AgriChat: A Multimodal Large Language Model for Agriculture Image Understanding
The deployment of Multimodal Large Language Models (MLLMs) in agriculture is currently stalled by a critical trade-off: the existing literature lacks the large-scale agricultural datasets required for robust model development and evaluation, while cu…
Authors: Abderrahmene Boudiaf, Irfan Hussain, Sajid Javed
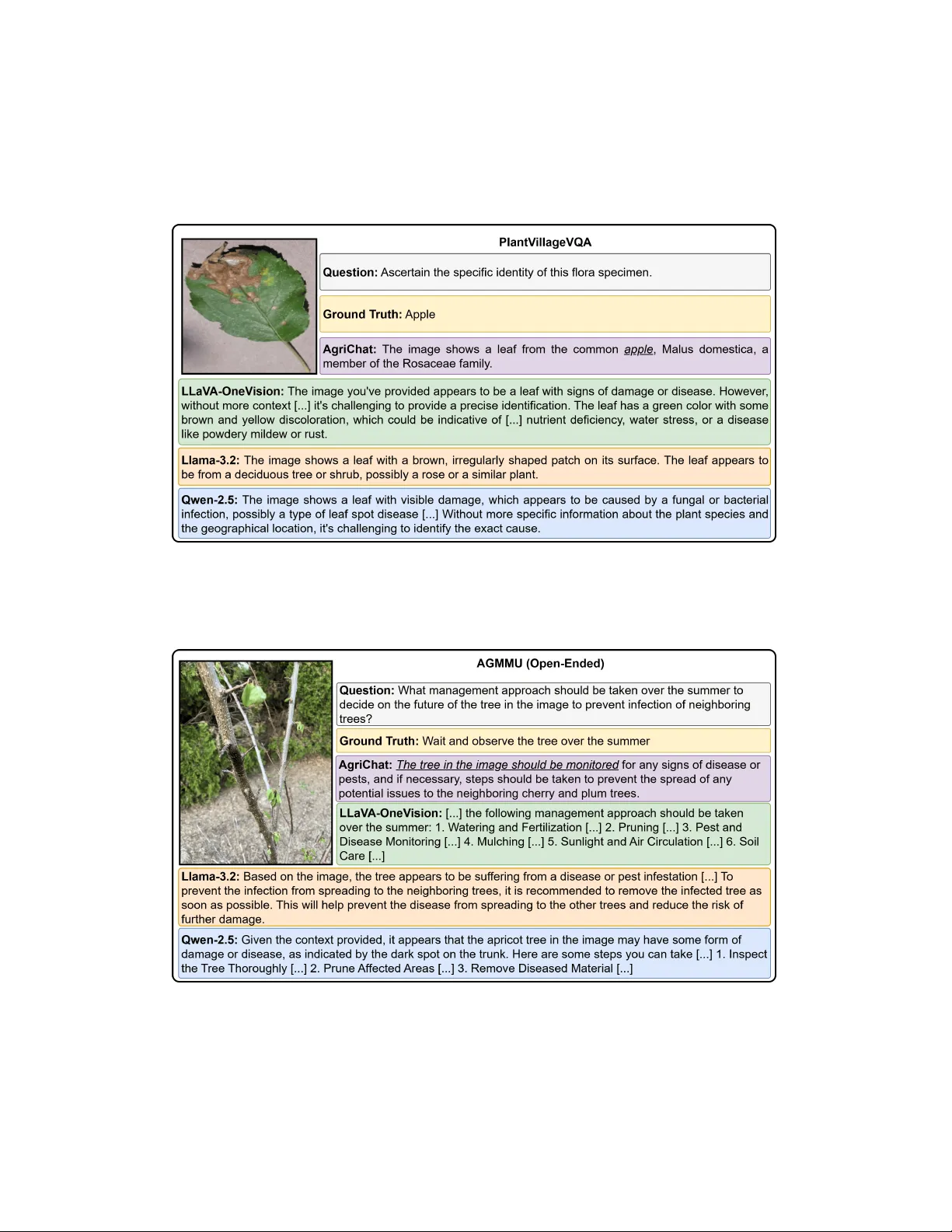
AgriChat: A Multimodal Lar ge Language Model for Agriculture Image Understanding Abderrahmene Boudiaf a, ∗ , Irfan Hussain a , Sajid Jav ed a a Department of Computer Science, Khalifa University of Science and T echnology , Abu Dhabi, United Arab Emir ates Abstract The deployment of Multimodal Lar ge Language Models (MLLMs) in agriculture is currently stalled by a critical trade-o ff : the existing literature lacks the lar ge- scale agricultural datasets required for robust model de velopment and e v aluation, while current state-of-the-art models lack the verified domain expertise necessary to reason across di verse taxonomies. T o address these challenges, we propose the V ision-to-V erified-Knowledge (V2VK) pipeline, a nov el generativ e AI-driv en annotation framework that integrates visual captioning with web-augmented sci- entific retrie v al to autonomously generate the AgriMM benchmark, e ff ecti vely eliminating biological hallucinations by grounding training data in verified phy- topathological literature. The AgriMM benchmark contains ov er 3,000 agricul- tural classes and more than 607k VQAs spanning multiple tasks, including fine- grained plant species identification, plant disease symptom recognition, crop count- ing, and ripeness assessment. Le veraging this v erifiable data, we present AgriChat , a specialized MLLM that presents broad knowledge across thousands of agricul- tural classes and provides detailed agricultural assessments with extensi v e expla- nations. Extensi ve e v aluation across di verse tasks, datasets, and e v aluation condi- tions rev eals both the capabilities and limitations of current agricultural MLLMs, while demonstrating AgriChat’ s superior performance over other open-source mod- els, including internal and e xternal benchmarks. The results v alidate that pre- serving visual detail combined with web-v erified kno wledge constitutes a reliable pathway toward robust and trustworth y agricultural AI. The code and dataset are publicly av ailable at https://github.com/boudiafA/AgriCha t . ∗ Corresponding author Email addr ess: 100058322@ku.ac.ae (Abderrahmene Boudiaf) K e ywor ds: Multimodal Large Language Models, F oundation models, Agricultural, Plant Identification, Plant Disease Analysis, Crop counting. 1. Introduction Figure 1: Illustrati ve examples of AgriChat’ s con versational diagnostic capabilities across three core agricultural tasks. Left: Fine-grained plant species identification with follo w-up knowledge queries. Center: Plant disease symptom recognition, correctly diagnosing Sigatoka disease from visible leaf lesions. Right: Fruit counting and ripeness assessment from a single field image. These examples highlight AgriChat’ s ability to support interactive, expert-lev el agricultural rea- soning beyond simple classification. Agriculture remains the cornerstone of global economic stability and food se- curity , sustaining approximately 80% of the world’ s rural poor and serving as the primary instrument for poverty alleviation [6, 7]. As the global population is projected to approach 10 billion by 2050, the agriculture sector faces mounting pressure to increase production by an estimated 70% while simultaneously na vi- gating the existential threats of climate change, soil degradation, and diminishing arable land [8]. Artificial Intelligence (AI) has emer ged as a transformati ve force, shifting traditional farming paradigms tow ard data-driven precision agriculture [9]. Recent studies indicate that AI-inte grated systems ranging from predicti v e analytics for yield optimization to computer vision for early disease detection en- hance crop producti vity by up to 20% while reducing w ater and fertilizer inputs by nearly 30% [10, 11]. Consequently , the integration of AI tools is no longer merely adv antageous but essential for fostering resilient agricultural systems capable of meeting future nutritional demands within planetary boundaries [12]. 2 Figure 2: (a) Comparison between existing foundation models (Llama-3.2 [1], LLav a-OneV ision [2], Qwen-2.5 [3]) and our proposed agriculture-focused AgriChat model which achiev ed su- perior performance across four agriculture benchmarks. In comparison to existing agriculture benchmark datasets (PlantV illageVQA [4], A GMMU [5]), our proposed large-scale benchmark AgriMM exceeds the state-of-the-art benchmarks in terms of (b) number of images, (c) number of classes, and (d) number of VQA pairs. 3 In the literature, the Con v olutional Neural Networks (CNNs) ha ve dri v en much of this progress, they typically function as “black boxes, ” providing classification labels without the explanatory reasoning required for e ff ecti v e decision support [13]. This limitation has catalyzed interest in Multimodal Large Language Mod- els (MLLMs), which o ff er the ability to reason ov er visual inputs and provide interacti ve diagnostics [14]. Howe v er , the e ffi cacy of MLLMs is strictly bound by the quality of the data upon which the y are trained [15]. Existing agricultural V isual Question Answering (VQA) datasets su ff er from a fundamental trade-o ff : they either rely on expensi v e human annotation that limits scale and taxonomic di versity (e.g., PlantV illage with only 38 diseases [16], CDDM with 60 disease types biased toward commodity crops [17]), or they employ synthetic generation using frozen LLMs that may introduce incorrect hallucinated information without real-time verification. Historical expert logs like AgMMU, while factually ac- curate at creation, su ff er from kno wledge staleness as agricultural best practices, pesticide re gulations, and pathogen management protocols continuously ev olve [5]. T o address this gap, we introduce the Agriculture Multi-Modal dataset (AgriMM), which resolves both limitations simultaneously through a novel approach to data curation. Unlike previous benchmarks, AgriMM w as created by systematically aggregating and filtering 63 distinct agricultural datasets to ensure broad taxo- nomic cov erage, spanning more than 3,000 classes and 121,000 images, con- taining questions related to fine-grained species identification, disease recogni- tion, and crop counting tasks. The core nov elty lies in our V ision-to-V erified- Kno wledge (V2VK) pipeline, a three-stage synthesis process designed to achiev e di verse, accurate, and verifiably grounded knowledge. First, we employ Gemma 3 [18] to generate detailed visual captions for each agricultural image, captur - ing ke y visual attributes such as plant morphology , gro wth stage, visible symp- toms of disease when present, and other agronomically rele v ant features. Sec- ond, utilizing a Retriev al-Augmented Generation (RA G) framew ork, we lev erage Gemini 3 Pro [19] with real-time web access to produce a comprehensi ve class- le vel description covering taxonomy , biology , and known diseases, drawing from authoritati ve sources to filter out hallucinations and ensure factual accuracy . Fi- nally , LLaMA 3.1 [1] synthesizes both the image caption and the class description into 607,000 di v erse instruction-tuning question-answer pairs. This rigorous cura- tion ensures that the ground truth te xtual descriptions are not merely linguistically plausible, but scientifically current. Despite the av ailability of such data, current model architectures present a secondary bottleneck. Existing agricultural MLLMs including Agri-LLaV A [20], 4 AgroGPT [14], and LLaV A-PlantDiag [21] univ ersally su ff er from limited and narro w fine-tuning data: they are trained on small-scale, task-specific corpora that cov er only a handful of crops and diseases, restricting their ability to generalize across the v ast taxonomic and pathological di v ersity encountered in real-world agricultural settings [14, 20]. For instance, models fine-tuned on fewer than 200 classes inevitably fail when confronted with underrepresented re gional culti v ars, emerging pathogens, or multi-task scenarios requiring simultaneous species iden- tification, disease diagnosis, and yield estimation [22]. In this work, we propose AgriChat , an MLLM framew ork that fundamentally ov ercomes this data bottle- neck. AgriChat is fine-tuned on our lar ge-scale AgriMM dataset comprising o ver 121k images and over 607k QA pairs spanning more than 3,000 classes, repre- senting the widest and most di verse agricultural fine-tuning corpus to date. T o e ff ecti v ely adapt the base architecture to the agricultural domain while pre- serving its general-purpose reasoning capabilities, AgriChat employs a parameter- e ffi cient fine-tuning strategy using Lo w-Rank Adaptation (LoRA) [23]. Lightweight trainable adapters are injected into both the vision encoder and the language model decoder , while all pre-trained weights remain frozen. The model undergoes su- pervised fine-tuning on the AgriMM corpus, optimizing an autoregressi ve objec- ti ve with role-aware masking. This training paradigm allo ws AgriChat to acquire expert-le v el agricultural diagnostic kno wledge, covering fine-grained species iden- tification, disease recognition, and crop counting, while retaining the vast world kno wledge inherent in the base model, all within a computationally e ffi cient foot- print (inference in ∼ 2.3 seconds on consumer hardware). W e v alidate the e ffi cacy of our approach through rigorous experiments across four di verse benchmarks, including AgriMM, CDDM [17], PlantV illageVQA [4], and A GMMU [5]. Our results demonstrate that AgriChat not only achiev es state- of-the-art performance on in-domain tasks b ut also exhibits superior zero-shot generalization to unseen datasets compared to larger open-source models. The main contributions of this paper are summarized as follo ws: 1. Agriculture Multimodal Dataset (AgriMM): W e introduce a high-quality instruction-tuning dataset comprising ov er 121k images and 607k e xpert- aligned QA pairs. By employing a web-search augmented generation pipeline, we solv e the issue of kno wledge staleness and hallucination, ensuring taxo- nomic di versity across 3,000 + agricultural classes. 2. AgriChat MLLM: W e introduce the first multimodal lar ge language model (MLLM) purpose-built for agriculture and fine-tuned on the widest and 5 most di verse range of agricultural species to date, enabling state-of-the- art performance on fine-grained species identification, disease classification and crop counting tasks. 3. Comprehensiv e Benchmarking: Extensi ve ev aluation on four agriculture benchmarks demonstrates that our proposed AgriChat achie ves superior performance in both in-domain tasks and zero-shot generalization, outper- forming state-of-the-art generalist baselines. The rest of this paper is organized as follo ws: Section 2 revie ws the e v olution of agricultural AI and analyzes the limitations of current VQA datasets. Section 3 details the construction of the AgriMM dataset through our V ision-to-V erified- Kno wledge pipeline and presents the AgriChat architecture. Section 4 presents the experimental setup, quantitati v e comparisons, and qualitati v e analysis. Finally , Section 5 concludes the paper and outlines future directions. 2. Related W ork 2.1. Existing Agricultur e Datasets and T asks The dev elopment of rob ust agricultural AI has been historically constrained by the av ailability and quality of training data. Early benchmarks like PlantV illage [24] provided a foundational resource with 54,306 images co vering 38 disease classes. Howe v er , its reliance on laboratory-controlled imaging limits its applica- bility to complex field environments. More recent e ff orts hav e attempted to scale up visual div ersity . The CDDM (Crop Disease Diagnosis Multimodal) bench- mark [17] expanded the scope to 137,000 images and 1 million QA pairs. Y et, as noted in recent critiques, its taxonomic scope remains constrained to 60 dis- ease types across 16 major crop categories, reflecting a fundamental bias tow ard economically significant commodity crops (e.g., maize, rice, wheat) while leaving regional heirlooms and emer ging pathogens underrepresented [22]. T o address the prohibiti ve cost of manual annotation, researchers hav e turned to synthetic gen- eration and historical log mining. AgroInstruct [14] utilized div erse vision-only datasets to synthesize 70,000 instructions. While scalable, this approach relies on frozen language models that su ff er from hallucination, generating plausible but factually incorrect statements about disease etiology or management proto- cols that were ne ver true [25]. Con v ersely , the A GMMU benchmark [5] curated 57,079 e xpert-farmer dialogues from historical extension logs. While factually accurate at the time of creation, these datasets su ff er from “kno wledge staleness”: 6 agricultural best practices ev olve continuously as pesticide regulations change and ne w pathogen strains appear . Expert dialogues from 2020 may no longer reflect current recommendations in 2026 [26]. Furthermore, a critical barrier to adv anc- ing agricultural AI is the repr oducibility crisis . As detailed in T able 1, the largest corpus, Agri-3M-VL, remains unpublished. This pre vents independent verifica- tion of reported performance and hinders regional adaptation, where researchers might otherwise fine-tune models for local crops. T o bridge these gaps, we intro- duce AgriMM. Unlike prior datasets, AgriMM is a Multi-Source dataset consol- idating 63 data sources into 121,425 images and 607,125 QA pairs. It integrates three critical tasks: fine-grained species identification, disease diagnosis, and crop counting spanning 3,099 classes. 2.2. V ision Models in Agriculture Deep learning in agriculture initially focused on pure visual classification us- ing Con v olutional Neural Networks (CNNs). Architectures such as ResNet [27], DenseNet [28], and E ffi cientNet [29] established the dominant paradigm, achiev- ing impressiv e accurac y rates exceeding 95% on controlled datasets. Recent ad- v ancements ha ve introduced hybrid CNN-T ransformer architectures to capture global conte xt. Notable e xamples include LGNet [30], which emplo ys dual- branch adaptiv e feature fusion, and ST -CFI [31], which implements bidirectional fusion of CNN spatial features with Swin T ransformer [32] tokens. Howe v er , despite their superior classification performance, all such systems fundamentally operate as black-box discriminati ve models. They accept an image as input and return only a categorical label (e.g., “T omato Late Blight”) without explanatory reasoning, contextual information, or interacti ve diagnostic capabilities. This lack of interpretability limits their utility in decision support systems, where farmers require not just a diagnosis, but an understanding of the cause and appropriate management strategies. 2.3. V ision Language Models in Agricultur e T o ov ercome the limitations of closed-set classification, researchers began ex- ploring V ision-Language Models (VLMs) that align image features with natural language representations. Early attempts such as PlantV illageVQA [4] introduced question-answer pairs but su ff ered from rigid templates (e.g., “What disease does this plant hav e?”), resulting in limited linguistic di v ersity and shallo w reason- ing depth. More recent work has lev eraged large-scale pretraining. ALiv e [33] provides approximately 600,000 image-te xt pairs covering crops and li v estock to train AgriCLIP . By aligning visual encoders with agricultural text, AgriCLIP 7 demonstrated significant gains (9.07%) in zero-shot classification ov er standard CLIP models. Ho we ver , these models are primarily designed for retriev al and matching tasks. While they can associate an image of a leaf with the text “boron deficiency , ” they lack the generativ e capacity to engage in multi-turn dialogue or provide nuanced reasoning about symptom progression required for interacti v e agricultural decision support. 2.4. MLLMs in Agricultur e The most recent paradigm shift in volves Multimodal Lar ge Language Models (MLLMs), which integrate visual encoders with Large Language Models (LLMs) to enable con v ersational diagnostics. Agri-LLaV A [20] pioneered kno wledge- infused training by incorporating agricultural domain expertise into the LLaV A architecture. Similarly , AgroGPT [14] and LLaV A-PlantDiag [21] hav e pushed the boundaries of interacti ve diagnosis, with the latter achie ving BLEU-4 scores of 48.7% on con v ersational diagnostic tasks. Howe ver , a critical limitation shared by all e xisting agricultural MLLMs is their reliance on narro w , small-scale fine- tuning datasets that cov er only a limited number of crops and disease classes. This data bottleneck constrains their capacity to generalize across the v ast taxo- nomic and pathological div ersity encountered in real-world agricultural deploy- ments, particularly for underrepresented regional cultiv ars, emerging pathogens, and multi-task scenario s requiring simultaneous species identification, disease di- agnosis, and quantitati ve yield estimation. In this work, we propose AgriChat to address this data bottleneck. As illustrated in Figure 1, AgriChat supports interacti ve, e xpert-le v el diagnostic reasoning across a di v erse range of agricul- tural tasks from fine-grained plant species identification and multi-turn knowl- edge queries, to pathogen-le vel disease diagnosis and fruit counting with ripeness assessment. AgriChat is fine-tuned via Low-Rank Adaptation (LoRA) [23] on our proposed AgriMM dataset. This parameter-e ffi cient adaptation strategy in- jects domain-specific agricultural kno wledge into both the vision encoder and the language backbone while preserving the general-purpose reasoning capabilities of the base model. 3. Methodology 3.1. AgriMM Dataset T o address the scarcity of high-quality , verifiable agricultural data, we in- troduce AgriMM, a large-scale visual benchmark that represents a robust con- solidation of div erse agricultural en vironments. AgriMM aggregates 63 source 8 datasets, comprising 1 fine-grained taxonomic dataset (iNatAg subset [34]), 33 counting / detection datasets, and 29 disease classification datasets (see Appendix A for complete source list), to create a total of 121,425 images annotated with 607,125 V isual Question Answering (VQA) pairs, covering over 3,000 distinct classes. The dataset architecture is built upon three complementary components, ensuring coverage across species-le vel identification, pathological diagnosis, and quantitati ve crop monitoring. Unlike existing datasets that often su ff er from label noise and taxonomic inconsistencies, AgriMM is created through a rigorous cu- ration pipeline that balances broad taxonomic cov erage with specific, high-v alue agronomic tasks. T o contextualize the contribution of AgriMM, T able 1 com- pares it against prominent agricultural VQA benchmarks. Existing datasets such as CDDM [17] and PlantV illageVQA [4] are limited by narrow task definitions (lacking counting or fine-grained classification) or rely on static, potentially out- dated knowledge bases. While Agri-3M-VL [35] o ff ers scale, it remains unav ail- able to the research community . AgriMM addresses these limitations by o ff ering the first publicly av ailable, multi-source benchmark that integrates fine-grained taxonomy , counting capabilities, and a dynamic W eb-RA G verification pipeline. T able 1: Comparison of Agricultural VQA Datasets. AgriMM stands out by integrating 63 data sources and utilizing a W eb-RAG pipeline to ensure knowledge is factual and up-to-date, unlike static or hallucination-prone alternativ es. Dataset Name # Images # QA Pairs # Classes Multi-Source Fine-Grained Counting W eb-V erified A vailable CDDM [17] 137,000 1,000,000 76 ✗ ✗ ✗ ✗ ✓ PlantV illageVQA [4] 55,448 193,609 52 ✗ ✗ ✗ ✓ ∗ ✓ Agri-3M-VL [35] ∼ 1.0M ∼ 3.0M 42,253 ✓ ✓ ✗ ✗ ✗ AgroInstruct [14] 108,701 70,000 202 ✓ ✗ ✗ ✗ ✗ A GMMU [5] 57,079 58,571 5 ✗ ✗ ✗ ✓ ∗ ✓ AgriMM (Ours) 121,425 607,125 3,099 ✓ ✓ ✓ ✓ ✓ ∗ V erified by humans but static / outdated (no live web retrie val). 3.1.1. Dataset Statistics Component Distribution: The dataset is stratified into three major subsets: 1. Fine-Grained Identification: The largest component, comprising 48,580 images spanning 2,956 distinct species. This subset is org anized into nine categories (e.g., Cereals , Le gumes , Medicinal ) to ensure taxonomic breadth. 2. Disease Classification: Consisting of 49,348 images, this subset targets 110 specific disease conditions across 33 major crops (a veraging ∼ 1,701 images per crop). It cov ers critical staples such as Wheat , Rice , and Corn , alongside cash crops like Co ff ee and Sugar cane . 9 3. Crop Counting & Detection: A specialized set of 23,497 images across 33 crops (av eraging ∼ 712 images per crop), designed for spatial reasoning tasks. Categorical Aggregation: By integrating these three sources, AgriMM achiev es a robust distrib ution across nine agricultural categories. The aggregated totals re- veal a focus on high-economic value and high-biodi versity segments: Ornamen- tal / Other (48,112 images), Fruits (25,710), V e getables & T ubers (19,317), and In- dustrial Crops (14,954). The remaining distribution ensures ecological complete- ness with Cereals & Grasses (13,528), W eeds / W ild (10,902), Medicinal & Spices (5,195), Forestry & Timber (4,723), and Legumes / Pulses (3,572). This distribu- tion (visualized in Figure 3) prev ents the common bias to wards solely commercial crops by maintaining significant representation of wild and forestry species. 3.1.2. Dataset T asks AgriMM is designed to train models capable of addressing three critical agri- cultural AI capabilities, each reflecting real-world deployment requirements: 1. Di ff erential Pathology Diagnosis. Moving beyond binary healthy / sick clas- sification, this task requires models to distinguish between visually similar con- ditions across 29 geographically di v erse disease datasets. The collection spans multiple continents, from bean rust in Uganda to gua v a canker in P akistan to vine viruses in Europe, ensuring models learn to generalize across en vironmental and culti v ar v ariations. Critical diagnostic challenges include di ff erentiating between ov erlapping symptomatology (e.g., Bacterial Spot vs. Fungal Blight in tomatoes), identifying co-occurring stresses (e.g., nutritional deficiency vs. pathogen infec- tion in chilli), and classifying physical damage distinct from biological stress (e.g., ’crushed’ vs. ’crack ed’ sugarcane stalks). This capability is essential for preci- sion agriculture decision support systems where misdiagnosis can lead to incorrect treatment protocols. 2. Species-Lev el T axonomic Recognition. The fine-grained identification task challenges models to perform botanical discrimination using standardized scien- tific nomenclature rather than common names. Sourced from iNaturalist Agricul- ture (iNatAg), this subset includes observ ations from e very continent, reflecting real-world en vironmental variability , from Abies balsamea (balsam fir) in North American boreal forests to Acacia erioloba (camelthorn) in African sav annas. The task requires distinguishing between morphologically similar species within eco- nomically important genera ( Acacia , Eucalyptus ) and identifying in v asi v e or nox- ious weeds in complex field backgrounds (e.g., Abutilon theophrasti among cash 10 (a) Hierarchical composition of AgriMM. (b) Quantitativ e distribution across cate gories. Figure 3: AgriMM dataset statistics sho wing (a) taxonomic hierarchy and (b) class balance across functional categories. 11 crops). This capability supports biodi versity monitoring, in v asi v e species detec- tion, and regulatory compliance in agricultural trade. 3. Quantitative Spatial Reasoning f or Y ield Estimation. Unlike classification tasks, counting and detection require models to perform spatial analysis under highly variable field conditions. The detection subset includes annotations for fruits and crops captured at di ff erent times of day (morning / night illumination), maturity stages (unripe / ripe / occluded stra wberries), and density lev els (sparse to heavily o verlapping). Models must detect and count indi vidual agricultural units, wheat heads in dense canopies, apples with varying occlusion, tomatoes at mix ed ripeness, while reasoning about spatial relationships and yield potential. This capability directly supports automated harvest planning, quality grading systems, and economic forecasting for gro wers. 3.1.3. V ision-to-V erified-Knowledge Synthesis Generating high-quality instruction-following data at scale requires overcom- ing the biological hallucinations common in frozen language models. W e intro- duce a three-stage V ision-to-V erified-Knowledge (V2VK) pipeline (Figure 4) that grounds visual observations in scientifically verified literature (full prompts provided in Appendix B). Stage I: V isual Gr ounding via Image Captioning . W e employ Gemma 3 (12B) [18] to generate structured natural language descriptions. T o prev ent label drift, we condition the generation on the ground-truth labels from AgriMM (e.g., injecting “ Solanum lycopersicum ”). The prompt enforces the extraction of agri- cultural metadata, including growth stage, planting density , and en vironmental context. Stage II: Knowledge Retriev al and V erification. V isual captions lack deep botanical knowledge. W e address this using Gemini 3 Pro [19] with a Retrie v al- Augmented Generation (RA G) approach. The model retrieves contemporary sci- entific descriptions, management protocols, and phenological data from trusted databases. Stage III: Instruction Synthesis. The final stage synthesizes the visual cap- tions (Stage I) and verified knowledge (Stage II) into 607,125 div erse QA pairs using LLaMA-3.1-8B-Instruct [1]. W e utilize a constrained prompting strategy that mandates fiv e distinct question types per image: Identification, V isual Rea- soning, Health Condition, Culti v ation Knowledge, and Quantification. For images belonging to the Crop Counting & Detection subset, the ground-truth crop count is derived by tallying the number of bounding box annotations per image and in- jected directly into the generation prompt. This ensures that Quantification QA 12 Figure 4: Overview of the V ision-to-V erified-Knowledge Synthesis Pipeline. V isual features are grounded in verified scientific literature through a three-stage generation and v erification process. 13 pairs reflect verifiable, annotation-grounded values rather than model estimates, pre venting the hallucination of incorrect counts. 3.1.4. Pipeline V erification and Quality Assur ance T o guarantee the biological v alidity and semantic accuracy of the generated instructions, we implemented a rigorous human-in-the-loop verification protocol. While the pipeline automates the synthesis of scale, manual oversight is critical to pre vent the propagation of hallucinations common in Lar ge Language Models. Manual Knowledge verification: W e conducted a comprehensi ve revie w of the scientific kno wledge retrie v ed in Stage II. Expert annotators manually fact- checked the detailed class descriptions against established agricultural taxonomies and phytopathology literature. This process ensured that the retrieved biological traits, disease symptoms, and phenological data accurately corresponded to the visual e vidence (Stage I). 3.2. AgriChat Ar c hitectur e As illustrated in Figure 5, AgriChat is a domain-specialized Multimodal Lar ge Language Model (MLLM) tailored for high-precision agricultural diagnostics. The architecture is composed of four primary modules: a high-resolution vi- sion encoder , a cross-modal projection network, a Large Language Model (LLM) decoder , and a parameter -e ffi cient adaptation mechanism. In contrast to broad- scope MLLMs that prioritize con versational v ersatility , AgriChat focuses on fine- grained visual discrimination, a prerequisite for detecting minute pest damage, early-onset pathologies, and subtle phenotypic traits. 3.2.1. V ision Encoder: SigLIP-SO400M For visual feature extraction, we utilize SigLIP-SO400M (Sigmoid Loss for Language-Image Pre-training) [36]. This contrasti ve model was chosen ov er stan- dard CLIP v ariants for its superior ability to preserve spatial details essential for agricultural tasks. Formally , let T ∈ R H v × W v × 3 be an input image tile, where the nativ e resolution is fixed at H v = W v = 384. The encoder , denoted as E ω +∆ ω , retains its pre-trained base parameters ω in a frozen state while introducing a set of trainable LoRA parameters ∆ ω to adapt the encoder to agricultural visual features. It processes the tile to produce a spatial feature map F s p , defined as: F s p = E ω +∆ ω ( T ) ∈ R h × w × d v (1) 14 Figure 5: Overview of the AgriChat Architecture. The model utilizes an adaptive resolution strategy to process high-quality agricultural imagery . Input images are split into local patches and a resized global thumbnail, encoded by SigLIP equipped with LoRA adapters, and aligned via a projection layer . The visual tokens are concatenated with text instructions and processed by LLM backbone equipped with LoRA adapters (SwiGLU + Multi-Head Self-Attention) to generate verifiable diagnostic responses. 15 where h = w = 27 represents the spatial grid size, and d v = 1152 is the visual hidden dimension. Flattening this map yields a sequence of N v = h × w = 729 tokens per tile. 3.2.2. High-Resolution V isual Encoding T o mitigate the resolution loss common in standard resizing, we adopt the adapti ve resolution strategy [2]. This allo ws the model to process high-resolution field photography by decomposing images into nati ve-resolution patches. Stage 1: Adaptive Grid Generation. Giv en a high-resolution input image I ∈ R H × W × 3 , we calculate a grid configuration ( n h , n w ) that best approximates the orig- inal aspect ratio. This grid is constrained by a pre-defined token budget N max . In our implementation, we set N max = 8748, allo wing the processing of images up to 1344 × 1344 resolution. Stage 2: Dual-P ath Featur e Extraction. The encoding process operates on two parallel tracks: • Local Path: The image is di vided into n h × n w non-ov erlapping patches {T i , j } . Each patch is encoded indi vidually and flattened to e xtract local de- tails: H i , j = Flatten( E ω +∆ ω ( T i , j )) ∈ R N v × d v . • Global Path: A downsampled thumbnail T global is encoded to capture holis- tic context: H global = Flatten( E ω +∆ ω ( T global )) ∈ R N v × d v . Stage 3: Adaptiv e P ooling. Let L denote the total number of visual tokens gen- erated by the dual paths, calculated as L = ( n h · n w + 1) · N v . T o handle v ariable resolutions e ffi ciently , if L exceeds the sequence limit N max , we apply bilinear in- terpolation, denoted as Interp( · ), in the feature space. Let F i , j ∈ R h × w × d v be the unflattened feature map of a specific tile. The final local features ˜ H i , j are com- puted as: ˜ H i , j = Flatten Interp( F i , j , size = ( h ′ , w ′ )) if L > N max H i , j otherwise (2) where ( h ′ , w ′ ) represents the reduced spatial dimensions calculated to satisfy the condition that the ne w total token count ˜ L ≤ N max . 3.2.3. Cr oss-Modal Pr ojection Network T o bridge the modality gap between the visual encoder ( d v = 1152) and the language model ( d llm = 3584), we employ a Multi-Layer Perceptron (MLP) pro- jector P ψ . W e use the bracket notation [ · , · ] to denote tensor concatenation along the sequence dimension. 16 The complete sequence of visual features is formed by concatenating the global thumbnail features with the processed local patch features ˜ H i , j : H visual = [ H global , ˜ H 1 , 1 , . . . , ˜ H n h , n w ] ∈ R ˜ L × d v (3) Let ψ = { W 1 , b 1 , W 2 , b 2 } be the trainable projection weights, and GELU( · ) be the Gaussian Error Linear Unit acti v ation function. The projected visual tokens Z v are obtained via: Z v = P ψ ( H visual ) = GELU( H visual W 1 + b 1 ) W 2 + b 2 (4) This results in Z v ∈ R ˜ L × d llm , aligning the visual representations with the LLM’ s embedding space. 3.2.4. Language Model Decoder The core reasoning engine is Qwen-2-7B [3], a transformer-based decoder D ϕ +∆ ϕ whose pre-trained weights ϕ are kept frozen while trainable LoRA param- eters ∆ ϕ are injected to enable domain adaptation. Multimodal Fusion: Fusion is achie ved through early embedding inte gration [2]. Let E text ( · ) denote the word embedding layer of the LLM which maps discrete tokens to R d llm . For a tokenized instruction sequence X text = { x 1 , . . . , x n } contain- ing a special placeholder token x k = , the input embeddings S are created by replacing the placeholder with the visual sequence: S = [ E text ( x 1 ) , . . . , E text ( x k − 1 ) , Z v , E text ( x k + 1 ) , . . . , E text ( x n )] (5) This unified sequence S serves as the input to the decoder . 3.3. T raining Strate gy 3.3.1. P arameter -E ffi cient Domain Adaptation (LoRA) T o adapt the model to the agricultural domain, we utilize Low-Rank Adap- tation (LoRA) [23] across both the vision encoder and the language model de- coder . This strategy keeps all pre-trained base parameters frozen while injecting lightweight, trainable low-rank adapters, enabling e ffi cient domain specialization with minimal computational ov erhead. 17 V ision Encoder Adaptation.. The pre-trained SigLIP-SO400M [36] parameters ω are frozen, and trainable LoRA adapters ∆ ω are injected into the linear lay- ers of the vision encoder . For a frozen linear layer defined by weights W (vis) 0 ∈ R D (vis) in × D (vis) out , we define adapter matrices A (vis) ∈ R D (vis) in × r v and B (vis) ∈ R r v × D (vis) out , where r v = 32 is the lo w-rank dimension for the vision encoder . The forw ard pass for an input x ∈ R 1 × D (vis) in is modified as: h (vis) = xW (vis) 0 + xA (vis) B (vis) · α v r v (6) where α v is the scaling factor for the vision encoder adapters. Language Model Adaptation.. W e inject trainable Low-Rank Adapters into the linear layers of the decoder (specifically the Attention and SwiGLU projection layers). The base LLM parameters ϕ are kept frozen in their nativ e precision. Let D = 3584 be the hidden dimension of the Qwen-2-7B backbone. For a frozen linear layer defined by weights W 0 ∈ R D in × D out , we define adapter matrices A ∈ R D in × r and B ∈ R r × D out , where r = 128 is the low-rank dimension. The forward pass for an input x ∈ R 1 × D in is modified as follo ws: h = xW 0 + xAB · α r (7) where α = 256 is the scaling factor . For both components, A matrices are ini- tialized from a Gaussian distribution, while B matrices are initialized to zero to ensure training starts from the identity function. 3.3.2. Optimization Objective The model is optimized using the standard autoregressi v e cross-entropy loss. Let Θ = ψ ∪ { A (vis) l , B (vis) l } ∀ l ∪ { A l , B l } ∀ l represent the union of all trainable param- eters (projector weights and LoRA adapters across both the vision encoder and language model decoder). Gi ven a ground-truth response sequence Y of length T , we apply a mask M t ∈ { 0 , 1 } to exclude instruction tok ens from the loss calculation. The optimization objecti ve is defined as: L ( Θ ) = − T X t = 1 M t log P ( y t | Y < t , S ; Θ ) (8) This ensures the gradients are backpropagated only through Θ based on the like- lihood of the correct agricultural diagnostic tokens y t . 18 4. Experimental Evaluations 4.1. T raining and Implementation Details AgriChat was fine-tuned e xclusi vely on the AgriMM dataset using a consumer - grade NVIDIA R TX 3090 GPU (24GB VRAM). The AgriMM dataset was par- titioned using an 80:20 train / test split, with no ov erlap between the training and e v aluation subsets. Fine-tuning was performed for 1 epoch with a per -de vice batch size of 1 and gradient accumulation steps of 16, yielding an e ff ecti ve batch size of 16. A learning rate of 2 × 10 − 4 was used with bfloat16 mix ed precision. LoRA adapters were applied to both the Qwen2 LLM backbone (rank r = 128, α = 256) and the SigLIP vision encoder (rank r = 32, α = 64), targeting the attention pro- jections ( q_proj , k_proj , v_proj ) in both components, the LLM-specific lay- ers ( o_proj , gate_proj , up_proj , down_proj ), and the vision encoder-specific layers ( out_proj , fc1 , fc2 ). Inference benchmarks were conducted using 4-bit quantization, a batch size of 1, and a standardized test image (1,328 KB) with the prompt “What is the name of the plant in the image?” repeated across 100 infer- ence cycles. Giv en that comparable domain-specific models such as Agri-LLaV A and AgroGPT are not publicly released as of this writing, AgriChat (7B) was e v aluated against three state-of-the-art generalist multimodal models: LLaV A- OneV ision (7B) [2], Llama-3.2 (V ision-11B) [1], and Qwen-2.5 (VL-7B) [3]. These baselines were selected as representati v e of the current frontier in open- weight vision-language models across a range of parameter scales. 4.2. Evaluation Metrics Ev aluating generativ e con v ersational agents requires a multi-faceted frame- work, as no single metric captures all dimensions of response quality [37, 38]. W e therefore combine lexical ov erlap metrics, semantic similarity measures, and a multimodal alignment score. 4.2.1. Lexical and N-Gr am Metrics Lexical metrics measure surface-le vel textual overlap and provide a standard baseline for text generation e v aluation: • BLEU-4: Measures n -gram precision (unigrams through 4-grams) against the reference, with a bre vity penalty [39]. • ROUGE-2: Measures bigram overlap between system output and refer- ence, capturing basic phrasal coherence [40]. 19 • METEOR: Extends exact matching with stemming and synonymy via W ord- Net, and penalizes word order violations [41]. These metrics are nonetheless limited by their semantic blindness: a model generating a paraphrase of the correct answer is penalized despite factual equiv a- lence [42]. This moti v ates the semantic and judge-based metrics belo w . 4.2.2. Semantic and Embedding-Based Metrics • BER TScore (F1): Computes token-le v el cosine similarity between candi- date and reference using contextual BER T embeddings [43]. • Long-CLIP Cosine Similarity: Ev aluates semantic alignment between generated text and reference te xt within CLIP’ s shared embedding space [44], providing a text-to-text similarity measure that captures conceptual and se- mantic relationships beyond simple le xical ov erlap. • T5 Cosine Similarity (T5 Cos): Measures semantic similarity using the T5 encoder’ s latent representations [45]. • SBER T Similarity: Computes cosine similarity between sentence-le v el dense embeddings from a Siamese BER T architecture [46]. 4.2.3. LLM-as-a-J udge Our analysis rev eals a critical limitation of the lexical and embedding-based metrics described abov e, particularly salient for con v ersational diagnostic agents: a systematic bias toward brevity , which we term the verbosity penalty . This phe- nomenon results in misleadingly lo w scores for models providing deeper reason- ing, e v en when diagnostic accurac y remains correct. This issue arises from a mis- match between reference formatting and assistant-optimized generation. Ground truth annotations in standard diagnostic datasets typically provide minimal, bi- nary responses (e.g., “No”), whereas models trained for con versational assistance generate contextual e xplanations. Example of Metric Misalignment: Pr ompt: “Can you detect signs indicating absence of pathogens?” Gr ound T ruth: “No” Concise Pr ediction: “No” → BER TScore: 1.0 V erbose Pr ediction: “No, the image does not provide information about the presence or absence of pathogens. ” → BER TScore: ≈ 0.50 20 Both predictions are f actually correct and clinically equiv alent, yet surface- le vel metrics hea vily penalize the more informativ e response. This is not an iso- lated observation: [38] hav e formally demonstrated that con ventional reference- based metrics such as BLEU and R OUGE exhibit relativ ely lo w correlation with human judgments, particularly on tasks requiring contextual, open-ended gener- ation—precisely the setting of our diagnostic assistant. This limitation is further corroborated by [47], who identify verbosity bias as a documented failure mode in automated e v aluation, wherein longer b ut substanti v ely equi v alent responses recei ve systematically lower scores despite no loss in factual accuracy . More broadly , [48] highlight in their comprehensiv e surve y that traditional metrics fail to capture ke y aspects such as logical coherence and semantic correctness in gen- erati ve tasks, motiv ating a paradigm shift tow ard LLM-based ev aluation. These con v er ging findings motiv ate the adoption of the LLM-as-a-J udg e paradigm [49], which enables e v aluation based on semantic intent rather than surface form and correlates more strongly with human judgment than n-gram matching. W e em- ploy Qwen3-30B-A3B-Instruct as our ev aluator . Our selection is grounded in the “Judge’ s V erdict Benchmark” recently introduced by NVIDIA [50]. In their extensi v e analysis of 54 LLM judges, Qwen3-30B w as identified as a T ier 1 e v al- uator , demonstrating exceptional alignment with human annotators. Specifically , it achie ved a Z-score of | z | = 0 . 04 in the “T uring T est for Judges, ” classifying it as a Human-Like Judg e capable of preserving the nuances of expert ev aluation, outperforming larger proprietary models in agreement stability . Follo wing the prompt design strategies outlined in [49] regarding Criteria Decomposition , our e v aluation prompt separates assessment into four distinct axes: Correctness, Com- pleteness, Clarity , and Conciseness (full prompt can be found in Appendix C). W e utilize a 4-point Likert scale (1–4) rather than the traditional 5-point scale. This design choice draws upon the methodology in [50], which emphasizes the utility of discrete, normalized scoring to improve reliability . A 4-point scale forces the e v aluator to make a definiti v e decision (positi v e vs. negati v e) by removing the neutral middle option, thereby reducing the central tendenc y bias often observed in automated judges. By lev eraging a model prov en to exhibit human-like agree- ment patterns [50] and a constrained scoring rubric, we ensure that higher scores reflect genuine diagnostic utility rather than stochastic generati ve fluenc y . 4.3. Evaluation Datasets Our experimental framework e v aluates performance across a div erse set of agricultural benchmarks, ranging from controlled laboratory settings to complex field en vironments. 21 1. AgriMM (Ours): The primary training and in-domain e v aluation set. It consolidates 63 distinct data sources into 121,425 images and 607,125 QA pairs, verified via a V ision-to-V erified-Kno wledge pipeline. It cov ers fine- grained species identification, disease diagnosis, and crop counting. 2. PlantV illage [24]: A foundational dataset comprising 54,306 images of 38 disease classes. W e use this as a zero-shot transfer benchmark to ev aluate performance on laboratory-controlled imagery . 3. CDDM (Crop Disease Diagnosis Multimodal) [17]: A lar ge-scale dataset focusing on 16 major crop categories. W e utilize this to test generalization on web-crawled field images. 4. A GMMU [5]: A benchmark derived from 58,571 expert-f armer dialogues. W e utilize both the Multiple Choice Question (MCQ) and Open-Ended sub- sets to ev aluate con v ersational reasoning and management advice capabili- ties. 4.4. Qualitative Analysis T o complement the quantitative e valuation, we present representativ e exam- ples that illustrate the strengths and limitations of AgriChat relati v e to generalist baselines. These cases were selected to demonstrate ke y findings across a variety of agricultural vision-language tasks, including counting, species identification, open-ended decision-making, multiple-choice reasoning, and disease diagnosis. It is important to note that AgriChat was fine-tuned exclusi v ely on the AgriMM dataset; all remaining examples reflect zero-shot generalization to unseen bench- marks. 4.4.1. Example 1: Pr ecise Agricultur al Counting (AgriMM) As sho wn in Figure 6, AgriChat is the only model to produce the exact nu- merical answer , correctly identifying 61 wheat heads in the image. This reflects the direct benefit of fine-tuning on AgriMM, which exposes the model to pre- cise, count-oriented agricultural queries. The generalist baselines fail substan- tially: LLaV A-OneV ision resorts to a vague quantifier ("numerous"), while both Llama-3.2 and Qwen-2.5 incorrectly assert that no wheat heads are present at all, misidentifying the crop entirely . This stark contrast highlights how domain- specific fine-tuning equips AgriChat with the visual grounding necessary for fine- grained agricultural counting, a task that generalist models, trained without such priors, are ill-equipped to handle. 22 Figure 6: Qualitativ e comparison on an AgriMM wheat-head counting task. 4.4.2. Example 2: Zer o-Shot Species Identification (PlantV illageVQA) Figure 7 illustrates a zero-shot species identification task in which AgriChat correctly identifies the specimen as Malus domestica and situates it within the appropriate taxonomic family . The generalist baselines, by contrast, uniformly fixate on the visible pathology and treat the query as a disease diagnosis task rather than a species identification task. This reflects a fundamental misalign- ment between generalist model beha vior and agricultural workflo ws, where cor - rectly identifying “what plant” must logically precede determining “what ails it. ” AgriChat’ s agricultural fine-tuning appears to instil the correct task prioritization e ven in zero-shot settings. 4.4.3. Example 3: Open-Ended Management Reasoning (A GMMU) The open-ended management scenario presented in Figure 8 tests the mod- els’ ability to pro vide conte xtually appropriate, cautious advice under uncertainty . AgriChat’ s response aligns closely with the ground truth by recommending moni- toring and w atchful observ ation rather than immediate interv ention, appropriately ackno wledging the risk to neighboring trees. The generalist baselines lean in op- posite directions: LLaV A-OneV ision produces a generic horticultural checklist that ignores the specific decision context, while Llama-3.2 recommends immedi- ate remov al of the tree, a drastic and potentially unnecessary interv ention. Qwen- 2.5 similarly defaults to aggressi v e pruning and remediation steps. AgriChat’ s measured, context-sensiti v e response suggests that domain fine-tuning fosters a 23 Figure 7: Qualitativ e comparison on a PlantV illageVQA plant identification task. Figure 8: Qualitativ e comparison on an A GMMU open-ended tree management task. 24 more calibrated reasoning style aligned with real-world agricultural decision-making. 4.4.4. Example 4: Multiple-Choice V isual Reasoning (AGMMU) Figure 9: Qualitativ e comparison on an A GMMU multiple-choice visual reasoning task. In the zero-shot multiple-choice task depicted in Figure 9, AgriChat selects the correct answer (A) while all three generalist baselines fail. LLaV A-OneV ision selects option D without justification, Llama-3.2 cites low image resolution as an e xcuse before incorrectly choosing C, and Qwen-2.5 constructs a plausible- sounding but factually incorrect rationale in fav our of D. AgriChat, despite not being trained on A GMMU, correctly interprets the visual relationship between the vine and the wisteria as detrimental, demonstrating that its agricultural priors transfer to structured reasoning tasks. The pattern of confident but incorrect re- sponses from the baselines underscores the risk of deploying generalist models in agricultural advisory contexts without domain adaptation. 4.4.5. Example 5: Disease Diagnosis (CDDM) As illustrated in Figure 10, AgriChat correctly identifies T omato Y ellow Leaf Curl V irus (TYLCV) in this zero-shot scenario, matching the ground truth pre- cisely . The generalist baselines each propose di ff erent, incorrect diagnoses: LLaV A- OneV ision names Early Blight, Llama-3.2 suggests Bacterial Leaf Spot, and Qwen- 2.5 declines to commit to any diagnosis at all. The div ergence in the baselines’ 25 Figure 10: Qualitativ e comparison on a CDDM tomato disease diagnosis task. outputs, confidently wrong or deliberately ev asi v e, illustrates the lack of reliable plant pathology knowledge in generalist models. AgriChat’ s correct identification, achie ved without any CDDM training data, suggests that agricultural fine-tuning culti v ates transferable visual–semantic associations for disease recognition that extend well be yond the training distrib ution. 4.5. Quantitative Results Gi ven that comparable domain-specific models (e.g., Agri-LLaV A, AgroGPT) are not publicly released as of this writing, we compared AgriChat (7B) against state-of-the-art generalist multimodal models: LLaV A-OneV ision (7B), Llama- 3.2 (V ision-11B), and Qwen-2.5 (VL-7B). 4.5.1. Benchmark P erformance T able 2 and Figure 11 summarize the results. AgriChat demonstrates dominant performance on the in-domain AgriMM dataset (LLM Judge: 77.43%) and strong zero-shot transfer to PlantV illage (74.26%) and CDDM Diagnosis (69.94%). 26 Figure 11: Radar chart ev aluation of AgriChat against baselines. AgriChat sho ws distinct advan- tages in diagnostic tasks (AgriMM, CDDM, PlantV illage) while generalist models remain com- petitiv e in open-ended reasoning. 27 T able 2: Comprehensi ve benchmark on multiple datasets. All metrics are scaled to 0–100 for clarity . Bold indicates the best performance; underline indicates the second best. Di ff reports the absolute di ff erence in percentage points between AgriChat and the best-performing baseline. Dataset Metric LLaV A-OneVision Llama-3.2 Qwen-2.5 AgriChat Di ff (7B) (11B) (7B) (7B) (vs Best) AGMMU (MCQs) Lexical / Math Metrics BLEU-4 4.20 2.49 3.29 64.94 + 60.7 ↑ ROUGE-2 7.81 9.73 15.36 50.98 + 35.6 ↑ METEOR 8.88 28.06 31.70 63.87 + 32.2 ↑ Semantic Metrics BER TScore 84.57 83.89 84.53 91.22 + 6.7 ↑ LongCLIP 71.66 79.24 79.98 87.38 + 7.4 ↑ T5 Cos 25.81 51.32 53.83 72.61 + 18.8 ↑ SBER T 20.25 53.01 56.90 73.92 + 17.0 ↑ Accuracy Percentage Accuracy 70.13 65.73 70.94 70.19 − 0.8 ↓ AGMMU (Open-Ended) Lexical / Math Metrics BLEU-4 0.35 0.11 0.09 0.43 + 0.1 ↑ ROUGE-2 2.60 1.04 0.76 2.83 + 0.2 ↑ METEOR 12.45 7.69 6.62 13.44 + 1.0 ↑ Semantic Metrics BER TScore 83.74 80.24 80.07 82.81 − 0.9 ↓ LongCLIP 75.91 75.68 75.72 75.08 − 0.8 ↓ T5 Cos 47.17 44.40 43.47 49.95 + 2.8 ↑ SBER T 39.65 36.30 36.82 39.23 − 0.4 ↓ LLM-based LLM Judge (%) 52.49 55.70 59.93 46.93 − 13.0 ↓ PlantV illage- VQA Lexical / Math Metrics BLEU-4 0.14 0.08 0.03 2.00 + 1.9 ↑ ROUGE-2 0.65 0.50 0.22 3.18 + 2.5 ↑ METEOR 17.25 6.72 3.43 19.52 + 2.3 ↑ Semantic Metrics BER TScore 86.02 80.37 79.12 83.58 − 2.4 ↓ LongCLIP 80.65 79.11 79.89 75.20 − 5.5 ↓ T5 Cos 51.65 37.85 28.20 44.80 − 6.9 ↓ SBER T 41.64 20.95 10.90 32.13 − 9.5 ↓ LLM-based LLM Judge (%) 57.41 54.44 53.21 74.26 + 16.9 ↑ CDDM (Diagnosis) Lexical / Math Metrics BLEU-4 0.45 0.56 0.57 6.42 + 5.9 ↑ ROUGE-2 2.28 2.62 2.52 17.16 + 14.5 ↑ METEOR 17.17 18.63 18.11 39.59 + 21.0 ↑ Semantic Metrics BER TScore 84.91 85.06 84.82 89.60 + 4.5 ↑ LongCLIP 76.30 76.03 77.21 84.50 + 7.3 ↑ T5 Cos 54.47 57.21 55.13 69.50 + 12.3 ↑ SBER T 52.06 52.52 52.92 67.92 + 15.0 ↑ LLM-based LLM Judge (%) 55.53 53.03 59.51 69.94 + 10.4 ↑ AgriMM Lexical / Math Metrics BLEU-4 14.13 4.24 2.29 49.34 + 35.2 ↑ ROUGE-2 24.61 11.84 9.24 57.29 + 32.7 ↑ METEOR 37.89 32.43 29.11 66.70 + 28.8 ↑ Semantic Metrics BER TScore 88.52 87.02 85.83 94.71 + 6.2 ↑ Continued on next page . . . 28 Dataset Metric LLaV A-OneVision Llama-3.2 Qwen-2.5 AgriChat Di ff (7B) (11B) (7B) (7B) (vs Best) LongCLIP 87.05 86.10 0.00 93.97 + 6.9 ↑ T5 Cos 77.23 74.58 74.02 90.68 + 13.5 ↑ SBER T 67.79 66.41 65.47 83.60 + 15.8 ↑ LLM-based LLM Judge (%) 55.12 57.18 65.77 77.43 + 11.7 ↑ 4.5.2. Infer ence P erformance T o assess real-world deployment viability , we benchmarked AgriChat on a consumer-grade NVIDIA R TX 3090 GPU (24GB VRAM), representati ve of hard- ware accessible to agricultural research institutions and extension services. All models were e v aluated using 4-bit quantization, batch size of 1, and a standard- ized test image (1328 KB) with the prompt “What is the name of the plant in the image?” repeated across 100 inference cycles. T able 3: Inference performance benchmarks on NVIDIA R TX 3090 (24GB VRAM). All models ev aluated at 4-bit precision with batch size 1 on a 1328 KB agricultural field image. Metrics av eraged ov er 100 repetitions. Bold indicates best performance; Underline indicates second best. Model Load Time (s) A vg Inf Time (s) It / sec T okens / sec Mem Load (GB) Mem Peak (GB) Llama-3.2 (V ision-11B) 29.05 9.938 0.101 25.86 7.18 8.16 Qwen-2.5 (VL-7B) 22.45 29.769 0.034 8.60 11.33 11.76 LLaV A-OneV ision (7B) 9.98 1.546 0.647 12.94 9.99 10.99 AgriChat (7B) 9.27 2.315 0.432 9.50 10.71 12.32 As shown in T able 3, AgriChat achie ves an av erage inference time of 2.315 seconds per query , enabling real-time diagnostic applications while remaining 4.3 × faster than Llama-3.2-11B and 12.9 × faster than Qwen-2.5-7B. The model’ s throughput of 0.432 iterations per second translates to approximately 1,555 diag- nostic queries per hour , su ffi cient for field deployment via mobile applications. W ith a memory footprint of 10.71–12.32 GB and rapid load time of 9.27 seconds, AgriChat runs comfortably on mid-range GPUs (R TX 4060 T i, A4000) without requiring specialized data center hardware. While the 50% latency increase rela- ti ve to base LLaV A-OneV ision reflects the additional computational cost of spe- cialized adapters, this trade-o ff deliv ers a 21.4 percentage point improvement in diagnostic accuracy on AgriMM (T able 2), making it acceptable for agricultural applications where misdiagnosis carries significant economic consequences. 4.6. Discussion 4.6.1. Generalization to Unseen Domains A critical finding is the model’ s ability to generalize to datasets not seen dur- ing fine-tuning. Despite being trained exclusi vely on AgriMM, AgriChat achiev ed 29 strong performance on CDDM (Diagnosis), leading all baselines across both lex- ical and semantic metrics by substantial margins (e.g., BLEU-4: + 5.9%, LLM Judge: + 10.4%). On PlantV illageVQA, AgriChat led on lexical metrics and LLM-judged quality ( + 16.9%), though it trailed LLaV A-OneV ision on semantic similarity metrics (BER TScore, SBER T), suggesting its outputs are structurally well-formed but phrased di ff erently from reference answers. T ogether , these re- sults suggest that training on AgriMM’ s div erse 63-source corpus helps prev ent ov erfitting to specific visual conditions, enabling the model to learn robust repre- sentations of crop pathology features across di ff erent data distributions. 4.6.2. The Specialist-Generalist T rade-o ff The results highlight an inherent trade-o ff . While AgriChat excels in visual diagnosis, generalist baselines retained an advantage in tasks requiring broad, open-ended kno wledge retriev al, such as A GMMU (Open-Ended). AgriMM’ s training pipeline prioritizes “V ision-to-V erified-Knowledge, ” creating a specialist diagnostician . Consequently , the model is highly accurate at identification but conserv ati ve in generating treatment protocols where its training data did not pro- vide verified advice. This behavior acts as a safety mechanism, reducing the risk of hallucinating management strategies in high-stak es agricultural contexts. 4.6.3. Metric Sensitivity and Alignment The discrepancy between le xical and semantic metrics in T able 2 v alidates our multi-faceted e valuation approach. Generalist models achiev ed near-zero BLEU- 4 scores on structured tasks such as A GMMU (MCQs) (e.g., LLaV A-OneV ision: 0.0420, Llama-3.2: 0.0249), reflecting v erbose or ill-formatted outputs, despite maintaining moderate BER TScore v alues. On structured diagnostic datasets (A G- MMU MCQs, CDDM, AgriMM), AgriChat achie v es consistently high scores across both lexical and semantic metrics, demonstrating that domain-specific fine- tuning successfully aligns both output content and format . 4.7. Ablation Studies W e conduct a series of controlled ablation e xperiments to isolate the contrib u- tion of each design decision in our fine-tuning frame work. Specifically , we e xam- ine three ax es of v ariation: (i) the rank of the LoRA adapter matrices, (ii) whether the vision encoder benefits from lightweight adaptation alongside the language de- coder , and (iii) single-stage versus two-stage curriculum training. All fine-tuning runs and e v aluations are carried out on our dataset AgriMM. Unless otherwise 30 stated, each e xperiment varies a single factor while holding all remaining hyperpa- rameters at their baseline v alues. Results are reported across three complementary e v aluation dimensions: lexical ov erlap (BLEU-4, R OUGE-2, METEOR), seman- tic similarity (BER TScore, LongCLIP , T5 Cosine, SBER T), and holistic judge- ment quality (LLM Judge). 4.7.1. E ff ect of LoRA Rank The rank r of the lo w-rank adapter matrices gov erns the capacity av ailable for domain adaptation: an insu ffi ciently lo w rank risks underfitting the special- ized v ocab ulary and reasoning patterns of agricultural advisory dialogue, whereas an excessiv ely high rank in vites overfitting giv en the limited size of the train- ing corpus. W e ev aluate r ∈ { 32 , 64 , 128 } , maintaining the relationship α = 2 r throughout so that the e ff ecti v e learning-rate scaling remains constant across con- figurations. As shown in T able 4, increasing the rank from 32 to 64 yields sub- stantial g ains across all metrics, with BLEU-4 impro ving by roughly 10% relati v e and LLM Judge accurac y rising by ov er 3 percentage points. Further doubling the rank to 128 produces marginal additional improvements on the semantic metrics while leaving LLM Judge essentially unchanged (76.39% vs. 76.54%). These re- sults suggest that r = 64 captures the bulk of the adaptation benefit; nonetheless, we adopt r = 128 for subsequent experiments to maximise headroom for the more challenging ablation conditions that follo w . T able 4: E ff ect of LLM LoRA rank on AgriMM. Bold denotes the best result per metric; underline denotes the second best. Metric Rank 32 Rank 64 Rank 128 Lexical BLEU-4 0.4422 0.4860 0.4860 R OUGE-2 0.5146 0.5598 0.5663 METEOR 0.6172 0.6599 0.6588 Semantic BER TScore 0.9377 0.9447 0.9461 LongCLIP 0.9258 0.9365 0.9378 T5 Cos 0.8850 0.9027 0.9035 SBER T 0.8064 0.8302 0.8317 LLM-based LLM Judge (%) 73.46 76.54 76.39 31 4.7.2. E ff ect of V ision Encoder Adaptation In the default configuration the SigLIP vision encoder remains fully frozen and LoRA adapters are injected exclusi vely into the Qwen-2 language decoder . This design assumes that the general-purpose visual features learned during large-scale pre-training transfer adequately to the agricultural domain. T o test this assump- tion, we introduce an additional set of rank-32 LoRA adapters into the vision encoder’ s linear projection layers and retrain under otherwise identical condi- tions (LLM rank 128, single-stage training). T able 5 reports the results. Adapt- ing the vision encoder yields consistent improv ements across ev ery metric, with the lar gest relati ve gains observed on the le xical measures (BLEU-4: + 1 . 52%; R OUGE-2: + 1 . 17%) and on the LLM Judge score ( + 1 . 36%). The semantic simi- larity metrics also improv e, albeit more modestly (BER TScore: + 0 . 11%; SBER T : + 0 . 52%). These findings indicate that domain-specific visual representations con- stitute a meaningful, if secondary , performance bottleneck: fine-grained agri- cultural features such as lesion morphology , growth-stage appearance, and plant anatomical detail benefit from lightweight encoder adaptation be yond what frozen pre-trained features provide. T able 5: E ff ect of vision encoder adaptation on AgriMM. Both configurations use LLM rank 128 with single-stage training. Metric w / o Vision LoRA w / Vision LoRA Lexical BLEU-4 0.4860 0.4934 R OUGE-2 0.5663 0.5729 METEOR 0.6588 0.6670 Semantic BER TScore 0.9461 0.9471 LongCLIP 0.9378 0.9397 T5 Cos 0.9035 0.9068 SBER T 0.8317 0.8360 LLM-based LLM Judge (%) 76.39 77.43 4.7.3. E ff ect of T raining Strate gy Our default protocol trains the model in a single stage on the full multimodal dataset. W e compare this against a two-stage curriculum motiv ated by the hy- 32 pothesis that establishing linguistic grounding before introducing visual alignment may yield more stable optimisation. In Stage 1, LoRA adapters are applied solely to the language decoder and trained on text-only agricultural QA pairs, allo wing the model to internalise domain terminology , disease nomenclature, and advisory reasoning patterns. In Stage 2, adapters are additionally introduced into the vi- sion encoder and training proceeds on the complete multimodal corpus. Contrary to expectation, the two-stage curriculum substantially underperforms the single- stage baseline across all ev aluation axes (T able 6). The de gradation is particu- larly pronounced on lexical metrics, where BLEU-4 drops by 15.12% relati ve and R OUGE-2 by 16.32%. Semantic scores and the LLM Judge similarly decline. W e attribute this to catastrophic for getting during the stage transition: the adapter weights optimised for te xt-only supervision in Stage 1 provide a suboptimal ini- tialisation for the multimodal objecti v e in Stage 2, e ff ecti vely undoing a portion of the linguistic adaptation. These results firmly justify the simpler single-stage pro- tocol, which benefits from joint vision–language optimisation from the outset. T able 6: E ff ect of training strate gy on AgriMM. Both configurations use LLM rank 128 and vision rank 32. Metric Single-stage T wo-stage Lexical BLEU-4 0.4934 0.4286 R OUGE-2 0.5729 0.4925 METEOR 0.6670 0.6103 Semantic BER TScore 0.9471 0.9364 LongCLIP 0.9397 0.9274 T5 Cos 0.9068 0.8869 SBER T 0.8360 0.8057 LLM-based LLM Judge (%) 77.43 73.26 5. Conclusion This work addressed two fundamental bottlenecks limiting the deployment of MLLMs in agriculture: the scarcity of scientifically verified training data and the lack of domain-specialized models capable of fine-grained agricultural reasoning. 33 T o overcome the first, we introduced the V ision-to-V erified-Knowledge (V2VK) pipeline, which synthesizes high-quality VQA pairs by grounding the outputs of multiple generativ e models in v erified phytopathological literature. This produced AgriMM, a publicly av ailable benchmark of 121,425 images and 607,125 VQA pairs spanning more than 3,000 agricultural classes across 63 source datasets, v al- idated through a rigorous human-in-the-loop protocol. T o ov ercome the second, we presented AgriChat, a domain-specialized MLLM b uilt on LLaV A-OneV ision and adapted through parameter-e ffi cient fine-tuning of both the vision encoder and language decoder . AgriChat achiev es state-of-the-art in-domain performance in both internal and external benchmarks and strong zero-shot generalization, outper- forming open-source models, while w orking on consumer -grade hardware. Future work will focus on three directions: expanding AgriMM to cov er pest identifica- tion, increasing per-class image representation to support more robust learning, and incorporating more recent generativ e models into both the V2VK pipeline and the AgriChat architecture. All data, code, and model weights are publicly released to support reproducibility and to serve as a foundation for future research in agricultural multimodal intelligence. 34 A ppendix A. Dataset Composition This section details the specific source datasets aggregated to construct AgriMM. T o ensure reproducibility , we list the e xact dataset identifiers used in our aggrega- tion pipeline. Appendix A.1. Cr op Counting and Spatial Reasoning Sour ces The 33 object detection datasets used to construct the quantitativ e reasoning component of AgriMM are listed below . These datasets were processed to gener- ate ground-truth counts for instruction tuning. • Fruit & Nut Detection: almond_bloom_2023 [34], almond_harvest_2021 [34], apple_detection_drone_brazil [34], apple_detection_spain [34], apple_detection_usa [34], embrapa_wgisd_grape_detection [34], fruit_detection_w orldwide [34], grape_detection_california day [34], grape_detection_californianight [34], grape_detection_syntheticday [34], mango_detection_australia [34], Orange_dataset, stra wberry_detection_2022 [34], strawberry_detection_2023 [34], tomato_ripeness_detection [34], wGrapeUNIPD- DL [51]. • Field Crops & V egetables: ghai_broccoli_detection [34], ghai_green_cabbage_detection [34], ghai_iceberg_lettuce_detection [34], ghai_romaine_detection [34], GWHD2021 (Global Wheat Head Detection) [52], wheat_head_counting [34]. • Specialized & General Agriculture: CBD A [53, 54], DRPD [55], gemini_flower_detection_2022 [34], gemini_leaf_detection_2022 [34], gemini_plant_detection_2022 [34], gem- ini_pod_detection_2022 [34], MTDC [56], plant_doc_detection [34], SHC [57], WEDD [58], Y OLOPOD [59]. Appendix A.2. Disease and Str ess Classification Sour ces The 29 classification datasets comprising the pathological component of the benchmark are listed belo w . Each dataset includes a “Healthy” baseline class alongside specific biotic and abiotic stress classes. arabica_co ff ee_leaf_disease_classification [34], banana_leaf_disease_classification [34], bean_disease_uganda [34], betel_leaf_disease_classificatio n [34], blackgram_plant_leaf_disease_classification [34], chilli_leaf_classification [34], coconut_tree_disease_classification [34], corn_maize_leaf_disease [34], crop_weeds_greece [34], cucumber_disease_classification [34], gua v a_disease_pakistan [34], jav a_plum_leaf_disease_classifi cation [34], leaf_counting_denmark [34], onion_leaf_classification [34], orange_leaf_disease_classification [34], paddy_disease_classification [34], papaya_leaf_disease_classification [34], plant_doc_classification [34], plant_seedlings_aarhus [34], plant_village_classification 35 [34], rangeland_weeds_australia [34], rice_leaf_disease_classification [34], rise- holme_strawberry_classification_2021 [34], so ybean_weed_uav_brazil [34], sug- arcane_damage_usa [34], sunflower_disease_classification [34], tea_leaf_disease_classification [34], tomato_leaf_disease [34], vine_virus_photo_dataset [34]. A ppendix B. Synthesis Pipeline Prompt T emplates W e provide the exact system prompts used in our V ision-to-V erified-Knowledge pipeline. These prompts were designed to minimize hallucination by enforcing strict constraints on the language models. Appendix B.1. Stage 1: V isual Gr ounding (Ima ge Captioning) Model: Gemma 3 (12B) Purpose: T o generate structured visual metadata rather than generic captions. Write a descriptive caption of about 3-5 sentences given that the image contains {extra_deta ils}. Include these aspects if clearly visible: - Crop name and type - Growth stage (seedling/veget ative/flowering/fruiti ng/harvest) - Ground cover and plant density - Image perspective (top-down/ oblique/side/macro/unk nown) - Environmental conditions (fi eld/greenhouse/laborat ory) - Plant health indicators Rules for caption: - Use clear, neutral language - No speculation - only describe what is visible - If something cannot be determined, use ’unknown’ - Write as natural sentences Appendix B.2. Stage 2: Knowledge Retrieval Model: Gemini 3 Pro (W eb Search Enabled) Purpose: T o retriev e v erified botanical and phytopathological kno wledge. Appendix B.2.1. Species Identification Pr ompt For the class name {class_names}, generate a detailed botanical description paragraph (~300 words) covering: - Taxonomic classification (fa mily, genus, species) - Morphological characteristics (leaf shape, stem structure, 36 inflorescence, fruit morphology) - Native habitat and biogeographic distribution - Cultivation requirements (so il, climate, water) - Ecological significance and agricultural use Format your output as {"class_name":"detailed discription"} Appendix B.2.2. Disease Classification Pr ompt For each class name in {disease_class_names}, generate a detailed paragraph (~300 words ) providing an integrated account of: - Plant taxonomy, morphology, and natural habitat - Disease etiology (causal agent, pathogen taxonomy) - Visible symptoms (lesion morphology, discoloration patterns, necrosis, wilting) - Affected plant organs (leaves, stems, fruits, roots) - Pathogen biology and infection cycle - Environmental factors influe ncing disease development - Comparison with healthy plant phenotype When the class represents "Healthy", describe the ideal botanical state emphasizing vi gor, normal morphology, and optimal appearance. Appendix B.3. Stage 3: Instruction Generation Model: LLaMA 3.1 8B Instruct Purpose: T o synthesize visual captions and retriev ed kno wledge into QA pairs. You are an expert agricultural AI trainer. Generate exactly 5 high-quality, diverse QA pairs. **SOURCE DATA:** - Additional Info: {class_info } - Image Caption: {caption} **STRICT RULES:** 1. GROUNDING: Use ONLY provided info. If the image/info doesn’t mention a disease, don’t invent one. 2. FORMAT: Output a single JSON array of 5 objects. 3. ANSWER STYLE: Use full, professional sentences. Instead of "okra," say "The image shows an okra plant (Abelmoschus esculentus)." 37 **REQUIRED QUESTION CATEGORIES (One per slot):** 1. Identification: Identify th e plant and its variety. 2. Visual Reasoning: Ask HOW the plant can be identified (e.g., "What visual features distinguish this species?"). 3. Condition & Health: Ask about leaf/fruit/stem state (color, spots, growth stage). 4. Cultivation Knowledge: Conn ect visuals to agronomic requirements (e.g., "What are this plant’s soil pH needs?"). 5. Anatomy/Detail: Ask about a specific visible part (flower, fruit, leaf structure) . A ppendix C. Evaluation Pr ompts Appendix C.1. LLM-as-a-J udge Pr ompt Model: Qwen3-30B-A3B-Instruct-2507 Purpose: T o e v aluate response quality while penalizing verbosity and hallucina- tion. You are an expert evaluator assessing an AI model’s response. Evaluate systematically and ob jectively. **QUESTION**: {question} **GROUND TRUTH (Correct Answer )**: {ground_truth} **MODEL OUTPUT (To Evaluate)** : {model_output} --- **EVALUATION CRITERIA** 1. **Correctness**: Does the output contain the correct information from the Ground Truth? 2. **Completeness**: Does the output include all important information from Ground Truth? 3. **Clarity**: Is the output clear, well-organized, and easy to understand? 4. **Conciseness**: Is the output appropriately concise without unnecessary content? --- 38 **SCORING RUBRIC** (1-4 scale) **Score 1 (Poor)**: Major deficiencies. Factually incorrect or missing multiple key facts. **Score 2 (Fair)**: Significan t issues. Missing 1-2 important facts or minor inaccuracies. **Score 3 (Good)**: Solid with minor issues only. Factually accurate but maybe slightly verbose or misses tiny details. **Score 4 (Excellent)**: Outst anding quality. Perfectly accurate, complete, clear, and concise. --- **OUTPUT**: Provide evaluation in this EXACT JSON format: {{ "score": , "justification": "1-2 sentence summary" }} Begin evaluation: References [1] A. Dubey , A. Jauhri, A. Pandey , A. Kadian, A. Al-Dahle, A. Letman, A. Mathur , A. Schelten, A. Y ang, A. F an, et al., The llama 3 herd of models, arXi v e-prints (2024) arXi v–2407. [2] B. Li, Y . Zhang, D. Guo, R. Zhang, F . Li, H. Zhang, K. Zhang, P . Zhang, Y . Li, Z. Liu, et al., Lla v a-one vision: Easy visual task transfer , arXi v preprint arXi v:2408.03326 (2024). [3] A. Y ang, et al., Qwen2 technical report, arXiv preprint arXi v:2407.10671 (2024). [4] S. N. Sakib, N. Haque, M. Z. Hossain, S. E. Arman, Plantvillage vqa: A visual question answering dataset for benchmarking vision-language models in plant science, arXi v preprint arXi v:2508.17117 (2025). [5] A. Gauba, I. Pi, Y . Man, Z. Pang, V . S. Adve, Y .-X. W ang, Agmmu: A comprehensiv e agricultural multimodal understanding benchmark, 2025. 39 arXiv:2504.10568 . URL [6] W orld Bank, Agriculture and food: Overvie w, accessed: 2024-10-24 (2024). URL https://www.worldbank.org/en/topic/agriculture/ overview [7] International Fund for Agricultural De velopment, Ifad annual report 2024 (2024). URL https://www.ifad.org/en/annual- report- 2024 [8] F A O, The State of F ood Security and Nutrition in the W orld 2024: Financing to End Hunger , F ood Insecurity and Malnutrition in All Its F orms, F ood and Agriculture Org anization of the United Nations, Rome, 2024. [9] A. O. Adewusi, O. F . Asuzu, T . Olorunsogo, C. Iwuanyanwu, E. Adaga, D. O. Daraojimba, Ai in precision agriculture: A revie w of technologies for sustainable f arming practices, W orld Journal of Adv anced Research and Re- vie ws 21 (1) (2024) 2276–2285. doi:10.30574/wjarr.2024.21.1.0314 . [10] L. Zhang, H. W ang, Y . Li, Adv anced artificial intelligence: A rev olution for sustainable agriculture, Nature Sustainability 8 (2) (2025) 112–125. [11] P . Kumar , R. Singh, Ai-dri ven support tools for crop yield optimization: A re vie w , Smart Agricultural T echnology 7 (2024) 100391. [12] M. El Jarroudi, L. Kouadio, B. Mercatoris, et al., Le veraging edge artificial intelligence for sustainable agriculture, Nature Sustainability 7 (2024) 846– 854. doi:10.1038/s41893- 024- 01352- 4 . [13] L. Nieradzik, H. Stephani, J. Siebur g-Rockel, S. Helmling, A. Olbrich, J. K e- uper , Challenging the black box: A comprehensive e v aluation of attribu- tion maps of cnn applications in agriculture and forestry , arXiv preprint arXi v:2402.11670 (2024). [14] M. A wais, A. H. S. A. Alharthi, A. Kumar , H. Cholakkal, R. M. Anwer , Agrogpt: E ffi cient agricultural vision-language model with expert tuning, in: 2025 IEEE / CVF W inter Conference on Applications of Computer V ision (W A CV), IEEE, 2025, pp. 5687–5696. 40 [15] S. Y in, Y . Xi, X. Zhang, C. Sun, Q. Mao, Foundation models in agriculture: A comprehensi ve re vie w , Agriculture 15 (8) (2025) 847. doi:10.3390/ agriculture15080847 . [16] D. P . Hughes, M. Salathé, An open access repository of images on plant health to enable the dev elopment of mobile disease diagnostics, arXiv preprint arXi v:1511.08060 (2015). [17] X. Liu, Z. Liu, H. Hu, Z. Chen, K. W ang, K. W ang, S. Lian, A multi- modal benchmark dataset and model for crop disease diagnosis, in: Com- puter V ision – ECCV 2024, Springer Nature Switzerland, 2024, p. 157–170. doi:10.1007/978- 3- 031- 73016- 0_10 . URL http://dx.doi.org/10.1007/978- 3- 031- 73016- 0_10 [18] G. T eam, A. Kamath, J. Ferret, S. P athak, N. V ieillard, R. Merhej, S. Perrin, T . Matejovicov a, A. Ramé, M. Rivière, et al., Gemma 3 technical report, arXi v preprint arXi v:2503.19786 (2025). [19] Gemini T eam, Google, Gemini 3: Frontier multimodal intelligence, T echni- cal report, Google DeepMind (2025). URL https://deepmind.google/technologies/gemini/ [20] L. W ang, T . Jin, J. Y ang, A. Leonardis, F . W ang, F . Zheng, Agri-lla v a: Kno wledge-infused lar ge multimodal assistant on agricultural pests and dis- eases, arXi v preprint arXi v:2412.02158 (2024). [21] K. Sharma, V . V ats, A. Singh, R. Sahani, D. Rai, A. Sharma, Llav a- plantdiag: Integrating large-scale vision-language abilities for con versa- tional plant pathology diagnosis, in: 2024 International Joint Conference on Neural Networks (IJCNN), IEEE, 2024, pp. 1–7. [22] Y . Lu, S. Y oung, A survey of public datasets for computer vision tasks in precision agriculture, Computers and Electronics in Agriculture 178 (2020) 105760. [23] E. J. Hu, Y . Shen, P . W allis, Z. Allen-Zhu, Y . Li, S. W ang, L. W ang, W . Chen, LoRA: Lo w-rank adaptation of large language models, in: International Conference on Learning Representations, 2022. URL https://openreview.net/forum?id=nZeVKeeFYf9 41 [24] D. Hughes, M. Salathé, et al., An open access repository of images on plant health to enable the development of mobile disease diagnostics, arXiv preprint arXi v:1511.08060 (2015). [25] Y . Zhang, Y . Li, L. Cui, D. Cai, L. Liu, T . Fu, X. Huang, E. Zhao, Y . Zhang, Y . Chen, et al., Siren’ s song in the ai ocean: A survey on hallucination in large language models, Computational Linguistics 51 (4) (2025) 1373–1418. [26] M. L. Gullino, R. Albajes, I. Al-Jboory , F . Angelotti, S. Chakraborty , K. Gar - rett, B. Hurley , P . Juroszek, K. Makkouk, X. Pan, et al., Scientific revie w of the impact of climate change on plant pests: A global challenge to prev ent and mitigate plant-pest risks in agriculture, forestry and ecosystems, T ech. rep. (2021). [27] K. He, X. Zhang, S. Ren, J. Sun, Deep residual learning for image recogni- tion, in: Proceedings of the IEEE conference on computer vision and pattern recognition, 2016, pp. 770–778. [28] G. Huang, Z. Liu, L. V an Der Maaten, K. Q. W einberger , Densely connected con v olutional networks, in: Proceedings of the IEEE conference on com- puter vision and pattern recognition, 2017, pp. 4700–4708. [29] M. T an, Q. Le, E ffi cientnet: Rethinking model scaling for con v olutional neural networks, in: International conference on machine learning, PMLR, 2019, pp. 6105–6114. [30] J. Lin, X. Zhang, Y . Qin, S. Y ang, X. W en, T . Cernav a, Q. Migheli, X. Chen, Local and global feature-a ware dual-branch networks for plant dis- ease recognition, Plant Phenomics 6 (2024) 0208. [31] S. Y u, L. Xie, L. Dai, St-cfi: Swin transformer with con volutional feature interactions for identifying plant diseases, Scientific Reports 15 (1) (2025) 25000. [32] Z. Liu, Y . Lin, Y . Cao, H. Hu, Y . W ei, Z. Zhang, S. Lin, B. Guo, Swin trans- former: Hierarchical vision transformer using shifted windo ws, in: Proceed- ings of the IEEE / CVF international conference on computer vision, 2021, pp. 10012–10022. 42 [33] U. Nawaz, A. Muhammad, H. Gani, M. Naseer , F . S. Khan, S. Khan, R. Anwer , Agriclip: Adapting clip for agriculture and liv estock via domain- specialized cross-model alignment, in: Proceedings of the 31st International Conference on Computational Linguistics, 2025, pp. 9630–9639. [34] A. Joshi, D. Guev ara, M. Earles, Standardizing and centralizing datasets for e ffi cient training of agricultural deep learning models, Plant Phenomics 5 (2023) 0084. doi:https://doi.org/10.34133/plantphenomics.0084 . URL https://www.sciencedirect.com/science/article/pii/ S2643651524001432 [35] B. Y ang, Y . Chen, L. Feng, Y . Zhang, X. Xu, J. Zhang, N. Aierken, R. Huang, H. Lin, Y . Y ing, S. Li, Agrigpt-vl: Agricultural vision-language understand- ing suite (2025). . URL [36] X. Zhai, B. Mustaf a, A. K olesnikov , L. Beyer , Sigmoid loss for language im- age pre-training, in: Proceedings of the IEEE / CVF International Conference on Computer V ision, 2023, pp. 11975–11986. [37] E. Reiter , A structured re vie w of the v alidity of bleu, Computational Lin- guistics 44 (3) (2018) 393–401. [38] Y . Liu, D. Iter , Y . Xu, S. W ang, R. Xu, C. Zhu, G-e v al: Nlg e v aluation using gpt-4 with better human alignment, in: Proceedings of the 2023 Conference on Empirical Methods in Natural Language Processing (EMNLP), 2023, pp. 2511–2522. [39] K. Papineni, S. Roukos, T . W ard, W .-J. Zhu, Bleu: A method for automatic e v aluation of machine translation, in: Proceedings of the 40th Annual Meet- ing of the Association for Computational Linguistics, Association for Com- putational Linguistics, 2002, pp. 311–318. [40] C.-Y . Lin, Rouge: A package for automatic ev aluation of summaries, in: T ext summarization branches out, 2004, pp. 74–81. [41] S. Banerjee, A. Lavie, Meteor: An automatic metric for mt ev aluation with improv ed correlation with human judgments, in: Proceedings of the A CL W orkshop on Intrinsic and Extrinsic Ev aluation Measures for Machine T ranslation and / or Summarization, 2005, pp. 65–72. 43 [42] A. Nainia, Llm e v aluation is broken: Why bleu and rouge don’t measure real understanding, Medium (2024). URL https://medium.com/@nainia_ayoub/ llm- evaluation- is- broken [43] T . Zhang, V . Kishore, F . W u, K. Q. W einberger , Y . Artzi, Bertscore: Evalu- ating text generation with bert, 2020. . URL [44] A. Radford, J. W . Kim, C. Hallac y , A. Ramesh, G. Goh, S. Agarw al, G. Sas- try , A. Askell, P . Mishkin, J. Clark, et al., Learning transferable visual mod- els from natural language supervision, in: International conference on ma- chine learning, PMLR, 2021, pp. 8748–8763. [45] C. Ra ff el, N. Shazeer , A. Roberts, K. Lee, S. Narang, M. Matena, Y . Zhou, W . Li, P . J. Liu, Exploring the limits of transfer learning with a unified text- to-text transformer (2023). . URL [46] N. Reimers, I. Gure vych, Sentence-bert: Sentence embeddings using siamese bert-networks, 2019. . URL [47] L. Zheng, W .-L. Chiang, Y . Sheng, S. Zhuang, Z. W u, Y . Zhuang, Z. Lin, Z. Li, D. Li, E. P . Xing, H. Zhang, J. E. Gonzalez, I. Stoica, Judging llm-as- a-judge with mt-bench and chatbot arena, 2023. . URL [48] H. Li, Q. Dong, J. Chen, H. Su, Y . Zhou, Q. Ai, Z. Y e, Y . Liu, Llms-as- judges: A comprehensi ve surve y on llm-based e v aluation methods (2024). arXiv:2412.05579 . URL [49] J. Gu, X. Jiang, Z. Shi, H. T an, X. Zhai, C. Xu, W . Li, Y . Shen, S. Ma, H. Liu, S. W ang, K. Zhang, Y . W ang, W . Gao, L. Ni, J. Guo, A survey on llm-as-a-judge (2025). . URL [50] S. Han, G. T . Junior , T . Balough, W . Zhou, Judge’ s v erdict: A comprehensiv e analysis of llm judge capability through human agreement, arXi v preprint arXi v:2510.09738 (2025). 44 [51] M. Sozzi, S. Cantalamessa, A. Cogato, A. Kayad, F . Marinello, wGrapeUNIPD-DL: an open dataset for white grape bunch detection (2022). doi:10.5281/zenodo.4066730 . URL https://doi.org/10.5281/zenodo.4066730 [52] E. David, M. Serouart, D. Smith, S. Madec, K. V elumani, S. Liu, X. W ang, F . Pinto, S. Shafiee, I. S. T ahir , H. Tsujimoto, S. Nasuda, B. Zheng, N. Kirchgessner , H. Aasen, A. Hund, P . Sadhegi-T ehran, K. Nagasa wa, G. Ishikawa, S. Dandrifosse, A. Carlier , B. Dumont, B. Mercatoris, B. Evers, K. Kuroki, H. W ang, M. Ishii, M. A. Badhon, C. Pozniak, D. S. LeBauer , M. Lillemo, J. Poland, S. Chapman, B. de Solan, F . Baret, I. Stavness, W . Guo, Global wheat head detection 2021: An improv ed dataset for benchmarking wheat head detection methods, Plant Phenomics 2021 (2021) 9846158. doi:https://doi.org/10.34133/2021/9846158 . URL https://www.sciencedirect.com/science/article/pii/ S2643651524000591 [53] D. Lu, J. Y e, Y . W ang, Z. Y u, Plant detection and counting: Enhancing preci- sion agriculture in ua v and general scenes, IEEE Access 11 (2023) 116196– 116205. doi:10.1109/ACCESS.2023.3325747 . [54] J. Y e, Z. Y u, Y . W ang, D. Lu, H. Zhou, Wheatlfanet: In-field detection and counting of wheat heads with high-real-time global regression network, Plant Methods 19 (103) (2023). doi:10.1186/s13007- 023- 01079- x . [55] Z. T eng, J. Chen, J. W ang, S. W u, R. Chen, Y . Lin, L. Shen, R. Jackson, J. Zhou, C. Y ang, Panicle-cloud: An open and ai-powered cloud computing platform for quantifying rice panicles from drone-collected imagery to en- able the classification of yield production in rice, Plant Phenomics 5 (2023) 0105. doi:10.34133/plantphenomics.0105 . URL https://doi.org/10.34133/plantphenomics.0105 [56] H. Zou, H. Lu, Y . Li, L. Liu, Z. Cao, Maize tassels detection: a benchmark of the state of the art, Plant Methods 16 (1) (2020) 1–15. [57] H. Lu, Z. Cao, T asselnetv2 + : A fast implementation for high-throughput plant counting from high-resolution rgb imagery, Frontiers in Plant Science V olume 11 - 2020 (2020). doi:10.3389/fpls.2020.541960 . URL https://www.frontiersin.org/journals/plant- science/ articles/10.3389/fpls.2020.541960 45 [58] S. Madec, X. Jin, H. Lu, B. De Solan, S. Liu, F . Duyme, E. Heritier , F . Baret, Ear density estimation from high resolution RGB imagery using deep learn- ing technique, Agricultural and Forest Meteorology 264 (2019) 225–234. doi:10.1016/j.agrformet.2018.10.013 . [59] S. Xiang, S. W ang, M. Xu, W . W ang, W . Liu, Y olo pod: a fast and accu- rate multi-task model for dense so ybean pod counting, Plant methods 19 (1) (2023) 8. 46
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment