FairMed-XGB: A Bayesian-Optimised Multi-Metric Framework with Explainability for Demographic Equity in Critical Healthcare Data
Machine learning models deployed in critical care settings exhibit demographic biases, particularly gender disparities, that undermine clinical trust and equitable treatment. This paper introduces FairMed-XGB, a novel framework that systematically de…
Authors: Mitul Goswami, Romit Chatterjee, Arif Ahmed Sekh
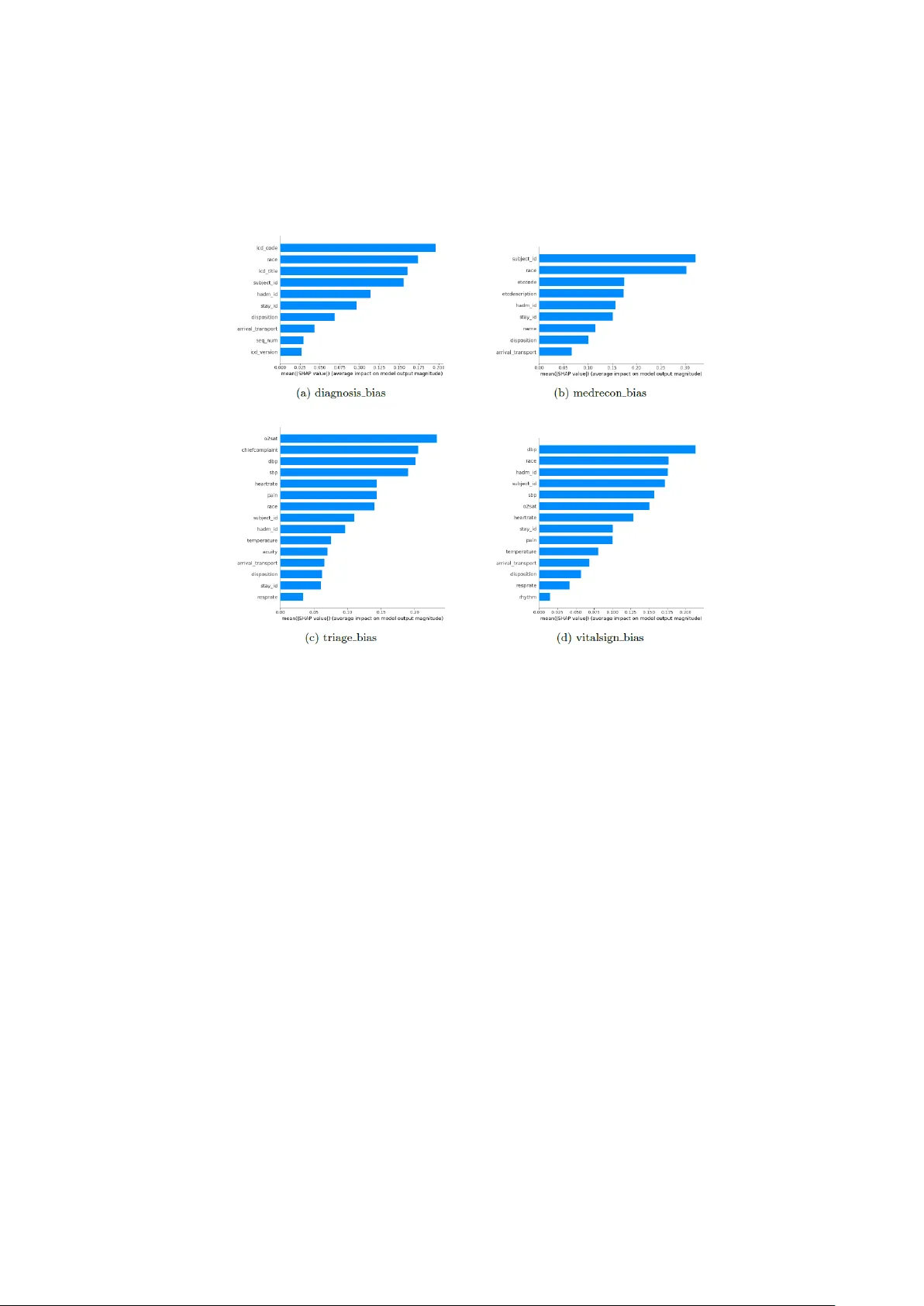
FairMed - XGB: A Bayesian-Optimised Multi-Metric Frame work with Exp lainability for Demographic Equ ity in Critica l Healthcare Data Mitul Goswami 1 , Romit Chatter jee 1 and Arif Ahmed Sekh 2 1 School of Computer Engineering, Kalinga Institute of Industrial Technology, 751024, Bh uba- neswar, India 2 Bio-AI Lab, UiT The Arctic University of Norway, Tromso, Norway Abstract. Machine learning m odels deployed i n critical care settings exhibit demographic biases, particularly gender disparities, that undermine clinical trust and equitable treatment. This paper introduces FairMed -XGB, a novel frame- work that systematically detects and mitigates ge nder -based prediction bias while p reserving model performance and transparency. The framework inte- grates a fairness-aware loss function combining Statistical Parity Difference, Theil Index, and Wasserstein Distance, join tly op timised via Bayesian search into an XGBoost classifier. Post-mitigation evaluation on seven clinically dis- tinct cohorts d erived from the MIMIC- IV -ED and eICU databases demonstrates substantial bias reduction: S tatistical Parity Difference decreases b y 40 – 51% on MIMIC- IV -ED and 10 – 19% on eICU; Th eil Index collapses by four to five or- ders o f magnitude to near-zero values; Wasserstein Distance is reduced by 20 – 72%. Th ese gains are achieved with n egligible degradation in p redictive accu- racy (AUC-ROC drop <0.02). SHAP-based explainability reve als that the framework diminishes reliance on gender -proxy features, providing clinicians with actionable insights into how and wh ere bias is correc ted. FairMed -XGB offers a ro bust, interpretable, and ethically aligned solution for equitable clini- cal decision-making, p aving the way for trustworthy deployment of AI in h igh - stakes healthcare environments. Keywords: Fair Machine Learning, Gender Bias Mitigation, Bayesian Optimi- sation, XGBoost, SHAP, Critical Care, MIMIC-IV, eICU, Algorithmic Fair- ness, Explainable AI. 1 Introduction Machine learn ing (ML) has emerg ed as a transform ative force in health care, particu- larly in high-stakes critical care settings such as intensive care units (ICUs) and emer- gency departments. Advance d predictiv e mod els are increasing ly dep loyed to forec ast patient outco mes, optimise resource allocatio n, and guide treatment protocols, lever - aging vast datasets to enhance clinical decision -m aking [1][2 ]. For instan ce, algo- rithms trained on electronic health recor ds ( EHRs) can predict sepsis onset, mortality risk, or rea dmission likelihood with r emarkable ac curacy, enabling p roactive interven- 2 tions th at save lives and reduce costs [3]. These models hold imm ense p otential t o streamline workflows, minimise human error, and improve equity in ca re delivery. However, this gr owing reliance on d ata -driven systems und erscores a pressing eth ical challenge: biases embedd ed in training data or alg orithmic design can p erpetuate or exacerbate d isparities among v ulnerable popu lations. Studies revea l that demogr aphic imbalances, such as the underrepresen tation of female patients in cardio vascular da- tasets or r acial dispar ities in pain assessment algorithm s, often lead to skewe d predic- tions, misdiagnoses, o r inequitable resou rce distribu tion [4][5]. In cr itical car e, where timely and unbiased d ecisions are par amount, such flaws risk d eepening existing healthcare in equities, disproportion ately affecting marg inalised groups. Thus, en sur- ing fairness in ML -driven clinical too ls is not merely a technical concern but a moral imperative to uphold equity and trust in m odern healthcare systems. The ef ficacy o f machine learning models i n healthcar e hinges on the representa- tiveness and quality of training data, yet pervasive demographic biases in clinical datasets often underm ine their fairness. Biases arise from systemic ineq uities in data collection, such as the under representation of minority gr oups, imbalan ced outcomes across demo graphics, or incomplete do cumentation of socioeconomic f actors [6, 7]. For instance, models train ed on ICU datasets with skewed gender ratios may system- atically misestimate mo rtality r isks for fem ale patients, while alg orithms lever aging racially homog eneous data might inaccurately triage patients from u nderrepresented backgrounds [8]. These biases p ropagate th rough the predictive pipeline, amplifying disparities in cr itical o utcomes. A 2022 study of eI CU data revea led that m odels pr e- dicting sepsis exhibited significantly lower recall for Black patients compared to White p atients, exacerb ating d elays in life -saving interven tions [9 ]. Similarly, socio- economic b iases em bedded in triage algor ithms have been shown to disproportionate- ly divert resources away from low -income populations, compou nding existing access barriers [10]. Such failures not only erode tru st in clinical AI but also risk perpetuat- ing cy cles of inequity, where marginalised groups fac e compou nded disad vantages in acute care settings. Ad dressing these challenges requires frameworks that explicitly identify and mitigate data-driven biases while preserving model utility — a gap this work seeks to bridge. Existing bias mitigation techniques, such as reweighting an d adversarial d ebiasing, often prioritise isolated fairness metr ics, such as statist ical parity , while neglecting the multifaceted nature of d emographic equity in healthca re [11, 1 2]. For example, re - weighting m ethods may b alance dataset rep resentation across gr oups bu t fail to ad - dress disparities in m odel performan ce metrics such as false -negative rates, wh ich are critical in life-thr eatening conditions. Similarly, adversarial ap proaches that suppre ss sensitive attrib ute co rrelations risk degrading predictive accuracy or ignoring intersec- tional biases arising from overlapping demographic factors (e.g., race and gender) [13]. Furthermor e, current frameworks seldo m harmonise competing fair ness objec- tives, such as equalised o dds (equality in error rates) and distribu tional fairness (alignment o f prediction distributions acro ss g roups), lead ing to suboptimal trad e -offs in clinical settings [14]. Compo unding this issue is the lack of explain ability in fair - 3 ness-aware mod els: many m ethods o perate as "b lack boxes," offering clinicians lim- ited insight into how bias mitigation impacts decision logic or feature importance [15]. This opacity hind ers trust an d ad option, as healthcare pro viders r equire tran spar- ent tools to v alidate model behavio ur against ethical an d clinical standards. This work intro duces FairMed -XGB, a nov el framework designed to advance de- mographic equity in critical healthca re prediction models through three synergistic innovations. First, it em ploys Bayesian-op timised multi-metr ic f airness, dynamically balancing equalised odds, Theil index, an d Wasserstein distance to harmonise accura- cy with subgr oup equity. Unlike prior method s that fix fairness constraints statically, this ap proach ad aptively tunes hyperpar ameters to optimize trade -offs across heter o- geneous ICU datasets, su ch as eICU and MIMIC -IV ED. Second, the framework pri- oritizes demo graphic equity by explicitly mitigatin g gender -based dispar ities in clini- cal outcomes — a critical gap in ex isting critical care models — through rigorou s evalu - ation of pr ediction d istributions and error rates across subgroups. Third, FairMed – XGB integrates SHAP to provide gran ular, clinician -actionable in sights into feature contributions, bridging th e explainability gap in fairness -aware ML. By linking SHAP values to bias, the f ramework ensures transpar ency in b oth model decisions and miti- gation strategies. 2 Related Works 2.1 Medical Tabular Da ta for Machine and Deep Learn ing Medical tabular datasets, such as MIMIC -IV E D [16] and eICU [1 7], ar e pivotal for training ML models in critical care. Th ese datasets captur e stru ctured EHRs, includ- ing demograp hics, vitals, and lab results, enablin g predictive task s like sepsis detec- tion. Howev er, inherent biases — such as underrepresen tation of minority groups or imbalanced outcomes — comprom ise mod el fairness. For instance, MI MIC -IV ED exhibits gender d isparities in cardiovascular entries, while eICU data reflects racial imbalances in sepsis prediction recall. Recent studies high light the prevalence of so- cioeconom ic and rac ial b ias v ariables in su ch datasets, n ecessitating prepro cessing or algorithmic mitigation. Table 1 summarises common datasets an d associated bias variables. Table. 1. Bias Variables in Medical Tabular Datasets Dataset Description Bias Variables MIMIC-I V [16] ICU data from Beth Israel Hosp ital Gender, Race, Ag e eICU [17] Multi-center I CU data Race, Socioeco nomic status NHANES [18] National health s urvey data Ethnicity, Income , Education UK Biobank [1 9] Large-scale UK b iomedical datab ase Ethnicity, Socio economic status All of Us [20] NIH-funded div erse cohort data Geographic div ersity, Race Framingham Hear t [21] Longitudinal car diovascular stud y Historical cohort BRFSS [22] CDC behavioral ri sk factor sur vey Self-reporting bias, Income 4 2.2 Bias Detection Method s in Medical Data Detecting biases in medical datasets requires systematic method ologies to id entify and quan tify inequities. Disp arate Im pact Analysis [23] compares f avorable outcome ratios across privileged and marg inalized groups (e.g ., ICU adm ission rates by race) to flag systemic disparities. C ausal Grap h An alysis [24] uses directed acyclic graph s (DAGs) to uncov er confounding variab les (e.g., socioeconomic status) an d isolate bias pathway s. Adv ersarial Debiasing [25] trains models with ad versarial n etworks to minimize bias during lear ning, thou gh at times it m ay re duce p redictive accurac y. Fairness Au dits [26 ] assess sub group perfor mance using metrics like False Po sitive Rate (FPR) an d False Negative Rate (FNR) to reveal disparities (e.g ., in sep sis predic- tions for Black patients). Counterfactual Testing [27] alter s sensitive attributes (e.g., gender) to ch eck prediction invariance, exposin g causal biases. Reweighting Tech- niques [28] a djust data weights (e.g., oversampling female cardiovascu lar cases) to address represen tation im balance. Lastly, Subgroup Performance Analysis [29] evalu- ates models across dem ographic splits (e.g., age, eth nicity), as sh own in MIMIC -IV studies [30]. 2.3 Bias Mitigation Method s in Medical Data Bias mitigation in m edical data emp loys techniqu es to reduce disparities in m odel outcomes across dem ographic g roups. Reweigh ting [31] a djusts sample weights to balance underrepresented cohorts, such as o versampling female patients in cardiovas- cular d atasets. Adversarial debiasing [32] trains mod els with ad versarial n etworks to minimize corr elations between pr edictions and sen sitive attributes (e.g., r ace in sepsis risk models). Fairness constraints [33] integrate f airness metrics (e.g., eq ualised odds) into the loss function, penalising d iscriminatory prediction s during training. Data augmentatio n [34], including synthetic minority oversampling (SMOT E), addresses class imbalance in rare disease datasets. Causal mitigation methods [ 35] leverage directed acyclic graphs to adjust for con founders like socioecon omic status, ensuring unbiased causal relationships in outcomes. Post-p rocessing techniques [36 ] recalibrate decision thresholds (e.g., ICU discharge prediction s) to alig n error rates acr oss sub - groups. Fair meta- learning [37] adapts mo dels to heterogeneous population s by trans- ferring fairness -aware k nowledge acro ss datasets. While these m ethods reduce bias, challenges persist in preserv ing clinical accuracy and addressing in tersectional dispar- ities (e.g., race -gender interactions). 2.4 Explainable Bias Detec tion and Mitigation Met hods Explainable m ethods enhance transparen cy in identifying and addressing b iases in medical AI. SHAP (SHap ley Additive ex Planations) [38] quantifies feature contribu- tions to pr edictions, ex posing biases in variab les like race or insurance type. LIME (Local In terpretable Mod el-agnostic Exp lanations) [39] generates local ex planations to high light b iased decision logic (e.g., triage disparities in eICU data). FairML [ 40] audits models via input perturbation to trace biases to specific features (e.g., ZIP code 5 proxies for race). Interpretable f airness co nstraints [4 1], such as equalized odds penal- ties, in tegrate fairness metrics in to model train ing while providin g dec ision - boundary transparency . Coun terfactual exp lanations [ 42] test prediction invariance under hy po- thetical chan ges to sensitive a ttributes (e. g., alter ing gen der in mortality risk models), isolating causal bias pathway s. Rule- based models [4 3], like fairness -aware d ecision trees, enforce interpretable fairness rules (e.g. , equitable ICU discharge cr iteria). 3 Methodology This section details the FairMed - XGB framework, a systematic approach de- signed to detect and mitigate gender -based predictio n bias in clin ical machine learn- ing models while preserv ing predictive p erformance and explainability. Th e method- ology compr ises fo ur core stages: (1) d ata preproce ssing, (2) pre -mitigation bias de- tection and an alysis, (3) the formulation and integration of a fairness -aware loss func- tion, and (4) post-mitigation training and ev aluation. The mathematical fo rmulation of the framewo rk is presented in Algorithm 1 . Algorithm 1 : FairMed – XGB: Bayesian Optimised Mu lti-Metric Fairn ess Learning INPUT: Clinical dataset where feature v ector binary clin ical label sensitive attribute (gender) OUTPUT: Fairness-aware mo del Data Prepro cessing Encode ca tegorical features: Standardise co ntinuous features: Split into and (stratified on ) Baseline Training Initialise model: to Compute pr edictions: Compute gr adient: 6 Fit regression tree to Update mod el: Bias Quantification Define group s: Compute Statistical Parity Dif feren ce: Compute Theil Index: Compute Wasserstein Di stance: Fairness Aware Object ive Define fairn ess loss: Define total loss: Bayesian Optimiza tion Initialize search spac e: Train model min imizing Evaluate valid ation objective: Update using Gaussian Process surro gate Select optimal Final Training Retrain model o n using Obtain final m odel Explainability each featur e Compute v alue Compute group disparity: 7 3.1 Data Prepro cessing We utilise two p ublicly availab le critical -care datasets: MIMIC- IV - ED [16] and the eICU Collaborative Rese arch Database [17] . To ensu re con sistency and focu s, gender is treated as a binary sensitive attribute , where de- notes Male an d d enotes Female . All records with undefined or non -binary gender values are excluded from the analysis. Categorical features are encod ed using label encoding, and continuous features are normalised. Each dataset is p artitioned into train ing and testing subsets using an 80/20 stratified split, preserv ing the original distribution of th e sensitive attribute and tar get classes. Formally , let rep- resent th e fea ture matrix, the binary outcome labels, and the protected attr ibute vector for samples. 3.2 Pre-m itigation Bias Detection A standard XGBoost classifie r is initially trained on the preprocessed data using th e binary logistic lo ss in equation (1), where is the predicted prob- ability, for instance . (1) 3.2.1 Model In terpretation via SHAP To diagnose the sour ce and magnitude of po tential bias, we em ploy SHAP ( Shapley Additive exPlanations) [38]. For a g iven prediction, SHA P attributes a contribution value to each feature based on cooperative gam e theory. We use the Tree SHAP [44] varian t fo r eff icient computation with tree-b ased m odels. Summary plo ts (e.g. , beeswarm plots) are generated to rank features by their average ab solute SHAP value ( ), providing an interpretab le view of the model's d ecision logic and identify ing features that may act as proxies for gend er. 3.2.2 Quantifica tion of Bias We quan tify the baseline d emographic disparity using three complemen tary fairness metrics: • Statistical Parity Differen ce (SPD): Me asures the differen ce in positive pre- diction rates b etween groups using equatio n ( 2 ). (2) • Theil In dex: An information-theoretic measure of in equality in the distribu- tion of predicted outcomes calculated u sing equation ( 3) 8 (3) • Wasserstein Distance ( W): Qu antifies th e distan ce between the predicted probability distribu tions of the two demo graphic groups using equation ( 4) (4) where and are the c umulative distribution functions for groups an d , respectively. 3.3 Fairness-Aware Los s Function To mitigate the identifie d biases, we formulate a custom loss function that aug- ments the standard predictive loss with a fair ness penalty. Th e total loss is d e- fined using equation (5) (5) Here, is th e b inary cross-en tropy loss, is a regu larisation hyperparameter controlling the overall fairness penalty stren gth, and are weights balancing the contribution of each fairness metric (SPD, Theil Index, Wasserstein Distance). 3.4 Post -mitigation Training and Evalua tion The fairness-awar e lo ss is integrated in to a custom XGBoost training loop. The hy- perparam eters are optimised using Bayesian optimisation with cross- validation to find the optimal trade -off between pred ictive accuracy and f airness. The final model is evalu ated o n the held -out test set. We compare the post-mitigatio n fair - ness metr ics (SPD, Theil Index , Wasserstein Distance) against their pre -mitigatio n values to quan tify bias reduction . SHAP analysis i s rep eated o n the debiased mod el to demonstrate the shift in feature imp ortance an d the reductio n of relian ce on gender - proxy featur es, thereby providing exp lainable insights into the mitigatio n process. 4. Experimenta l Setup, Results, and Discu ssion This section p resents the em pirical ev aluation of th e FairMed - XGB framework. We detail the experimental setup, report quantitative and qualitative results on bias miti- gation, and provide a comprehensive discussion of the findings, their implications, and future d irections. 9 4.1 Experiment al Setup 4.1.1 Datasets an d Cohort Derivation The fram ework was evalu ated on two lar ge- scale, publicly av ailable critical ca re databases: • MIMIC- IV -ED [16]: Contains app roximately 280,000 emergency depart- ment visits from 2 011 -2019, includin g demographics, vital sign s, laboratory results, triage scores, diagnoses, and p atient outcomes. • eICU Collaborative Research Database [17]: Contains data from over 200,000 ICU stays across th e U.S. (20 14 -2015), with g ranular clinical meas- urements, inter ventions, and outcomes. To assess m odel behav iour across dif ferent clinical contexts, we deriv ed seven dis- tinct cohorts. Fro m MIMIC- IV -ED: , , , and . From eICU: , , and . All cohorts were filtered to include on ly adult patients (ag e ≥ 18) with a v alid binary gender label. Categorical variables wer e label -encoded , and continuous var iables were normalised. 4.1.2 Implemen tation and Evaluation Protocol Each co hort was split into training (80%) and test (20%) sets u sing stratified sampling on the gen der attr ibute to preserv e distribution. Th e base classifier wa s XGBoost, and all exper iments wer e conducted on an NVI DIA RTX 3050 GPU. The f airness -aware loss fun ction (Section 3.3) was integ rated into the training loop. T he hyperparame- ters — regularisation factor and fairness metric weights — were optimised using Bayesian Optimisation with 5-fo ld cross-valid ation, max imising a combined objective of accuracy and fairness. Model per formance was evaluated using : • Fairness Metrics: Statistical P arity Differen ce (SPD), Theil Ind ex, and Was- serstein Distance (W) on the test set. • Explainability: SHAP (TreeSHAP) an alysis was performed pre - and post- mitigation to in terpret feature contributions. • Predictive Per formance: Overall accu racy and AUC -ROC wer e mon itored to assess the fairness -utility trade -off. 10 4.2 Results 4.2.1 Pre-mitiga tion Baseline Disparities The stan dard XGBoost mo del ex hibited significant gender -based predictio n d ispar- ities across all cohor ts. Table . 2 summarises the b aseline fairness metrics. • MIMIC- IV -ED Cohorts: Th e cohor t sh owed the most se- vere initial bias, with an SPD of 0.63 and a Th eil Index of ~4×10 ⁴, indicating a highly skewed pred iction distribution. The and cohorts also showed substantial imbalances (SPD = 0 .37 and 0.52, respectively). • eICU Cohor ts: Bias wa s ev en mo re pr onounced. All three cohor ts exhibited extreme SPD values (~0 .98- 0.99) and Theil Indices on the order of 10⁵, con- firming a strong sy stematic bias toward on e gender group in the baseline model. These results q uantitatively confirm th at mod els train ed on imbalan ced clinical data without f airness constraints can perpetuate and amplify ex isting demogr aphic dispari- ties. 4.2.2 Efficacy o f the Fairness-Awa re Mitigation Applying the FairMed - XGB f ramework with the Bay esian-o ptimised fairness loss significantly redu ced bias across all datasets. Tab le 2 presents a comparative sum- mary of key fairn ess metrics before and af ter mitigation. Table 2. Summary of Fairness Metrics Pre-Mitigation and Post-Mitigation for Representative Cohorts Cohorts Female Predicted % Male Predicted % SPD Theil Wasserstein Female Predicted % Male Predicted % SPD Theil Wasserstein Pre – Mitigation St age Pre – Mitigati on Stage MIMIC – IV ED diagnosis 59.36 40.63 0.37 6444.92 0.16 49.71 50.28 0.24 0.05 0.07 medrecon 62.77 37.22 0.63 40344.23 0.32 63.71 36.29 0.31 0.28 0.11 triage 59.81 40.18 0.24 3778.09 0.12 52.97 47.02 0.20 0.05 0.071 vitalsign 56.45 43.54 0.52 13589.49 0.23 52.87 47.12 0.21 0.06 0.07 e - ICU care 45.13 54.86 0.99 181473.06 0.95 44.86 55.13 0.80 0.53 0.65 diagnosis 43.54 56.45 0.98 163423.39 0.96 43.32 56.67 0.87 0.62 0.75 treatment 43.45 56.54 0.99 226915.1 0.97 43.56 56.73 0.89 0.64 0.76 • Bias Reduction: Post-mitigati on, SPD decreased by 40 -51% for MI MIC- IV - ED and 10 - 19% for eICU cohorts. T he most d ramatic improvem ent was in the Theil Index, which collapsed b y 4-5 o rders of magnitude to near-zero 11 values (~0.06-0 .65), indicating achievement of near -perfect distributio nal parity. Wasserstein Dis tance was also reduced by approximately 20 -72%, signifying a mu ch closer overlap in the pr ediction score distributions be- tween gend er groups. • Bias Reduction: Post-m itigation, SPD decreased by 40 -51% for MIMIC- IV - ED and 10 -19% for eICU co horts. T he most dramatic improvement was in the Theil Index, which collapsed by 4-5 orders of magnitude to near-zero values (~0.06-0 .65), indicating achievement of near -perfect distributio nal parity. Wasserstein Distance was also reduce d by approximately 2 0 -72%, signifying a mu ch closer overlap in the pr ediction score distributions be- tween gend er groups. Fig. 1: SHAP Visualizations for all the custom cohorts of eICU dataset. 4.2.3 Explainab le Insights from SHAP An alysis 12 Fig. 2: SHAP Visualizations for all the custom cohorts of MIMIC- IV -ED database. SHAP analysis provid ed crucial, interpretable evidence of how bias was manifest- ed and subsequ ently mitigated (Fig. 1 & 2 ). • Pre-mitigation: SHAP summary plots for the b aseline models identified spe- cific clinical and administrative features (e.g., specific diagn ostic codes, ag e, triage score, max imum h eart rate) as top contributor s. Th ese f ea- tures often ser ved as strong proxies for gender, driving disparate p redictions. • Post-mitigation: After applying FairMed - XGB, the SHAP value d istribu- tions for these pr oxy features becam e markedly more un iform acro ss gen der groups. The overall ranking of feature importan ce shifted , reducin g the mod- el's reliance on spuriou s gender -co rrelated signals an d promoting a more bal- anced use of clin ically relevant features fo r prediction. 13 4.3 Discussion 4.3.1 Interpretatio n of Findings The results validate the co re hypothesis of th is wo rk: a multi -metric, penalty -based approach gu ided by Bayesian optimisation can effectively mitigate gender bias in clinical pr ediction models. The con current op timisation of SPD (parity), Theil (distri- butional eq uality), and Wasserstein Distan ce (score distribu tion align ment) addresses bias from complementary angles, leading to robust de -biasing. The preservation of accuracy confirms that fair ness and utility ar e not mutually exclusive but can b e joint- ly optimised with a carefully designed objective. The SHAP visualisatio ns tr ansform the m itigation p rocess from a "black b ox" into an auditable, transparent o ne. Clinicians and data stewards can see n ot just that bias was red uced, but how — by observing wh ich feature contributions were recalibrated. This is vital for b uilding trust and facilitating model audits in regulated healthcare environments. 4.3.2 Implicatio ns for Clinical AI The pervasive b aseline biases uncovered in stan dard models und erscore a signifi- cant risk in deplo ying "off - the-shelf" ML in healthcar e. Frameworks like FairMed - XGB ar e essen tial fo r dev eloping tru stworthy AI that aligns with ethical principles of equity. By provid ing bo th quantitative fairness guar antees an d ex plainable insights, such framewo rks enable: • Regulatory Compliance: Me eting em erging s tandard s for algorithmic fair - ness in medical d evices. • Clinical Ad option: Equipping healthcare providers with understand able tools, increasing their confidence in AI -assisted decision-making . • Health Equity: Actively con tributing to reducing outco me dispar ities b y en- suring models per form equitably acr oss demographic groups. 4.3.3 Limitation s and Future Work This study has several limitations that p oint to productive future research direc - tions: • Binary Gend er Framewor k: The cur rent work ad dresses gender as a binar y construct. Future work mu st expan d to multi-class and non-bin ary sen sitive attributes an d address in tersectional fairn ess (e.g., considering interac tions between gender , race, and socioecon omic status). • Task Gen eralisation: While tested on multiple prediction tasks, ap plying FairMed - XGB to a wider r ange o f clinical outcomes (e.g., length of stay, specific comp lication risks) is necessary. 14 • Real-time Deplo yment: Developing mech anisms for continuous o r real- time fairness mo nitoring in live clinical decision -support system s is a crucial nex t step for operatio nal deployment. • Causal Perspectiv es: Integratin g causal inference methods more deeply co uld help distinguish between unf air statistical associatio ns and legitimate clinical correlates of g ender, leading to mor e nuanced mitigation. 5. Conclusion This study pr esented FairMed - XGB, a comprehensive fram ework for achieving fairness in critical care ML mo dels. By in tegrating Bayesian -optimised multi-metric fairness p enalties with SHAP- based ex plainability, FairMed successfully reduced gender-based prediction disparities in MIMIC - IV -ED an d eICU datasets wh ile main- taining high predictive accuracy. The framework moves beyond mere bias correction to provide ac tionable transparency, showing how biases are m itigated. This wo rk provides a practical, ethical, and tran sparent pathway toward d eploying mor e equita- ble and trustwor thy machin e learning models in high - stakes healthca re settings, con- tributing to the broader goal of algor ithmic fairness in medicine. References 1. Olalekan Kehind e, A. (2025). Leveraging machine learning for predictive models in healthcare to enhance patient outcome management. Int Res J Mod Eng Technol Sci , 7 (1), 1465. 2. Ghimire, P ., & Adhikari, S. (2025). Advancing Smarter Healthcare Through Data Analyt- ics: A Study on the In tegration of Machine Learn ing and P redictive Models in Cli nical Decision-Making. Advances in Computational Systems, Alg orithms, and Emerging Tech- nologies , 10 (4), 21-36. 3. Desautels, T., Calvert, J., Hoffman, J., Jay, M., Kerem, Y., Sh ieh, L., ... & Das, R. (2016). Prediction of sepsis in the intensive care unit with minimal electronic health record data: a machine learning approach. JMIR medical informatics , 4 (3), e5909. 4. Awwad, Y., Fletcher, R., Frey, D., Ga ndhi, A ., Najafia n, M., & Teodore scu, M. (2020). Exploring fairness in machine learning for international development . CITE MIT D-Lab. 5. Hoffmann, D. E., & Goodman, K. E. (2022). Allocating Scarce Medical Resources During a Public Health Emergency: Can We Consider Sex?. Hous. J. Health L. & Pol'y , 22 , 175. 6. Qi, M. (2022). Tackling health inequity using machine learning fairness, AI, and optimiza- tion . Rensselaer Polytechnic Institute. 7. Andrich, A. (2025). Computational and longitudinal approaches to detecting gender bias in p olitical co mmunication (Doctoral dissertation, Dissertation, Ilmenau, TU Ilmenau, 2025). 8. Long, E. F., & M athews, K. S. (2018). The b oarding patient: effects of ICU and hospital occupancy surges on patient flow. Pro duction and op erations management , 27 (12), 2122- 2143. 15 9. Mohammed, S., Matos, J., Doutrelign e, M., Celi, L. A., & Struja, T. (2023). Racial dispari- ties in invasive ICU treatments among septic patients: high reso lution electronic health records analysis from MIMIC-IV. The Yale journal of biology and medicine , 96 (3), 293. 10. O'Brien, N., Van Dael, J., Clarke, J., Gardner, C., O'Shaughnessy, J., Darzi, A., & Ghafur, S. (2022). Addressing racial and ethnic inequities in data-driven health technologies. 11. Li m, J., Kim, Y., Kim, B., Ahn, C., Shin, J., Yang, E., & Han, S. (2023). Biasadv: Bias - adversarial augmentation for model debiasing. In Proceedings of the I EEE/CVF co nfer- ence on computer vision and pattern recognition (pp. 3832-3841). 12. Zh ang, Y., & Sang, J. (2020, October). Towards ac curacy -fairness p aradox: Adversarial example-based data augmentation for visual debiasing. In Proceedings of the 28th ACM International Conference on Multimedia (pp. 4346-4354). 13. Ch en, R. J., Wang, J. J., Williamson, D. F., Chen, T. Y., Li pkova, J., Lu, M. Y., ... & Mahmood, F. (2023). Algorithmic fairness in artificial in telligence for medicine and healthcare. Nature biomedical engineering , 7 (6), 719-742. 14. M oreno-Sánchez, P. A. , De l Ser, J., v an Gils, M., & Hernesniemi, J. (2025). A Design Framework for operationalizing Trustworthy Artificial Intelligence in Healthcare: Re- quirements, Tradeoffs and Challenges for it s Clinical Adoption. arXiv preprint arXiv:2504.19179 . 15. Greeng rass, C. J. (2025). Transf orming clinical reasoning — the r ole of AI in supporting human cognitive limitations. Frontiers in Digital Health , 7 , 1715440. 16. Jo hnson, A., Bulgarelli, L., P ollard, T., Celi, L. A., Mark, R., & Horng IV, S . (2021) . Mimic- iv -ed. PhysioNet . 17. P ollard, T. J., Johnson, A. E. , Raffa, J. D., Celi, L. A., Mark, R. G., & Badawi, O. (2018). The eICU C ollaborative Research Database, a free ly availa ble multi -center database for critical care research. Scientific data , 5 (1), 1-13. 18. F ain, J. A. (2017). NHANES: u se of a free public data set. The Diabetes Educator , 43 (2 ), 151 -151. 19. Allen , N. E., Sudlow, C., Peakman, T. , Collins, R., & Uk biobank. (2014). UK biobank da- ta: come and get it. Science translational medicine , 6 (224), 224ed4-224ed4. 20. Bio bank, Mayo Blegen Ashley L. 18 Wirkus Samantha J. 18 Wagner Victoria A. 18 Mey- er Jeffrey G. 18 Cicek Mine S . 10 18, & All of Us Research Demonstration Project Teams Choi Seung H oan 14 http://orcid. org/0000 -0002-0322-8970 Wang Xin 14 http://orcid. org/0000-0001-6042-4487 Rosenthal Elisabeth A. 15. (2024). Genomic data in the all of us research program. Nature , 627 (8003), 340-346. 21. An dersson, C. , Johnson, A. D., Benjamin, E. J., Levy, D., & Vasan, R. S. (2019). 70 -year legacy of the Framingham Heart Study. Nature Reviews Cardiology , 16 (11), 687-698. 22. Iachan, R., Pierannunzi, C., Healey, K., Greenlund, K. J., & Town, M. (2016). National weighting of data fro m the behavioral risk factor surveillance system (BRFSS). BMC med- ical research methodology , 16 (1 ), 155. 23. Tau b, N. (1995). The relevance of disparate impact analysis in reaching for gend er equali- ty. Seton Hall Const. LJ , 6 , 941. 24. Li ang, X. S. (2021). No rmalized multivariate time series causality analysis and causal graph reconstruction. Entropy , 23 (6), 679. 25. Zh ang, B. H., Lemoin e, B., & Mitchell, M. (2018, December). M itigating unwanted biases with adv ersarial learning. In Proceedings of the 2018 AAAI/ACM Conference o n AI, Eth- ics, and Society (pp. 335-340). 26. Kamalaruban, P., Pi, Y., Burrell, S., Drage, E., Skalski, P., Wong, J., & Sutton, D. (2024, November). Evaluating fairness in transaction fraud models: Fairness metrics, bias audit s, 16 and challenges. In Proceedings of th e 5th ACM International Conference on AI in Fi- nance (pp. 555-563). 27. Veitch, V., D'Amour, A., Yadlowsky, S., & Eisenstein, J. (2021). Counterfactual invari- ance to spurious co rrelations: Why and how to pass stress tests. arXiv preprint arXiv:2106.00545 . 28. Yazdi, F., & Asadi, S. (2025). Enhancing Cardiovascular Disease Diagn osis: The Power of Optimised Ensemble Learning. IEEE Access , 13 , 46747-46762. 29. M eng, C., Trinh, L. , Xu, N., Enouen , J., & Liu, Y . (2022). I nterpretability and fairness evaluation of deep learning models on MIMIC-IV dataset. Scientific Reports , 12 (1), 7166. 30. M ohammed, S., Matos, J., Doutreligne, M., Celi, L. A., & Struja, T. (2023). Racial dispari- ties in invasive ICU treatments among septic patients: high reso lution electronic health records analysis from MIMIC-IV. The Yale journal of biology and medicine , 96 (3), 293. 31. M iller, S. (2023). The Use of Data Balancing Algorithms to Correct fo r the Under - Representation of Female Patients in a Cardiovascular Dataset. 32. Yang , J., So ltan, A. A., Eyre, D. W. , Yang, Y., & Clifton, D. A. (2023 ). An adversarial training framework for mitigating algorithmic biases in c linical machine learning . NPJ digital medicine , 6 (1), 55. 33. P agano, T. P., Loureiro, R. B., Lisboa, F . V., Peixoto, R. M., Guimarães, G. A., Cruz, G. O., ... & Nascimento, E. G. (2023). Bias and u nfairness in machine learning models: a sys- tematic review on d atasets, tools, fairness metrics, and identification and mitigation meth- ods. Big data and cognitive computing , 7 (1), 15. 34. M ures, O. A., Taibo, J., Padrón, E. J., & Iglesias-Guitian, J. A. (2025). Mitigating Class Imbalance in Tabular Data through Neural Network -based Synthetic Data Generatio n: A Comprehensive Survey and Library. Authorea Preprints . 35. M ulomba, C. M., Kiketa, V. M., Kutangila, D. M., M ampuya, P. H., Mukenze, N., Kasun- zi, L. M., ... & Kasereka, S. K. (2025). Applying causal machine learnin g to spatiotem- poral data analysis: An investigation of opportunities and challenges. IEEE Access . 36. Hü ser, M. (2 021). Machine Learning App roaches for Pa tient Monitoring in the intensive care unit (Doctoral dissertation, ETH Zurich). 37. Li u, Z., Sarkani, S., & Mazzuchi, T. Fairness -Optimized Dynamic Aggregation (Foda): A Novel Approach to E quitable Federated Learning in Heterogeneous Environ- ments. Available at SSRN 5145003 . 38. Lu ndberg, S. M., & Lee, S. I. (2017). A unified approach to interpreting m odel predic- tions. Advances in neural information processing systems , 30 . 39. Rib eiro, M. T., Singh, S., & Guestrin, C. (2016, August). " Why sho uld i trust y ou?" Ex- plaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD inter- national conference on knowledge discovery and data mining (pp. 1135-1144). 40. Ad ebayo, J. A. (2016). FairML: ToolBox for diagnosing bias in p redictive mod el- ing (Doctoral dissertation, Massachusetts Institute of Technology). 41. Jo , N., Agh aei, S., Benson, J., Gomez, A., & Vayanos, P. (2023, Augu st). Learning opti- mal fair decision trees: Trade-offs b etween interpretability, fairness, and accuracy. In Proceedings of the 2023 AA AI/ACM Conference on AI , Ethics, and S ociety (pp. 181- 192). 42. M othilal, R. K., Sharma, A., & Tan, C. (2020, January). Explaining machine learning clas- sifiers through div erse counterfactual explanations. In Proceedings of the 2020 conference on fairness, accountability, and transparency (pp. 607-617). 43. Ab ualrous, R., Zouzou, H., Zgheib, R., Hasan, A., Hijazi, B., & Kermani, A. (2025). Fair- ness-Aware In telligent Reinforceme nt (FAIR): An AI-Powered Hospital Sche duling Framework. Information , 16 (12), 1039. 17 44. Bifet, A., Read, J., & Xu, C. (2022). Linear tree shap. Advan ces in Neura l Information Processing Systems , 35 , 25818-25828.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment