Echo-E$^3$Net: Efficient Endocardial Spatio-Temporal Network for Ejection Fraction Estimation
Objective To develop a robust and computationally efficient deep learning model for automated left ventricular ejection fraction (LVEF) estimation from echocardiography videos that is suitable for real-time point-of-care ultrasound (POCUS) deployment. Methods We propose Echo-E$^3$Net, an endocardial spatio-temporal network that explicitly incorporates cardiac anatomy into LVEF prediction. The model comprises a dual-phase Endocardial Border Detector (E$^2$CBD) that uses phase-specific cross attention to localize end-diastolic and end-systolic endocardial landmarks and to learn phase-aware landmark embeddings, and an Endocardial Feature Aggregator (E$^2$FA) that fuses these embeddings with global statistical descriptors of deep feature maps to refine EF regression. Training is guided by a multi-component loss inspired by Simpson’s biplane method that jointly supervises EF and landmark geometry. We evaluate Echo-E$^3$Net on the EchoNet-Dynamic dataset using RMSE and R$^2$ while reporting parameter count and GFLOPs to characterize efficiency. Results On EchoNet-Dynamic, Echo-E$^3$Net achieves an RMSE of 5.20 and an R$^2$ score of 0.82 while using only 1.55M parameters and 8.05 GFLOPs. The model operates without external pre-training, heavy data augmentation, or test-time ensembling, supporting practical real-time deployment. Conclusion By combining phase-aware endocardial landmark modeling with lightweight spatio-temporal feature aggregation, Echo-E$^3$Net improves the efficiency and robustness of automated LVEF estimation and is well-suited for scalable clinical use in POCUS settings. Code is available at https://github.com/moeinheidari7829/Echo-E3Net
💡 Research Summary
This paper introduces Echo‑E³Net, a lightweight yet accurate deep‑learning framework for automated left‑ventricular ejection fraction (LVEF) estimation from echocardiography videos, specifically targeting real‑time point‑of‑care ultrasound (POCUS) deployments. The authors argue that most existing approaches either rely heavily on dense global features or require computationally intensive segmentation pipelines, which hampers their applicability in resource‑constrained settings. To address this, Echo‑E³Net explicitly incorporates cardiac anatomy by focusing on endocardial borders at the two clinically critical phases: end‑diastole (ED) and end‑systole (ES).
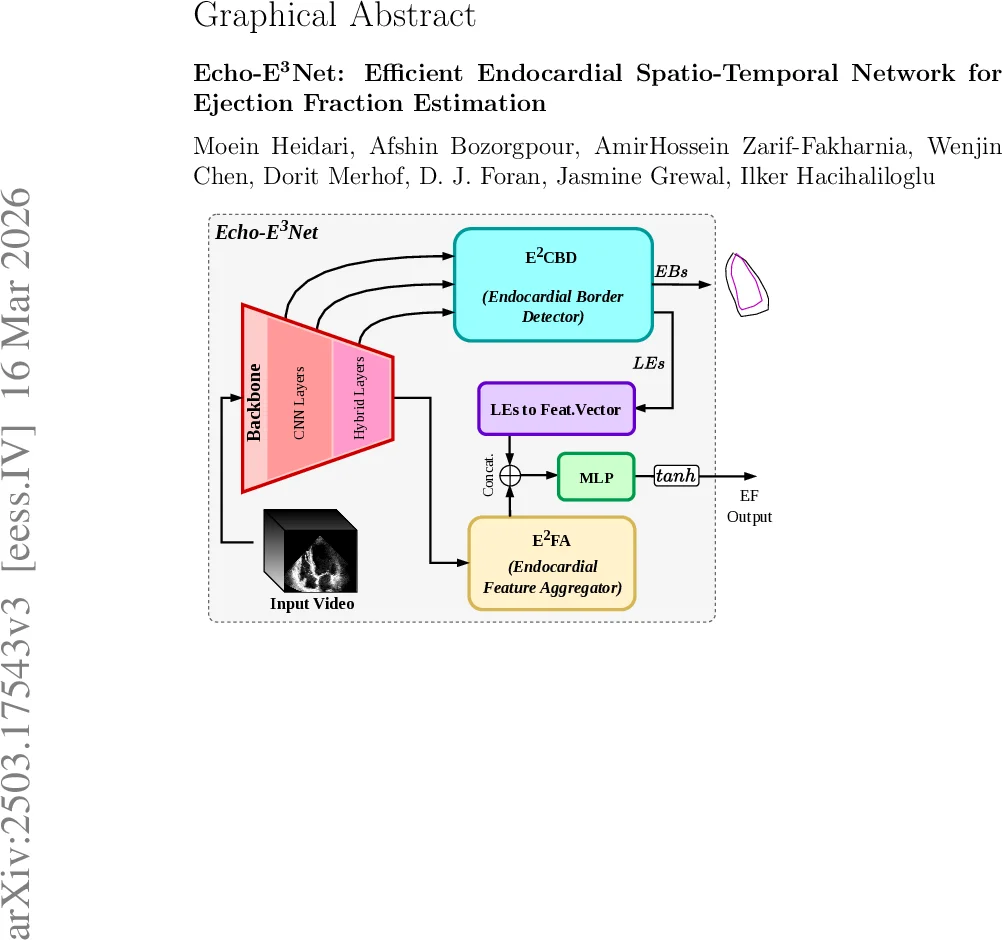
The architecture consists of three main components. First, a 3‑D encoder based on the LHUNet backbone extracts multi‑scale spatio‑temporal features from the input video. These features are tokenized, enriched with learned positional and level embeddings, and then uniformly subsampled to respect a predefined token budget, thereby limiting memory consumption. Second, the Dual‑Phase Endocardial Border Detector (E²CBD) employs phase‑specific query banks that attend to the compressed token set via multi‑head cross‑attention. Each query produces a 32‑dimensional embedding for a landmark; a small MLP followed by a gated linear unit decodes the embedding into a pair of (x, y) coordinates representing an endocardial chord. The coordinates are constrained to the normalized image grid, yielding explicit ED and ES border predictions that align with the landmarks used in Simpson’s biplane method. Third, the Endocardial Feature Aggregator (E²FA) gathers global statistics (average, max, variance) from the deepest feature map, flattens them into a vector, and concatenates this with a compressed representation of the landmark embeddings (obtained via an MLP projection). The combined descriptor is fed to a final MLP head that outputs the EF value, scaled to the 0‑100 % range.
Training is guided by a multi‑component loss. The primary term is mean‑squared error between predicted and reference EF. A landmark regression loss penalizes deviations of the predicted chords from ground‑truth annotations. Crucially, a Simpson‑inspired geometric regularizer is introduced: each chord is interpreted as a disk cross‑section; its width provides the disk diameter, while the perpendicular distance between successive chords estimates disk height. By stacking these disks, an approximate end‑diastolic volume (EDV) and end‑systolic volume (ESV) are computed, yielding a surrogate EF (f_EF). The loss enforces consistency between f_EF and the ground‑truth EF, thereby encouraging the network to learn anatomically plausible geometry without requiring explicit volume computation at inference time.
Empirical evaluation on the large EchoNet‑Dynamic dataset demonstrates that Echo‑E³Net achieves an RMSE of 5.20 % and an R² of 0.82 while using only 1.55 M parameters and 8.05 GFLOPs. This represents a >5‑fold reduction in model size and computational demand compared with prior state‑of‑the‑art methods, yet performance remains competitive. Notably, the model shows robust accuracy across the low‑EF range (≤30 %), a clinically important segment often under‑represented in training data. The authors also highlight that no external pre‑training, heavy data augmentation, or test‑time ensembling is required, simplifying deployment pipelines.
Because of its modest computational footprint, Echo‑E³Net can run at real‑time frame rates (≈30 fps) on mobile GPUs or embedded processors, making it suitable for bedside POCUS where rapid decision‑making is essential. The codebase, including training scripts and pretrained weights, is publicly released on GitHub, facilitating reproducibility and further research.
In summary, Echo‑E³Net advances the field by (1) integrating phase‑aware cross‑attention for precise endocardial landmark detection, (2) fusing these anatomical cues with lightweight global feature statistics, (3) employing a differentiable Simpson‑based geometric loss that aligns learning with clinical measurement practice, and (4) delivering a compact model capable of real‑time EF estimation on low‑resource ultrasound devices. This work paves the way for scalable, accurate cardiac function assessment in point‑of‑care environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment