Vision-Language Model Based Multi-Expert Fusion for CT Image Classification
Robust detection of COVID-19 from chest CT remains challenging in multi-institutional settings due to substantial source shift, source imbalance, and hidden test-source identities. In this work, we propose a three-stage source-aware multi-expert fram…
Authors: Jianfa Bai, Kejin Lu, Runtian Yuan
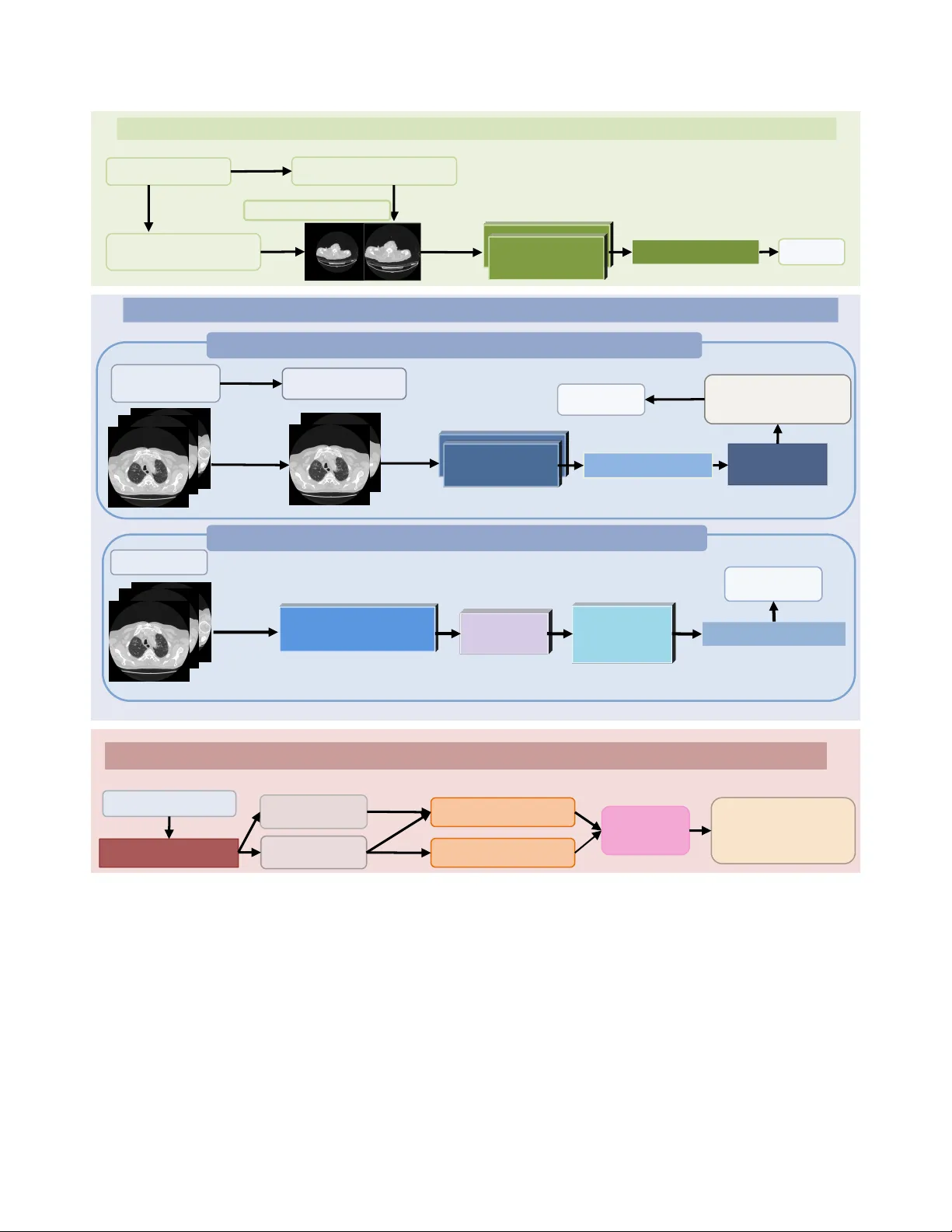
V ision-Language Model Based Multi-Expert Fusion f or CT Image Classification Jianfa Bai 1 , K ejin Lu 1 , Runtian Y uan 1 , Qingqiu Li 1 , Jilan Xu 2 , Junlin Hou 3 ∗ , Y uejie Zhang 1 ∗ , Rui Feng 1 ∗ 1 College of Computer Science and Artificial Intelligence, Shanghai K ey Laboratory of Intelligent Information Processing, Fudan Univ ersity 2 Uni versity of Oxford 3 The Hong K ong Univ ersity of Science and T echnology Abstract Robust detection of CO VID-19 fr om chest CT r emains chal- lenging in multi-institutional settings due to substantial sour ce shift, source imbalance, and hidden test-sour ce iden- tities. In this work, we pr opose a thr ee-stage sour ce- awar e multi-expert frame work for multi-sour ce CO VID-19 CT classification. F irst, we build a lung-aware 3D expert by combining original CT volumes and lung-extracted CT volumes for volumetric classification. Second, we develop two MedSigLIP-based e xperts: a slice-wise r epr esentation and pr obability learning module, and a T ransformer -based inter-slice conte xt modeling module for capturing cross- slice dependency . Thir d, we train a sour ce classifier to pr e- dict the latent sour ce identity of each test scan. By lever- aging the pr edicted sour ce information, we perform model fusion and voting based on differ ent experts. On the val- idation set covering all four sour ces, the Stage 1 model achie ves the best macr o-F1 of 0.9711, ACC of 0.9712, and A UC of 0.9791. Stage 2a and Stage 2b achieve the best A UC scor es of 0.9864 and 0.9854, r espectively . Stage 3 sour ce classifier reac hes 0.9107 A CC and 0.9114 F1. These re- sults demonstrate that source-awar e expert modeling and hierar chical voting pr ovide an effective solution for r obust CO VID-19 CT classification under hetero geneous multi- sour ce conditions. 1. Introduction Robust detection of CO VID-19 from chest CT is not merely a classification problem, but a domain-generalization prob- lem under substantial variation between institutions [ 1 – 5 , 10 – 22 , 25 – 28 ]. In multi-center clinical settings, CT scans may differ markedly in scanner vendors, reconstruc- tion kernels, slice counts, field-of-view , and background ar - tifacts, while patient populations and disease presentations also vary across hospitals. As a result, a model trained on one source can perform well in-distrib ution yet degrade no- ticeably on unseen or shifted sources. This issue becomes ev en more critical in the Multi-Source CO VID-19 Detection Challenge, where the task explicitly inv olves four distinct medical sources and therefore directly ev aluates whether a method can maintain reliable performance under source shift rather than simply fit a single pooled dataset. T o address these limitations, we propose a three-stage source-awar e multi-expert framework for multi-source CO VID-19 CT classification. As illustrated in Fig. 1 , Stage 1 b uilds a lung-aw are 3D expert that performs lung extrac- tion and unified volume canonicalization before 3D classi- fication, reducing irrelev ant peripheral background and pre- serving volumetric morphology . Stage 2 introduces two complementary 2D experts based on a pretrained Med- SigLIP [ 23 ] encoder: Stage 2a performs slice-wise prob- ability learning by randomly sampling contiguous slices during training and av eraging per-slice predictions, while Stage 2b removes the original classification head and adds lightweight Transformer [ 24 ] encoder blocks to refine inter- slice context before classification. Stage 3 further trains a source classifier to estimate the latent source identity of each test scan and route samples to appropriate experts. T o improve stability , we also train multiple model variants within each step and aggregate their outputs by v oting. Our framework is designed together with challenge- specific data handling. W e first perform lung e xtraction for all scans and suppress excessiv e peripheral dark regions. W e then apply a unified preprocessing rule that discards the first and last 15% of slices when the slice count ex- ceeds 150, followed by stage-specific canonicalization for 3D and 2D pipelines. T o alleviate the zero-positiv e vali- dation issue of source 2, we include its 39 positive train- ing cases in validation-oriented source analysis. W e also correct the source 0 non-CO VID training count based on folder-le vel inspection and predict test-source identities for source-aware inference. These steps are important for reli- able model dev elopment under scarce and imperfect source supervision. Extensiv e experiments validate the effecti veness of the proposed design. Stage 1 achieves a best macro-F1 of 1 COVID VS. Non-COVID L ctx Stage 1. Lung-Aware 3D T rain ing for C OVID-19 Classific ation Stage 3. Source D iscriminat ion and S ource-Spec ific Expe rt Inferen ce Stage 2b. Transformer-Based Inter-Slice Context Modeling Source Classifier Lung Region Extraction Original Training Se t source 1/2/3 Stage 1 3D Ex pert MedSigLI P Encoder source 0 Canonicaliz e 128 × 256 × 256 Input CT Volum e Raw CT Volum es 3D Backbone Predict s ource Classification Head L 3D Slice Embeddings Transformer Encoder Blocks Augment Stage 2 2D Ex pert Stage 2. MedSigLIP -Based 2D Learning and Inter- Slice Co ntext Ref inement (24 × 448 × 448) L slice Mean Probability Across Slices Random Contiguous Sampling (24 × 448 × 448) Per-Slice Probabilities (12 × 448 × 448) Pretrained MedSigLIP (2D) Augment Stage 2a. Slice-Wise Representation and Probability Learning Classification Head Mixed Training Se t Expert Fusion Route Canonicaliz e 128 × 256 × 256 Classification Head Figure 1. Overvie w of the proposed three-stage source-aware multi-expert framework for multi-source CO VID-19 CT classification. Stage 1 builds a lung-aware 3D expert for volumetric classification. Stage 2 introduces two MedSigLIP-based 2D experts for slice-wise probability learning and inter-slice context modeling. Stage 3 performs source discrimination and source-specific expert inference, where source 0 is handled by the 3D expert and source 1/2/3 are jointly inferred by multiple e xperts through fusion and voting. 97.11% , A UC of 98.29% , and accuracy of 97.12% . Stage 2a reaches a best macro-F1 of 94.50% , A UC of 98.64% , and accuracy of 94.81% , while Stage 2b further improves to 95.82% macro-F1, 98.54% A UC, and 95.97% accurac y . The Stage 3 source classifier achie ves 91.07% accuracy and 91.14% F1, providing a reliable basis for source-aw are routing. Notably , the Stage 1 3D expert attains perfect accu- racy on source 0, which directly moti vates our final source- specific expert allocation strate gy . Our main contributions are summarized as follo ws: • W e reformulate multi-source CO VID-19 CT detection as a source-aware domain-rob ust classification problem 2 and explicitly address hidden test sources, source imbal- ance, and validation sparsity . • W e propose a three-stage multi-expert framework that combines a lung-aware 3D e xpert, two MedSigLIP-based 2D experts, and a source classifier for source-specific in- ference. • W e introduce a hierarchical voting and fusion strat- egy that integrates multiple models within each step and jointly exploits heterogeneous experts for source 1/2/3 prediction. • W e achiev e strong validation performance across stages, with the best Stage 1 model reaching 97.10% macro-F1, demonstrating the ef fectiveness of source-aware expert modeling for robust multi-source CO VID-19 CT classi- fication. 2. Methodology 2.1. Overview As illustrated in Fig. 1 , we propose a three-stage source- aware multi-expert frame work for rob ust multi-source CO VID-19 CT classification. Giv en a chest CT scan X with binary label y ∈ { 0 , 1 } , where y = 1 denotes CO VID and y = 0 denotes Non-CO VID , our goal is to learn comple- mentary experts under heterogeneous source distributions and perform source-aware inference at test time. All scans are first processed by lung region extraction to suppress excessi ve irrele v ant dark background around the body region. In addition, when the slice number exceeds 150, we discard the first 15% and the last 15% of slices to reduce unstable peripheral slices and excessi ve redun- dancy . After preprocessing, two stage-specific canonical- ization strate gies are adopted. For the 3D branch, each scan is con verted to a volume of size 128 × 256 × 256 . For the 2D branch, each scan is canonicalized to 24 × 448 × 448 . Our framew ork contains four trainable models. The first is a lung-aw are 3D classifier in Stage 1, which directly pre- dicts CO VID/Non-CO VID from canonicalized 3D CT vol- umes. The second is the Stage 2a model, which fine-tunes a pretrained MedSigLIP encoder by slice-wise representation and probability learning. The third is the Stage 2b model, which reuses the MedSigLIP encoder from Stage 2a, dis- cards its original classification head, and introduces two T ransformer encoder blocks for inter-slice context model- ing. The fourth is a source classifier in Stage 3, which pre- dicts the source identity of each scan and enables source- aware e xpert routing. At inference time, the final prediction is determined ac- cording to the predicted source. Since the Stage 1 3D ex- pert achiev es perfect accuracy on source 0 in our v alidation analysis, scans predicted as source 0 are classified by the Stage 1 expert alone. For scans predicted as source 1/2/3, the predictions from Stage 1 and Stage 2 are combined by expert voting and fusion. Moreov er , multiple model vari- ants are trained within each stage, and their predictions are aggregated by v oting to further improve rob ustness. 2.2. Stage 1: Lung-A ware 3D T raining f or CO VID- 19 Classification Stage 1 is designed to learn a robust volumetric expert from both ra w CT scans and lung-extracted CT scans, following the line of research established by prior CMC-based meth- ods [ 6 – 9 ]. After preprocessing and canonicalization, each scan is represented as a 3D volume V ∈ R 128 × 256 × 256 . T o improv e robustness against source-dependent background variation, we construct a mixed training set that con- tains both canonicalized original volumes and canonical- ized lung-extracted v olumes. Giv en an input volume V , a 3D ResNet-style back- bone [ 24 ] extracts a volumetric representation, which is then passed to a binary classification head for CO VID/Non- CO VID prediction. The output logit is denoted by z 3 D ∈ R 2 , and the corresponding probability is computed by soft- max. The Stage 1 objective is standard cross-entrop y: L 3 D = − 1 X c =0 1 ( y = c ) log p 3 D ( y = c | V ) . (1) During training, scan-level augmentation is applied to the 3D volumes, including cropping, resizing, and random rotation. Importantly , the same augmentation parameters are consistently applied to all slices within the same CT scan, while dif ferent scans use different random parameters. During validation, only deterministic resizing/rescaling is used. The optimized Stage 1 model serves not only as a strong 3D CO VID-19 expert, but also as the feature back- bone of the source classifier in Stage 3. 2.3. Stage 2a: Slice-Wise Repr esentation and Prob- ability Learning Stage 2a adapts a pretrained MedSigLIP encoder to the CO VID-19 CT task in a memory-ef ficient slice-wise man- ner . Each scan is first canonicalized to 24 × 448 × 448 . During training, instead of using all 24 slices at once, we randomly sample a contiguous 12-slice subsequence from the scan, denoted by ˜ U = { u τ , u τ +1 , . . . , u τ +11 } , where each slice u t ∈ R 448 × 448 . Each sampled slice is independently fed into the pre- trained MedSigLIP image encoder, followed by a binary classification head to obtain a slice-level probability . These 12 slice-lev el probabilities are then av eraged to produce a scan-lev el prediction. Formally , if p t ( y = c | u t ) de- notes the predicted probability of class c for the t -th sam- pled slice, then the scan-lev el probability is ¯ p slice ( y = c | ˜ U ) = 1 12 12 X t =1 p t ( y = c | u t ) . (2) 3 The corresponding training loss is L slice = − 1 X c =0 1 ( y = c ) log ¯ p slice ( y = c | ˜ U ) . (3) This design allo ws Stage 2a to adapt the pretrained Med- SigLIP encoder using scan-le vel supervision while av oiding excessi ve GPU memory consumption. At inference time, all 24 slices are used, and their probabilities are averaged in the same way to obtain the final Stage 2a prediction. 2.4. Stage 2b: T ransformer -Based Inter -Slice Con- text Modeling Although Stage 2a provides strong slice-lev el supervision, it does not explicitly model contextual dependency across adjacent slices. Therefore, in Stage 2b, we further build a sequence-lev el expert on top of the Stage 2a model. Specifically , the MedSigLIP encoder trained in Stage 2a is reused as the visual backbone. Its original classification head is discarded, and most encoder layers are frozen, with only the last two layers kept trainable. Each of the 24 canon- icalized slices is passed through this partially frozen en- coder to obtain a sequence of slice embeddings. These em- beddings are then fed into two Transformer encoder blocks to model inter-slice contextual relationships. The result- ing contextualized slice features are aggre gated into a scan- lev el representation, which is finally fed to a new binary classification head for CO VID/Non-CO VID prediction. Let U denote the full 24-slice scan and let p ctx ( y = c | U ) denote the final scan-le vel probability predicted by this module. The Stage 2b objective is L ctx = − 1 X c =0 1 ( y = c ) log p ctx ( y = c | U ) . (4) Only the last two unfrozen MedSigLIP layers, the two T ransformer encoder blocks, and the newly added classifi- cation head are updated in this stage. In this way , Stage 2b preserves the strong pretrained slice representation from Stage 2a while further introducing explicit cross-slice con- text modeling. 2.5. Stage 3: Source Discrimination and Source- Specific Expert Inference The purpose of Stage 3 is to predict the hidden source iden- tity of each scan and enable source-a ware inference. T o this end, we reuse the 3D backbone trained in Stage 1 and re- place its binary classification head with a new four-class source classification head. The Stage 1 3D backbone is kept fixed and only the source classification head is trained. The resulting model predicts the source label s ∈ { 0 , 1 , 2 , 3 } for each scan. T able 1. Official dataset split. Split Class S0 S1 S2 S3 T otal Train COVID 175 175 39 175 564 Train Non-COVID 165 165 165 165 660 V al CO VID 43 43 0 42 128 V al Non-CO VID 45 45 45 45 180 T est – – – – – 1488 Because the official validation split of source 2 con- tains no positiv e scans, we additionally include the 39 pos- itiv e training scans from source 2 in source-aware valida- tion analysis and model selection. After training, the source classifier is applied to the test set to estimate source identi- ties. At test time, source-aware e xpert inference is performed according to the predicted source. If a scan is predicted as source 0, the final label is directly determined by the Stage 1 3D expert, since this expert achieves perfect accuracy on source 0 in our v alidation experiments. If a scan is predicted as source 1/2/3, the predictions from Stage 1, Stage 2a, and Stage 2b are integrated by expert voting. Denoting the cor - responding step-level voted predictions by ˜ y (1) , ˜ y (2 a ) , and ˜ y (2 b ) , the final prediction can be written as ˆ y = V ote ˜ y (1) , ˜ y (2 a ) , ˜ y (2 b ) . (5) In practice, each stage contains multiple model v ariants with slightly dif ferent data processing settings but compa- rable performance, and their outputs are first aggregated within each step before cross-expert fusion. This hierarchi- cal voting strategy improves robustness and helps stabilize prediction under source heterogeneity and source-specific data imbalance. 3. Datasets and Experiments 3.1. Datasets The Multi-Source CO VID-19 Detection Challenge pro vides a multi-institutional chest CT dataset collected from four different sources. The of ficial split is summarized in T a- ble 1 . The training and validation sets contain binary la- bels ( CO VID vs. Non-CO VID ), whereas the test set contains 1488 unlabeled scans. A major challenge of this benchmark is that the data are highly heterogeneous across sources, not only in acquisition and appearance, but also in class com- position and source-specific sample balance. Howe ver , we found that directly using the official split is suboptimal for reliable model dev elopment and source- aware e valuation. Therefore, we performed three practical corrections. First, since the official validation split of Source 2 con- tains no positi ve CO VID cases, source-specific validation on this source is ill-posed. T o alleviate this issue, we addi- 4 T able 2. Revised train/v al statistics used in our experiments. Split Class S0 S1 S2 S3 T otal Train COVID 175 175 39 175 564 Train Non-COVID 230 165 165 165 725 V al CO VID 43 43 39 42 167 V al Non-CO VID 45 45 45 45 180 tionally include the 39 positiv e Source 2 training scans in validation-oriented source analysis and model selection. Second, for the Source 0 non-CO VID training data, the official ct scan 8 entry was found to contain multiple folders, each representing an individual CT sample. Based on a folder-le vel inspection, the number of ef fectiv e Source 0 non-COVID training scans was initially corrected from 165 to 231. Since ct scan 0 was confirmed to be absent, this number was subsequently adjusted to 230. Third, since the challenge does not provide source la- bels for the test set, we estimate test-source identities us- ing the Stage 3 source classifier . The raw predicted number of Source 0 test scans is 549, but because ct scan 492 also contains multiple CT samples and introduces ambigu- ity , we discard ct scan 492 and finally use 548 Source 0 test scans. The final predicted test-source distribution is 548/314/245/380 for Source 0/1/2/3, respectiv ely , resulting in 1487 vali d test scans used in our source-aw are inference pipeline. The corrected dataset statistics used in our experiments are summarized in T able 2 . 3.2. Experiments W e ev aluate each stage of the proposed framework sepa- rately and report the best-performing configurations. Since the challenge emphasizes robustness across heterogeneous sources, we mainly focus on macro-F1, while also report- ing accuracy (A CC) and area under the R OC curve (A UC) when av ailable. 3.2.1. Stage 1: Lung-A ware 3D Classification Stage 1 directly trains a 3D CO VID classifier on canon- icalized volumetric CT scans. W e compare sev eral input settings, including original CT volumes, lung-extracted CT volumes, and their combination, with or without additional rotation-based augmentation. T able 3 summarizes the re- sults. As shown in T able 3 , the best Stage 1 performance is obtained by combining Orig and Lung , which achie ves the highest accuracy ( 0.9712 ) and macro-F1 ( 0.9711 ). This re- sult indicates that the original scans and lung-focused scans capture complementary cues, and that their combination is more effecti ve than either input alone. Notably , Stage 1 reaches a perfect score on Source 0, providing strong jus- tification for using the Stage 1 expert alone on Source 0 in the final source-aware inference frame work. 3.2.2. Stage 2a: Slice-Wise Representation and Proba- bility Learning Stage 2a adapts a pretrained MedSigLIP encoder via slice- wise probability learning. W e ev aluate se veral input con- struction strategies, including contiguous random depth sampling from canonicalized 24 × 448 × 448 scans, con- tiguous random depth sampling from 128 × 256 × 256 scans, and a depth-random-sampling variant applied to the 24 × 448 × 448 setting. The quantitati ve results are reported in T able 4 . Overall, Stage 2a achiev es strong performance, with the best macro-F1 reaching 0.9450 under contiguous random sampling and the highest A UC reaching 0.9864 . These re- sults confirm that pretrained MedSigLIP can be effecti vely adapted to chest CT classification through slice-wise prob- ability supervision. 3.2.3. Stage 2b: T ransf ormer-Based Inter-Slice Context Modeling Stage 2b further models inter-slice dependency on top of the pretrained MedSigLIP encoder from Stage 2a. W e compare three settings: (1) training only the T ransformer blocks and the new classification head, (2) jointly training the Trans- former blocks, the classification head, and the last two Med- SigLIP layers, and (3) directly flattening visual features fol- lowed by classification. The results are summarized in T a- ble 5 . Among the three settings, Flat + cls deliv ers the best overall performance, achieving the highest macro-F1 ( 0.9582 ) and accuracy ( 0.9597 ). This result suggests that the pretrained MedSigLIP encoder already produces suf- ficiently discriminative slice representations, such that a lightweight classifier is enough to obtain strong classifica- tion performance. In contrast, T rans + last2 , which addi- tionally trains the Transformer encoder, the last two Med- SigLIP layers, and the ne wly introduced classification head, achiev es the highest A UC ( 0.9854 ), indicating improv ed ranking ability . 3.2.4. Stage 3: Source Classification Stage 3 predicts the source identity of each scan for source- aware expert routing. The source classifier is built on top of the Stage 1 3D backbone and ev aluated on the validation split with the additional Source 2 positiv e samples described abov e. The final Stage 3 source classifier achie ves an A CC of 0.9107 and an F1 score of 0.9114. These results indicate that the source classifier can pro- vide suf ficiently reliable source predictions for downstream source-aware expert inference. Combined with the obser- vation that Stage 1 achie ves perfect performance on Source 0, this makes it possible to route Source 0 scans directly to the 3D expert, while using multi-expert fusion for Source 1/2/3. 5 T able 3. Stage 1 results under different input configurations. Lung denotes the lung-extracted CT dataset, Orig denotes the original CT dataset, and Rot denotes random rotation augmentation. S0–S3 denote source-wise F1 scores on the four sources. Setting A CC Macro-F1 A UC S0 S1 S2 S3 Lung 0.9683 0.9682 0.9829 0.9886 0.9545 0.9523 0.9769 Lung + Rot 0.9568 0.9567 0.9803 1.0000 0.9204 0.9519 0.9540 Orig + Lung 0.9712 0.9711 0.9791 1.0000 0.9545 0.9642 0.9654 Orig 0.9625 0.9625 0.9776 0.9773 0.9773 0.9152 0.9770 T able 4. Stage 2a results using Orig + Lung inputs. “CRS” denotes contiguous random sampling and “DRS” denotes depth-random sampling. Setting A CC Macro-F1 A UC CRS (24×448×448) 0.9481 0.9450 0.9864 CRS (128×256×256) 0.9481 0.9425 0.9812 DRS (24×448×448) 0.9452 0.9440 0.9821 T able 5. Stage 2b results. Setting Macro-F1 ACC A UC T rans-only 0.9511 0.9539 0.9837 T rans + last2 0.9516 0.9539 0.9854 Flat + cls 0.9582 0.9597 0.9847 4. Conclusion In this work, we presented a three-stage source-aware multi- expert framew ork for robust multi-source CO VID-19 CT classification. Instead of relying on a single unified clas- sifier , our method explicitly addresses the ke y challenges of this benchmark, including strong inter-source heterogene- ity , source imbalance, source-specific v alidation difficulty , and hidden test-source identities. The proposed framew ork integrates a lung-aware 3D e xpert for volumetric modeling, two MedSigLIP-based 2D experts for slice-wise adaptation and inter-slice context modeling, and a source classifier for source-aware routing and e xpert fusion. The experimental results validate the effecti veness of this design. Stage 1 provides the strongest overall per- formance and achiev es perfect accuracy on Source 0, which supports source-specific expert allocation. Stage 2 further contributes complementary discriminativ e cues through pretrained 2D visual modeling and inter-slice re- finement. Stage 3 enables reliable source prediction and makes source-aw are inference feasible on the unlabeled test set. By combining multiple experts and aggregating their outputs through hierarchical voting, our frame work achiev es robust and stable performance under challenging multi-source conditions. Overall, our results suggest that multi-source CO VID- 19 CT detection should be addressed not only as a bi- nary classification problem, but also as a source-aware domain-robust inference problem. W e believ e the proposed framew ork provides a practical and ef fective paradigm for building reliable medical imaging systems under real-w orld cross-institutional heterogeneity . Acknowledgements. This work was supported by National Natural Science Foundation of China (No. 62576107), and the Shanghai Municipal Commission of Economy and Informatization, Corpus Construction for Large Language Models in Pediatric Respiratory Diseases (No.2024-GZL-RGZN-01013, and the Science and T echnology Commission of Shanghai Munici- pality (No.24511104200), and 2025 National Major Science and T echnology Project - Noncommunicable Chronic Diseases-National Science and T echnology Ma- jor Project, Research on the Pathogenesis of Pancreatic Cancer and Novel Strategies for Precision Medicine (No.2025ZD0552303). References [1] Anastasios Arsenos, Dimitrios K ollias, and Stefanos Kol- lias. A large imaging database and novel deep neural ar- chitecture for covid-19 diagnosis. In 2022 IEEE 14th Im- age , V ideo, and Multidimensional Signal Processing W ork- shop (IVMSP) , pages 1–5. IEEE, 2022. 1 [2] Anastasios Arsenos, Andjoli Davidhi, Dimitrios K ollias, Panos Prassopoulos, and Stefanos Kollias. Data-driven covid-19 detection through medical imaging. In 2023 IEEE International Conference on Acoustics, Speech, and Signal Pr ocessing W orkshops (ICASSPW) , pages 1–5. IEEE, 2023. [3] Jiawang Cao, Lulu Jiang, Junlin Hou, Longquan Jiang, Rui- wei Zhao, W eiya Shi, Fei Shan, and Rui Feng. Exploiting deep cross-slice features from ct images for multi-class pneu- monia classification. In 2021 IEEE International Conference on Image Pr ocessing (ICIP) , pages 205–209. IEEE, 2021. [4] Demetris Gerogiannis, Anastasios Arsenos, Dimitrios K ol- lias, Dimitris Nikitopoulos, and Stefanos K ollias. Covid- 19 computer-aided diagnosis through ai-assisted ct imaging 6 analysis: Deploying a medical ai system. In 2024 IEEE In- ternational Symposium on Biomedical Imaging (ISBI) , pages 1–4. IEEE, 2024. [5] Hayden Gunraj, Ali Sabri, Da vid Kof f, and Alexander W ong. Covid-net ct-2: Enhanced deep neural networks for detec- tion of covid-19 from chest ct images through bigger , more div erse learning. F rontier s in Medicine , 8:729287, 2022. 1 [6] Junlin Hou, Jilan Xu, Rui Feng, Y uejie Zhang, Fei Shan, and W eiya Shi. Cmc-cov19d: Contrastiv e mixup classification for covid-19 diagnosis. In Proceedings of the IEEE/CVF In- ternational Confer ence on Computer V ision , pages 454–461, 2021. 3 [7] Junlin Hou, Jilan Xu, Longquan Jiang, Shanshan Du, Rui Feng, Y uejie Zhang, Fei Shan, and Xiangyang Xue. Periphery-aware covid-19 diagnosis with contrasti ve repre- sentation enhancement. P attern Recognition , 118:108005, 2021. [8] Junlin Hou, Jilan Xu, Nan Zhang, Y i W ang, Y uejie Zhang, Xiaobo Zhang, and Rui Feng. Cmc v2: T ow ards more accu- rate covid-19 detection with discriminative video priors. In Eur opean Confer ence on Computer V ision , pages 485–499. Springer , 2022. [9] Junlin Hou, Jilan Xu, Nan Zhang, Y uejie Zhang, Xiaobo Zhang, and Rui Feng. Boosting covid-19 sev erity detec- tion with infection-aware contrastiv e mixup classification. In Eur opean Confer ence on Computer V ision , pages 537–551. Springer , 2022. 3 [10] Dimitrios Kollias, Athanasios T agaris, Andreas Stafylopatis, Stefanos Kollias, and Georgios T agaris. Deep neural archi- tectures for prediction in healthcare. Complex & Intelligent Systems , 4(2):119–131, 2018. 1 [11] Dimitrios Kollias, N Bouas, Y Vlaxos, V Brillakis, M Se- feris, Ilianna K ollia, Lev on Sukissian, James W ingate, and S K ollias. Deep transparent prediction through latent represen- tation analysis. arXiv pr eprint arXiv:2009.07044 , 2020. [12] Dimitris Kollias, Y Vlaxos, M Seferis, Ilianna K ollia, Le von Sukissian, James Wingate, and S K ollias. Transparent adap- tation in deep medical image diagnosis. In International W orkshop on the F oundations of T rustworthy AI Inte grat- ing Learning, Optimization and Reasoning , pages 251–267. Springer , 2020. [13] Dimitrios Kollias, Anastasios Arsenos, Lev on Soukissian, and Stefanos Kollias. Mia-cov19d: Covid-19 detection through 3-d chest ct image analysis. In Pr oceedings of the IEEE/CVF International Confer ence on Computer V ision , pages 537–544, 2021. [14] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. Ai-mia: Covid-19 detection and sev erity analysis through medical imaging. In European conference on computer vi- sion , pages 677–690. Springer , 2022. [15] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. Ai-enabled analysis of 3-d ct scans for diagnosis of co vid-19 & its se verity . In 2023 IEEE International Confer ence on Acoustics, Speec h, and Signal Pr ocessing W orkshops (ICAS- SPW) , pages 1–5. IEEE, 2023. [16] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. A deep neural architecture for harmonizing 3-d input data analysis and decision making in medical imaging. Neuro- computing , 542:126244, 2023. [17] Dimitrios Kollias, Anastasios Arsenos, and Stefanos K ol- lias. Domain adaptation explainability & fairness in ai for medical image analysis: Diagnosis of covid-19 based on 3-d chest ct-scans. In Proceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition , pages 4907– 4914, 2024. [18] Dimitrios K ollias, Anastasios Arsenos, James W ingate, and Stefanos Kollias. Sam2clip2sam: V ision language model for segmentation of 3d ct scans for covid-19 detection. arXiv pr eprint arXiv:2407.15728 , 2024. [19] Dimitrios K ollias, Anastasios Arsenos, and Stefanos K ollias. Pharos-afe-aimi: Multi-source & fair disease diagnosis. In Pr oceedings of the IEEE/CVF International Conference on Computer V ision , pages 7265–7273, 2025. [20] David Krueger , Ethan Caballero, Joern-Henrik Jacobsen, Amy Zhang, Jonathan Binas, Dinghuai Zhang, Remi Le Priol, and Aaron Courville. Out-of-distribution general- ization via risk extrapolation (rex). In International confer - ence on machine learning , pages 5815–5826. PMLR, 2021. [21] Qingqiu Li, Runtian Y uan, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Advancing covid-19 de- tection in 3d ct scans. In Pr oceedings of the IEEE/CVF Con- fer ence on Computer V ision and P attern Recognition , pages 5149–5156, 2024. [22] Qingqiu Li, Runtian Y uan, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Adv ancing lung disease diagnosis in 3d ct scans. In Proceedings of the IEEE/CVF International Conference on Computer V ision , pages 7377– 7382, 2025. 1 [23] Andrew Sellergren, Sahar Kazemzadeh, T iam Jaroen- sri, Atilla Kiraly , Madeleine Tra verse, Timo Kohlber ger, Shawn Xu, Fayaz Jamil, C ´ ıan Hughes, Charles Lau, et al. Medgemma technical report. arXiv pr eprint arXiv:2507.05201 , 2025. 1 [24] Ashish V aswani, Noam Shazeer , Niki Parmar , Jakob Uszko- reit, Llion Jones, Aidan N Gomez, Łukasz Kaiser, and Illia Polosukhin. Attention is all you need. Advances in neural information pr ocessing systems , 30, 2017. 1 , 3 [25] Zifeng W ang, Zhenbang W u, Dinesh Agarwal, and Jimeng Sun. Medclip: Contrastiv e learning from unpaired medical images and text. In Pr oceedings of the 2022 Conference on Empirical Methods in Natural Language Pr ocessing , pages 3876–3887, 2022. 1 [26] Runtian Y uan, Qingqiu Li, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Domain adaptation us- ing pseudo labels for covid-19 detection. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition , pages 5141–5148, 2024. [27] Runtian Y uan, Qingqiu Li, Junlin Hou, Jilan Xu, Y uejie Zhang, Rui Feng, and Hao Chen. Multi-source covid-19 detection via variance risk extrapolation. In Pr oceedings of the IEEE/CVF International Conference on Computer V i- sion , pages 7304–7311, 2025. [28] Hongyi Zhang, Moustapha Cisse, Y ann N Dauphin, and David Lopez-Paz. mixup: Beyond empirical risk minimiza- tion. arXiv pr eprint arXiv:1710.09412 , 2017. 1 7
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment