SynLeaF: A Dual-Stage Multimodal Fusion Framework for Synthetic Lethality Prediction Across Pan- and Single-Cancer Contexts
Accurate prediction of synthetic lethality (SL) is important for guiding the development of cancer drugs and therapies. SL prediction faces significant challenges in the effective fusion of heterogeneous multi-source data. Existing multimodal methods…
Authors: Zheming Xing, Siyuan Zhou, Ruinan Wang
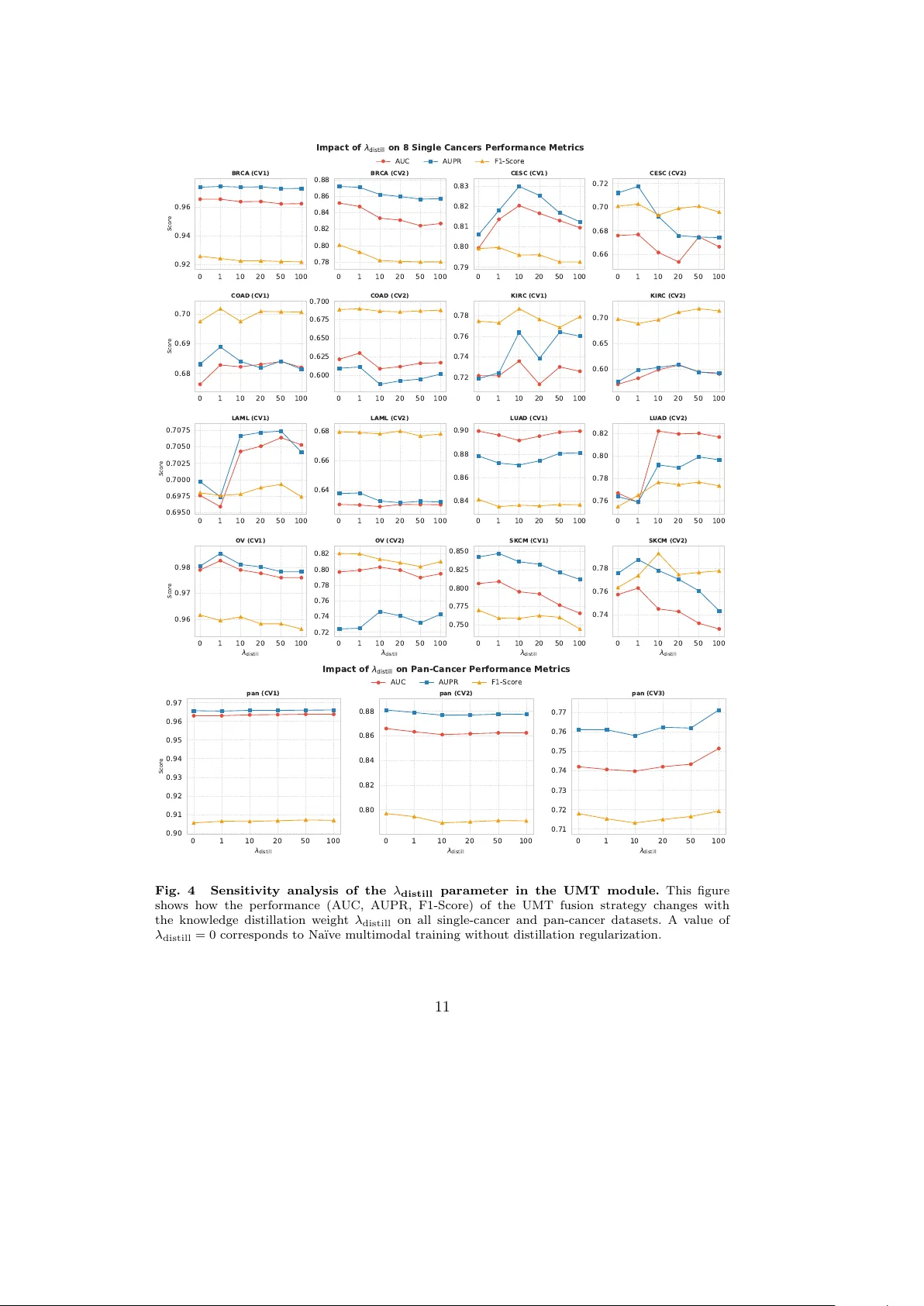
SynLeaF: A Dual-Stage Multimo dal F usion F ramew ork for Syn thetic Lethalit y Prediction Across P an- and Single-Cancer Con texts Zheming Xing 1 , Siyuan Zhou 1 , Ruinan W ang 1 , Rui Han 1 , Shiming Zhang 1 , Shiqu Chen 1 , Y urui Huang 1 , Jiahao Ma 3 , Yifan Chen 4 , Xuan W ang 1 , Y adong W ang 2 , Jun yi Li 1,2* 1* Sc ho ol of Computer Science and T ec hnology , Harbin Institute of T echnology (Shenzhen), Shenzhen, Guang Dong 518055, China. 2 Key Lab oratory of Biological Bigdata, Ministry of Education, Harbin Institute of T echnology , Harbin, Heilongjiang 150001, China. 3 Sc ho ol of Biomedical Sciences, The Universit y of Hong Kong, Hong Kong SAR, China. 4 Departmen ts of Mathematics and Computer Science, Hong Kong Baptist Universit y , Hong Kong SAR, China. *Corresp ondence should be addressed to lijun yi@hit.edu.cn Abstract Accurate prediction of syn thetic lethalit y (SL) is important for guiding the dev el- opmen t of cancer drugs and therapies. SL prediction faces significant challenges in effectiv ely fusing heterogeneous multi-source data. Existing multimodal metho ds often suffer from “mo dalit y laziness” due to disparate conv ergence sp eeds, which hinders the exploitation of complementary information and causes most SL pre- diction mo dels to p erform p oorly for b oth pan-cancer and single-cancer SL pair predictions. In this study , we prop ose SynLeaF, a dual-stage m ultimo dal fusion framew ork for SL prediction in pan-cancer and single-cancer contexts. The frame- w ork employs a V AE-based cross-enco der with a Pro duct of Exp erts mec hanism to fuse four omics data t ypes (gene expression, mutation, methylation, and CNV), sim ultaneously utilizing a relational graph conv olutional netw ork to capture structured gene representations from biomedical knowledge graphs. T o mitigate mo dalit y laziness, SynLeaF in tro duces a dual-stage training mec hanism that emplo ys a feature-level kno wledge distillation. In extensiv e exp erimen ts across eigh t sp ecific cancer types and a pan-cancer dataset, SynLeaF achiev ed sup erior 1 p erformance in 17 of the 19 scenarios. Ablation studies and gradient analyses further v alidate the critical contributions of the prop osed fusion and distillation mec hanisms for mo del robustness and generalization. T o facilitate communit y use, a web server is av ailable at https://synleaf.bioinformatics- lilab.cn . Keyw ords: Synthetic Lethality , Cancer Specific Prediction, Multimo dal Learning, V ariational Autoenco der, Knowledge Distill ation 1 In tro duction Syn thetic Lethality (SL) characterizes a sp ecific genetic relationship wherein the defi- ciency of an individual gene remains viable for the cell, whereas concurren t impairmen t or inactiv ation of a gene pair results in cell death [ 1 ]. As a promising targeted an ti- cancer therapy , SL can eliminate malignant cells while preserving health y tissues [ 2 ], and expands druggable targets for genes that are difficult to target directly [ 3 ]. A clas- sic clinical success is the P ARP inhibitors, approv ed by the FDA in 2014 for o v arian cancer with BR CA1/2 mutations [ 4 , 5 ]. Recently , computationally designed aptamers w ere shown to induce SL b y blo cking the RAD51–BRCA2 interaction [ 6 ]. ADAR1 w as also identified as a new SL target in BR CA-mutan t tumors, where its inhibition acti- v ates innate immunit y via auto crine in terferon p oisoning [ 7 ]. These adv ances contin ue to expand druggable targets and accelerate their translation into therapies [ 8 ]. Despite this p oten tial, iden tifying clinically relev ant SL pairs remains a c hallenge [ 9 ]. W et-lab screens, including y east, RNAi, and CRISPR, are accurate but costly , time-consuming, and suffer from significan t off-target effects [ 10 , 11 ]. The sheer v ol- ume of p ossible gene pairs makes exhaustive screening infeasible [ 12 ]. T o ov ercome the limitations of wet-lab methods, computational methods, which can b e group ed in to statistics-based, netw ork-based, traditional machine learning, and deep learning metho ds, hav e emerged as effective complements [ 3 ]. Statistics and net work approaches for synthetic lethal prediction rely on h yp othe- ses and domain kno wledge and often ignore other data types such as sequences or functional attributes [ 13 ]. T raditional machine learning methods in tegrate m ulti- source data but still dep end on feature engineering, which can in tro duce noise [ 14 , 15 ]. T o address these limitations, deep learning metho ds hav e b een introduced to automatically learn complex representations. Deep learning metho ds learn complex represen tations automatically . Early net w ork represen tation mo dels such as GRSMF and SL2MF use matrix factorization [ 16 , 17 ]. GRSMF introduces graph-regularized self-represen tation to alleviate data sparsity , and SL2MF uses logistic matrix fac- torization with differen t w eights for known and unknown pairs. How ever, matrix factorization metho ds are essentially a form of shallo w embedding and may not fully exploit the structural information and no de features of the netw ork [ 18 ]. The application of Graph Neural Netw orks (GNN) has significantly enhanced the accuracy of predicting syn thetic lethalit y . DDGCN [ 19 ] uses a dual-dropout strat- egy to address sparsity and o verfitting, and GCA TSL [ 20 ] applies dual atten tion at the no de and feature levels. Ho wev er, these GNN-based methods mainly rely on 2 homogeneous netw orks, which only contain gene no des and p ossess limited expres- siv e p o wer. The incorp oration of Knowledge Graphs (K G) has further broadened the use of deep learning techniques for syn thetic lethality prediction. Basically , a K G represen ts a heterogeneous net work that con tains multiple t yp es of en tities (such as genes, pathw ays, diseases, etc.) and relations, that can pro vide richer biological back- ground information [ 21 ]. K G4SL [ 22 ] automatically generates gene features using a kno wledge graph conv olutional netw ork. PiLSL [ 23 ] adopts a gene pair-based metho d, extracting enclosing subgraphs from the kno wledge graph to capture pairwise in ter- actions betw een gene pairs. SLGNN [ 24 ] mo dels relation com binations as factors and impro ves the interpretabilit y of the mo del through no de- and factor-level attention mec hanisms. KR4SL [ 25 ] introduces a reasoning metho d based on paths, utilizing rela- tional digraphs to extract structural seman tic information within the knowledge graph. Mean while, MP ASL [ 26 ] prop osed a hybrid in teraction framew ork in volving gene- en tity and en tity-en tity relationships to improv e gene representations from v arious viewp oin ts. Although kno wledge graphs provide rich structured biological knowledge, KGs solely from a single mo dalit y often cannot fully capture the complex mec hanisms of syn thetic lethality . Consequently , fusing data from multiple sources has emerged as a crucial approach to enhance the accuracy of synthetic lethalit y prediction. PTGNN [ 27 ] uses Conv olutional Neural Netw ork (CNN) features from protein sequences and a graph reconstruction pre-training task on Protein-Protein Interaction and Gene Ontol- ogy (GO) graphs. PiLSL [ 23 ] in tegrates explicit omics features with KG embeddings. T ARSL [ 28 ] applies non-negative matrix tri-factorization with triple atten tion, and Struct2SL [ 29 ] combines protein sequences, protein 3D structures, and PPI net works. Although these methods improv e prediction accuracy b y in tro ducing m ulti-source information, they face a common and critical limitation, as in pure K G metho ds, the neglect of c ontext-sp e cificity . The realization of generalized syn thetic lethal effects is often imp eded by sp ecific tumor-asso ciated factors, including cellular heterogeneity , metab olic status, and the complexities of the tumor microenvironmen t [ 30 ]. Man y syn thetic lethal in teractions are observ ed in only a few specific cancers. An in vestigation utilizing CRISPR-Cas9 screening to pinp oin t synthetic lethal gene pairs across three cell lines revealed a min- imal intersection: merely 10% of the in teractions w ere shared b et ween any t wo lines, while no single pair w as consistently observed across all three [ 31 , 32 ]. Adequately addressing tumor heterogeneity can mitigate certain challenges associated with the translation of syn thetic lethalit y strategies from cellular mo dels to in vivo systems and, ultimately , clinical applications [ 33 ]. Context-specific synthetic lethal effects hav e gar- nered significan t interest within the medicinal chemistry communit y , as inv estigating cancer-sp ecific synthetic lethalit y offers nov el a ven ues for pharmaceutical dev elop- men t. Although several studies ha ve attempted to address the sp ecificity problem, they still exhibit significant shortcomings. While ELISL [ 34 ] pioneered the in tegra- tion of context-free protein sequence asso ciations with context-specific omics data, its reliance on shallow models, suc h as random forests, limits its ability to capture the non-linear relationships in high-dimensional data. Based on cancer-sp ecific p ositive 3 syn thetic lethalit y datasets, SLGNNCT [ 35 ] divides the knowledge graph in to differ- en t subgraphs and p erforms cancer-sp ecific syn thetic lethality prediction on the small kno wledge graphs separately via factor attention modeling, such as SLGNN. Ho wev er, the resulting kno wledge subgraphs ha ve v ery few no des, and the small graph size leads to p o or generalization p erformance in deep learning mo dels. T o address these obstacles, w e propose SynLeaF, a dual-stage m ultimo dal fusion framew ork for SL prediction across pan- and single-cancer contexts. The method com- bines a Cross-V AE with Product-of-Exp erts early fusion for four omics mo dalities and an RGCN-based K G enco der, and introduces an adaptive t wo-stage fusion/training paradigm (Uni-Mo dal T eacher, UMT; Uni-Modal Ensem ble, UME) to mitigate mo dal- it y laziness [ 36 – 39 ]. In extensive exp erimen ts co vering eight sp ecific cancer types and a pan-cancer dataset, SynLeaF demonstrated excellen t generalization capabilit y and robustness, surpassing existing state-of-the-art techniques across the ma jorit y of core ev aluation metrics. 2 Results 2.1 SynLeaF: A Dual-Stage Multimo dal F usion F ramework As sho wn in Figure 1 , SynLeaF is a dual-stage m ultimo dal fusion framew ork, designed for the prediction of cancer synthetic lethalit y by integrating heterogeneous data from m ultiple sources. The mo del accepts tw o streams of input: first, four t yp es of omics data, including gene expression, m utation, DNA methylation, and cop y n umber v ari- ation (CNV) 1 ; and second, a biomedical K G that contains v arious en tities such as genes, pathw a ys, and diseases, along with their relationships. The omics data are enco ded through a V AE-based early fusion mo dule, while the KG is processed by an R GCN to extract structured representations. The high-dimensional nature and significant mo dal heterogeneit y of omics data hinder the effectiv e fusion of m ulti-source information. Dra wing inspiration from the framew ork established b y W ang et al. [ 40 ], we emplo y ed a V ariational Auto enco der (V AE) architecture [ 36 ] and innov atively designed a Cross-V AE-based early fusion mo dule. As sho wn in the Omics Enco der module of Figure 1 , unlik e traditional feature concatenation, our metho d constructs an N × N enco der matrix, which skillfully bal- ances in tra-mo dal feature self-learning (via diagonal auto encoders) with inter-modal in teractive inference (via off-diagonal cross-enco ders). By in tro ducing the Pro duct of Exp erts (PoE) mechanism [ 37 ] to aggregate multi-source information, SynLeaF can generate unified and robust gene representations in the latent space. Notably , this architecture naturally addresses the problem of missing data through its cross- inference mechanism. Ev en if some omics data are unav ailable, the mo del can still reconstruct the missing features based on other mo dalities. This greatly improv es the mo del’s generalization abilit y when dealing with fragmented clinical data. In m ultimo dal learning, features are generally categorized in to tw o distinct classes. The former comprises unimo dal attributes acquirable via isolated training, whereas the latter consists of paired characteristics that necessitate cross-mo dal in teraction 1 Due to data source limitations, the pan-cancer dataset uses only three types of omics and does not include DNA methylation data. 4 cn v ex p m y l m u t cn v ex p m y l m u t Po E Mean L = 2 L = 1 R GC N Om ics E n co d er KG E n co d er 𝜇 UM T 🧊 🧊 K G D a t a O mics Da t a 🧊 🧊 🧊 : F r o ze n wh en u s i n g UM T /UME SynLe aF F ramewo rk c n v e x p m y l m u t (Ge n e p a ir , ) (En ti ty p a ir , ) UM E Fig. 1 Ov erview of the SynLeaF F ramework . SynLeaF constitutes a dual-stage m ultimo dal integration architecture designed for syn thetic lethalit y prediction, taking cancer-sp ecific omics profiles and biomedical knowledge graphs as inputs. The Omics Enco der uses a cross-encoder archi- tecture based on a V ariational Autoenco der (V AE) and p erforms early fusion on four t yp es of omics data: copy num b er v ariation (cn v), gene expression (exp), DNA meth ylation (m yl), and m utation (mut), through a Pro duct of Exp erts (PoE) mechanism. The Knowledge Graph Enco der utilizes a Relational Graph Conv olutional Netw ork (RGCN) to extract structural features asso ciated with genes within the biological netw ork. With these enco ders, SynLeaF first indep enden tly pre-trains tw o unimodal mo dels and then constructs tw o base estimators respectively under tw o complemen tary fusion strategies in further training. F or b oth the UMT and UME strategies, the parameters of the pre-trained unimodal enco ders are strictly frozen. Sp ecifically , under the UMT strategy , SynLeaF treats the pre-trained unimo dal enco ders as teac hers, guiding the training of a m ultimo dal student model through feature-level kno wledge distillation. Under the UME strategy , SynLeaF directly inte- grates the prediction results ( p o and p k ) from the tw o pre-trained unimo dal mo dels. Here, p o and p k denote the predicted probabilities from the Omics and Knowledge Graph models, respectively . Finally , SynLeaF adaptively selects the optimal strategy b et ween UMT and UME according to observed v al- idation efficacy . for extraction. Optimally , the ob jective is for a multimodal model to capture paired features via cross -modal mechanisms, while ensuring that it also learns sufficient uni- mo dal features. How ever, Du et al. [ 39 ] found that traditional multimodal late-fusion training metho ds suffer from the problem of “Mo dalit y Laziness”. This means that enco ders trained join tly on multiple m odalities p erform worse on unimo dal feature learning than enco ders trained unimodally , and this phenomenon is particularly evi- den t in tasks where unimo dal priors are meaningful. T o mitigate this problem, Du et al. [ 39 ] prop osed tw o complemen tary multimodal fusion strategie s, designated as 5 Uni-Mo dal T eacher (UMT) and Uni-Modal Ensem ble (UME), and achiev ed go od p erformance in m ultimo dal audio-visual classification tasks. The UMT strategy uses pre-trained unimo dal enco ders as “teachers” to guide a multimodal “student” model to learn the teac hers’ feature representations through feature-level Knowledge Distil- lation. The UME strategy , on the other hand, directly in tegrates the prediction results of the tw o pre-trained unimo dal mo dels. Inspired by the UMT/UME framework, this pap er prop oses a dual-stage train- ing strategy , which is adapted and extended for the characteristics of the omics-graph bimo dal setting to ensure the slo w mo dalit y is fully trained. During the initial stage, w e conduct independent pre-training for b oth the omics enco der and the kno wledge graph enco der to ensure that each unimo dal enco der can fully learn the feature rep- resen tations within its modality . During the second stage, considering the significant differences in inter-modal interactions across different cancer types and data split- ting strategies, we designed an adaptive selection mec hanism. After training, the mo del automatically selects the b est strategy betw een UMT and UME based on the Area Under the R OC Curv e (A UC) metric obtained from the v alidation dataset. UMT usually p erforms better when the unimodal features from b oth omics and the kno wledge graph are strong, and the cross-modal interaction provides additional infor- mation. How ever, when one mo dalit y is clearly dominant or the cross-mo dal interaction in tro duces noise, the simple ensemble strategy of UME is more effectiv e. 2.2 SynLeaF Surpasses Current State-of-the-Art T ec hniques in Both Pan-Cancer and Cancer-Sp ecific Settings T o ev aluate the effectiv eness and robustness of SynLeaF, we conducted a comprehen- siv e comparison with four state-of-the-art metho ds on a collection of datasets, which includes eigh t cancer-sp ecific datasets and one large pan-cancer dataset, under v arious splitting settings. The eigh t sp ecific cancer t yp es are: breast cancer (BR CA), cervi- cal cancer (CESC), colon cancer (COAD), kidney renal clear cell carcinoma (KIRC), acute m yeloid leuk emia (LAML), lung adeno carcinoma (LUAD), o v arian cancer (O V), and skin cutaneous melanoma (SK CM). These settings include CV1 (Random Split), CV2 (Semi-New Gene Split), and CV3 (New Gene Split) [ 23 ]. The four state-of-the-art metho ds compared w ere SLGNN, ELISL, PTGNN, and MP ASL. T o guarantee a fair ev aluation, we standardized the data loading part for all base- line metho ds to mak e sure they all used the same datasets, including the SL gene pairs, protein sequences, omics features, and knowledge graph data. The implementation details for baselines and all training configurations are describ ed in the Supplemen- tary Material (see Exp erimental Design ). Empirical outcomes indicate that SynLeaF exhibits a distinct p erformance edge. On the core comparison metrics across a total of 19 scenarios, SynLeaF ac hieved state-of-the-art (SOT A) results in 17 of these instances. T able 1 details the comparativ e outcomes across cancer-sp ecific datasets, where we observ e that SynLeaF has a significan t lead in most cancer types. Notably , regarding the SKCM dataset, when ev aluated against the second-b est mo del, SynLeaF achiev ed h uge impro vemen ts of 17.71% and 6.89% under the CV1 and CV2 settings, resp ec- tiv ely . Since the ELISL metho d relies on cancer-specific clinical omics data and cell 6 T able 1 Exp eriment results on cancer-sp ecific datasets. The standard deviations of the readings are reported in parentheses. The best p erforming method is highlighted in bold and the second best is underlined. The last column indicates the improv ement made b y SynLeaF. Cancer Split Metric SLGNN ELISL PTGNN MP ASL SynLeaF ↑ (%) BRCA CV1 AUC 0.8847(0.0130) 0.9041(0.0104) 0.9453(0.0093) 0.8187(0.0173) 0.9654(0.0038) 2.13 AUPR 0.8949(0.0169) 0.9196(0.0079) 0.9600(0.0084) 0.8508(0.0118) 0.9743(0.0020) 1.49 CV2 AUC 0.7147(0.1036) 0.7615(0.0470) 0.8153(0.0911) 0.6660(0.0701) 0.8474(0.0502) 3.94 AUPR 0.7358(0.1114) 0.7894(0.0664) 0.8316(0.0973) 0.6888(0.0763) 0.8707(0.0563) 4.70 CESC CV1 AUC 0.6251(0.0795) 0.7136(0.0797) 0.7765(0.0480) 0.7321(0.0946) 0.8136(0.0505) 4.78 AUPR 0.6543(0.0846) 0.7466(0.0484) 0.7915(0.0621) 0.7299(0.0847) 0.8180(0.0650) 3.35 CV2 AUC 0.5957(0.0489) 0.5280(0.0374) 0.6834(0.0948) 0.6297(0.0750) 0.6845(0.0444) 0.16 AUPR 0.5834(0.0443) 0.5753(0.0384) 0.6946(0.0967) 0.6412(0.0272) 0.7190(0.0530) 3.51 COAD CV1 AUC 0.5933(0.0291) 0.6894(0.0150) 0.6212(0.0498) 0.6278(0.0172) 0.7162(0.0263) 3.89 AUPR 0.5979(0.0416) 0.6645(0.0382) 0.6071(0.0592) 0.6535(0.0257) 0.7088(0.0287) 6.67 CV2 AUC 0.5404(0.0237) 0.6206(0.0391) 0.5157(0.0364) 0.5702(0.0385) 0.6317(0.0244) 1.79 AUPR 0.5397(0.0280) 0.6064(0.0316) 0.5181(0.0279) 0.6031(0.0397) 0.6120(0.0316) 0.92 KIRC CV1 AUC 0.6459(0.1381) 0.6754(0.0886) 0.7161(0.0957) 0.6739(0.0601) 0.6940(0.0990) - AUPR 0.6668(0.1067) 0.7238(0.0806) 0.7327(0.0908) 0.7059(0.0835) 0.7162(0.0855) - CV2 AUC 0.5417(0.1334) 0.6599(0.1128) 0.6430(0.1319) 0.6317(0.1601) 0.5822(0.0573) - AUPR 0.5647(0.1054) 0.6646(0.1106) 0.6441(0.1123) 0.6191(0.1975) 0.5978(0.0770) - LAML CV1 AUC 0.5793(0.0224) 0.6267(0.0061) 0.6960(0.0303) 0.6306(0.0256) 0.6980(0.0156) 0.29 AUPR 0.5914(0.0247) 0.6310(0.0177) 0.6925(0.0533) 0.6605(0.0151) 0.7002(0.0239) 1.11 CV2 AUC 0.5467(0.0258) 0.5810(0.0218) 0.6188(0.0366) 0.5977(0.0468) 0.6296(0.0195) 1.75 AUPR 0.5648(0.0200) 0.5930(0.0222) 0.6220(0.0518) 0.6297(0.0326) 0.6378(0.0205) 1.29 LUAD CV1 AUC 0.8254(0.0229) 0.8513(0.0295) 0.8865(0.0254) 0.7945(0.0584) 0.9000(0.0259) 1.52 AUPR 0.8372(0.0132) 0.8571(0.0336) 0.8753(0.0356) 0.7951(0.0471) 0.8873(0.0273) 1.37 CV2 AUC 0.5858(0.1520) 0.7465(0.0920) 0.7678(0.0982) 0.6336(0.1427) 0.8161(0.0854) 6.29 AUPR 0.6365(0.1477) 0.7407(0.1208) 0.7676(0.1140) 0.6636(0.1326) 0.7924(0.1110) 3.23 OV CV1 AUC 0.9201(0.0209) 0.7790(0.0683) 0.9824(0.0116) 0.8555(0.0341) 0.9827(0.0144) 0.03 AUPR 0.9426(0.0166) 0.7812(0.0867) 0.9590(0.0384) 0.8325(0.0413) 0.9855(0.0103) 2.76 CV2 AUC 0.5894(0.0828) 0.7087(0.0300) 0.7416(0.0813) 0.6846(0.0601) 0.7990(0.0850) 7.74 AUPR 0.6136(0.0598) 0.6812(0.0202) 0.6694(0.0854) 0.6867(0.0620) 0.7252(0.1149) 5.61 SKCM CV1 AUC 0.6692(0.1494) 0.6871(0.0248) 0.6469(0.0788) 0.5849(0.1209) 0.8088(0.0307) 17.71 AUPR 0.7106(0.1710) 0.7173(0.0451) 0.6815(0.0737) 0.6745(0.0609) 0.8471(0.0436) 18.10 CV2 AUC 0.6777(0.0862) 0.6302(0.1424) 0.7138(0.2257) 0.5760(0.1715) 0.7630(0.1576) 6.89 AUPR 0.7059(0.0661) 0.6216(0.1371) 0.7251(0.2111) 0.6150(0.1551) 0.7876(0.1566) 8.62 T able 2 Comparison of exp erimen tal results on the pan-cancer dataset Cancer Split Metric SLGNN PTGNN MP ASL SynLeaF ↑ (%) pan CV1 A UC 0.9550(0.0021) 0.9315(0.0011) 0.9336(0.0037) 0.9652(0.0012) 1.07 AUPR 0.9616(0.0018) 0.9386(0.0024) 0.9425(0.0033) 0.9669(0.0008) 0.55 F1 0.8964(0.0023) 0.8894(0.0014) 0.8692(0.0041) 0.9099(0.0018) 1.51 CV2 A UC 0.7736(0.0305) 0.7684(0.0657) 0.4900(0.0234) 0.8624(0.0236) 11.48 AUPR 0.8050(0.0255) 0.8104(0.0448) 0.6118(0.0232) 0.8754(0.0226) 8.02 F1 0.6006(0.0394) 0.7189(0.0433) 0.3858(0.1325) 0.7955(0.0246) 10.66 CV3 A UC 0.5757(0.0367) 0.5128(0.0576) 0.5379(0.0469) 0.7407(0.0271) 28.66 AUPR 0.6050(0.0414) 0.5213(0.0494) 0.5416(0.0413) 0.7611(0.0417) 25.80 F1 0.1163(0.0944) 0.6673(0.0012) 0.0000(0.0000) 0.7153(0.0156) 7.19 line omics data, it cannot b e applied to the pan-cancer synthetic lethalit y prediction task. Therefore, it was not included in the comparison. The experimental data confirm that our prop osed method ac hieved the highest performance metrics under all split- ting strategies on the pan-cancer dataset. Particularly in the CV3 (zero-shot) setting, whic h sim ulates the prediction of unknown genes, SynLeaF still ac hiev ed an AUC of 0.7407, represen ting an improv ement of up to 28.66% ov er the second-place SLGNN. PTGNN show ed strong comp etitiveness in the single-cancer exp erimen ts, ranking sec- ond after SynLeaF on most metrics. Ho wev er, this adv antage has its limitations. By lo oking at T able 1 and T able 2 together, we can see that although PTGNN can fit the single distribution of a sp ecific cancer, its p erformance dropp ed significantly when faced with the pan-cancer scenario, which has more complex data and greater distri- bution differences. In contrast, SynLeaF demonstrated an all-around adaptabilit y , as 7 it maintained the b est p erformance across b oth the smaller single-cancer datasets and the larger pan-cancer dataset. Although SynLeaF p erformed excellently on the v ast ma jorit y of datasets, the framew ork failed to secure the leading p osition on the KIRC dataset. W e conducted an in-depth analysis of this phenomenon and found that the core reason is the extremely small sample size (only ab out 120 samples). This caused a serious distri- bution shift betw een the v alidation and test sets, which in turn led to a misjudgment b y the adaptive selection strategy . This statistical deviation caused the v alidation- set-based adaptiv e selection strategy to b e conserv ative. Sp ecifically , the sub-mo dules of SynLeaF actually hav e the p oten tial to reach SOT A on KIRC, esp ecially in the CV1 setting. On the testing partition, the UMT branc h ac hieved an AUC of 0.7218 ( ± 0.0923) and an A UPR of 0.7246 ( ± 0.0737), metrics that align clos ely with the effi- cacy exhibited b y the top-ranked PTGNN. This result indicates that in small-sample and highly heterogeneous cancer datasets, relying solely on v alidation set metrics for mo del selection can b e risky . The result on KIRC rev eals that how to ov ercome distri- bution shift and ev aluation bias in extremely data-sparse scenarios remains a common c hallenge for the entire field of computational biology . 2.3 The Dual-Stage Adaptive F usion Strategy Effectively Addresses Mo dality Dep endency and Heterogeneity Challenges T o inv estigate the contribution of multimodal data to synthetic lethality prediction and to v alidate the effectiveness of the SynLeaF dual-stage fusion strategy , w e executed an in-depth ablation analysis fo cusing on the Only Omics, Only KG, and the full version of SynLeaF. 2.3.1 Multimo dal In tegration Sho ws an “En velope Effect” and Robustness Sup erior to Unimo dal Benc hmarks As shown in Figure 2 , SynLeaF forms a clear “env elop e effect” ov er the unimo dal mo dels, indicating stable gains from multimodal fusion. Notably , mo dal adv antages shift drastically depending on the data split. F or SKCM in CV1, KG provides the main predictiv e signal (AUC ≈ 0.78) compared to Omics (AUC ≈ 0.64). How ev er, in CV2 (unseen genes), a modal reversal occurs: KG drops sharply to 0.60 due to sparse graph connections, while Omics rises to 0.74. Despite this fluctuation, SynLeaF adaptiv ely shifts its fo cus to Omics, main taining a robust AUC of 0.76. It is worth noting that this modal adv antage sho ws a different pattern from the pan-cancer p erspective. In the CV1 setting on the pan-cancer dataset, b oth Omics and KG show ed very high p erformance. But in the more c hallenging zero-shot condi- tion of the CV3 setting, the situation was reversed. The p erformance of Only Omics dropp ed significantly to 0.6259, while Only K G still maintained a robust AUC of 0.7391. This reveals that in complex scenarios that span m ultiple cancer types, simple omics features are easily affected by heterogeneity noise. The global knowledge graph, on the other hand, provides a stronger inductive bias through the biological net work top ology , thus enabling structural inference on completely unkno wn genes. 8 BRCA CESC CO AD KIRC LAML L U AD O V SK CM pan 0.6 0.7 0.8 0.9 1.0 Performance Comparison (CV1) BRCA CESC CO AD KIRC LAML L U AD O V SK CM pan 0.6 0.7 0.8 0.9 1.0 Performance Comparison (CV2) Only Omics Only KG SynL eaF Fig. 2 Radar chart comparing the p erformance of SynLeaF and unimo dal baseline v ariants. This chart shows the AUC p erformance comparison of SynLeaF against the two unimo dal baseline v ariants, Only Omics and Only KG, on the pan-cancer and eight cancer-specific datasets, under the two data splitting strategies of CV1 and CV2. The p erformance curve of SynLeaF forms an “env elop e effect” ov er the unimo dal models on almost all datasets, demonstrating the consistent adv antage of m ultimo dal fusion. In summary , the lo cal case of SK CM and the global case of pan-cancer together pro ve that no single modality can excel under all splits, and SynLeaF’s multimodal fusion mechanism is an essen tial approac h to handle this complexity . 2.3.2 The Adaptiv e Selection Mec hanism Captures the Differen tial Mo dalit y Dep endency of Different Cancers Although m ultimo dal fusion is generally effective, w e observed a key phenomenon that differen t cancer datasets show very different modality dep endency under different split- ting strategies. In the synthetic lethality prediction task, deep cross-mo dal interaction is v ery imp ortan t in some scenarios, while in other scenarios, forced in teraction can instead in tro duce noise. Therefore, SynLeaF in tro duces an adaptive selection mec ha- nism to address the significan t differences in in ter-mo dal interactions under differen t data splitting strategies. T o accommo date the heterogeneit y within data distribu- tions, the model identifies the most effective fusion path by ev aluating outcomes on the v alidation set. W e plotted bar charts to ev aluate the effectiveness of the mo dal fusion strategies. Due to space limitations, this section only discusses t wo representa- tiv e datasets, CESC CV1 and COAD CV1, as sho wn in Figure 3 . Detailed outcomes of the full exp erimen ts are presented in Supplemen tary Figure S3. In the CESC (CV1) dataset, unimo dal Omics and KG yielded AUCs of 0.7679 and 0.7899, resp ectively . A simple late fusion (UME) achiev ed only 0.7959, struggling to capture deep interactions. How ev er, consisten t with the first scenario in Du et al. [ 39 ], SynLeaF’s UMT strategy achiev ed a significant adv antage (AUC=0.8136). By 9 Only Omics Only KG UMT UME 0.63 0.66 0.69 0.72 0.75 0.78 0.81 0.84 0.87 A UC Score Performance on CESC (CV1) Only Omics Only KG UMT UME 0.630 0.645 0.660 0.675 0.690 0.705 0.720 0.735 0.750 A UC Score Performance on CO AD (CV1) Fig. 3 Performance comparison of SynLeaF baseline v ariants on tw o cancer datasets. This figure shows the AUC scores for the unimo dal baseline v ariants (Only Omics, Only KG) and the baseline v ariants resp ectiv ely employing tw o multimodal fusion strategies (UMT, UME) on the CESC and COAD cancer datasets under the CV1 split. The height of the bars corresp onds to the mean AUC obtained via 5-fold cross-v alidation, while the standard deviations are denoted by the error bars. The star indicates the optimal fusion strategy , which is finally adopted in the SynLeaF adaptive selection mechanism on that dataset. utilizing feature-lev el distillation, UMT effectiv ely forces the netw ork to learn from b oth mo dalities and retain key cross-mo dal interactions. Conv ersely , COAD (CV1) presen ts a “strong omics (0.7069) and weak graph (0.6776)” scenario. Here, forcing feature alignmen t via UMT in tro duced noise, dropping the A UC to 0.6828 (lo wer than Only Omics). As prop osed by Du et al. [ 39 ] for cases with insignificant paired interac- tions, the UME strategy selected b y SynLeaF performed b est (AUC=0.7162). UME straigh tforwardly aggregates unimo dal results, effectiv ely av oiding mo dalit y laziness or negative transfer caused b y forced cross-mo dal in teractions. 2.3.3 Parameter Sensitivit y Analysis Confirms the Complemen tary Stabilit y of UMT and UME Strategies T o ev aluate the con tribution of the feature-lev el distillation mo dule and determine the optimal hyperparameter configuration, a sensitivity analysis was p erformed regarding the distillation weigh t λ distill in equation ( 10 ) for the UMT module, within the range of [0 , 1 , 10 , 20 , 50 , 100] (as shown in Figure 4 ). Here, λ distill = 0 is equiv alent to Na ¨ ıv e early fusion without distillation regularization. The exp erimen tal results sho w that for most datasets, in tro ducing mo derate distil- lation regularization ( λ distill = 1) significantly outp erforms the no-distillation baseline ( λ distill = 0). F or example, in CESC (CV1), the A UC increased from 0.7995 at λ distill = 0 to 0.8136 at λ distill = 1. This indicates that light weigh t feature alignment 10 0 1 10 20 50 100 0.92 0.94 0.96 Scor e BRCA (CV1) 0 1 10 20 50 100 0.78 0.80 0.82 0.84 0.86 0.88 BRCA (CV2) 0 1 10 20 50 100 0.79 0.80 0.81 0.82 0.83 CESC (CV1) 0 1 10 20 50 100 0.66 0.68 0.70 0.72 CESC (CV2) 0 1 10 20 50 100 0.68 0.69 0.70 Scor e CO AD (CV1) 0 1 10 20 50 100 0.600 0.625 0.650 0.675 0.700 CO AD (CV2) 0 1 10 20 50 100 0.72 0.74 0.76 0.78 KIRC (CV1) 0 1 10 20 50 100 0.60 0.65 0.70 KIRC (CV2) 0 1 10 20 50 100 0.6950 0.6975 0.7000 0.7025 0.7050 0.7075 Scor e LAML (CV1) 0 1 10 20 50 100 0.64 0.66 0.68 LAML (CV2) 0 1 10 20 50 100 0.84 0.86 0.88 0.90 L U AD (CV1) 0 1 10 20 50 100 0.76 0.78 0.80 0.82 L U AD (CV2) 0 1 10 20 50 100 d i s t i l l 0.96 0.97 0.98 Scor e O V (CV1) 0 1 10 20 50 100 d i s t i l l 0.72 0.74 0.76 0.78 0.80 0.82 O V (CV2) 0 1 10 20 50 100 d i s t i l l 0.750 0.775 0.800 0.825 0.850 SK CM (CV1) 0 1 10 20 50 100 d i s t i l l 0.74 0.76 0.78 SK CM (CV2) I m p a c t o f d i s t i l l o n 8 S i n g l e C a n c e r s P e r f o r m a n c e M e t r i c s AUC AUPR F1-Scor e 0 1 10 20 50 100 d i s t i l l 0.90 0.91 0.92 0.93 0.94 0.95 0.96 0.97 Scor e pan (CV1) 0 1 10 20 50 100 d i s t i l l 0.80 0.82 0.84 0.86 0.88 pan (CV2) 0 1 10 20 50 100 d i s t i l l 0.71 0.72 0.73 0.74 0.75 0.76 0.77 pan (CV3) I m p a c t o f d i s t i l l o n P a n - C a n c e r P e r f o r m a n c e M e t r i c s AUC AUPR F1-Scor e Fig. 4 Sensitivit y analysis of the λ distill parameter in the UMT mo dule. This figure shows how the p erformance (AUC, AUPR, F1-Score) of the UMT fusion strategy changes with the kno wledge distillation weigh t λ distill on all single-cancer and pan-cancer datasets. A v alue of λ distill = 0 corresponds to Na ¨ ıve multimodal training without distillation regularization. 11 can effectively mitigate mo dalit y laziness while a voiding the risk of excessiv e regular- ization. Based on the ma jorit y principle and considerations for model generality , we uniformly set λ distill to 1 in the final SynLeaF mo del. Ho wev er, this fixed-parameter strategy inevitably faces c hallenges in some highly heterogeneous datasets. W e observ ed that LAML (CV1) and LUAD (CV2) show ed a sp ecial decrease-then-increase trend. A w eak distillation ( λ distill = 1) actually in ter- fered with the model’s feature learning, leading to performance lo wer than the baseline with λ distill = 0. F or example, in LUAD (CV2), the AUC dropp ed from 0.7672 to 0.7589 when λ distill = 1. Although the data trend shows that the mo del could o vercome this obstacle and achiev e b etter performance if the distillation weigh t were further increased (for example, λ distill ≥ 10), under the unified setting of λ distill = 1, the UMT module did indeed reach a p erformance low. But during the v alidation phase, SynLeaF’s adaptiv e strategy successfully iden tified the performance decline of UMT and selected UME as the final inference mo del. This allow ed the final model to still main tain a comp etitive performance (the final AUC for LUAD CV2 was 0.8161). This result precisely v alidates the necessity of the adaptiv e selection mechanism and the high fault-tolerance of SynLeaF’s dual-stage architecture. It allo ws the model to use a single set of general hyperparameters for most scenarios, while relying on the adap- tiv e switc hing mec hanism to provide a robust fallback for the few scenarios that are sensitiv e to parameters. 2.4 Gradien t Dynamics Analysis Rev eals the Mec hanism for Mitigating Mo dality Laziness T o b etter understand how the UMT strategy effectively mitigates the modality laziness problem in multimodal learning, The CESC (CV1, F old2) dataset, whic h shows a t ypical UMT adv antage, was employ ed as a representativ e case to study the gradient dynamics across the training phase (as shown in Figure 5 ). 2.4.1 The UMT Strategy Significan tly Impro ves Both Ov erall and Unimo dal Performance Figure 5 (a) sho ws the comparison of o verall test p erformance. In the early stage of training (ab out the first 50 ep ochs), the Na ¨ ıv e no-distillation baseline conv erges faster b ecause it lacks additional regularization. How ever, the cost of this rapid conv ergence is o verfitting. After reaching its peak, the p erformance fluctuations follow ed by a decline are observ ed in the baseline model. In contrast, although the UMT strategy rises more slo wly at the b eginning, it shows a contin uous and stable learning ability , and con tinues to climb after surpassing the baseline around ep och 60, even tually stabilizing at a high level ab o ve 0.86. The light green shaded area visually illustrates the substantial p erformance adv antage achiev ed by the UMT strategy during the adv anced training ep ochs. Figure 5 (b) further reveals that this ov erall p erformance improv ement comes from impro vemen ts at the unimo dal level. W e observ ed t wo key phenomena. First, for the Omics mo dalit y , the Na ¨ ıve no-distillation baseline ( λ distill = 0) exp erienced a significan t p erformance collapse in the later stages of training. The AUC dropp ed 12 0 25 50 75 100 125 150 175 200 Epoch 0.50 0.55 0.60 0.65 0.70 0.75 0.80 0.85 AUC (a) Overall T est Performance N a ï v e ( d i s t i l l = 0 ) U M T ( d i s t i l l = 5 0 ) Selected Ckpt 0 25 50 75 100 125 150 175 200 Epoch 0.50 0.55 0.60 0.65 0.70 0.75 0.80 0.85 AUC (b) Unimodal T est Performance O m i c s ( d i s t i l l = 0 ) K G ( d i s t i l l = 0 ) O m i c s ( d i s t i l l = 5 0 ) K G ( d i s t i l l = 5 0 ) Selected Ckpt 0 25 50 75 100 125 150 175 200 Epoch 0.02 0.03 0.04 0.05 0.06 0.07 Gradient Nor m (c) Gradient Norm Comparison K G ( d i s t i l l = 0 ) O m i c s ( d i s t i l l = 0 ) K G ( d i s t i l l = 5 0 ) O m i c s ( d i s t i l l = 5 0 ) 0 25 50 75 100 125 150 175 200 Epoch 0.5 0.6 0.7 0.8 0.9 1.0 R atio (d) Gradient Norm Ratio d i s t i l l = 0 ( N a ï v e ) d i s t i l l = 1 0 d i s t i l l = 5 0 Fig. 5 Gradient dynamics analysis on the CESC (CV1, F old2) dataset. (a) Overall test performance comparison: T est AUC curves for UMT (solid line) and the Na ¨ ıv e no-distillation base- line (dashed line). The red dots mark the chec kp oin ts selected based on the v alidation set. (b) improv ements in unimo dal scenarios: change in test AUC for the Omics and KG mo dalities under the UMT and Na ¨ ıve no-distillation strategies (denoted as Omics/KG ( λ distill = 50) and Omics/KG ( λ distill = 0), resp ectiv ely). The vertical line marks the UMT chec kp oin t p osition. (c) Gradient norm comparison: change in the gradient norms of the tw o mo dalit y enco ders ov er training ep ochs under different distillation weigh ts λ distill . (d) Gradient norm ratio: trend of the gradien t balance betw een the t wo mo dalities. Here, the gradien t norm ratio is defined as the L 2 norm of the gradients with respect to the Omics enco der’s parameters divided b y that of the KG enco der’s parameters. sharply from its p eak to about 0.75, which is a typical phenomenon of ov erfitting. In con trast, the UMT strategy ( λ distill = 50) not only increased steadily but also success- fully a voided p erformance degradation in the later stages, alw ays staying ab o ve 0.80. Second, for the K G modality , the UMT strategy also maintained a small but consis- ten t adv antage compared to the baseline, indicating that the knowledge distillation mec hanism had a p ositiv e regularization effect on b oth mo dalities. Notably , the chec kp oin ts denoted by red dots within Figure 5 (a) are not the global optimal p oin ts of their resp ectiv e curves on the test set, and there is still room for impro vemen t after these p oin ts. This observ ation once again confirms the previous discussion ab out KIRC, sho wing that the distribution shift b et ween the v alidation and test sets is a common challenge in biomedical data analysis [ 41 ]. Selecting mo dels based on v alidation set metrics may fail to capture the true optimal state on the test set. 13 2.4.2 Gradient Norm Analysis: Kno wledge Distillation Enhances the Learning Signal of the W eak er Mo dalit y T o explain the ab o v e phenomena from an optimization dynamics p ersp ectiv e, w e analyzed the c hanges in the gradient norms of the tw o mo dal enco ders during the training pro cess (Figure 5 (c)), follo wing the analytical proto cols established in recent m ultimo dal learning studies [ 42 ]. In the Na ¨ ıve no-distillation baseline ( λ distill = 0), we observed a significan t asym- metry in gradient decay . In the early stage of training, the gradient norms of b oth mo dalities w ere at a relatively high level of ab out 0.07. How ever, as the training pro- gressed, the gradient norm of the Omics mo dalit y dropp ed sharply , ev entually falling b elo w ab out 0.02. On the other hand, the decrease for the KG mo dalit y was relativ ely gen tle, finally staying at a level of ab out 0.03. This difference means that in the later stages of training, the Omics enco der almost stopp ed learning effectively . The mo del mainly relied on the KG mo dalit y for prediction, which is direct evidence of mo dality laziness. After in tro ducing the knowledge distillation regularization ( λ distill = 50), the situ- ation changed significantly . The ligh t green shaded area clearly sho ws ho w the UMT strategy successfully increased the gradient norm of the Omics mo dalit y . Although the gradient for Omics was still decreasing, the rate of decrease slow ed do wn signifi- can tly . This means that the distillation loss pro vided an additional sup ervision signal for the Omics enco der, forcing it to contin ue learning to align with the feature repre- sen tations deriv ed from the pre-trained teacher mo del and th us a voiding early gradien t v anishing. Notably , under b oth strategies, the gradients for the KG modality in the later training stages almost o v erlapp ed. This indicates that the main role of UMT is to selectively enhance the w eaker mo dalit y , rather than in terfering with the normal learning of the stronger mo dality . Figure 5 (d) more directly quan tifies the degree of learning balance betw een the t w o mo dalities through the gradien t ratio (Omics/KG). A ratio closer to 1.0 indicates that the learning dynamics of the tw o mo dalities are more balanced, while a decreasing ratio means that the relative contribution of Omics is w eakening. In the Na ¨ ıv e no- distillation baseline ( λ distill = 0), the gradient ratio starts from a p erfectly balanced 1.0 and then rapidly decreases, dropping to 0.50 in the later training stages. This means that b y the end of training, the gradient con tribution of the Omics mo dalit y w as only half that of the KG mo dalit y , causing a serious learning im balance. As the distillation weigh t λ distill increases, this im balance is gradually mitigated. A v alue of λ distill = 10 keeps the ratio at a higher level, and the effect of λ distill = 50 is the most significan t. Ev en at the end of training, the ratio remains at ab out 0.58, sho wing a clear improv ement in balance compared to the baseline’s v alue of b elo w 0.50. The gradient dynamics analysis abov e provides direct mec hanical evidence for the effectiv eness of the UMT strategy . Through the knowledge distillation framework, the pre-trained Uni-mo dal T eacher mo del provides contin uous feature-level sup ervision for the student enco ders in the join t training. This effectively enhances the gradien t signal of the w eaker mo dality and reduces the learning gap with the stronger modality . W e also honestly p oint out that the UMT strategy did not completely eliminate the gradien t gap betw een modalities. Even with λ distill = 50, the final gradient ratio was 14 still 0.58, which is still far from the ideal 1.0. This suggests that mo dalit y laziness is a deep-rooted optimization problem that is difficult to solve completely with a single tec hnique. How ever, the exp erimen tal results sho w that ev en mitigating rather than completely eliminating the problem is enough to bring substan tial impro vemen ts in predictiv e p erformance, which v alidates the rationale of our metho d’s design. 2.5 SynLeaF W eb Server and Case Study Applications T o facilitate the exploration of syn thetic lethality in teractions, we dev elop ed a w eb- based query platform 2 . The back end uses an ensemble of fiv e CV2 mo dels p er cancer t yp e and av erages their outputs to impro ve robustness. The in terface visualizes predic- tions across eight cancers and the pan-cancer context, alongside ground-truth labels from existing datasets for direct comparison. T able 3 Prediction scores retrieved from the SynLeaF w eb server for two case studies. Gene Pairs BRCA CESC COAD KIRC LAML LUAD OV SKCM pan RAD51–BRCA1 0.9928 0.3817 0.6562 0.6038 0.5830 0.4169 0.3849 0.5850 0.9748 RAD51–BRCA2 0.9213 0.4462 0.4572 0.6160 0.5406 0.5259 0.3624 0.6082 0.9596 ADAR-BR CA1 0.9307 0.4876 0.5443 0.3883 0.5413 0.4984 0.2611 0.5221 0.8667 ADAR-BR CA2 0.7750 0.4697 0.3759 0.3465 0.5174 0.4815 0.1832 0.5595 0.7112 Milordini et al. [ 6 ] demonstrated the synthetic lethalit y of targeting the RAD51– BR CA2 in teraction, particularly in pancreatic cancer. Although these pairs are labeled as p ositive in our pan-cancer dataset, SynLeaF provides detailed context-specific insigh ts. As sho wn in T able 3 , the prediction scores in the BRCA column are excep- tionally high, significan tly surpassing those in other cancer t yp es. This strong signal suggests that breast cancer ma y share the same RAD51-mediated synthetic lethalit y mec hanism observ ed in pancreatic cancer. Chabanon et al. [ 7 ] identified ADAR1 as a k ey synthetic lethal target in BRCA-m utan t cancers. Notably , despite the absence of AD AR-BRCA* pairs from our dataset, SynLeaF successfully predicted this latent rela- tionship, crucially exhibiting high efficacy in the relev ant cancer context: the prediction score in the BRCA column reached 0.9307 for AD AR-BRCA1, whic h is remark ably higher than in other unrelated cancer types. This result demonstrates SynLeaF’s capa- bilit y to generalize and discov er no vel, clinically relev ant synthetic lethalit y pairs in sp ecific cancer t yp es. 3 Discussion Syn thetic lethality is a key mechanism in precision oncology , y et identifying robust SL pairs remains c hallenging [ 9 ]. Our results show that SynLeaF improv es prediction p erformance across pan- and single-cancer settings and adapts to modality dep endency differences. A key finding is that balancing modality in teraction and a voiding mo dalit y laziness are crucial for multimodal SL prediction [ 39 ]. SynLeaF com bines unimo dal 2 The web serv er is accessible at: h ttps://synleaf.bioinformatics- lilab.cn 15 pre-training with tw o complementary fusion strategies: UMT for deep interaction and UME for conserv ative ensem bling, providing robustness across div erse cancer contexts and alleviating negative transfer [ 43 ]. Although SynLeaF has ac hieved a breakthrough in prediction accuracy , the current w ork still faces limitations regarding the practical requiremen ts for clinical translation. First, our mo del lac ks interpretabilit y regarding multimodal interactions. F uture work will b e devoted to clarifying how con tinuous omics features enhance or suppress KG top ology to provide a complete biological evidence chain. Second, the 1:1 class bal- ancing strategy deviates from the highly im balanced biological realit y . F urthermore, distribution shifts b et ween v alidation and test sets in small-sample cancers hinder optimal mo del selection. F uture research could reframe SL prediction as a recom- mendation task to handle long-tail distributions and dev elop more robust ev aluation strategies. Finally , while Cross-V AE handles missing in tra-omics data, SynLeaF still requires b oth omics and K G mo dalities. In clinical practice, a patient may completely lac k sequencing data, or some new genes ma y not b e included in existing kno wledge graphs. F uture studies could explore cross-mo dal completion techniques based on gen- erativ e modeling to enable flexible prediction when only a single fundamen tal modality is av ailable. 4 Conclusion This study in tro duces SynLeaF, a dual-stage m ultimo dal fusion framew ork for syn thetic lethality prediction across pan- and single-cancer con texts. Through V AE- enhanced omics enco ding and a kno wledge distillation strategy , SynLeaF effectiv ely o vercomes the c hallenges of mo dalit y laziness and heterogeneit y in multi-source data fusion. Extensive exp erimen ts show that the prop osed framew ork exhibits state-of- the-art capabilit y within prediction scenarios targeting for both kno wn and new genes. SynLeaF not only pro vides a p o werful computational to ol for discov ering synthetic lethalit y targets, but its approach to handling dynamic multimodal dep endencies also offers a new p erspective for general link prediction tasks in the biomedical field. 5 Metho ds 5.1 Data Acquisition and Prepro cessing W e constructed a strictly filtered and m ulti-source in tegrated pan-cancer and cancer- sp ecific dataset. The data cov ers SL gene pairs, multi-omics data, biomedical kno wledge graphs, and protein sequence information. W e in tegrated data from author- itativ e databases suc h as SynLethDB 2.0 [ 3 ], ELISL [ 34 ], TCGA [ 44 ], and UniProt [ 45 ], and we designed a standardized prepro cessing workflo w to eliminate noise and heterogeneit y . 5.1.1 Data Sources Syn thetic Lethality Data. The synthetic lethality lab el data comes from tw o main sources. F or the pan-cancer prediction task, we use the SynLethDB 2.0 database, which 16 pro vides general synthetic lethality gene pairs across cancer types. F or the cancer- sp ecific prediction task, we adopted the dataset organized b y the ELISL study . ELISL in tegrates high-confidence SL gene pairs from previous studies such as Disco verSL [ 14 ], ISLE [ 46 ], and EXP2SL [ 15 ]. It cov ers eight sp ecific cancer types, and the num b er of syn thetic lethality pairs b efore preprocessing is shown in Supplementary T able S3. Multi-Omics Data. The omics data comes from the TCGA database curated b y the cBioP ortal platform [ 47 ]. W e collected four key omics features for the ab ov e eigh t cancer types: Cop y Num b er V ariation, which represents the relativ e linear copy n umber v ariation v alues of genes; Gene Expression, whic h uses mRNA z-score data pro cessed by RNA-Seq V2 RSEM normalization; DNA Methylation, which reflects the epigenetic regulation status of genes (HM27 or HM450); and Mutation data, which records the mutation status of each gene in every sample, where we only consider the mutation counts of genes in each sample. F or the pan-cancer dataset, due to data limitations, the dataset is restricted to three data modalities consisting of gene expression, mutation, and copy n umber v ariation. Kno wledge Graph. W e adopted the biomedical knowledge graph (SLKG 2.0) pro vided b y SynLethDB 2.0. This kno wledge graph con tains 37,341 entities of 11 t yp es (including genes, Gene Ontology , pathw ays, drugs, and diseases) and 1,405,652 relationships of 27 t yp es, forming a rich biomedical knowledge netw ork. Protein Sequences. T o supp ort comparative exp eriments, we downloaded review ed h uman protein sequence data from the UniProt database (as of Octob er 26, 2024), whic h contains a total of 20,428 sequences where eac h gene corresp onds to a represen tative protein sequence consisting of 21 types of amino acids. 5.1.2 Data Prepro cessing W e executed rigorous proto cols for gene filtering and data cleaning to guaran tee the uniformit y and superior quality of the multimodal data. Gene Filtering and Alignmen t. W e p erformed m ulti-condition in tersection filtering on the genes. Genes included in the study must meet the following conditions sim ultaneously: they must ha ve corresp onding review ed protein sequences in UniProt, b e recorded in at least one type of omics data, and hav e at least one accessible neigh b or no de in the knowledge graph. Through this strict filtering mechanism, we filtered out genes with incomplete information and ensured that ev ery gene used for mo del training has a complete multimodal feature representation. The num b er of synthetic lethalit y pairs after filtering is shown in T able 4 . SL Data Construction and Balancing. T o address the noise and imbalance issues in the original SL data, w e performed the follo wing processing. First, in the pan- cancer dataset, we treated pairs explicitly lab eled as Non-SL and Synthetic Rescue as negativ e samples. Second, for conflicting gene pairs that app eared in b oth the p ositiv e and negativ e sample sets, we adopted a conserv ativ e strategy and uniformly classi- fied them as negative samples. Additionally , w e corrected errors in the ELISL data where some self-loops were incorrectly identified as p ositiv e samples. Consequently , w e reassigned them to the negative category . Considering the inherent sparsity of gen uine synthetic lethality associations, the quantit y of negative samples within the dataset typically v astly surpasses the count of p ositiv e ones. T o av oid the mo del biasing 17 T able 4 Statistics of synthetic lethalit y pair counts after data processing Cancer Ty p e # T otal Genes # Pos. # Neg. BRCA 17965 1349 990 CESC 17977 144 4738 COAD 17961 1560 70982 KIRC 17963 60 2514 LAML 17944 1147 18912 LUAD 17954 582 5460 OV 17958 253 556 SKCM 17969 101 16157 pan 17690 33746 3509 # Pos. denotes the coun t of p ositiv e samples; # Neg. denotes the coun t of negative samples. to wards the ma jorit y class, w e adopted a 1:1 positive-negativ e sample balance strat- egy . When there were to o many negative samples, we used random undersampling; when kno wn negative samples were insufficient in specific cancer types, we randomly generated unlab eled gene pairs from the filtered gene p o ol as supplementary nega- tiv e samples, and we ensured that these generated pairs did not ov erlap with kno wn p ositiv e samples. Omics Data Normalization. The sample counts for different features across v arious cancer types in the original omics data are shown in Figure 6 . W e p erformed sample-lev el alignment on the omics data for eac h cancer to ensure that the four omics features for each case sample are one-to-one matc hed. T o address the sparsity and distribution characteristics of the data, the pro cessing w orkflow is as follo ws. Due to the sparsit y of omics data, w e filled all missing v alues and non-numeric v alues with 0 to represen t no abnormality or a default state; for gene expression data, w e truncated v alues smaller than -10 to -10 b ecause w e observed that extremely small v alues could cause n umerical instabilit y; regarding mutation profiles, we calculated the mutation frequency for every individual gene and applied a logarithmic transformation to map it to the interv al [0 , 1] using the following form ula: x ′ = ( ln(1+ x ) ln(1+ M ) , if M > 0 0 , if M = 0 (1) where x is the original m utation coun t and M is the maxim um m utation coun t in that cancer type. F urthermore, giv en that syn thetic lethalit y effects often in volv e in teractions b et ween normal genes and abnormal genes [ 48 ], we did not exclude genes with low v ariation based on the c ase sample distribution but instead retained all genes that met the omics alignmen t requiremen ts. W e retained only unique records for duplicate gene en tries app earing in single-omics data. Kno wledge Graph Refinemen t. T o construct cancer-sp ecific knowledge graphs, w e p erformed subgraph extraction on the original graph based on cancer types. F or eac h sp ecific cancer, we only retained its corresp onding “Disease” no de and the edges 18 BR CA CESC CO AD KIR C L AML L UAD O V SK CM pan Cancer T ype 0 500 1000 1500 2000 2500 3000 Sample Count 1080 295 616 528 191 516 579 367 2703 1100 306 382 534 173 517 307 472 1210 788 309 396 320 194 460 592 473 0 982 194 223 451 197 230 316 368 2683 Sample Statistics of 4 Omics T ypes CNV Gene Expr ession Methylation Mutation Fig. 6 Statistics of sample counts for original omics data across cancer types. This figure displays the n umber of raw av ailable samples b efore prepro cessing for four omics data types (Copy Number V ariation (CNV), Gene Expression, DNA Methylation, and Mutation) across the eight specific cancer types and the pan-cancer dataset used in this study . Noting that the pan-cancer (pan) dataset does not include DNA methylation data due to data source limitations. directly connected to it, removing other irrelev an t cancer nodes to reduce noise inter- ference. Additionally , for the unidirectional relationships in the graph, w e generated corresp onding reversed edges to transform the directed graph into a heterogeneous graph containing bidirectional information. 5.1.3 Dataset Splitting Strategies In order to thoroughly assess the robustness and generalization capacit y of our frame- w ork, we referred to the work of PiLSL [ 23 ] and adopted three cross-v alidation (CV) splitting strategies. The partitioning of all datasets adhered to a fixed proportion of 7:1:2 for the training, v alidation, and testing subsets, resp ectiv ely . • CV1 (Random Split) : All samples (gene pairs) are randomly sh uffled and split. Under this setting, genes present in the testing phase migh t also exist within the training set. This primarily ev aluates the inductive p otential of the mo del concerning no vel combinations of kno wn genes. • CV2 (Semi-New Gene Split) : The gene set is partitioned to guarantee that for ev ery gene pair situated in the v alidation or testing subsets, precisely a single gene is found in the training data, while the other gene is completely new to the training set. This simulates the real-world scenario of finding kno wn targets for new genes, and it ev aluates the mo del’s generalization ability for semi-new gene pairs. • CV3 (New Gene Split) : This is the strictest splitting method, ensuring a com- plete absence of test-set genes within the training set. This tests the mo del’s inferen tial capability in a completely unexplored gene space (zero-shot setting). 19 It is w orth noting that due to the small sample size of some cancer-sp ecific datasets, p erforming CV3 splitting ma y result in extremely few samples in the test set, rendering it statistically insignifican t. Therefore, in subsequent exp eriments, we only rep ort the results of CV1 and CV2 for single-cancer prediction tasks, while w e use all three splitting strategies for comprehensiv e ev aluation in the pan-cancer prediction task. 5.2 The Sy nLeaF F ramew ork This pap er in tro duces the SynLeaF arc hitecture, a deep learning system grounded in dual-stage m ultimo dal fusion strategies. This framework aims to accurately predict SL in teractions b y in tegrating omics features from genomics and structured kno wledge from biomedical KGs. The arc hitecture of the mo del is illustrated in Figure 1 . 5.2.1 Problem Definition W e define the synthetic lethality prediction task as a binary classification problem. Giv en a sp ecific instance of a gene pair ( g i , g j ), the ob jective is to predict whether an SL relationship exists b et ween them, which corresp onds to outputting a lab el y ∈ { 0 , 1 } . Our metho d follo ws a unified Siamese-like netw ork architecture [ 49 ] for representation learning of gene pairs, regardless of whether it is in a single-mo dal or multimodal setting. Specifically , for each gene in the pair, we first utilize its multimodal features to learn its embedding representation through a corresp onding mo dalit y-sp ecific enco der φ m (where m ∈ { o, k } represents the omics or knowledge graph mo dalit y): h m i = φ m ( f m i ) , h m j = φ m ( f m j ) (2) where f m i denotes the feature input corresponding to the m -th mo dality for the gene g i . The obtained embedding vectors of the tw o genes are subsequen tly used as input features to output the final prediction logits through a shared classifier F m : ˆ y m i,j = F m ([ h m i ; h m j ]) (3) where [ · ; · ] denotes the concatenation op eration. W e aim to optimize the model by minimizing the Binary Cross-Entrop y (BCE) loss, which is formulated as: L BCE = − 1 |D | X ( g i ,g j ) ∈D y i,j log( ς ( ˆ y m i,j )) + (1 − y i,j ) log(1 − ς ( ˆ y m i,j )) (4) where D represents the set of all gene pairs within the training set, y i,j denotes the ground truth annotation, and ς ( · ) serves as the sigmoid activ ation function for transforming logits into probability scores. W e denote the predicted probability after sigmoid transformation as p m i,j = ς ( ˆ y m i,j ), whic h represents the probability that the gene pair ( g i , g j ) is predicted to hav e a syn thetic lethality relationship under mo dalit y m . 20 5.2.2 Omics Enco der with V AE Early F usion The encoding of the omics modality adopts an early fusion mo dule (OmicsEncoder, φ o ). W e constructed an N × N V AE enco der matrix (where N = 4 denotes the total coun t of omics categories) and lev eraged the Pro duct of Exp erts (PoE) [ 37 ] mec hanism to execute early fusion on the omics features. Eac h V AE enco der is implemen ted b y a Multi-La yer Perceptron (MLP), whic h maps input features to the mean µ and log-v ariance log σ 2 in the latent space. Let the N t yp es of omics features for gene g i b e denoted as F i = { f (1) i , . . . , f ( N ) i } . As shown in the Omics Enco der mo dule of Figure 1 , the enco der matrix consists of self-enco ders (Self-V AE) on the diagonal and cross-enco ders (Cross-V AE) off the diagonal. F or the k -th omics data f ( k ) i of gene i , the self-enco der V AE k,k maps it to the parameters of the latent distribution, µ ( k, self) i and log( σ ( k, self) i ) 2 . F or the j -th ( j = k ) omics data f ( j ) i of gene i , the cross-enco der V AE k,j attempts to infer the laten t distribution of the k -th omics type, with parameters denoted as µ ( k, cross ,j ) i and log( σ ( k, cross ,j ) i ) 2 . T o aggregate features from differen t p erspectives and address the deficiency of missing mo dalities for certain genes, w e utilize the P oE mechanism to calculate a join t p osterior distribution for each target omics type k of gene i , where the parameters ( µ ( k, PoE) i , σ ( k, PoE) i ) are given by the following formulas: 1 ( σ ( k, PoE) i ) 2 = X j = k 1 ( σ ( k, cross ,j ) i ) 2 , µ ( k, PoE) i = ( σ ( k, PoE) i ) 2 X j = k µ ( k, cross ,j ) i ( σ ( k, cross ,j ) i ) 2 (5) Then, we sample from the posterior distributions of the self-enco der path and the P oE path using the reparameterization trick to obtain the respective sets of laten t v ariables Z self i = { z ( k, self) i } N k =1 and Z PoE i = { z ( k, PoE) i } N k =1 : z ( k, path) i = µ ( k, path) i + σ ( k, path) i ⊙ ϵ , path ∈ { self , PoE } (6) where ϵ ∼ N (0 , I ) denotes noise sampled from a standard normal distribution. Finally , the latent v ariable sets from the tw o paths are sub jected to mean p o oling resp ectiv ely and then concatenated. They are then pro jected via a fully connected la yer to yield the final omics embedding h o i : h o i = F C " 1 N N X k =1 z ( k, self) i ; 1 N N X k =1 z ( k, PoE) i #! (7) The training pro cess adopts a V ariational Information Bottleneck (VIB) loss func- tion [ 50 ], which includes reconstruction error and a KL div ergence regularization term: L omics = L BCE + λ self X k D K L ( q ( k ) self || p ) + λ cross X k D K L ( q ( k ) PoE || p ) (8) where p is the standard normal prior distribution N (0 , I ). The w eights of the t wo KL div ergences, λ self and λ cross , are set to 0.1 and 0.5, respectively . 21 5.2.3 Knowledge Graph Enco der with RGCN F or the KG mo dalit y , w e adopt R GCN [ 38 ] as the enco der (KGEncoder, φ k ) to capture the top ological structures and semantic relationships of genes in the biological netw ork. Let G = ( V , E , R ) b e a knowledge graph, comprising the collections of entities V , edges E , and relations R . Regarding a sp ecific target gene g i (corresp onding to graph node i ), we first emplo y an L -hop subgraph sampling strategy to extract its lo cal neighborho o d subgraph to reduce computational complexit y . The initial no de features are obtained through an Em b edding La yer. The RGCN lay er updates no de represen tations b y aggregating neigh b or information under different relationship t yp es r ∈ R : h ( l +1) i = ReLU X r ∈R X j ∈N r i 1 c i,r W ( l ) r h ( l ) j + W ( l ) 0 h ( l ) i (9) where N r i constitutes the neighbor set for no de i asso ciated with relationship r , while c i,r serv es as a normalization factor equal to |N r i | . After multi-la yer RGCN aggregation, w e extract the final representation h k i of the cen tral gene no de. Optimization of the knowledge graph branc h is likewise ac hiev ed through the minimization of the binary cross-entrop y loss, denoted as L kg = L BCE . 5.2.4 Dual-Stage Multimo dal F usion Strategy The multimodal fusion adopts a dual-stage training strategy , which includes tw o com- plemen tary sc hemes named UMT and UME [ 39 ]. In the first stage, we indep enden tly train the omics prediction mo del (Only Omics) and the knowledge graph prediction mo del (Only K G). Let the omics teacher enco der obtained from the first stage training b e φ T o , and the knowledge graph teacher enco der b e φ T k . This stage ensures that each single-mo dal enco der can fully explore the feature representations within its mo dalit y without interference from other mo dalities. In the second stage, we designed tw o com- plemen tary fusion strategies and dynamically identified the optimal scheme according to the results observ ed in the v alidation subset. (1) Uni-Mo dal T eac her (UMT) : This strategy employs a Knowledge Distil- lation framework to address the issue of mo dalit y laziness and guarantee that the m ultimo dal architecture comprehensively captures the unimo dal feature representa- tions sp ecific to each mo dalit y . W e freeze the single-mo dal enco ders pre-trained in the first stage as the “T eac her” and initialize a new multimodal mo del as the “Student”, whic h includes an omics studen t enco der φ S o and a knowledge graph student enco der φ S k . While learning the classification task, the student mo del is also required to simu- late the intermediate feature represen tations of the teacher mo del by minimizing the distillation loss (Mean Squared Error, MSE). The ov erall loss function is form ulated as follows: L UMT = L BCE + λ distill X m ∈{ o,k } ∥ h m,S i − h m,T i ∥ 2 + L KL (10) 22 Here, h o,T i and h k,T i are the omics and graph em b eddings output by the frozen teac her enco ders for gene i , resp ectively . While h o,S i and h k,S i are the corresp onding outputs of the student enco ders. The classifier F UMT receiv es the genomic and knowledge graph features [ h o,S i ; h k,S i ] extracted by the student enco ders to mak e predictions. The h yp erparameter λ distill is set to 1, and this feature-level distillation forces eac h branch of the student mo del to main tain strong feature extraction capabilities, whic h effectiv ely alleviates mo dalit y laziness. (2) Uni-Mo dal Ensem ble (UME) : As a simpler late fusion baseline, UME directly integrates the prediction results of the tw o pre-trained mo dels from the first stage. The final predicted probability p UME is the av erage of the output probabilities from the tw o single-mo dal models: p UME = 1 2 ( ς ( ˆ y o ) + ς ( ˆ y k ) ) (11) UME requires no additional training and completely a voids gradien t interference issues during joint training, making it more robust in cases where there are significant differences b etw een mo dalities. Considering the differences in the effects of cross-mo dal in teractions across differ- en t datasets, we adopted a data-driven adaptiv e selection strategy . After training is completed, we ev aluate the AUC metrics of UMT and UME on the v alidation set separately and select the b etter-p erforming metho d as the final mo del: p i,j = ( p UMT i,j , if A UC v al UMT > A UC v al UME p UME i,j , otherwise (12) UMT typically p erforms b etter when b oth omics and knowledge graph single- mo dal features are strong and their interaction pro vides additional information. Con versely , when one mo dalit y is significan tly dominant or when cross-modal inter- action introduces noise, the simple ensemble strategy of UME prov es to b e more effectiv e. It is w orth noting that compared to traditional complex fusion metho ds, our metho d is simple to implement and easy to tune. UMT requires tuning only one addi- tional h yp erparameter (the distillation loss w eight λ distill ), while UME do es not even require extra training. This simplicit y not only impro ves the practicalit y of the method but also enhances its p ortability to new datasets. Co de a v ailability The data and co de can be accessed at the following GitHub rep ository: https://gith ub. com/Jmpax404/SynLeaF . Ac kno wledgemen ts This w ork was supp orted b y the gran ts from the National Key R&D Program of China (2024YF A0919600) and National Natural Science F oundation of China (32470704). 23 Author con tributions J.L. conceived and designed the pro ject and sup ervised the w ork. Z.X. developed the metho ds, p erformed bioinformatics analysis and drafted the man uscript. S.Z., R.W., and R.H. prepared the data and p erformed the benchmarks. S.Z., S.C., Y.H., and J.M. con tributed to the b enc hmarks. Y.C., X.W., and Y.W. participated in pro ject design and co ordination. Comp eting in terests The authors declare no comp eting in terests. References [1] Huang, A., Garraw a y , L.A., Ash worth, A., W eb er, B.: Syn thetic lethality as an engine for cancer drug target discov ery. Nature Reviews Drug Disco v ery 19 (1), 23–38 (2020) https://doi.org/10.1038/s41573- 019- 0046- z [2] Ashw orth, A., Lord, C.J.: Synthetic lethal therapies for cancer: what’s next after P ARP inhibitors? Nature Reviews Clinical Oncology 15 (9), 564–576 (2018) https: //doi.org/10.1038/s41571- 018- 0055- 6 [3] W ang, J., Zhang, Q., Han, J., Zhao, Y., Zhao, C., Y an, B., Dai, C., W u, L., W en, Y., Zhang, Y., Leng, D., W ang, Z., Y ang, X., He, S., Bo, X.: Computa- tional methods, databases and to ols for synthetic lethalit y prediction. Briefings in Bioinformatics 23 (3), 106 (2022) https://doi.org/10.1093/bib/bbac106 [4] T opatana, W., Juengpanich, S., Li, S., Cao, J., Hu, J., Lee, J., Suliyan to, K., Ma, D., Zhang, B., Chen, M., Cai, X.: Adv ances in synthetic lethality for cancer therap y: cellular mec hanism and clinical translation. Journal of Hematology & Oncology 13 (1), 118 (2020) https://doi.org/10.1186/s13045- 020- 00956- 5 [5] Lord, C.J., Ash worth, A.: P ARP inhibitors: Syn thetic lethality in the clinic. Science 355 (6330), 1152–1158 (2017) https://doi.org/10.1126/science.aam7344 [6] Milordini, G., Zacco, E., Masi, M., Armaos, A., Di Palma, F., Oneto, M., Gilo di, M., Rupert, J., Broglia, L., V arignani, G., Scotto, M., Marotta, R., Girotto, S., Ca v alli, A., T artaglia, G.G.: Computationally-designed aptamers targeting rad51- brca2 in teraction impair homologous recombination and induce synthetic lethal- it y . Nature Communications (2025) h ttps://doi.org/10.1038/s41467- 025- 66694- 9 [7] Chabanon, R.M., Shcherbak ov a, L., Lacroix-T riki, M., Agla ve, M., Zeghondy , J., Kriaa, V., Goug´ e, A., Garrido, M., Edmond, E., Bigot, L., Krastev, D.B., Brough, R., Pettitt, S.J., Thomas-Bonafos, T., Samstein, R., Massard, C., Deloger, M., T utt, A.N., Barlesi, F., Loriot, Y., Delaloge, S., T awk, M., Degern y , C., Lin, Y.-L., Pistilli, B., P asero, P ., Lord, C.J., P ostel-Vinay , S.: Auto crine interferon p oisoning 24 mediates adar1-dep enden t synthetic lethality in brca1/2-m utant cancers. Nature Comm unications 16 , 6972 (2025) https://doi.org/10.1038/s41467- 025- 62309- 5 [8] Gon¸ calv es, E., Ryan, C.J., Adams, D.J.: Synthetic lethalit y in cancer drug dis- co very: challenges and opp ortunities. Nature Reviews Drug Discov ery 25 , 22–38 (2026) https://doi.org/10.1038/s41573- 025- 01273- 7 [9] O’Neil, N.J., Bailey , M.L., Hieter, P .: Syn thetic lethality and cancer. Nature Reviews Genetics 18 (10), 613–623 (2017) https://doi.org/10.1038/nrg.2017.47 [10] Hao, Z., W u, D., F ang, Y., W u, M., Cai, R., Li, X.: Prediction of Syn thetic Lethal In teractions in Human Cancers Using Multi-View Graph Auto-Enco der. IEEE Journal of Biomedical and Health Informatics 25 (10), 4041–4051 (2021) h ttps://doi.org/10.1109/JBHI.2021.3079302 [11] F ath, M.K., Na jafiyan, B., Morov atshoar, R., Khorsandi, M., Dashtizadeh, A., Kiani, A., F arzam, F., Kazemi, K.S., Afjadi, M.N.: P otential promising of syn thetic lethality in cancer researc h and treatment. Naunyn-Sc hmiedeb erg’s Arc hives of Pharmacology 398 (2), 1403–1431 (2025) h ttps://doi.org/10.1007/ s00210- 024- 03444- 6 [12] Horlb eck, M.A., Xu, A., W ang, M., Bennett, N.K., P ark, C.Y., Bogdanoff, D., Adamson, B., Chow, E.D., Kampmann, M., P eterson, T.R., Nak amura, K., Fis- c hbac h, M.A., W eissman, J.S., Gilb ert, L.A.: Mapping the Genetic Landscap e of Human Cells. Cell 174 (4), 953–96722 (2018) https://doi.org/10.1016/j.cell.2018. 06.010 [13] Nijman, S.M.B.: Synthetic lethality: general principles, utility and detection using genetic screens in human cells. FEBS Letters 585 (1), 1–6 (2011) https://doi.org/ 10.1016/j.febslet.2010.11.024 [14] Das, S., Deng, X., Camphausen, K., Shank av aram, U.: Discov erSL: an R pack- age for multi-omic data driv en prediction of synthetic lethality in cancers. Bioinformatics 35 (4), 701–702 (2019) https://doi.org/10.1093/bioinformatics/ bt y673 [15] W an, F., Li, S., Tian, T., Lei, Y., Zhao, D., Zeng, J.: EXP2SL: A Machine Learn- ing F ramew ork for Cell-Line-Sp ecific Syn thetic Lethalit y Prediction. F rontiers in Pharmacology 11 , 112 (2020) https://doi.org/10.3389/fphar.2020.00112 [16] Huang, J., W u, M., Lu, F., Ou-Y ang, L., Zhu, Z.: Predicting synthetic lethal in teractions in human cancers using graph regularized self-representativ e matrix factorization. BMC Bioinformatics 20 (Suppl 19), 657 (2019) https://doi.org/10. 1186/s12859- 019- 3197- 3 [17] Liu, Y., W u, M., Liu, C., Li, X.-L., Zheng, J.: SL2MF: Predicting Syn thetic Lethalit y in Human Cancers via Logistic Matrix F actorization. IEEE/ACM 25 T ransactions on Computational Biology and Bioinformatics 17 (3), 748–757 (2020) https://doi.org/10.1109/TCBB.2019.2909908 [18] Hamilton, W.L.: Graph Represen tation Learning. Syn thesis Lectures on Artificial In telligence and Mac hine Learning, vol. 14, pp. 1–159. Springer, Cham (2020) [19] Cai, R., Chen, X., F ang, Y., W u, M., Hao, Y.: Dual-drop out graph con volu- tional net work for predicting syn thetic lethality in h uman cancers. Bioinformatics 36 (16), 4458–4465 (2020) h ttps://doi.org/10.1093/bioinformatics/btaa211 [20] Long, Y., W u, M., Liu, Y., Zheng, J., Kwoh, C.K., Luo, J., Li, X.: Graph con textualized attention netw ork for predicting synthetic lethality in h uman cancers. Bioinformatics 37 (16), 2432–2440 (2021) h ttps://doi.org/10. 1093/bioinformatics/btab110 [21] Y e, Q., Hsieh, C.-Y., Y ang, Z., Kang, Y., Chen, J., Cao, D., He, S., Hou, T.: A unified drug-target in teraction prediction framework based on kno wledge graph and recommendation system. Nature Comm unications 12 (1), 6775 (2021) https: //doi.org/10.1038/s41467- 021- 27137- 3 [22] W ang, S., Xu, F., Li, Y., W ang, J., Zhang, K., Liu, Y., W u, M., Zheng, J.: K G4SL: kno wledge graph neural net work for synthetic lethality prediction in h uman cancers. Bioinformatics 37 (Supplemen t 1), 418–425 (2021) https://doi. org/10.1093/bioinformatics/btab271 [23] Liu, X., Y u, J., T ao, S., Y ang, B., W ang, S., W ang, L., Bai, F., Zheng, J.: PiLSL: pairwise in teraction learning-based graph neural netw ork for syn thetic lethality prediction in human cancers. Bioinformatics 38 (Supplemen t 2), 106–112 (2022) h ttps://doi.org/10.1093/bioinformatics/btac476 [24] Zhu, Y., Zhou, Y., Liu, Y., W ang, X., Li, J.: SLGNN: synthetic lethality prediction in h uman cancers based on factor-aw are knowledge graph neural net- w ork. Bioinformatics 39 (2), 015 (2023) h ttps://doi.org/10.1093/bioinformatics/ btad015 [25] Zhang, K., W u, M., Liu, Y., F eng, Y., Zheng, J.: KR4SL: kno wledge graph reasoning for explainable prediction of synthetic lethality . Bioinformat- ics 39 (Supplemen t 1), 158–167 (2023) h ttps://doi.org/10.1093/bioinformatics/ btad261 [26] Zhang, G., Chen, Y., Y an, C., W ang, J., Liang, W., Luo, J., Luo, H.: MP ASL: m ulti-p ersp ectiv e learning knowledge graph atten tion netw ork for synthetic lethalit y prediction in human cancer. F rontiers in Pharmacology 15 , 1398231 (2024) https://doi.org/10.3389/fphar.2024.1398231 [27] Long, Y., W u, M., Liu, Y., F ang, Y., Kwoh, C.K., Chen, J., Luo, J., Li, X.: Pre-training graph neural netw orks for link prediction in biomedical netw orks. 26 Bioinformatics 38 (8), 2254–2262 (2022) https://doi.org/10.1093/bioinformatics/ btac100 [28] Li, J., Lu, X., Jiang, K., T ang, D., Ning, B., Sun, F.: T ARSL: T riple-Atten tion Cross-Net work Representation Learning to Predict Synthetic Lethality for Anti- Cancer Drug Discov ery . IEEE Journal of Biomedical and Health Informatics 29 (3), 1680–1691 (2025) h ttps://doi.org/10.1109/JBHI.2023.3306768 [29] Huang, Y., Y uan, R., Li, Y., Xing, Z., Li, J.: Struct2SL: Synthetic lethality prediction based on AlphaF old2 structure information and Multilay er Percep- tron. Computational and Structural Biotechnology Journal 27 , 1570–1577 (2025) h ttps://doi.org/10.1016/j.csb j.2025.04.012 [30] Previtali, V., Bagnolini, G., Ciamarone, A., F errandi, G., Rinaldi, F., Myers, S.H., Rob erti, M., Cav alli, A.: New Horizons of Synthetic Lethality in Cancer: Current Dev elopment and F uture P ersp ectiv es. Journal of Medicinal Chemistry 67 (14), 11488–11521 (2024) https://doi.org/10.1021/acs.jmedc hem.4c00113 [31] Shen, J.P ., Zhao, D., Sasik, R., Lueb ec k, J., Birmingham, A., Bo jorquez-Gomez, A., Licon, K., Klepp er, K., Pekin, D., Beck ett, A.N., Sanchez, K.S., Thomas, A., Kuo, C.-C., Du, D., Roguev, A., Lewis, N.E., Chang, A.N., Kreisb erg, J.F., Krogan, N., Qi, L., Idek er, T., Mali, P .: Com binatorial CRISPR–Cas9 screens for de nov o mapping of genetic interactions. Nature Metho ds 14 (6), 573–576 (2017) h ttps://doi.org/10.1038/nmeth.4225 [32] T ang, S., G¨ okba˘ g, B., F an, K., Shao, S., Huo, Y., W u, X., Cheng, L., Li, L.: Syn- thetic lethal gene pair s: Exp erimen tal approaches and predictive mo dels. F rontiers in Genetics 13 , 961611 (2022) https://doi.org/10.3389/fgene.2022.961611 [33] Ryan, C.J., Ba jrami, I., Lord, C.J.: Synthetic Lethality and Cancer - Penetrance as the Ma jor Barrier. T rends in Cancer 4 (10), 671–683 (2018) https://doi.org/ 10.1016/j.trecan.2018.08.003 [34] T ep eli, Y.I., Seale, C., Gon¸ calv es, J.P .: ELISL: early–late integrated synthetic lethalit y prediction in cancer. Bioinformatics 40 (1), 764 (2023) h ttps://doi.org/ 10.1093/bioinformatics/btad764 [35] Chen, J., P an, J., Zh u, Y., Li, J.: SLGNNCT: Syn thetic lethality prediction based on kno wledge graph for different cancers t yp es. In: International Conference on In telligent Computing, pp. 159–170 (2024) [36] T u, X., Cao, Z.-J., Xia, C., Mostafa vi, S., Gao, G.: Cross-linked unified em b edding for cross-mo dalit y representation learning. In: Ko y ejo, S., Mohamed, S., Agarwal, A., Belgrav e, D., Cho, K., Oh, A. (eds.) Adv ances in Neural Information Pro cess- ing Systems, vol. 35, pp. 15942–15955. Curran Associates, Inc., Red Hook, NY (2022) 27 [37] Kutuzov a, S., Krause, O., McCloskey , D., Nielsen, M., Igel, C.: Multimodal v aria- tional auto enco ders for semi-supervised learning: In defense of pro duct-of-exp erts. arXiv, 2101–07240 (2021) [38] Schlic htkrull, M., Kipf, T.N., Blo em, P ., Berg, R., Tito v, I., W elling, M.: Mo deling relational data with graph conv olutional net works. arXiv, 1703–06103 (2017) [39] Du, C., T eng, J., Li, T., Liu, Y., Y uan, T., W ang, Y., Y uan, Y., Zhao, H.: On uni-mo dal feature learning in sup ervised multi-modal learning. Computer Vision and Pattern Recognition (2023) [40] W ang, F.-A., Zh uang, Z., Gao, F., He, R., Zhang, S., W ang, L., Liu, J., Li, Y.: Tmo-net: an explainable pretrained multi-omics mo del for m ulti-task learning in oncology . Genome Biology 25 (1), 149 (2024) [41] Subbaswam y , A., Saria, S.: F rom dev elopment to deploymen t: dataset shift, causalit y , and shift-stable models in health ai. Biostatistics 21 (2), 345–352 (2020) h ttps://doi.org/10.1093/biostatistics/kxz041 [42] Peng, X., W ei, Y., Deng, A., W ang, D., Hu, D.: Balanced m ultimo dal learning via on-the-fly gradien t mo dulation. In: Proceedings of the IEEE/CVF Conference on Computer Vision and P attern Recognition (CVPR), pp. 18228–18237 (2022). h ttps://doi.org/10.1109/CVPR52688.2022.01772 [43] W ang, Z., Dai, Z., P o czos, B., Carb onell, J.: Characterizing and av oiding negativ e transfer. In: Proceedings of the IEEE/CVF Conference on Computer Vision and P attern Recognition (CVPR), pp. 11285–11294 (2019). https://doi.org/10.1109/ CVPR.2019.01155 [44] Cerami, E., Gao, J., Dogrusoz, U., Gross, B.E., Sumer, S.O., Aksoy , B.A., Jacob- sen, A., Byrne, C.J., Heuer, M.L., Larsson, E., Antipin, Y., Rev a, B., Goldberg, A.P ., Sander, C., Sch ultz, N.: The cbio cancer genomics p ortal: an op en plat- form for exploring multidimensional cancer genomics data. Cancer Discov ery 2 (5), 401–404 (2012) [45] Consortium, T.U.: Uniprot: the universal protein knowledgebase in 2021. Nucleic Acids Research 49 (D1), 480–489 (2020) [46] Lee, J.S., Das, A., Jerby-Arnon, L., Arafeh, R., Auslander, N., Davidson, M., McGarry , L., James, D., Amzallag, A., Park, S.G., Cheng, K., Robinson, W., A tias, D., Stossel, C., Buzhor, E., Stein, G., W aterfall, J.J., Meltzer, P .S., Golan, T., Hannenhalli, S., Gottlieb, E., Benes, C.H., Samuels, Y., Shanks, E., Ruppin, E.: Harnessing syn thetic lethalit y to predict the resp onse to cancer treatmen t. Nature Comm unications 9 (1), 2546 (2018) h ttps://doi.org/10.1038/ s41467- 018- 04647- 1 28 [47] Cerami, E., Gao, J., Dogrusoz, U., Gross, B.E., Sumer, S.O., Aksoy , B.A., Jacob- sen, A., Byrne, C.J., Heuer, M.L., Larsson, E., Antipin, Y., Rev a, B., Goldberg, A.P ., Sander, C., Sc hultz, N.: The cbio cancer genomics p ortal: An open plat- form for exploring multidimensional cancer genomics data. Cancer Discov ery 2 (5), 401–404 (2012) https://doi.org/10.1158/2159- 8290.CD- 12- 0095 [48] Kaelin, W.G.: The concept of synthetic lethality in the context of an ticancer therap y . Nature Reviews Cancer 5 (9), 689–698 (2005) https://doi.org/10.1038/ nrc1691 [49] Chicco, D.: Siamese neural netw orks: An ov erview. In: Cart wright, H. (ed.) Artificial Neural Net works, pp. 73–94. Springer, New Y ork, NY (2021). https: //doi.org/10.1007/978- 1- 0716- 0826- 5 3 [50] Alemi, A.A., Fischer, I., Dillon, J.V., Murph y , K.: Deep v ariational information b ottlenec k. arXiv, 1612–00410 (2016) 29
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment