Clinical Priors Guided Lung Disease Detection in 3D CT Scans
Accurate classification of lung diseases from chest CT scans plays an important role in computer-aided diagnosis systems. However, medical imaging datasets often suffer from severe class imbalance, which may significantly degrade the performance of d…
Authors: Kejin Lu, Jianfa Bai, Qingqiu Li
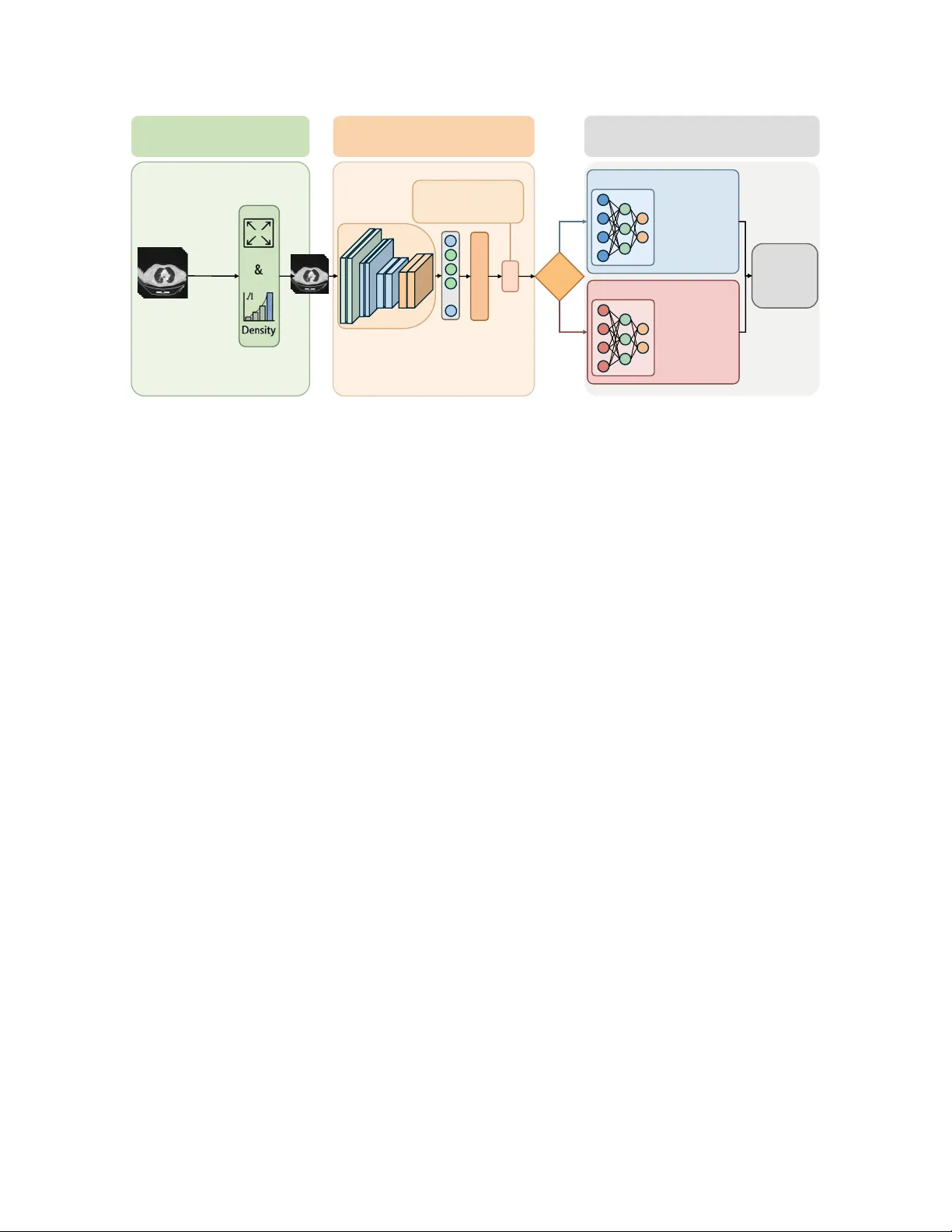
Clinical Priors Guided Lung Disease Detection in 3D CT Scans K ejin Lu 1 , Jianfa Bai 1 , Qingqiu Li 1 , Runtian Y uan 1 , Jilan Xu 2 , Junlin Hou 3 ∗ , Y uejie Zhang 1 ∗ , Rui Feng 1 ∗ 1 College of Computer Science and Artificial Intelligence, Shanghai K e y Laboratory of Intelligent Information Processing, Fudan Uni versity 2 Uni versity of Oxford 3 The Hong K ong Uni v ersity of Science and T echnology Abstract Accurate classification of lung diseases fr om chest CT scans plays an important r ole in computer -aided diagnosis sys- tems. However , medical imaging datasets often suf fer from sever e class imbalance, which may significantly de gr ade the performance of deep learning models, especially for minor- ity disease cate gories. T o addr ess this issue, we pr opose a gender -awar e two-stage lung disease classification frame- work. The proposed appr oach e xplicitly incorporates gen- der information into the disease reco gnition pipeline. In the first stage, a gender classifier is trained to predict the patient’ s gender fr om CT scans. In the second stage , the in- put CT image is r outed to a corresponding g ender-specific disease classifier to perform final disease pr ediction. This design enables the model to better capture gender -r elated imaging char acteristics and alleviate the influence of imbal- anced data distribution. Experimental results demonstrate that the pr oposed method impr oves the r ecognition perfor- mance for minority disease categories, particularly squa- mous cell car cinoma, while maintaining competitive per- formance on other classes. 1. Introduction Lung diseases remain one of the leading causes of mortality worldwide. Accurate and early diagnosis of lung diseases is crucial for improving patient outcomes and guiding clin- ical treatment. Computed tomography (CT) has become an important imaging modality for lung disease screening and diagnosis due to its high spatial resolution and ability to re- veal detailed anatomical structures. In recent years, deep learning methods have shown remarkable performance in medical image analysis and hav e been widely applied to au- tomated lung disease classification tasks [ 1 – 22 ]. Despite these advances, dev eloping robust deep learn- ing models for lung disease classification remains challeng- ing. One major issue is the severe data imbalance that fre- quently exists in real-world medical datasets. Such imbal- ance can lead to biased model training, causing the classifier to fa vor majority classes while underperforming on minor - ity classes. Another challenge arises from demographic het- erogeneity in medical datasets. Clinical studies ha ve sho wn that the pre valence and imaging characteristics of lung dis- eases may vary across different patient groups. This may introduce additional bias during training and limit the gen- eralization ability of con ventional classification models. T o address these issues, we propose a two-stage gender - awar e lung disease classification frame work . The proposed method explicitly incorporates gender information into the disease recognition pipeline. In the first stage, a gender clas- sifier is trained to predict the patient’ s gender from the CT scan. In the second stage, the input CT image is routed to a gender-specific disease classifier based on the predicted gender . Each classifier is trained using samples from the corresponding gender group, allowing the model to better capture gender -related imaging characteristics and alleviate the impact of imbalanced data distribution. Extensiv e e xperiments demonstrate that the proposed gender-a ware two-stage framework improves the recogni- tion performance for minority disease categories, particu- larly squamous cell carcinoma, while maintaining competi- tiv e performance on other classes. The main contrib utions of this work can be summarized as follows: • W e analyze the impact of gender imbalance on lung disease classification and highlight its influence on the recognition of squamous cell carcinoma. • W e propose a two-stage gender-aw are classification framew ork that first predicts patient gender and then per- forms gender-specific disease classification. • Experimental results demonstrate that the proposed ap- proach ef fectiv ely impro ves the robustness and classifica- tion performance under highly imbalanced data distribu- tions. 1 Prepr ocessing Input: 3D Chest CT Sc an( 𝑋 ) Discard Irrelevant Slices Resi zin g & Normalization Processed CT V olume Stage I: Gender Cla ssifica tion Backbone 𝑓(⋅) Feat ure s 𝑧 … FC Laye r + Softmax Gender Prediction ' 𝑦 ! Gender Loss: ℒ !"#$"% = − $ &'( ) 𝑦 & log ( * 𝑦 & ) Gender Rou te Stage II: Gender - Specific Classification Male - Specific Classi fier ( 𝐷 , ) Fema le - Specific Classifi er ( 𝐷 - ) Adenocarcinom a Squamous Ce ll Carcinoma Normal COVID - 19 Final Disease Prediction " 𝑦 . Adenocarcinom a Squamous Ce ll Carcinoma Normal COVID - 19 Figure 1. Ov erview of the proposed gender -aware two-stage frame work. 2. Methodology Chest CT based lung disease classification often suffers from severe data imbalance, especially for squamous cell carcinoma cases. In our dataset, the number of squamous cell carcinoma samples is relativ ely small, and the distribu- tion between male and female patients is highly imbalanced, with male samples significantly outnumbering female sam- ples. Such imbalance may lead to biased learning and de- graded performance for minority groups. T o address this issue, we propose a two-stage gender- aware disease classification framew ork. The model first predicts the patient’ s gender from the CT scan, and then routes the sample to a gender-specific disease classifier . Each classifier is trained using only samples from the cor- responding gender group, allo wing the model to better cap- ture gender-specific imaging patterns and mitigate the im- balance problem. As shown in Figure 1 , The overall pipeline consists of three main components: 1. CT preprocessing and lung region extraction 2. Stage I: Gender classification 3. Stage II: Gender-specific disease classification Similar to pre vious CT -based diagnosis frameworks, we first perform preprocessing on the 3D CT volumes to re- mov e irrelev ant regions. Since the upper neck and lo wer ab- dominal slices contain little diagnostic information for lung diseases, these slices are discarded to focus the model on lung-related structures. The remaining CT slices are then resized to a fixed spatial resolution and normalized before being fed into the network. In the first stage, we train a gender classifier to predict whether the patient is male or female from the CT scan. This design is moti vated by the observation that the dataset exhibits a significant gender imbalance, particularly in the Squamous Cell Carcinoma category . Explicitly modeling gender information allows the subsequent disease classi- fication stage to account for potential demographic differ - ences in imaging characteristics. Giv en an input CT v olume X , a backbone network f ( · ) is used to extract deep features: z = f ( X ) , (1) where z represents the high-le vel semantic representa- tion of the CT scan. The extracted feature vector is then fed into a fully connected layer to produce the gender predic- tion: ˆ y g = Softmax ( W g z + b g ) , (2) where W g and b g denote the learnable parameters of the gender classification head. The gender classifier is opti- mized using the standard cross-entropy loss: L g ender = − 2 X c =1 y c log( ˆ y c ) , (3) which encourages correct discrimination between male and female categories. After predicting the gender, the CT scan is routed to the corresponding disease classifier . This routing strategy enables the model to learn gender-specific disease repre- sentations rather than relying on a single shared classifier , thereby reducing the potential bias introduced by unev en gender distributions in the dataset. T wo disease classifiers are trained independently: • D m : disease classifier trained using male samples • D f : disease classifier trained using female samples 2 Each classifier predicts one of four disease categories: Adenocarcinoma, Squamous Cell Carcinoma, CO VID-19, and Normal. By separating the training process according to gender , the model can better capture subtle differences in disease manifestation across demographic groups. Giv en the predicted gender g , the final disease prediction is defined as: ˆ y d = ( D m ( X ) , g = male D f ( X ) , g = female (4) where the inference process dynamically selects the cor - responding classifier according to the predicted gender . This conditional inference mechanism forms the core of the proposed two-stage frame work. T o further mitigate the impact of class imbalance in dis- ease categories, the disease classifiers are trained using a weighted cross-entropy loss: L disease = − C X c =1 w c y c log( ˆ y c ) , (5) where C denotes the number of disease categories and w c represents the class-specific weight. The weights are introduced to penalize misclassification of minority classes more heavily , thereby improving the overall robustness of the model under imbalanced data distributions. 3. Datasets The dataset used in this study consists of chest CT scans collected for lung disease classification. Each CT volume is annotated with a disease label and patient gender informa- tion. The dataset contains four disease categories. Adeno- carcinoma, CO VID-19, and Normal categories contain rel- ativ ely balanced gender distributions, while the Squamous Cell Carcinoma category exhibits a strong gender imbal- ance. In particular, the number of male Squamous Cell Carcinoma cases is significantly larger than that of female cases. T able 1 summarizes the distrib ution of samples in the dataset. The imbalance in the Squamous Cell Carcinoma category motiv ates the design of the proposed gender -aw are classification framew ork. T able 1. Dataset distribution Disease T rain V al F M F M Adenocarcinoma 125 125 25 25 Squamous Cell Carcinoma 5 79 13 12 CO VID-19 100 100 20 20 Normal 100 100 20 20 Algorithm 1 T wo-Stage Gender-A ware Lung Disease Clas- sification Require: Training dataset D = { ( X i , y i , g i ) } Ensure: Gender classifier G , disease classifiers D m , D f 1: Stage I: T rain Gender Classifier 2: for each epoch do 3: for mini-batch ( X, g ) do 4: z = f ( X ) 5: ˆ g = G ( z ) 6: Compute L g ender 7: Update parameters 8: end for 9: end for 10: Split dataset by gender 11: D m = { ( X , y ) | g = male } 12: D f = { ( X , y ) | g = f emal e } 13: T rain D m on D m 14: T rain D f on D f 15: Inference 16: for test sample X do 17: ˆ g = G ( X ) 18: if ˆ g = mal e then 19: ˆ y = D m ( X ) 20: else 21: ˆ y = D f ( X ) 22: end if 23: end for 4. Experiments 4.1. Implementation Details All experiments are implemented using the PyT orch deep learning framew ork. CT volumes are first resized to a fixed spatial resolution and normalized before being fed into the network. The backbone network is optimized using the Adam optimizer with an initial learning rate of 1 × 10 − 4 . A warmup strategy is employed during the early training phase, followed by cosine learning rate decay . The batch size is set to 8, and the models are trained for 100 epochs. The best-performing model is selected based on v alidation performance. 4.2. Evaluation Metrics T o ev aluate the classification performance, we adopt sev- eral commonly used metrics, including Accuracy , Preci- sion, Recall, F1-score, and A UC. Specifically , class-wise metrics are calculated for each disease category to measure the model’ s performance on individual classes. Further- more, macro-averaged metrics are reported to summarize the ov erall classification performance across all categories. 3 4.3. Experimental Results T able 2 reports the classification performance of the pro- posed method compared with a baseline model that does not incorporate gender information. Overall, the proposed gender-a ware two-stage framework achie ves improv ed clas- sification performance. T able 2. Comparison with baseline Method Accuracy Macro-F1 Macro-A UC Baseline 86.45 0.8223 0.9394 Gender-A ware 86.49 0.8482 0.9361 As shown in the table, the F1 score increases from 0.8223 to 0.8482, demonstrating a noticeable improvement in the balance between precision and recall across cate- gories. Meanwhile, the overall accuracy slightly improves from 86.45% to 86.49%. Although the A UC of the pro- posed method is slightly lower than that of the baseline, the difference is marginal. In contrast, the improv ement in F1 score indicates that the proposed framew ork is more effec- tiv e in handling class imbalance and enhancing recognition of difficult categories, particularly Squamous Cell Carci- noma, which suffers from limited training samples. These results suggest that incorporating gender information helps the model learn more discriminativ e disease representations and improves classification robustness without sacrificing ov erall performance. 5. Conclusion In this paper, we proposed a gender-a ware two-stage lung disease classification framework for chest CT analysis. Considering the severe class imbalance in squamous cell carcinoma and its sk ewed gender distribution in the dataset, the proposed approach e xplicitly incorporates gender infor - mation into the disease recognition pipeline. Specifically , a gender classifier is first trained to predict the patient’ s gen- der from CT scans, and the input image is then routed to a corresponding gender-specific disease classifier for final diagnosis. Experimental results demonstrate that the pro- posed method improv es the recognition performance of mi- nority disease categories, particularly squamous cell carci- noma, while maintaining competitive performance for other disease classes. These results indicate that incorporating de- mographic information such as gender can provide useful prior knowledge for medical image analysis and help alle- viate the impact of imbalanced data distributions. Acknowledgements. This work was supported by National Natural Science Foundation of China(No. 62576107), and the Shanghai Municipal Commission of Economy and Informatization, Corpus Construction for Large Language Models in Pediatric Respiratory Diseases(No.2024-GZL-RGZN-01013), and the Science and T echnology Commission of Shanghai Munici- pality(No.24511104200), and 2025 National Major Science and T echnology Project — Noncommunicable Chronic Diseases-National Science and T echnology Ma- jor Project, Research on the Pathogenesis of Pancreatic Cancer and Nov el Strategies for Precision Medicine( No.2025ZD0552303) References [1] Anastasios Arsenos, Dimitrios Kollias, and Stefanos Kol- lias. A large imaging database and novel deep neural ar- chitecture for covid-19 diagnosis. In 2022 IEEE 14th Im- age , V ideo, and Multidimensional Signal Pr ocessing W ork- shop (IVMSP) , page 1–5. IEEE, 2022. 1 [2] Anastasios Arsenos, Andjoli Davidhi, Dimitrios K ollias, Panos Prassopoulos, and Stefanos Kollias. Data-driv en covid-19 detection through medical imaging. In 2023 IEEE International Confer ence on Acoustics, Speech, and Signal Pr ocessing W orkshops (ICASSPW) , page 1–5. IEEE, 2023. [3] Jiawang Cao, Lulu Jiang, Junlin Hou, Longquan Jiang, Rui- wei Zhao, W eiya Shi, Fei Shan, and Rui Feng. Exploiting deep cross-slice features from ct images for multi-class pneu- monia classification. In 2021 IEEE International Confer ence on Image Pr ocessing (ICIP) , pages 205–209. IEEE, 2021. [4] Demetris Gerogiannis, Anastasios Arsenos, Dimitrios Kol- lias, Dimitris Nikitopoulos, and Stefanos Kollias. Covid- 19 computer-aided diagnosis through ai-assisted ct imaging analysis: Deploying a medical ai system. In 2024 IEEE In- ternational Symposium on Biomedical Ima ging (ISBI) , pages 1–4. IEEE, 2024. [5] Junlin Hou, Jilan Xu, Rui Feng, Y uejie Zhang, Fei Shan, and W eiya Shi. Cmc-cov19d: Contrastive mixup classification for covid-19 diagnosis. In Pr oceedings of the IEEE/CVF In- ternational Confer ence on Computer V ision , pages 454–461, 2021. [6] Junlin Hou, Jilan Xu, Longquan Jiang, Shanshan Du, Rui Feng, Y uejie Zhang, Fei Shan, and Xiangyang Xue. Periphery-aware covid-19 diagnosis with contrastive repre- sentation enhancement. P attern Recognition , 118:108005, 2021. [7] Junlin Hou, Jilan Xu, Nan Zhang, Y i W ang, Y uejie Zhang, Xiaobo Zhang, and Rui Feng. Cmc v2: T owards more accu- rate covid-19 detection with discriminativ e video priors. In Eur opean Confer ence on Computer V ision , pages 485–499. Springer , 2022. [8] Junlin Hou, Jilan Xu, Nan Zhang, Y uejie Zhang, Xiaobo Zhang, and Rui Feng. Boosting covid-19 sev erity detec- tion with infection-aware contrastive mixup classification. In Eur opean Confer ence on Computer V ision , pages 537–551. Springer , 2022. [9] Dimitrios K ollias, Athanasios T agaris, Andreas Stafylopatis, Stefanos K ollias, and Georgios T agaris. Deep neural archi- tectures for prediction in healthcare. Complex & Intelligent Systems , 4(2):119–131, 2018. [10] Dimitrios K ollias, N Bouas, Y Vlaxos, V Brillakis, M Se- feris, Ilianna K ollia, Levon Sukissian, James W ingate, and S 4 K ollias. Deep transparent prediction through latent represen- tation analysis. arXiv pr eprint arXiv:2009.07044 , 2020. [11] Dimitris K ollias, Y Vlaxos, M Seferis, Ilianna K ollia, Lev on Sukissian, James Wingate, and Stefanos D K ollias. T ranspar- ent adaptation in deep medical image diagnosis. In T AILOR , page 251–267, 2020. [12] Dimitrios Kollias, Anastasios Arsenos, Lev on Soukissian, and Stefanos Kollias. Mia-cov19d: Covid-19 detection through 3-d chest ct image analysis. In Proceedings of the IEEE/CVF International Conference on Computer V ision , page 537–544, 2021. [13] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. Ai-mia: Covid-19 detection and sev erity analysis through medical imaging. In Eur opean Conference on Computer V i- sion , page 677–690. Springer , 2022. [14] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. Ai-enabled analysis of 3-d ct scans for diagnosis of covid-19 & its sev erity . In 2023 IEEE International Confer ence on Acoustics, Speec h, and Signal Pr ocessing W orkshops (ICAS- SPW) , page 1–5. IEEE, 2023. [15] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. A deep neural architecture for harmonizing 3-d input data analysis and decision making in medical imaging. Neur o- computing , 542:126244, 2023. [16] Dimitrios Kollias, Anastasios Arsenos, and Stefanos Kol- lias. Domain adaptation explainability & fairness in ai for medical image analysis: Diagnosis of covid-19 based on 3-d chest ct-scans. In Pr oceedings of the IEEE/CVF Conference on Computer V ision and P attern Recognition , pages 4907– 4914, 2024. [17] Dimitrios Kollias, Anastasios Arsenos, James W ingate, and Stefanos Kollias. Sam2clip2sam: V ision language model for segmentation of 3d ct scans for covid-19 detection. arXiv pr eprint arXiv:2407.15728 , 2024. [18] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. Pharos-afe-aimi: Multi-source & fair disease diagnosis. In Pr oceedings of the IEEE/CVF International Conference on Computer V ision , pages 7265–7273, 2025. [19] Qingqiu Li, Runtian Y uan, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Adv ancing covid-19 detec- tion in 3d ct scans. arXiv pr eprint arXiv:2403.11953 , 2024. [20] Qingqiu Li, Runtian Y uan, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Adv ancing lung disease diagnosis in 3d ct scans. arXiv preprint , 2025. [21] Runtian Y uan, Qingqiu Li, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Domain adaptation us- ing pseudo labels for covid-19 detection. arXiv pr eprint arXiv:2403.11498 , 2024. [22] Runtian Y uan, Qingqiu Li, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Multi-source covid-19 detection via variance risk extrapolation. In Proceedings of the IEEE/CVF International Conference on Computer V i- sion , pages 7304–7311, 2025. 1 5
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment