Manifold-Matching Autoencoders
We study a simple unsupervised regularization scheme for autoencoders called Manifold-Matching (MMAE): we align the pairwise distances in the latent space to those of the input data space by minimizing mean squared error. Because alignment occurs on …
Authors: Laurent Cheret, Vincent Létourneau, Isar Nejadgholi
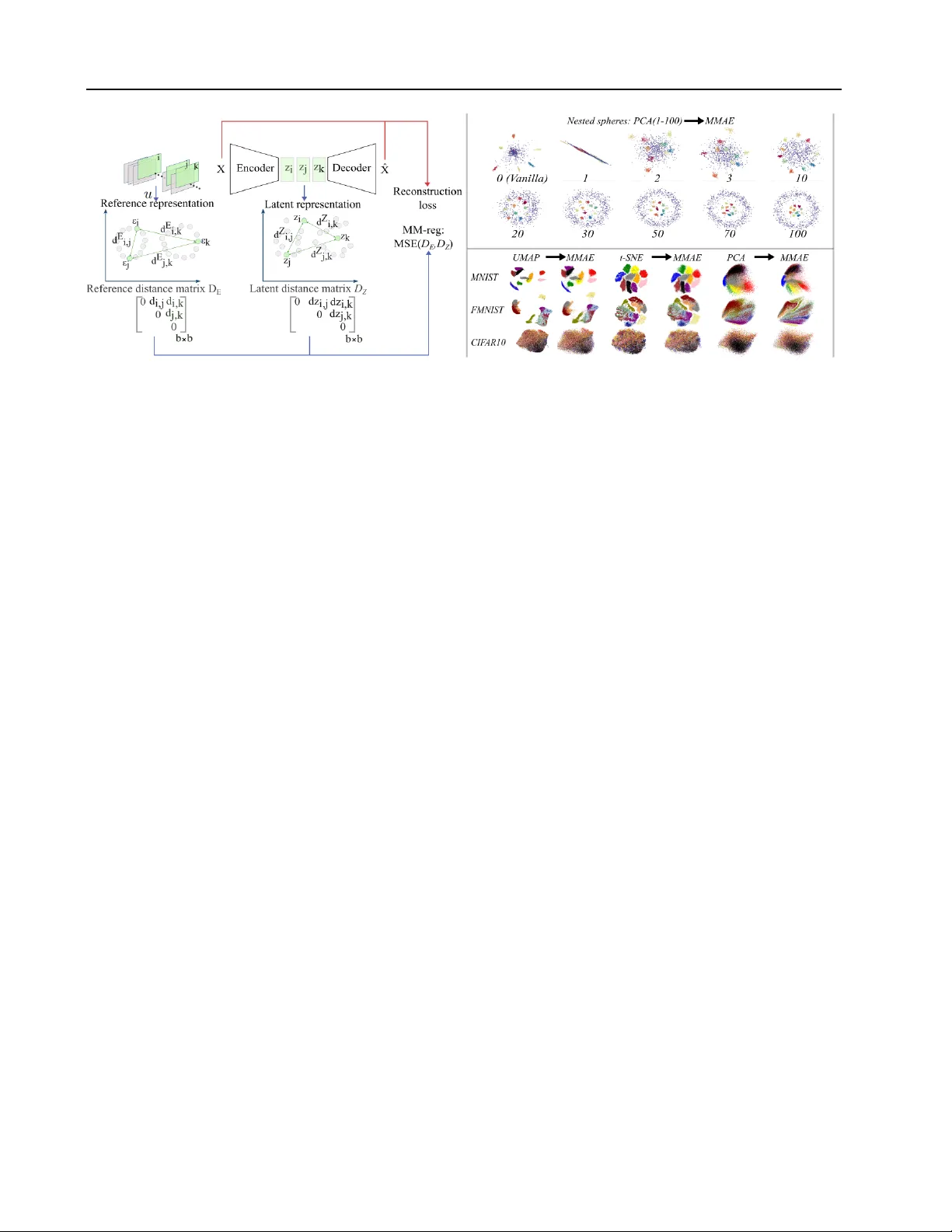
Manif old-Matching A utoencoders Laurent Cheret 1 V incent L ´ etourneau 2 Isar Nejadgholi 3 Chris Drummond 3 Hussein Al Osman 1 Maia Fraser 1 Abstract W e study a simple unsupervised regulariza- tion scheme for autoencoders called Manifold- Matching (MMAE): we align the pairwise dis- tances in the latent space to those of the input data space by minimizing mean squared error . Because alignment occurs on pairwise distances rather than coordinates, it can also be extended to a lower dimensional representation of the data, adding flexibility to the method. W e found that this regularization outperforms similar methods on metrics based on preservation of closest neigh- bors distances and persistence homology based measures. W e also observe MMAE provides a scalable approximation of ‘Multi-Dimensional Scaling” (MDS). 1. Introduction Dimensionality reduction is fundamental to modern data analysis, enabling visualization and interpretation of high- dimensional datasets. Autoencoders ( Hinton & Salakhutdi- nov , 2006 ; Chen & Guo , 2023 ) learn compressed representa- tions by minimizing reconstruction error , but this objecti ve alone does not guarantee preservation of any particular geo- metric or topological structure. When the encoder ignores these structures, similar objects in the input space may be mapped to distinct regions of the latent space, creating dis- continuities that negati vely affect the decoder’ s ability to reconstruct ( Batson et al. , 2021 ). This problem can also affect other downstream tasks. F or example, in anomaly detection or when visualizing dev elopmental trajectories in single-cell data ( Chari & Pachter , 2023 ) or e xploring latent spaces in generative models ( Chadebec & Allassonni ` ere , 2022 ; Xu et al. , 2024 )—additional re gularization becomes necessary . 1 Department of Computer Science, Uni versity of Otta wa, Ot- tawa, Canada 2 MILA, Montr ´ eal, Canada 3 National Research Coun- cil of Canada, Ottaw a, Canada. Correspondence to: Laurent Cheret < lcher021@uottawa.ca > . Pr eprint. Marc h 18, 2026. 1.1. T opology and Geometry in A utoencoders Follo wing the success of statistical methods using topologi- cal data analysis tools like persistence diagrams ( Su et al. ), there has been a recent effort to improv e the preservation of topological features by autoencoders, which we classify as topological or geometrical methods. T opological methods ( Moor et al. , 2020b ; T rofimov et al. , 2023 ) use persistent homology to identify and preserve multi-scale structural features such as connected components, loops, and voids. Geometric methods ( Singh & Nag , 2021 ; Nazari et al. , 2023 ; Lim et al. , 2024 ) focus on preserving local angles and distances. T ake, for example, the nested spheres dataset ( Moor et al. , 2020b ), a simple synthetic yet highly nonlinear case: ten 100-dimensional small spheres are nested inside a larger 100D enclosing sphere. A topologically accurate 2D repre- sentation should preserve this nesting relationship, with the outer sphere cluster surrounding the inner sphere clusters. T o date, only topological autoencoder v ariants consistently recov er this structure in 2D/3D. Other autoencoder variants and nonparametric methods such as UMAP ( McInnes et al. , 2018 ), t-SNE ( van der Maaten & Hinton , 2008 ), and PHA TE ( Moon et al. , 2017 ) fail in this case. Interestingly , we found that the classical method Multidimensional Scaling (MDS) ( T or gerson , 1952 ) also successfully recovers the nesting re- lationship—a result that, to the best of our kno wledge, has not been reported in previous w ork. 1.2. Our Appr oach: Alignment of Pairwise Distances Classical MDS ( T or gerson , 1952 ) utilizes the pairwise dis- tance matrix of the entire dataset to preserve global geom- etry . In contrast, topological autoencoders like T opoAE ( Moor et al. , 2020b ) and R TD-AE ( T rofimov et al. , 2023 ) employ persistent homology signatures on pairwise dis- tance matrices at the mini-batch level. While the former focuses on geometric distances—which may implicitly cap- ture topology—the latter prioritize multi-scale structural connectivity , a focus that has been shown to result in supe- rior preservation of the manifold’ s global geometry . One issue arises with these methods, MDS scales poorly with data size due to memory requirements to compute the n × n pairwise distance matrix, and topological v ariants of au- 1 Manifold-Matching A utoencoders F igur e 1. Left: Overvie w of the current approach. The Manifold-Matching regularization MM-re g is added to the objective function of the standard AE, forming MMAEs. T op Right: 2D latent spaces of the Nested Spheres dataset ( Moor et al. , 2020b ). Standard AE (V anilla) using no MM-reg and 9 MMAE models using dif ferent number of PCA components in their regularization ( 1 → 100 ). Bottom Right: MMAE 2D latent spaces “copying” 2D embeddings from UMAP , t-SNE, and PCA across MNIST , F-MNIST , and CIF AR10 datasets. toencoders scale poorly with the batch size b due to the persistent homology computations batch-wise. A central question emerges: What happens to latent spaces when global geometry pr eservation is imposed in autoencoders? W e address this by introducing a re gularization term called Manifold-Matching (MM-re g), defined as the MSE between the pairwise distance matrix D Z of the latent space and a reference distance matrix D E computed from either the input data X itself or its respective embedding. Crucially , since both D Z and D E are b × b matrices (where b is the batch size), the dimensionality of the reference space is decoupled from the bottleneck dimensionality . This means, for example, that a 2D latent space can be regularized using distances from a 50D or 100D reference representation. Note there is strong theoretical justification in the choice to replace the data by its distance matrix. In short, dis- tance preservation implies topology preservation. This is the content of section 2.2 . Figure 1 illustrates this effect on the nested spheres dataset. W ithout regularization, the standard AE projects the inner spheres outside the cluster representing the outer sphere, consistent with prior literature. Howe ver , with MM-reg, as the number of PCA components in the reference increases, the nesting structure present in the original data emerges, with the inner sphere clusters progressiv ely pulled inside the outer sphere cluster . In the specific case in which the reference embeddings are 2D, a “copying” effect can be seen where the 2D latent space approximates the reference, allowing the autoencoder to extend kno wn representation to new data points. Specifically , our contributions are: (1) W e introduce the Manifold-Matching A utoencoder (MMAE) , an unsuper- vised framew ork for global structure-aware dimensionality reduction. (2) W e study its visualization effects on synthetic datasets where the topology is intuitiv ely understood. (3) W e extend e xperiments to real-world benchmarks, showing competitiv e performance against topological and geomet- rical autoencoder variants. (4) W e provide discussions on global geometry preservation as a proxy for topology preser- vation. 2. Background W e re view the key concepts underlying our approach: per- sistent homology as the gold standard for topological com- parison, and the relation between distance preservation and topology . 2.1. Persistent Homology The importance of understanding data topology has been recognized since the 1960s ( Rosenblatt , 1962 ). This con- cern is tied to the manifold hypothesis: high-dimensional data X = { x i } k i =1 with x i ∈ R n typically lies on or near a lower-dimensional manifold M ⊂ R n . Persistent ho- mology provides a principled way to detect the topological features of this manifold across scales ( Edelsbrunner et al. , 2002 ; Carlsson , 2009 ). 2.2. From Distance Preser vation to T opology Preser vation Multidimensional Scaling (MDS) ( T or gerson , 1952 ) pro- vides a classical approach that finds a low-dimensional con- figuration of points whose pairwise distances best preserve those of the input distance matrix. The key insight is that 2 Manifold-Matching A utoencoders while points x i , x j ∈ R n may hav e many coordinates, their Euclidean distance d ij = ∥ x i − x j ∥ 2 reduces their rela- tionship to a single scalar value. Remarkably , collecting all such pairwise distances into a matrix D contains suf- ficient information to recover the original geometric con- figuration. Classical MDS formalizes this by con verting distance relationships into geometric configurations through eigendecomposition of the associated Gram matrix ( Borg & Groenen , 2005 ; Schoenberg , 1935 ). This distance-centric view connects naturally to topology preservation through the stability theorem. For finite metric spaces with V ietoris-Rips persistence diagrams: Theorem 2.1 (Stability ( Cohen-Steiner et al. , 2007 ; Chazal et al. , 2016 )) . d B ( Dgm p ( X ) , Dgm p ( Y )) ≤ 2 · d GH ( X, Y ) (1) for all homology dimensions p ≥ 0 , wher e d B is the bottle- neck distance and d GH the Gr omov-Hausdorf f distance. Since uniform distance preservation bounds d GH , we ob- tain: Corollary 2.2 (Distance Preservation Implies T opology Preservation) . If an encoder f θ satisfies | d X ( x i , x j ) − d Z ( f θ ( x i ) , f θ ( x j )) | ≤ ϵ for all pairs, then d B ( Dgm p ( X ) , Dgm p ( f θ ( X ))) ≤ 2 ϵ for all p ≥ 0 . This result rev eals our path forward: rather than computing persistent homology during training, we preserve topology by preserving distances. Manifold-Matching Autoencoders operationalize this principle by aligning the latent space to a reference geometry through pairwise distances. In practice, MMAE applies this principle at the minibatch level, lever - aging the theoretical justification that batch-wise topology approximates global topology as batch size increases. A practical consideration is that training operates on mini- batches rather than the full dataset. T opoAE ( Moor et al. , 2020b ) provides theoretical justification for this approach through two ke y results. First, they establish that the proba- bility of batch topology de viating from full set topology is bounded by geometric sampling: P ( d B ( Dgm ( X ) , Dgm ( X ( b ) )) > ϵ ) ≤ P ( d H ( X, X ( b ) ) > ϵ 2 ) (2) where X ( b ) is a mini-batch of size b and d H is the Hausdorf f distance. Second, they sho w that as batch size approaches dataset size, the expected Hausdorf f distance conv erges to zero, meaning batch-level topology increasingly mirrors global topology . This justifies using batch-wise distance preservation as a proxy for global structure preserv ation. 3. Related W ork The challenge of learning topologically correct representa- tions has motiv ated se veral autoencoder variants. W e re view these methods through our core question: ho w can we cap- ture global structure efficiently? 3.1. T opological A utoencoders T opoAE ( Moor et al. , 2020b ) pioneered using persistent homology as a training signal. Gi ven distance matrices D X and D Z in input and latent space, it penalizes discrepancies between topologically significant point pairs: L topo = 1 2 X ( i,j ) ∈P X ( D ij X − D ij Z ) 2 + 1 2 X ( k,l ) ∈P Z ( D kl Z − D kl X ) 2 (3) where P X , P Z denote topologically significant pairs (births/deaths in persistence diagrams). While theoretically generalizable, the implementation focuses on H 0 (connected components) via minimum spanning trees for efficienc y . T wo limitations arise: (1) the loss is discontinuous under point perturbations, as small changes can abruptly alter the spanning tree ( T rofimov et al. , 2023 ); (2) L topo = 0 is nec- essary but not suf ficient for topological equiv alence, failing to capture higher-order features lik e loops. 3.2. RTD-AE: Representation T opology Divergence A utoencoders R TD-AE ( Trofimo v et al. , 2023 ) addresses these limitations with stronger guarantees: nullity of R TD ensures persistence barcodes coincide across all homology de grees. Crucially , the loss is continuous and accounts for feature localiza- tion. The method constructs a joint distance matrix over 2 n points: D joint = 0 n × n D T X D X min( D X , D Z ) (4) with loss L R TD = P ( b,d ) ∈ Dgm ( D joint ) | d − b | p summing per- sistence lifetimes. Notably , their experiments sho w the gap between PCA and topological methods narrows at higher latent dimensions (64–128D). Ho wev er , R TD incurs high computational cost with batch size, often requiring two- stage training (reconstruction, then topology). 3.3. Structure Preserving A utoencoders The Structure-Preserving Autoencoder (SP AE) ( Singh & Nag , 2021 ) learns lo w-dimensional representations where pairwise distances are a linearly scaled version of the input space distances. The method defines a distance ratio r ij = d Z ( z i , z j ) /d X ( x i , x j ) and regularizes by minimizing the variance of log-ratios: L SP AE = L recon + λ · V ar [log r ij ] i
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment