Hierarchy-Guided Multimodal Representation Learning for Taxonomic Inference
Accurate biodiversity identification from large-scale field data is a foundational problem with direct impact on ecology, conservation, and environmental monitoring. In practice, the core task is taxonomic prediction - inferring order, family, genus,…
Authors: Sk Miraj Ahmed, Xi Yu, Yunqi Li
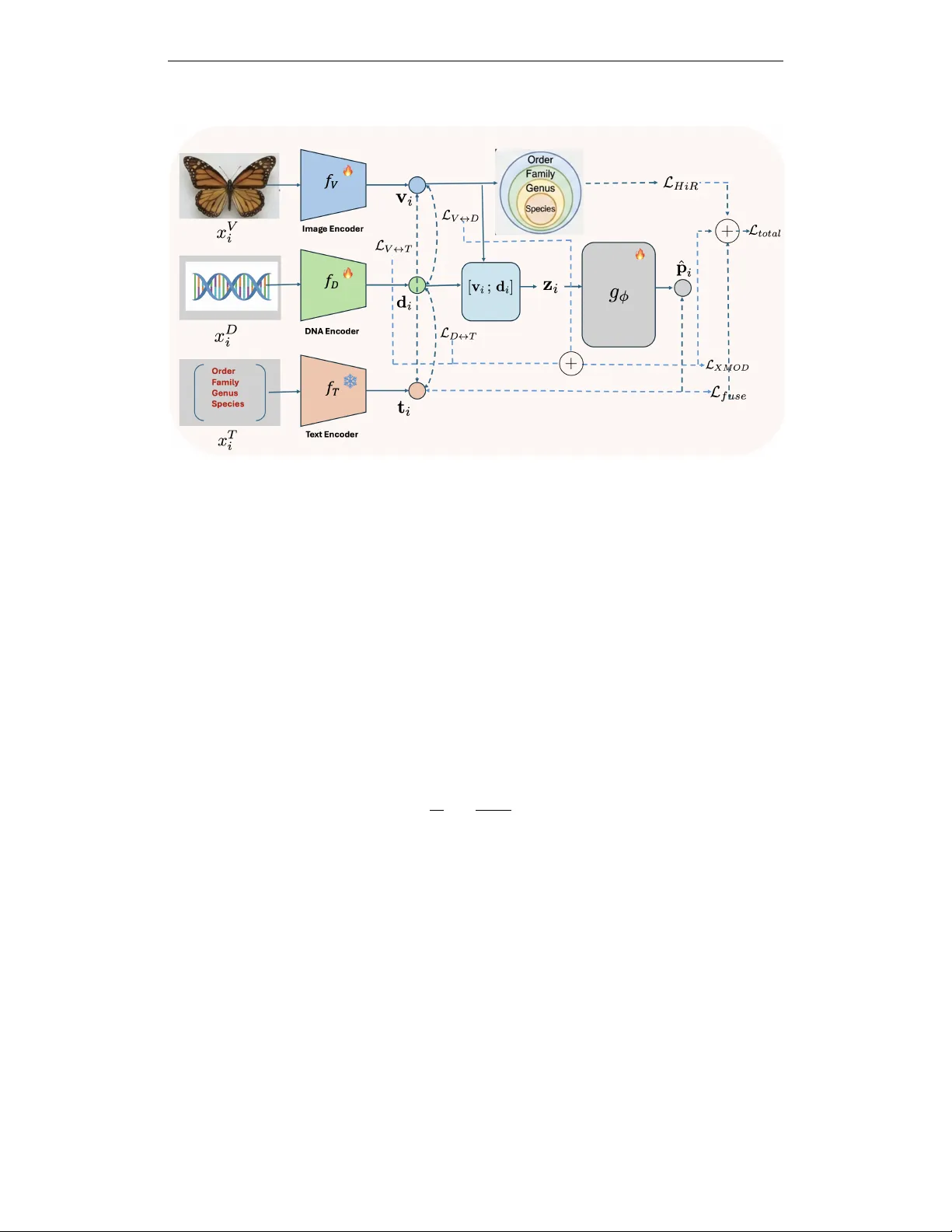
Published as a conference paper at ICLR 2026 H I E R A R C H Y - G U I D E D M U L T I M O DA L R E P R E S E N T A - T I O N L E A R N I N G F O R T A X O N O M I C I N F E R E N C E Sk Miraj Ahmed 1 , Xi Y u 1 , Y unqi Li 2 ∗ , Y uewei Lin 1 , W ei Xu 1 1 Computing and Data Sciences, Brookhav en National Laboratory , Upton, NY 11973, USA 2 Rutgers Univ ersity , Ne w Brunswick, NJ, USA 1 { sahmed3, xyu1, ywlin, xuw } @bnl.gov 2 yunqi.li@rutgers.edu A B S T R AC T Accurate biodiv ersity identification from large-scale field data is a foundational problem with direct impact on ecology , conservation, and en vironmental monitor- ing. In practice, the core task is taxonomic prediction —inferring order , family , genus, or species from imperfect inputs such as specimen images, DN A barcodes, or both. Existing multimodal methods often treat taxonomy as a flat label space and therefore fail to encode the hierarchical structure of biological classification, which is critical for robustness under noise and missing modalities. W e present two end-to-end variants for hierarchy-aw are multimodal learning: CLiBD-HiR , which introduces Hierar c hical Information Regularization (HiR) to shape em- bedding geometry across taxonomic levels, yielding structured and noise-rob ust representations; and CLiBD-HiR-Fuse , which additionally trains a lightweight fusion pr edictor that supports image-only , DN A-only , or joint inference and is re- silient to modality corruption. Across large-scale biodi versity benchmarks, our approach improves taxonomic classification accuracy by ov er 14% compared to strong multimodal baselines, with particularly large gains under partial and cor- rupted DN A conditions. These results highlight that explicitly encoding biological hierarchy , together with fle xible fusion, is key for practical biodi versity foundation models. 1 I N T RO D U C T I O N Biodiv ersity research V an Horn et al. (2018) increasingly relies on large-scale multimodal data, in- cluding specimen images, DN A barcodes, and auxiliary taxonomic metadata. While curated datasets enable controlled benchmarking, real-w orld deployments are far less controlled. In particular , DN A barcodes obtained through large-scale sequencing pipelines (e.g., BOLD Ratnasingham & Hebert (2007)) can exhibit partial reads, ambiguous bases, and sequencing artifacts M ´ edigue et al. (1999); Meiklejohn et al. (2019), and field-collected specimen images are often degraded by cluttered back- grounds, occlusions, lighting variation, motion blur Nguyen et al. (2024), or lo w signal-to-noise acquisition. Bridging this gap between curated ev aluation and imperfect operational inputs makes robust taxonomic inference from heterogeneous and noisy modalities a central and unresolv ed chal- lenge in applied biodiv ersity science. A ke y recent step tow ard unifying these modalities is CLIBD Gong et al. (2024), which to our knowledge is the only prior work that explicitly aligns images, DNA bar codes, and taxonomic text in a shared embedding space at very larg e scale , and is therefore our primary state-of-the-art ref- erence. In practice, ho wev er, cross-modal retrie val typically surfaces candidate matches that still require downstream v erification by human experts, whereas operational pipelines ultimately benefit from reliable taxonomic pr ediction from one or more av ailable modalities. More importantly , ex- isting multimodal objecti ves commonly treat taxonomy as a flat label space and rely on standard contrastiv e learning without explicitly encoding biological hierarchy . As a result, learned embed- dings may lack hierarchy-consistent geometry: closely related taxa are not guaranteed to be nearby , and perturbations from noise or missing data can lead to unpredictable errors across taxonomic ∗ This author contributed to this w ork while at Brookhaven National Laboratory . 1 Published as a conference paper at ICLR 2026 Figure 1: Effect of hierarchical r egularization on embedding geometry and noise rob ustness (CLIBD-HiR, variant 1). Left: without the HiR loss, standard contrastiv e training treats mis- matched taxa uniformly , yielding no explicit geometric relationship between intra-genus distances ( d 1 ; different species within the same genus) and inter-genus / higher-le vel distances ( d 2 , d 3 ). Un- der realistic noise, a perturbed query embedding may drift across arbitrary clusters, leading to errors that can propagate to higher taxonomic ranks. Right: with HiR, the loss explicitly enforces a hierarchy-consistent structure ( d 1 < d 2 < d 3 ), so nearby neighborhoods reflect taxonomic proxim- ity . Consequently , even when noise causes a species-level mistake, predictions are more likely to remain correct at coarser lev els (genus/family/order), improving rob ustness. lev els. This issue is particularly acute when one modality—most commonly DNA—is partially corrupted or una vailable, a routine scenario in large-scale biodi versity repositories. Finally , most existing approaches do not explicitly model adaptive image–DN A fusion , even though DNA alone may be insufficient in practice and complementary morphological cues from images can be critical for resolving fine-grained taxa Cong et al. (2017). In this work, we b uild on CLIBD Gong et al. (2024) and present a taxonomy-aw are multimodal framew ork for rob ust taxonomic prediction from images, DN A barcodes, and their combination un- der realistic noise. Our approach is instantiated as two complementary end-to-end variants of CLIBD (Algo 1 and 2) that (i) inject taxonomic hierarchy into representation learning via a hierarchy-a ware objectiv e, and (ii) optionally train an explicit image–DN A fusion predictor for joint inference. Algo 1: CLIBD-HiR (structured, noise-rob ust repr esentation learning). Our first v ariant targets a core failure mode of prior contrasti ve methods: the lack of hierarchy-consistent structure in the learned representation space. W e propose Hierar chical Inf ormation Regularization (HiR) , which injects taxonomic hierarchy directly into representation learning and explicitly shapes the embed- ding geometry during training. Samples sharing coarser taxa (e.g., family or genus) are encouraged to remain close, while finer distinctions (e.g., species) are learned without collapsing higher-le vel neighborhoods. This hierarchical organization acts as an intrinsic noise stabilizer: when a noisy sample drifts a way from its species cluster due to image corruption or barcode degradation, HiR still anchors it to the correct higher-le vel neighborhood, limiting catastrophic semantic drift (Fig. 1). Algo 2: CLIBD-HiR-Fuse (adaptive fusion for variable modality quality). Our second variant extends CLIBD-HiR with a lightweight fusion predictor trained jointly with the encoders. This is motiv ated by deployment realities where av ailable evidence varies by sample—some specimens hav e only images, some only barcodes, and some both, often with differing degrees of corruption. CLIBD-HiR-Fuse (Fig. 2) supports image-only , DNA-only , and fused image+DNA inference, and lev erages the hierarchy-aware aligned space to better combine complementary signals when one modality is unreliable. T ogether, hierarchy-guided regularization (Algo 1) and adapti ve fusion (Algo 2) produce noise- resilient multimodal models that better match real-world biodiv ersity workflo ws. Across large-scale benchmarks, our approach improves taxonomic prediction accuracy over CLIBD and strong fusion baselines, with particularly large gains under DN A corruption and low-quality imaging. 2 Published as a conference paper at ICLR 2026 Our contributions ar e threef old: • W e introduce Hierarchical Information Regularization (HiR) , a taxonomy-a ware objectiv e that explicitly shapes embedding geometry and improves robustness to noisy and partially corrupted in- puts. • W e present two end-to-end variants: CLIBD-HiR (Algo 1), a structured embedding learner opti- mized for hierarchical taxonomic prediction, and CLIBD-HiR-Fuse (Algo 2), which adds an adap- tiv e fusion predictor supporting image-only , DNA-only , and image+DN A inference under varying modality quality . • W e demonstrate consistent improvements in taxonomic prediction across biodiv ersity benchmarks, with especially large gains in noise-dominated re gimes. 2 R E L A T E D W O R K Foundation models f or biodiversity and multimodal alignment. Recent progress in foundation models has enabled transferable representations via large-scale pretraining and multimodal align- ment. CLIP-style training aligns vision and language to support zero-shot transfer (Radford et al., 2021; Cherti et al., 2023), and ne wer multimodal alignment frame works extend this idea beyond image–text to jointly embed heterogeneous modalities. For example, ImageBind learns a shared space across multiple sensory streams (e.g., image, text, audio, depth, IMU) using paired data (Girdhar et al., 2023), and related ef forts such as “X-CLIP”/“LanguageBind”-style models extend CLIP-like alignment to additional modalities. In biodiv ersity , BioCLIP adapts CLIP-style pretrain- ing to ward organism-centric visual recognition (Stev ens et al., 2024), while CLIBD is a key step tow ard multimodal biodiv ersity foundation modeling by aligning specimen images, DNA barcodes, and taxonomic text in a shared embedding space (Gong et al., 2024; Gharaee et al., 2023). These backbones provide strong representations, but are typically optimized for retriev al-style alignment and do not explicitly enforce hierarchy-consistent geometry or rob ustness to modality degradation. T axonomy-aware r epresentation learning. T axonomic labels are inherently hierarchical (order – family–genus–species), motiv ating objectives that respect coarse-to-fine structure beyond flat clas- sification. Prior work has explored hierarchy-aw are losses and hierarchical contrasti ve learning to impose semantic structure directly in the embedding space (Khosla et al., 2020; Zhang et al., 2022). Our HiR regularization builds on this line by injecting taxonomic hierarchy into multimodal align- ment, shaping neighborhoods so that nearby embeddings reflect biological relatedness and improv- ing robustness when fine-grained cues are noisy or incomplete. Multimodal fusion under noisy modalities. Beyond alignment, practical biodiversity applica- tions often benefit from combining complementary e vidence across modalities. Recent fusion strate- gies include uncertainty-aware fusion that explicitly improves robustness to noisy unimodal repre- sentations (Gao et al., 2024), as well as dynamic routing / mixture-of-experts style fusion with learned gating that adapts fusion beha vior to the input (Cao et al., 2023; Han et al., 2024). W e build on these ideas with a lightweight gated fusion head that adaptively mixes image and DN A embeddings, and ev aluate it against naive a veraging under clean and degraded modality conditions. Robust modeling of barcodes and field imagery . DN A barcodes encountered in operational pipelines can be imperfect (e.g., substitutions/indels, ambiguous bases, partial reads), motiv ating noise-aware preprocessing and modeling M ´ edigue et al. (1999); similarly , field imagery is affected by background clutter , occlusion, illumination/pose changes, and motion blur , which can degrade fine-grained recognition Nguyen et al. (2024). Our work is complementary: rather than relying on noise-specific training data, we stabilize multimodal prediction by enforcing hierarchy-consistent geometry and enabling adapti ve fusion to lev erage complementary evidence when one modality is degraded. 3 M E T H O D W e consider triplets ( x V i , x D i , x T i ) consisting of an image, a DNA barcode, and a textual taxonomy description for specimen i . Our goal is twofold: (1) learn a shared multimodal embedding space 3 Published as a conference paper at ICLR 2026 where visual, DN A, and text representations are aligned in a hierarchy-a ware, noise-robust manner; and (2) learn a fusion network that directly predicts taxonomy from jointly using image and DNA features. W e denote the encoders by v i = f V ( x V i ) , d i = f D ( x D i ) , t i = f T ( x T i ) , where v i , d i , t i ∈ R d are ℓ 2 -normalized embeddings. Each specimen is also annotated with hierar- chical taxonomic labels y i = y (1) i , y (2) i , . . . , y ( L ) i , e.g., order , family , genus, species for L = 4 . Our framework is trained in two stages: (1) multimodal pretraining with symmetric cross-modal contrastiv e losses plus an image-only Hierarchical Information Regularization (HiR) loss; and (2) post-hoc training of an MLP fusion head that predicts taxonomy from visual and DN A embeddings. 3 . 1 C RO S S - M O D A L C O N T R A S T I V E O B J E C T I V E S Follo wing CLIBD Gong et al. (2024), we align modalities with symmetric InfoNCE. For a batch of size N , define s ( a i , b j ) = a ⊤ i b j /τ . The directed loss is L A → B = − 1 N N X i =1 log exp( s ( a i , b i )) P N j =1 exp( s ( a i , b j )) , (1) and L A ↔ B = L A → B + L B → A . W e use pairs ( V , T ) , ( V , D ) , and ( D , T ) , and combine them as L XMOD = λ V T L V ↔ T + λ V D L V ↔ D + λ DT L D ↔ T . (2) 3 . 2 H I E R A R C H I C A L I N F O R M AT I O N R E G U L A R I Z A T I O N F O R I M AG E S While the cross-modal objecti ve encourages alignment between modalities, it does not explicitly encode the taxonomy hierarchy , nor does it guarantee robustness when one modality is noisy or par - tially corrupted. T o address this, we introduce an image-only hierarchical contrastiv e loss inspired by the HiConE objectiv e from Zhang et al. (2022). For each taxonomic lev el ℓ ∈ { 1 , . . . , L } (e.g., order , family , genus, species), we treat images sharing the same label y ( ℓ ) i as positi ves at level ℓ . Let P ( ℓ ) i be the set of indices j = i such that y ( ℓ ) j = y ( ℓ ) i , and let N ( ℓ ) i be the remaining images in the batch. For an anchor i and a positiv e j ∈ P ( ℓ ) i , we define the pair-wise supervised contrasti ve loss at lev el ℓ as ℓ ( ℓ ) ( i, j ) = − log exp s ( v i , v j ) P k ∈P ( ℓ ) i ∪N ( ℓ ) i exp s ( v i , v k ) . (3) The standard lev el- ℓ supervised contrasti ve loss is then L ( ℓ ) SupCon = 1 N N X i =1 1 |P ( ℓ ) i | X j ∈P ( ℓ ) i ℓ ( ℓ ) ( i, j ) . (4) Follo wing HiConE, we further enforce that finer taxonomic le vels cannot be optimized in a way that violates coarser-le vel structure. T o this end, we compute, for each lev el ℓ , the maximum pair-wise loss ov er all positiv e pairs at that level: m ( ℓ ) = max i max j ∈P ( ℓ ) i ℓ ( ℓ ) ( i, j ) , (5) and define a hierar chically r ectified pair loss ˜ ℓ (1) ( i, j ) = ℓ (1) ( i, j ) , ˜ ℓ ( ℓ ) ( i, j ) = max ℓ ( ℓ ) ( i, j ) , m ( ℓ − 1) for ℓ > 1 . (6) 4 Published as a conference paper at ICLR 2026 Figure 2: CLIBD-HiR-Fuse framework (Algorithm variant 2). Giv en a specimen image and its DN A barcode, we encode each modality with an image encoder and a DNA encoder , and embed the taxonomy prompt with a frozen BioCLIP text encoder . W e align image–text and DN A–text representations using CLIP-style contrasti ve learning, and enforce hierarchy-aware structure with a hierarchical loss over augmented image views. A lightweight GatedFusion module adaptively combines image and DN A embeddings into a fused representation, which is additionally aligned to the fixed te xt embedding space via a fused-to-text contrasti ve objectiv e. Intuitiv ely , this loss prev ents the model from minimizing fine-grained losses while coarser-le vel structure is still poorly organized. Concretely , if the loss of a fine-lev el positive pair (e.g., same species) becomes smaller than the lar gest loss among positiv es at the immediately coarser lev el (e.g., same genus), we clamp the fine-le vel loss to that coarser-le vel maximum. This ensures that fine-lev el positiv es are not allowed to be optimized “ahead” of the coarser lev el: the model must first reduce the worst-case within-genus (or within-family) positi ve loss before further tightening species-level clusters. Our final Hierarchical Information Regularization (HiR) loss aggregates these rectified pair losses across lev els: L HiR = L X ℓ =1 α ℓ 1 N N X i =1 1 |P ( ℓ ) i | X j ∈P ( ℓ ) i ˜ ℓ ( ℓ ) ( i, j ) , (7) where α ℓ are non-negati ve weights (we use uniform weights in our experiments). This hierarchical structure makes the visual encoder noise-r obust : if an embedding is perturbed such that it drifts away from its species cluster, the loss still anchors it using genus and family supervi- sion, and the max-rectification in Eq. 6 prev ents the optimizer from o verfitting to noisy fine-grained labels while ignoring coarser , more reliable signals. As a result, e ven under noisy supervision or mis- matched modalities, the image representation preserves higher -lev el semantic consistency (Fig. 1). 3 . 3 O B J E C T I V E S : T W O E N D - T O - E N D V A R I A N T S ( A L G O 1 V S . A L G O 2 ) W e instantiate our framework in two variants that share the same multimodal alignment and hierar- chical regularization backbone, b ut differ in whether an e xplicit fusion predictor is trained. Algo 1: CLiBD-HiR (structured, noise-rob ust repr esentation learning). In Algo 1, we train the encoders end-to-end using the sum of cross-modal contrasti ve alignment and hierarchical image regularization: L (1) total = L XMOD + λ HiR L HiR , (8) 5 Published as a conference paper at ICLR 2026 where λ HiR balances hierarchical image re gularization ag ainst multimodal alignment. This objecti ve encourages a hierarchy-consistent embedding geometry , improving robustness when one modality is noisy or partially corrupted, and yielding strong taxonomy classification via nearest-neighbor or linear probing in the learned space. (Full pseudocde in Algorithm 1). Algo 2: CLiBD-HiR-Fuse (flexible fusion with missing-modality support). Algo 2 augments Algo 1 with a lightweight fusion module trained jointly with the encoders under a single objectiv e: L (2) total = L XMOD + λ HiR L HiR + λ fuse L fuse , (9) where λ fuse controls the contribution of fusion loss L fuse (Eq. 11). This variant provides a direct prediction interface that is applicable when only one modality is av ailable (image-only or DNA- only) and when both modalities are a vailable, while remaining noise-resilient due to the shared aligned and hierarchy-aw are representation. (Full pseudocde in Algorithm 2) 3 . 4 F L E X I B L E M U LT I M O D A L F U S I O N F O R T A X O N O M Y P R E D I C T I O N ( A L G O 2 ) Algo 2 introduces a lightweight fusion predictor g ϕ that maps av ailable non-text modality em- beddings to taxonomy logits. Giv en image and DNA embeddings, we concatenate them as z i = [ v i ; d i ] ∈ R 2 d and define the fused representation as ˆ p i = v i (image-only) , d i (DN A-only) , g ϕ ( z i ) (image+DN A) . (10) W e train g ϕ with a supervised cross-entropy loss at le vel ℓ ⋆ , L fuse = − 1 N N X i =1 log ˆ p i y ( ℓ ⋆ ) i . (11) Since the encoders are aligned across modalities and regularized to respect taxonomy , the fusion predictor is more robust when one modality is de graded. Degradation Models. T o assess robustness under realistic perturbations, we introduce degradation strategies for both modalities. For images, we simulate optical blur and defocus by con volving the input x with a normalized averaging kernel h ∈ R k × k , where the kernel size k controls the sev erity of high-frequency attenuation. For DN A, we model sequencing errors and partial reads by transforming a clean sequence s into a corrupted observation ˜ s via a stochastic pipeline comprising fiv e operations: (1) substitution of nucleotides with probability p sub ; (2) ambiguous masking where bases are replaced by ‘N’ with probability p mask ; (3) insertions and deletions ( p ins , p del ) to simulate frameshifts; (4) contiguous dr opout of a subsequence with relativ e length ρ ; and (5) tail truncation ( τ ) to mimic incomplete reads. 4 E X P E R I M E N T S Dataset and split. W e use the BIOSCAN-1M Gharaee et al. (2023) insect dataset and construct paired samples consisting of a specimen image, a COI DNA barcode sequence, and a textual tax- onomy description deri ved from the hierarchical labels (order , family , genus, species). Each sample includes both string taxonomy fields and consistent integer label IDs at each taxonomic lev el. Our final split contains 903,536 training samples and 224,777 test samples. The split is closed-set across all taxonomic le vels: ev ery order , family , genus, and species that appears in the test set also appears in the training set (no unseen taxa in test). T axonomic completeness varies across specimens (man y are labeled only up to coarser levels such as order or family); accordingly , we report ev aluation at each lev el using all av ailable labels for that lev el. 4 . 1 B A S E L I N E S A N D E V A L U A T I O N M E T R I C S W e use CLIBD Gong et al. (2024) as the primary reference and construct all comparisons to isolate the effects of hierarchical regularization and fusion. (1) No-fusion comparison (T able 1). W e report CLIBD and CLIBD-HiR under clean and noisy settings, ev aluating Image → T ext and DNA → T ext T op-1/T op-5 taxonomic classification accuracy without any Image–DNA fusion. (2) Fusion com- parison within CLIBD-HiR (T able 2). Here we keep the same CLIBD-HiR training setup and 6 Published as a conference paper at ICLR 2026 Algorithm 1 CLiBD-HiR: End-to-End Hierarchy-Guided Multimodal Contrasti ve T raining Require: Encoders f V , f D , f T ; temperature τ ; weights λ V T , λ V D , λ DT , λ HiR ; hierarchy weights { α ℓ } L ℓ =1 1: while not con verged do 2: Sample mini-batch { ( x V i , x D i , x T i , y i ) } N i =1 3: for i = 1 to N do 4: v i ← f V ( x V i ) , d i ← f D ( x D i ) , t i ← f T ( x T i ) 5: (Optional) ℓ 2 -normalize v i , d i , t i 6: end for 7: Compute pairwise similarities with temperature τ 8: Compute symmetric CLIP-style losses L V ↔ T , L V ↔ D , L D ↔ T 9: L CLIBD ← λ V T L V ↔ T + λ V D L V ↔ D + λ DT L D ↔ T 10: L HiR ← 0 , m (0) ← 0 11: for ℓ = 1 to L do ▷ taxonomy lev els: order , family , genus, species 12: for i = 1 to N do 13: P ( ℓ ) i ← { j = i | y ( ℓ ) j = y ( ℓ ) i } , N ( ℓ ) i ← { k = i | y ( ℓ ) k = y ( ℓ ) i } 14: end for 15: Compute ℓ ( ℓ ) ( i, j ) using Eq. 3 16: m ( ℓ ) ← max i,j ∈P ( ℓ ) i ℓ ( ℓ ) ( i, j ) 17: if ℓ = 1 then 18: ˜ ℓ ( ℓ ) ( i, j ) ← ℓ ( ℓ ) ( i, j ) 19: else 20: ˜ ℓ ( ℓ ) ( i, j ) ← max ℓ ( ℓ ) ( i, j ) , m ( ℓ − 1) 21: end if 22: L ( ℓ ) HiR ← 1 N P N i =1 1 |P ( ℓ ) i | P j ∈P ( ℓ ) i ˜ ℓ ( ℓ ) ( i, j ) 23: L HiR ← L HiR + α ℓ L ( ℓ ) HiR 24: end for 25: L total ← L CLIBD + λ HiR L HiR 26: Update parameters of f V , f D , f T using ∇L total 27: end while 28: r eturn T rained encoders f V , f D , f T compare three variants: CLIBD-HiR (no fusion; unimodal I → T and D → T), CLIBD-HiR + A vg (na ¨ ıve averaging of image and DN A embeddings for I+D → T), and CLIBD-HiR + Fusion (our learned GatedFusion head for I+D → T). W e e valuate rob ustness under both DNA-only noise ( Noisy D ) and joint image+DN A noise ( Noisy I+D ). This isolates the benefit of adapti ve fusion beyond hierarchy-aw are representation learning. As per the metrics, we report T op-1 and T op-5 taxonomic prediction accuracy at four hierarchical lev els (order , family , genus, species). For each query (im- age, DNA, or fused image+DNA), we rank candidate taxonomy text prompts by similarity in the shared embedding space and measure whether the ground-truth label appears at rank 1 or within the top 5. Global denotes an aggregate accurac y across the four taxonomic lev els. Models and training setup. W e adopt a three-encoder architecture consisting of an image en- coder , a DNA encoder , and a text encoder . The image and text encoders are initialized from a pretrained vision–language model, either standard OpenCLIP V iT -L/14 (Cherti et al., 2023) or Bio- CLIP V iT -L/14 (Stevens et al., 2024). The DN A branch is built upon DNABER T2 (Zhou et al., 2024). When BioCLIP is used, we employ a modified DNABER T2 v ariant by adding a learnable linear projection layer on top of the DN ABER T2 embedding to match the embedding dimension of the image–text backbone. Follo wing Algo. 1 and Algo. 2, we train the image and DN A encoders end-to-end. The optimization strategy for the text encoder depends on the chosen backbone. W ith BioCLIP , we freeze the entire text encoder . W ith standard OpenCLIP , we fine-tune the text en- coder during training. Empirically , freezing BioCLIP text encoder improves performance, likely because BioCLIP already encodes strong biological language priors (Stevens et al., 2024); this dif- fers from the original CLIBD training recipe, which fine-tunes the text encoder (Gong et al., 2024). Unless stated otherwise, we report results using this fixed-BioCLIP-text variant. T raining uses con- trastiv e alignment losses together with a hierarchy-aw are loss on augmented image features (Khosla 7 Published as a conference paper at ICLR 2026 Algorithm 2 CLiBD-HiR-Fuse: End-to-End Multimodal Contrasti ve + HiR + Fusion-Supervised T raining Require: Encoders f V , f D , f T ; fusion head g ϕ ; temperature τ ; weights λ V T , λ V D , λ DT , λ HiR , λ fuse ; hierarchy weights { α ℓ } L ℓ =1 1: while not con verged do 2: Sample mini-batch { ( x V i , x D i , x T i , y i ) } N i =1 3: for i = 1 to N do 4: v i ← f V ( x V i ) , d i ← f D ( x D i ) , t i ← f T ( x T i ) 5: (Optional) ℓ 2 -normalize v i , d i , t i 6: z i ← [ v i ; d i ] ▷ fusion input 7: ˆ p i ← g ϕ ( z i ) ▷ taxonomy logits/probabilities 8: end for 9: Compute similarities with temperature τ 10: Compute symmetric CLIP-style losses L V ↔ T , L V ↔ D , L D ↔ T 11: L CLIBD ← λ V T L V ↔ T + λ V D L V ↔ D + λ DT L D ↔ T 12: Compute hierarchy-guided loss L HiR exactly as in Alg. 1 13: Compute fusion classification loss L fuse (Eq. 11) 14: L total ← L CLIBD + λ HiR L HiR + λ fuse L fuse 15: Update parameters of f V , f D , f T , g ϕ using ∇L total 16: end while 17: r eturn T rained encoders f V , f D , f T and fusion head g ϕ et al., 2020; Zhang et al., 2022). For the fusion v ariant, we add a trainable GatedFusion head, a lightweight 2-layer MLP (Linear–ReLU–Dropout–Linear–Sigmoid) that predicts a per-dimension gate from concatenated image and DN A embeddings and mixes them before normalization, super- vised by an additional fused-to-text contrasti ve loss. Implementation and noise settings. W e train with distributed data parallel on 4 NVIDIA A100 GPUs for ∼ 1 day (batch size 30, 10 epochs). Images are loaded from an HDF5 container and paired with DNA barcodes and taxonomy text from CSV metadata. Optimization uses AdamW with a OneCycle schedule (base LR 1 × 10 − 6 , max LR 5 × 10 − 5 ), with λ HiR = 0 . 99 and α = 0 . 1 in the gathered contrastiv e loss; the fusion variant adds a fused-to-text term weighted by λ fuse = 0 . 7 (omit- ted for the no-fusion baseline). F or robustness ev aluation, we apply inference-time modality degra- dation only (no noisy data during training): DNA is corrupted with substitutions ( p sub =0 . 01 ), inser- tions/deletions ( p ins = p del =0 . 002 ), masking to N ( p mask =0 . 003 ), contiguous N -dropout (run fraction 0 . 05 ), and tail truncation (10%), while images use blur noise with a 7 × 7 kernel. W e report clean, DN A-noisy , and joint image+DNA noisy results. Results. T able 1 shows that adding HiR to CLIBD impro ves no-fusion taxonomic prediction, with the largest gains under noise. For I → T , CLIBD-HiR increases Global T op-1 from 75.5 to 78.2 on clean data and from 40.0 to 46.6 on noisy data (T op-5: 58.4 to 59.6), dri ven mainly by im- prov ed coarse-level accurac y (e.g., noisy Family T op-1: 66.8 to 70.1). For D → T , HiR yields a small clean gain (Global T op-1: 94.8 to 95.6) but a substantial robustness impro vement under noisy DN A (Global T op-1: 52.4 to 66.0; Global T op-5: 91.5 to 96.9), with a notable increase at the family lev el (57.1 to 70.3 T op-1). Overall, HiR consistently improves global performance and noise robust- ness without using fusion. Moreover , T able 2 analyzes fusion within the same CLIBD-HiR training setup. The unimodal baselines (I → T and D → T) establish modality-specific performance, while the third block, I+D → T (A vg) , fuses image and DNA by simple embedding averaging (no fusion mod- ule). Our learned fusion model, I+D → T (Ours) , impro ves over averaging in the most realistic setting where both modalities are noisy: Global accurac y increases from 85.5/96.5 to 88.0/97.5 (T op-1/T op- 5), with the largest gain at species le vel (54.6/79.9 to 57.4/81.7). Under DNA-only noise, a veraging and learned fusion are comparable globally (91.3/97.7 vs. 91.4/98.0), indicating that the main benefit of the fusion module is robustness when image and DN A quality vary simultaneously . Limitations. W e do not use the original CLIBD split, which is designed around seen/unseen ev alu- ation across multiple taxonomic lev els and is highly imbalanced. In our initial experiments, this split substantially reduces the effecti ve supervision av ailable at deeper lev els (genus/species) and makes 8 Published as a conference paper at ICLR 2026 T able 1: Algo 1 (CLIBD-HiR): no-fusion evaluation. Comparison of CLIBD and CLIBD-HiR without any image–DNA fusion module. W e report T op-1 / T op-5 taxonomic prediction accu- racy (%) for Image → T ext and DN A → T ext under clean inputs and under noisy inputs (syntheti- cally degraded at inference). Global denotes an aggregate across order, family , genus, and species. Highlighted rows indicate our proposed HiR model. Setting Method Cond. T op-1 / T op-5 Accuracy Order Family Genus Species Global I → T CLIBD Clean 98.8 / 99.2 93.3 / 95.8 77.8 / 90.4 46.6 / 71.4 75.5 / 89.8 Noisy 88.8 / 92.1 66.8 / 74.7 48.7 / 65.6 22.2 / 42.0 40.0 / 58.4 CLIBD-HiR Clean 99.3 / 99.5 94.3 / 96.1 76.8 / 89.1 46.6 / 68.3 78.2 / 89.2 I → T CLIBD-HiR Noisy 88.7 / 91.4 70.1 / 77.0 42.9 / 59.0 15.5 / 31.8 46.6 / 59.6 D → T CLIBD Clean 99.9 / 99.9 99.4 / 99.8 94.7 / 98.1 68.1 / 88.5 94.8 / 98.3 Noisy 99.7 / 99.9 57.1 / 97.8 81.6 / 94.2 48.6 / 76.3 52.4 / 91.5 CLIBD-HiR Clean 100.0 / 100 99.3 / 99.9 95.9 / 98.8 71.6 / 91.3 95.6 / 98.7 D → T CLIBD-HiR Noisy 99.8 / 99.9 70.3 / 99.2 81.5 / 95.1 51.6 / 79.3 66.0 / 96.9 T able 2: Algo 2 (CLIBD-HiR-Fuse): fusion evaluation. W e report T op-1 / T op-5 taxonomic prediction accurac y (%). Highlighted rows indicate our proposed fusion model. Bold indicates the best performance for that specific condition (Clean, Noisy D, or Noisy I+D) across all methods. Method Cond. T op-1 / T op-5 Accuracy Order F amily Genus Species Global I → T (Ours) Clean 99.3 / 99.5 93.5 / 99.9 74.7 / 98.9 40.4 / 91.8 77.0 / 98.7 Noisy I 88.3 / 91.8 67.0 / 75.9 37.0 / 55.7 11.7 / 26.7 50.2 / 62.1 D → T (Ours) Clean 100.0 / 100 99.5 / 99.9 96.1 / 98.9 74.4 / 92.7 95.7 / 98.7 Noisy D 99.8 / 99.9 87.3 / 99.3 86.1 / 95.6 57.8 / 82.0 82.3 / 97.1 I+D → T (A vg) Clean 99.9 / 100 99.6 / 99.9 95.5 / 98.6 71.2 / 89.8 95.7 / 98.6 Noisy D 99.9 / 100 96.8 / 99.7 90.6 / 97.1 60.1 / 82.9 91.3 / 97.7 Noisy I+D 99.6 / 100 91.3 / 99.3 87.9 / 96.1 54.6 / 79.9 85.5 / 96.5 I+D → T (Ours) Clean 100.0 / 100 99.7 / 99.9 96.0 / 98.9 73.5 / 91.2 96.1 / 98.8 (GatedFusion) Noisy D 100.0 / 100 96.2 / 99.7 88.7 / 96.9 60.8 / 83.2 91.4 / 98.0 Noisy I+D 99.9 / 100 93.0 / 99.5 87.0 / 96.0 57.4 / 81.7 88.0 / 97.5 hierarchy learning less informativ e, leading to performance that is close to the baseline. T o clearly demonstrate the impact of hierarchy-aware re gularization and fusion, we therefore report results on our split where all taxonomic lev els are consistently represented. Developing training and ev alu- ation protocols that better handle the se vere long-tail imbalance and le vel-dependent seen/unseen structure in CLIBD remains an important direction for future work. 5 C O N C L U S I O N W e presented HiR-Fusion, a hierarchy-guided multimodal framew ork for robust taxonomic predic- tion from images, DNA barcodes, and their combination. Our first variant (CLIBD-HiR) improves noise robustness by e xplicitly shaping embedding geometry to respect biological hierarchy , and our second variant (CLIBD-HiR-Fuse) further enhances performance by learning an adaptiv e fusion module that outperforms naiv e averaging, particularly when both modalities are degraded. Across clean and noisy settings, our results show that incorporating taxonomic structure and reliability- aware fusion yields more rob ust and practically useful biodiversity recognition models. 9 Published as a conference paper at ICLR 2026 R E F E R E N C E S Bing Cao, Y iming Sun, Pengfei Zhu, and Qinghua Hu. Multi-modal gated mixture of local-to-global experts for dynamic image fusion. In Pr oceedings of the IEEE/CVF International Conference on Computer V ision (ICCV) , pp. 23555–23564, 2023. Mehdi Cherti, Romain Beaumont, Ross W ightman, Mitchell W ortsman, Gabriel Ilharco, Cade Gor- don, Christoph Schuhmann, Ludwig Schmidt, and Jenia Jitse v . Reproducible scaling laws for contrastiv e language-image learning. In Proceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pp. 2818–2829, 2023. doi: 10.1109/CVPR52729.2023. 00276. Qian Cong, Jinhui Shen, Dominika Borek, Robert K Robbins, Paul A Opler , Zbyszek Otwinowski, and Nick V Grishin. When coi barcodes deceiv e: complete genomes rev eal introgression in hairstreaks. Pr oceedings of the Royal Society B: Biological Sciences , 284(1848):20161735, 2017. Zixian Gao, Xun Jiang, Xing Xu, Fumin Shen, Y ujie Li, and Heng T ao Shen. Embracing unimodal aleatoric uncertainty for robust multimodal fusion. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pp. 26876–26885, 2024. Zahra Gharaee, ZeMing Gong, Nicholas Pellegrino, Iuliia Zarubiie va, Joakim Bruslund Haurum, Scott C. Lowe, Jaclyn T . A. McKeo wn, Chris C. Y . Ho, Joschka McLeod, Y i-Y un C. W ei, Jireh Agda, Sujeev an Ratnasingham, Dirk Steinke, Angel X. Chang, Graham W . T aylor, and Paul Fieguth. A step towards worldwide biodiv ersity assessment: The BIOSCAN-1M insect dataset. In Advances in Neural Information Pr ocessing Systems , volume 36, pp. 43593–43619. Curran Associates, Inc., 2023. Rohit Girdhar , Alaaeldin El-Nouby , Zhuang Liu, Mannat Singh, Kalyan V asude v Alwala, Armand Joulin, and Ishan Misra. Imagebind: One embedding space to bind them all. In Pr oceedings of the IEEE/CVF confer ence on computer vision and pattern r ecognition , pp. 15180–15190, 2023. ZeMing Gong, Austin T . W ang, Xiaoliang Huo, Joakim Bruslund Haurum, Scott C. Lo we, Gra- ham W . T aylor, and Angel X. Chang. CLIBD: Bridging vision and genomics for biodi versity monitoring at scale. arXiv preprint , 2024. doi: 10.48550/arXiv .2405.17537. Xing Han, Huy Nguyen, Carl Harris, Nhat Ho, and Suchi Saria. Fusemoe: Mixture-of-experts trans- formers for fleximodal fusion. In Advances in Neural Information Pr ocessing Systems (NeurIPS) , 2024. Prannay Khosla, Piotr T eterwak, Chen W ang, Aaron Sarna, Y onglong T ian, Phillip Isola, Aaron Maschinot, Ce Liu, and Dilip Krishnan. Supervised contrastive learning. In Advances in Neural Information Pr ocessing Systems , 2020. Claudine M ´ edigue, Matthias Rose, Alain V iari, and Antoine Danchin. Detecting and analyzing dna sequencing errors: to ward a higher quality of the bacillus subtilis genome sequence. Genome r esear ch , 9(11):1116–1127, 1999. Kelly A Meiklejohn, Natalie Damaso, and James M Robertson. Assessment of bold and genbank– their accuracy and reliability for the identification of biological materials. PloS one , 14(6): e0217084, 2019. Thi Thu Thuy Nguyen, Anne C Eichholtzer, Don A Driscoll, Nathan I Semianiw , Dean M Corva, Abbas Z Kouzani, Thanh Thi Nguyen, and Duc Thanh Nguyen. Sawit: A small-sized animal wild image dataset with annotations. Multimedia T ools and Applications , 83(11):34083–34108, 2024. Alec Radford, Jong W ook Kim, Chris Hallacy , Aditya Ramesh, Gabriel Goh, Sandhini Agar- wal, Girish Sastry , Amanda Askell, Pamela Mishkin, Jack Clark, Gretchen Krueger , and Ilya Sutske ver . Learning transferable visual models from natural language supervision. In Proceedings of the 38th International Confer ence on Machine Learning (ICML) , volume 139 of Proceedings of Machine Learning Resear ch , pp. 8748–8763. PMLR, 2021. Sujeev an Ratnasingham and Paul DN Hebert. Bold: The barcode of life data system (http://www . barcodinglife. org). Molecular ecology notes , 7(3):355–364, 2007. 10 Published as a conference paper at ICLR 2026 Samuel Ste vens, Jiaman W u, Matthe w J. Thompson, Elizabeth G. Campolongo, Chan Hee Song, David Edward Carlyn, Li Dong, W asila M. Dahdul, Charles Stew art, T anya Berger -W olf, W ei- Lun Chao, and Y u Su. BioCLIP: A vision foundation model for the tree of life. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , 2024. Grant V an Horn, Oisin Mac Aodha, Y ang Song, Y in Cui, Chen Sun, Alex Shepard, Hartwig Adam, Pietro Perona, and Ser ge Belongie. The inaturalist species classification and detection dataset. In Pr oceedings of the IEEE confer ence on computer vision and pattern r ecognition , pp. 8769–8778, 2018. Shu Zhang, Ran Xu, Caiming Xiong, and Chetan Ramaiah. Use all the labels: A hierarchical multi- label contrastiv e learning framew ork. In Pr oceedings of the IEEE/CVF Conference on Computer V ision and P attern Recognition (CVPR) , 2022. doi: 10.48550/arXi v .2204.13207. Zhihan Zhou, Y anrong Ji, W eijian Li, Pratik Dutta, Ramana V . Davuluri, and Han Liu. DNABER T -2: Efficient foundation model and benchmark for multi-species genomes. In International Confer- ence on Learning Repr esentations (ICLR) , 2024. doi: 10.48550/arXiv .2306.15006. 11
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment