3D-LLDM: Label-Guided 3D Latent Diffusion Model for Improving High-Resolution Synthetic MR Imaging in Hepatic Structure Segmentation
Deep learning and generative models are advancing rapidly, with synthetic data increasingly being integrated into training pipelines for downstream analysis tasks. However, in medical imaging, their adoption remains constrained by the scarcity of reliable annotated datasets. To address this limitation, we propose 3D-LLDM, a label-guided 3D latent diffusion model that generates high-quality synthetic magnetic resonance (MR) volumes with corresponding anatomical segmentation masks. Our approach uses hepatobiliary phase MR images enhanced with the Gd-EOB-DTPA contrast agent to derive structural masks for the liver, portal vein, hepatic vein, and hepatocellular carcinoma, which then guide volumetric synthesis through a ControlNet-based architecture. Trained on 720 real clinical hepatobiliary phase MR scans from Samsung Medical Center, 3D-LLDM achieves a Fréchet Inception Distance (FID) of 28.31, improving over GANs by 70.9% and over state-of-the-art diffusion baselines by 26.7%. When used for data augmentation, the synthetic volumes improve hepatocellular carcinoma segmentation by up to 11.153% Dice score across five CNN architectures.
💡 Research Summary
**
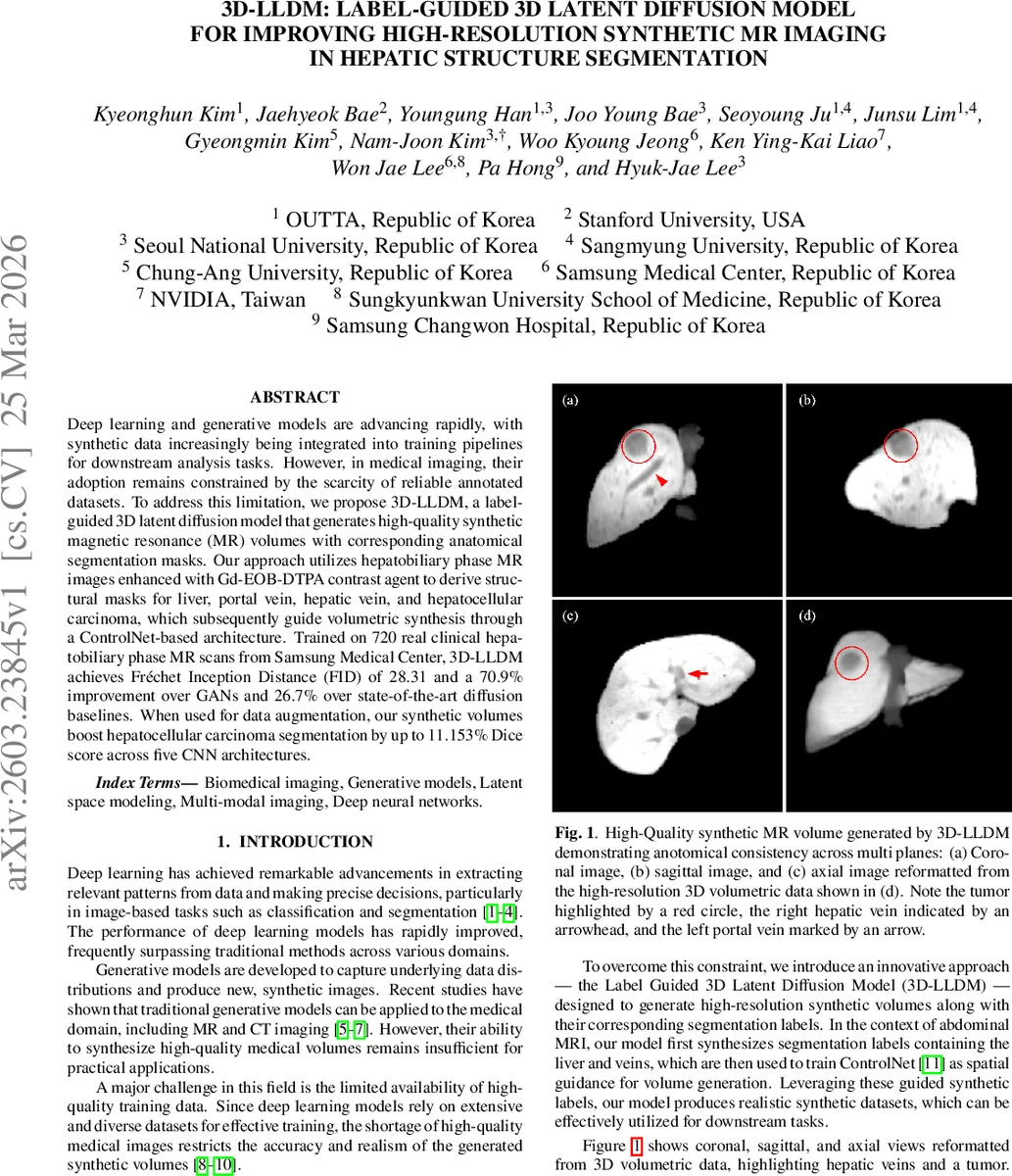
This paper introduces 3D‑LLDM, a label‑guided three‑dimensional latent diffusion model designed to generate high‑resolution synthetic magnetic resonance (MR) volumes together with accurate anatomical segmentation masks for hepatic structures. The authors address the chronic shortage of annotated medical imaging data, which hampers the training of deep learning models for tasks such as organ and tumor segmentation. By leveraging hepatobiliary phase (HBP) MR images enhanced with the Gd‑EOB‑DTPA contrast agent, they obtain ground‑truth masks for the liver, portal vein, hepatic vein, and hepatocellular carcinoma (HCC). These masks serve as spatial guidance for a ControlNet‑augmented diffusion process, ensuring that the generated volumes are anatomically consistent across all three orthogonal planes.
The methodology consists of four main stages. First, a label auto‑encoder (Eₗ, Dₗ) compresses the binary masks into a latent representation zₗ. Second, a separate image auto‑encoder (E, D) encodes real MR volumes into latent space z. Third, a latent diffusion model ε_θ is trained to denoise zₜ while being conditioned on the label latent code via ControlNet, which injects the structural information directly into the diffusion steps. Finally, during inference, random noise is transformed into synthetic labels, which are then fed to the conditioned diffusion model to synthesize a paired MR volume. This pipeline yields perfectly aligned label‑volume pairs without requiring post‑hoc registration.
The model is trained on a sizable internal dataset of 720 HBP MR scans from Samsung Medical Center, each cropped to a fixed size of 160 × 160 × 64 voxels and normalized to
Comments & Academic Discussion
Loading comments...
Leave a Comment