On the Degrees of Freedom of Gridded Control Points in Learning-Based Medical Image Registration
Many registration problems are ill-posed in homogeneous or noisy regions, and dense voxel-wise decoders can be unnecessarily high-dimensional. A sparse control-point parameterisation provides a compact, smooth deformation representation while reducin…
Authors: Wen Yan, Qianye Yang, Yipei Wang
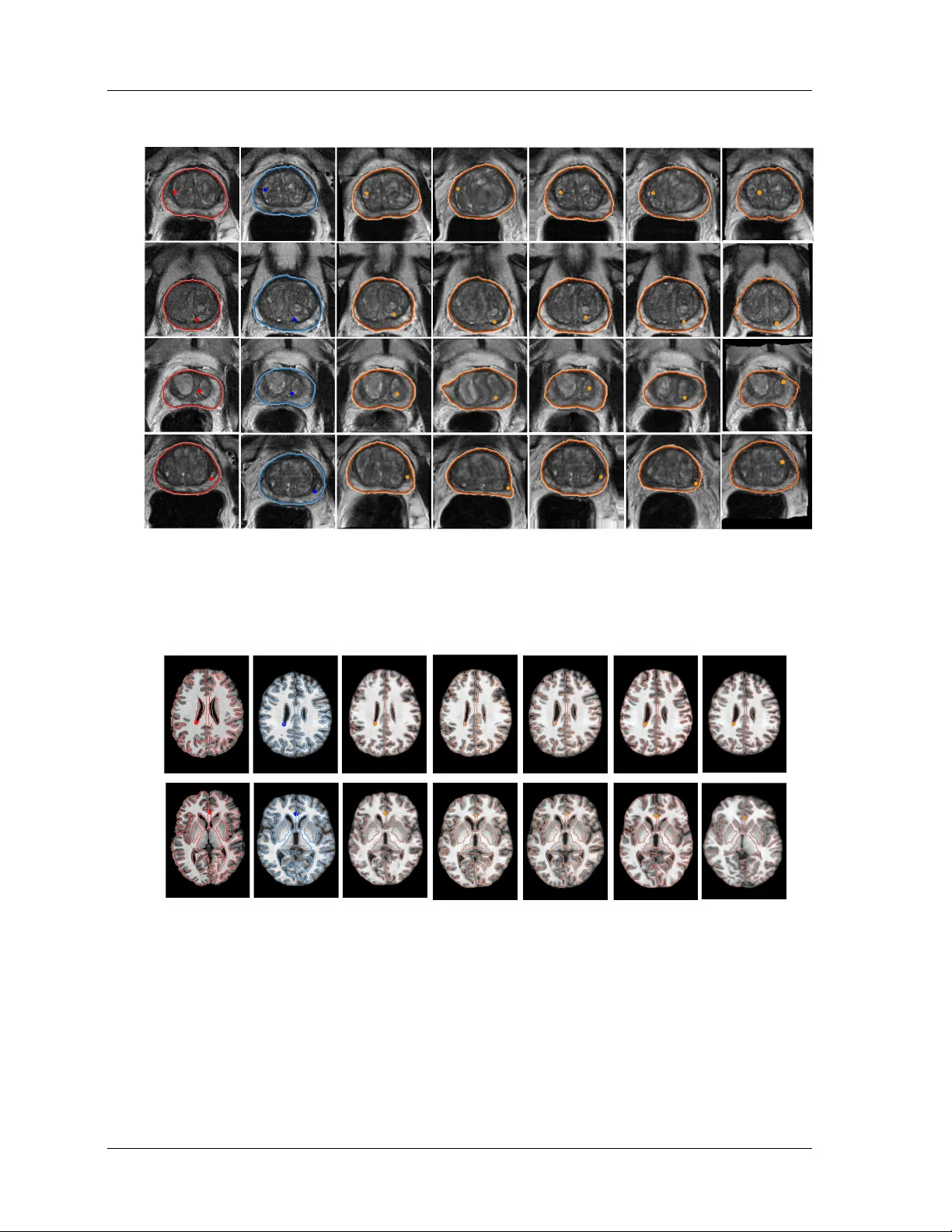
On the Degrees of F reedom of Gridded Control P oints in Lea rning-Based Medical Image Registration W en Y an 1 , Qiany e Y ang 1 , Yip ei W ang 1 , Shonit Punw ani 2 , Ma rk Emb erton 3 , V asilis Stavrinides 4 , 5 , Yip eng Hu 1 , Dean Ba rratt 1 1 UCL Ha wkes Institute; Depa rtment of Medical Physics and Biomedical Engineering, University College London, Gow er St, London WC1E 6BT, London, U.K. 2 Centre for Medical Imaging, Division of Medicine, University College London, Gow er St, London WC1E 6BT, London, U.K. 3 Division of Surgery and Interventional Science, University College London, 10 Pond St, London NW3 2PS, U.K. 4 Cancer Institute, Urology Department, UCL Hospital, University College London, 16-18 W estmoreland St, London W1G 8PH, U.K 5 Radiology Depa rtment, Imp erial College Healthca re, The Ba ys, S Wha rf Rd, London W2 1NY, U.K. corr email: w en-y an@ucl.ac.uk Abstract Bac kground: Man y registration problems are ill-p osed in homogeneous/noisy re- gions, and dense v oxel-wise decoders can b e unnecessarily high-dimensional. A sparse con trol-p oint parameterisation provides a compact, smo oth deformation represen ta- tion while reducing memory and improving stability . Purp ose: This work in vestigates the required control points for learning-based regis- tration netw ork developmen t. In particular, as sparse as 5 × 5 × 5 control p oints are congured and compared with alternative approaches, including those using scattered con trol points and displacemen ts sampled at ev ery v o xel, i.e. dense displacement elds. Metho d: W e presen t GridReg, a learning-based registration framework that replaces dense vo xel-wise deco ding with displacemen t predictions at a sparse grid of con trol p oin ts. This design substantially cuts the parameter count and memory while retain- ing registration accuracy . Multiscale 3D enco der feature maps are attened into a 1D tok en sequence with positional encoding to retain spatial con text. The mo del then predicts a sparse gridded deformation eld using a cross-attention module: Eac h con- trol point attends to enco der tokens within its lo cal grid neigh b orho o d to estimate its displacemen t, whic h is sub-sequently interpolated to a dense eld. W e further intro- duce grid-adaptiv e training, enabling an adaptiv e mo del to op erate at m ultiple grid sizes at inference without retraining. Results: This w ork quantitativ ely demonstrates the b enets of using sparse grids. Using three data sets for registering prostate gland, pelvic organs and neurological structures, the experimental results suggest a m uc h improv ed computational eciency , due to the prediction of sparse-grid-sampled displacemen ts. Alternatively , the sup erior i R unning title here: Printed Ma rch 19, 2026 registration p erformance w as obtained using the prop osed approac h, with the similiar or less compute cost, compared with existing algorithms that predict DDF s (e.g., V ox- elMorph/T ransMorph) or displacemen ts sampled on scattered k ey points (KeyMorph). Conclusion: W e conclude that predicting sparsely gridded displacemen ts pro- vides reduced computational cost and/or impro ved p erformance, independent of the enco der arc hitecture, and can b e readily implemented. Therefore, GridReg should p oten tially b e considered for man y registration tasks with adaptive grid sizes. The co de is av ailable via git@github.com:yanwenCi/GridReg.git . Keyw ords: Medical image registration; Gridded control deformation; Computa- tional Eciency I . In tro duction Medical image registration aligns images from dierent mo dalities, patients, or time p oints to the same spatial co ordinates, with many clinical utilities 1 . T raditional pairwise image reg- istration metho ds optimise parametric or non-parametric transformation mo dels iterativ ely giv en a pair of images 2 . Learning-based image registration enables a data-driven approach with fast inference to predict transformations that represen t spatial corresp ondences 3 , 4 , 5 , 6 , 7 . Examples are discussed in Sec. I I . . Moving Image Fixed Image Moving Image Fixed Image 1 2 3 4 1 2 3 4 5 Sparse grid Dense grid Figure 1: Demonstration of an illustrative example of false-p ositive corresp ondence (orange stars), p ossibly caused by similar in tensity patterns (e.g., unsup ervised loss similarity) or lab el errors (e.g., segmen tation noise). A sparse lo w-resolution grid helps lter out suc h a noise, as stronger “true” corresp ondences (four corners of the shaded ROIs) dominate. In con trast, a ner high-resolution grid may amplify the noise, distorting additional control p oin ts within the ROIs and leading to inaccurate registration. An interesting observ ation in the developmen t of learning-based image registration tech- niques is that early deep learning approac hes largely bypassed the use of gridded con trol p oin ts for mo deling deformation 3 , 4 , 5 , 6 , 8 . Instead, these metho ds commonly predicted dense displacemen t elds (DDF s), in whic h a displacemen t v ector is estimated at ev ery v oxel lo- cation. This arc hitectural choice is likely inuenced by the ev olution of computing capabili- Last edited D ate : page 2 I . INTR ODUCTION ties, particularly the adv ent of high-p erformance GPUs and parallelised training framew orks, whic h ha v e substan tially reduced the computational burden asso ciated with predicting and optimising p er-v oxel displacements. Dense displacemen t elds can b e view ed as an extreme case of grid-based mo dels where the grid resolution equals the image resolution — eectiv ely treating ev ery vo xel as a control p oint. A higher-resolution control grid, dened by a larger num b er of sampled con trol p oin ts, indeed allows the mo del to represent more lo calised, ne-scale transformations. How ev er, this increase in resolution do es not necessarily improv e registration p erformance 9 , 10 . When the transform has excessiv ely high DoF in medical imaging and similar domains, many re- gions of interest — suc h as homogeneous tissue areas or regions with strong random noise — inheren tly lack well-dened lo cal corresp ondences. Allowing the netw ork to resolve extremely lo calised deformations under dense transformation ma y b e unnecessary or ev en detrimen tal. It can introduce noise, ov ertting, or anatomically implausible corresp ondences, esp ecially if the similarity metrics used during training fail to distinguish meaningful lo cal v ariation from spurious noise. Moreo v er, increasing the grid densit y comes at a substantial compu- tational cost. This burden is particularly observ ed in the deco der, which is resp onsible for reconstructing laten t representations in to high-resolution structured deformation outputs. Suc h deco ders t ypically rely on multi-scale feature integration and upsampling mec hanisms, whic h b ecome progressively more complex and exp ensiv e as the output resolution increases 11 . Coarser con trol grids reduce the degrees of freedom, acting as an implicit regulariser that limits high-frequency , vo xel-level w arps and can reduce sensitivity to unreliable lo cal corre- sp ondences. As illustrated in Figure 1 , the c hoice of grid resolution pla ys a crucial role in con trolling the granularit y of the estimated transformation eld. A coarser grid ma y act as a form of implicit denoising or regularisation, eectiv ely ltering out high-frequency similar- it y that is not consisten tly supp orted b y the underlying anatom y or image evidence. This insigh t suggests that, depending on the application con text and anatomical region, using a coarser con trol grid can lead to more anatomically meaningful and computationally ecient deformation estimates. Our approach diers from p yramidal registration, in which do wnsampled images still yield a high-DoF dense eld via a learned high-resolution decoder 3 , 12 . Instead, w e predict a sparse con trol-p oint displacemen t eld from attened 1D feature tokens with p ositional enco ding, and then reconstruct a dense v oxel-wise displacement eld via interpolation (e.g., I. INTRODUCTION R unning title here: Printed Ma rch 19, 2026 trilinear or B-spline 13 ). Constraining DoF of the predicted transformation acts as built-in spatial regularisation, reducing ov ertting in noisy or homogeneous regions. Crucially , lo w DoF is not low expressivity , where spline interpolation can pro duce smo oth y et highly non- linear, non-rigid deformations. W e do not claim that high-resolution displacemen t elds are nev er useful; rather, their necessity is often ov erestimated. Our main contributions are: • W e in tro duce a ligh tw eight deco der that pro jects m ulti-scale 3D features in to control- p oin t embeddings and predicts sparse displacements via lo cal cross-attention. • W e prop ose a grid-adaptiv e framework that trains across multiple con trol-grid resolu- tions and selects an eective grid densit y for eac h dataset via v alidation-driv en mo del selection. • W e ac hieve comparable or improv ed registration accuracy relative to v oxel-wise DDF deco ders, while substan tially reducing memory and computation and impro ving defor- mation regularity , which is attractiv e for computation-limited settings. I I . Related W orks The degree-of-freedom (DoF) of gridded con trol p oints reects the resolution of correspon- dence in registration, discussed ab o ve. A higher DoF (dense) implies the abilit y to capture ner, lo cal corresp ondence, while a low er DoF represents coarser corresp ondence. F or scat- tered control p oin ts, corresp ondence resolution v aries spatially , dep ending on its lo cal sam- pling mec hanisms for allow ed DoF p er unit space (volume in 3D). The spatial adaptability from scattered points allows v ersatilit y and, arguably , eciency , but requires optimisation or predened with prior knowledge. In non-learning-based algorithms based on iterativ e optimisation, these tw o categories of transformation representation are closely related to free-form deformation 14 and point-feature-based metho d 15 . W e discuss example studies in the context of learning-based algorithms as follows. I I . A . Gridded con trol p oin ts Spatial T ransformation Net work (STN) 16 applies a predicted dense displacement eld through a dieren tiable sampling op erator, enabling end-to-end optimisation using image- Last edited D ate : page 4 I I . RELA TED WORKS domain losses b etw een the w arp ed source and the target 17 . Based on this, muc h work using enco der-deco der structured net works predicts the transfoThese w orks utilised unsup ervised metho ds to predict the displacement eld b etw een tw o real-world images by minimising im- age similarity loss betw een the warped source image and the target image.e w orks utilized unsup ervised metho ds to predict the displacemen t led b et ween t wo real-w orld images by minimizing image similarity loss b et ween the warped source image and the target image. In the case of complex tasks lik e m ulti-mo dality registration, rening the dense deformation eld (DDF) requires a more focused in v estigation in to training strategies. Y ang proposed a w ork 8 that b enets from exploiting additional images during training for cross-mo dalities registration. Zhou 18 prop osed a structured cross-modality laten t space to represen t 2D- pixel features and 3D features via a dieren tiable probabilistic Perspective-n-P oin ts solver. Hussain 19 prop oses a hierarchical cross-attention-based transformer structure to extract fea- tures of xed and mo ving images at multiple scales, enabling more precise alignment and impro ved registration accuracy . This structure strengthens the ability of deco der, enabling it to facilitate progressive deformation eld renement across multiple resolution lev els. I I . B . Scattered con trol p oin ts The scattered control p oin ts method can utilise manually selected anatomical landmarks, aiming to register sp ecic regions of in terest 20 , 21 . Otherwise, it can automatically detect k ey points using feature extraction metho ds. Keymorph 5 utilised a centre of mass lay er to generate corresp onding key p oin ts for source and target images, resp ectively , and pro duced a deformation eld based on these paired key points. W ang 22 extended KeyMorph as a to ol that supp orts m ulti-mo dal, pairwise, and scalable groupwise registration, solving 3D rigid, ane, and nonlinear registration in brain registration. Ekv all et al. 23 presen ted an unsu- p ervised spatial landmark detection and registration netw ork to address the c hallenges in nonlinear deformations b etw een histological tissue sections. Zac hary et al. 24 presen t a F ree P oint T ransformer (FPT) in which the scattered p oints are sampled to represent the surface of the organ via a separate segmentation pro cess for non-rigid point-set registration. F u et al. 25 generate volumetric prostate p oin t clouds from segmented prostate masks using tetra- hedron meshing and then use a p oint cloud matching netw ork to obtain the deformation eld for registration. SuperPoin t 26 op erates on full-size images and jointly computes pixel-level I I. RELA TED WORKS I I.B. Scattered control p oints R unning title here: Printed Ma rch 19, 2026 in terest p oint lo cations and asso ciated descriptors in one forward pass using homographic adaptation. Zhang et al. 27 prop osed a tw o-stage metho d for multi-model image registration. In the coarse stage, the metho d uses a key-point based metho d inherited from 26 to generate a course deformation eld from the segmen tation map, and in the ne stage, the metho d uses a vo xel-based metho d to rene the deformation eld. I I I . Metho ds I I I . A . Predicting gridded displacemen t elds Let I = { ( F j , M j ) | j = 1 , . . . , J } b e a dataset of J source–target image pairs, where eac h image is a 3D tensor F j , M j ∈ R W × H × D . Dene the vo xel domain Ω = { 1 , . . . , W } × { 1 , . . . , H } × { 1 , . . . , D } and N = | Ω | = W H D . W e write the vectorized forms as f j ∈ R N , m j ∈ R N , so that f j = [ f j ( x 1 ) , . . . , f j ( x N ) ] ⊤ with vo xel co ordinates x n = ( x n , y n , z n ) ∈ Ω . A spatial transform is represented b y a dense displacemen t eld T j : Ω → R 3 o ver the v oxel domain Ω = { 1: W } × { 1: H } × { 1: D } . W e denote warping by W ( M j , T j ) = M j ◦ T j , where ◦ denotes spatial transform op eration 16 . Learning-based image registration optimises the mo del weigh ts to nd transformations that align eac h pair of source and target images. This can b e form ulated as an optimisation problem: arg min T j P N n =0 L ( M j ◦ T j , F j ) , where and L is a registration loss function of the transformed moving image and the xed image. In this study , w e propose a registration metho d based on a sparse gridded displacemen t eld (GDF) with size G = g w × g h × g d . As illustrated in Figure 2 , w e describe a simple, easy-to-implemen t registration netw ork, GridReg, denoted by g θ ( · ) , with net w ork parameter θ = { θ 0 , θ 1 , θ 2 } , g θ ( F j , M j ) = g θ 2 prd g θ ( l ) 1 pro j ( E ( l ) j ) L l =1 , { E ( l ) j } L l =1 = g θ 0 enc ( F j , M j ) , GridReg can b e viewed as three sub-mo dules: (1) g θ 0 enc ( · ) extract feature E ( l ) j ∈ R C l × D l × H l × W l , (2) m ultiple pro jection lay ers g θ ( l ) 1 proj ( · ) , whic h pro ject 3D features at l -th lay er of the enco der in to compact 1D vectors, and g θ 2 dec ( · ) is used to aggregate features from enco der and pro jector and pro duce the gridded displacement µ j ∈ R 3 × g w × g h × g d , as shown in Figure 2 . In detail, eac h la y er in the enco der consists of a 3D conv olutional la yer with ReLU activ ation and do wnsampling b y a stride of 2. W e set a base C c hannels and doubled the Last edited D ate : page 6 I I I . METHODS Figure 2: Illustration of GridReg. The enco der extracts m ulti-scale 3D feature maps; at eac h scale, a skip-pro jection attens features in to 1D tok ens that are fused with grid-cell queries via (local) atten tion. The resulting features are mapp ed to a sparse con trol-p oint displacemen t eld via Bay esian in tegration, and a dense displacement eld (DDF) is obtained b y in terp olation (e.g., trilinear, B-spline, or transp osed conv olution). n umber of c hannels for each subsequent lay er in the enco der. Then, the features at l − th lay er from the enco der are passed through a projection la yer, whic h includes 3D con v olutional la y ers with xed 8C output c hannels and attening op eration, resulting in a vector at l -th la yer: P ( l ) j = g θ ( l ) 1 proj ( E j ( l ) ) ∈ R N p × C p with a length of N p = p w · p h · p d . Instead of using a symmetric decoder to reconstruct the output, as in previous w ork 3 , we use a 1D cross-atten tion predictor g θ 2 dec ( · ) to progressiv ely rene the deco der features, from bottom to top in Figure 2 . The deco der at l -th la yer is expressed as: Y − ( l − 1) j = g θ ( l ) 2 dec ( ˆ Y − ( l ) j ) , where ˆ Y − ( l ) j = [( P j ( l ) ) , ( Y − ( l ) j )] ∈ R N p × ( C p + C − ( l ) d ) , where C − ( l ) d is deco der c hannel n umber at l -th la yer. Let G = g w g h g d b e the num b er of control p oin ts (queries) and control p oints co ordinates stac ked as R ∈ R G × 3 and p ositional enco ding ψ ( · ) pro duce ψ ( R ) ∈ R G × C pe , where C pe is the p osition embedding channels. Giv en a multi-atten tion head and dimension in eac h head as H and d , dene learned pro jections: W Q ∈ R C pe × H d , W K , W V ∈ R ( C p + C − ( l ) d )) × H d , W O ∈ I I I. METHODS I I I.A. Predicting gridded displacement elds R unning title here: Printed Ma rch 19, 2026 I I I . A . Predicting gridded displacement elds R H d × C − ( l ) out . Queries, keys, and v alues are formed b y matrix multiplication: Q = ψ ( R ) W Q ∈ R G × H d , K = ˆ Y W K ∈ R N × H d , V = ˆ Y W V ∈ R N × H d . Reshap e in to H heads: Q → Q ( h ) ∈ R G × d , K → K ( h ) ∈ R N p × d , V → V ( h ) ∈ R N p × d , h = 1 , . . . , H . The pro do ct-atten tion is computed as: A ( h ) = softmax Q ( h ) ( K ( h ) ) ⊤ √ d ∈ R G × N p , Y ( h ) = A ( h ) V ( h ) ∈ R G × d , (1) where softmax( · ) is applied row-wise ov er the N p tok ens. Concatenating heads and pro- jecting giv es the deco der output: Y = Concat Y (1) , . . . , Y ( H ) W O ∈ R G × C − ( l ) out , whic h is reshap ed to C − ( l ) out × g w × g h × g d at last lay er of deco der. Registration mo dels with adaptiv e grid sizes T o a void training one net work p er con trol–p oint grid, w e implement a single model that adapts to multiple grid sizes during training and selects an optimised grid size on the v alidation set. W e mo died ho w the query p ositional enco dings (PEs) are generated as describ ed ab o ve to adapt the atten tion-based deco der to dynamic grid sizes. In addition, the dynamic grid implementation calculates and cac hes the PEs for use at run time, enabling exibilit y in target grid size. In summary , the enco der is shared across all settings. The PE width C pe sets the em b edding feature length of each query . In deco der, the grid size ( g w , g h , g d ) changes only the n umber of queries G = g w g h g d . According to ( I I I . A . ), the pro jection weigh ts W Q , W K , W V , W O and head sizes ( H , d ) are indep enden t of ( g w , g h , g d ) . Th us, the mo del can adapt to dierent grid sizes without additional computational cost. T o support a dynamic grid, we treat the grid resolution as a discrete hyperparameter and, during training, sample ( g w , g h , g d ) ∼ U { (5 , 5 , 5) , (8 , 8 , 8) , (10 , 10 , 10) , (15 , 15 , 15) } , . The sampled grid is em b edded b y a p osition em b edding ϕ ( g w , g h , g d ) . The cross atten tion mo dule tak es a p osition embedding as the query and produces deformation parameters µ and σ on a exible, dynamic grid. This approach oers computational eciency comparable to xed-grid mo dels while signicantly reducing storage and deploymen t ov erhead. During inference, w e select the optimal, xed grid size based on v alidation metrics for each Dataset. Last edited D ate : I I I.A. Predicting gridded displacement elds page 8 I I I . METHODS I I I . B . T ransformation sampling Ba yesian grid transformer W e adopted a previously prop osed probabilistic registration algorithm 12 , 28 and learn a grid-based p osterior ov er deformations, to estimate uncertaint y in medical image registration. This allows the metho d to assess uncertaint y for deformation as w ell as impro ve the p erformance in 12 , 28 . Th us, we add a Ba y esian head after the decoder, whic h predicts a mean and v ariance p er control p oin t, µ j , η j ∈ R 3 × g w × g h × g d , σ 2 j = softplus( η j ) , for eac h pair j . During training, w e use the reparameterization tric k to dra w S Monte–Carlo samples at the grid, T ( s ) j = µ j + σ j ⊙ ϵ ( s ) , ϵ ( s ) ∼ N ( 0 , I ) , s = 1 , . . . , S, W e optimise the similarit y o bjective with an uncertaint y-w eighting: L uncert = 1 S S X s =1 X x ∈ Ω ℓ sim ( m j ◦ T ( s ) j )( x ) , f j ( x ) 2 σ 2 j ( x ) + λ 0 X x ∈ Ω log σ 2 j ( x ) , (2) σ 2 j ( x ) is the spatially v arying uncertain ty , and λ 0 con trols the uncertain ty p enalty . Intu- itiv ely , σ j gro ws in am biguous (e.g. homogeneous or artefact–corrupted) regions and reduces where corresp ondences are clear. This pathwise sampling provides an un biased Monte Carlo estimate of the exp ectation, discouraging sharp, unstable deformations in homogeneous ar- eas. In inference, we use only the mean eld to warp the moving image, T µ,j ≡ ( µ j ) , while σ j is used solely for uncertain ty visualisation. T ransformation in terp olation Given a predicted GDF T ( x ) ∈ R 3 × g w × g h × g d , three interpo- lation metho ds are in vestigated to upsample the sparse GDF to image size, obtaining T ↑ ( x ) , where ↑ represen ts upsampling op eration. T o interpolate at each vo xel x n = ( x i , y ν , z k ) , the lo cations of GDF are represen ted as x ij k . W e describ e three upsampling strategies that can consisten tly b e considered by using a basis function to conv olve with the GDF, denoted as T ↑ ( x n ) = l X i =1 l X ν =1 l X k =1 T ( x iν k ) B x ( x i ) B y ( y ν ) B z ( z k ) (3) , where the terms B x ( x i ) , B y ( x i ) , and B z ( x i ) represent the basis functions in three dimen- sions. These basis functions share the same functional form across all three dimensions ( x , y , I I I. METHODS I I I.B. T ransfo rmation sampling R unning title here: Printed Ma rch 19, 2026 I I I . C . Loss function and z ). F or clarit y , let us consider B x ( x i ) as an example to illustrate the three interpolation metho ds: (1) T rilinear in terp olation: The basis function in trilinear interpolation is dened as B x ( x i ) = 1 − | x − x i | along 1st dimension (similarly for the other t wo dimensions). (2) Cubic B-spline interpolation: The base function of cubic B-spline interpolation is dened recursiv ely from 0-degree to 3-degree, i.e., the 1st dimension basis function at 0- degree is dened as: B 0 i ( x ) = ( 1 if t i ≤ x < t i +1 0 otherwise , where t 0 < · · · < t i < · · · < t I is a linearly sampled knots v ector t = [0 , 0 , 0 , t 0 , ..., t i , ..., t I , 1 , 1 , 1] . Then, basis functions at p = { 1 , 2 , 3 } degree are dened recursively by: B p x ( x i ) = x − t i t i + p − t i B p − 1 x ( x i ) + t i + p +1 − x t i + p +1 − t i +1 B p − 1 x ( x i +1 ) (similarly for the other t wo dimensions). (3) T ransp osed con volution upsampling: The 3D transp osed con volutions were used to implemen t a Gaussian spline-based transformation resampling. In this case, the basis function of 1st dimension is dened as: B x ( x i ) = 1 √ 2 π σ 2 e − ( x − x i ) 2 2 σ 2 , where σ is the standard deviation con trolling the width of the k ernel, which can also b e empirically congured to eciently approximate the B-spline interpolation, should it b e b enecial. All implementations for the trilinear, B-spline in terp olation and its approximation are dieren tiable during bac kpropagation, whic h allows the net work to learn the optimal in terp olation metho d for the task at hand 29 . I I I . C . Loss function W e dene image-level and grid-lev el similarit y as the displacemen t eld sampled at full resolution and a sparse, low er resolution using mean squared error, denoted as L imag e sim , where L img sim = 1 | Ω | X x ∈ Ω sim ( m j ◦ T ↑ j )( x ) , f j ( x ) , (4) Last edited D ate : I I I.C. Loss function page 10 IV . EXPERIMENTS and ↑ denotes an upsampling op eration (describ ed in Sec. I I I . B . ). S im is a similarit y function, suc h as m utual information, MSE and cross-correlation functions. Here, w e adopted MSE for faster training. Additionally , L img sim is adapted into an uncertaint y-w eighted loss, enabling the mo del to identify and adjust for regions with unreliable registration in ( 2 ). W e additionally calculated Dice loss function b etw een xed mask s fix and warped mask from mo ving mask s mov : L Dice = 1 − 2 P x ∈ Ω s fix ( x ) s mov ◦ T ↑ ( x ) P x ∈ Ω s fix ( x ) + P x ∈ Ω s mov ◦ T ↑ ( x ) + ϵ . (5) to measure the o verlap b et ween the predicted segmen tation mask and the ground truth segmen tation mask. A b ending energy loss is dened as: L bend = 1 X X i ∇ 2 t i 2 + 2 X ρ = ϱ ∂ 2 t i ∂ ρ∂ ϱ 2 ! (6) , where ∇ 2 is the Laplacian op erator, and t i is the displacemen t eld at vo xel lo cation i , ρ, ϱ ∈ { x, y , z } denote 3 spatial dimensions. The b ending energy loss p enalises the second deriv ative of the displacemen t eld, encouraging the netw ork to generate smo other deformation elds 30 . The nal loss can b e written as: L = λ 1 L uncertainty + λ 2 L DS C + λ 3 L bend , where λ 1 , λ 2 and λ 3 are hyperparameters that control the relative imp ortance of the dierent loss terms. Dice loss is only included when segmentation masks for training data are av ailable, i.e., Dataset 1 and Dataset 2. IV . Exp erimen ts IV . A . Datasets Dataset 1: Longitudinal prostate T2-weigh ted MR images were acquired from 86 patients at Univ ersity College London Hospitals NHS F oundation. All patien ts provided written consen t. Among them, 60 patien ts had 2 visits, 8 patien ts had 3 visits, and 18 patien ts had 4 visits, yielding a total of 216 T2-weigh ted volumes. This dataset w as used to p erform intr a-p atient registration. All the image and mask v olumes w ere resampled to 0 . 7 × 0 . 7 × 0 . 7 mm 3 isotropic v o xels. The in tensit y of all image volumes is normalised to a range of [0 , 1] . T o facilitate computation, all images and masks were also cropp ed from the cen tre IV. EXPERIMENTS R unning title here: Printed Ma rch 19, 2026 IV . A . Datasets Ldmk 1 Ldmk 2 Ldmk 3 Ldmk 4 Ldmk 5 Figure 3: Examples of anatomical landmarks annotated for the same patien t at tw o dieren t time p oints. The rst and second rows show 5 landmarks at the rst and second time p oin ts, resp ectiv ely . of v olume to 128 × 128 × 102 vo xels with preserved prostate glands. The data set w as divided in to 70, 6 and 10 patients for training, v alidation, and holdout test sets, comp osing 156, 32 and 38 in tra-patient pairs, resp ectively . F or images in the holdout test set, pairs of corresp onding anatomical and pathological landmarks are man ually identied on moving and xed images by a radiologist with more than 5 y ears of exp erience in reading prostate cancer mpMR images. Examples are shown in Figure 3 , including patient-specic uid-lled cysts, calcication, and cen troids of zonal b oundaries. The landmarks dier from patien t to patien t b ecause of the complex anatomical structure of the prostate. Dataset 2: W e used p elvis data from 850 prostate cancer patients at Universit y Col- lege London Hospital for inter-p atient registration. Details of the data can b e found in 31 , 32 , 33 , 34 , 35 . All patien ts pro vided written consent, and ethics appro v al was granted as per trial proto cols in 36 . W e use 313, 58, and 69 patients for training, v alidation and holdout test sets, resp ectively . This dataset spans several clinical trials conducted at UCLH, with images acquired from v arious clinical scanners. The patient group includes b oth biopsy and therapy cases. Sp ecics on vendors and imaging proto cols for eac h study can b e found in 37 . All pa- tien ts provided written consent and ethics appro v al w as granted as p er trial proto cols 36 . W e p erform inter-patien t registration on Dataset 2. The 850 patients (30 patien ts hav e m ultiple T2 mo dalities) w ere randomly partitioned in to train, v alidation and holdout test sets, on patien t-level, resulting in 626, 116 and 118 patien ts, resp ectiv ely . Within each set, image pairs are randomly formed and sampled without replacement, yielding 313, 58, and 59 pairs for train, v alidation, and holdout test sets, resp ectively . T2-w eighted images ha ve in-plane Last edited D ate : IV.A. Datasets page 12 IV . EXPERIMENTS dimensions ranging from 180 × 180 to 640 × 640, with a resolution of 1.31 × 1 . 31 mm 2 to 0.29 × 0 . 29 mm 2 , and slice thickness b et ween 0.82 and 1 mm. All image mo dalities w ere resam- pled to isotropic vo xels of 1 × 1 × 1 mm 3 using linear in terp olation, with in tensit y normalised b et ween 0 and 1 per mo dality . Prostate masks are lab elled b y radiologists with more than 5 years of exp erience in in terpreting prostate mpMR images. Inter-subject corresponding landmarks are unav ailable in this dataset. Dataset 3: W e used a public brain MR dataset LUMR from Learn2Reg 38 to p erform inter-p atient registration, whic h contains 3384 training and 40 v alidation sets, respectively . All image mo dalities were resampled to isotropic vo xels of 1 × 1 × 1 mm 3 using linear in ter- p olation, with in tensity normalised b et ween 0 and 1 p er mo dalit y . W e split the dataset into 3000, 384 and 40 for training, v alidation and holdout test, resp ectiv ely . W e used F reeSurfer 39 to generate brain zonal segmen tation, and 7 landmarks were lab eled by exp erienced exp erts for holdout test data as shown in Figure 4 . No segmen tation lab el was provided for training; L1 L2 L3 L4 L5 L6 L7 Figure 4: Examples of 7 landmarks lab eled by exp erts in brain dataset. th us, the dice loss was not used during training for this dataset. It is w orth noting that only training and v alidation sets are used in up dating weigh ts and tuning hyperparameters. holdout test sets are only used for mo del inference; all the results in this pap er are based on the holdout test set. IV . B . Comparison and ablation studies IV . B . 1 . Comparison studies T o rigorously assess the p erformance of the prop osed metho d, we conducted extensiv e exp er- imen ts across three datasets, comparing against represen tative state-of-the-art approac hes: dense deformation eld estimation and keypoint-based methods. F or dense-grid-based ap- IV. EXPERIMENTS IV.B. Compa rison and ablation studies R unning title here: Printed Ma rch 19, 2026 IV . C . Net work and baselines implementations proac hes, w e selected V o xelMorph 3 , a CNN-based metho d that predicts vo xel-wise displace- men t elds using U-Net architectures, and T ransMorph 4 , a recent transformer-based v ariant that lev erages self-atten tion mec hanisms to mo del long-range spatial dep endencies. In the k eyp oint-based category , we b enchmark ed against KeyMorph 5 , which emplo ys a sparse set of automatically detected anatomical keypoints to driv e the registration. T o further elucidate the eciency adv an tages of our metho d, we p erformed a focused arc hitectural comparison with KeyMorph, analysing the relationship b et w een model complexit y (quantied b y en- co der depth) and registration accuracy . W e included a strong iterativ e baseline: ANT s SyN(dieomorphic) 3 with matc hed pre-pro cessing and masks, tuned on the v alidation set. IV . B . 2 . Ablation studies W e ev aluated the signicance of key comp onen ts by isolating these elements. W e identied their individual con tributions to mo del p erformance and v alidated the eectiv eness of our arc hitectural decisions. Ablation studies in volv ed the following exp eriments: (1) Pro jected- skip connection: W e compared the p erformance of the proposed method with and without the skip-projection connection, which is used to project 3D features in to 1D vectors. (2) Grid size: W e in vestigated the impact of v arying grid sizes on registration p erformance, sp ecically using grid sizes of 5, 8, 10, and 15 in eac h dimension. (3) Enco der c hannel g θ enc : W e assessed the inuence of dieren t base channels (8, 16, and 32) in the enco der on registration p erformance. (4) Hyp erparameters in loss function: W e ev aluated the eect of dieren t h yp erparameters in the loss function. (5) Ba y esian head: W e ev aluate how the Ba yesian head aects registration accuracy and deformation regularity . IV . C . Net w ork and baselines implemen tations W e implemented all experiments with PyT orc h 2.0 under CUD A 12.4 on NVIDIA-SMI 550.107.02 with 32GB memory . The optimiser was A dam, with a learning rate of 10 − 4 and a batch size of 4. The net work w as trained for 300 ep o chs. All comparison metho ds w ere trained with their ocial implemen tation. Last edited D ate : IV.C. Net wo rk and baselines implementations page 14 V . RESUL TS IV . D . Ev aluation metrics Registration accuracy metrics The Dice similarit y co ecien t (DSC) was used to measure the o verlap b etw een the xed and warped prostate masks. Cen troid distance is adopted to ev aluate the mean Euclidean distance b etw een corresp onding cen troids: D CD = 1 K P K i =1 c fix i − c warp i 2 , where c ( · ) i denotes the i -th landmark centre or region of in terest mask cen tre in the xed/warped image, denoted as D ldmk CD and D msk CD , resp ectively . the centroids of the xed and warped R OI masks, computed as c ( · ) = ∑ x ∈ Ω x s ( · ) ( x ) ∑ x ∈ Ω s ( · ) ( x ) , with s ( x ) ∈ { 0 , 1 } the mask v alue at vo xel x . Jacobian-based deformation regularit y . Let ϕ : Ω ⊂ R 3 → R 3 denote the estimated spatial transformation. The Jacobian matrix of ϕ at vo xel x is J ϕ ( x ) = ∇ ϕ ( x ) ∈ R 3 × 3 . W e quantify lo cal volume change using the log-Jacobian determinan t log det( J ϕ ( x )) , where v alues near 0 indicate near-incompressible mappings, p ositive v alues indicate lo cal expansion, and negative v alues indicate lo cal con traction, denoted as log detJ . W e also rep orted the folding rate, dened as the percentage of vo xels with a negativ e Jacobian determinan t, det( J ϕ ( x )) < 0 , which corresp onds to non-inv ertible (folded) transformations. F or the o v erall methods comparison, w e compared GridReg to each baseline on the same cases using paired, one-sided tests and controlled multiplicit y within eac h dataset × metric family using Benjamini–Ho c hberg FDR correction 40 at 5%; w e report q -v alues. F or the ablation study , as they are indep enden t one-b y-one comparisons, we p erformed a paired t-test with a signicance lev el α = 0 . 05 . V . Results V . A . Comparison results In T able 1 , ANT s SyN w as 40 × slo wer than learning-based metho ds and yielded low er Dice/cen troid distance. V o xelMorph and T ransmorph, which used DDF, emphasised global shap e registration and achiev ed m uch larger landmark TRE (8.76mm and 7.73mm) in T able 1 with Dataset 1. V. RESUL TS IV.D. Evaluation metrics R unning title here: Printed Ma rch 19, 2026 V . A . Comparison results F or Dataset 2, GridReg achiev ed a DSC of 0.78, sligh tly lo w er than KeyMorph’s score of 0.79 but higher than T ransMorph and V o xelMorph. Both GridReg and KeyMorph ac hieved comparable p erformance in mask-based CD for the p elvis, with GridReg at 4.52 mm and Key- Morph at 4.14 mm, suggesting reliable mask alignment. F or Dataset3, GridReg maintained a DSC of 0.75, outp erforming V oxelMorph and T ransMorph. GridReg matched KeyMorph on mask-based CD (2.97 mm vs 3.05 mm, p = 0 . 832 ) and trailed sligh tly on Dice (0.77 vs 0.78, p = 0 . 471 ) but using less computation cost. These results suggested that when data features w ere rich in corresp ondence, architectural or transformation constrain ts mat- tered less, and p erformance dierences were narrow. Figure 5 show ed prostate registration, where large homogeneous regions made dense DDF mo dels (e.g., V o xelMorph, T ransMorph) more prone to physically unlikely lo cal distortions, despite comparable Dice scores. Despite ne-tuning the hyperparameters for b ending energy loss, the net work struggled to preserve the prostate structure. This highligh ts the imp ortance of method selection in registering v arious medical images. In con trast, Figure 6 shows tw o xed–mo ving brain pairs (rst t wo columns) and the corresp onding results from four metho ds, i.e. GridReg, V oxelMorph, KeyMorph and T ransMorph, resp ectively , without implausible deformation readily visible across all metho ds. T able 1: Performance and memory across metho ds. V alues are mean ± SD o ver identical holdout test cases. Prop osed metho ds app ear in the last columns (GridReg). F or eac h dataset, we compared GridReg to eac h baseline using paired, one-sided t-tests and controlled m ultiplicity within the family using BH-FDR ( 5% ); w e rep ort q-v alues. Bold indicates the b est mean. ‡ denotes GridReg is signicantly b etter than all baselines ( q < 0 . 05 ); † denotes it is signicantly b etter than three of baselines ( q < 0 . 05 ). Slash / indicates that a result is not av ailable. Dataset Metric Original ANT s SyN V o xelMorph T ransMorph KeyMorph GridReg(Ours) Prostate Dice ↑ 0.70 ± 0.10 0 . 81 ± 0 . 05 0.86 ± 0.02 0.84 ± 0.03 0.86 ± 0.05 0.88 ± 0.02 ‡ D ldmk C D ↓ (mm) 9.60 ± 4.09 9.09 ± 8.09 8.76 ± 8.59 7.73 ± 6.98 3.98 ± 2.14 4.54 ± 2.77 † D msk C D ↓ (mm) 8.76 ± 4.04 3.38 ± 0.78 2.70 ± 0.54 3.05 ± 0.84 2.56 ± 0.66 1.87 ± 1.16 ‡ GPU(Mb) / / 208 377 274 146 P elvis Dice ↑ 0 . 53 ± 0 . 17 0.69 ± 0 . 09 0.73 ± 0.16 0.73 ± 0.09 0.79 ± 0.07 0.78 ± 0.09 † D msk C D ↓ (mm) 10.65 ± 5 . 85 6.03 ± 2 . 78 5.99 ± 5 . 68 4.47 ± 2 . 95 4.14 ± 2.47 4.52 ± 2.88 † GPU(Mb) / / 168 333 225 128 Brain Dice ↑ 0.70 ± 0.05 0.69 ± 0 . 12 0.78 ± 0.07 0.77 ± 0.05 0.77 ± 0.06 0.78 ± 0.06 D ldmk C D ↓ (mm) 8.67 ± 6.58 8 . 92 ± 6 . 10 8.45 ± 6.64 8.70 ± 6.64 8.35 ± 6.85 8.21 ± 6.58 † D msk C D ↓ (mm) 4.16 ± 1.94 3.98 ± 1 . 79 3.11 ± 1.73 3.80 ± 1 . 14 2.97 ± 1.72 3.05 ± 1.47 GPU(Mb) / / 317 486 378 240 T rainable Parameters (bytes) / / 524181 41476659 1410329 252960 Inference Time (case/s) / 8.08 0.23 0.25 0.24 0.23 As shown in the last t wo rows of T able 1 , The inference time w ere: GridReg 0.23 s, V oxelMorph 0.23 s, KeyMorph 0.24 s, T ransMorph 0.25 s, resp ectiv ely . Although GridReg Last edited D ate : V.A. Compa rison results page 16 V . RESUL TS used 2×–163× few er parameters than the baselines, single-image inference time is similar b ecause GPU execution is dominated b y reading the input images and cop ying them from memory to the compute units. Our metho d substantially reduces the num b er of parameters, enabling deploymen t on smaller GPUs, but it do es not noticeably reduce inference time due to the nature of GPU computing. Keymorph achiev ed sligh tly lo w er DSC than the prop osed metho d. It ac hiev ed higher accuracy in terms of centroid distances of landmarks, but it re- quired more computation cost, as shown in T able 2 . W e compared KeyMorph and GridReg with v arying encoder lay ers. KeyMorph’s DSC improv ed from 0.54 at 4 la y ers to 0.86 at 9 la yers, requiring deep er netw orks for impro ved p erformance. GridReg consisten tly outp er- forms KeyMorph across all congurations, with DSC from 0.88 to 0.90. This highlighted GridReg’s ability to main tain high p erformance with few er parameters. Though GridReg ac hieved sligh tly lo w er landmark distances compared to KeyMorph with more than 7 lay- ers, it ac hieved more stable CD across all la y ers, further demonstrating its eciency and reduced computation cost. KeyMorph demonstrated go o d p erformance on Dataset 3 (brain imaging), where structural consistency supports its k eyp oin t-based design, but p erformed less eectiv ely on prostate datasets, where such consistency is lacking, as shown in T able 1 . V . B . Ablation studies W e conducted ablation studies to assess the eectiveness of the prop osed metho d on Dataset 1. (1), remo ving the skip-pro jection module resulted in a lo wer DSC of 0.86 com- pared to GridReg ( p = 2 . 02 × 10 − 234 ). The skip-pro jection and deco der rene the deformation T able 2: KeyMorph vs. GridReg: Performance comparison across v arying Enco der la yer coun ts (Dataset 1). Results sho w Dice, D ldmk C D (mm), and inference memory cost. An asterisk ( ∗ ) indicates a statistically signicant improv emen t ov er the comparator metho d at the same la yer count ( p < 0 . 05 ). La yers KeyMorph GridReg Dice D ldmk C D (mm) Cost(Mb) Dice D ldmk C D (mm) Cost(Mb) 4 la y ers 0.54 ± 0.14 14.74 ± 9.46 169 0.88 ± 0.03 ∗ 6.05 ± 3.43 ∗ 145 5 la y ers 0.56 ± 0.14 14.69 ± 12.40 171 0.88 ± 0 . 02 ∗ 4.54 ± 2.77 ∗ 146 6 la y ers 0.58 ± 0.19 10.97 ± 7.52 174 0.89 ± 0.03 ∗ 3.97 ± 2.38 ∗ 147 7 la y ers 0.82 ± 0.18 4.42 ± 3.24 182 0.89 ± 0.02 ∗ 4.59 ± 3.09 148 8 la y ers 0.84 ± 0.07 4.33 ± 2.84 195 0.89 ± 0.04 ∗ 4.25 ± 2.46 152 9 la y ers 0.86 ± 0.05 3.98 ± 2.14 ∗ 208 0.90 ± 0.03 ∗ 4.64 ± 2.82 157 V. RESUL TS V.B. Ablation studies R unning title here: Printed Ma rch 19, 2026 V . B . Ablation studies Fixed Moving (A) GridReg (B) Vo x e l M o r p h (C) KeyM orph (D) Tra n sM o rp h (E) ANT s SyN (1) (2) (3) (4) Figure 5: Visualisation of prostate registration. The rst column and the second column sho w four examples of the xed and mo ving images, resp ectively . The next four columns show the registration results from(A)GridReg, (B) V oxelMorph, (C) KeyMorph, (D)T ransMorph and (E) ANT s SyN, resp ectiv ely . (C) KeyMorph (D) Tra ns Mo r ph (B) Vo x e l M o r p h Fixed Moving (A) GridReg (E) ANTs SyN (1) (2) Figure 6: Visualisation of brain registration. The rst column and the second column show four examples of the xed and mo ving images, resp ectively . The next four columns sho w the registration results from (A)GridReg, (B) V o xelMorph, (C) KeyMorph, (D)T ransMorph and (E) ANT s SyN, resp ectiv ely . eld, capturing b oth global and lo cal information for b etter accuracy . (2), v arying the grid con trol p oin ts show ed that (10,10,10) control p oints achiev ed the best accuracy (T able 3 ). (3), increasing base channels impro ved accuracy , with 32 c hannels performing b est Dice, Last edited D ate : V.B. Ablation studies page 18 V . RESUL TS T able 3: Ablation study of prop osed metho d with v arying conguration of grid n umber, base c hannel of enco der, upsampling metho d and usage of pro jection mo dule. The ablation studies are p erformed on holdout test set in Dataset 1. The sym b ol * represents that the b est results ha ve a statistical dierence b etw een Cong 4 and other metho ds (all p − v al ues < 0.050). Setting Grid Base-NC Upsample Pro jector Ba yesian Dice D lmdk C D (mm) Cost(Mb) Cong 1 5 32 trilinear ✓ ✓ 0.86 ± 0.05 ∗ 5.29 ± 2.85 146 Cong 2 10 32 trilinear ✓ ✓ 0.89 ± 0.03 4.89 ± 3.19 148 Cong 3 10 8 trilinear ✓ ✓ 0.87 ± 0.04 5.88 ± 3.27 144 Cong 4 10 16 trilinear ✓ ✓ 0.88 ± 0.02 4.54 ± 2.77 146 Cong 5 10 16 trilinear × ✓ 0.86 ± 0.03 ∗ 6.06 ± 3.23 ∗ 145 Cong 6 10 16 decon v ✓ ✓ 0.89 ± 0.04 4.18 ± 2.88 171 Cong 7 10 16 bspl ✓ ✓ 0.89 ± 0.03 4.45 ± 2.95 290 Cong 8 10 16 trilinear ✓ × 0.88 ± 0.03 4.90 ± 3.65 146 Cong 9 15 16 trilinear ✓ ✓ 0.90 ± 0.03 ∗ 5.29 ± 3.09 148 though the dierence from 16 channels w as not signicant (DSC p = 0 . 053 ). (4), cubic B-spline (via transp osed con volution) achiev ed higher cen troid distances (p = 0.800) than trilinear in terp olation, while trilinear w as slightly faster. The transposed-conv olutional B- spline is markedly more ecient than the zero-insertion B-spline, which inates memory by expanding the grid with zero b efore ltering. fixed moving Uncertainty map fixed moving Uncertainty map 1 2 3 4 Figure 7: This gure shows four examples of uncertaint y maps generated by the Ba yesian Grid transformer. A lighter area in the Uncertaint y map represen ts lo wer uncertaint y , while a dark er area represents higher uncertain ty . (5), The Ba yesian head has b een used as a regulariser in prior w ork 12 , 28 . In our ex- p erimen ts, adding this head did not signican tly improv e registration accuracy ( p = 0 . 815 ). Lik ewise, Jacobian-based regularit y metrics show ed no signicant dierences, although we observ ed a near-signican t trend on Brain (Prostate: p = 0 . 513 ; Brain: p = 0 . 063 ; T able 4 ). Figure 7 presents tw o representativ e cases from the holdout test set, sho wing the xed im- V. RESUL TS V.B. Ablation studies R unning title here: Printed Ma rch 19, 2026 V . B . Ablation studies age, moving image, and corresp onding uncertaint y maps ( σ 2 ), where homogeneous regions exhibit higher uncertaint y than high-contrast b oundaries. (6), Figure 8 (1) sho ws that bal- ancing b ending energy loss w eight ( 2 × 10 5 ) optimally reduces distortion and aligns regions of in terest, ac hieving DSC=0.90 and D ldmk C D = 6 . 14 mm. Figure 8 (2–3) compares the sepa- rately trained and the grid-v ariable mo del across dierent grid sizes on Dataset 1. Using the same grid sizes, p erformance is similar: no signican t dierences w ere detected ( p > 0 . 205 ). The only visible gap is at grid = 5, where the grid-adaptiv e model sho ws a 0 . 39 mm lo wer cen troid distance (CD) than that of the mo del with prexed grid size ( p = 0 . 570 ) and 0.02 higher Dice than the prexed grid size ( p = 0 . 043 ). T able 4: Jacobian statistics across methods. log det( J ) is reported as mean ± standard de- viation; folding rate is the p ercentage of v o xels with det( J ) < 0 . F or each dataset, we rep ort paired, one-sided t-tests with BH-FDR(5%) with condence α = 0 . 05 . † and ∗ denote GridReg Bay esian and GridReg is signicantly b etter than other metho ds, resp ectively . Dataset Metric GridReg Bay esian GridReg KeyMorph V oxelMorph T ransMorph Prostate log det( J ) 0 . 0352 ± 0 . 0193 † 0 . 0446 ± 0 . 0228 ∗ 13 . 3064 ± 32 . 1452 1 . 9258 ± 1 . 0610 0 . 0975 ± 0 . 0532 det( J ) < 0 (%) 0 0 3 . 8 0 . 4 0 . 03 Brain log det( J ) 0 . 0022 ± 0 . 0015 † 0 . 0038 ± 0 . 0014 ∗ 55 . 2012 ± 133 . 4529 1 . 0793 ± 0 . 3254 15 . 9102 ± 6 . 5335 det( J ) < 0 (%) 0 0 11 . 0 0 . 2 4 . 5 Last edited D ate : V.B. Ablation studies page 20 V . RESUL TS Dice ↑ (Se parate vs. A daptive) 𝐷 !" #$%& 𝑚𝑚 ↓ (Separate vs . Adapti ve) 𝑤 ( 𝐵𝐷𝐸 )× 10 ' (1) (2) (3) Grid size Grid size Adaptive Adaptive DSC and CD of land marks v s. weigh t of b ending energy loss Figure 8: (1)Eect of Bending Energy W eight on Segmentation Accuracy (DSC) and Con tour Distance (CD); (2)-(3)Comparisons on Dataset1 showing Dice and cen troid distance b etw een separately trained and auto-adapted mo dels with the same grid size range from 5 to 15. V. RESUL TS V.B. Ablation studies R unning title here: Printed Ma rch 19, 2026 VI . Discussion Learning-based registration trades hand-crafted ob jectives for data-driven mo dels that learn complex, non-linear corresp ondences and pro vide single-pass fast inference with sucien t data requirements. Besides, with the domain-shift strategy , it could transfer across do- mains 41 . In data-scarce settings, classical metho ds with strong inductive priors may remain comp etitiv e. With moderately more data, as demonstrated in all our exp erimen ts, ho w- ev er, learning-based registration attains state-of-the-art accuracy with substan tially lo wer run time. Across three datasets, our results highligh t complementary strengths and failure mo des of current 3D registration strategies, and motiv ate design c hoices b ehind GridReg. Dense DDF s can ov ert lo cally in homogeneous tissue. On Dataset 1 (prostate), V o xelMorph and T ransMorph achiev ed reasonable Dice but large landmark TRE and visibly distorted gland interiors (Fig. 5 ). Increasing the b ending-energy weigh t reduced lo cal distortion but also imp eded con vergence (Fig. 8 (1)), reecting a known smo othness–delity trade-o: excessiv e regularisation suppresses deformations, whereas weak regularisation permits im- plausible w arps 9 , 10 . Dice alone did not fully exp ose these issues, underscoring the need to consider geometric plausibility alongside o verlap. Prostate MR images often exhibit relativ ely homogeneous in tensity with the prostate gland and substantial anatomical v ariation across patients. These c haracteristics mak e it c hallenging for deep netw orks to reliably identify consistent keypoints or corresp ondences b et ween dieren t sub jects. Constraining the degrees of freedom at prediction time using a simple coarse grid reduces the hypothesis space of admissible ows (often unnecessary lo cal deformation), limiting spurious lo cal distortions without sacricing mask o verlap. In con trast, brain MR (Dataset 3) provides abundan t, consistent edges; all metho ds pro duced visually plausible deformations with similar centroid distances. In prostate registration, the b est-p erforming grids w ere o ver 20 × coarser than vo xel- lev el DDF s, pro viding no evidence that a full-resolution parameterisation is necessary for these datasets. Instead, coarse grids b enet from inheren t spatial regularisation, and p erfor- mance was relatively insensitiv e to the exact grid size within a reasonable (alb eit p otentially application-dep enden t) range, from 5 × 5 × 5 to 15 × 15 × 15 . W e observ ed that using a grid size of approximately 10% of the image size, in each dimension, led to generally go o d registration p erformance, based on Dataset 1. Last edited D ate : page 22 VI I . CONCLUSION Although Bay esian head did not improv e registration p erformance, w e included this uncertain ty estimation for reference purp oses for those interested in Figure 7 and T able 3 . These visualisations highlight t wo main observ ations: regions with weak lo cal features (e.g. homogeneous areas suc h as ven tricles) are assigned high uncertain ty due to the lack of dis- tinctiv e gradients for unam biguous alignment, whereas contours and high-con trast structures exhibit lo w er uncertain ty . Clinical relev ance Higher DSCs in the prostate and p elvis regions suggest that GridReg could supp ort future clinical applications in organ trac king and atlas construction, p ending prosp ectiv e v alidation and task-sp ecic accuracy b enc hmarks. The low memory consumption of GridReg mak es it suitable for use in resource-constrained en vironments, suc h as mobile devices and edge computing systems. VI I . Conclusion This work provides a systematic comparison of learning-based registration parameterisations that predict deformation from either (1) sparse, coarse regular grids of control points, (2) scattered non-gridded con trol points and (3) dense displacement elds. W e show that reg- ular grids oer a practical adv an tage: they enable fast and stable reconstruction of dense deformation elds via standard in terp olation (e.g., trilinear) or transp ose-con volution-based spline appro ximations, av oiding the additional complexity and ov erhead of resampling from irregular p oin t sets. Imp ortantly , a sparse deformation parameterisation pro vides implicit capacit y control: fewer degrees of freedom suppress high-frequency vo xel-wise warps driven b y noisy or ambiguous corresp ondences, while reducing memory and compute. GridReg fur- ther supp orts grid-adaptiv e training, allowing a single mo del to op erate across m ultiple grid resolutions and to select an eective grid densit y for eac h dataset through v alidation, rather than committing to a xed resolution a priori. Finally , the approach is enco der-agnostic: it can b e attached to dieren t bac kb one enco ders from existing registration net works with minimal changes. Overall, GridReg pro vides a simple, general, and ecien t path to sparse deformation mo delling, impro ving the eciency–accuracy trade-o and broadening applica- bilit y across div erse medical image registration tasks. VI I. CONCLUSION R unning title here: Printed Ma rch 19, 2026 A c kno wledgmen t This work is supp orted by the In ternational Alliance for Cancer Early Detection, an alliance b et ween Cancer Research UK [C28070/A30912; C73666/A31378; EDD AMC-2021/100011], Canary Cen ter at Stanford Universit y , the Universit y of Cambridge, OHSU Knigh t Cancer Institute, Universit y College London and the Univ ersity of Manc hester. This work is also supp orted by the National Institute for Health Research (NIHR) Universit y College London Hospitals (UCLH) Biomedical Research Cen tre (BRC). Conict of In terest Statemen t Mark Emberton receiv es researc h support from the United Kingdom’s National Institute of Health Researc h (NIHR) UCLH/UCL Biomedical Researc h Cen tre. He acts as consul- tan t/lecturer/trainer to Sonacare Inc. Angio dynamics Inc. Early Health Ltd and Alb emarle Medical Ltd The remaining authors declare that they ha ve no kno wn comp eting nancial in terests or p ersonal relationships that could hav e app eared to inuence the work rep orted in this pap er. Data a v ailabilit y Dataset 1 and Dataset 2 are retrosp ectiv e datasets, whic h are not publicly av ailable. Part of Dataset 2 is av ailable for academic use up on formal application at https://ncita.org. uk/promis- data- set- open- access- request . Dataset 3 is publicly a v ailable at https: //learn2reg.grand- challenge.org 1 D. L. Hill, P . G. Batchelor, M. Holden, and D. J. Hawk es, Medical image registration, Ph ysics in medicine & biology 46 , R1 (2001). 2 J. M. Fitzpatric k et al., Image registration, Handb o ok of medical imaging 2 , 447–513 (2000). Last edited D ate : page 24 3 G. Balakrishnan, A. Zhao, M. R. Sabuncu, J. Guttag, and A. V. Dalca, V oxelmorph: a learning framew ork for deformable medical image registration, IEEE transactions on medical imaging 38 , 1788–1800 (2019). 4 J. Chen, E. C. F rey , Y. He, W. P . Segars, Y. Li, and Y. Du, T ransmorph: T ransformer for unsup ervised medical image registration, Medical image analysis 82 , 102615 (2022). 5 M. Y. Ev an, A. Q. W ang, A. V. Dalca, and M. R. Sabuncu, KeyMorph: Robust multi- mo dal ane registration via unsup ervised keypoint detection, in Me dic al imaging with de ep le arning , 2022. 6 M. Homann, B. Billot, D. N. Grev e, J. E. Iglesias, B. Fischl, and A. V. Dalca, Syn th- Morph: learning con trast-inv ariant registration without acquired images, IEEE trans- actions on medical imaging 41 , 543–558 (2021). 7 J. Chen, Y. Liu, S. W ei, Z. Bian, S. Subramanian, A. Carass, J. L. Prince, and Y. Du, A survey on deep learning in medical image registration: New technologies, uncertaint y , ev aluation metrics, and b ey ond, Medical Image Analysis , 103385 (2024). 8 Q. Y ang, D. Atkinson, Y. F u, T. Sy er, W. Y an, S. Pun wani, M. J. Clarkson, D. C. Bar- ratt, T. V ercauteren, and Y. Hu, Cross-Mo dalit y Image Registration Using a T raining- Time Privileged Third Mo dality , IEEE T ransactions on Medical Imaging 41 , 3421–3431 (2022). 9 elastix team, elastix, the manual , 2024, Accessed: 2025-11-01. 10 M. W ang and P . Li, A review of deformation mo dels in medical image registration, Journal of Medical and Biological Engineering 39 , 1–17 (2019). 11 J. Leng, G. Xu, and Y. Zhang, Medical image interpolation based on m ulti-resolution registration, Computers & Mathematics with Applications 66 , 1–18 (2013). 12 A. V. Dalca, G. Balakrishnan, J. Guttag, and M. R. Sabuncu, Unsup ervised learning of probabilistic dieomorphic registration for images and surfaces, Medical image analysis 57 , 226–236 (2019). 13 N. J. T ustison and B. B. A v ants, Explicit B-spline regularization in dieomorphic image registration, F rontiers in neuroinformatics 7 , 39 (2013). R unning title here: Printed Ma rch 19, 2026 14 D. Ruec k ert, L. Sono da, C. Hay es, D. Hill, M. Leach, and D. Hawk es, Nonrigid registra- tion using free-form deformations: application to breast MR images, IEEE T ransactions on Medical Imaging 18 , 712–721 (1999). 15 M. A. A udette, F. P . F errie, and T. M. P eters, An algorithmic o v erview of surface registration techniques for medical imaging, Medical Image Analysis 4 , 201–217 (2000). 16 M. Jaderb erg et al., Spatial transformer net works, Adv ances in neural information pro cessing systems 28 (2015). 17 S. Shan, W. Y an, X. Guo, E. I.-C. Chang, Y. F an, and Y. Xu, Unsup ervised End-to-end Learning for Deformable Medical Image Registration, 2018. 18 J. Zhou, B. Ma, W. Zhang, Y. F ang, Y.-S. Liu, and Z. Han, Dieren tiable registration of images and lidar p oin t clouds with v oxelpoint-to-pixel matching, A dv ances in Neural Information Pro cessing Systems 36 (2024). 19 N. Hussain, Z. Y an, W. Cao, and M. Anw ar, Hierarc hical renemen t with adaptive deformation cascaded for m ulti-scale medical image registration, Magnetic Resonance Imaging , 110449 (2025). 20 D. P antazis, A. Joshi, J. Jiang, D. W. Shattuc k, L. E. Bernstein, H. Damasio, and R. M. Leahy , Comparison of landmark-based and automatic metho ds for cortical surface registration, Neuroimage 49 , 2479–2493 (2010). 21 T. Lange, N. P ap enberg, S. Heldmann, J. Mo dersitzki, B. Fisc her, H. Lamec ker, and P . M. Schlag, 3D ultrasound-CT registration of the liv er using com bined landmark- in tensity information, In ternational journal of computer assisted radiology and surgery 4 , 79–88 (2009). 22 A. Q. W ang, R. Saluja, H. Kim, X. He, A. Dalca, and M. R. Sabuncu, BrainMorph: A F oundational Keyp oint Mo del for Robust and Flexible Brain MRI Registration, arXiv preprin t arXiv:2405.14019 (2024). 23 M. Ekv all, L. Bergenstråhle, A. Andersson, P . Czarnewski, J. Olegård, L. Käll, and J. Lundeberg, Spatial landmark detection and tissue registration with deep learning, Nature Metho ds 21 , 673–679 (2024). Last edited D ate : page 26 24 Z. M. Baum, Y. Hu, and D. C. Barratt, Real-time m ultimo dal image registration with partial intraoperative p oint-set data, Medical image analysis 74 , 102231 (2021). 25 Y. F u, Y. Lei, T. W ang, P . P atel, A. B. Jani, H. Mao, W. J. Curran, T. Liu, and X. Y ang, Biomec hanically constrained non-rigid MR-TR US prostate registration using deep learning based 3D p oint cloud matching, Medical image analysis 67 , 101845 (2021). 26 D. DeT one, T. Malisiewicz, and A. Rabino vich, Sup erp oint: Self-sup ervised interest p oin t detection and description, in Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition workshops , pages 224–236, 2018. 27 J. Zhang, Y. W ang, J. Dai, M. Ca vichini, D.-U. G. Bartsc h, W. R. F reeman, T. Q. Nguy en, and C. An, T wo-Step Registration on Multi-Modal Retinal Images via Deep Neural Netw orks, IEEE T ransactions on Image Pro cessing 31 , 823–838 (2022). 28 X. Gong, L. Khaidem, W. Zh u, B. Zhang, and D. Do ermann, Uncertain ty learning to wards unsup ervised deformable medical image registration, in Pr o c e e dings of the IEEE/CVF Winter Confer enc e on A pplic ations of Computer V ision , pages 2484–2493, 2022. 29 M. F ey , J. E. Lenssen, F. W eichert, and H. Müller, Splinecnn: F ast geometric deep learning with con tinuous b-spline kernels, in Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition , pages 869–877, 2018. 30 D. R ueck ert, L. I. Sono da, C. Ha y es, D. L. Hill, M. O. Leach, and D. J. Hawk es, Non- rigid registration using free-form deformations: application to breast MR images, IEEE transactions on medical imaging 18 , 712–721 (1999). 31 S. Hamid et al., The SmartT arget biopsy trial: a prosp ective, within-p erson randomised, blinded trial comparing the accuracy of visual-registration and magnetic resonance imag- ing/ultrasound image-fusion targeted biopsies for prostate cancer risk stratication, Eu- rop ean urology 75 , 733–740 (2019). 32 L. A. Simmons et al., The PICTURE study: diagnostic accuracy of m ultiparametric MRI in men requiring a rep eat prostate biopsy , British journal of cancer 116 , 1159–1165 (2017). R unning title here: Printed Ma rch 19, 2026 33 C. Orczyk et al., Prostate Radiofrequency F o cal Ablation (ProRAFT) T rial: A Prosp ec- tiv e Developmen t Study Ev aluating a Bip olar Radiofrequency Device to T reat Prostate Cancer, The Journal of Urology 205 , 1090–1099 (2021). 34 L. Dic kinson et al., A m ulti-centre prospective dev elopment study ev aluating fo cal therap y using high intensit y fo cused ultrasound for lo calised prostate cancer: the INDEX study , Contemporary clinical trials 36 , 68–80 (2013). 35 A. E.-S. Bosaily et al., PR OMIS—prostate MR imaging study: a paired v alidating cohort study ev aluating the role of m ulti-parametric MRI in men with clinical suspicion of prostate cancer, Con temp orary clinical trials 42 , 26–40 (2015). 36 S. Hamid et al., The SmartT arget biopsy trial: a prosp ective, within-p erson randomised, blinded trial comparing the accuracy of visual-registration and magnetic resonance imag- ing/ultrasound image-fusion targeted biopsies for prostate cancer risk stratication, Eu- rop ean urology 75 , 733–740 (2019). 37 W. Y an et al., Combiner and h yp ercombiner netw orks: Rules to com bine multimodality MR images for prostate cancer lo calisation, Medical Image Analysis 91 , 103030 (2024). 38 A. Hering et al., Learn2Reg: comprehensive multi-task medical image registration chal- lenge, dataset and ev aluation in the era of deep learning, IEEE T ransactions on Medical Imaging 42 , 697–712 (2022). 39 B. Fischl, F reeSurfer, Neuroimage 62 , 774–781 (2012). 40 Y. Benjamini and Y. Ho ch b erg, Controlling the false disco v ery rate: a practical and p o werful approach to multiple testing, Journal of the Ro yal statistical so ciet y: series B (Metho dological) 57 , 289–300 (1995). 41 S. Zheng, X. Y ang, Y. W ang, M. Ding, and W. Hou, Unsup ervised cross-modality domain adaptation netw ork for x-ray to CT registration, IEEE Journal of Biomedical and Health Informatics 26 , 2637–2647 (2021). Last edited D ate :
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment