Multimodal Connectome Fusion via Cross-Attention for Autism Spectrum Disorder Classification Using Graph Learning
Autism spectrum disorder (ASD) is a complex neurodevelopmental condition characterized by atypical functional brain connectivity and subtle structural alterations. rs-fMRI has been widely used to identify disruptions in large-scale brain networks, wh…
Authors: Ansar Rahman, Hassan Shojaee-Mend, Sepideh Hatamikia
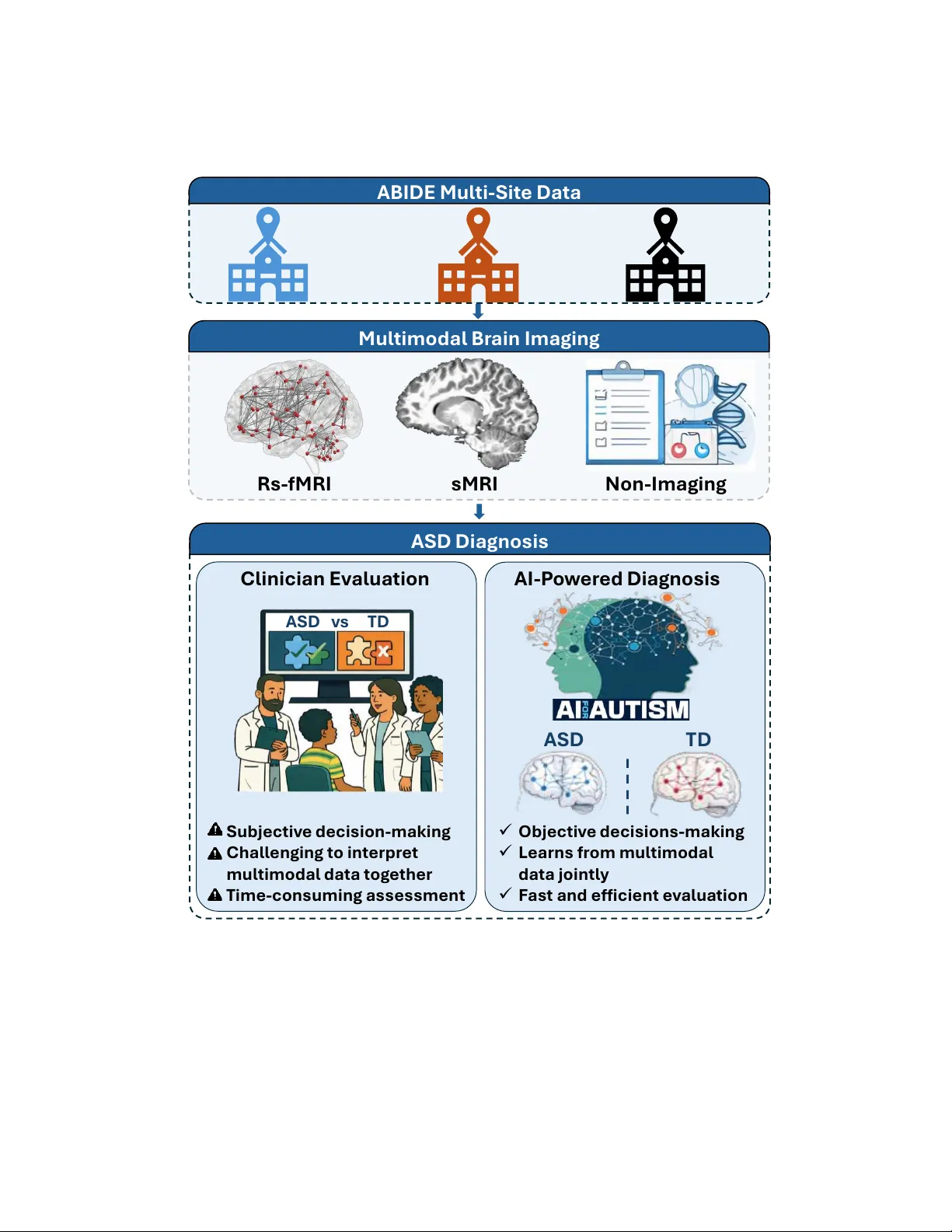
Multimodal Connectome Fusion via Cross-Attention for Autism Spectrum Disorder Classification Using Graph Learning Ansar Rahman 1 a,b , Hassan Shojaee-Mend c , Sepideh Hatamikia a,b,d a Department of Medicine, Danube Private University (DPU), Kr ems, Austria b Department of Medical Physics and Biomedical Engineering, Medical University of V ienna, V ienna, Austria c Department of Medical Informatics, F aculty of Medicine, Gonabad University of Medical Sciences, Gonabad, Iran d Austrian Center for Medical Innovation and T ec hnology (ACMIT), W iener Neustadt, Austria Abstract Autism spectrum disorder (ASD) is a complex neurode velopmental condition characterized by atyp- ical functional brain connectivity and subtle structural alterations. Resting-state fMRI (rs-fMRI) has been widely used to identify disruptions in large-scale brain networks, while structural MRI (sMRI) provides complementary information about morphological or ganization. Despite their com- plementary nature, e ff ecti vely inte grating these heterogeneous imaging modalities within a unified framew ork remains challenging. This study proposes a multimodal graph learning framew ork that preserves the dominant role of functional connecti vity while inte grating structural imaging and phe- notypic information for ASD classification. The proposed multimodal graph-based framew ork is ev aluated on the multi-site ABIDE-I dataset. Each subject is represented as a node within a population graph. Functional and structural features are extracted as modality-specific node attributes, while inter-subject relationships are modeled us- ing a pairwise association encoder (P AE) based on phenotypic information. T wo edge-variational graph conv olutional networks (EV -GCNs) are trained to learn subject-level embeddings. T o enable e ff ectiv e multimodal integration, we introduce a novel asymmetric transformer-based cross-attention mechanism that allows functional embeddings to selectively incorporate complementary structural information while preserving functional dominance. The fused embeddings are then passed to a feed-forward multilayer perceptron (MLP) for ASD classification. Using stratified 10-fold cross-validation, the framework achiev ed an A UC of 87.3% and an ac- curacy of 84.4%. Under leav e-one-site-out cross-validation (LOSO-CV), the model achiev ed an av erage cross-site accuracy of 82.0%, outperforming existing methods by approximately 3% under 10-fold cross-validation and 7% under LOSO-CV . The proposed framew ork e ff ecti vely integrates heterogeneous multimodal data from the multi-site ABIDE-I dataset, improving automated ASD classification across imaging sites. K eywor ds: Autism Spectrum Disorder, Resting-State fMRI, Structural MRI, Multimodal Data Fusion, Cross-Attention, Graph Learning 1. Introduction The comple x neurode velopmental disorder known as ASD is characterized by limited and repet- itiv e patterns of behavior , as well as ongoing impairments in social interaction and communication [1]. ASD manifests clinically in a very diverse way , with significant indi vidual di ff erences in in- tensity of symptoms, developmental pathways, and underlying neurobiological processes. Because of this heterogeneity , objecti ve diagnosis is severely hampered, and there is increasing interest in data-driv en methods that can identify di ff use and subtle neural patterns. Neuroimaging research has increasingly shown that ASD is linked to widespread changes in both brain structure and functional connecti vity rather than isolated abnormalities in particular brain regions [2]. Anatomical organization and morphological characteristics can be thoroughly under - stood through sMRI, whereas rs-fMRI records spontaneous neural activity and functional interac- tions throughout dispersed brain networks. No single imaging modality can adequately capture the intricate neurobiological signatures of ASD, ev en though each one provides insightful information. Thus, one of the main challenges in ASD research continues to be the e ffi cient integration of com- plementary multimodal information. For the classification and characterization of ASD, automated analysis of high-dimensional neu- roimaging data has become more popular due to adv ancements in machine learning, especially deep learning [3, 4]. Single-modality functional connectivity was the main focus of early deep learning techniques. For instance, the e ffi cacy of transfer learning on extensi ve ASD datasets and using au- toencoders to learn latent representations of whole-brain functional connectivity was shown [5, 6]. In order to supplement imaging data and enhance diagnostic performance, researchers [7] started adding non-imaging v ariables within a machine learning frame work, such as age, sex, genetic infor - mation, and cognitiv e measures, as larger cohorts and richer non-imaging phenotypic data became av ailable. Howe ver , the representational capacity of con ventional machine learning techniques is frequently limited by the high dimensionality and heterogeneous nature of such multimodal data 2 [8]. Thus, deep-learning-based multimodal fusion techniques hav e attracted interest for their ability to jointly model heterogeneous data sources. Previous research has demonstrated the e ffi cacy of deep architectures in related neuroimaging tasks, including sex prediction and brain age estimation [9]. Using large, div erse datasets, con volutional neural networks (CNNs) hav e also been in vestigated for the classification of ASD [10]. Regardless of their achievements, these non-graph-based methods are frequently limited to single-modality inputs and usually work with Euclidean data representa- tions, which limits their capacity to model population-le vel structure and inter-subject relationships explicitly . At the same time, although multimodal learning has attracted increasing attention in ASD research, most existing approaches remain limited in how multiple data sources are integrated. In practice, man y studies primarily combine rs-fMRI with non-imaging or demographic v ariables such as age, sex, and acquisition site. While these v ariables pro vide important conte xtual information, they do not fully exploit the complementary neurobiological insights available from sMRI. In par- ticular , sMRI captures morphological and anatomical variations that are not directly observ able in functional connectivity alone. T o date, multimodal inte gration of rs-fMRI and sMRI remains a rarely explored area in ASD classification. T o address the limitations of con ventional multimodal approaches, graph-based learning frame- works have emerged as a promising alternative. Unlike standard deep learning models that process subjects independently , graph models e xplicitly encode relationships between indi viduals at the pop- ulation level. By representing subjects as nodes and modeling inter -subject relationships through edges, graph neural networks enable the joint use of imaging and non-imaging information within a unified representation [11]. Graph con volutional networks (GCNs) further extend conv olutional operations to non-Euclidean domains, making them well-suited for capturing subject associations and brain connectivity patterns [12]. Early work demonstrated that population graphs constructed using non-imaging phenotypic similarities could improve ASD classification performance [11], mo- tiv ating a growing body of graph-based ASD research. Subsequent studies have explored increas- ingly sophisticated graph architectures, including hierarchical GCNs that jointly model subject-le vel associations and network topology [13], as well as adaptive graph constructions such as the Edge- V ariational GCN (EV -GCN) by [14], which learns edge weights through a pairwise association encoder . While these methods have shown promising results, most graph-based ASD studies re- 3 main limited in two important aspects. First, they primarily rely on a single imaging modality , typically rs-fMRI augmented with non-imaging phenotypic information. Second, ev en when deeper GCN architectures are employed [15], increasing network depth alone does not address the chal- lenge of e ff ecti vely integrating complementary structural and functional information across di ff erent modalities. Therefore, despite these advances, multimodal graph-based ASD classification remains relativ ely underexplored, particularly with respect to joint rs-fMRI and sMRI integration. T o the best of our knowledge, only a limited number of studies have explicitly incorporated both structural MRI and functional MRI alongside non-imaging phenotypic data within the ABIDE dataset. Dong et al. [16] conducted a comprehensi ve and standardized comparison of multiple machine learning models using functional connectivity , structural volumetric measures, and non-imaging phenotypic features. Howe ver , this study primarily relies on machine-learning-based feature-lev el integration and en- semble strate gies, and also does not explicitly modeled cross-modal interactions or joint learning population graph structure in an end-to-end multimodal framework. Similarly , Ashrafi et al. [17] proposed a multi-branch GCN architecture that integrates rs-fMRI, sMRI, and non-imaging phe- notypic information from the ABIDE dataset, demonstrating improved classification performance ov er several baselines. While e ff ectiv e, the proposed approach processes each modality largely in- dependently before fusion through concatenation, which limits explicit modeling of cross-modal dependencies and constrains joint optimization of multimodal representations. Advances in multimodal learning have highlighted the importance of modeling interactions be- tween heterogeneous data sources rather than relying on simple feature concatenation. Fixed fusion strategies often fail to capture nuanced, conte xt-dependent relationships between modalities, partic- ularly when functional and structural brain representations encode distinct but interrelated aspects of neurobiology . Recently , attention-based mechanisms hav e emerged as an e ff ectiv e approach for selectiv ely integrating complementary information across modalities by explicitly modeling inter- modality dependencies [18]. W ithin a graph learning context, such mechanisms o ff er the potential to align modality-specific node representations while preserving population-le vel relational struc- ture. T o address the existing challenges, we propose a multimodal graph learning framework for ASD classification that jointly integrates functional and structural brain connectivity deri ved from rs-fMRI and sMRI using an asymmetric cross-attention-based fusion mechanism. By treating modality- 4 specific population graphs as complementary representations of brain organization, the proposed approach explicitly learns ho w information from one modality can guide and refine feature repre- sentations in the other . This design enables adaptive weighting of modality-specific features, facil- itates the extraction of discriminative connectivity patterns, and enhances robustness to site-related variability . T o e valuate the proposed frame work, we conduct e xtensive experiments on the large and heterogeneous multi-site ABIDE-I neuroimaging dataset, comprising data collected from 17 acqui- sition centers (CAL TECH, CMU, KKI, LEUVEN, MAX-MUN, NYU, OHSU, OLIN, PITT , SBL, SDSU, ST ANFORD, TRINITY , UCLA, UM, USM, and Y ALE). The dataset includes multimodal brain imaging and phenotypic information acquired across diverse clinical sites, reflecting the het- erogeneity commonly encountered in real-world ASD studies. In clinical practice, ASD diagnosis primarily relies on beha vioral observ ations and interview-based assessments as illustrated in Fig. 1, which can be subjectiv e and time-consuming [19]. In contrast, recent adv ances in machine learning demonstrate that AI-based analysis of neuroimaging data can provide more objectiv e and e ffi cient diagnostic support by automatically learning discriminativ e patterns from multimodal brain data [20, 21]. Experimental results show that the proposed framework e ff ectively leverages multimodal information and consistently outperforms existing state-of-the-art graph-based and multimodal fu- sion approaches for ASD classification. 5 Mult i mod a l B r a i n I ma ging Rs - fMRI s MRI N on - I ma ging A SD Diag nosi s Cl i nic i an E v al ua t i o n AI - P o w er ed Diagnosi s ✓ O b je ct iv e d e ci s ions - ma kin g ✓ L e arns f r o m mu l t imod al d a t a joint l y ✓ F a s t a nd eff icie nt ev a l u a t ion Sub je ct iv e d e ci s ion - mak in g Ch all e nging t o int e rp r e t mul t imod al d a t a t o g e t he r T ime - co nsuming a ss es sme nt ASD v s T D A SD T D A B I DE Mult i - Sit e Data Figure 1: Overvie w of multimodal ASD diagnosis using heterogeneous ABIDE multi-site data, highlighting the transition from clinician-based assessment to AI-driv en multimodal analysis The main contributions of the proposed w ork are summarized as follows: 6 • W e propose a multimodal population graph learning framew ork for ASD classification that models rs-fMRI and sMRI connectivity using separate modality-specific population graphs with shared phenotypic-driv en edge weights, enabling robust integration of structural and functional information across sites. • W e introduce a nov el transformer-based asymmetric cross-attention–based fusion mechanism that captures directional inter-modality dependencies by using rs-fMRI features as queries and sMRI features as keys and values, enabling adaptiv e integration of complementary structural information while preserving the dominant role of functional connectivity . • W e perform comprehensi ve ev aluations on the multi-site ABIDE-I dataset using both stratified 10-fold cross-v alidation and LOSO-CV to rigorously assess classification performance and cross-site generalization. Compared to existing state-of-the-art multimodal and graph-based approaches, the proposed method demonstrates performance improvements of approximately 3% (in 10-fold cross-validation) and 7% (in LOSO-CV), highlighting enhanced discriminati ve capability , improved rob ustness, and superior generalization across imaging sites. 2. Proposed Methodology In this study , we propose an end-to-end population-graph–based multimodal learning frame- work for ASD classification that integrates rs-fMRI, sMRI, and non-imaging phenotypic informa- tion. Each subject is represented as a node in modality-specific population graphs, where node attributes are deri ved from imaging-based connecti vity features and inter -subject relationships are modeled through adaptive, learnable edge weights computed from non-imaging attributes. T o this end, we used a Pairwise Association Encoder (P AE) that constructs a data-driven population graph by learning non-imaging phenotypic a ffi nities in a latent space optimized for the do wnstream classi- fication task. Modality-specific EV -GCNs [14] are employed to learn complementary subject-lev el represen- tations from rs-fMRI and sMRI population graphs, jointly optimizing node embeddings and graph structure in an end-to-end manner . The resulting modality-specific embeddings are then integrated using an asymmetric cross-attention fusion mechanism, in which rs-fMRI representations act as queries, and sMRI representations serve as keys and values. This design explicitly prioritizes func- tional connectivity while allowing structural information to selecti vely modulate functional repre- 7 sentations. Finally , the fused embeddings are passed to a feed-forward MLP for binary ASD classi- fication. An ov erview of the complete frame work is illustrated in Fig. 2. P henotypic Da t a Age Gen der Si t e St ruct ur al MR I s - Conn ec t i v i t y ma t ri x St ru c t u ral M R I R es t ing - s t a t e f MR I R es ti n g Stat e fM R I f - Conn ec t i v i t y mat rix Data P r ep r oce s s i ng 𝑾 𝒊 . 𝒋 Adapt iv e P o pul a t ion Gr a ph F eat u r e Ve c t o r 𝑿 𝒊 L in ear L ayer L in ear L ayer P AE N e t w o rk C o sine S imi l ar it y C o s ( 𝒉 𝒊 , 𝒉 𝒋 ) 𝑿 𝒋 𝑾 𝒊 . 𝒋 ED Gr a ph Con vol u t ion a l L a ye r s 𝑯 𝒔 𝑴 𝑹 𝑰 𝑯 𝒇 𝑴 𝑹 𝑰 EV - GC N 𝑸 𝒇 𝑭𝒖𝒔 𝒆 𝒅 𝑬 𝒎 𝒃𝒆 𝒅𝒅 𝒊𝒏 𝒈 Cr o s s - At t e nt ion F usi o n 𝑲 𝒔 𝑽 𝒔 EV - GC N & Cr o s s - a t t e nt ion F usi o n R e L U,B at c hN o r m D r o p o u t Si mi lari t y Sc o re Cl a s s if ic a t ion M L P A UC A cc u r a cy Eva lua t ion ASD T D 𝑭𝒖 𝒔 𝒆𝒅 𝑬 𝒎𝒃 𝒆 𝒅 𝒅 𝒊𝒏 𝒈 Se n s it ivi t y Cl ass i fi c at i o n & E v alu at i o n 𝑮 𝒇𝑴 𝑹𝑰 = ( 𝑽 , 𝑬 , 𝑾 ) 𝑽 𝟑 𝑽 𝟐 𝑽 𝑵 𝑽 𝟏 𝑽 𝟓 𝑽 𝟒 P o p ula t ion gr a ph f MR I R F E R i dge Clas s i fi er 𝑽 𝟑 𝑽 𝟐 𝑽 𝑵 𝑽 𝟏 𝑽 𝟓 𝑽 𝟒 𝑮 𝒔 𝑴 𝑹𝑰 = ( 𝑽 , 𝑬 , 𝑾 ) P o pul a t ion gr aph s MR I R F E R i dge Clas s i fi er 𝑮 𝒇 𝑴 𝑹 𝑰 𝑮 𝒔 𝑴 𝑹 𝑰 (a) (b) (c) G C L ayer G C L ayer G C L ayer G C L ayer Figure 2: End-to-End W orkflow Diagram(a) rs-fMRI, sMRI, and phenotypic data are processed to extract functional and structural connectivity features; (b) modality-specific population graphs are constructed and processed using EV -GCN to learn functional and structural node embeddings;(c) the resulting embeddings are integrated through asymmetric cross-attention fusion and passed to a Feed Forward MLP classifier to predict ASD vs. TD 2.1. Dataset and Pr epr ocessing The proposed framework was ev aluated on the ABIDE-I dataset [22], which provides rs-fMRI, sMRI, and non-imaging phenotypic data for ASD and typically developing (TD) subjects. Following the curation protocol in [23] to ensure fair comparison with prior population-graph approaches [23, 24], subjects with incomplete cov erage or inconsistent scanner information were excluded, yielding 871 participants (403 TD, 468 ASD). Experiments were conducted using both stratified 10-fold and LOSO-CV . In each fold, feature selection and population graph construction were performed exclusi vely on the training set to prev ent information leakage. T est subjects’ imaging and non-imaging phenotypic information were not used during training. rs-fMRI: Preprocessed rs-fMRI data were obtained from the ABIDE Preprocessed Connec- tomes Project [25]. Regional mean BOLD time series were extracted via atlas-based parcellation, 8 and Pearson correlation was used to compute symmetric functional connectivity matrices as sho wn in Fig.2 (a). Brain regions were defined using the Harvard–Oxford (HO) atlas (111 R OIs), whose probabilistic definition accommodates inter-subject anatomical v ariability in multi-site data. sMRI: sMRI data were processed using FreeSurfer [26] with the standard r econ-all pipeline for automated segmentation of cortical, subcortical, and white matter regions. Each brain was parcel- lated using the AAL atlas (116 ROIs), and region-wise morphological connectivity matrices were computed to capture inter -regional structural relationships. Only the upper-triangular values of these symmetric matrices were extracted as node features for each subject, as shown in Fig.2 (a). This representation encodes morphological v ariations across cortical thickness, subcortical volumes, and white matter properties while maintaining a compact feature set. Non-Imaging Phenotypic Data: The most commonly used non-imaging features in literature [16, 14, 27] Age, gender , and acquisition site were selected to capture inter-subject similarity in the multi-site setting. These v ariables were numerically encoded and used solely for population graph construction through the Pairwise Association Encoder (P AE) as depicted in Fig.2 (b), which learns adaptiv e edge weights. They were not used as node features or direct inputs to the classifier , thereby av oiding information leakage while incorporating non-imaging phenotypic context into graph learn- ing. 2.2. Multimodal Brain Network Construction In this study , multimodal brain networks are constructed from standardized, preprocessed con- nectivity features deriv ed from the ABIDE-I dataset, as described in Section 2.1. Rather than op- erating on raw neuroimaging v olumes, we focus on population-le vel graph modeling [14], which improv es reproducibility across acquisition sites and enables e ff ective inte gration of multimodal in- formation. Each subject is represented as a node in a population graph, with node attrib utes deriv ed from modality-specific connecti vity features. F or rs-fMRI and sMRI modalities based on connectome data, distinct graphs are constructed as shown in Fig.2 (b), allowing each modality to maintain its inherent representational properties while sharing a connectivity structure driv en by non-imaging phenotypic data. For each modality , vectorized connecti vity features are used as node attributes, while inter-subject similarities are encoded through adaptiv e edge weights deri ved from non-imaging 9 phenotypic information. These population graphs serve as the basis for graph conv olutional repre- sentation learning. 2.2.1. F eatur e Selection and Standar dization Connectivity matrices deriv ed from both resting-state functional and structural imaging are in- herently high-dimensional due to the large number of inter-re gional connections. For each modality , the symmetric connecti vity matrix C ∈ R R × R was transformed into a feature vector by e xtracting the upper triangular elements while excluding the diagonal, where R represents the number of brain regions. This procedure retains all unique pairwise relationships and remo ves redundant information arising from matrix symmetry . T o reduce the risk of ov erfitting, feature selection was performed separately for each modality within ev ery training fold. Recursive Feature Elimination (RFE) [28] with ridge regression was used to rank features based on their contrib ution to classification performance. At each iteration, features with the smallest weight magnitudes were remov ed, and a fixed number of features ( D = 2400) was retained for each modality . The selected features were subsequently standardized using z-score normalization and used as node attrib utes in the corresponding modality-specific population graphs. 2.2.2. Adaptive P opulation Graph Construction Let G = ( V , E , W ) denote a population graph, where V represents a set of N nodes corre- sponding to subjects, E is the set of edges, and W contains edge weights. Each node v i ∈ V is associated with an imaging-deriv ed feature vector z i ∈ R D , which captures discriminati ve diagnostic information from imaging modalities such as rs-fMRI and sMRI. T o model inter-subject relationships, we define graph connectivity using non-imaging attributes (sex, age, site), which pro vide complementary conte xtual information. Instead of relying on fixed or statistically defined similarities, we used a learnable function to model edge weights. Specifically , the edge weight w i j ∈ W between subjects i and j is computed as w i j = f ( x i , x j ; Ω ) , (1) where x i and x j denote non-imaging features and Ω represents trainable parameters of the P airwise Association Encoder (P AE) . 10 2.2.3. P airwise Association Encoder (P AE) The P AE first normalizes each modality of the non-imaging features to zero mean and unit vari- ance to reduce scale discrepancies across modalities [11]. The normalized features are then projected into a shared latent space via a multi-layer perceptron (MLP). In this w ork, the latent dimensionality is set to D h = 128. The association strength between subjects is computed using cosine similarity in the latent space: w i j = h ⊤ i h j 2 ∥ h i ∥∥ h j ∥ + 0 . 5 , (2) which rescales similarity scores to ensure numerical stability . Defining edge weights in the latent space yields more robust graph connecti vity than directly operating on raw non-imaging features. 2.2.4. Edge-V ariational Graph Con volution on Adaptive P opulation Graphs Giv en the adaptively constructed population graph G = ( V , E , W ), EV -GCN performs edge- weighted graph con volution to propagate information across subjects and learn population-aw are node representations. Each node corresponds to a subject and is initialized with modality-specific connectivity features, while edges encode adaptiv e inter-subject relationships learned from non- imaging phenotypic information via the Pairwise Association Encoder (P AE). Graph con volution is performed using spectral filtering based on Chebyshev polynomial approx- imation, which enables e ffi cient aggregation of information from multi-hop neighborhoods without explicitly computing eigen-decompositions. In this formulation, learned edge weights directly mod- ulate the message-passing process, controlling the influence of neighboring subjects during feature aggregation. Since these edge weights are generated by the di ff erentiable P AE, gradients from the classifi- cation objecti ve propagate jointly through both the graph conv olution layers and the edge-weight learning module. This joint optimization enables simultaneous learning of node embeddings and population graph structure, allo wing inter-subject relationships to adapt dynamically to the down- stream ASD classification task. 2.2.5. EV -GCN Encoder For each imaging modality , a dedicated EV -GCN encoder [14] learns population-aware subject representations from graph-structured connectivity data. Each encoder consists of multiple stacked 11 edge-weighted graph conv olution layers with shared hidden dimensionality . Nonlinear activ ation functions and feature-le vel dropout are applied after each layer to impro ve generalization and reduce ov erfitting. EV -GCN block diagram is depicted in Fig.2 (c). T o preserve information across multiple neighborhood scales and pre vent over -smoothing, node embeddings from all graph con volution layers are concatenated using a multi-scale aggregation strat- egy . This allows the model to integrate both local connectivity patterns and global population-lev el structures, capturing diagnostically relev ant di ff erences across subjects. The resulting embeddings provide a rich, structured representation for each subject and serve as the input for the subsequent cross-attention-based multimodal fusion module. 2.3. T ransformer-based Asymmetric Cr oss-Attention for Multimodal Fusion Building upon the modality-specific embeddings learned from the EV -GCN encoders, we pro- posed a nov el transformer-based asymmetric cross-attention fusion mechanism to integrate func- tional and structural brain connectivity . This mechanism enables rs-fMRI embeddings to selectiv ely attend to complementary information from the sMRI, capturing cross-modal interactions more ef- fectiv ely than con ventional feature concatenation or averaging, and forming a ke y component of our multimodal framew ork as shown in Fig.3. Let H f ∈ R N × D and H s ∈ R N × D denote the node embeddings learned from the rs-fMRI and sMRI population graphs, respecti vely , where N is the number of subjects and D is the embedding dimensionality produced by the EV -GCN encoders. 12 𝑸 𝒇 Mu l t i - He ad Cr o s s - At t e nt ion 𝑯 𝒇 𝑴 𝑹𝑰 𝑵 × 𝑫 A s ymmet ri c Cr os s A t t ent i on f MR I - E mb e d d in g 𝑯 𝒔 𝑴 𝑹𝑰 𝑵 × 𝑫 s MR I - E mb e d d in g 𝑲 𝒔 𝑽 𝒔 Linear P ro j ec t i o n 𝒁 ( 𝑭𝒖 𝒔𝒆𝒅 𝑬 𝒎 𝒃 𝒆𝒅 𝒅 𝒊𝒏𝒈 𝒔 ) 𝒁 + 𝑯 𝒇 𝒑𝒓𝒐 𝒋 𝑹𝒆 𝒔𝒊𝒅 𝒖 𝒂𝒍 𝑨𝒅 𝒅 𝒊 𝒕𝒊𝒐𝒏 𝑭𝒆𝒆𝒅 𝑭𝒐 𝒓 𝒘𝒂𝒓 𝒅 𝑵𝒆𝒕𝒘 𝒐𝒓 𝒌 𝑳 𝒊𝒏𝒆𝒂𝒓 𝑺 𝒐𝒇𝒕𝒎 𝒂𝒙 Figure 3: T ransformer-based asymmetric cross-attention fusion integrating modality-specific EV -GCN embeddings from functional and structural brain connectivity . 13 2.3.1. Asymmetric Cr oss-Attention F ormulation T o prioritize functional connectivity while incorporating structural context, in this study , we propose the rs-fMRI embeddings to be treated as queries, while the sMRI embeddings serve as keys and values. The functional embeddings are first projected into a shared latent space, while the structural embeddings are directly used as keys and values without an additional projection to retain their original relational structure, enabling them to act as stable contextual references during attention. Formally , the attention inputs are defined as Q f = H f W Q , K s = H s , V s = H s (3) where W Q ∈ R D × D are learnable projection matrices. Multi-head attention is then applied to compute fused representations: Z = MultiHeadAttn ( Q f , K s , V s ) , (4) allowing each subject’ s functional representation to selectively attend to structurally rele vant pat- terns across the population graph. Because the attention mechanism is driven by the functional query representation, the resulting fusion process is naturally guided by functional connecti vity pat- terns, while structural embeddings provide complementary contextual information. This asymmetric formulation ensures that functional connectivity remains the principal source of information in the fused representation while structural connecti vity contributes additional context that enhances cross- modal interaction. 2.3.2. Residual Refinement T o stabilize training and preserve the discriminative properties of the functional embeddings, a residual connection is applied between the projected functional embeddings and the attention output, maintaining the intended asymmetry of the fusion process: Z r = Z + Q f . (5) 14 The fused representations are subsequently refined using a feed-forward MLP with layer normaliza- tion and non-linear activ ation: ˜ Z = MLP ( Z r ) , (6) where the MLP consists of a sequence of layer normalization, linear transformation, GELU acti va- tion, dropout, and a final linear projection. This refinement step enhances the e xpressiv e capacity of the fused embeddings while maintaining the original functional signal. 2.4. Classification Module The refined embeddings ˜ Z ∈ R N × D are then passed to a linear classification layer: ˆ Y = ˜ Z W c + b , (7) where W c ∈ R D × C and b ∈ R C are learnable parameters, and C denotes the number of diagnos- tic classes. The entire framework, comprising the dual EV -GCN encoders, the asymmetric cross- attention fusion module, and the classification layer , is trained end-to-end using the cross-entropy loss: L = − N X i = 1 C X c = 1 y ic log ˆ y ic . (8) This joint optimization enables coordinated learning of modality-specific embeddings and cross- modal interactions, e ff ectiv ely capturing complex relationships in functional and structural connec- tivity for ASD classification. 3. Experiments and Results This section presents a comprehensive e valuation of the proposed multimodal population-graph framew ork for ASD classification using the ABIDE-I dataset. The experimental analysis in vestigates the e ff ectiv eness of the proposed framework under di ff erent modality configurations and examines the contribution of the proposed asymmetric cross-attention–based fusion strate gy . Experiments are performed using both stratified 10-fold and LOSO cross-validation methods to ev aluate robustness and generalization across the sites. The stratified 10-fold cross-v alidation setup assesses subject-lev el generalization by preserving class balance across folds, while LOSO- CV specifically examines the model’ s capacity to generalize to unseen acquisition sites. These 15 assessment techniques provide a comprehensive e valuation of performance even when multi-center neuroimaging datasets exhibit inter -site variation. In our experiments, the spectral graph con volution is implemented using a Chebyshev polyno- mial approximation of order K = 3, enabling localized yet expressi ve graph filtering, while the network depth is defined by L G = 4 graph con volutional layers to facilitate hierarchical feature ag- gregation across the population graph. All models are optimized using the Adam optimizer with an initial learning rate of 0 . 01 and a weight decay of 5 × 10 − 5 to control model comple xity . T o further reduce overfitting, a dropout rate of 0 . 2 is applied during training. Each model is trained for 300 epochs. All models are de veloped on Python with experiments conducted on the open-source PyT orch framew ork, with graph computations handled via PyT orch Geometric, and e xperiments run on an NVIDIA R TX 4000 Ada Generation GPU. 3.1. P erformance Evaluation The proposed model was ev aluated by using Accurac y , Area under the Curve, Sensitivity , Speci- ficity , and F1-score ev aluation metrics. T ogether , these metrics provide a comprehensi ve assessment of classification performance, particularly in multi-site neuroimaging settings where data hetero- geneity must be carefully considered. Accuracy is defined as: Acc = T P + T N T P + T N + F P + F N . (9) Sensitivity (T rue Positive Rate) and Specificity (T rue Negati ve Rate) are computed as: Sensitivity = T P T P + F N , Specificity = T N T N + F P . (10) The F1-score, which provides a balanced measure of precision and recall, is gi ven by: F1 = 2 · Precision · Recall Precision + Recall . (11) The Area Under the Receiver Operating Characteristic Curve (A UC) is computed by plotting the true positiv e rate against the False Positi ve Rate. All metrics are computed exclusi vely on held-out test subjects in each validation split and re- ported as averages across cross-validation folds and acquisition sites, ensuring an unbiased and sta- 16 tistically reliable assessment of model performance. 3.2. E ff ect of Multimodal Learning T o analyze the contribution of di ff erent imaging modalities and non-imaging phenotypic in- formation, we ev aluated sev eral configurations of the proposed framework using stratified 10-fold cross-validation while keeping the baseline EV -GCN architecture unchanged. The results are sum- marized in T able 1. T able 1: E ff ect of Multimodal Learning, ev aluated using stratified 10-fold cross-validation Model Modality A UC Acc Sensitivity Specificity F1-Score rs-fMRI 79.40 77.40 76.70 84.41 80.00 Proposed rs-fMRI* 84.80 82.60 83.90 85.30 84.10 Method sMRI 73.90 73.30 73.00 81.00 76.40 sMRI* 82.30 81.00 86.73 76.50 80.90 Multimodal 87.30 84.40 85.20 87.00 85.70 Note: (*) indicates the inclusion of non-imaging data alongside the specified imaging modality . rs-fMRI by itself already produces competitive results, demonstrating the ability of functional connectivity patterns to discriminate in the categorization of ASD. Incorporating non-imaging phe- notypic data regularly enhances performance, suggesting that adaptive modeling of inter-subject connections o ff ers useful context at the population le vel. sMRI alone achie ves lower performance compared to rs-fMRI, reflecting its limited sensitivity to functional brain dynamics. Howe ver , when structural connecti vity is combined with non-imaging phenotypic information, classification performance improves substantially , demonstrating the ef- fectiv eness of the adaptive population graph in leveraging complementary non-imaging attributes. Moreov er, when compared with rele vant state-of-the-art approach for the sMRI modality , the pro- posed model achiev es superior performance, as summarized in T able 2. T able 2: Results comparison for sMRI + non-imaging Data , e valuated using stratified 10-fold cross-v alidation Reference Model A UC Acc [16] FCN 69.60 66.20 Proposed Method 82.30 81.00 17 The best ov erall performance is achie ved through multimodal fusion, where functional and struc- tural information are jointly modeled within the proposed asymmetric cross-attention frame work. This configuration attains an A UC of 87.30%, an accurac y of 84.40%, a sensiti vity of 85.20%, a specificity of 87.00%, and an F1-score of 85.70%, confirming that multimodal integration enables richer and more robust subject representations than any single imaging modality . For results visual- ization of the proposed model, R OC curves are plotted in Fig.4 0.0 0.2 0.4 0.6 0.8 1.0 F alse P ositive R ate 0.0 0.2 0.4 0.6 0.8 1.0 T rue P ositive R ate ROC Curves 10-F old Cr oss V alidation F old 0 (AUC = 0.8267) F old 1 (AUC = 0.8727) F old 2 (AUC = 0.9274) F old 3 (AUC = 0.9303) F old 4 (AUC = 0.7617) F old 5 (AUC = 0.8032) F old 6 (AUC = 0.9798) F old 7 (AUC = 0.9590) F old 8 (AUC = 0.8984) F old 9 (AUC = 0.7649) Mean ROC (AUC = 0.8724) Figure 4: R OC–A UC curve for the proposed model under 10-fold cross-validation 3.3. E ff ect of asymmetric Cr oss-Attention Fusion T o ev aluate the e ff ecti veness of the proposed asymmetric cross-attention mechanism for mul- timodal feature fusion, we compare it against a conv entional concatenation [18] and symmetric cross-attention-based [29] fusion strategies, using stratified 10-fold cross-validation. In the concate- nation approach, modality-specific embeddings are directly merged and passed to the classification head, without e xplicitly modeling inter-modality interactions, and for symmetric (two-stream) cross 18 attention, the query is exchanged from each modality . In contrast, the proposed asymmetric cross- attention fusion selecti vely integrates structural information by allo wing rs-fMRI to attend sMRI, enabling adaptive and context-a ware feature alignment. The quantitative results are reported in T a- ble 3. Across all assessment measures, the asymmetric cross-attention-based fusion outperforms simple concatenation and symmetric cross-attention. T able 3: E ff ect of the proposed asymmetric cross-attention–based fusion strategy on multimodal ASD classification perfor- mance, ev aluated using stratified 10-fold cross-validation Multimodal Fusion Strategy A UC Acc Sens Spec F1-Score Concatenation 84.90 83.00 84.90 84.20 84.30 Symmetric Cross-Attention 82.30 81.30 84.40 82.30 82.90 Asymmetric Cross-Attention 87.30 84.40 85.20 87.00 85.70 3.4. Comparison with State-of-the-Art Methods W e compare the proposed model with existing population-graph and deep learning–based ap- proaches for ASD classification using stratified 10-fold cross-validation. In particular , the ev aluation includes comparisons with several state-of-the-art methods, including Brain Network T ransformer [30], Com-BrainTF [31], ASDFormer [32], DeepGCN [27], MMGCN [33], GCN [16], EV -GCN [14], and Enhanced EV -GCN [17]. All these studies used the same 871 number of images from the ABIDE-I dataset within consistent data pre-processing. 19 T able 4: Stratified 10-fold cross-validation based results comparison with SO T A ASD classification methods Reference Model Modality A UC A CC [30] Brain Network T ransformer rs-fMRI* 80.2 71.0 [31] Com-BrainTF rs-fMRI* 79.6 72.5 [32] ASDF ormer rs-fMRI* 81.17 74.60 [27] DeepGCN rs-fMRI* 82.59 77.27 [33] MMGCN rs-fMRI* 84.00 78.31 [14] EV -GCN rs-fMRI* 84.72 81.06 [16] GCN Multimodality 78.00 71.30 [17] Enhanced EvGCN Multimodality 82.00 74.81 Proposed Method Multimodality 87.30 84.40 Note:(*) r epresents inte gration of non-imaging data into ima ging modality Multimodality r epresents all modalities, i.e ., rs-fMRI, sMRI, and Non-Imaging. Our proposed multimodal architecture performs better overall across all provided measures, as shown in T able 4. Although current graph learning techniques and transformer-based models en- hance performance by modeling population-level interactions and long-range dependencies, they are still limited by their dependence on a single imaging modality [30, 31, 32, 27, 33, 14]. Among population-graph approaches, EV -GCN [14] has sho wn strong performance by jointly learning node representations and adapti ve graph structures from resting-state functional connecti v- ity and non-imaging phenotypic information. Building on this foundation, the proposed framew ork further enhances classification accuracy and area under the curve by explicitly incorporating struc- tural connectivity alongside functional and non-imaging attrib utes. Through end-to-end optimiza- tion of adaptiv e edge weights, graph conv olutional representations, and a novel asymmetric cross- attention–based fusion mechanism instead of simple concatenation-based fusion like [17], helps the model to e ff ectiv ely capture both intra-subject characteristics and inter -subject relationships. This multimodal design enables the e xtraction of complementary aspects of brain organization that are not fully accessible from functional data alone, resulting in consistent improv ements ov er strong baseline methods and o ff ering a robust solution for multi-site autism classification. 20 3.5. Leave-One-Site-Out Based Evaluation A LOSO-CV was performed using the ABIDE-I dataset to assess the robustness of the proposed framew ork to site-specific variability and its capacity to generalize across varied acquisition condi- tions. Under the LOSO ev aluation protocol, data from a single imaging site is held out entirely for testing, while the model is trained on all remaining sites. This process is repeated until each site serves once as the unseen test set, allowing a comprehensi ve assessment of cross-site generalization. This assessment procedure rev eals the model to di ff erences in scanner hardware, acquisition meth- ods, and demographic distributions, o ff ering a rigorous e xamination of cross-site generalization. Under LOSO-CV , the proposed approach attains the highest a verage accurac y of 82.0%, out- performing existing LOSO-CV style e valuation methods [34], [35], and [27] as shown in T able 5. It achie ves superior performance in 12 of the 17 sites as shown in Bar Plots in Fig 4, indicating improv ed robustness to variability across imaging centers. Importantly , the performance gains are evident not only in larger cohorts such as NYU, TRINITY , and UCLA, but also in smaller sites, including CMU, ST ANFORD, and Y ALE. This consistent improv ement across both lar ge and small datasets suggests that the adapti ve population graph construction, together with multimodal fusion, enables the model to better utilize heterogeneous training data and generalize more reliably across di ff erent acquisition settings. 21 T able 5: Site-wise classification accuracy (%) under LOSO-CV e valuation Site Size ASD- ASD- DeepGCN Our DiagNet SAENet Proposed CAL TECH 15 52.8 56.7 71.7 86.7 CMU 11 68.5 70.6 80.5 81.8 KKI 33 69.5 72.6 76.2 63.6 LEUVEN 56 61.3 64.6 74.3 71.4 MAX_MUN 46 48.6 47.5 69.7 87.0 NYU 172 68.0 72.0 79.1 91.9 OHSU 25 82.0 72.0 78.6 72.0 OLIN 28 65.1 66.6 80.0 82.1 PITT 50 67.8 73.1 72.4 88.0 SBL 26 51.6 56.6 79.7 76.9 SDSU 27 63.0 64.2 67.7 81.5 ST ANFORD 25 64.2 53.2 68.3 92.0 TRINITY 44 54.1 57.5 66.9 95.5 UCLA 85 73.2 68.3 72.8 83.5 UM 120 63.8 67.8 81.2 78.3 USM 67 68.2 70.0 67.5 79.1 Y ALE 41 63.6 66.0 75.6 82.9 A verage 51.2 63.8 64.7 74.2 82.0 While the model demonstrates consistent impro vements across most sites, some performance variability across sites is evident, particularly for smaller cohorts such as KKI, OHSU, and SBL. This trend aligns with previous LOSO studies on ABIDE and is likely due to limited sample sizes and site-specific distribution shifts. 22 0 20 40 60 80 1 0 0 1 2 0 Ch art Titl e AS D -D iag N E T AS D -S AE N e t D ee p GCN Pro pos ed Metho d Figure 5: Bar plot comparing the performance of the proposed model with SO T A methods under LOSO cross-validation 4. Discussion The e xperimental findings demonstrate that the proposed multimodal graph-based fusion frame- work e ff ectiv ely captures complementary functional and structural brain patterns for ASD classifi- cation. In particular, the integration of adaptive population graph modeling with asymmetric cross- attention-based multimodal fusion improv ed performance compared to single-modality and e xisting state-of-the-art approaches, as shown in T ables 1 and 4, respectiv ely . These results suggest that jointly modeling inter -subject relationships and modality-specific representations pro vides a more discriminativ e and biologically meaningful characterization of brain connectivity alterations. The ablation analysis indicates that rs-fMRI provides stronger discriminative power than sMRI when used independently , which is consistent with prior literature highlighting the sensitivity of functional connectivity to atypical network synchronization in ASD. Transformer -based and graph- based models operating on rs-fMRI, such as Brain Network T ransformer [30], Com-BrainTF [31], and ASDFormer [32], hav e demonstrated that modeling long-range dependencies enhances classi- fication performance. Howe ver , these approaches are primarily constrained to functional data. The results of the present study suggest that although rs-fMRI alone is competitiv e, incorporating struc- tural connectivity further enriches the representation space by capturing anatomical organization that functional measures alone cannot fully characterize. 23 For sMRI combined with non-imaging data, the proposed framework substantially outperforms previously reported model FCN [16]. This impro vement highlights the importance of population- graph modeling when structural connecti vity is considered. Unlike con ventional deep learning mod- els that operate at the subject level, the adaptiv e graph construction enables non-imaging phenotypic attributes to guide inter -subject relationships, providing conte xtual information that enhances dis- crimination ev en when the imaging modality itself is less sensitive than rs-fMRI. These findings re- inforce the rele vance of incorporating demographic and acquisition-related factors into graph-based neuroimaging framew orks. The superiority of the multimodal configuration further confirms that functional and structural connectivity encode complementary aspects of brain organization. While functional connectiv- ity reflects dynamic synchronization patterns, structural connectivity represents relativ ely stable anatomical pathways. The asymmetric cross-attention mechanism enables selective information exchange between modalities, allowing functional embeddings to attend to structural representa- tions in a context-aw are manner . Compared to simple concatenation based fusion [17], which treats modality-specific embeddings independently before classification, cross-attention explicitly models inter-modality dependencies, leading to more expressi ve fused representations. This observ ation aligns with emerging e vidence that attention-based fusion strategies provide advantages ov er static fusion schemes in multimodal neuroimaging analysis. In comparison with state-of-the-art methods, the proposed framew ork achie ves improved per- formance across both transformer and graph-based approaches. Population-graph models such as EV -GCN [14] demonstrated the e ff ectiveness of jointly learning node embeddings and adaptiv e graph structures from rs-fMRI and non-imaging phenotypic data. Building upon this foundation, the present study extends the modeling capacity by incorporating structural connectivity and introduc- ing asymmetric cross-attention for multimodal integration. Similarly , multimodal graph frame works such as those reported in [16] and enhanced EV -GCN variants [17] impro ve representation learning through multi-source information; ho wev er, they rely on simpler fusion strategies. The consistent gains observed in the current experiments suggest that explicitly modeling modality interaction, in addition to adaptiv e graph learning, contributes to impro ved discriminativ e performance. The LOSO-CV ev aluation provides further insight into the generalization capability of the frame- work under realistic multi-center conditions. Compared with ASD-DiagNet [34], SAENet [35], and 24 DeepGCN [27], the proposed model achiev es the highest average accuracy and demonstrates supe- rior performance across the majority of acquisition sites. This indicates that adapti ve phenotypic- driv en edge construction, combined with multimodal feature integration, enhances robustness to inter-site variability . The improved performance across both large cohorts and several smaller sites suggests that the model e ff ecti vely le verages heterogeneous training data to learn more sta- ble population-lev el representations. Nev ertheless, variability across certain smaller sites remains observable, particularly in cohorts with limited sample sizes. Such fluctuations are consistent with previously reported LOSO findings on ABIDE, where site-specific distribution shifts and demographic imbalance influence generaliza- tion performance. Although the proposed framework improves the overall av erage accuracy , these observations highlight the ongoing challenge of domain heterogeneity in multi-site neuroimaging studies. 5. Conclusion In this study , we proposed a multimodal population-graph framew ork for ASD classification that integrates rs-fMRI, sMRI, and non-imaging phenotypic information within a unified learning strategy . By modeling subjects as nodes in an adaptiv e graph and learning inter -subject relation- ships through non-imaging phenotypic a ffi nity , the frame work e ff ectiv ely captures both indi vidual connectivity patterns and population-le vel structure. The introduction of an asymmetric cross-attention fusion mechanism allows functional repre- sentations to selectively incorporate complementary structural information, leading to improved discriminativ e performance compared to single imaging modality and simple fusion approaches. Experimental results under both 10-fold and LOSO cross-v alidation demonstrate the robustness and generalization of the proposed model across multi-site data. Overall, this work emphasizes ho w crucial it is to create fusion strategies that use complementary information while respecting modality hierarchy . The proposed framework provides a scalable and interpretable approach for multimodal neuroimaging analysis and may serve as a foundation for future research in computer-aided diagnosis of neurode velopmental and psychiatric disorders. 25 Acknowledgements This study was funded by Gesellschaft für Forschungsförderung Niederösterreich m.b.H. under project number: FTI24-G-024 References [1] C. Lord, T . S. Brugha, T . Charman, J. Cusack, G. Dumas, T . Frazier , E. J. Jones, R. M. Jones, A. Pickles, M. W . State, et al., Autism spectrum disorder , Nature re views Disease primers 6 (1) (2020) 5. [2] C. Ecker , S. Y . Bookheimer , D. G. Murphy , Neuroimaging in autism spectrum disorder: brain structure and function across the lifespan, The Lancet Neurology 14 (11) (2015) 1121–1134. [3] S. S. Dcouto, J. Pradeepkandhasamy , Multimodal deep learning in early autism detec- tion—recent advances and challenges, Engineering Proceedings 59 (1) (2024) 205. [4] N. Qiang, J. Gao, Q. Dong, J. Li, S. Zhang, H. Liang, Y . Sun, B. Ge, Z. Liu, Z. W u, et al., A deep learning method for autism spectrum disorder identification based on interactions of hierarchical brain networks, Beha vioural Brain Research 452 (2023) 114603. [5] A. S. Heinsfeld, A. R. Franco, R. C. Craddock, A. Buchweitz, F . Meneguzzi, Identification of autism spectrum disorder using deep learning and the abide dataset, NeuroImage: clinical 17 (2018) 16–23. [6] Z. Sherkatghanad, M. Akhondzadeh, S. Salari, M. Zomorodi-Moghadam, M. Abdar, U. R. Acharya, R. Khosrowabadi, V . Salari, Automated detection of autism spectrum disorder using a con volutional neural network, Frontiers in neuroscience 13 (2020) 1325. [7] M. Quaak, L. V an De Mortel, R. M. Thomas, G. V an W ingen, Deep learning applications for the classification of psychiatric disorders using neuroimaging data: systematic revie w and meta-analysis, NeuroImage: Clinical 30 (2021) 102584. [8] L. Zhang, M. W ang, M. Liu, D. Zhang, A surv ey on deep learning for neuroimaging-based brain disorder analysis, Frontiers in neuroscience 14 (2020) 779. 26 [9] H. Peng, W . Gong, C. F . Beckmann, A. V edaldi, S. M. Smith, Accurate brain age prediction with lightweight deep neural networks, Medical image analysis 68 (2021) 101871. [10] M. Khosla, K. Jamison, A. Kuceyeski, M. R. Sabuncu, Ensemble learning with 3d con volu- tional neural networks for functional connectome-based prediction, NeuroImage 199 (2019) 651–662. [11] S. P arisot, S. I. Ktena, E. Ferrante, M. Lee, R. Guerrero, B. Glocker , D. Rueckert, Disease prediction using graph con volutional networks: application to autism spectrum disorder and alzheimer’ s disease, Medical image analysis 48 (2018) 117–130. [12] F . M. Bianchi, D. Grattarola, L. Livi, C. Alippi, Graph neural networks with con volutional arma filters, IEEE transactions on pattern analysis and machine intelligence 44 (7) (2021) 3496–3507. [13] H. Jiang, P . Cao, M. Xu, J. Y ang, O. Zaiane, Hi-gcn: A hierarchical graph con volution network for graph embedding learning of brain network and brain disorders prediction, Computers in Biology and Medicine 127 (2020) 104096. [14] Y . Huang, A. C. Chung, Edge-variational graph con volutional networks for uncertainty-aw are disease prediction, in: International Conference on Medical Image Computing and Computer- Assisted Intervention, Springer , 2020, pp. 562–572. [15] M. Cao, M. Y ang, C. Qin, X. Zhu, Y . Chen, J. W ang, T . Liu, Using deepgcn to identify the autism spectrum disorder from multi-site resting-state data, Biomedical Signal Processing and Control 70 (2021) 103015. [16] Y . Dong, D. Batalle, M. Deprez, A framework for comparison and interpretation of machine learning classifiers to predict autism on the abide dataset, Human Brain Mapping 46 (5) (2025) e70190. [17] A. F . Ashrafi, M. H. Kabir , Enhanced graph con volutional network with chebyshev spectral graph and graph attention for autism spectrum disorder classification, in: 2025 40th Interna- tional Conference on Image and V ision Computing New Zealand (IVCNZ), IEEE, 2025, pp. 1–6. 27 [18] P . Xu, X. Zhu, D. A. Clifton, Multimodal learning with transformers: A surve y , IEEE Trans- actions on Pattern Analysis and Machine Intelligence 45 (10) (2023) 12113–12132. [19] D.-Y . Song, C.-C. T opriceanu, D. C. Ilie-Ablachim, M. Kinali, S. Bisdas, Machine learning with neuroimaging data to identify autism spectrum disorder: a systematic revie w and meta- analysis, Neuroradiology 63 (12) (2021) 2057–2072. [20] C. P . Santana, E. A. de Carv alho, I. D. Rodrigues, G. S. Bastos, A. D. de Souza, L. L. de Brito, rs-fmri and machine learning for asd diagnosis: a systematic re view and meta-analysis, Scien- tific reports 12 (1) (2022) 6030. [21] L. Pang, X. Zhao, L. Zhao, J. Li, F . Kuo, H. W ang, C. Liu, Multi-modal data analysis for autism spectrum disorder in children: State of the art and trends, EngMedicine 3 (1) (2026) 100117. [22] A. Di Martino, C.-G. Y an, Q. Li, E. Denio, F . X. Castellanos, K. Alaerts, J. S. Anderson, M. As- saf, S. Y . Bookheimer , M. Dapretto, et al., The autism brain imaging data exchange: towards a large-scale e v aluation of the intrinsic brain architecture in autism, Molecular psychiatry 19 (6) (2014) 659–667. [23] A. Abraham, M. P . Milham, A. Di Martino, R. C. Craddock, D. Samaras, B. Thirion, G. V aro- quaux, Deriving reproducible biomarkers from multi-site resting-state data: An autism-based example, NeuroImage 147 (2017) 736–745. [24] A. Kazi, S. Shekarforoush, S. Arvind Krishna, H. Burwinkel, G. V iv ar, K. Kortüm, S.-A. Ah- madi, S. Albarqouni, N. Nav ab, Inceptiongcn: recepti ve field aware graph conv olutional net- work for disease prediction, in: International conference on information processing in medical imaging, Springer , 2019, pp. 73–85. [25] R. C. Craddock, S. Jbabdi, C.-G. Y an, J. T . V ogelstein, F . X. Castellanos, A. Di Martino, C. Kelly , K. Heberlein, S. Colcombe, M. P . Milham, Imaging human connectomes at the macroscale, Nature methods 10 (6) (2013) 524–539. [26] B. Fischl, Freesurfer , Neuroimage 62 (2) (2012) 774–781. [27] M. W ang, J. Guo, Y . W ang, M. Y u, J. Guo, Multimodal autism spectrum disorder diagnosis method based on deepgcn, IEEE Transactions on Neural Systems and Rehabilitation Engineer- ing 31 (2023) 3664–3674. 28 [28] X. Ding, F . Y ang, F . Ma, An e ffi cient model selection for linear discriminant function-based recursiv e feature elimination, Journal of Biomedical Informatics 129 (2022) 104070. [29] J. Lu, D. Batra, D. Parikh, S. Lee, V ilbert: Pretraining task-agnostic visiolinguistic represen- tations for vision-and-language tasks, Adv ances in neural information processing systems 32 (2019). [30] X. Kan, W . Dai, H. Cui, Z. Zhang, Y . Guo, C. Y ang, Brain network transformer , Advances in Neural Information Processing Systems 35 (2022) 25586–25599. [31] A. Bannadabhavi, S. Lee, W . Deng, R. Y ing, X. Li, Community-aware transformer for autism prediction in fmri connectome, in: International conference on medical image computing and computer-assisted interv ention, Springer , 2023, pp. 287–297. [32] M. Izadi, M. Safayani, Asdformer: A transformer with mixtures of pooling-classifier experts for rob ust autism diagnosis and biomarker disco very , arXi v preprint arXi v:2508.14005 (2025). [33] T . Song, Z. Ren, J. Zhang, M. W ang, Multi-view and multimodal graph con volutional neural network for autism spectrum disorder diagnosis, Mathematics 12 (11) (2024) 1648. [34] T . Eslami, V . Mirjalili, A. Fong, A. R. Laird, F . Saeed, Asd-diagnet: a hybrid learning approach for detection of autism spectrum disorder using fmri data, Frontiers in neuroinformatics 13 (2019) 70. [35] F . Almuqhim, F . Saeed, Asd-saenet: a sparse autoencoder, and deep-neural network model for detecting autism spectrum disorder (asd) using fmri data, Frontiers in Computational Neuro- science 15 (2021) 654315. 29
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment