023-SIMS [ 416 ] 2023 2D Endoscopy N A 0 Y es Seg Endoscopy 440 EndoVis2023-Syn-ISS [ 417 ] 2023 2D Endoscopy NA 0 Y es Seg NA 441 P2ILF [ 418 ] 2022 2D+3D Endoscopy NA 15 Y es Reg Multi-organ 442 EndoVis2023-Sur gRIPE[ 419 ] 2023 2D Endoscopy NA 0 Y es Est NA 443 m2caiSeg[ 420 ] 2020 2D Endoscopy Instrument 614 Y es Seg NA 444 CVC-EndoSceneStill[ 406 ] NA 2D Endoscopy NA 3.4k Y es Seg Polyp 445 Endo-FM[ 421 ] NA 2D+V ideo Endoscopy NA 0 Y es Seg, Cls, Det NA 446 SegSTRONG-C[ 422 ] NA 2D+Video Endoscopy NA 17 Y es Seg NA 447 SegCol[ 423 ] NA 2D+Video Colposcopy , Endoscopy NA 78 Y es Seg N A 448 FedSurg [ 424 ] 2024 2D+V ideo Endoscopy NA 30 Y es Cls Laparoscopic appendectomy Overall 2015 ∼ 2024 2D Multi Full Body 288.5k Y es Multi Multi a Multi-modalities of ImageCLEF 2016: MR, Ultrasound, Histopathology , X-Ray , CT , PET , Endoscopy , Dermoscopy , Others, EEG, ECG, EMG, Electron Microscop y , Fundus Photo graph y . Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Recon=Reconstruction, Reg=Registration, Loc=Localization, Est=Estimation, GI=Gastrointestinal, H&N=H ead & Neck. T able 20: 2D datasets of the other modalities. # Dataset Y ear Dim Modality Structure Images Label T ask Diseases 449 Intel & MobileODT Cervical Cancer Screening [ 425 ] 2017 2D Colposcopy V agina 1,993 Y es Cls Cervical cancer 450 ADDI ALZHEIMER’S DETECTION CHALLENGE [ 426 ] 2021 2D Series Data NA 34,614 Y es Cls Lung adenocarcinoma 451 The Digital Mammography DREAM Challenge [ 427 ] 2016 2D Mammography Breast 640,000 Y es Cls Atrophic AMD 452 BigNeuron [ 428 ] 2016 2D NA Brain 2,166 Y es Recon Diabetic foot ulcer 453 Human Activity Classification with Radar [ 429 ] 2019 2D Others NA 1,854 Y es Cls NA 454 KvasirCapsule-SEG [ 430 ] 2021 2D Colposcopy (Capsule) Polyp 55 Y es Seg Polyp 455 SCDB [ 431 ] 2020 2D Others Skin 4 Y es Cls Skin lesion 456 R OSE [ 432 ] NA 2D OCT A (CT) Eye a 229 Y es Seg NA 457 CMMD [ 433 ] 2021 2D Mammography Breast 1,775 Y es Seg, Cls Breast Cancer 458 Br35H [ 434 ] 2022 2D MR Brain 3,000 Y es Cls Brain Tumor 459 CDD-CESM [ 435 ] 2021 2D Mammography NA 2,006 Y es Seg, Cls Breast Cancer 460 OralCancer [ 436 ] 2020 2D Digital Photography Lip, T ongue 131 Y es Cls Oral Cancer 461 Oral_Diseases [ 437 ] 2023 2D Digital Photography NA 12,944 Y es Cls, Det Dental conditions 462 MRL Eye Gender [ 438 ] 2018 2D Infrared Reflectance Imaging Retina 84,898 Y es Cls NA 463 MITOS-A TYPIA-14 [ 439 ] 2013 2D Biopsy slides Breast 1,420 Y es Cls Breast Cancer 464 V inDr-Mammo [ 440 ] NA 2D Mammography Breast 19,992 Y es Det Breast Cancer 465 V inDr-SpineXR [ 441 ] NA 2D X-ray Spine 10,469 Y es NA Spinal Lesions 466 V inDr-PCXR [ 442 ] N A 2D X-ray Chest 9,125 Y es NA NA 467 V inDr-CXR [ 443 ] NA 2D X-ray Chest 18,000 Y es NA NA 468 HAM10000 [ 444 ] NA 2D Dermoscopy Skin 10,015 Y es Cls Skin Cancer 469 RFMiD [ 445 ] NA 2D Fundus Retina 3,200 Y es Cls Ophthalmic Diseases 470 Chaksu [ 446 ] N A 2D Fundus Retina 1,345 Y es Seg Glaucoma 471 PBC_dataset_normal_DIB [ 447 ] NA 2D Microscopy Blood 0 Y es Cls NA Overall 2013 ∼ 2023 2D Multi Full Body 859.2k Y es Multi Multi a Includes anatomical structures like optic nerve, e yeball, etc. Abbre viations: Seg=Se gmentation, Det=Detection, Cls=Classification, Recon=Reconstruction. 142 T able 21: 3D CT datasets # Dataset Y ear Dim Modality Structure V olumes Label T ask Diseases 472 T otalSe gmentator[ 67 ] 2022 3D CT Full Body 1204 Y es Seg V aried pathologies 473 AutoPET[ 448 ] 2022 3D CT/PET Whole-body 1014 Y es Seg Cancer 474 AutoPET II[ 449 ] 2023 3D CT/ PET Whole-body 1219 Y es Seg Cancer 475 ULS23[ 450 ] 2023 3D CT Chest, Ab- domen, Pelvis 39468 Y es Seg Oncological lesions 476 DeepLesion[ 451 ] 2018 2D/3D CT Bone, Ab- domen, Medi- astinum, Liver , Lung, Kidney ... 32735 Y es Det V arious lesions 477 CT -ORG[ 452 ] 2019 3D CT Bladder, Bone, Brain, Kidney , Liver , Lung 140 Y es Seg Liver lesions, Metastatic disease 478 SegTHOR[ 453 ] 2019 3D CT Heart, Aorta, Trachea, Esophagus 60 Y es Seg Lung cancer 479 AbdomenAtlas[ 454 ] 2024 3D CT Abdomen 20460 Y es Seg Multi-organ 480 CT -RA TE[ 36 ] 2024 3D CT Chest 50188 Y es Cls Chest abnormalities 481 M3D[ 66 ] 2024 3D CT/MR Whole-body 120000 Y es Seg/Rec/Reg/Loc Multiple pathologies 482 AMOS22[ 70 ] 2022 3D CT/MR Abdomen 600 Y es Seg Multi-disease 483 AMOS-MM[ 455 ] 2024 3D CT Abdomen 2300 Y es syn/Rec Abdominal diseases 484 FLARE21[ 456 ] 2021 3D CT Liver , Kid- ney , Spleen, Pancreas 511 Y es Seg Multi-organ 485 FLARE22[ 457 ] 2022 3D CT Abdomen 2300 Y es Seg Pan-cancer 486 FLARE23[ 458 ] 2023 3D CT Abdomen 4500 Y es Seg Pan-cancer 487 WORD[ 459 ] 2021 3D CT Abdomen 150 Y es Seg Cancer 488 RA OS[ 460 ] 2024 3D CT/MR Abdomen 4130 Y es Seg Abdominal cancers 489 SLIVER07[ 461 ] 2007 3D CT Liver 30 Y es Seg Liver 490 LiTS[ 68 ] 2017 3D CT Liver 201 Y es Seg Liver tumors 491 Pancreas-CT[ 462 ] 2016 3D CT Pancreas 80 Y es Seg Healthy controls 492 CHA OS CT[ 463 ] 2019 3D CT Liv er 40 Y es Seg Healthy subjects 493 CHA OS MRI[ 463 ] 2019 3D CT/MR Abdomen 160 Y es Seg Healthy 494 KiTS19[ 464 ] 2019 3D CT Kidneys 300 Y es Seg Kidney cancer 495 KiTS21[ 465 ] 2021 3D CT Kidneys 400 Y es Seg Kidney diseases 496 KiTS23[ 466 ] 2023 3D CT Kidneys 599 Y es Seg Kidney cancer 497 KiP A22[ 467 ] 2022 3D CT Kidney 130 Y es Seg Renal Cancer 498 AbdomenCT -1K[ 468 ] 2021 3D CT Abdomen 1062 Y es Seg Multi-disease cases 499 MSD03-Liver[ 469 ] 2018 3D CT Liver 210 Y es Seg Liver cancer 500 MSD06-Lung[ 469 ] 2018 3D CT Lung 96 Y es Seg Lung cancer 501 MSD07-Pancreas[ 469 ] 2018 3D CT Pancreas 420 Y es Seg Pancreatic masses 502 MSD08- HepaticV essel[ 469 ] 2018 3D CT Liver 443 Y es Seg Liver tumors 503 MSD09-Spleen[ 469 ] 2018 3D CT Spleen 61 Y es Seg 504 MSD10-Colon[ 469 ] 2018 3D CT Colon 190 Y es Seg Colorectal Cancer 505 EXACT09[ 470 ] 2009 3D CT Lung 40 Y es Seg Lung diseases 506 LOLA11[ 471 ] 2011 3D CT Chest 55 Y es Seg Lung abnormalities 507 LUNA16[ 472 ] 2016 3D CT Lung 888 Y es Det/Cls Lung cancer 508 A TM’22[ 473 ] 2022 3D CT Lung 500 Y es Seg Pulmonary diseases 509 AIIB23[ 474 ] 2023 3D CT Airway 312 Y es Seg Fibrotic lung disease 510 BIMCV CO VID- 19+[ 475 ] 2020 2D/ 3D CT/ X-RA Y Lung 2428 Y es Seg/ Cls/ Loc CO VID-19 pneumonia 511 CO VID-19-AR[ 476 ] 2020 2D/ 3D CT/ X-RA Y Chest 105 Y es Cls COVID-19 512 CT Images in CO VID- 19[ 477 ] 2020 3D CT Chest 771 No Cls CO VID-19 pneumonia 513 Chest CT Scans with CO VID-19[ 478 ] 2020 3D CT Chest 20 Y es Cls COVID-19 514 MIDRC-RICORD- 1a[ 479 ] 2020 3D CT Chest 120 Y es Seg/Cls CO VID-19 515 CO VID-19-20 Lung CT Lesion Segmentation Challenge[ 480 ] 2020 3D CT Lung 295 Y es Seg CO VID-19 516 CO VID-19-NY - SBU[ 481 ] 2021 2D/ 3D CT/ MR/ PET/ X-RA Y Brain/ Chest 1384 No Cls COVID-19 517 CO VID19-CT -1000[ 482 ] 2021 2D/ 3D CT Lung 1000 Y es Cls COVID-19 518 MIDRC-RICORD- 1B[ 483 ] 2021 3D CT Chest 120 Y es Cls COVID-19 ne gative 519 STOIC2021[ 484 ] 2021 3D CT Lung 10735 Y es Cls COVID-19 520 CO VID-19 CT scans[ 485 ] 2021 3D CT Lung 20 Y es Seg COVID-19 infection 521 CO V19-CT -DB[ 486 ] 2022 3D CT Chest 7750 Y es Cls CO VID-19 522 LNQ2023[ 487 ] 2023 3D CT Chest 513 Y es Seg L ymph node metas- tases 523 Parse2022[ 488 ] 2022 3D CT Pulmonary artery 200 Y es Seg Pulmonary hyperten- sion 524 LNDb[ 489 ] 2019 3D CT Lung 294 Y es Cls/Det/Seg Lung cancer 525 FUMPE[ 490 ] 2018 3D CT Lung 35 Y es Seg/Det Pulmonary embolism 526 InST ANCE2022[ 491 ] 2022 3D CT Brain 200 Y es Seg Intracranial Hemor- rhage 143 527 ISLES 2024[ 492 ] 2024 3D CT/MR Brain 250 Y es Seg Acute/subacute ischemic stroke 528 HaN-Seg[ 493 ] 2023 3D CT/MR Head and Neck 42 Y es Seg Head and Neck Cancer 529 SegRap2023[ 494 ] 2023 3D CT Head, Neck 400 Y es Seg Nasopharyngeal carcinoma 530 PDDCA[ 495 ] 2015 3D CT Brainstem, Mandible, Optic nerves, Chiasm, Paro... 48 Y es Seg Head and neck cancer 531 StructSeg2019 T ask 1[ 496 ] 2019 3D CT Head and Neck 60 Y es Seg Head and Neck Cancer 532 StructSeg2019 T ask 2[ 496 ] 2019 3D CT Head and Neck 60 Y es Seg Nasopharyngeal carcinoma 533 StructSeg2019 T ask 3[ 496 ] 2019 3D CT Chest 60 Y es Seg Lung cancer 534 StructSeg2019 T ask 4[ 496 ] 2019 3D CT Lung 60 Y es Seg Lung cancer 535 Learn2Reg Lung CT[ 497 ] 2020 3D CT Thorax 30 Y es Reg Respiratory motion 536 Learn2Reg NLST[ 69 ] 2022 3D CT Thorax 420 Y es Reg Lung cancer 537 Learn2Reg Abdomen CT -CT[ 497 ] 2020 3D CT Abdomen 50 Y es Re g Multi-organ 538 Learn2Reg Abdomen MR-CT[ 497 ] 2021 3D CT/MRI Abdomen 122 Y es Reg Multi-modal 539 Continuous Registration[ 498 ] 2019 3D CT/MR Lung/Brain 142 Y es Reg Respiratory mo- tion/COPD 540 Continuous Registration DIRLAB[ 499 ] 2018 3D CT Lungs 20 Y es Reg/lmk Respiratory-induced motion 541 Continuous Registration EMPIRE[ 500 ] 2010 3D CT Lungs 60 Y es Re g Lung conditions 542 HECKTOR 2020[ 501 ] 2020 3D CT/ PET Head and Neck 254 Y es Seg Head and Neck Cancer 543 HECKTOR 2021[ 502 ] 2021 3D CT/ PET Head and Neck 325 Y es Seg/ Reg Head and Neck Cancer 544 HECKTOR 2022[ 503 ] 2022 3D CT/ PET Head and Neck 883 Y es Seg/ Reg Head and Neck Cancer 545 V erSe19[ 504 ] 2019 3D CT Spine 160 Y es Seg/Lab Spine diseases 546 V erSe20[ 505 ] 2020 3D CT Spine 300 Y es Seg/Lab Spine diseases 547 CTSpine1K[ 506 ] 2021 3D CT Spine 1005 Y es Seg Spine diseases 548 CTPelvic1K[ 507 ] 2020 3D CT Pelvic 1184 Y es Seg Pelvic bone conditions 549 RibFrac2020[ 508 ] 2020 3D CT Ribs 660 Y es Seg/Cls/Det Rib fractures 550 BTCV Abdomen[ 509 ] 2015 3D CT Abdomen 50 Y es Seg 551 BTCV Cervical[ 509 ] 2015 3D CT Cervix 50 Y es Seg Cervical cancer 552 ACRIN-HNSCC- FDG-PET -CT (ACRIN 6685)[ 510 ] 2016 3D CT/MR/NM/PET Head and Neck 260 Y es Cls Head and Neck Cancer 553 ACRIN-DSC-MR-Brain (ACRIN 6677)[ 511 ] 2019 2D/3D CT/MR Brain 123 Y es Cls Glioblastoma 554 ACRIN-FL T -Breast (ACRIN 6688)[ 512 ] 2017 3D CT/ PET Breast 83 Y es Cls Breast Cancer 555 ACRIN-FMISO-Brain (ACRIN 6684)[ 513 ] 2016 3D CT/MR/ PET Brain 45 Y es Se g/Cls Glioblastoma 556 ACRIN-NSCLC-FDG- PET (ACRIN 6668)[ 511 ] 2020 3D CT/ PET Lung 242 Y es Cls Lung cancer 557 AREN0532[ 514 ] 2022 2D/3D CR/ CT/ MR/ PET/ R TIMA GE/ US Kidney 544 No Cls Wilms tumor 558 AREN0532-T umor- Annotations[ 515 ] 2023 3D CT/MR Kidney 543 Y es Seg Wilms’ T umor 559 AREN0533[ 516 ] 2022 3D CR/ CT/ MR/ US Kidney 294 No Cls Wilms tumor 560 AREN0533-T umor- Annotations[ 517 ] 2023 3D CR/ CT/ MR/ US Kidney 294 Y es Seg Wilms tumor 561 AREN0534[ 518 ] 2021 2D/ 3D CT/ MR/ PET/ US Kidney 239 Y es Seg Wilms tumor 562 AHOD0831[ 519 ] 2022 2D/3D CR/ CT/ DX/ MR/ NM/ OT/ PET/ SC/ XA L ymphatic 165 Y es Seg Hodgkin Lymphoma 563 AHOD0831-T umor- Annotations[ 520 ] 2023 3D CT/ PET L ymph nodes, spleen, sali- vary glands, W alde yer’... 165 Y es Seg Hodgkin Lymphoma 564 HNSCC-3DCT -RT[ 521 ] 2018 3D CT Head and Neck 31 Y es Seg Head and Neck Cancer 565 HNSCC[ 522 ] 2020 3D CT/MR/PET Head and Neck 627 Y es Seg Head and Neck Cancer 566 CT COLONOGRAPHY[ 523 ] 2015 3D CT Colon 825 Y es Cls Colon Cancer 567 CT -Lymph-Nodes[ 524 ] 2015 2D/3D CT Mediastinum / Abdomen 176 Y es Det/Seg Lymphadenopathy 568 CC-Radiomics- Phantom[ 525 ] 2017 3D CT Phantom 17 No Cls Lung cancer 569 CC-Radiomics-Phantom- 2[ 526 ] 2019 3D CT Phantom 251 No - Phantom study 570 CC-Radiomics-Phantom- 3[ 527 ] 2019 3D CT Head/Chest 275 Y es Seg 144 571 CC-T umor Heterogeneity[ 528 ] 2023 3D CT/MR/ PET Cervix 23 Y es Seg/Cls Cervical cancer 572 TCGA-BLCA[ 529 ] 2014 3D CT/ MR/ PET/ X-RA Y Bladder 120 No Cls Bladder carcinoma 573 TCGA-COAD[ 530 ] 2014 3D CT Colon 25 No Cls Colon cancer 574 TCGA-ESCA[ 531 ] 2014 3D CT Esophagus 16 No Cls Esophageal carcinoma 575 TCGA-GBM[ 532 ] 2014 3D CT/ DX/ MR Brain 575 No Cls Glioblastoma Multi- forme 576 TCGA-HNSC[ 533 ] 2014 3D CT/ MR/ PET Head and Neck 479 No Cls Head and Neck Cancer 577 TCGA-KICH[ 534 ] 2016 3D CT/MR Kidney 15 No Cls Kidney cancer 578 TCGA-KIRC[ 535 ] 2016 3D CR/CT/MR Kidney 2654 Y es Cls Kidney cancer 579 TCGA-KIRC- Radiogenomics[ 536 ] 2015 3D CT/MR Kidney 103 Y es Cls Renal cell carcinoma 580 TCGA-KIRP[ 534 ] 2014 3D CT/ MR/ PET Kidney 33 No Cls Kidney cancer 581 TCGA-LGG[ 537 ] 2014 3D CT/MR Brain 199 Y es Cls Lower Grade Glioma 582 TCGA-LIHC[ 538 ] 2014 2D/3D CT/MR/ PET Liver 97 No Cls Liv er cancer 583 TCGA-LUAD[ 539 ] 2016 3D CT/ NM/ PT Lung 69 No Cls Lung cancer 584 TCGA-LUSC[ 540 ] 2016 3D CT/ PET Lung 37 No Cls Lung cancer 585 TCGA-O V[ 541 ] 2016 3D CT/ MR/ OT Ovary 143 No Cls Ovarian cancer 586 TCGA-O V - Radiogenomics[ 542 ] 2016 3D CT Ovary 93 Y es Cls Ovarian cancer 587 TCGA-O V - Proteogenomics[ 543 ] 2020 3D CT Ovary 20 No Seg Ovarian cancer 588 TCGA-PRAD[ 533 ] 2015 3D CT/MR/PET Prostate 14 No Cls Prostate cancer 589 TCGA-READ[ 544 ] 2014 3D CT/MR Rectum 3 No Cls Rectum adenocarci- noma 590 TCGA-SARC[ 545 ] 2014 3D CT/MR Chest/ Ab- domen/ Leg/ Spine 33 No Cls Sarcoma 591 TCGA-UCEC[ 538 ] 2020 3D CT/ MR/ PET/ X-RA Y Uterus 65 No Cls Uterine cancer 592 CPT AC-CCRCC[ 127 ] 2018 2D/ 3D CT/ MI- CROSCOPY/ MR Kidney 262 Y es Cls Renal cell carcinoma 593 CPT AC-CM[ 127 ] 2018 2D/ 3D CT/ MR/ PT Skin 95 No Cls/ Seg Cutaneous Melanoma 594 CPT AC-GBM[ 127 ] 2018 2D/ 3D CT/ MI- CROSCOPY/ MR Brain 200 No Cls Glioblastoma 595 CPT AC-HNSCC[ 127 ] 2018 3D CT/MR/WSI Head and Neck 207 Y es Cls Head and Neck Cancer 596 CPT AC-LSCC[ 127 ] 2018 2D/3D CT/ MI- CROSCOPY/ PET Lung 212 No Cls Lung cancer 597 CPT AC-LUAD[ 546 ] 2018 2D/ 3D CT/ MR/ PET Lung 244 No Cls Lung cancer 598 CPT AC-PDA[ 547 ] 2018 3D CT/ MR/ PET/ US Pancreas 168 No Cls Pancreatic cancer 599 CPT AC-SAR[ 548 ] 2019 2D/ 3D CT/ MR/ PET/ US Abdomen/ Arm/ Bladder/ Chest/ Head–Neck/ Kidney/ Leg/... 88 No Cls Sarcomas 600 CPT AC-UCEC[ 548 ] 2019 2D/ 3D CT/ MR/ PET/ US Uterus 250 No Cls Endometrial Carci- noma 601 NSCLC-Radiomics[ 549 ] 2014 3D CT Lung 422 Y es Se g/Cls Lung cancer 602 NSCLC-Radiomics- Genomics[ 549 ] 2014 3D CT Lung 89 Y es Cls Lung cancer 603 NSCLC- Radiogenomics[ 550 ] 2015 3D CT/ PET Chest 211 Y es Seg/ Cls Lung cancer 604 NSCLC-Cetuximab (R TOG-0617)[ 551 ] 2018 3D CT Chest 490 No Cls Lung cancer 605 NSCLC-Radiomics- Interobserver1[ 552 ] 2019 3D CT Lung 22 Y es Seg Lung cancer 606 CMB-CRC[ 553 ] 2022 2D/3D CT/ MR/ PET/ US/ WSI Colon 12 No Cls Colorectal Cancer 607 CMB-GEC[ 553 ] 2022 2D/3D CT/ MI- CROSCOPY/ MR/ PET Esophagus 17 No Seg/Cls Gastroesophageal Cancer 608 CMB-LCA[ 553 ] 2022 2D/3D CT/ DX/ MR/ NM/ PT/ US Lung 16 No Cls Lung cancer 609 CMB-MEL[ 553 ] 2022 2D/ 3D CT/ PET/ US/ WSI Skin 40 No Cls Melanoma 610 CMB-MML[ 553 ] 2022 2D/3D CR/ CT/ DX/ HISTOP A THOL- OGY/ MR/ PET/ XA Blood/Bone 138 No Cls Multiple Myeloma 611 CMB-PCA[ 553 ] 2022 2D/3D CT/ DX/ MR/ NM/ PET/ RF Prostate 50 No Cls Prostate cancer 612 Crowds-Cure-2017[ 554 ] 2017 3D CT Lung, Kidney , Liver , Ov ary 352 Y es Seg/Det Cancer 613 Crowds-Cure-2018[ 555 ] 2019 3D CT Bladder/ Brain/ Colon/ Head- Neck/ Kidney/ Lung/ Pancr 324 Y es Det/Reg Metastatic diseases 614 QIN-Breast[ 556 ] 2015 3D CT/ MR/ PET Breast 68 Y es Cls Breast cancer 145 615 QIN-HEADNECK[ 557 ] 2015 3D CT/ PET Head and Neck 279 Y es Seg Head and neck carcino- mas 616 QIN LUNG CT[ 558 ] 2015 3D CT Lung 47 No Cls Lung cancer 617 QIN-LungCT -Seg[ 559 ] 2015 3D CT Chest 31 Y es Seg Lung cancer 618 QIBA CT -1C[ 560 ] 2011 3D CT Phantom 1 Y es Seg 619 QIBA-V olCT -1B[ 561 ] 2020 3D CT Lung 40 Y es Seg Lung cancer 620 QIBA-CT -Liver - Phantom[ 562 ] 2021 3D CT Liver 684 No Se g/Det Liver pathology 621 Multi-Modality V ertebra Recognition[ 563 ] 2015 2D/3D CT/MR Spine 0 Y es Loc/Reg/Rec Spine diseases 622 xV ertSeg[ 564 ] 2016 3D CT Lumbar verte- brae 25 Y es Seg/Cls V ertebral fractures 623 Computational Methods and Clinical Applications for Spine Imaging[ 565 ] 2015 3D CT Spine 312 Y es Seg/Loc Spinal diseases 624 ACNS0332[ 566 ] 2021 3D CT/MR Brain/Spine 85 Y es Seg Brain cancer 625 AHEP0731[ 567 ] 2021 3D CT/ MR/ PET/ US/ XA Liver/ Chest 190 No Seg/ Cls Liver Cancer 626 Anti-PD-1 Lung[ 568 ] 2019 3D CT/ PET/ SC Lung 46 No Cls Lung cancer 627 Anti-PD-1 Immunother- apy Melanoma[ 569 ] 2019 3D CT/ MR/ PT Skin 47 No Cls Melanoma 628 BREAST - DIAGNOSIS[ 570 ] 2011 2D/ 3D CT/ MG/ MR/ PT Breast 88 Y es Cls Breast cancer 629 CALGB50303[ 571 ] 2021 3D CT/ PET Chest/ Ab- domen/ Pelvis 155 Y es Cls Diffuse Lar ge B-Cell L ymphoma 630 ELCAP Public Lung Image Database[ 572 ] 2003 3D CT Lung 50 Y es Det Lung nodules 631 GLIS-R T[ 573 ] 2021 2D/3D CT/MR Brain 230 Y es Seg/Reg Brain tumors 632 HEAR-EU[ 574 ] 2017 3D CT Cochlea 80 Y es Seg Sensorineural hearing loss 633 Head-Neck Cetuximab (R TOG 0522)[ 575 ] 2013 3D CT/ PET Head and Neck 111 No Cls Head and Neck Carcinomas 634 Head-Neck-PET -CT[ 576 ] 2017 3D CT/ PET Head and Neck 298 Y es Seg/ Cls Head and Neck Cancer 635 Head-Neck-Radiomics- HN1[ 549 ] 2019 3D CT/ PET Head/ Neck 137 Y es Seg Head and Neck Cancer 636 LCTSC[ 577 ] 2017 3D CT Lung 60 Y es Seg Lung cancer 637 LDCT -and-Projection- data[ 578 ] 2020 3D CT Head, Chest, Abdomen 299 Y es Rec V arious pathologies 638 LIDC-IDRI[ 579 ] 2011 3D CT Lung 1018 Y es Seg/Det Lung cancer 639 Lung-PET -CT -Dx[ 580 ] 2020 3D CT/ PET Lung 355 Y es Cls/ Det Lung cancer 640 LungCT -Diagnosis[ 581 ] 2014 3D CT Lung 61 Y es Cls Lung cancer 641 National Lung Screening Trial (NLST)[ 69 ] 2013 3D CT/ MI- CROSCOPY Chest 26254 Y es Cls Lung cancer 642 OPC-Radiomics[ 582 ] 2020 3D CT Head and Neck 606 Y es Seg/Cls Oropharyngeal Carci- noma 643 Parkinson’s Progression Markers Initiativ e (PPMI)[ 583 ] 2010 3D MR/ PET/ SPECT Brain 683 Y es Cls Parkinson’s Disease 644 Pediatric-CT -SEG[ 584 ] 2021 3D CT Chest/ Ab- domen/ Pelvis 359 Y es Seg Non-cancer pediatric exams 645 Public Lung Database T o Address Drug Response[ 572 ] 2009 3D CT Lung 93 Y es Seg/Det Lung cancer 646 QIDW[ 585 ] 2015 3D CT/ MR/ PET/ US Lung 52000 No Cls Quality assurance 647 RIDER Lung PET - CT[ 134 ] 2015 3D CT/ PET Lung 243 No Cls Lung cancer 648 SMIR Full Body CT[ 586 ] 2013 3D CT Skeleton 50 No Seg - 649 SPIE-AAPM Lung CT Challenge[ 587 ] 2015 3D CT Lung 70 Y es Cls Lung cancer 650 V estibular-Schwannoma- SEG[ 588 ] 2021 3D MR/ R T - DOSE/ R TPLAN/ R TSTR UCT Brain 242 Y es Seg V estibular schwan- noma 651 MM-WHS[ 589 ] 2017 3D CT/MR Heart 120 Y es Seg Cardiac conditions 652 Soft-tissue-Sarcoma[ 590 ] 2015 3D CT/ MR/ PET Extremities 51 Y es Se g/Cls Soft-tissue sarcoma 653 Seg Soft T issue[ 591 ] 2021 3D CT/MR/PET Soft tissue 51 Y es Seg Soft-tissue sarcomas (preprocessed) 654 Left Atrial W all Thick- ness Challenge[ 592 ] 2016 3D CT/MR Heart 20 Y es Seg Atrial Fibrillation 655 MELA22[ 593 ] 2022 3D CT Mediastinum 1100 Y es Det Mediastinal lesions 656 Head and Neck Auto Segmentation Challenge[ 495 ] 2015 3D CT Brainstem, Mandible, Chiasm, Optic Nerves, Paro... 48 Y es Seg Head and Neck Neoplasms 657 RSNA STR Pulmonary Embolism Detection[ 594 ] 2020 3D CT Lung 12195 Y es Cls/Det Pulmonary embolism 658 KNIGHT[ 595 ] 2021 3D CT Kidney 400 Y es Cls Renal cancer 659 Data Science Bowl 2017[ 596 ] 2017 3D CT Lungs 2101 Y es Cls Lung cancer 660 OSIC Pulmonary Fibrosis Progression[ 597 ] 2020 3D CT Lung 200 Y es Reg Pulmonary fibrosis 146 661 The V isible Human Project[ 598 ] 1994 2D/3D CT/MR Full Body 2 No Atlas Anatomical reference 662 ABCs[ 599 ] 2020 3D CT/MR Brain 75 Y es Seg Glioma 663 MA TCH[ 600 ] 2021 2D/3D CT/X-RA Y Lung 9 Y es Loc/Trk Lung cancer 664 CTVIE19[ 601 ] 2019 3D CT Lung 64 Y es Seg Lung cancer 665 ImageCLEFtuberculosis 2018[ 602 ] 2018 3D CT Chest 1513 Y es Cls Tuberculosis 666 ImageCLEF 2017 T uberculosis[ 603 ] 2017 3D CT Chest 944 Y es Cls Tuberculosis 667 fastPET -LD[ 604 ] 2021 3D CT/ PET Whole Body 68 Y es Det Oncologic Imaging 668 Low Dose CT[ 605 ] 2016 3D CT Liver 30 Y es Det Liv er lesions 669 LUMIC[ 606 ] 2018 3D CT Lung 13 Y es Reg/Seg Pulmonary perfusion defects 670 CAD-PE[ 607 ] 2013 3D CT Liver 91 Y es Seg Pulmonary Embolism 671 ANODE09[ 608 ] 2009 3D CT Lung 55 Y es Det Lung cancer 672 VESSEL12[ 609 ] 2012 3D CT Lung 20 Y es Seg Respiratory diseases 673 RIDER Lung CT[ 610 ] 2015 3D CT Lung 32 Y es Seg Lung cancer 674 OpenKBP[ 611 ] 2020 3D CT Head and Neck 340 Y es Reg Head and Neck Cancer 675 CT -vs-PET -V entilation- Imaging[ 612 ] 2022 3D CT/ PET Lung 20 No Cls Lung cancer 676 HCC-T ACE-Seg[ 613 ] 2021 3D CT Liver 105 Y es Seg Liver cancer 677 MOOD 2022- abdominal[ 614 ] 2022 3D CT Abdomen 550 Y es Seg/Cls Anomalies 678 StageII-Colorectal- CT[ 615 ] 2022 3D CT Abdomen 230 No Cls Colorectal cancer 679 Pseudo-PHI-DICOM- Data[ 616 ] 2021 2D/3D CT/MR/PET/X- RA Y V arious 21 No Reg V arious cancers 680 4D-Lung[ 617 ] 2016 3D CT Lung 20 Y es Seg Lung cancer 681 Lung Phantom[ 618 ] 2015 3D CT Thorax 1 No Seg Synthetic lung nodules 682 NaF PROST A TE[ 619 ] 2013 3D CT/ PET Prostate 9 No Cls Prostate cancer 683 EndoV is 2022 - P2ILF 3D-CT[ 620 ] 2022 3D CT/ENDOSCOPY Liver 15 Y es Seg Liver T umor 684 Finding and Measuring Lungs in CT Data[ 621 ] 2019 2D/3D CT Lungs 538 Y es Seg Lung diseases 685 SynthRAD2023[ 622 ] 2023 3D CT/MR Brain, Pelvis 1080 Y es Syn Oncological 686 IACT A-EST2023[ 623 ] 2023 3D CT Brain 402 Y es Cls Acute Ischemic Stroke 687 ISBI2023 challenge - APIS[ 624 ] 2023 3D CT Brain 96 Y es Seg 688 A UTO-R TP[ 625 ] 2022 3D CT Prostate, Sem- inal V esicles, Pelvic L ymph Nodes 10 Y es Seg Prostate cancer 689 Subsolid Nodules[ 626 ] 2025 3D CT Lung 59 Y es Seg Lung cancer 690 DICOM-LIDC-IDRI- Nodules[ 627 ] 2018 3D CT Lung 875 Y es Seg Lung cancer 691 Burdenko-GBM- Progression[ 628 ] 2023 3D CT/MR Brain 180 Y es Seg/Cls Glioblastoma 692 PDMR-833975-119- R[ 629 ] 2020 3D CT/MR/PET Pancreas 20 No Cls Pancreatic adenocarci- noma 693 APOLLO-5-LSCC[ 630 ] 2021 3D CT/PET Lung 36 Y es Seg Lung squamous cell carcinoma 694 APOLLO-5-LUAD[ 631 ] 2021 3D CT Lung 5 Y es Seg Lung adenocarcinoma 695 APOLLO-5-ESCA[ 632 ] 2021 3D CT Esophagus 4 Y es Seg Esophageal squamous cell carcinoma 696 APOLLO-5-P AAD[ 633 ] 2021 3D CT Pancreas 1 Y es Seg Pancreatic adenocarci- noma 697 APOLLO-5-THYM[ 634 ] 2021 3D CT Thymus 4 Y es Seg Thymoma 698 Lung-Fused-CT - Pathology[ 635 ] 2018 2D/3D CT/MICROSCOPY Lung 6 Y es Seg Lung cancer 699 LUAD-CT -Surviv al[ 636 ] 2017 3D CT Lung (Thorax) 40 Y es Seg Lung Adenocarcinoma 700 ARAR0331[ 637 ] 2022 3D CT/MR/ PET Head 108 Y es Seg Nasopharyngeal cancer 701 GammaKnife- Hippocampal[ 638 ] 2022 3D CT/ MR/ R TSTR UCT Hippocampus 390 Y es Seg Brain conditions 702 A091105[ 639 ] 2023 3D CT/MR Soft tissue 83 No Seg Desmoid tumors 703 Colorectal-Liver - Metastases[ 640 ] 2023 3D CT/SEG Liv er 197 Y es Seg Colorectal cancer liver metastases 704 Head-Neck-CT - Atlas[ 641 ] 2017 3D CT/ MR/ PET Head/Neck 215 Y es Seg Head and neck cancer 705 NRG-1308 (R TOG 1308)[ 642 ] 2015 3D CT Lung 12 Y es Seg Lung cancer 706 Phantom FDA[ 643 ] 2015 3D CT Lungs 7 No Reg Lung cancer 707 CTpred-Sunitinib- panNET[ 644 ] 2022 3D CT Pancreas 38 Y es Cls Pancreatic neuroen- docrine tumors 708 CALGB50303-T umor- Annotations[ 520 ] 2023 3D CT/ PET L ymphatic system 155 Y es Seg/ Cls Diffuse Lar ge B-Cell L ymphoma 709 RIDER-LungCT - Seg[ 645 ] 2020 3D CT Lung 31 Y es Seg Lung cancer 710 Radiomic-Feature- Standards[ 646 ] 2020 3D CT Chest 13 Y es Seg Lung cancer 711 PleThora[ 647 ] 2020 3D CT Thoracic 402 Y es Seg Lung cancer 712 MRI-DIR[ 648 ] 2018 3D CT/MR Head-Neck 9 No Reg Head and Neck Cancer 147 713 Ivy Glioblastoma Atlas Project (IvyGAP)[ 649 ] 2016 3D CT/MR Brain 39 Y es Se g/Cls Glioblastoma 714 RIDER Pilot[ 650 ] 2023 2D/3D CR/CT/DX Lung 8 No Cls Lung cancer 715 RADCURE[ 651 ] 2024 3D CT Head and Neck 3346 Y es Seg Head and Neck Cancer 716 Adrenal-ACC-Ki67- Seg[ 652 ] 2023 3D CT Adrenal 53 Y es Seg Adrenocortical carci- noma 717 Prostate-Anatomical- Edge-Cases[ 653 ] 2023 3D CT Prostate, Rectum, Bladder, Femoral Heads 131 Y es Seg Prostate Cancer 718 3D-IRCADb[ 65 ] 2010 3D CT Liver 20 Y es Seg Liv er tumors 719 SEG.A. 2023[ 654 ] 2023 3D CT Aorta 56 Y es Seg Aortic diseases 720 P2ILF[ 655 ] 2022 2D/3D CT/MR Liver 186 Y es Seg/Reg Liver cancer 721 SynthStrip[ 656 ] 2022 2D/3D CT/MR/PET Brain 622 Y es Seg Glioblastoma 722 MOOD[ 657 ] 2024 3D CT/MR Brain/Abdomen 1358 Y es Det/Loc Incidental pathologies 723 PENGWIN2024- T ask1[ 658 ] 2024 3D CT Pelvis 150 Y es Seg Pelvic fractures 724 TriALS2024-T ask1[ 659 ] 2024 3D CT Liver 201 Y es Seg Liver tumors 725 TriALS2024-T ask2[ 660 ] 2024 3D CT Liver 240 Y es Seg Hepatocellular carci- noma 726 National Lung Screening Trial (NLST) 2D- Pathology[ 69 ] 2013 2D/3D CT/ MI- CROSCOPY Chest 451 Y es Cls Lung cancer 727 QUBIQ2021 3D CT[ 661 ] 2021 3D CT Pancreas 118 Y es Seg Pancreatic lesions 728 orCaScore[ 662 ] 2014 3D CT Heart 72 Y es Det/Seg Cardiovascular dis- eases 729 INSPECT[ 663 ] 2023 3D CT Lungs 23248 Y es Cls Pulmonary embolism 730 BIMCV -R[ 664 ] 2024 2D/3D CT Thoracic 8069 Y es Rec Multiple diseases 731 Mandibular-CT - Dataset[ 665 ] 2018 3D CT Mandible 10 Y es Seg 732 Semi-T eethSeg[ 666 ] 2023 2D/3D CT/X-RA Y T eeth 38000 Y es Seg Dental issues T otal: 516,087+ T able 22: 3D MR datasets # Dataset Y ear Dim Modality Structure V olumes Label T ask Diseases 733 T otalSe gmentator MRI[ 667 ] 2024 3D MR Whole- body 616 Y es Seg V arious pathologies 734 ACDC[ 668 ] 2017 3D MR Heart 150 Y es Seg/ Cls Cardiac conditions 735 M&Ms[ 669 ] 2020 3D MR Heart 375 Y es Seg Cardiac diseases 736 M&Ms-2[ 670 ] 2021 3D MR Heart 360 Y es Seg Cardiac conditions 737 LAScarQS 2022[ 671 ] 2022 3D MR Heart 194 Y es Seg/Quan Left atrial scar quantifi- cation and segmenta- tion 738 LAScarQS++ 2024[ 672 ] 2024 3D MR Heart 200+ Y es Seg/Quan Multi-center left atrial and scar segmentation 739 MyoPS2020[ 673 ] 2020 3D MR Heart 45 Y es Seg Myocardial infarction 740 MyoPS++ 2024[ 674 ] 2024 3D MR Heart 200+ Y es Seg Multi-center myocar- dial pathology 741 MM-WHS[ 589 ] 2017 3D CT/MR Heart 120 Y es Seg Cardiac conditions 742 WHS++ 2024[ 675 ] 2024 3D CT/MR Heart 200+ Y es Seg Multi-center whole heart segmentation 743 CuRIOUS2018-MR FLAIR[ 676 ] 2018 3D MR/ US Brain 33 Y es Reg Brain tumors 744 CuRIOUS2018-US[ 77 ] 2018 3D MR/ US Brain 32 Y es Reg Brain tumors 745 CuRIOUS2018-MR T1W[ 77 ] 2018 3D MR/ US Brain 33 Y es Reg Brain tumor 746 CuRIOUS2019-MR- FLAIR[ 677 ] 2019 3D MR/ US Brain 32 Y es Reg Low-grade gliomas 747 CuRIOUS2019[ 77 ] 2019 3D MR/ US Brain 33 Y es Reg Brain tumor 748 CuRIOUS2019 US[ 677 ] 2019 3D MR/ US Brain 33 Y es Reg Low-grade gliomas 749 CuRIOUS2019-MR T1W[ 677 ] 2019 3D MR/ US Brain 33 Y es Reg Brain tumor 750 CuRIOUS2022[ 678 ] 2022 3D MR/ US Brain 33 Y es Seg Low-grade gliomas 751 CrossMoDA2021[ 679 ] 2021 3D MR Brain 347 Y es Seg V estibular Schwan- noma 752 CrossMoDA2022[ 679 ] 2022 3D MR Brain 347 Y es Seg/ Cls V estibular Schwan- noma 753 CrossMoDA2023[ 679 ] 2023 3D MR Brain 544 Y es Seg V estibular Schwan- noma 754 OASIS-1[ 680 ] 2007 3D MR Brain 416 Y es Seg/ Cls Alzheimer’s Disease 755 OASIS-2[ 681 ] 2010 3D MR Brain 1200 Y es Cls Alzheimer’s Disease 756 OASIS-3[ 74 ] 2019 3D CT/MR/ PET Brain 5699 Y es Seg/Cls Alzheimer’ s Disease 757 Learn2Reg O ASIS[ 497 ] 2020 3D MR Brain 416 Y es Reg Alzheimer’ s Disease 758 Learn2Reg Hippocampus MR[ 469 ] 2020 3D MR Brain 263 Y es Reg Anatomical segmenta- tion 759 Learn2Reg LUMIR[ 497 ] 2024 3D MR/US Brain 269 Y es Reg Multi-modal 760 PROMISE09[ 682 ] 2009 3D MR Prostate 15 Y es Seg Prostate cancer 761 PROMISE12[ 683 ] 2012 3D MR Prostate 50 Y es Seg Prostate cancer 762 Prostate-3T[ 684 ] 2013 3D MR Prostate 64 Y es Seg Prostate cancer 148 763 Prostate-Diagnosis[ 685 ] 2015 3D MR Prostate 92 Y es Seg/ Cls Prostate cancer 764 PROST A TEx[ 686 ] 2016 3D MR Prostate 204 Y es Cls Prostate cancer 765 PROST A TEx-2[ 687 ] 2017 3D MR Prostate 162 Y es Cls Prostate cancer 766 QIN Breast DCE-MR[ 688 ] 2014 3D MR Breast 10 Y es Seg Breast cancer 767 QIN-SARCOMA[ 689 ] 2014 3D MR Breast, Calf, Chest, Elbow , Knee, Leg, Shoulder ... 15 No Cls Soft-tissue sarcoma 768 QIN GBM Treatment Response[ 690 ] 2015 2D/ 3D MR Brain 54 No Cls Glioblastoma Multi- forme 769 QIN-BRAIN-DSC-MR[ 691 ] 2016 3D MR Brain 49 Y es Seg Glioma 770 QIN-PROST A TE- Repeatability[ 692 ] 2018 3D MR Prostate 15 Y es Seg Prostate cancer 771 QIN-BREAST -02[ 693 ] 2019 3D MR Breast 13 No Cls Breast cancer 772 Prostate-MR-US-Biopsy[ 694 ] 2020 3D MR/ US Prostate 1151 Y es Reg/Seg Prostate Cancer 773 Prostate MR Segmentation Dataset[ 695 ] 2020 3D MR Prostate 116 Y es Seg Prostate cancer 774 PROST A TEx-Seg-Zones[ 696 ] 2020 3D MR Prostate 98 Y es Seg Prostate cancer 775 PROST A TEx-Seg-HiRes[ 697 ] 2020 3D MR Prostate 66 Y es Seg Prostate cancer 776 PI-CAI[ 698 ] 2022 3D MR Prostate 1500 Y es Seg/ Cls Prostate cancer 777 µ -RegPro2023[ 699 ] 2023 3D MR/ US Prostate 108 Y es Reg/lmk Prostate cancer 778 SKI10[ 700 ] 2010 3D MR Bone, Cartilage 150 Y es Seg Osteoarthritis 779 T ADPOLE[ 701 ] 2017 3D MR/ PET Brain 1667 Y es Cls/ Reg Alzheimer’s Disease 780 MSSEG-2[ 702 ] 2021 3D MR Brain 100 Y es Seg Multiple sclerosis 781 MSSEG 2016[ 703 ] 2016 2D/ 3D MR Brain 15 Y es Seg Multiple sclerosis 782 MSSEG 2008[ 704 ] 2008 3D MR Brain 38 Y es Seg Multiple Sclerosis 783 Cam-CAN[ 705 ] 2015 3D MEG/ MR Brain 3000 No Cls Healthy ageing 784 ISLES 2015[ 706 ] 2015 3D MR Brain 114 Y es Seg Ischemic stroke 785 ISLES 2016[ 707 ] 2016 3D MR Brain 75 Y es Seg Ischemic stroke 786 ISLES 2017[ 707 ] 2017 3D MR Brain 75 Y es Seg Ischemic stroke 787 ISLES 2018[ 708 ] 2018 3D MR Brain 103 Y es Seg Ischemic stroke 788 ISLES 2022[ 709 ] 2022 3D MR Brain 400 Y es Seg Ischemic stroke 789 ISLES 2024[ 492 ] 2024 3D MR Brain 250 Y es Seg Acute/subacute ischemic stroke 790 WMH[ 710 ] 2017 3D MR Brain 170 Y es Seg White matter hyperin- tensities 791 BraTS 2012[ 73 ] 2012 3D MR Brain 50 Y es Seg Glioma 792 BraTS 2013[ 73 ] 2013 3D MR Brain 60 Y es Seg Glioma 793 BraTS 2014[ 73 ] 2014 3D MR Brain 238 Y es Seg Glioma 794 BraTS 2015[ 73 ] 2015 3D MR Brain 253 Y es Seg/Prog Glioma 795 BraTS 2016[ 711 ] 2016 3D MR Brain 391 Y es Seg/Prog Glioma 796 BraTS 2017[ 537 ] 2017 3D MR Brain 477 Y es Seg/Surv Glioma 797 BraTS 2018[ 73 ] 2018 3D MR Brain 542 Y es Seg/Surv Glioma 798 BraTS 2019[ 73 ] 2019 3D MR Brain 626 Y es Seg/Surv Glioma 799 BraTS 2020[ 712 ] 2020 3D MR Brain 660 Y es Seg/Surv Glioma 800 BraTS 2021[ 713 ] 2021 3D MR Brain 2040 Y es Seg/MGMT Glioma 801 BraTS 2022[ 537 ] 2022 3D MR Brain 1470 Y es Seg Glioma 802 BraTS 2023[ 714 ] 2023 3D MR Brain 5880 Y es Seg Glioma 803 MSD01 BrainT umour[ 469 ] 2018 3D MR Brain 750 Y es Seg Brain tumor 804 MSD02 Heart[ 469 ] 2018 3D MR Heart 30 Y es Seg Cardiac conditions 805 MSD04 Hippocampus[ 469 ] 2018 3D MR Hippocampus 394 Y es Seg Anatomical segmenta- tion 806 MSD05 Prostate[ 469 ] 2018 3D MR Prostate 48 Y es Seg Prostate cancer 807 FeT A 2021[ 715 ] 2021 3D MR Brain 50 Y es Seg Congenital Disorders 808 FeT A 2022[ 716 ] 2022 3D MR Brain 80 Y es Seg Neurodevelopment disorders 809 fastMR[ 717 ] 2020 2D/ 3D MR Brain/ Knee/ Prostate/ Breast 1594 No Rec Multi-organ 810 High Anisotropy MR[ 718 ] 2013 3D MR Spine 17 No Rec Spine conditions 811 1000 Functional Connectomes Project[ 719 ] 2010 3D MR Brain 1414 No Cls Adult ADHD 812 Language Processing Children[ 720 ] 2022 3D MR Brain 322 Y es Cls Language impairment 813 ISPY1 (ACRIN 6657)[ 721 ] 2016 3D MR Breast 847 Y es Seg/ Cls Breast cancer 814 ACRIN-Contralateral-Breast- MR[ 722 ] 2021 3D CR/MR Breast 984 No Cls Breast Cancer 815 ACRIN 6698/I-SPY2 Breast DWI[ 723 ] 2021 3D MR Breast 385 Y es Seg/ Cls Breast cancer 816 ADNI[ 76 ] 2017 3D MR/ PET Brain 2500 No Cls Alzheimer’s Disease 817 ADNIDOD[ 724 ] 2017 3D MR/ PET Brain 195 No Cls Alzheimer’s Disease 818 ABVIB[ 725 ] 2017 3D MR Brain 280 No Cls Alzheimer’ s Disease 819 AIBL[ 726 ] 2017 3D MR/ PET Brain 278 Y es Cls Alzheimer’s Disease 820 A OMIC-ID1000[ 727 ] 2021 3D MR Brain 928 No Reg Healthy adults 821 BOLD V erb Generation[ 728 ] 2020 4D MR Brain 143 No Reg Developmental lan- guage disorder 149 822 Brain Correlates of Math Development[ 729 ] 2018 3D MR Brain 132 No Cls Developmental disor - ders 823 Brain Segmentation T esting Protocol[ 730 ] 2011 3D MR Brain 312 Y es Seg Alzheimer’s Disease 824 BrainMetShare[ 731 ] 2020 3D MR Brain 156 Y es Seg Brain metastases 825 Breast-MR-NA CT -Pilot[ 732 ] 2016 3D MR Breast 64 Y es Seg Breast cancer 826 CAMR Rat Brain MR Data[ 733 ] 2020 3D MR Brain 264 No Reg Animal model study 827 Caltech Conte Center[ 734 ] 2022 3D MR Brain 117 No Cls Healthy 828 Cognitive T raining[ 735 ] 2020 3D MR Brain 166 No Cls Cognitive training study 829 Colin 3T/7T High-resolution Atlas[ 736 ] 2014 3D MR Brain 19 No Reg Brain diseases 830 Concrete Permuted Rule Operations[ 737 ] 2021 3D MR Brain 96 No Cls Cognitive function 831 Cortical Myelin T1w/T2w[ 738 ] 2021 3D MR Brain 86 No Cls Unipolar depressive disorders 832 Cross-Sectional Multidomain Lexical Processing[ 739 ] 2019 3D MR Brain 91 No Cls Language processing study 833 Prefrontal Cortex Development[ 740 ] 2021 3D MR Brain 90 No Cls Developmental study 834 Duke-Breast-Cancer-MR[ 741 ] 2021 3D MR Breast 922 Y es Seg/ Cls Invasi ve breast cancer 835 Dynamic Passiv e Threat[ 742 ] 2019 3D MR Brain 295200 No Cls Threat processing study 836 Emotion Regulation Ageing Brain[ 743 ] 2020 3D MR Brain 34 No Cls Normal ageing 837 Resting State Bilinguals[ 744 ] 2019 3D MR Brain 823 No Cls Healthy adults 838 Brain Genomics Superstruct Project (GSP)[ 745 ] 2015 3D MR Brain 1570 No Seg Healthy adults 839 Human Connectome Project (HCP)[ 75 ] 2017 3D MR Brain 1206 No Cls/Rec Healthy young adults 840 IXI Dataset[ 746 ] 2024 3D MR Brain 600 No Reg/Rec Healthy adults 841 KNOAP2020[ 747 ] 2020 2D/ 3D MR/X-RA Y Knee 453 Y es Cls Knee Osteoarthritis 842 LGG-1p19qDeletion[ 748 ] 2017 3D MR Brain 159 Y es Seg/ Cls Low Grade Glioma 843 Lausanne TOF-MRA Aneurysm Cohort[ 749 ] 2021 3D MR Brain 284 Y es Det Brain aneurysm 844 MASiV ar[ 750 ] 2021 3D MR Brain 319 No T rk Healthy subjects 845 MIRIAD dataset[ 751 ] 2013 3D MR Brain 708 No Reg Alzheimer’s Disease 846 MPI-Leipzig Mind-Brain- Body[ 752 ] 2019 3D EEG/MR Brain 318 No Cls Healthy cohort 847 Cue Induced Craving MR[ 753 ] 2020 3D MR Brain 598 No Rec Normal neurophysio- logical states 848 Children Adults Animated Film MR[ 754 ] 2018 3D MR Brain 155 No Rec Developmental study 849 Mouse rest multicentre[ 755 ] 2019 3D MR Brain 255 No Cls Healthy mouse model 850 Multi-echo Cambridge[ 756 ] 2018 3D MR Brain 89 No Rec Healthy neurotypical 851 NARPS[ 757 ] 2019 3D MR Brain 108 No Cls Healthy participants 852 Narratives[ 758 ] 2019 3D MR Brain 891 No Cls Healthy participants 853 Naturalistic Neuroimaging Database[ 759 ] 2021 3D MR Brain 86 No Cls Healthy controls 854 Neurocognitive aging data release with behavioral[ 760 ] 2022 3D MR Brain 301 No Cls Healthy cognitiv e aging 855 PETfrog[ 761 ] 2020 3D MR/ PET Brain 238 No Cls Brain development 856 Pragmatic Language[ 762 ] 2021 3D MR Brain 145 No Loc Pragmatic comprehen- sion deficits 857 REMBRANDT[ 763 ] 2014 3D MR Brain 130 Y es Seg/ Cls Gliomas 858 SIMON Dataset[ 764 ] 2019 3D MR Brain 73 No Reg Healthy Control 859 SUDMEX CONN[ 765 ] 2021 3D MR Brain 138 No Cls Cocaine use disorder 860 Serum Grey Matter Cortical Thickness[ 766 ] 2020 3D MR Brain 143 No Reg Brain morphometry study 861 Speech disfluencies: Neuro- physiological aspect in normal population[ 767 ] 2021 3D MR Brain 81 No Cls Speech disorders 862 T1 Chronotype Sleep Study[ 768 ] 2021 3D MR Brain 136 No Cls Healthy 863 TCGA-GBM-QI- Radiogenomics[ 769 ] 2014 3D MR Brain 55 Y es Seg Glioblastoma 864 TCGA-GBM- Radiogenomics[ 769 ] 2014 3D MR Brain 75 Y es Seg Glioblastoma 865 TCGA-BRCA[ 770 ] 2014 2D/ 3D MG/MR Breast 139 No Cls Breast cancer 866 TCGA-CESC[ 531 ] 2014 3D MR Cervix 54 No Cls Cervical cancer 867 TCGA-Breast- Radiogenomics[ 771 ] 2015 3D MR Breast 84 Y es Seg/ Cls Breast cancer 868 BraTS-TCGA-GBM[ 537 ] 2017 3D MR Brain 135 Y es Seg Glioma 869 BraTS-TCGA-LGG[ 537 ] 2017 3D MR Brain 108 Y es Seg Glioma 870 TCGA-LGG-Mask[ 537 ] 2017 3D MR Brain 188 Y es Seg Low Grade Glioma 871 Stockholm Sleepy Brain Study[ 772 ] 2018 3D MR Brain 84 No Cls Sleep deprivation 872 Harm A v oidance Gray Matter[ 773 ] 2016 3D MR Brain 95 No Cls Personality traits 873 Human V oice Areas[ 774 ] 2015 3D MR Brain 218 Y es Cls Healthy 150 874 UCLA Consortium for Neuropsychiatric Phenomics LA5c Study[ 775 ] 2016 3D MR Brain 273 No Cls Neuropsychiatric Disorders 875 W ashington Uni versity 120[ 776 ] 2017 3D MR Brain 120 No Rest Healthy young adults 876 White matter deficits in co- caine use disorder V1.0[ 777 ] 2021 3D MR Brain 133 No Cls Cocaine Use Disorder 877 W orking memory in healthy and schizophrenic individuals[ 778 ] 2016 3D MR Brain 99 No Cls Schizophrenia 878 rsfMR comorbidity SmokingandSchizophrenia[ 779 ] 2018 3D MR Brain 92 Y es Cls Schizophrenia and nicotine dependence 879 IVDM3Seg[ 780 ] 2018 3D MR Intervertebral Discs 96 Y es Seg/Loc Spine diseases 880 MRNet[ 781 ] 2018 3D MR Knee 1370 Y es Cls Knee MR abnormali- ties 881 EMIDEC[ 782 ] 2020 3D MR Heart 150 Y es Seg/ Cls Myocardial infarction 882 iSeg2017[ 783 ] 2017 3D MR Brain 23 Y es Seg Neurodevelopmental conditions 883 UW -Madison GI T ract Image Segmentation[ 784 ] 2022 3D MR Stomach, Small Bowel, Large Bowel 467 Y es Seg Gastrointestinal cancers 884 MUDI2019[ 785 ] 2019 3D MR Brain 1344 Y es Rec Healthy 885 iSeg-2019[ 786 ] 2019 3D MR Brain 39 Y es Seg Healthy development 886 Longitudinal Multiple Sclero- sis Lesion Segmentation[ 787 ] 2015 3D MR Brain 82 Y es Seg Multiple sclerosis 887 COSMOS 2022[ 788 ] 2022 3D MR Carotid Artery 75 Y es Seg Atherosclerosis 888 cSeg-2022[ 789 ] 2022 3D MR Cerebellum 33 Y es Seg Normal development 889 Brain T umor Progression Prediction[ 790 ] 2021 3D MR Brain 40 Y es Cls Brain cancer 890 Heart Segmentation in MR Images 2021 3D MR Heart 30 Y es Seg Cardiac conditions 891 VWS 2021[ 791 ] 2021 3D MR Carotid Arteries 50 Y es Seg Atherosclerosis 892 Atrial Segmentation Challenge[ 792 ] 2018 3D MR Heart 154 Y es Seg Atrial fibrillation 893 IronTract Challenge 2019[ 793 ] 2019 3D MR Brain 2 Y es Trk Anatomical structure 894 DiSCo 2021[ 794 ] 2021 3D MR Brain 3 Y es Reg Tractography challenge 895 RealNoiseMR 2021[ 795 ] 2021 3D MR Brain 25 Y es Rec Denoising challenge 896 MOOD 2022-brain[ 657 ] 2022 3D MR Brain 800 Y es Cls/Loc General pathologies 897 AAPM-R T -MAC[ 796 ] 2019 3D MR Head-Neck 55 Y es Seg Head and Neck Cancer 898 HVSMR 2016[ 797 ] 2016 3D MR Heart 20 Y es Seg Congenital heart disease 899 MRBrainS13[ 71 ] 2013 3D MR Brain 20 Y es Seg Age-related brain conditions 900 HARDI Reconstruction Challenge Dataset[ 798 ] 2013 3D MR Brain 6 Y es Rec Diffusion imaging 901 CA USE07[ 799 ] 2007 3D MR Brain 38 Y es Seg Neurological Disorders 902 PROMISE09[ 682 ] 2009 3D MR Prostate 15 Y es Seg Prostate cancer 903 ISMRM2015[ 800 ] 2015 3D MR Brain 34 Y es Rec/Trk Tractography challenge 904 Where is V ALDO?[ 801 ] 2021 3D MR Brain 306 Y es Seg/Det/Loc Cerebral Small V essel Disease 905 NEA TBrainS15[ 71 ] 2015 3D MR Brain 20 Y es Seg Age-related brain conditions 906 MRBrainS18[ 802 ] 2018 3D MR Brain 30 Y es Seg Diabetes, Dementia, Alzheimer’ s 907 ST ACOM 2011[ 803 ] 2011 3D MR/ US Heart 1158 Y es Reg/Trk Healthy volunteers 908 3T Brain-Behavior MR[ 804 ] 2014 3D MR Brain 36 No Reg Healthy subjects 909 Connectivity T est-Retest MR[ 805 ] 2015 3D MR Brain 342 No T rk Healthy volunteers 910 Mindboggle-101[ 806 ] 2012 3D MR Brain 101 Y es Seg Anatomical segmenta- tion 911 Individual Brain Charting (IBC)[ 807 ] 2020 3D MR Brain 600 Y es Cls Healthy 912 Raider[ 808 ] 2015 3D MR Brain 11 No Cls Healthy 913 Diffusion MR Data Harmonisation[ 809 ] 2017 3D MR Brain 14 Y es Reg Cross-scanner harmo- nization 914 MEMENTO[ 810 ] 2019 3D MR Brain 1536 Y es Reg Neurodegenerativ e diseases 915 MUDI2019[ 785 ] 2019 3D MR Brain 6720 No Reg/Rec Microstructure imaging 916 CMRxMotion[ 811 ] 2022 3D MR Heart 360 Y es Seg/ Cls Not specified 917 PDMR-BL0293-F563[ 812 ] 2019 3D MR Liv er/Bone 19 No Cls Bladder cancer metas- tasis 918 PDMR-292921-168-R[ 629 ] 2020 3D MR/SR Abdomen 20 No Cls Pancreatic adenocarci- noma 919 PDMR-997537-175-T[ 629 ] 2020 3D MR/SR Colon 24 No Cls Colon adenocarcinoma 920 PDMR-425362-245-T[ 629 ] 2021 3D MR/SR Abdomen 20 No Cls Melanoma 151 921 PDMR-521955-158-R4[ 813 ] 2022 3D MR/SR Pancreas/ Lung 20 No Cls Pancreatic adenocarci- noma 922 ICDC-Glioma[ 814 ] 2021 2D/ 3D MICROSCOPY/ MR Brain 78 No Seg Glioma 923 Generation R Pediatric MR Resources[ 815 ] 2014 3D MR Brain 666 No Cls Normative de velop- mental 924 High-quality diffusion- weighted imaging of Parkin- son’s disease[ 816 ] 2014 3D MR Brain 53 No Cls Parkinson’ s disease 925 MGH Neonatal/Pediatric ADC Atlases[ 817 ] 2015 3D MR Brain 201 Y es Reg Acute brain injury 926 RIDER Phantom MR[ 818 ] 2011 3D MR Phantom 10 No QA Phantom study 927 RIDER Breast MR[ 819 ] 2011 3D MR Breast 40 No Cls Breast cancer 928 A TLAS2023[ 820 ] 2023 3D MR Liver 90 Y es Seg Hepatocellular carci- noma 929 SMILE-UHURA2023[ 821 ] 2023 3D MR Brain 25 Y es Seg Cerebral Small V essel Diseases 930 CMRxRecon[ 822 ] 2023 2D/ 3D MR Heart 300 Y es Rec/Seg Cardiac diseases 931 CAS2023 2023 3D MR Brain 150 Y es Seg Anatomical segmenta- tion 932 CRO WN2023[ 823 ] 2023 3D MR Brain 600 Y es Cls/ Reg Multiple pathologies 933 QuantConn[ 824 ] 2023 3D MR Brain 206 Y es Rec Microstructure quan- tification 934 BONBID-HIE2023[ 825 ] 2023 3D MR Brain 133 Y es Seg Hypoxic Ischemic Encephalopathy 935 LLD-MMR2023[ 826 ] 2023 3D MR Liver 498 Y es Cls Liver diseases 936 WBMR-NF[ 827 ] 2023 3D MR Whole- body 400 Y es Seg/Det Neurofibromatosis 937 SLCN[ 828 ] 2023 3D MR Brain 514 Y es Reg/Cls Neurodevelopmental disorders 938 SPPIN2023[ 829 ] 2023 3D MR Abdomen 96 Y es Seg Neuroblastoma 939 Shifts Challenge 2022[ 830 ] 2022 3D MR Brain 172 Y es Seg Multiple sclerosis 940 Mouse-Astrocytoma[ 831 ] 2017 3D MR Brain 48 No Cls Glioblastoma Multi- forme 941 GBM-MR-NER- Outcomes[ 832 ] 2014 3D MR Brain 45 No Reg Glioblastoma 942 UPENN-GBM[ 833 ] 2022 3D MR Brain 3680 Y es Seg Glioblastoma 943 ISPY1-T umor-SEG- Radiomics[ 834 ] 2022 3D MR Breast 163 Y es Seg Breast cancer 944 I-SPY2 Trial[ 835 ] 2022 3D MR Breast 719 Y es Seg/ Cls Breast cancer 945 IvyGAP-Radiomics[ 836 ] 2020 3D MR Brain 37 Y es Seg Glioblastoma 946 Brain-TR-GammaKnife[ 837 ] 2023 3D MR Brain 47 Y es Seg/ Cls Brain cancer 947 ExACT[ 838 ] 2023 3D MR Anus 30 Y es Cls/Loc Anal cancer 948 ACNS0332-T umor - Annotations[ 839 ] 2022 3D MR Brain 85 Y es Seg Brain tumor 949 UCSF-PDGM[ 840 ] 2022 3D MR Brain 501 Y es Seg Diffuse Gliomas 950 RHUH-GBM[ 841 ] 2023 3D MR Brain 40 Y es Seg Glioblastoma 951 RIDER Neuro MR[ 842 ] 2011 3D MR Brain 19 No Cls Brain cancer 952 Meningioma-SEG- CLASS[ 843 ] 2023 3D MR Brain 96 Y es Seg/ Cls Intracranial menin- giomas 953 Mouse-Mammary[ 831 ] 2015 3D MR Mammary 32 No Cls Breast Cancer 954 MRQy-Quality- Measures[ 844 ] 2020 3D MR Brain, Cervix 233 No QA Brain and Cervical Cancers 955 DICOM-Glioma-SEG[ 845 ] 2020 3D MR Brain 167 Y es Seg Brain cancer 956 ADAM2020[ 846 ] 2020 3D MR Brain 255 Y es Seg/Det/Cls Intracranial aneurysms 957 CSI15[ 847 ] 2015 2D/ 3D MR/X-RA Y Spine 345 Y es Cls/Seg Spine conditions 958 LPBA40[ 848 ] 2001 3D MR Brain 40 Y es Seg Healthy 959 Continuous Registration ISBR18[ 849 ] 2018 3D MR Brain 18 Y es Seg Registration challenge 960 CUMC12[ 850 ] 2018 3D MR Head and Neck 18 Y es Seg Not specified 961 MGH10[ 497 ] 2018 3D MR Brain 10 Y es Seg Anatomical segmenta- tion 962 BrainPTM 2021[ 851 ] 2021 3D MR Brain 75 Y es Seg Brain tumors 963 OpenMind[ 72 ] 2024 3D MR Head and Neck 114570 Y es Seg/Rec Health Status T otal: 523,847+ T able 23: 3D US datasets # Dataset Y ear Dim Modality Structure V olumes Label T ask Diseases 964 TDSC-ABUS2023[ 852 ] 2023 3D US Breast 200 Y es Seg/ Cls/ Det Breast cancer 965 CETUS2014[ 853 ] 2014 3D US Heart 45 Y es Seg Cardiac conditions 966 MVSeg- 3DTEE2023[ 854 ] 2023 3D US Mitral valv e 175 Y es Seg Mitral valve disease 967 AREN0532[ 514 ] 2022 2D/3D US/ CR/ CT/ MR/ PET/ R TIMA GE Kidney 544 No Cls W ilms tumor 968 AREN0533[ 516 ] 2022 3D US/CR/CT/MR Kidne y 294 No Cls Wilms tumor 152 969 AREN0533-T umor- Annotations[ 517 ] 2023 3D US/CR/CT/MR Kidney 294 Y es Seg Wilms tumor 970 AREN0534[ 518 ] 2021 2D/3D US/ CT/ MR/ PET Kidney 239 Y es Seg Wilms tumor 971 CPT AC-PDA[ 547 ] 2018 3D US/ CT/ MR/ PET Pancreas 168 No Cls Pancreatic cancer 972 CPT AC-SAR[ 548 ] 2019 2D/3D US/ CT/ MR/ PET Abdomen/ Arm/ Bladder/ Chest/ Head–Neck/ Kidney/ Leg/ ... 88 No Cls Sarcomas 973 CPT AC-UCEC[ 548 ] 2019 2D/3D US/ CT/ MR/ PET Uterus 250 No Cls Endometrial Carcinoma 974 CMB-CRC[ 553 ] 2022 2D/3D US/ CT/ MR/ PET/ WSI Colon 12 No Cls Colorectal Cancer 975 CMB-LCA[ 553 ] 2022 2D/3D US/ CT/ DX/ MR/ NM/ PT Lung 16 No Cls Lung cancer 976 CMB-MEL[ 553 ] 2022 2D/3D US/ CT/ PET/ WSI Skin 40 No Cls Melanoma 977 AHEP0731[ 567 ] 2021 3D US/ CT/ MR/ PET/ XA Liver/Chest 190 No Seg/Cls Liver Cancer 978 QIDW[ 585 ] 2015 3D US/ CT/ MR/ PET Lung 52000 No QA Quality assurance 979 CuRIOUS2018-MR FLAIR[ 676 ] 2018 3D US/MR Brain 33 Y es Reg Brain tumors 980 CuRIOUS2018-US[ 77 ] 2018 3D US/MR Brain 32 Y es Reg Brain tumors 981 CuRIOUS2018-MR T1W[ 77 ] 2018 3D US/MR Brain 33 Y es Reg Brain tumor 982 CuRIOUS2019-MR- FLAIR[ 677 ] 2019 3D US/MR Brain 32 Y es Reg Low-grade gliomas 983 CuRIOUS2019[ 77 ] 2019 3D US/MR Brain 33 Y es Reg Brain tumor 984 CuRIOUS2019 US[ 677 ] 2019 3D US/MR Brain 33 Y es Reg Low-grade gliomas 985 CuRIOUS2019-MR T1W[ 677 ] 2019 3D US/MR Brain 33 Y es Reg Brain tumor 986 CuRIOUS2022[ 678 ] 2022 3D US/MR Brain 33 Y es Seg Low-grade gliomas 987 Learn2Reg LUMIR[ 497 ] 2024 3D US/MR Brain 269 Y es Reg Multi-modal 988 Prostate-MR-US- Biopsy[ 694 ] 2020 3D US/MR Prostate 1151 Y es Reg/Seg Prostate Cancer 989 µ -RegPro2023[ 699 ] 2023 3D US/MR Prostate 108 Y es Reg/lmk Prostate cancer 990 ST ACOM 2011[ 803 ] 2011 3D US/MR Heart 1158 Y es Reg/Trk Healthy v olunteers T otal: 56,609+ T able 24: 3D PET datasets # Dataset Y ear Dim Modality Structure V olumes Label T ask Diseases 991 QIN PET Phantom[ 855 ] 2014 3D PET Body 2 Y es Seg Phantom study 992 AutoPET[ 448 ] 2022 3D PET/CT Whole-body 1014 Y es Seg Cancer 993 AutoPET II[ 449 ] 2023 3D PET/CT Whole-body 1219 Y es Seg Cancer 994 CO VID-19-NY -SBU[ 481 ] 2021 2D/ 3D PET/CT/ MR/ X-RA Y Brain/ Chest 1384 No Cls COVID-19 995 HECKTOR 2020[ 501 ] 2020 3D PET/CT Head and Neck 254 Y es Seg Head and Neck Cancer 996 HECKTOR 2021[ 502 ] 2021 3D PET/CT Head and Neck 325 Y es Seg/ Reg Head and Neck Cancer 997 HECKTOR 2022[ 503 ] 2022 3D PET/CT Head and Neck 883 Y es Seg/ Reg Head and Neck Cancer 998 ACRIN-HNSCC- FDG-PET -CT (ACRIN 6685)[ 510 ] 2016 3D PET/CT/MR/NM Head and Neck 260 Y es Cls Head and Neck Cancer 999 ACRIN-FL T -Breast (ACRIN 6688)[ 512 ] 2017 3D PET/CT Breast 83 Y es Cls Breast Cancer 1000 ACRIN-FMISO-Brain (ACRIN 6684)[ 513 ] 2016 3D PET/CT/MR Brain 45 Y es Seg/Cls Glioblastoma 1001 ACRIN-NSCLC-FDG- PET (ACRIN 6668)[ 511 ] 2020 3D PET/CT Lung 242 Y es Cls Lung cancer 1002 AREN0532[ 514 ] 2022 2D/3D PET/CR/ CT/ MR/ R TIMA GE/ US Kidney 544 No Cls Wilms tumor 1003 AREN0534[ 518 ] 2021 2D/ 3D PET/CT/ MR/ US Kidney 239 Y es Seg Wilms tumor 1004 AHOD0831[ 519 ] 2022 2D/3D PET/CR/ CT/ DX/ MR/ NM/ OT/ SC/ XA L ymphatic 165 Y es Seg Hodgkin Lymphoma 1005 AHOD0831-T umor- Annotations[ 520 ] 2023 3D PET/CT L ymph nodes, spleen, sali- vary glands, W alde yer’... 165 Y es Seg Hodgkin L ymphoma 1006 HNSCC[ 522 ] 2020 3D PET/CT/MR Head and Neck 627 Y es Seg Head and Neck Cancer 1007 CC-T umor Heterogeneity[ 528 ] 2023 3D PET/CT/MR Cervix 23 Y es Seg/Cls Cervical cancer 1008 TCGA-BLCA 2014 3D PET/CT/ MR/ X-RA Y Bladder 120 No Cls Bladder carcinoma 1009 TCGA-HNSC[ 533 ] 2014 3D PET/CT/ MR Head and Neck 479 No Cls Head and Neck Cancer 1010 TCGA-KIRP[ 534 ] 2014 3D PET/CT/ MR Kidney 33 No Cls Kidney cancer 1011 TCGA-LIHC[ 538 ] 2014 2D/3D PET/CT/MR Li ver 97 No Cls Liver cancer 1012 TCGA-LUSC 2016 3D PET/CT Lung 37 No Cls Lung cancer 1013 TCGA-PRAD[ 533 ] 2015 3D PET/CT/MR Prostate 14 No Cls Prostate cancer 153 1014 TCGA-UCEC[ 538 ] 2020 3D PET/CT/ MR/ X-RA Y Uterus 65 No Cls Uterine cancer 1015 CPT AC-LSCC[ 127 ] 2018 2D/3D PET/CT/ MI- CROSCOPY Lung 212 No Cls Lung cancer 1016 CPT AC-LUAD[ 546 ] 2018 2D/ 3D PET/CT/ MR Lung 244 No Cls Lung cancer 1017 CPT AC-PDA[ 547 ] 2018 3D PET/CT/ MR/ US Pancreas 168 No Cls Pancreatic cancer 1018 CPT AC-SAR[ 548 ] 2019 2D/ 3D PET/CT/ MR/ US Abdomen/ Arm/ Bladder/ Chest/ Head–Neck/ Kidney/ Leg/... 88 No Cls Sarcomas 1019 CPT AC-UCEC[ 548 ] 2019 2D/ 3D PET/CT/ MR/ US Uterus 250 No Cls Endometrial Carci- noma 1020 NSCLC- Radiogenomics[ 550 ] 2015 3D PET/CT Chest 211 Y es Seg/ Cls Lung cancer 1021 CMB-CRC[ 553 ] 2022 2D/3D PET/CT/ MR/ US/ WSI Colon 12 No Cls Colorectal Cancer 1022 CMB-GEC[ 553 ] 2022 2D/3D PET/CT/ MI- CROSCOPY/ MR Esophagus 17 No Seg/Cls Gastroesophageal Cancer 1023 CMB-MEL[ 553 ] 2022 2D/ 3D PET/CT/ US/ WSI Skin 40 No Cls Melanoma 1024 CMB-MML[ 553 ] 2022 2D/3D PET/CR/ CT/ DX/ HISTOP A THOL- OGY/ MR/ XA Blood/Bone 138 No Cls Multiple Myeloma 1025 CMB-PCA[ 553 ] 2022 2D/3D PET/CT/ DX/ MR/ NM/ RF Prostate 50 No Cls Prostate cancer 1026 QIN-Breast[ 556 ] 2015 3D PET/CT/ MR Breast 68 Y es Cls Breast cancer 1027 QIN-HEADNECK[ 557 ] 2015 3D PET/CT Head and Neck 279 Y es Seg Head and neck carcino- mas 1028 AHEP0731[ 567 ] 2021 3D PET/CT/ MR/ US/ XA Liver/ Chest 190 No Seg/ Cls Liver Cancer 1029 Anti-PD-1 Lung[ 568 ] 2019 3D PET/CT/ SC Lung 46 No Cls Lung cancer 1030 CALGB50303[ 571 ] 2021 3D PET/CT Chest/ Abdomen/ Pelvis 155 Y es Cls Diffuse Lar ge B-Cell L ymphoma 1031 Head-Neck Cetuximab (R TOG 0522)[ 575 ] 2013 3D PET/CT Head and Neck 111 No Cls Head and Neck Carcinomas 1032 Head-Neck-PET -CT[ 576 ] 2017 3D PET/CT Head and Neck 298 Y es Seg/ Cls Head and Neck Cancer 1033 Head-Neck-Radiomics- HN1[ 549 ] 2019 3D PET/CT Head/ Neck 137 Y es Seg Head and Neck Cancer 1034 Lung-PET -CT -Dx[ 580 ] 2020 3D PET/CT Lung 355 Y es Cls/ Det Lung cancer 1035 Anti-PD-1 Immunotherapy Melanoma[ 569 ] 2019 3D PET/CT/MR Skin 47 No Cls Melanoma 1036 BREAST - DIAGNOSIS[ 570 ] 2011 2D/ 3D PET/CT/MG/MR Breast 88 Y es Cls Breast cancer 1037 Parkinson’s Progres- sion Markers Initiativ e (PPMI)[ 583 ] 2010 3D PET/MR/SPECT Brain 683 Y es Cls Parkinson’ s Disease 1038 QIDW[ 585 ] 2015 3D PET/CT/MR/US Lung 52000 No QA Quality assurance 1039 RIDER Lung PET - CT[ 134 ] 2015 3D PET/CT Lung 243 No Cls Lung cancer 1040 Soft-tissue-Sarcoma[ 590 ] 2015 3D PET/CT/MR Extremities 51 Y es Seg/Cls Soft-tissue sarcoma 1041 Seg Soft T issue[ 591 ] 2021 3D PET/CT/MR Soft tissue 51 Y es Seg Soft-tissue sarcomas (preprocessed) 1042 fastPET -LD[ 604 ] 2021 3D PET/CT Whole Body 68 Y es Det Oncologic Imaging 1043 CT -vs-PET -V entilation- Imaging[ 612 ] 2022 3D PET/CT Lung 20 No Cls Lung cancer 1044 NaF PROST A TE[ 619 ] 2013 3D PET/CT Prostate 9 No Cls Prostate cancer 1045 Pseudo-PHI-DICOM- Data[ 616 ] 2021 2D/3D PET/CT/MR/X- RA Y V arious 21 No Reg V arious cancers 1046 PDMR-833975-119- R[ 629 ] 2020 3D PET/CT/MR Pancreas 20 No Cls Pancreatic adenocarci- noma 1047 APOLLO-5-LSCC[ 630 ] 2021 3D PET/CT Lung 36 Y es Seg Lung squamous cell carcinoma 1048 ARAR0331[ 637 ] 2022 3D PET/CT/MR Head 108 Y es Seg Nasopharyngeal cancer 1049 CALGB50303-T umor- Annotations[ 520 ] 2023 3D PET/CT L ymphatic system 155 Y es Seg/ Cls Diffuse Lar ge B-Cell L ymphoma 1050 OASIS-3[ 74 ] 2019 3D PET/CT/MR Brain 5699 Y es Seg/Cls Alzheimer’ s Disease 1051 T ADPOLE[ 701 ] 2017 3D PET/MR Brain 1667 Y es Cls/ Reg Alzheimer’s Disease 1052 ADNI[ 76 ] 2017 3D PET/MR Brain 2500 No Cls Alzheimer’s Disease 1053 ADNIDOD[ 724 ] 2017 3D PET/MR Brain 195 No Cls Alzheimer’s Disease 1054 AIBL[ 726 ] 2017 3D PET/MR Brain 278 Y es Cls Alzheimer’ s Disease 1055 PETfrog[ 761 ] 2020 3D PET/MR Brain 238 No Cls Brain development T otal: 95,456+ T able 25: 3D Other datasets # Dataset Y ear Dim Modality Structure V olumes Label T ask Diseases 1056 MitoEM[ 856 ] 2020 3D 3D MICROSCOPY Brain 2 Y es Seg Mitochondrial ultrastructure 1057 3D Platelet EM[ 857 ] 2021 3D 3D MICROSCOPY Platelet 2 Y es Seg Platelet ultrastruc- ture 154 1058 PCa Bx 3Dpathology[ 858 ] 2023 3D 3D MICROSCOPY Prostate 50 Y es Seg/ Loc Prostate cancer 1059 CADA[ 859 ] 2020 3D 3D DSA Brain 131 Y es Det/ Seg/ Cls Cerebral aneurysms 1060 ISBI2023 challenge - SHINY - ICARUS[ 860 ] 2022 3D 3D DSA Internal Carotid Artery 82 Y es Seg Internal carotid artery aneurysms 1061 CADA-AS[ 861 ] 2020 3D 3D DSA Brain 110 Y es Seg Cerebral aneurysms 1062 CADA-RRE[ 861 ] 2020 3D 3D DSA Brain 131 Y es Seg/Cls Cerebral aneurysm 1063 XPRESS2023[ 862 ] 2023 3D 3D X-RA Y Brain 3 Y es Se g Neuroanatomy 1064 Learn2Reg ThoraxCBCT[ 617 ] 2023 3D CBCT/FBCT Thorax 40 Y es Reg Lung cancer 1065 Pancreatic-CT -CBCT - SEG[ 863 ] 2021 3D CBCT/CT Pancreas 40 Y es Se g Pancreatic cancer 1066 Pelvic-Reference-Data[ 864 ] 2019 3D CBCT/CT Pelvis 58 Y es Reg Prostate cancer 1067 T oothF airy2023[ 865 ] 2023 3D CBCT Inferior Alveolar Nerve 443 Y es Seg Dental surgery planning 1068 RETOUCH[ 866 ] 2017 3D OCT Retina 112 Y es Seg Retinal diseases 1069 ROCC[ 867 ] 2017 3D OCT Retina 72 Y es Cls Diabetic Retinopathy 1070 OCT Glaucoma Detection[ 868 ] 2018 3D OCT Optic nerve head 1110 Y es Cls Glaucoma 1071 OLIVES[ 869 ] 2022 2D/3D OCT Eye 1268 Y es Cls Diabetic conditions 1072 GAMMA[ 870 ] 2021 2D/3D OCT Retina 300 Y es Cls/Seg/Loc Glaucoma 1073 Farsiu Ophthalmology 2013[ 871 ] 2013 3D OCT Retina 384 Y es Seg/Cls Age-related macular degeneration 1074 Chiu IO VS 2011[ 872 ] 2012 3D OCT Retina 25 Y es Seg/Cls Macular Degenera- tion 1075 Chiu BOE 2014 dataset[ 873 ] 2014 3D OCT Retina 16 Y es Seg Diabetic macular edema 1076 Sriniv asan BOE 2014[ 874 ] 2014 3D OCT Retina 45 Y es Cls Eye diseases 1077 Soltanian Optica 2021[ 875 ] 2021 3D OCT Retina 8 Y es Seg Glaucoma 1078 ST AGE[ 876 ] 2023 3D OCT Retina 400 Y es Reg/Cls Glaucoma 1079 Eye OCT Datasets[ 877 ] 2021 3D OCT Retina 148 Y es Cls/Seg Retinal diseases 1080 OCT A-500[ 878 ] 2024 3D OCT/OCT A Retina 500 Y es Cls/Seg Retinal diseases 1081 OCT A2024[ 227 ] 2024 3D OCT/OCT A Retina TBD Y es Rec/Trans OCT to OCT A translation T otal: 5,381+ C T ables of Medical V ideo Datasets T able 26: V ideo datasets. # Dataset Y ear Dim Modality Structure Num of samples Label T ask Diseases 1082 CholecT50 [ 879 ] 2023 Video Endoscopy Gallbladder 50 Y es Cls, Det surgical instru- ment, action, target 1083 CholecTriplet 2021 [ 880 ] 2021 V ideo Endoscopy Gallbladder 45 Y es Cls, Det surgical instru- ment, action, target 1084 SurgV isDom [ 881 ] 2020 V ideo Endoscopy Bowel 488 Y es Cls skin lesion 1085 CA T ARACTS [ 54 ] 2017 V ideo Microscopy Retina 50 Y es Cls, Det surgical work- flow 1086 EndoVis 2018-SW AS [ 882 ] 2018 V ideo Endoscopy Colon 42 Y es Cls surgical phase 1087 EndoVis 2019-SWSA [ 883 ] 2019 Video Endoscopy Gallbladder 30 Y es Cls surgical phase, action, instru- ment 1088 EndoV is 2020- CA T ARA CTS W ork- flow [ 54 ] 2020 Video Microscopy Retina 50 Y es Cls surgical work- flow 1089 EndoV is 2020- MISA W [ 884 ] 2020 Video RGB Artificial vessel 27 Y es Cls surgical phase 1090 EndoV is 2021- PETRA W [ 885 ] 2021 Video Endoscopy NA 150 Y es Cls surgical work- flow 1091 EndoV is 2022- SurgT oolLoc [ 886 ] 2022 Video Endoscopy NA 24695 Y es Cls surgical instru- ment 1092 T3 Challenge [ 887 ] 2023 V ideo RGB N A 200 Y es Cls, Det, VQA life-saving intervention procedure 1093 Endo-FM [ 421 ] 2023 2D, V ideo Endoscopy NA 32896 No NA NA 1094 OSS [ 888 ] 2025 Video Endoscopy NA 330 Y es Cls surgical suturing skill 1095 FedSurg [ 889 ] 2024 V ideo Endoscopy NA 30 Y es Cls laparoscopic grading of the appendicitis 1096 CardiacUDC [ 890 ] 2023 Video Ultrasound Heart 992 Y es Seg, Cls cardiac anatomi- cal structures 1097 m2cai16-tool [ 402 ] 2016 V ideo Endoscopy Gallbladder 15 Y es Cls, Det surgical instru- ment 1098 Cholec80 [ 891 ] 2016 V ideo Endoscopy Gallbladder 80 Y es Cls surgical phase 155 1099 SAGES-CVS [ 892 ] 2024 Video Endoscopy Gallbladder, Cystic Duct, Cystic Artery , Hepatocystic Triangle, Li ver 18000 Y es Cls surgical skill 1100 CatRelDet [ 893 ] 2020 V ideo Microscopy Retina 8 Y es Cls, Det surgical phase 1101 SurgicalActions160 [ 894 ] 2017 V ideo Endoscopy Female Repro- ductive System 160 Y es Cls, Retrieval sur gical phase 1102 OphNet [ 895 ] 2024 V ideo Microscopy Retina 743 Y es Cls surgical phase 1103 NurVid [ 896 ] 2023 Video RGB NA 1,538 Y es Cls nursing proce- dure 1104 m2cai16-workflow [ 897 ] 2016 Video Endoscopy Gallbladder 41 Y es Cls surgical phase 1105 EndoCV 2021 [ 898 ] 2021 Video Endoscopy Polyp 4019 Y es Det surgical actions 1106 AdaptOR2021 [ 899 ] 2021 Video Endoscopy Heart 5584 Y es Det 2D landmarks 1107 CholecTriplet2022 [ 879 ] 2022 V ideo Endoscopy Gallbladder 45 Y es Cls, Det surgical instru- ment, action, target 1108 m2cai16-tool- locations [ 402 ] 2016 Video Endoscopy Gallbladder 15 Y es Det, Tracking surgical instru- ment 1109 A-AFMA [ 900 ] 2020 Video Ultrasound Bladder NA Y es Det amniotic fluid 1110 GIANA [ 901 ] 2017 V ideo Endoscopy Colon 3500 Y es Seg, Det angiodysplasia 1111 EndoV is 2021- SimSurgSkill [ 902 ] 2021 Video Endoscopy NA 321 Y es Det, Cls surgical tool clevis and needle, surgical skill 1112 A VOS [ 84 ] 2024 Video Endoscopy NA 1997 Y es Det, Tracking surgical instru- ment, action 1113 EndoV is 2022-SimCol-to- 3D [ 903 ] 2022 Video Endoscopy Colon 15 Y es Est depth, camera pose 1114 KBD [ 79 ] 2017 V ideo Endoscopy Kidney 4 Y es Seg kidney boundary 1115 EndoVis15-IST [ 904 ] 2015 2D, V ideo Endoscopy NA 100 Y es Seg, T racking surgical instru- ment 1116 Robotic Instrument Seg- mentation [ 905 ] 2017 Video Endoscopy Bowel 18 Y es Seg surgical instru- ment 1117 ROBUST -MIS [ 906 ] 2019 Video Endoscopy Colon 10 Y es Seg, Det surgical instru- ment 1118 EndoV is20- CA T ARA CTS [ 907 ] 2020 Video Microscopy Retina 50 Y es Seg surgical instru- ment 1119 EndoVis21-HeiSurf [ 908 ] 2021 V ideo Endoscopy NA 33 Y es Seg, Det, T rack- ing surgical phase, action, instru- ment, organ 1120 EndoVis22-P2ILF [ 91 ] 2022 V ideo, 3D Endoscopy , CT Liver 167 Y es Seg, Reg liver 1121 SegSTRONG-C [ 909 ] 2024 2D, Video Endoscopy NA 17 Y es Seg Surgical instru- ment 1122 SegCol [ 423 ] 2024 2D, V ideo Endoscopy NA 78 Y es Seg surgical instru- ment, colon folds 1123 FetReg [ 910 ] 2021 V ideo Endoscopy Placenta 2060 Y es Seg vessel 1124 EndoVis23-PitV is [ 911 ] 2024 Video Endoscopy Brain, Pituitary gland 25 Y es Seg, Det, Cls surgical phase 1125 EndoV is23- SurgT oolLoc [ 912 ] 2023 Video Endoscopy Pituitary , Ab- domen 949 Y es Seg, Det, Cls surgical instru- ment 1126 EndoVis18-RSS [ 913 ] 2018 Video Endoscopy Abdomen 15 Y es Seg, Tracking surgical instru- ment 1127 EndoV is22-SAR- RARP50 [ 914 ] 2022 Video Endoscopy Prostate 50 Y es Seg, Cls surgical instru- ment, action 1128 PhaKIR [ 915 ] 2024 V ideo Endoscopy Gallbladder 13 Y es Seg, Det, Cls surgical phase, instrument, instrument keypoint 1129 SurgVU [ 90 ] 2024 V ideo Endoscopy Abdomen 155 Y es Seg, Det, Cls surgical phase, instrument 1130 Cataract-1K [ 93 ] 2023 Video Microscopy Retina, Iris, Pupil 2256 Y es Seg, Det, Cls surgical phase, instrument, abnormality 1131 LensID [ 916 ] 2021 Video Microscopy Retina 2589 Y es Det, Seg surgical phase, instrument, anatomy 1132 AutoLaparo [ 917 ] 2022 V ideo Endoscopy Uterus 21 Y es Seg surgical phase, action, instru- ment and key anatomy 1133 CholecSeg8k [ 918 ] 2020 V ideo Endoscopy Gallbladder 17 Y es Seg surgical elements 1134 CholecInstanceSeg [ 919 ] 2024 Video Endoscopy Gallbladder 85 Y es Seg surgical instru- ment 1135 CaDIS [ 920 ] 2019 V ideo Microscopy Retina, Iris, Pupil 25 Y es Seg surgical full scene 1136 Endoscapes2023 [ 921 ] 2023 V ideo Endoscopy Gallbladder 201 Y es Cls, Det, Seg surgical anatomy , instrument, skill 1137 The Dresden Surgical Anatomy Dataset [ 922 ] 2023 Video Endoscopy Abdominal organs, vessel structures 32 Y es Seg surgical anatomy 1138 PolypGen [ 923 ] 2023 V ideo Endoscopy Polyp 2,225 Y es Seg polyp 156 1139 ASU-Mayo polyp database [ 924 ] 2022 Video Endoscopy Polyp 38 Y es Seg polyp 1140 GynSurg [ 925 ] 2025 V ideo Endoscopy Uterus 15 Y es Cls, Seg surgical instru- ment, anatomy 1141 SurgT: Sur gical Track [ 926 ] 2022 video Endoscopy N A 137 Y es Tracking soft tissue 1142 STIR [ 927 ] 2023 V ideo Endoscopy NA 60 Y es Tracking tissue keypoint 1143 HiSW A-RLLS [ 928 ] 2024 Video Endoscopy Liver 50 Y es Det, Cls surgical phase, action, instru- ment 1144 Egosurgery [ 929 ] 2024 V ideo Endoscopy NA 27000 Y es Cls, Det surgical phase, instrument 1145 TN-SCUI2020 [ 930 ] 2020 V ideo Ultrasound Thyroid gland 637 Y es Cls thyroid nodules 1146 EchoNet-Dynamic [ 931 ] 2020 V ideo Ultrasound Heart 10030 Y es Measurement heart 1147 Gastrointestinal Atlas [ 932 ] 2000 V ideo Endoscopy Bowel, Stomach 5142 No NA NA 1148 Cataract-101 [ 933 ] 2018 Video Microscopy Retina 101 Y es Cls surgical phase 1149 MedVidQA [ 934 ] 2022 V ideo RGB N A 3010 Y es VQA visual question answering 1150 HMC-QU [ 935 ] 2021 Video Ultrasound Heart 162 Y es Cls myocardial infarction 1151 Endovis 2019-SCRE [ 936 ] 2019 V ideo Endoscopy Porcine cadaver 9 Y es Recon, Est, Stereo Matching depth 1152 HyperKvasir [ 83 ] 2020 V ideo Endoscopy Colon, Esopha- gus, Stomach 373 Y es Seg, Det, Cls polyp 1153 ERS [ 937 ] 2022 V ideo Endoscopy Gastrointestinal tract 1520 Y es Cls, Seg Abnormality 1154 SUN-SEG [ 409 ] 2022 V ideo Endoscopy Colon 1106 Y es Seg polyp 1155 SARAS-MESAD [ 88 ] 2021 Video Endoscopy Prostate, Bladder 4 Y es Det surgical action 1156 Ophora-160K [ 81 ] 2025 Video Microscopy Retina 9819 Y es Video generation Video caption 1157 POCUS [ 938 ] 2020 V ideo Ultrasound Lung 64 Y es Cls COVID-19 1158 CLUST [ 154 ] 2014 V ideo Ultrasound Liver 63 Y es Tracking NA T otal: 166,691 Abbre viations: Seg=Se gmentation, Cls=Classification, Pred=Prediction, Det=Detection, Recon=Reconstruction, Reg=Registration, Est=Estimation, VQA=V isual Question Answering. 157
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment