xplainfi: Feature Importance and Statistical Inference for Machine Learning in R
We introduce xplainfi, an R package built on top of the mlr3 ecosystem for global, loss-based feature importance methods for machine learning models. Various feature importance methods exist in R, but significant gaps remain, particularly regarding c…
Authors: Lukas Burk, Fiona Katharina Ewald, Giuseppe Casalicchio
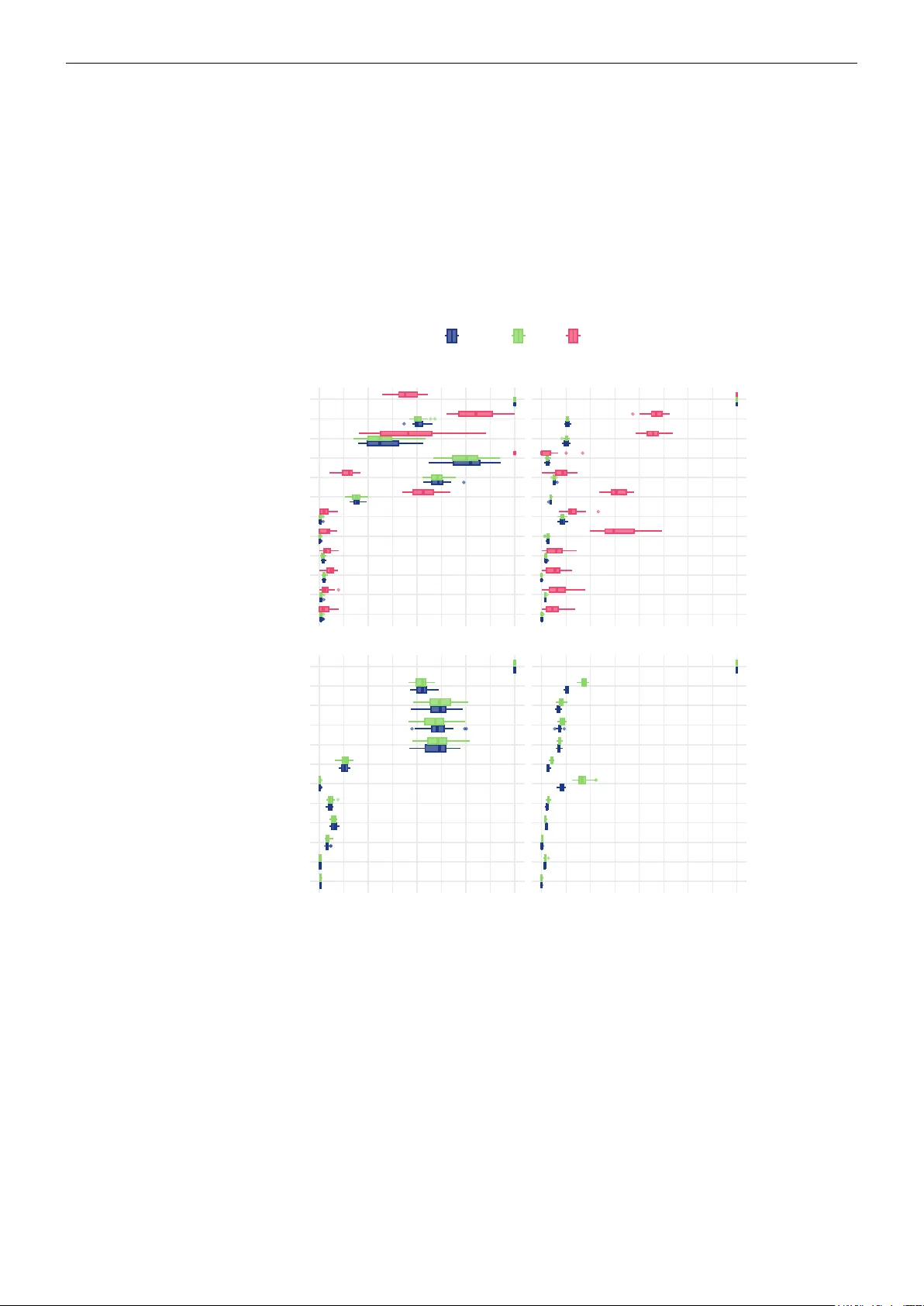
C O N T R I B U T E D R E S E A R C H A RT I C L E 1 xplainfi: Feature Importance and Statistical Inference for Machine Learning in R by Lukas Burk, Fiona Katharina Ewald, Giuseppe Casalicchio, Marvin N. Wright, and Bernd Bischl Abstract W e introduce xplainfi, an R package built on top of the mlr3 ecosystem for global, loss-based feature importance methods for machine learning models. V arious featur e im- portance methods exist in R, but significant gaps remain, particularly regar ding conditional importance methods and associated statistical infer ence procedures. The package imple- ments permutation feature importance, conditional feature importance, relative feature importance, leave-one-covariate-out, and generalizations ther eof, and both mar ginal and conditional Shapley additive global importance methods. It pr ovides a modular conditional sampling architectur e based on Gaussian distributions, adversarial random for ests, condi- tional inference tr ees, and knockof f-based samplers, which enable conditional importance analysis for continuous and mixed data. Statistical inference is available thr ough multiple approaches, including variance-corr ected confidence intervals and the conditional predictive impact framework. W e demonstrate that xplainfi pr oduces importance scor es consistent with existing implementations across multiple simulation settings and learner types, while offering competitive r untime performance. The package is available on CRAN and pr ovides resear chers and practitioners with a compr ehensive toolkit for featur e importance analysis and model interpretation in R. 1 Introduction In machine learning (ML), understanding featur e-target r elationships is increasingly valued alongside predictive accuracy . Complex models can capture nonlinear patterns, but are typically considered “black box” models due to their opaque internal mechanisms, which has motivated the development of interpretable machine learning (IML) methods ( Molnar , 2020 ; Mur doch et al. , 2019 ). Among these, featur e importance (FI) methods quantify the relevance of input featur es for a model’s predictions, pr oviding insight into which features drive model behavior ( Murdoch et al. , 2019 ; Fisher et al. , 2019 ). The applications range from increasing our understanding of a given data-generating pr ocess (DGP) to practical issues such as feature selection ( Guyon and Elisseef f , 2003 ; Guidotti et al. , 2018 ). W e pr esent xplainfi , an R package that provides a unified interface for computing and comparing global, loss-based FI methods, which measure the change in predictive performance when features are r emoved, perturbed, or mar ginalized ( Ewald et al. , 2024 ). Alternative approaches out of scope include local FI methods (i.e., FI values for individual predictions such as Shapley values (see Shapley , 1953 ; Rozemberczki et al. , 2022 )) and variance-based sensitivity measures, such as Sobol indices ( Sobol’ , 2001 ). When we refer to FI methods from here on, we mean global, loss-based FI methods unless otherwise stated. V arious FI methods are implemented across packages in both the R and Python ecosystems, but in the R ecosystem, some methods are completely absent. xplainfi aims to fill this gap by pr oviding a larger collection of FI methods than previously available, along with p-values and confidence intervals for FI uncertainty quantification, which are often needed in applications. The package is built on top of the mlr3 ecosystem ( Lang et al. , 2019 ), which allows devel- opment to focus on the FI methods themselves without re-implementing common building blocks such as abstractions for learning algorithms, resampling, tuning, and pipelines ( Binder et al. , 2021 ). Because many real-world datasets exhibit dependent and correlated features, mar ginal perturbation-based FI can yield misleading attributions by breaking the dependence structur e and redistributing importance acr oss corr elated predictors ( Hooker et al. , 2021 ; Nicodemus et al. , 2010 ; Debeer and Strobl , 2020 ). Conditional importance methods address this by assessing performance changes under interventions that preserve feature dependencies ( W atson and W right , 2021 ), yet r emain underrepr esented in available The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 2 implementations. One challenge with these methods is the need for suitable conditional sampling methods, which is why xplainfi pr ovides multiple methods for continuous and mixed data. Both conditional importance methods and the ability to handle mixed data ar e motivated by common issues arising in practical statistical work and data analysis, and they are also ar eas of active resear ch (see also Blesch et al. , 2024 ; Redelmeier et al. , 2020 ). T o that end, we offer xplainfi as a compr ehensive, extensible tool to support these ef forts. Contributions xplainfi pr ovides a unified framework for FI methods built on the mlr3 ecosystem. Key contributions include: 1) Implementation of many standar d FI methods, including permuta- tion feature importance (PFI) ( Breiman , 2001 ), conditional featur e importance (CFI) ( Str obl et al. , 2008 ; Debeer and Strobl , 2020 ), leave-one-covariate-out (LOCO) ( Lei et al. , 2018 ), relative feature importance (RFI) ( König et al. , 2021 ), and both marginal and conditional SAGE (Shapley Additive Global importancE) ( Covert et al. , 2020 ); 2) Seamless integration with mlr3 ’s learners, tasks, measures, and r esampling strategies; 3) A modular conditional sampling interface supporting Gaussian, adversarial random for ests (ARF) ( W atson et al. , 2023 ), conditional inference tr ees ( Hothorn et al. , 2006 ), and knockoff-based samplers ( Can- dès et al. , 2018 ); 4) The option to tar get model, learner , or DGP importance, by leveraging mlr3 ’s support for r esampling, ensembling, and tuning; 5) Uncertainty quantification via variance-corrected confidence intervals ( Nadeau and Bengio , 2003 ), observation-level LOCO inference ( Lei et al. , 2018 ), and the CPI testing framework ( W atson and W right , 2021 ). The paper is structur ed as follows: First, we give a brief overview of the implemented global, loss-based FI methods and ways to quantify their uncertainty in Section 2 . In Section 3 , we pr esent other R and Python packages that implement FI methods. Section 4 gives an introduction to xplainfi and showcases cor e functionality . Example scenarios, along with comparisons to existing implementations based on FI values and runtime, are provided in Section 5 . W e conclude with a discussion and outlook of xplainfi in Section 6 . Reproducibility and availability: xplainfi is on CRAN and maintained on GitHub . The GitHub repository contains all code r equired to r eproduce the r esults. The code and results are also included in the online supplement. 2 Implemented feature importance methods W e briefly review the FI methods implemented in xplainfi , drawing on the compr ehensive overview in Ewald et al. ( 2024 ), to which we refer for a br oader discussion and additional refer ences. W e also summarize the statistical infer ence procedur es supported by the package. Rather than providing a full theoretical treatment, we focus on the estimands and quantities that are r equir ed for computation and implementation. Notation : W e assume the supervised learning setting with n observations of p features X = ( X 1 , . . . , X p ) and a target Y . A model ˆ f is trained to predict Y from X , and its perfor- mance is evaluated using a loss function L ( Y , ˆ f ( X )) . The feature of interest (FOI) is denoted X j , with X − j repr esenting all r emaining features. Mor e generally , X S denotes a subset of features indexed by S ⊆ { 1, . . . , p } . Feature importance for feature j is denoted FI j , with specific methods indicated by subscripts (e.g., PFI j , LOCO j ). One central distinction between FI methods is whether they target marginal (uncon- ditional) association between a feature and the target, X j ⊥ ⊥ Y , or conditional association given a set of features G , X j ⊥ ⊥ Y | X G ( Strobl et al. , 2008 ; W atson and W right , 2021 ). Many well-established model-independent methods target mar ginal association (e.g., correlation- or information-theoretic measur es) ( Li et al. , 2018 ; Bommert et al. , 2020 ). In the paper and xplainfi , we primarily focus on conditional association of X j with Y , often with the important special case of conditioning on all remaining features X − j . This captures the incr emental predictive value of a featur e when the others are alr eady available, and it entails a substantially harder estimation problem than marginal association, typically The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 3 requiring pr edictive models and pr ocedures that r espect the joint structure of the featur es. Conditioning on arbitrary feature subsets is also supported by xplainfi , e.g., via relative feature importance (RFI) (see Section 2.2 ). Importantly , the loss-based FI methods considered below quantify pr edictive importance (or r eliance) under a particular intervention scheme (e.g., permutation, conditional r esampling, refitting, or marginalization) and are ther efor e not equivalent to generic measures of statistical dependence. For certain population-level (oracle) importance parameters—including refitting-based measures in the framework of W illiamson et al. ( 2023 ) under suitable losses—a zero importance parameter can be equivalent to conditional independence, but FI does not imply a causal effect of X j on Y , and estimated FI values can depend on the chosen intervention distribution as well as the presence of corr elated or r edundant predictors. 2.1 Estimands and statistical inference There are three possible tar gets of an FI analysis, corresponding to distinct estimands: a fitted model , a given learner , or the true pr ediction function regarding the DGP , also called population level inference (see also Chiabur u et al. , 2024 ; Molnar et al. , 2023 ). Model importance analyzes a single, fixed model and quantifies how it uses features for prediction. Learner importance targets the learning algorithm, and the model it produces, when training data is sampled from the DGP of a given sample size. It usually r equires refitting models via resampling to measur e importance acr oss multiple model instantiations, thereby capturing how much models of this class r ely on each featur e. DGP importance aims to explain the true feature-tar get r elationship, by analyzing the population level prediction function (or Bayes- optimal predictor), e.g., f 0 ( x ) = E [ Y | X = x ] under ℓ 2 loss. As we usually never have direct access to this Bayes-optimal predictor in practical applications, this function must then be approximated by a strong learner in an AutoML-like fashion, e.g., by optimizing over multiple model classes and hyperparameters ( Thornton et al. , 2013 ) or by constr ucting heterogenous stacking ensembles ( Erickson et al. , 2020 ; van der Laan et al. , 2007 ). In practice this is very similar to Learner importance , while model building is usually more expensive. Statistical infer ence can be performed at each of the thr ee estimand levels. At the model level, hypothesis tests can be based on observation-wise losses on a single test set. At the learner level, variance-corrected tests can be applied to paired loss differ ences across r esampling iterations. Since FI estimation inher ently involves random components (e.g., from data resampling, permutation, or stochastic learners), repeated evaluation is generally needed to quantify this uncertainty , just as cross-validation is needed for reliable performance estimation. xplainfi provides dedicated infer ence methods at the model and learner levels, which therefor e also includes population level infer ence. When evaluating multiple FOIs simultaneously , any of these infer ence procedur es intro- duces a multiple-testing problem. Depending on the use case, it is necessary to control either the family-wise error rate (FWER) via methods such as the Bonferr oni-Holm corr ection, or to control the false-discovery rate (FDR) with methods such as the Benjamini-Hochber g or Benjamini–Y ekutieli procedur es, where the latter is valid under arbitrary dependence structur es ( Holm , 1979 ; Benjamini and Hochberg , 1995 ; Benjamini and Y ekutieli , 2001 ). In an exploratory setting, it is often suf ficient to contr ol the FDR, which controls the expected pro- portion of false positives among all rejections (e.g., 1 out of 20 features deemed important), whereas contr olling the FWER is more suitable for confirmatory settings in which avoiding false positives takes priority . Regardless of the estimand, loss-based FI methods are usually constr ucted by “r emoving” the information about the FOI(s) and calculating the differ ence in expected loss (with vs. without this information). Following Ewald et al. ( 2024 ), we group the FI methods by the strategy used to remove information: feature perturbations, model refitting, or Shapley values. The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 4 2.2 Methods based on feature perturbations The most well-known method in this category is permutation featur e importance (PFI), origi- nally introduced by Breiman ( 2001 ) for random for ests. PFI targets the effect of breaking the link between X j and the rest of the joint distribution at evaluation time, holding ˆ f fixed. The model is fit once but evaluated twice: once with the original features and once with the FOI X j replaced by a perturbed version ˜ X j . In PFI, ˜ X j is derived by simply shuffling X j in-place, which keeps the marginal distribution of X j intact. Importance is then measured as the differ ence in performance between the original model prediction and the model pr ediction using the uninformative replacement for the FOI. Formally , PFI j = E h L Y , ˆ f ( ˜ X j , X − j ) i − E h L Y , ˆ f ( X ) i . While PFI is computationally cheap, requiring only model pr edictions and simple shuf- fling of feature vectors, it breaks dependencies not only between the FOI and the target, but also between the FOI and all other featur es. This can yield implausible feature combinations and misleading attributions under feature dependence (see Hooker et al. , 2021 ). Conditional permutation feature importance (CFI), intr oduced by Strobl et al. ( 2008 ), addresses this by per - turbing the FOI conditional on a set of other features: X j is replaced by ˜ X j sampled from the conditional distribution of X j | X − j , thereby pr eserving (parts of) the dependence structure among the predictors. König et al. ( 2021 ) generalize PFI and CFI to relative feature importance (RFI) by specifying a conditioning set G for which ˜ X j retains conditional dependencies, i.e., ˜ X j is sampled from the conditional distribution of X j | X G ; this yields CFI for G = − j and PFI for G = ∅ . Since the permutation (or sampling) step intr oduces randomness, multiple r epetitions should be used to stabilize the estimate of the importance score. In xplainfi , this is controlled via the n_repeats parameter , with the final importance scor e averaged across r epetitions. In terms of inference, xplainfi allows applying the corrected t-test appr oach proposed by Nadeau and Bengio ( 2003 ) to paired loss differences acr oss resampling iterations, which can be used to quantify uncertainty of (learner-level) FI estimates based on repeated data splits. Molnar et al. ( 2023 ) evaluate and recommend this appr oach for PFI in combination with subsampling or bootstrapping with 10–15 iterations. Schulz-Kümpel et al. ( 2025 ) recommend for this test a train-test ratio of 0.9 and 25 subsampling iterations. xplainfi lets the user apply the inference method to all FI methods, while warning the user if fewer than 10 bootstrap or subsampling iterations are used. Beyond conditional permutation-based appr oaches, the conditional pr edictive impact (CPI) framework proposed by W atson and W right ( 2021 ) provides a dedicated hypothesis test for conditional featur e importance. xplainfi of fers both the original version based on the knockoff framework ( Candès et al. , 2018 ) and an ARF-based version for mixed data ( Blesch et al. , 2025 ). The conditional sampling required by CFI, RFI, and conditional SAGE (described below) can be performed using dif ferent approaches, each with distinct trade-offs. The Gaussian approach assumes multivariate normality , making it fast and straightforwar d but limited to continuous features. Conditional inference tr ees provide a nonparametric alternative that handles mixed feature types, as pr oposed by Redelmeier et al. ( 2020 ) for conditional distribution estimation (e.g., for conditional Shapley-style estimands) and implemented in shapr . Adversarial random forests offer flexible density estimation and conditional sampling for mixed data at a higher computational cost ( W atson et al. , 2023 ). Finally , knockoff sampling provides a specialized approach that enables valid inference through the CPI framework, but implementations for mixed data are less common ( Candès et al. , 2018 ; Blesch et al. , 2024 ). Gaussian, conditional infer ence tree, and ARF samplers can be applied to all conditional importance methods in xplainfi , while knockoff sampling is only compatible with CFI. The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 5 2.3 Methods based on model refitting A more intuitive approach to remove an FOI’s information is to r efit a model without the FOI (using the same learner) and then measure the performance dif ference between the reduced and the original (full) model. The best-known method following this appr oach is the leave-one-covariate-out (LOCO) ( Lei et al. , 2018 ), which analyzes a single FOI X j . If one is interested in the importance of a set of featur es at once, this is known as leave-one-group- out (LOGO) ( Au et al. , 2022 ). W illiamson et al. ( 2023 ) pr oposed a general framework for refitting-based methods, including a dedicated statistical inference method. In xplainfi , we use the terms LOCO and WVIM ( Williamson’ s variable importance measure ; Ewald et al. ( 2024 ) proposed the acr onym). This approach is conceptually simple: T o evaluate an FOI’s importance, the model in question is fit twice, once with and once without the FOI. Again, the r esulting performance differ ence gives a straightforwar d indication of the predictive value “lost” by leaving out feature X j : LOCO j = E h L Y , ˆ f − j ( X − j ) i − E h L Y , ˆ f ( X ) i , where ˆ f − j denotes the model fitted without feature X j . Model r efitting, of course, increases the computational cost in comparison to FI methods based on perturbations, where only one model needs to be fit as the basis for all subsequent calculations. This can be problematic when using LOCO (or a generalization thereof) on datasets with many features, especially when combined with expensive learners. When using stochastic learners, it is also recommended to perform multiple refits (in xplainfi via n_repeats ) to obtain stable importance estimates, further increasing the r equir ed computation time. The inverse operation, leave-one-covariate-in (LOCI), trains models with only a single feature and compar es performance against a featureless baseline to measure the incr ease in performance. While theoretically consistent, LOCI is rarely useful in practice as it essentially investigates univariate associations between the target and a single feature, for which mor e appropriate methods are available (see above). The group version of LOCI (also included in the WVIM framework) is more useful, where a subset of the features is “left in”, chosen, for example, based on domain knowledge about the data. For statistical inference on LOCO, Lei et al. ( 2018 ) suggest an observation-level nonpara- metric test using the ℓ 1 loss differ ences on a test set, which is implemented in xplainfi in a generalized manner that also enables other losses and tests. The corr ected t-test approach mentioned in the previous section is also available for LOCO, but it has not been explicitly investigated in this context. 2.4 Methods based on Shapley values Shapley values originate in cooperative game theory (see Shapley , 1953 ) and, in ML, are often used for “local” explanations, i.e., for explaining individual model pr edictions for selected data instances. But several methods have also been proposed to apply the underlying principle to global FI, including SFIMP ( Shapley Feature IMPortance ) by Casalicchio et al. ( 2019 ) and SPVIM ( Shapley Population V ariable Importance Measure ) by W illiamson and Feng ( 2020 ), which is related to the more widely adopted SAGE ( Shapley Additive Global importancE ) by Covert et al. ( 2020 ). xplainfi implements SAGE, which distributes the overall model loss across the featur es based on their individual contributions, yielding importance scores that sum to the overall model loss. The SAGE value for feature X j is defined as the weighted average of its marginal contributions to featur e subsets S , also called coalitions, based on a value function v : SAGE j = ∑ S ⊆ P \ { j } | S | ! ( p − | S | − 1 ) ! p ! [ v ( S ∪ { j } ) − v ( S ) ] . The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 6 Here, v ( S ) measures how much better the model performs when the information of the features in S is used for prediction (while featur es outside S are integrated out) compar ed to a featureless baseline. Marginal SAGE (mSAGE) uses the mar ginal distribution, resulting in v m ( S ) = E h L ( Y , ˆ f ∅ ) i − E h L Y , E X − S h ˆ f ( X S , X − S ) i i , and conditional SAGE (cSAGE) uses the conditional distribution, r eplacing E X − S with E X − S | X S in the equation above to yield v c ( S ) . Since exact computation r equires evaluating all possible featur e coalitions (2 p ), imple- mentations must rely on approximations. xplainfi implements the permutation estima- tor described by Covert et al. ( 2020 ), where the feature coalitions are built from one of n_permutations shuffles of the feature vector . The empty coalition is always evaluated, resulting in a total number of evaluated coalitions of 1 + p · n_permutations . The variance of SAGE value estimates along the evaluated permutations is also used to define an “early stopping” criterion to avoid excessive computations. Covert et al. ( 2020 ) use these variances to construct confidence-like intervals, but we do not include them as statistical infer ence methods because their coverage is unclear . Additionally , the marginalization step of the estimation r equires a subset of size n_samples for each data instance, and lar ger values yield more stable SAGE value estimates but incur additional time and memory costs for lar ge datasets. 3 Related work V arious R packages on CRAN implement one or more FI methods. W e compar e them along two dimensions: (1) the scope of FI methods they provide, and (2) whether they support model importance (analyzing a single pr e-fit model) or also learner importance via resampling and refitting (see Section 2 ). vip ( Gr eenwell and Boehmke , 2020 ) requires a pre-fit model and pr ovides model-specific importance extraction, PFI, a variance-based method (“FIRM”), and a Shapley-based mea- sure based on the mean absolute Shapley/SHAP values per featur e, i.e., an aggr egate of per-observation featur e attributions of the model prediction. Unlike loss-based FI methods such as PFI and SAGE, both FIRM and the Shapley-based importance in vip summarize im- portance on the model-output (pr ediction) scale rather than via changes in loss. It supports a wide range of model classes and offers repeated permutations and optional subsampling of the evaluation data, but does not refit models; accor dingly , it targets model importance. Simi- larly , iml (no longer actively developed) operates on pre-fit models via a Predictor wrapper and provides PFI alongside other IML methods such as featur e effects, interaction statistics, and local explanations. DALEX likewise wraps pr e-fit models via an explainer object; its PFI implementation is pr ovided by the companion package ingredients , complemented by local explanations and feature-ef fect visualizations. hstats provides PFI alongside H-statistics for interaction detection, also operating on pre-fit models. flashlight wraps pr e-fit models in an explainer abstraction and provides model-agnostic PFI, interaction-str ength measur es based on Friedman’s H -statistics for interaction detection, as well as Shapley-based importance via mean absolute (approximate) SHAP feature attributions aggregated across observations. It supports side-by-side comparison of multiple fitted models via the multiflashlight function, which can also be used to compare models that were manually refit on dif ferent resampling splits. However , it does not orchestrate r efitting/resampling or aggregate fold- wise importance into a single learner importance estimate. The mlr3 ecosystem includes mlr3filters , which provides PFI with r esampling support within its feature-selection frame- work, but is designed for feature ranking rather than importance analysis with uncertainty quantification. Generally , PFI is also available in many model-specific implementations (e.g., in ranger and randomForestSRC for random forests). T o the best of our knowledge, there is no general-purpose, model-agnostic R implemen- tation of CFI or RFI that mirr ors the kind of model-wrapping interfaces available for PFI. The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 7 cpi implements Conditional Pr edictive Impact (CPI), a dedicated conditional-importance testing framework based on knockoff sampling ( W atson and W right , 2021 ). It is tied to the knockoff constr uction rather than providing a general conditional-sampling interface for CFI/RFI and focuses on hypothesis testing (p-values and confidence intervals) rather than a broad FI computation framework. permimp implements a version of CFI but is restricted to tree-based methods and is ther efor e not model-agnostic. vimp takes a differ ent approach: rather than evaluating a pre-fit model, it refits learners for each evaluated feature subset using the SuperLearner framework, which automatically ensembles prediction models to appr oximate the Bayes-optimal pr edictor ( W illiamson et al. , 2023 ). This targets population-level importance with valid confidence intervals and hypothesis tests. 1 It implements what it calls “conditional VIM” (equivalent to LOCO for individual FOIs), “marginal VIM” (equivalent to LOCI), and SPVIM ( Shapley Population V ariable Importance Measure ; W illiamson and Feng ( 2020 )). SPVIM is a Shapley-based method that, like SAGE, distributes the overall model performance across features accor ding to their marginal contributions. Unlike SAGE, it replaces the mar ginalization or conditional sampling step with model r efitting for each feature coalition, which can be slow , but uses Kernel SHAP ( Covert and Lee , 2021 ) to approximate the Shapley values. vimp also accepts pre-computed pr ediction vectors, which, in principle, allow a form of model importance, but with a notably differ ent API compar ed to the model-wrapping approach of the other packages. It is tightly coupled with the SuperLearner framework and offers less flexibility in resampling strategies. In the Python ecosystem, scikit-learn provides tr ee-based PFI and model-agnostic PFI, both operating on pre-fit estimators. The ELI5 project similarly provides PFI as its only model-agnostic FI method ( eli , 2017 ). sage offers the original (mar ginal) SAGE implementa- tion with no dedicated interface for conditional sampling, but it allows supplying external surrogate models for the same purpose ( Covert et al. , 2020 ; Covert and Lee , 2021 ). The fippy package in Python of fers the closest overlap with xplainfi in terms of perturbation-based methods, providing PFI, CFI, RFI, and both marginal and conditional SAGE using various sampling methods. It also supports simple W ald-type CIs for FI scor es fr om the observation- based losses on test data if the chosen performance measure is decomposable. Overall, fippy is mainly centered ar ound explaining pre-fit estimators on fixed evaluation data, but it also offers a (somewhat underdeveloped) LearnerExplainer which only supports CFI in combination with subsampling. In summary , xplainfi offers the most comprehensive coverage of model-agnostic FI methods in R, including PFI, CFI, RFI, LOCO, WVIM, and both mar ginal and conditional SAGE, with flexible systems covering learners, metrics, resampling, and model/learner importance, including tuning and complex ensembles for the latter . Among the compar ed R packages, only xplainfi and vimp provide statistical inference methods for their importance estimates, including confidence intervals and hypothesis tests. In the Python ecosystem, fippy also provides infer ence, but limited to the model-importance setting. T able 1 gives a comparative overview . 4 An introduction to xplainfi xplainfi ’s architectur e is heavily inspired by the mlr3 framework and its extension packages, which also form the foundation of the underlying computational infrastructur e. This means primarily two things for the user: 1. The package API is based on R6 classes with the corresponding object-oriented design, using mlr3 functions and concepts. 2. xplainfi ’s basic ML capabilities are determined by the mlr3 ecosystem. W e begin with a brief intr oduction to mlr3 and then build from simple to more advanced applications of xplainfi . For a complete overview , we refer to Bischl et al. ( 2024 ), and in particular to Foss and Kotthof f ( 2024 ) for a full introduction. 1 But note that the SuperLearner interface also allows using only a single learner , which can then be analyzed for learner importance with vimp The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 8 T able 1: Overview of FI methods and capabilities in xplainfi and related R packages and selected Python packages. A checkmark indicates an available feature, and a checkmark in parenthesis indicates a feature is not directly offered as such but available in either a limited or indirect fashion. Model importance refers to analyzing a pre-fit model; learner importance refers to refitting models via resampling. Statistical inference denotes the availability of confidence intervals or hypothesis tests for importance estimates. xplainfi vimp vip iml hstats flashlight DALEX fippy sage Methods PFI ✓ ✓ ✓ ✓ ✓ ✓ ✓ CFI ✓ ✓ RFI ✓ ✓ LOCO ✓ ✓ WVIM ✓ ✓ mSAGE ✓ ✓ ✓ cSAGE ✓ ✓ ( ✓ ) SPVIM ✓ Capabilities Model importance ✓ ✓ ✓ ✓ ✓ ✓ ✓ ✓ ✓ Learner importance ✓ ✓ ( ✓ ) Statistical inference ✓ ✓ ✓ 4.1 A brief introduction to mlr3 mlr3 is an R package and an associated ecosystem of extension packages born fr om method- ological and applied ML resear ch. Its cor e components are abstractions for the building blocks of ML pipelines. Most importantly for our purposes, these include the objects shown in this short application example: library(mlr3learners) # loads mlr3 rr <- resample( learner = lrn("classif.ranger", num.trees = 100), task = tsk("penguins"), resampling = rsmp("cv", folds = 5)) rr$score(msr("classif.ce")) #> task_id learner_id resampling_id iteration classif.ce #> 1: penguins classif.ranger cv 1 0.0000 #> 2: penguins classif.ranger cv 2 0.0290 #> 3: penguins classif.ranger cv 3 0.0000 #> 4: penguins classif.ranger cv 4 0.0000 #> 5: penguins classif.ranger cv 5 0.0294 #> Hidden columns: task, learner, resampling, prediction_test rr$aggregate(msr("classif.ce")) #> classif.ce #> 0.0117 Here we applied the random forest classification learner as implemented in ranger with 100 trees to the penguins dataset ( Horst et al. , 2020 ) in a 5-fold cr oss-validation pr ocedure, evaluated each iteration with the classification error (CE), and finally calculated the average CE across iterations. A Task ( tsk() ) encapsulates the data and the learning pr oblem (regression, classifica- tion, etc.). Built-in tasks for common datasets are available, e.g., tsk("penguins") , and custom tasks can be cr eated from any data.frame -like object, e.g., as_task_regr(mtcars, target = "mpg") . A Learner ( lrn() ) repr esents an ML algorithm with its hyperparameters. For example, lrn("regr.ranger", num.trees = 100) creates a random for est learner via The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 9 ranger using 100 trees. Learners can also be extended into full ML pipelines with pre- processing, featur e extraction, or hyperparameter optimization (see Thomas , 2024 , Becker et al. ( 2024 ), Binder et al. ( 2021 )). A Measure ( msr() ) quantifies prediction performance, e.g., msr("regr.mse") for MSE for a regression task. Some measures ar e decomposable into observation-wise losses, which is relevant for certain inference methods described below . A Resampling ( rsmp() ) defines train-test splitting strategies, such as holdout, k-fold CV , bootstrapping, or subsampling, e.g., rsmp("cv", folds = 5) . The general xplainfi API relies on these components and applies them to all implemented feature importance methods. For each feature importance method, the user first defines the method object by specifying a Task as the target for the analysis, a Learner for training and predictions, a Resampling strategy , and a Measure for evaluating predictions. The $compute() method is then called to perform the actual computational steps r equir ed for the individual importance method. Finally , importance scores can be accessed at dif ferent levels of aggr egation: $importance() returns importance values per feature, aggregated across resampling iterations and r epetitions (e.g., permutation repetitions in PFI ); $scores() returns importance values per feature and per r esampling iteration and repetition, allowing for custom aggregation or visualization; and $obs_loss() returns observation-wise loss scores and importance values, if available for the curr ent importance method and Measure . In the following, we showcase xplainfi on an included DGP , and r efer to the package website for additional tutorials and descriptions of this and other illustrative simulation settings. 4.2 Example 1: PFI with xplainfi W e start by calculating PFI on a synthetic task with four normally distributed features x1 through x4 , two of which are correlated ( x1 and x2 , r = 0.8) and two of which are independent ( x3 and x4 ). The DGP is y = 2 x 1 + x 3 + ε , ( ε ∼ N ( 0, 0.04 ) ), generated by the sim_dgp_correlated() simulation utility function included in the package, which pr oduces a Task object. Model importance The simplest use case is analyzing a single pre-trained model. W e train a ranger random forest on a holdout split and compute PFI on the corr esponding test set: library(xplainfi) task <- sim_dgp_correlated(n = 5000, r = 0.8) lrn_ranger <- lrn("regr.ranger") resampling_ho <- rsmp("holdout")$instantiate(task) lrn_ranger$train(task, row_ids = resampling_ho$train_set(1)) pfi_model <- PFI$new( task = task, learner = lrn_ranger, measure = msr("regr.mse"), resampling = resampling_ho, n_repeats = 10) pfi_model$compute() pfi_model$importance() #> feature importance #> 1: x1 6.67e+00 #> 2: x2 1.52e-01 #> 3: x3 1.82e+00 #> 4: x4 2.02e-05 When a pr e-trained learner is passed, xplainfi detects this automatically and skips the training step, using the fitted model dir ectly for prediction. The r esampling must be The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 10 instantiated with exactly one test set (i.e., holdout), as ther e is only one model to evaluate. This yields model importance : the importance scores r eflect the behavior of this specific model on this specific test set. Learner importance T o capture importance at the learner level, we pass an untrained learner together with a resampling strategy . xplainfi then trains a new model in each r esampling iteration, and importance scores ar e aggr egated across iterations: pfi <- PFI$new( task = task, learner = lrn("regr.ranger"), measure = msr("regr.mse"), resampling = rsmp("cv", folds = 3), n_repeats = 10) pfi$compute() pfi$importance() #> feature importance #> 1: x1 6.535166 #> 2: x2 0.163446 #> 3: x3 1.813342 #> 4: x4 -0.000126 The construction of the PFI object and the computation are separated: calling $new() de- fines the setup, while $compute() performs the actual work. T o access individual importance scores for each permutation and r esampling iteration, the $scores() method is available, which stores the corr esponding feature, iteration indices, and the associated measur e value of the original (baseline) model and the model post perturbation: head(pfi$scores(), 3) #> feature iter_rsmp iter_repeat regr.mse_baseline regr.mse_post importance #> 1: x1 1 1 0.0695 6.50 6.43 #> 2: x1 1 2 0.0695 6.46 6.39 #> 3: x1 1 3 0.0695 6.68 6.61 Using resampling not only captur es learner-level variation but also enables variance- corrected infer ence methods such as the Nadeau-Bengio corr ection (see Example 3), which requir e loss dif ferences across resampling iterations and are therefor e not available for model importance. All other components are easily swappable. Here we r erun PFI with XGBoost, R 2 , and subsampling instead: pfi <- PFI$new( task = task, learner = lrn("regr.xgboost", eta = 0.01, nrounds = 2000), measure = msr("regr.rsq"), resampling = rsmp("subsampling", repeats = 10), n_repeats = 15) pfi$compute() pfi$importance() #> feature importance #> 1: x1 1.62e+00 #> 2: x2 1.08e-03 #> 3: x3 3.81e-01 #> 4: x4 6.75e-05 The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 11 xplainfi ’s capabilities are extendable by the mlr3 ecosystem: any learner from mlr3learners or mlr3extralearners ( Fischer et al. , 2025 ) can be used, including “auto- tuned” learners, pipelines via mlr3pipelines or full AutoML systems ( Schneider and Becker , 2024 ; Binder et al. , 2024 ). Furthermore, any metric from mlr3measures is available. The trained models and baseline results ar e r etained via $resample_result for further analysis. 4.3 Example 2: CFI and conditional samplers As discussed in Section 2 , PFI can produce misleading results when featur es are corr elated. CFI addresses this by sampling from the conditional distribution, which requires a condi- tional sampler . xplainfi provides a modular abstraction for this. Several conditional samplers are available, but her e we focus on two: The ConditionalGaussianSampler assumes multi- variate normality and is fast but limited to numeric features. The ConditionalARFSampler uses adversarial random forests (ARF) and handles mixed data, but is computationally mor e expensive ( W atson et al. , 2023 ). Samplers are instantiated on a given task once, after which one can draw one or more observations, which CFI and related methods handle internally . Each conditional sampler allows the specification of an arbitrary conditioning set of features for sampling. W e continue with the corr elated DGP fr om Example 1, wher e x1 and x2 are corr elated ( r = 0.8), but only x1 affects the tar get. Using a Gaussian conditional sampler , we compute CFI and also request quantiles calculated acr oss r esampling iterations: cfi = CFI$new( task = task, learner = lrn("regr.ranger"), measure = msr("regr.mse"), resampling = rsmp("subsampling", repeats = 5), sampler = ConditionalGaussianSampler$new(task), n_repeats = 10) cfi$compute() cfi$importance(ci_method = "quantile", alternative = "two.sided") #> feature importance conf_lower conf_upper #> 1: x1 2.78e+00 2.724271 2.828378 #> 2: x2 -7.67e-04 -0.002407 0.000507 #> 3: x3 1.81e+00 1.764661 1.878462 #> 4: x4 -4.11e-05 -0.000561 0.000534 Compared to PFI, CFI identifies that x2 is not associated with y conditional on the other features. The sampler argument distinguishes CFI fr om PFI: while PFI uses marginal permutation, CFI requir es a conditional sampler . The "quantile" method shown above provides empirical quantiles fr om the distribution of importance scor es across r esampling iterations, which help to gauge the stability of the estimates. Note that via "alternative" , we explicitly requested the default of "two-sided" intervals, by analogy with two- or one- sided hypothesis tests, wher e "alternative = ' greater ' " would have given us only the 95% quantile as lower bound. 4.4 Example 3: Inference For principled infer ence beyond empirical quantiles, xplainfi supports two main approaches, depending on the chosen importance method. Nadeau-Bengio correction for PFI Molnar et al. ( 2023 ) r ecommend variance correction based on Nadeau and Bengio ( 2003 ) when using PFI with subsampling or bootstrapping. The correction accounts for dependence between resampling iterations that share training observations, yielding confidence intervals The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 12 x1 x3 0.3 0.6 0.9 1.2 1.5 Impor tance (PFI) F eature Nadeau & Bengio Quantiles Unadjusted Figure 1: Comparison of uncertainty quantification methods using PFI on the correlated task (omit- ting noise features). Empirical 95% quantiles fall between the very narrow unadjusted confidence intervals and the wider Nadeau-Bengio-corr ected intervals, which show the uncertainty masked by the unadjusted method. with improved (though still imperfect) coverage. The recommended setup uses appr oxi- mately 15 subsampling iterations. Since our earlier PFI example already uses subsampling, we can request corr ected confidence intervals via ci_method = "nadeau_bengio" : head(pfi$importance(ci_method = "nadeau_bengio"), 3) #> feature importance se statistic p.value conf_lower conf_upper #> 1: x1 1.62071 0.028601 56.67 8.36e-13 1.556005 1.68541 #> 2: x2 0.00108 0.000278 3.89 3.67e-03 0.000453 0.00171 #> 3: x3 0.38069 0.009970 38.18 2.88e-11 0.358136 0.40324 The output includes standar d errors and adjusted confidence intervals. Note that this approach assumes normally distributed importance scor es and was primarily evaluated for PFI; its use with other methods is experimental. Figure 1 compares the corr ected confidence intervals with the empirical 95% quantiles and also unadjusted confidence intervals. W e note that the latter ar e also available via ci_method = "raw" for comparison, but they ar e not valid for inference. CPI for conditional importance For CFI, inference is available through the CPI framework when using knockoff-based sampling, which leverages observation-wise losses to perform hypothesis tests for fea- ture importance ( W atson and W right , 2021 ). Using a knockoff sampler with CFI enables ci_method = "cpi" , for example in conjunction with a t-test: cfi_knockoff = CFI$new( task = task, learner = lrn("regr.ranger"), measure = msr("regr.mse"), sampler = KnockoffGaussianSampler$new(task), resampling = rsmp("holdout"), n_repeats = 1) cfi_knockoff$compute() cfi_knockoff$importance( ci_method = "cpi", alternative = "greater", p_adjust = "BH", test = "t") #> feature importance se statistic p.value conf_lower conf_upper #> 1: x1 3.061471 0.101382 30.197 3.88e-160 2.894620 Inf #> 2: x2 0.003612 0.002603 1.388 1.10e-01 -0.000672 Inf #> 3: x3 1.865136 0.061402 30.376 2.37e-161 1.764083 Inf #> 4: x4 0.000148 0.000569 0.261 3.97e-01 -0.000787 Inf The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 13 This CPI test provides p-values and one-sided confidence bounds for each featur e. Here we use the p_adjust argument to use R’s p.adjust() function to correct for the FDR with the Benjamini-Hochberg procedur e. W e note that this only corr ects the p-values, but not the confidence bounds, which would only be adjusted for p_adjust = "bonferroni" . Alternatively to the knockoff approach, ConditionalARFSampler can be used in place of knockoffs, enabling CPI-style infer ence for mixed data types as pr oposed by Blesch et al. ( 2025 ), though at a higher computational cost. Note that CPI was proposed in combination with a fixed model and a test set, whereas xplainfi also allows its use with cross-validation. 4.5 Example 4: LOCO and WVIM LOCO is implemented as a special case of the more general WVIM framework as described in Section 2 . W e apply it on the simulated task from Example 1 using a single refit iteration and show the same Bonferr oni-adjusted p-values and confidence intervals based on the W ilcoxon test proposed by Lei et al. ( 2018 ): loco = LOCO$new( task = task, learner = lrn("regr.ranger"), n_repeats = 1, measure = msr("regr.mse"), resampling = rsmp("holdout")) loco$compute() loco$importance( ci_method = "lei", test = "wilcox", p_adjust = "bonferroni")[, c(1:2, 5:7)] #> feature importance p.value conf_lower conf_upper #> 1: x1 0.75309 7.59e-243 1.05107 1.28582 #> 2: x2 0.00229 1.92e-08 0.00274 0.00732 #> 3: x3 0.49751 4.44e-224 0.66464 0.82465 #> 4: x4 0.00409 1.97e-33 0.00614 0.00997 WVIM supports arbitrary feature groups via the groups argument, which accepts a named list specifying features belonging to each gr oup (i.e., LOGO): groups = list(correlated = c("x1", "x2"), independent = c("x3", "x4")) wvim = WVIM$new( task = task, learner = lrn("regr.ranger"), groups = groups, direction = "leave-out", measure = msr("regr.mse"), resampling = rsmp("holdout"), n_repeats = 2) wvim$compute() wvim$importance() #> feature importance #> 1: correlated 4.49 #> 2: independent 1.05 Here, the "correlated" group contains featur es x1 and x2 , which are r emoved together . WVIM also supports a "leave-in" direction, which trains models with only the specified features and compar es against a featureless baseline (i.e., LOCI or “LOGI”). The groups argument is also available for PFI, CFI, and RFI, where the specified gr oups of features are then always perturbed at once. 4.6 Conditional samplers As described in Section 2 , conditional methods like CFI and cSAGE r equire a mechanism to sample from the conditional distribution P ( X j | X − j ) . xplainfi provides a modular sampler The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 14 architectur e that allows dif ferent sampling strategies to be instantiated and passed to the importance method. All samplers inherit fr om the FeatureSampler base class and shar e a common interface: Instantiate a sampler on a Task , then use the $sample() method to draw new values for specified features conditional on the remaining ones or a subset of featur es specified as conditioning_set . W e demonstrate the ARF sampler on the penguins task, which contains both numeric and categorical features. After instantiation, we sample new values for the body_mass feature for a few observations, conditional on all other features (including the categorical island ): task_penguins = tsk("penguins") sampler_arf = ConditionalARFSampler$new(task_penguins) # Original values for comparison task_penguins$data( rows = c(1, 20, 40), cols = c("island", "bill_length", "body_mass")) #> island bill_length body_mass #> 1: Torgersen 39.1 3750 #> 2: Torgersen 46.0 4200 #> 3: Dream 39.8 4650 # Sample new body_mass values conditional on all other features sampler_arf$sample(feature = "body_mass", row_ids = c(1, 20, 40))[, c(6, 3, 4)] #> island bill_length body_mass #> 1: Torgersen 39.1 4298 #> 2: Torgersen 46.0 3850 #> 3: Dream 39.8 3989 The sampler produces new values for body_mass drawn from the estimated conditional distribution P ( body_mass | X − body_mass ) . These sampled values replace the original feature when computing CFI or cSAGE, allowing importance to be assessed while pr eserving dependencies with other features. The choice of sampler involves trade-of fs between flexibility and computational cost. The ARF sampler handles mixed data types without making distributional assumptions but requir es fitting an adversarial random for est, which increases computational time and memory usage. 5 Benchmark experiments T o evaluate xplainfi , we performed benchmark experiments in two categories: Importance results : Since xplainfi re-implements methods available in other packages, we verify that it produces equivalent importance scores across shared methods and DGPs, establishing that implementations ar e faithful and that r esults ar e comparable acr oss packages. Runtime : W e assess whether xplainfi ’s implementations ar e competitive in terms of computational cost compared to existing single-method packages, across varying task dimensionalities and method parameters. W e compare against packages that implement the same methods in a model-importance setting, as this is the common denominator across implementations (see Section 3 ). For PFI, we compare with iml and vip . For marginal SAGE, we compar e with fippy and sage , the latter implementing kernel SAGE ( Covert and Lee , 2021 ). For CFI and conditional SAGE, we compare with fippy , using a Gaussian conditional sampler as the lowest common denominator of available sampling options. vimp is not included as it only shares the refitting-based methods (LOCO, WVIM) with xplainfi , and dif ferences in supported metrics, learner frameworks, and r esampling strategies make a fair apples-to-apples comparison The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 15 difficult. The MSE was used as the evaluation metric across all methods, except for vip , which only supports the root mean squared error . Since RMSE is a monotone transfor- mation of MSE, feature rankings are preserved but scaled importance magnitudes differ . Both benchmarks are defined in R using batchtools ( Lang et al. , 2017 ) for cluster-based parallelization, and reticulate is used to run Python implementations. Each method is evaluated using one of four learners to account for variability in learner capabilities: a linear model ( stats / scikit-learn ( Pedregosa et al. , 2011 )), a random for est ( ranger / scikit-learn , 500 trees), XGBoost ( Chen and Guestrin , 2016 ) (1000 rounds, early stopping after 50, η = 0.1), and an MLP ( mlr3torch / scikit-learn , 1 hidden layer , 20 neurons, 500 epochs with early stopping). Of these, XGBoost is the only learner with an identical underlying implementation across R and Python. The linear model is expected to yield approximately identical results in either language. Learner configurations wer e chosen for simplicity and robustness rather than predictive performance, i.e., learners were not tuned beyond built-in regularization mechanisms. 5.1 Importance benchmark W e compare importance scor es by scaling values for a given method and DGP to the unit interval to examine relative magnitudes. A rank-based comparison is available in the online supplement. W e compare five FI methods across one to four different implementations. T able 2 lists these methods and packages, alongside the parameters used for each method wher e applicable. T o obtain reliable FI estimates, we used 50 repetitions for PFI and CFI, and similarly a high number of permutations for mSAGE and cSAGE, while enabling the “early stopping” options in both xplainfi ’s and fippy ’s implementations to avoid high computational cost at diminishing returns. Since we prioritize comparability , we used Gaussian conditional sampling for the conditional methods CFI and cSAGE throughout the experiment, as this is the only sampling mechanism implemented by both xplainfi and fippy . Both packages offer additional sampling mechanisms with greater flexibility , but would have intr oduced more variability into the comparison. The tasks selected for this experiment consist of simulation settings with DGPs designed to showcase differences and similarities among feature importance methods; a full listing is provided in the online supplement. Here, we focus on the “correlated” DGP generated by xplainfi::sim_dgp_correlated() as introduced in the pr evious section ( Y = 2 X 1 + X 3 + ε , with cor ( X 1 , X 2 ) ∈ { 0.25, 0.75 } , p = 4), and the “bike sharing” dataset introduced by Fanaee-T and Gama ( 2014 ) ( p = 12), which is frequently used as an example for FI methods (e.g., Covert et al. , 2020 ; Blesch et al. , 2025 ). The bike sharing dataset’s categorical featur es wer e converted to numeric values, as this was requir ed to ensure comparability between xplainfi and fippy via the Gaussian conditional sampler . For simulated data, n = 5000 samples were generated and 2/3 of the observations were used for learner training and 1/3 for feature importance calculation as a test set. The train and test sets were created consistently across the different implementations, i.e., the linear model for xplainfi was trained on the same data as the one for vip or fippy within a replication. Importance scores shown are aggregated from 25 r eplications. This experiment was conducted on a shared Linux server running Ubuntu 24.04 on an AMD EPYC 9554 CPU with 1.41 T iB of RAM. For additional details on the simulated settings, see the online supplement or xplainfi ’s online documentation . Due to the large number of factors in the experiment, we pr esent a subset of results her e and refer to the online supplement for a complete overview . W e focus on the linear and boosting learners since they are the most comparable acr oss R and Python. Figure 2 shows almost identical scaled importance scores acr oss all implementing pack- ages for PFI and CFI. The exception is vip , which produces the same featur e ranking but with differing importance scor es, most likely due to the different evaluation metric. The feature ranking is equivalent between the two learner types. For mSAGE and cSAGE, similarly close agr eement is visible between xplainfi and fippy , while sage ’s kernel SAGE shows scores close to the other implementations with a slightly lar ger variance. The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 16 T able 2: Methods, implementations, and parameters for both benchmarks. Parameter names cor- respond to xplainfi’s API and are set equivalently in other implementations. For the importance benchmark, early stopping was enabled for mSAGE and cSAGE in xplainfi and fippy; for the runtime benchmark, it was disabled to ensure identical workloads. Method Packages Parameters PFI xplainfi, fippy , iml, vip Importance: n_repeats = 50 Runtime: n_repeats = {1, 50} CFI xplainfi, fippy Importance: n_repeats = 50, sampler: Gaussian Runtime: n_repeats = {1, 50}, sampler: Gaussian mSAGE xplainfi, fippy Importance: n_permutations = 100, n_samples = 100, min_permutations = 20 Runtime: n_permutations = {10, 20, 50}, n_samples = {10, 50, 100} mSAGE sage Importance: n_samples = 100 Runtime: n_samples = {10, 50, 100} cSAGE xplainfi, fippy Importance: n_permutations = 100, n_samples = 100, min_permutations = 20, sampler: Gaussian Runtime: n_permutations = {10, 20, 50}, n_samples = {10, 50, 100}, sampler: Gaussian mSAGE, linear mSAGE, boosting cSAGE, linear cSAGE, boosting PFI, linear PFI, boosting CFI, linear CFI, boosting 0 25 50 75 100 0 25 50 75 100 0 25 50 75 100 0 25 50 75 100 x4 x2 x3 x1 x4 x2 x3 x1 Impor tance (scaled, %) Feature xplainfi fippy vip iml sage Figure 2: Importance scores (scaled to per centages) for PFI, CFI, mSAGE, and cSAGE across imple- mentations on the correlated simulation setting with r = 0.75, based on either the linear model or the boosting learner . The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 17 CFI, linear CFI, boosting PFI, linear PFI, boosting 0 25 50 75 100 0 25 50 75 100 month weather holiday weekda y working_day windspeed temperature apparent_temperature season humidity year hour month weather holiday weekda y working_day windspeed temperature apparent_temperature season humidity year hour Impor tance (scaled, %) F eature xplainfi fippy vip iml Figure 3: Importance scores (scaled to per centages) for PFI and CFI across implementations on the bike sharing dataset, based on either the linear model or the boosting learner . Next, we consider the bike sharing task. Figur e 3 shows PFI and CFI using the linear model and boosting learner . For PFI, we see strong agreement between the methods similar to the previous setting. For CFI, xplainfi and fippy produce dif fer ent (scaled) importance scores for the year and working_day features. This is most likely explained by the Gaussian conditional sampler being implemented slightly differently in both packages, combined with its use here on effectively categorical features encoded as integers, which is not the most appropriate choice. A similar pattern is visible in Figure 4 for cSAGE with the same underlying cause but good agreement otherwise. For mSAGE, sage ’s importance scores produce a dif fer ent pattern which does not appear to agree with the other methods. Scores for the r emaining simulation settings and learners ar e available in the online supplement. In all settings, xplainfi ’s results agr ee with those of the reference implementations apart fr om minor deviations that did not affect the overall ranking, as is also shown by the rank-based analysis in the supplement. 5.2 Runtime benchmark Methods wer e run exclusively using linear models to isolate the computational cost of the FI method itself from that of the learner (see T able 2 for parameter ranges). W e used only the peak task from mlbench , which pr ovides a regr ession problem with user-definable sample The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 18 cSAGE, linear cSAGE, boosting mSAGE, linear mSAGE, boosting 0 25 50 75 100 0 25 50 75 100 holiday weekda y windspeed weather month working_day season humidity apparent_temperature temperature year hour holiday weekda y windspeed weather month working_day season humidity apparent_temperature temperature year hour Impor tance (scaled, %) F eature xplainfi fippy sage Figure 4: Importance scores (scaled to percentages) for mSAGE and cSAGE acr oss implementations on the bike sharing dataset, based on either the linear model or the boosting learner . The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 19 mSAGE cSAGE PFI CFI 5 10 20 5 10 20 1 3 10 30 100 300 1 000 3 000 1 3 10 30 100 1 000 10 000 Number of f eatures Runtime (seconds, log10) xplainfi fippy vip iml sage Figure 5: Median runtime in seconds with 25% and 75% quantiles for PFI and CFI with n_repeats = 50 and mSAGE and cSAGE with n_samples = 100 and n_permutations = 100 across implementations on the peak simulation setting with 5, 10, and 20 features and 5000 samples using a linear model across 25 replications. and feature sizes. Experiments were run for 25 r eplications on the Intel Xeon Platinum 8380 compute nodes of the Leibniz Supercomputing Centr e. Figure 5 shows median runtimes with 25% and 75% quantiles for n = 5000 with varying p . For PFI, iml is clearly the slowest implementation after fippy . vip shows only slightly slower times than xplainfi . For mSAGE, sage notably becomes faster with more featur es, likely because the kernel SAGE implementation converges faster in that case. Across all methods, fippy is notably slower than xplainfi . Overall, xplainfi was consistently faster than or comparable to the refer ence implementations under close-to-equal settings. Additional results for all methods and parameter configurations are available in the online supplement. 6 Discussion and conclusion W e introduced xplainfi , a package that provides a compr ehensive suite of global, loss-based FI methods within a unified framework built on the mlr3 ecosystem, by implementing methods previously unavailable or scattered acr oss packages, including conditional feature importance (CFI), relative feature importance (RFI), and both marginal and conditional SAGE. This allows xplainfi to focus primarily on FI methods without reimplementing com- mon ML building blocks such as model interfaces, resampling, and performance measures. However , this also means xplainfi is not compatible with related frameworks such as tidy- models or arbitrary “unwrapped” models and learner implementations. W e accept this compromise because we believe we would not be able to maintain or expand the current set of features otherwise. Our benchmark experiments demonstrate that xplainfi pr oduces importance scor es consistent with reference implementations across multiple DGPs and learner types, while offering competitive r untime performance. Choosing the “best” methods among those available depends on the resear ch question and the type of association of interest (marginal vs. conditional), and in no small part on the computational budget available, which can affect the choice between perturbation-based, The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 20 refit-based, and Shapley-based methods. For detailed guidance on underlying estimands and interpretation, we r efer to Ewald et al. ( 2024 ). The modular feature sampler ar chitecture is unique, enabling xplainfi to perform condi- tional FI analysis on continuous and mixed data alike via flexible samplers. This includes the ability to specify arbitrary conditioning sets for each sampler in the Conditional family , which enables conditional SAGE and RFI. Interest in conditional methods has been growing, and future work may benefit from the building blocks available with xplainfi . However , these methods come with trade-offs: the ARF sampler , while flexible, incurs higher compu- tational cost and memory usage than Gaussian sampling, and SAGE methods can be slow to compute due to the lar ge number of feature coalitions that must be evaluated. For the latter , we aim to implement corresponding improvements in the near futur e. The package could and should be extended to include additional confidence interval methods, or more generally , uncertainty quantification methods for FI. Although this is very desirable from a practical perspective, not enough (established) techniques and coverage studies exist, and we think exposing users to unvalidated techniques is not appr opriate; such dedicated studies are out of scope for the curr ent paper . The current package does not of fer any visualization options, as we have focused more on the computational aspects. Often, concr ete visualizations for r eports and publications have slightly differ ent requirements and are customized accor dingly . Because we provide well-structur ed container types for our r esults, the generated FI values (and other r esults) can be easily plotted using, e.g., ggplot2 . 7 Acknowledgements The authors gratefully acknowledge the computational and data resources pr ovided by the Leibniz Supercomputing Centr e (www .lrz.de). Lukas Burk is supported by the Federal Ministry of Resear ch, T echnology and Space (BMFTR), grant number 01EQ2409E. Marvin N. W right is supported by the German Research Foundation (DFG), grant numbers 437611051 and 459360854, and the Federal Ministry of Research, T echnology and Space (BMFTR), grant number 01EQ2409E. Bernd Bischl is supported by the German Research Foundation (DFG), grant number 460135501. References Eli5: Debug machine learning classifiers and explain their predictions, 2017. URL https: //github.com/eli5- org/eli5 . [p 7 ] Q. Au, J. Herbinger , C. Stachl, B. Bischl, and G. Casalicchio. Grouped featur e importance and combined features ef fect plot. Data Mining and Knowledge Discovery , 36(4):1401–1450, 2022. doi: 10.1007/s10618- 022- 00840- 5. [p 5 ] M. Becker , L. Schneider , and S. Fischer . Hyperparameter optimization. In B. Bischl, R. Son- abend, L. Kotthoff, and M. Lang, editors, Applied Machine Learning Using mlr3 in R . CRC Press, 2024. URL https://mlr3book.mlr- org.com/hyperparameter_optimization.html . [p 9 ] Y . Benjamini and Y . Hochberg. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple T esting. Journal of the Royal Statistical Society: Series B (Methodological) , 57(1):289–300, 1995. ISSN 0035-9246. doi: 10.1111/j.2517- 6161.1995. tb02031.x. [p 3 ] The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 21 Y . Benjamini and D. Y ekutieli. The contr ol of the false discovery rate in multiple testing under dependency . The Annals of Statistics , 29(4):1165–1188, 2001. ISSN 0090-5364, 2168-8966. doi: 10.1214/aos/1013699998. [p 3 ] M. Binder , F . Pfisterer , M. Lang, L. Schneider , L. Kotthoff, and B. Bischl. Mlr3pipelines - Flexible Machine Learning Pipelines in R. Journal of Machine Learning Research , 22(184): 1–7, 2021. ISSN 1533-7928. URL http://jmlr.org/papers/v22/21- 0281.html . [p 1 , 9 ] M. Binder , F . Pfisterer , M. Becker , and M. N. W right. Non-sequential pipelines and tun- ing. In B. Bischl, R. Sonabend, L. Kotthoff, and M. Lang, editors, Applied Machine Learning Using mlr3 in R . CRC Press, 2024. URL https://mlr3book.mlr- org.com/non- sequential_pipelines_and_tuning.html . [p 11 ] B. Bischl, R. Sonabend, L. Kotthoff, and M. Lang, editors. Applied Machine Learning Using mlr3 in R . CRC Pr ess, 2024. ISBN 978-1-032-50754-5. URL https://mlr3book.mlr- org.com . [p 7 ] K. Blesch, D. S. W atson, and M. N. W right. Conditional feature importance for mixed data. AStA Advances in Statistical Analysis , 108(2):259–278, 2024. ISSN 1863-818X. doi: 10.1007/s10182- 023- 00477- 9. [p 2 , 4 ] K. Blesch, N. Koenen, J. Kapar , P . Golchian, L. Burk, M. Loecher , and M. N. W right. Condi- tional Feature Importance with Generative Modeling Using Adversarial Random Forests. Proceedings of the AAAI Confer ence on Artificial Intelligence , 39(15):15596–15604, 2025. ISSN 2374-3468. doi: 10.1609/aaai.v39i15.33712. [p 4 , 13 , 15 ] A. Bommert, X. Sun, B. Bischl, J. Rahnenführer , and M. Lang. Benchmark for filter methods for featur e selection in high-dimensional classification data. Computational Statistics & Data Analysis , 143:106839, 2020. ISSN 0167-9473. doi: 10.1016/j.csda.2019.106839. [p 2 ] L. Breiman. Random Forests. Machine Learning , 45(1):5–32, 2001. ISSN 1573-0565. doi: 10.1023/A:1010933404324. [p 2 , 4 ] E. Candès, Y . Fan, L. Janson, and J. Lv . Panning for Gold: ‘Model-X’ Knockoffs for High Dimensional Controlled V ariable Selection. Journal of the Royal Statistical Society Series B: Statistical Methodology , 80(3):551–577, 2018. ISSN 1369-7412. doi: 10.1111/rssb.12265. [p 2 , 4 ] G. Casalicchio, C. Molnar , and B. Bischl. V isualizing the Featur e Importance for Black Box Models. In M. Berlingerio, F . Bonchi, T . Gärtner , N. Hurley , and G. Ifrim, editors, Machine Learning and Knowledge Discovery in Databases , pages 655–670, Cham, 2019. Springer International Publishing. ISBN 978-3-030-10925-7. doi: 10.1007/978- 3- 030- 10925- 7_40. [p 5 ] T . Chen and C. Guestrin. XGBoost: A Scalable T ree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining , KDD ’16, pages 785–794, New Y ork, NY , USA, 2016. Association for Computing Machinery . ISBN 978-1-4503-4232-2. doi: 10.1145/2939672.2939785. [p 15 ] T . Chiaburu, F . Haußer , and F . Bießmann. Uncertainty in xai: Human perception and modeling approaches. Machine Learning and Knowledge Extraction , 6(2):1170–1192, 2024. doi: 10.3390/make6020055. [p 3 ] I. Covert and S.-I. Lee. Improving KernelSHAP: Practical Shapley V alue Estimation Using Linear Regr ession. In Proceedings of The 24th International Confer ence on Artificial Intelligence and Statistics , pages 3457–3465. PMLR, 2021. URL https://proceedings.mlr.press/ v130/covert21a.html . [p 7 , 14 ] I. Covert, S. M. Lundberg, and S.-I. Lee. Understanding Global Feature Contributions W ith Additive Importance Measures. In Advances in Neural Information Processing Systems , volume 33, pages 17212–17223. Curran Associates, Inc., 2020. URL https://proceedings. The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 22 neurips.cc/paper/2020/hash/c7bf0b7c1a86d5eb3be2c722cf2cf746- Abstract.html . [p 2 , 5 , 6 , 7 , 15 ] D. Debeer and C. Strobl. Conditional permutation importance revisited. BMC Bioinformatics , 21(1):307, 2020. ISSN 1471-2105. doi: 10.1186/s12859- 020- 03622- 2. [p 1 , 2 ] N. Erickson, J. Mueller , A. Shirkov , H. Zhang, P . Larroy , M. Li, and A. Smola. AutoGluon- tabular: Robust and accurate AutoML for structured data. 2020. URL https://arxiv. org/abs/2003.06505 . [p 3 ] F . K. Ewald, L. Bothmann, M. N. W right, B. Bischl, G. Casalicchio, and G. König. A Guide to Feature Importance Methods for Scientific Inference. In L. Longo, S. Lapuschkin, and C. Seifert, editors, Explainable Artificial Intelligence , pages 440–464, Cham, 2024. Springer Nature Switzerland. ISBN 978-3-031-63797-1. doi: 10.1007/978- 3- 031- 63797- 1_22. [p 1 , 2 , 3 , 5 , 20 ] H. Fanaee-T and J. Gama. Event labeling combining ensemble detectors and backgr ound knowledge. Progress in Artificial Intelligence , 2(2):113–127, 2014. ISSN 2192-6360. doi: 10.1007/s13748- 013- 0040- 3. [p 15 ] S. Fischer , J. Zobolas, R. Sonabend, M. Becker , M. Lang, M. Binder , L. Schneider , L. Burk, P . Schratz, B. C. Jaeger , S. A. Lauer , L. A. Kapsner , M. Mücke, Z. W ang, D. Pulatov , K. Ganz, H. Funk, L. Harutyunyan, P . Camilleri, P . Kopper , A. Bender , B. Zhou, N. German, L. Koers, A. Nazarova, and B. Bischl. Mlr3extralearners: Expanding the mlr3 Ecosystem with Community-Driven Learner Integration. Journal of Open Source Software , 10(115):8331, 2025. ISSN 2475-9066. doi: 10.21105/joss.08331. [p 11 ] A. Fisher , C. Rudin, and F . Dominici. All Models are W rong, but Many are Useful: Learning a V ariable’s Importance by Studying an Entire Class of Prediction Models Simultaneously. Journal of machine learning research : JMLR , 20:177, 2019. ISSN 1532-4435. URL https: //www.ncbi.nlm.nih.gov/pmc/articles/PMC8323609/ . [p 1 ] N. Foss and L. Kotthoff. Data and basic modeling. In B. Bischl, R. Sonabend, L. Kotthoff, and M. Lang, editors, Applied Machine Learning Using mlr3 in R . CRC Press, 2024. URL https://mlr3book.mlr- org.com/data_and_basic_modeling.html . [p 7 ] B. M. Greenwell and B. C. Boehmke. V ariable importance plots—an introduction to the vip package. The R Journal , 12(1):343–366, 2020. doi: 10.32614/RJ- 2020- 013. [p 6 ] R. Guidotti, A. Monreale, S. Ruggieri, F . T urini, F . Giannotti, and D. Pedreschi. A survey of methods for explaining black box models. ACM computing surveys (CSUR) , 51(5):1–42, 2018. doi: 10.1145/3236009. [p 1 ] I. Guyon and A. Elisseeff. An Introduction to V ariable and Feature Selection. Journal of Machine Learning Research , 3(Mar):1157–1182, 2003. ISSN ISSN 1533-7928. URL https: //www.jmlr.org/papers/v3/guyon03a.html . [p 1 ] S. Holm. A Simple Sequentially Rejective Multiple T est Pr ocedure. Scandinavian Journal of Statistics , 6(2):65–70, 1979. ISSN 0303-6898. URL https://www.jstor.org/stable/ 4615733 . [p 3 ] G. Hooker , L. Mentch, and S. Zhou. Unrestricted permutation forces extrapolation: V ariable importance requir es at least one more model, or there is no free variable importance. Statistics and Computing , 31(6):82, Oct. 2021. ISSN 1573-1375. doi: 10.1007/s11222- 021- 10057- z. [p 1 , 4 ] A. M. Horst, A. P . Hill, and K. B. Gorman. Palmerpenguins: Palmer Archipelago (Antar ctica) Penguin Data , 2020. [p 8 ] T . Hothorn, K. Hornik, and A. Zeileis. Unbiased recursive partitioning: A conditional inference framework. Journal of Computational and Graphical Statistics , 15(3):651–674, 2006. ISSN 1061-8600, 1537-2715. doi: 10.1198/106186006X133933. [p 2 ] The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 23 G. König, C. Molnar , B. Bischl, and M. Gr osse-W entrup. Relative Feature Importance. pages 9318–9325, 2021. doi: 10.1109/ICPR48806.2021.9413090. [p 2 , 4 ] M. Lang, B. Bischl, and D. Surmann. Batchtools: T ools for R to work on batch systems. Journal of Open Sour ce Software , 2(10):135, Feb. 2017. ISSN 2475-9066. doi: 10.21105/joss.00135. [p 15 ] M. Lang, M. Binder , J. Richter , P . Schratz, F . Pfisterer , S. Coors, Q. Au, G. Casalicchio, L. Kotthoff, and B. Bischl. Mlr3: A modern object-oriented machine learning framework in R. Journal of Open Source Software , 4(44):1903, 2019. ISSN 2475-9066. doi: 10.21105/joss. 01903. [p 1 ] J. Lei, G. Max, R. Alessandro, T . R yan J., and L. W asserman. Distribution-Free Predictive Inference for Regr ession. Journal of the American Statistical Association , 113(523):1094–1111, 2018. ISSN 0162-1459. doi: 10.1080/01621459.2017.1307116. [p 2 , 5 , 13 ] J. Li, K. Cheng, S. W ang, F . Morstatter , R. P . T revino, J. T ang, and H. Liu. Feature Selection: A Data Perspective. ACM Computing Surveys , 50(6):1–45, Nov . 2018. ISSN 0360-0300, 1557-7341. doi: 10.1145/3136625. [p 2 ] C. Molnar . Interpr etable machine learning . Lulu. com, 2020. URL https://christophm.github. io/interpretable- ml- book/ . [p 1 ] C. Molnar , T . Freiesleben, G. König, J. Herbinger , T . Reisinger , G. Casalicchio, M. N. W right, and B. Bischl. Relating the Partial Dependence Plot and Permutation Feature Importance to the Data Generating Process. In L. Longo, editor , Explainable Artificial Intelligence , pages 456–479, Cham, 2023. Springer Natur e Switzerland. ISBN 978-3-031-44064-9. doi: 10.1007/978- 3- 031- 44064- 9_24. [p 3 , 4 , 11 ] W . J. Murdoch, C. Singh, K. Kumbier , R. Abbasi-Asl, and B. Y u. Definitions, methods, and applications in interpretable machine learning. Proceedings of the National Academy of Sciences , 116(44):22071–22080, 2019. doi: 10.1073/pnas.1900654116. [p 1 ] C. Nadeau and Y . Bengio. Inference for the Generalization Err or. Machine Learning , 52(3): 239–281, 2003. ISSN 1573-0565. doi: 10.1023/A:1024068626366. [p 2 , 4 , 11 ] K. K. Nicodemus, J. D. Malley , C. Str obl, and A. Ziegler . The behaviour of random forest permutation-based variable importance measures under predictor correlation. BMC Bioinformatics , 11(1):110, 2010. ISSN 1471-2105. doi: 10.1186/1471- 2105- 11- 110. [p 1 ] F . Pedr egosa, G. V aroquaux, A. Gramfort, V . Michel, B. Thirion, O. Grisel, M. Blondel, P . Pr et- tenhofer , R. W eiss, V . Dubourg, J. V anderplas, A. Passos, D. Cournapeau, M. Brucher , M. Perrot, and E. Duchesnay . Scikit-learn: Machine learning in Python. Journal of Machine Learning Research , 12:2825–2830, 2011. URL https://jmlr.org/papers/v12/ pedregosa11a.html . [p 15 ] A. Redelmeier , M. Jullum, and K. Aas. Explaining Predictive Models with Mixed Features Using Shapley V alues and Conditional Inference T rees. In A. Holzinger , P . Kieseberg, A. M. Tjoa, and E. W eippl, editors, Machine Learning and Knowledge Extraction , pages 117–137, Cham, 2020. Springer International Publishing. ISBN 978-3-030-57321-8. doi: 10.1007/978- 3- 030- 57321- 8_7. [p 2 , 4 ] B. Rozemberczki, L. W atson, P . Bayer , H.-T . Y ang, O. Kiss, S. Nilsson, and R. Sarkar . The Shapley V alue in Machine Learning. In Pr oceedings of the Thirty-First International Joint Conference on Artificial Intelligence , pages 5572–5579, V ienna, Austria, 2022. International Joint Conferences on Artificial Intelligence Organization. ISBN 978-1-956792-00-3. doi: 10.24963/ijcai.2022/778. [p 1 ] L. Schneider and M. Becker . Advanced tuning methods and black box optimization. In B. Bischl, R. Sonabend, L. Kotthoff, and M. Lang, editors, Applied Machine Learning Using mlr3 in R . CRC Pr ess, 2024. URL https://mlr3book.mlr- org.com/advanced_tuning_ methods_and_black_box_optimization.html . [p 11 ] The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 24 H. Schulz-Kümpel, S. Fischer , R. Hornung, A.-L. Boulesteix, T . Nagler , and B. Bischl. Con- structing Confidence Intervals for ’the’ Generalization Error – a Compr ehensive Bench- mark Study, 2025. [p 4 ] L. S. Shapley . A V alue for n-Person Games. In H. W . Kuhn and A. W . T ucker , editors, Contributions to the Theory of Games (AM-28), V olume II , pages 307–318. Princeton University Press, Dec. 1953. ISBN 978-1-4008-8197-0. doi: 10.1515/9781400881970- 018. [p 1 , 5 ] I. Sobol’. Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Mathematics and Computers in Simulation , 55(1–3):271–280, 2001. ISSN 03784754. doi: 10.1016/S0378- 4754(00)00270- 6. [p 1 ] C. Strobl, A.-L. Boulesteix, T . Kneib, T . Augustin, and A. Zeileis. Conditional variable importance for random forests. BMC Bioinformatics , 9(1):307, 2008. ISSN 1471-2105. doi: 10.1186/1471- 2105- 9- 307. [p 2 , 4 ] J. Thomas. Prepr ocessing. In B. Bischl, R. Sonabend, L. Kotthoff, and M. Lang, editors, Applied Machine Learning Using mlr3 in R . CRC Press, 2024. URL https://mlr3book.mlr- org.com/preprocessing.html . [p 9 ] C. Thornton, F . Hutter , H. H. Hoos, and K. Leyton-Br own. Auto-WEKA: Combined selection and hyperparameter optimization of classification algorithms. In Proceedings of the 19th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining , pages 847–855. Association for Computing Machinery , 2013. ISBN 978-1-4503-2174-7. doi: 10.1145/2487575.2487629. [p 3 ] M. J. van der Laan, E. C. Polley , and A. E. Hubbard. Super learner . Statistical Applications in Genetics and Molecular Biology , 6(1), 2007. doi: 10.2202/1544- 6115.1309. [p 3 ] D. S. W atson and M. N. W right. T esting conditional independence in supervised learning algorithms. Machine Learning , 110(8):2107–2129, 2021. ISSN 1573-0565. doi: 10.1007/ s10994- 021- 06030- 6. [p 1 , 2 , 4 , 7 , 12 ] D. S. W atson, K. Blesch, J. Kapar , and M. N. W right. Adversarial Random Forests for Density Estimation and Generative Modeling. In Proceedings of The 26th International Conference on Artificial Intelligence and Statistics , pages 5357–5375. PMLR, 2023. URL https://proceedings.mlr.press/v206/watson23a.html . [p 2 , 4 , 11 ] B. W illiamson and J. Feng. Efficient nonparametric statistical inference on population feature importance using Shapley values. In Proceedings of the 37th International Conference on Machine Learning , pages 10282–10291. PMLR, Nov . 2020. URL https://proceedings.mlr. press/v119/williamson20a.html . [p 5 , 7 ] B. D. W illiamson, P . B. Gilbert, N. R. Simon, and M. Carone. A general framework for inference on algorithm-agnostic variable importance. Journal of the American Statistical Association , 118(543):1645–1658, 2023. ISSN 0162-1459. doi: 10.1080/01621459.2021.2003200. [p 3 , 5 , 7 ] Lukas Burk 1) Leibniz Institute for Prevention Research and Epidemiology - BIPS 2) LMU Munich, Department of Statistics 3) Munich Center for Machine Learning (MCML) 4) University of Bremen Achterstraße 30, 28359 Bremen, Germany https://lukasburk.de ORCiD: 0000-0001-7528-3795 burk@leibniz- bips.de Fiona Katharina Ewald 1) LMU Munich, Department of Statistics 2) Munich Center for Machine Learning (MCML) Department of Statistics The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859 C O N T R I B U T E D R E S E A R C H A RT I C L E 25 Chair of Statistical Learning and Data Science ORCiD: 0009-0002-6372-3401 fiona.ewald@lmu.de Giuseppe Casalicchio 1) LMU Munich, Department of Statistics 2) Munich Center for Machine Learning (MCML) Department of Statistics Chair of Statistical Learning and Data Science ORCiD: 0000-0001-5324-5966 Marvin N. Wright 1) Leibniz Institute for Prevention Resear ch and Epidemiology - BIPS 2) University of Br emen Achterstraße 30, 28359 Bremen, Germany ORCiD: 0000-0002-8542-6291 Bernd Bischl 1) LMU Munich, Department of Statistics 2) Munich Center for Machine Learning (MCML) Department of Statistics Chair of Statistical Learning and Data Science ORCiD: 0000-0001-6002-6980 The R Journal V ol. XX/YY , AAAA 20ZZ ISSN 2073-4859
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment