Generative Replica-Exchange: A Flow-based Framework for Accelerating Replica Exchange Simulations
Replica exchange (REX) is one of the most widely used enhanced sampling methodologies, yet its efficiency is limited by the requirement for a large number of intermediate temperature replicas. Here we present Generative Replica Exchange (GREX), which…
Authors: Shengjie Huang, Sijie Yang, Jianqiao Yi
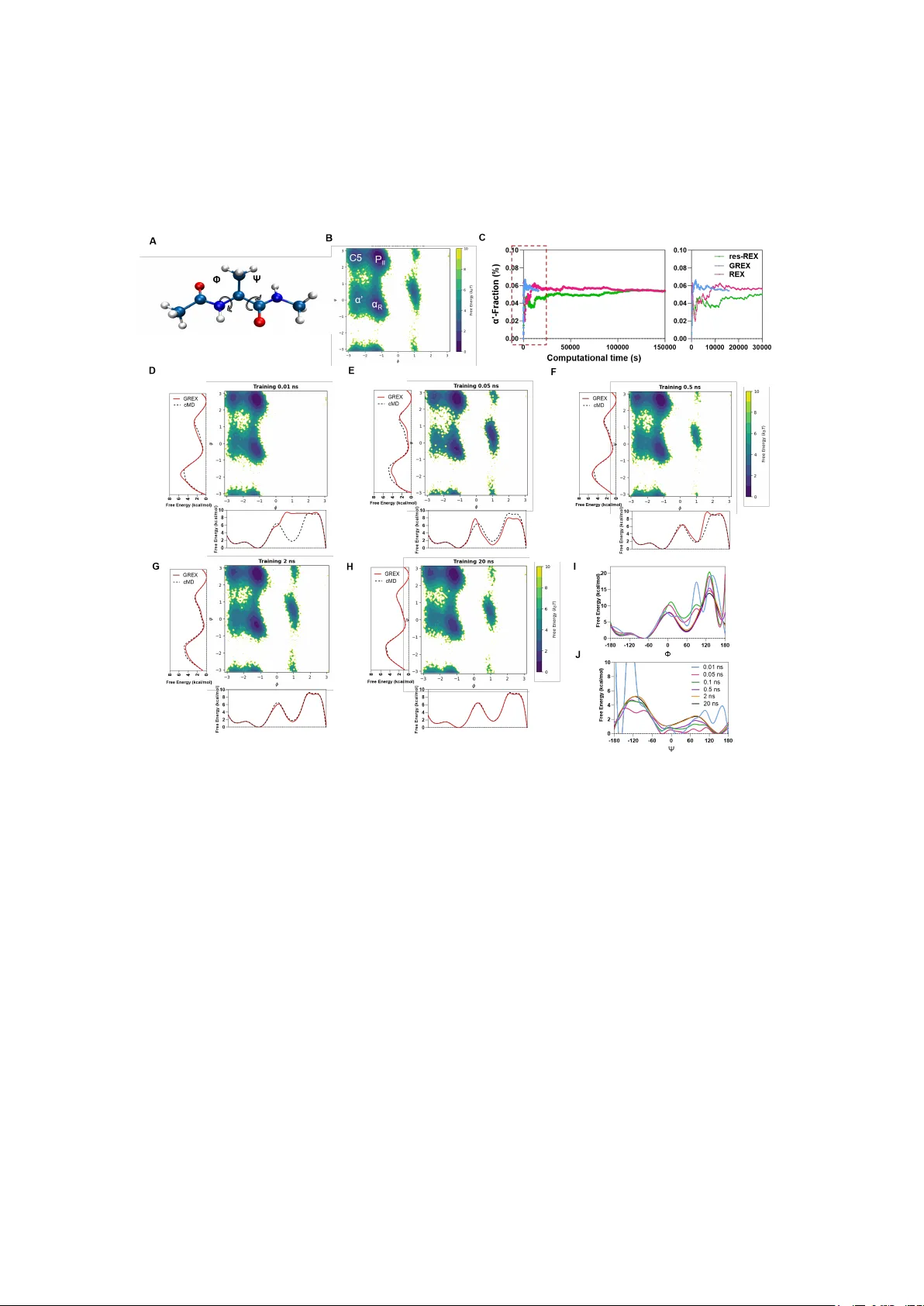
G enerat ive Replica - Exchange: A Flow - bas ed Framework for Accelerating Replica Excha nge Simulat ions Shengjie Huang 1 , Sijie Y ang 1 , Jianqiao Y i 1 , Rui Zheng 1 , Haocong Liao 1 , Muzammal Hussain 2 , 3 , Y aoquan T u 4 , Xiaoyun Lu 1 ,* , Y ang Zhou 1, * 1 State Key Laborator y of Bioactive Molecules and D ruggability Assessment, Internationa l Cooperative L aboratory of Trad itional Chinese Medicine Mode rnization and Innovative Drug Discovery of Chinese Ministry of Education (MOE), School of Pharmacy , Jinan University , #855 Xingye A venue, Guangzhou, 510632, China 2 Department of Biochemistry and Molecular Pharmacolog y , NYU Grossman School of Medicine, New Y ork, NY 10016, USA 3 Howard Hughes Medical Institute, NYU G rossman School of Medicine, New Y ork, NY 10016, USA 4 Departmen t of Theoretical C hemistry and Biology , K TH Royal Institu te of T echnology , Stockholm 1 14 28, Sweden Abst ract Replica exchange (REX) is one of the most widely used enhanced sampling methodologies , yet its ef ficiency is limited by th e requireme nt for a lar ge number o f intermediate temperatur e replicas. Here we present Generative Replica E xchange (GREX), which integrate s deep generativ e models into th e REX framework to eliminate th is temperatur e ladder . Drawing inspiration from reservoir replica exchange (res - REX), G REX utilizes trained norma lizing flows to generate h igh - tempera ture configurations on demand and map them directly to the tar get distribution using the potential energy as a constraint, withou t requiring t arget - temperature training data. This approach reduces production simul ations to a single replica at the tar get temperature while maintaining thermodynamic rigor through Metropolis exchange acceptance . We validate GREX on three benchmark systems of increasing complexity , highlighting its superior ef ficiency and practica l applicability for molecu lar simulations. 1. Introduction Molecular d ynamics (MD ) is a powerful computat ional techn ique for simu lating complex phenomena at the atomic level 1 . Recent advances in computing hardware and software have signific antly extended the accessibl e simulation timescales, r anging from nanoseconds to microseconds, thereby enabling researchers to investigate dynamic processes, such as small pep tide folding or ligand binding 2,3 . Despite these advancements, achieving efficient sampling of th e eq uilibrium distrib ution remain s a long- standing chal lenge in mole cular simu lations. This difficu lty arises bec ause systems often become trapped in lo cal ener gy mini ma, which lim its their abil ity to explore configurational space in short simulation t imes . Furthermore, achieving suf ficient samplin g on long time scales rema ins computation ally demandin g, especially for lar ge systems o r rare - event processes. T o address th ese challenge s, a variety of e nhanced sampling techniques have been developed to facilita te the traversal o f energy barriers and accele rate the explora tion of configurational space . These methodologies are generally categorized into two frameworks: collective variable (CV) - based and CV - free approaches . CV -based methods, such as metadynamic s 4,5 and umb re lla sampling 6 , rely on the selection of a few critical degrees of f reedo m to bias the simulation a nd promote sampling along relevant coo rdinates . However , the efficacy of these methods is heavily d ependent on the "quality" o f the selected CVs 7 . CV - free strategies , such as replica exchange molecular d ynamics (RE X) 8,9 , facilitate barrie r crossing with out relying on predefine d CV s. By allowing replicas to swap configurations between neighboring distributions, the system can ef fectively escape loc al minima. Driven by the need for better scalability , several variants of REX have emer ged such as Hamiltonian REX (HREX) 10,11 , Replica Exchange Solute T empering (REST) 12 , and reservoir replica exchange (res - REX) 13 . In particular , in res - REX a reservoir of configurations is pre - generated at a high temperature, a nd exchange attempts are performed b etween the reservoir and replicas. This method greatly enhances the convergence r ate and has been successfully used for a variety of small mo lecules and peptides such a s leucine tripeptide 14 , T rp zip2 15 , A β 21 - 30 peptide 16 . Nevertheless, conve ntional REX and its v ariants still req uire a lar ge number of intermediate t emperature replicas to maintain ad equate exchange accep tance probabilities, resulting in significant computational overhead that grows with system size and complexity . In this work, we present G enerative replica excha nge (GREX) , an enhanced sampling framework th at integrates n ormalizing flo ws into the conventional replica exchange framework to eliminate the need for a dense temperature ladder , reducing production simulation t o a single re plica at the t ar get temperature . T o demonstrate its ef fectiveness and generality , we apply GREX to three benchmark systems of increasing complexity: a double - well poten tial, alanin e dipeptide in explic it water , and the 10 - re sidue mini - pro tein chignolin. GREX achieves conver gence speedups of 5 - to 10 - f old relative to conventional REX while maintaining thermodynamic accuracy , with computational advantages that grow with system complexity . 2. Methods Reservoir Replica Exchange Simulations (r es - REX) W e briefly summarize the key a spects of REX and res - REX as they are r e lated to the present study . In standard REX , replicas ( to ) at dif ferent temperatu res are simulated simultaneously and independently for a chosen number of M D steps. Exchanges between a pair of replicas and (typically neighboring r eplicas where j = i + 1 ) are attempted periodically . T he acceptance probability for an exchange is defi ned by the Metropolis criterion, = min 1, exp 1 1 , (1) where and are the temperature s of and and and are the potential energies of the configurations at and , respectively . If the exchange is accepted, then the bath temperatures of these re plicas will be swapped. Otherwise, if the exchan ge is rejected, t hen each rep lica will continu e on its curren t trajectory wi th the same bath . Reservoir Replica Exch ange Simulations (res - REX) is a variant of standard REX. In this method, the highest temperatur e replica ( ) is replaced with a reser voir ( ), which is a set of structures previously ge nerated from MD simulations performed at the high tempera ture . The exchange attempt is mad e between the structure of and a randomly selected structure from the reservoir . If the exchange is accepted, the coordinates from are sent to replica and the chosen reservoir structure is left in the reservo ir , as it is assumed that the rese rvoir constitutes a complete represen tation of the ense mble and th at the inclu sion of the new coordin ates will h ave a neglig ible ef fect on the reservo ir . Concurrently , s tandard REX simulations were used for each of the lower temperatures (repl icas to , to ) and exchanges are attempted o n the basis of th e same crit erion as used fo r REX (eq 1). The standard REX process of s imulation acr oss the temp erature ladd er provide ex ploration/re finement of the basins present in the reservoir and also reweights the probability of observing these structures at diffe rent tempera tures. The e xchanges between and and the REX for to are repe ated multiple times and thermally reweighted man y times during the simulation, resulting in converged Boltzmann - weighted ensemb les at all temperat ures. GREX framewo rk GREX extends the res - REX concept by replacing the static high - temperature reservoir wi th a Generator Flow (GF) that generate configurations reproducing the high- temperature d istribution , and in troducing a Converter Flow (CF) that maps these configuratio ns directly to the tar get t emperature . T ogether, they eliminat e the intermediat e replica ladd er and redu cing the s imulation to a single tar get - t emperature replica. The GREX workflow has two main stages, as illust rated in Figu re 1 . In Stage 1 (training) , a short M D simulation is performed at the high te mperature to collect configurations . The se samples are used to train the GF to repr oduce the high - temperature dis tribution , while the CF is trained with the potential ener gy function as a physical constraint to learn the mapping from to the tar get distribution . In Stage 2 (production) , the GF generates high temperature configurations , which are then mapped to the tar get t emperature via the C F. Exchan ge attempts between t he mapped configurations and the ongoing MD trajectory are evaluated using the Metropolis criterion (eq 1) , ensuring thermodynamic consistency at the target temperature thro ughout the simulat ion. Figure 1. The GRE X workflow . The blue block represents t he generator flow (GF), and orange blocks represent the converter flow (CF). Red a rrows indica te State 1, while black arrows indicate St ate 2. Both the GF and CF are implemented as normalizing flows (as shown in F i gure 2) , which are t rainable in vertible neu ral networks ( and ) th at enable exact and tractable co mputation of p robability d ensities . Generator Flow (GF) The GF is a norma lizing flow th at learns to tra nsform samples from a prior Gaussian distribution ( ) into configurations dist ribut ed according to the high temperature Boltzmann distr ibution ( ) . O nce trained, the G F serves as an on - demand generator of high - temperatu re configurat ions, replacing the role of the pre - generated reservoir in res - REX. T he GF implemen ts this mappin g through a ser ies of reversib le coordin ate transformations 17 – 19 , expressed as follows: = ( ; ) (2) = ( ; ) (3) where, and represent samples fro m the high - temp erature distrib ution and the prior distrib ution, respec tively , and represents the trainable paramete rs of GF . Each transformation is assoc iated with a Jacobian matr ix: J ( ; ) = [ ( ; ) x , … , ( ; ) x ] (4) J ( ; ) = [ ( ; ) , … , ( ; ) z ] (5) The absolute value of the Jacobian determinant, det ( ; ) quantifies the degree of volume expansion or contraction induced by the transformation. Owing to the invertibility of the mapping, probability densities can be transformed between differ ent spaces as ( ) = ( ; ) | ( ; ) | (6) ( ) = ( ; ) | ( ; ) | (7) where denotes the distributions generated by the neural network in the corresponding spaces, which dif fer from the target distributions . The GF is built upon the af fine coupling layers of the RealNVP algorithm 20 proposed by Dinh et al, following t he design of Boltzmann generators 21 . Specifically , the input variables are partitioned into two channels, denoted as = ( , ) and = ( , ) . In each affine coupling layer , one part remains unchanged, while the other part is transformed using a scaling network and a translation network , both implemented as non-invertible neural networks. The correspo nding transformations are given by (( , ); ): = , = exp ( ( )) + ( ), (8) (( , ); ): = , = ( ( )) exp ( ( )), (9) where denotes element - wise multiplic ation. T o reconstruct the high - temperature distribution ( ) , the GF is trained using a “training by example” strategy 21 . High - temperature trajectory data from Stage 1 serve as the training data, and t he network parameters are opti mized by min imizing the negative log-likelihood (NLL) loss function 22,23 : L = ~ ( ) log p ( ; ) det J ( ; ) ( 10 ) Converter Flow (CF) T he CF is a normalizin g flow that map the hi gh - temperatu re distributi on ( ) directly to the ta rget low - temperature distrib ution ( ) , e liminating the need fo r intermediat e - temperature replicas. Following the LREX m ethod introduced by Invernizzi et al. 24 , t he CF achieves this distributio n transformatio n through : q ( ) = p ( ; ) |det J ( ; ) | ( 11 ) q ( ) = p ( ; ) |det J ( ; ) | ( 12 ) where represents samples from the low - temper ature distrib ution, and denotes the trainable parameters of CF . Unlike the GF , which is trained using a “train ing by example” strategy , the CF faces the major challenge th at samples from the tar get low - temperature d istribution ( ) are generally unavailable. T he C F therefore adopts a “training by ener gy” strategy 24 , in which the system’ s potential energy function ( ) is incorporated as a physical constraint. Specifical ly , the network parameters are opt imized by the Kullback – Leibler divergence (KL D) between the generated and target distributions 24 : L = ~ ( ) [ 1 k T ( ; ) 1 k T ( ) log |det J ( ; )|] ( 13 ) where is the Boltzmann constan t. Figure 2. Architecture of Gen erator Flow (G) an d Converter F low (C) A lanine dipeptide a nd Chignolin The structure of alanine dipeptide was modelled using AmberT ools23 25 and the structure of chignolin (PDB ID : 1 U AO ) was obtained from the RCSB Protein D ata Bank 26 . Both systems were simulated using OpenMM 27 , with the AMBER f f14SB force field 28 for th e protein an d the TIP3P 29 model for the solvent. Each system w as solvated in a cub ic simulation box with a min imum distan ce of 1.2 nm b etween the sy stems and the box boundaries. After solvat ion , an energy minimization process was carried out to remove the atomic c lashes and optimize th e geometry of al l molecules. The simulations were run in the constant n umber (N), pressur e (P), and tempe rature (T) (NP T) ensemble at 300 K and 1 bar using MonteCarlo algorithm . T he Particle Mesh Ewald (PME) 30 method was used for calculating long - range electrostatic i nteractions. The LangevinMiddleIntegrator 31 method was used with a time step of 2 fs. Following the simulation s , the MDtraj package 32 was used to ext ract protein atoms, and the trajectorie s were aligne d based on CA atom pairs to generate the da taset. For alanine dipeptide, the backbone dihedral angles Φ and Ψ were calculated. For chignolin, RMSD of simulatio n frames relativ e to the PDB native structure was ca lculated for the prot ein C α atoms with the terminal residues Gly1 and Gly10 excluded. 3. Results T o demonstrate the efficiency and capabilities of GREX, we apply it to three model systems of increasing co mplexity: parti cle in double-w ell potential, alanine dipeptide, and chignolin. 3.1 Double - well po tential T o validate the algorithmic correct ness and the sampling efficiency of GREX , we first applied it to a particle moving with Langevin dynamics in a double - well poten tial 33 ( see SI for details ). The tar get temperature was se t to T = 1, at which transitions betwee n the two basins are exceeding ly rare in cMD simulations. A separate cMD simulation at a high er tempera ture (T = 5), where barr ier crossing occu rs much more freque ntly (Figure S1A), was performed to generate training data for both the generator and converter flows. As illustrate d in Figure 3A , at the tar get tempe rature, the p article remains trapped in the left basin (x < 0) throughout the 20 ns cMD simulation . In contrast, frequent transitions between the two basins were observed in b oth GREX and REX (Figure s 3B and 3C). T he one - d imensional marginal probability distr ibutions along the x -coordinate demonstrate that both methods successfully recover the free ener gy dif ference between the two basins ( Δ G = 2.5 kcal/mol) in exce llent agree ment with the an alytical solu tion (Figure S1B), confirming that GREX reproduces the correct equilibrium free energy landscape. Next, to evaluate the efficiency of GR EX in comparison with REX, we followed the protocol of Invernizzi et al. by extending the double - well p otential system to N dimensions 24 . In conventional REX, the numb er of intermed iate temperature re plicas required to maintain adequate exch ange acceptance ratios scales wi th system size, leading to a dramatic increase in computationa l cost as dimen sionality g rows. In contrast, GR EX requires samplin g only at the tar get temperature, with the gener ator and converter flows establish the connection to the high - temperature distribution . To quantify th e resulting d if ference in sc alability , we measured th e computat ional time required to reach conver gence for systems of varying dimensionality , using the basin occupation fraction ( the fraction o f time with x > 0) as a conver gence indi cator . Fig ures 3D– 3H present the time evolution of this quant ity for systems with different d imensions. F or small systems (e.g., N = 2 ), GREX and REX perform comparably , with both methods requiring approximately 50 seconds to approach equilibrium . A s the system size increases, however , the conver gence ti me required for R EX increases drama tically , reaching approximately 300 s, 600 s, and 2000 s for N = 64, 512, and 1024, respectively . In contrast, GREX simulations reach equilibrium with minimal dependence on syste m size, maintaining convergence times ar ound 200 s for N ≤ 1024, with only modest increases observe d for l ar ger systems (F igure 3I). W e also examined the ef fect of system size on exchange a cceptance ratios. In RE X, maintaining reasonable accep tance ratios necessitates a pro portional increase in the number of intermediate repl icas as system size grows. Conversely , GREX consistently maint ains high accep tance ratios (> 20 %) acros s all system s izes without req uiring additional re plicas (Figu re 3J), demon strating sup erior scalabi lity . T aken together , t hese results establish that GREX of fers substantial computational advantages over convention al REX, pa rticularly for high - dimensi onal systems where the replica ladder approach becomes prohibitively expensive. Figure 3. Evaluation of the double - well potential model . (A - C) T he free ener gy surface and represen tative traject ories obtaine d from cMD, R EX and GREX simulat ions at the tar get temperature ( T = 1). (D - H) T ime evo lution of the basin occupation fraction (defined as the fraction of configurations with x > 0) as a function of computational time for N - dime nsional sy stems with in creasing di mensionality ( N = 2, 16, 64, 512, and 1024). (I) Computational time required for convergence as a function of system dimensionality N. (J) Exchan ge acceptance ratios increase with N for REX simu lations with dif ferent numb ers of replica s ( R ) and for GREX. 3.2 Alanine Dipe ptide W e next applied the method to alanine dipeptide in explicit water , a widely use d benchmark system whose conformationa l landscape is well - characterized by the t wo backbone dihedral angles Φ and Ψ ( Figure 4A) 34,35 . Five independent 2 μ s cMD simulations at 300 K were used to generate a refer ence free ener gy surface against which all methods were compared . These method included conventional REX using 32 replicas spanning 300 - 1000 K, res - REX using 31 replicas supplemented with a 1000 K reservoir constructed from 20 ns cMD data, and G REX using a single 300 K replica with generative models trained on the same 20 ns high- temperature t rajectory . All three methods reproduced the reference FE S obtained from cMD simulation s ( Figure 4A and S2), confirming tha t GREX achieves res ults comparab le to replica - based approaches de spite using only a single p roduction replica. T o quantify sampling efficiency , we monitored the converg ence of conformatio nal basin s {C5, P II , α R , and α’} as a function of computational time. Fo r the low- ener gy basins {C5, P II , and α R }, GREX converged within approximately 5000 s, roughly half the time required by both res - REX and REX (~10000 s; Figure S3). The advantage was even mor e pronounced for the high -energy α’ basin: GREX again conver ged within ~5000 s, whereas both res - REX and REX required ~30000 s , corresponding to an approximately sixfold dif ference (Figure 4C). Because GREX training depends entirely on high - tempera ture simulation da ta, the amount of training data required is a key practical consideration . T o investigate this, we trained GREX models on 1000 K trajectories ranging from 0.01 ns to 20 ns and performed 100 ns production simulations for each case. W ith only 0.01 ns of training data, GREX failed to rec over the reference free ener gy landscape, producing a distribution that was entirely absent in the region Φ > 0° ( Figure 4 D ). This failure arise because 0. 01 ns trajecto ry does not sample that region of co nformationa l space. Once the training trajectory covered the relevant conformational regions, however , GREX accurately reproduced the 300 K reference distribution. Notab ly , this did not require th e high- temperature FES itself to be converged . M odels trained on 0.05 ns and 0.5 ns trajectorie s , whose high - tempe rature FES remain far from the conver ged reference , still reproduce the 300 K reference landscape with reasonable accuracy (Figures 4 E to 4J ). W ith 2 ns of training data, GREX already matched the accuracy of both res - REX and GREX model trained on 20 ns of dat a (Figures 4G and 4H). Accounting for this training cost, the total computational expense of GREX was approxi mately ~16000 s (2 ns training plus 100 ns single - replica production ), compared with approximately ~157000 s for conventional REX (32 replicas × 100 ns), yielding an overall speedu p of roughly 10-fold. Figure 4. Evaluation of the alanine dipeptide model . (A) Illustration of the b ackbone dihedral angles Φ and Ψ of the alanine dipeptide. (B) R eference free ener gy surface at 300 K as a function of Φ and Ψ, obtained from cMD simul ations. Majo r conformation al basins {C5, P II , α R , and α’} are labeled. (C) Con verg ence of the α ’ basin population as a function o f computatio nal time for G REX, res - REX, and conventional REX. The dashed box highlights the early - time regime. (D - H) Free ener gy surfa ces reconstructed from GREX simulations at 300 K using high - temperatu re (1000 K) cM D training d ata of different lengths: (D) 0.01 ns, (E) 0.05 ns, (F) 0.5 ns, (G) 2 ns, and (H) 20 ns. (I) and (J) One - dimensional free energy profiles along Φ and Ψ obtained from high - temperature cMD simulation s with dif ferent t rajectory le ngths. 3.3. Chignolin T o further assess the applicability of GREX to more complex biomolecu lar systems, we examined chignolin, a 10 - residue mini - prot ein (GYDP ETGTWG) that folds in to a stable β - hairpin struc ture. Chignolin has been extensively characterized both experimentally and computationally , making it an ideal benchmark for enhanced sampling methods 36 – 39 . We generated training data from a 50 ns MD simulation at 500 K, with the target temperature set to 300 K. Three independent 10 μ s cMD simulations at 300 K served as reference data, and REX simulations with 24 replicas (300 – 500 K) were used for compariso n . W e first charact erized the overall conformational landscape by comput ing the t wo - dimensional FES as a function of backbone RMSD and radius of gyration (Rg). The reference F ES from 10 μ s cMD simulations (Fi gure 5 A ) reveal s a basin at (RMSD ≈ 1.2 Å, Rg ≈ 5.8 Å). Both GREX and REX successfully re produced this landscape (Figures 5 B and 5 C), confirming that both methods recover the correct equilibrium structural en semble at th e targ et temperatu re. We then assessed folding/unfolding transitions using backbone C α RMSD relative to the experimental structure. In the 10 μ s cMD simulations, chignolin exhibited characteristic two - sta te behavior , remaining in either the folded (RMSD < 2.0 Å) or unfolded states (RMSD > 4.0 Å) for hundreds of nanoseconds to microseconds (Figur e 5D ). In c ontrast, both GREX and REX sampled frequent folding/unfolding transitions within the 100 ns simulation s (Figure 5 E–F ). Notably , GREX also accessed near - native conformations with a minimum RMSD of 3.0 Å (Figure 5G ), prompting a quantitative assessment of folding accu racy . Therefore, to evaluate the accuracy of GRE X, we estimate the Gibbs free energy differ ence G between the unfolded and the folded states: = 1 ( 14 ) where is the Boltzmann constant, temperatu re is set to 300 K, is the probabilities of chignolin’ s conformations in the folded conformations . Honda et al. 40 measured the folding free energy G using circular dichroism and NMR techniques obtaining values between 0.2 6 and 0.4 5 kcal/mol at 298 K. GREX estimate o f = 0.47 kcal/mol that is in clos e agreement with the experimental range (Figure 5H, dashed line) , confir ming that GR EX accurately captu res the thermod ynamic stabi lity of chignolin at the targ et tempera ture. W e further compared conver gence ef ficiency by monitor over simulation time for all three metho ds (Figure 5 H ). In cMD simulatio n, converg es after approximately 600000 s (10 μ s) to a value of about 0. 43 kcal/mol (Figure 5 H) . In REX, converg ence reached 0.73 kcal/mol after 200000 s (24 replicas × 100 ns), significantly outside the experimental range , indicating incomple te conver gence despite t he higher computation al cost. GREX , in contrast, achieve d convergence to 0.47 kcal/mol with in only 20,000 s (100 ns), consistent with both the experimental value and the long- time cMD simulation data . These results demon strate that GRE X achieves superior conver gence ef ficiency related to REX while maintaining acc uracy compa rable to long - timescale cMD. Figure 5. Evaluation of the chignolin model. (A - C) Tw o - dimens ional FES as functions of RMSD and Rg from 10 μ s cMD (F), 100 ns GREX (G), and 100 ns REX (H) simulations. (D -F ) Backbone CA RMSDs of cMD (red), GREX (blue), and REX (green). (G) Starting from an unfo lded initial s tructure, GR EX reached a minimum RMSD of 0.30 nm, corresponding to native conformations (blue). ( H ) Convergence of the calculated as a function of simulation time. The dashed line in dicates the experimental referen ce. Beyond global folding thermodynamics, we als o examined local conformational features through the FES of backbone dihedral angles ( Φ₂, Ψ₂) at the centr al glutamate residue , following previous enhanced sampling studies 41 . The 10 μ s cMD referen ce simulation i dentified thr ee local minima in this la ndscape (Fig ure 6A): two dominant minima at ap proximately ( Φ₂ ≈ −70°, Ψ ₂ ≈ 140°) and ( Φ₂ ≈ −150°, Ψ₂ ≈ 140°) , and a third minim um at ( Φ₂ ≈ −70°, Ψ ₂ ≈ −30°) sepa rated by a free ener gy barrier of ~4 kcal/mol. This substantial barrier preve nts the metas t able basin from be ing sampl ed in a 100 ns cMD simulation (Figure 6B). B oth GREX and REX wer e able to capture this metastable basin within 100 ns (Figures 6C and 6D) , with s imilar behavior observed for FESs of other backbone dihedrals (Figure S 4 ). Importantly , GREX achieved this with only 34000 s (50 ns training + 100 ns production), representing a 5- fold speedup over REX ( 170000 s for 24 replicas × 100 ns) and an 18- fold speedup rel ative to the 1 0 μ s cMD reference. Figure 6. Tw o - dimensional FES of chignolin as functions of backbone dihedrals ( Φ 2 and Ψ 2 ) computed from (A) 10 μ s cMD , (B) 100 ns cMD, (C) 100 ns REX, and (D) 100 ns GREX simulations . 4. Discussion and Conclusion s In this study , we introduced GREX , an enhanced sampling framework that integrates no rmalizing f lows with t he replica exchange framework to eli minate the need for a dense temperature ladder . Instead of relying on the conventi onal online relay of configurations through intermediate replicas , GREX em ploys a pair of trained normalizing flo ws: a Generator Flow that learns the high - temperature d istribution fro m short MD data, and a Converter Flow that maps high - temperature configura tions directly to the target temperature using the potential energ y function. Generated configurations are introduced into the simulation through Metropo lis exchange attempts, which correct a ny inaccurac ies in the learned transformatio n s and ensure that the resulting ensemble remains fr om the tar get Boltz mann distribu tion. V alidation across three benchma rk systems of increasing complexity , from a two -dimensional model poten tial to the 10 - residue mini - protein chignolin, demonstrates that GREX is both thermodynamically rigorous and broadly ef fective, achieving 5- to 10 - fold f aster convergence speedups rela tive to conventional REX. Although GREX builds on ideas fro m reservoir - based REX and normalizing flow methods , it address es key limitation s of both . Reservoir REX 15 pre- gene rates a physical ensemble at high temperature and substitu tes it for the hig hest - temperature running replica, b ut still requires a full temperature ladder of N −1 intermed iate replica s to connect the reservoir to the tar get tempe rature . GREX advances this concept further by replacing the fixed res ervoir with a Generator Flow that learns the h igh - temperatu re distribution and produces new configurations on demand, while a Converter Flow maps them directl y to the tar get tempera ture. This eliminate s the intermed iate ladder enti rely , reducing th e simulation to a single replic a at the tar get tempe rature. Bo ltzmann Generators 21 use normalizing flows to bridge between distributions, but they are tra in ed on target - temperature data and rely on importance reweighting to ensure thermodyna mic rigor . For complex biomo lecular sys tems , however , obtaining suf ficient low - temperature tra ining data is itself the ce ntral sampling challenge , and a model trained on limited low- temperature co nfigurations risks inheriting th e very sampling gaps it is intende d t o overcome. GREX circumvents this limitation by training on the high - temperature ensemble, which is re adily easily explored in short MD simulations, and by using the potential energy fu nction rather than target - temp erature samples to guide th e Converter Flow . The learned replica ex change (LREX) method 24 shares with GREX both the use of normalizing flows and a Metropolis criterion to ensure thermodynamic rigor . However , it requires parallel high - temperature simulations to run concurrently throughout the production stage , with the flow updated iteratively as new samples accumul ate. GREX , in contrast, fully decouples training from produc tion, so th at a single o fflin e training st age suf fices fo r arbitrari ly long production runs without additional high- temperat ure replicas. T wo limitatio ns of the current imp lementation of GRE X should be noted. First, the performance of GREX depends on adequate sampling of rel evant co nformational states in the high - temperatu re simulations . Configurations that are not sampled at high temperature cannot be learned by the generative model and will therefore be absent in the resulting ensemble at the tar get temperatu r e, as illustrated by the missing conformational basin observed when only 0.01 ns of training data was used (Figure 4D) . Second, GREX share d a limitation common to oth er normalizin g flow -based methods: the scalabili ty of normaliz ing flow s to lar ger and more complex syst ems remains an open challenge 24 . Although GREX has been successfully applied here to systems with explicit solvent, extending the framework to significantly larg er complex biomolecular systems may require deeper and mo re expressive flow architectures . Su ch models are typically more demanding to train and m ay introduce additional computational overhead. Encouragingly , normalizing flow methods are advancing rapidly , and improvements in flow architecture s or training efficiency can be readi ly incorporated into GREX without altering the unde rlying exchange framework. Looking ahead , key extensio ns of GREX include adaptive strateg ies to detec t and address gaps in high - temperature sampling, as well as transfer learning across structurally related syste ms to reduce pe r - sy stem training cost s . In tegration with CV - based methods also of fers a promising direction , w ith GREX providing broad conformatio nal diversit y while CV - gui ded simulations refin e free energy estimates along specific reaction coordinates. In summary , GREX provides a p ractical route to efficient en hanced sampling by replacing the temperature ladder with learned normalizing flows, achieving accuracy comparable to conventional REX at a fraction of the computational cost, with advantage s that grow as system complexi ty increases. NOTES The code used in this work is publicly available at https://github.com/Hhuangsj/GREX . The normalizing flows have been implemented using the library https://github.com/noegroup/bgflow . ACKNOWLEDGMEN T The authors gratefully acknowledge the financial support from Guangdong Basic and Applied Basic Research Foundation (2025A15150121 14), Open Project of State Key Laboratory of Respiratory Disease ( SKLRD - OP - 202506), and Changjiang Scholars A ward Program of Ministry o f Education . S.H. would like to acknow ledge the s upported by the Excellent Graduate Student Cultivation Program of Jinan University (2025CXY353) and thank Dr . Michele Invernizzi for his helpful discussions on LREX simulations. The computational resources were provided by the High - Performance Public Computing Service Platform of Jinan University . REFERENCE (1) Karplus, M.; McCammon, J. A. Molecular Dynamics Simulations of Biomolecules. Nat Struct Mol Biol 2002 , 9 (9), 64 6 – 652. (2) Lindorf f - La rsen, K.; Pi ana, S.; Dror , R . O.; S haw , D. E. Ho w Fast - Foldi ng Prote ins Fol d. Science 201 1 , 334 (60 55), 517 – 520. (3) Liu, Y .; T an, J.; Hu, S.; Hussain, M .; Qiao, C.; T u, Y .; Lu, X.; Zhou, Y . Dynamics Playi ng a Key Role in the Co valent B inding o f Inhibi tors to Focal Adhesion Ki nase. J. Chem. Inf. Model. 2024 , 64 (15), 6053 – 6061. (4) Laio, A.; Parrine llo, M. Escapi ng Free - Ener gy Mi nima. Pr oceedings of the Nati onal Academy of Sciences 2002 , 99 (20), 12562 – 12566. (5) Ray , D.; Parrin ello, M. Kinetics from Metadynamics : Principles, Ap plications, and Outlook. J. Chem. The ory Comput . 2023 , 19 (17), 5649 – 5670. (6) T orrie, G. M.; V alleau , J. P . Nonphysical Sampling D istributions in Monte Carlo Free - Ener gy Estimation: Umbrella Sampling. Journal of C omputatio nal Physic s 1977 , 23 (2), 187 – 199 . (7) Pietrucc i, F . Strategi es for t he Expl oration of Free Ener gy Lands capes: Unity i n Dive rsity an d Challenges Ahead. Reviews in Physics 2017 , 2 , 32 – 45. (8) Earl, D. J.; Deem, M. W . Parallel T empering: Theory , Applications, and New Perspectives. Phys. Chem. Che m. Phys. 2005 , 7 (23), 3910 – 3916. (9) Swendsen, R . H.; W ang, J. - S. Replica Monte Carlo Simulation of Spin -Glasses. Phys. Rev . Lett. 1986 , 57 (21), 2 607 – 2609. (10) Sugita, Y .; Kitao, A .; Okamoto, Y . Multidimensional Replica - Exchange Method f or Free - Ene r gy Calculations. The Journal of Chemical Physics 20 00 , 11 3 (15), 6042 – 6051. ( 11 ) Fukunis hi, H.; W atanabe, O. ; T akada, S. On the Hamilt onian Repl ica Excha nge Metho d for Efficient Sampling of Biomolecular Systems: Application to Protein Structure Prediction . The Journal of Chemical Physics 2 002 , 11 6 (20), 9058 – 9067. (12) Liu, P .; Kim, B.; Friesn er , R. A.; Berne , B. J. Replica Exchange with Solute T emp ering: A Method for Sampling Biologica l Systems in Explicit W ater . Pr ocee dings of t he Natio nal Acade my of Sciences 2005 , 102 (39) , 13749 – 13754. (13) Kasavajha la, K.; Simmerling, C. Exploring the T ransferab ility of Replica Exchange Stru cture Reservoirs to Accelerate Generation of Ensembles for Alternate Hamiltonians or Protein Mutations. J. Chem. Theory Com put. 2023 , 19 (6), 19 31 – 1944. (14) L yma n, E.; Ytreber g, F . M.; Zuckerma n, D. M. Resoluti on Exchange Simulat ion. Phys. Rev . Lett. 2006 , 96 (2), 02 8105. (15) Okur , A.; Roe, D. R.; Cui , G.; Hor nak, V .; Simmerli ng, C. I mprovi ng Conve rge nce of R eplica - Exchange Simulat ions thro ugh Coupl ing to a High - T emper ature Structure Reservoir . J. Chem. Theory C omput. 2007 , 3 (2 ), 557 – 568. (16) Ruscio, J. Z.; Fawzi, N. L.; Head - Gordon, T . How Hot? Systematic Conver gence of the Replica Exchange Method Using M ultiple Reservoi rs. (17) Rezende, D. J.; Mohamed, S. V ariational Inference with Normalizing Flows. arXiv June 14, 2016. (18) Kingma, D. P .; Dhariwal, P . Glow: Generative Flow with Invertib le 1x1 Convolutions. arXiv July 10, 2018. (19) Grathwohl, W .; Chen, R. T . Q.; Bet tencourt, J.; Sutskeve r , I.; Duvenaud, D. FFJORD: Free - Form Continu ous Dynam ics for Scalabl e Revers ible Ge nerative Models. arXiv October 2 2, 2018 . (20) Dinh, L.; Sohl - Dickstein, J. ; Bengio, S. Density Estimation Using Real NVP . arXiv February 27, 2017. (21) Noé, F .; Olss on, S.; Köhler, J.; W u, H. Boltzmann Generators: Sampling Equilibr ium States of Many - Body Systems with Deep Learn ing. Science 2019 , 365 (6457), e aaw1 147. (22) T abak, E . G .; V anden - Eijnd en, E. Density Estimation by D ual Ascent of the Log - Likelihood. Comm. Mat h. Sci. 8 (1), 2 17 – 233. (23) Dinh, L.; Krueger , D.; Bengio, Y . NICE: Non - Li near Inde pendent C omponent s Estim ation. a rXiv April 10, 2015. (24) Invernizz i, M.; Kraemer , A.; Clementi, C.; No e, F . Sk ipping the Replica Exchange Ladder with Normalizi ng Flows. J. P hys. Chem. Lett. 2022 . (25) Case, D. A.; Aktulga, H. M.; B elfon, K.; C erutti, D. S.; Cisneros, G. A.; Cruzeiro, V . W . D.; Forouzesh, N.; Giese , T . J.; Göt z, A. W .; Gohlke, H.; Izadi, S.; Kasavaj hala, K.; Kaymak, M . C.; King, E.; Kurtzman, T .; Lee, T . - S.; Li, P .; Liu, J.; Luchko, T .; Luo , R.; M anathunga , M.; M achado, M. R.; N guyen, H. M.; O’Hea rn, K. A.; Onufriev , A. V .; Pan, F .; Panta no, S.; Qi, R.; Rah namoun, A.; Risheh, A.; Schott - V erdugo, S.; Shajan, A.; Swails, J.; W ang, J.; W ei, H.; W u, X.; Wu, Y .; Zhang, S .; Zha o, S.; Zh u, Q.; Cheatham, T . E. I.; Roe, D. R.; Ro itberg, A.; Simmerlin g, C.; Y ork, D. M.; Nagan, M. C. ; Merz, K. M. Jr . Amb erT ools . J. Chem. Inf. Mod el. 2023 , 63 (20), 6183 – 6191. (26) Berman, H. M.; W estbrook, J.; Feng, Z.; Gill iland, G.; Bha t, T . N.; W eissig, H.; Shindya lov , I. N.; Bourne, P . E. T he Prote in Data B ank. Nucleic Acids Resear ch 2000 , 28 (1), 235 – 242. (27) Eastman, P .; Galvelis, R. ; Peláez, R. P .; Abreu, C. R. A.; Farr , S. E.; Gallicchio, E.; Gor enko, A.; Henry , M. M.; Hu, F .; Huang, J.; K rämer , A.; Michel, J.; M itchell, J. A.; Pande, V . S.; Rodrigues, J. P . ; Rodriguez - Guerra, J.; Simm onett, A. C.; Singh, S .; Swail s, J.; T urner , P .; W ang, Y .; Zhang, I.; Chodera, J. D.; De Fabr itiis, G.; Markland, T . E. OpenMM 8: Molecular Dyn amics Simulation with Machine Learning Potentials. J. Phys. Chem. B 2024 , 128 (1), 109 – 11 6 . (28) Maier , J. A.; Martinez, C.; Kasavajhala, K.; W ickstrom, L.; Hauser , K. E.; Simmerling, C. ff14SB: Improvi ng the Accuracy o f Protei n Side C hain a nd Back bone Para meters from f f99SB . J. Chem . Theory C omput. 2015 , 11 (8), 3696 – 3713. (29) Jorge nsen, W . L.; Chandrase khar , J.; Madura , J. D.; Impey , R. W .; Klein, M. L. C omparison of Simple Potential Fu nctions for Simulating Liqu id W ater . The Journal of Chemic al Physic s 1983 , 79 (2), 926 – 935 . (30) Harvey , M. J.; De Fabritiis, G. A n Implementation of the Smooth Particle Mes h Ewald Method on GPU Hardwar e. J. Chem. Theory C omput. 2009 , 5 (9), 2371 – 2377. (31) Zhang, Z.; Liu , X.; Y an, K.; T uckerman, M. E.; L iu, J. Uni fied Ef ficient Ther mostat Sc heme for the Canonical Ens emble with Holonomic or I sokinetic Constraints via Mo lecular Dynamics. J. Phys. Chem. A 2019 , 123 (28), 6056 – 6079 . (32) McGibbon, R. T .; Beauchamp, K. A.; Harrigan, M. P .; Klein, C.; Swai ls, J. M.; He rnández, C. X.; Schwantes , C. R.; W ang, L. - P .; Lane, T . J.; Pande, V . S. MDT raj: A Mode rn Open Lib rary for the Analysis of Molecular Dynamics T rajectories. Biophysical Journal 2015 , 109 (8), 1528 – 1532. (33) Invernizz i, M.; Parrinello, M. Making the Best of a Bad Situation: A Multiscale Approach to Free Ener gy Calc ulation. J . Chem . Theory Comput. 2019 , 15 (4), 2187 – 2194. (34) Biswas, M. ; Lickert, B.; Stock , G. Metadynamics Enhanced Markov Mod eling of Protein Dynamics. J. Phys. Ch em. B 2018 , 122 (21 ), 5508 – 5514 . (35) Botan, V .; Backus, E. H. G.; Pfister , R.; Moretto, A.; Crisma, M.; T o niolo, C.; Nguyen, P . H.; Stock, G.; Hamm, P . En er gy T r ansport in Peptide Helices. Pr ocee dings of t he Nati onal Acade my of Sciences 2007 , 104 (31) , 12749 – 12754. (36) Lindorf f - Larse n, K.; Pia na, S.; Dror , R. O.; S haw , D. E. How Fast - Fol ding Prot eins Fol d. Science 201 1 , 334 (60 55), 517 – 520. (37) Honda, S.; Ak iba, T .; Kato, Y . S.; Sawada, Y .; Sekijim a, M.; Ishi mura, M.; Ooishi, A.; W atanabe, H.; Odaha ra, T .; Harata, K. Cryst al Struct ure of a T en - Amino Acid Prote in. J. A m. Chem. Soc. 2008 , 130 (46), 153 27 – 15331. (38) Miao, Y .; Feixas, F .; Eun, C.; McCammon, J. A. Accelerated Mo lecular Dynamics Simulations of Protein F oldin g. Journal of Comput ational Che mistry 2015 , 36 (20), 1536 – 1549. (39) Shaffer , P .; V alsson, O.; Parrinello, M. Enhanced, T argeted Sampling of High - Dimensi onal Fre e - Energy Landscapes Using V ariationally Enhanced Sampling, with an A pplication to Chignolin. Pr oc. Natl. Acad . Sci. U.S.A. 2016 , 11 3 (5), 1 150 – 1 155. (40) Okumura, H. T emperature and Pr essure Dena turati on of Chi gnolin: F olding and Unfoldin g Simulation by Multibaric‐multithermal Molecular Dynamics Method. Pr oteins 2 012 , 80 (10) , 2397 – 2416. (41) Shaffer , P .; V alsson, O.; Parrinello, M. Enhanced, T argeted Sampling of High - Dimensi onal Fre e - Energy Landscapes Using V ariationally Enhanced Sampling, with an A pplication to Chignolin. Pr ocee dings of the National Acade my of Scie nces 2016 , 11 3 (5) , 1 150 – 1 155. Generative Replica - Exchange: A Flow -based Framework for Accelerating Re plica Exchange Simulations Contents Double - well potential ........................................................................................................................ 2 List of Fi gures ................................................................................................................................... 2 Reference .......................................................................................................................................... 4 Double - well potential The N- dimensional double - well potential in troduced in Michele ’ s etc work 1 , shown in Figure 2. The system with a single parti cle moving w ith a Langevin dynamics with two minima. The translat ions between them are rare in low temperatu re . W e define the firs t two dimensions as coordinates x1 and x2, feel the double- well potential, while all the other are sub ject to a harmoni c potential : U (x) = U (x , x ) + 15 2 x W here U (x) is the modified W olfe -Quapp potential []: U ( x , x ) = 2 ( y + y 2y 4y + 2y y + 0.8y + 0.1y + 9. 28 ) And y [ , ] = y [ , ] ( x , x ) are rotated coordinates, y ( x , x ) = x cos ( ) x sin ( ) , = 0 . y ( x , x ) = x sin ( ) + x cos ( ) , = 0 . List of Figur es Figure S1 . (A) The free energy surface and rep resentative trajectories o btained from cMD at high te mperature ( T = 5) . (B) T he one - dimensio nal mar ginal probabil ity distributions along the x-coordinate. Figure S 2 . F ree ener gy surface at 300 K as a function of Φ a nd Ψ, obtained fro m Cmd ( 2 μ s), GREX ( 100 ns), REX (100 ns), and r es - REX ( 100 ns ). Figure S 3 . Conver gence of the {C5, P II , and α R } basin population as a function of computation al time for GRE X, res - REX, and conventional REX. The dashed box highlights th e early - time regime. Figure S 4 . Tw o - dimensional FES as functions of backbone dihedrals ( Ψ 3 , Ψ 4 ) and ( Φ 5 , Ψ 5 ) in chignolin 3 from (A) 10 μ s cMD, (B) 100 ns cMD, (C) 100 ns REX, and (D) 100 ns GREX. Refer ence (1) Invernizzi, M.; Parrinello, M. Mak ing the Best of a Bad Sit uation: A Multiscale Approach to Free Energy Calculation. J. Chem. Theory Comput. 2019 , 15 (4), 2187–2194. (2) Dinh, L.; Sohl - Dickstein, J.; Bengio, S. Density Estimation Using Real NVP . arXiv February 27, 2017. (3) Shaffer , P .; V alsson, O.; Parrinello, M. Enhanced, T argeted Sampling of High - Dimensional Free - Ener gy Landscapes Using V ariationally Enhanced Sampling, with an Appl ication to Chignolin. Pr oceedings of the National Academy of Sciences 2016 , 11 3 (5), 1 150–1 155.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment