Accelerating Stroke MRI with Diffusion Probabilistic Models through Large-Scale Pre-training and Target-Specific Fine-Tuning
Purpose: To develop a data-efficient strategy for accelerated MRI reconstruction with Diffusion Probabilistic Generative Models (DPMs) that enables faster scan times in clinical stroke MRI when only limited fully-sampled data samples are available. …
Authors: Yamin Arefeen, Sidharth Kumar, Steven Warach
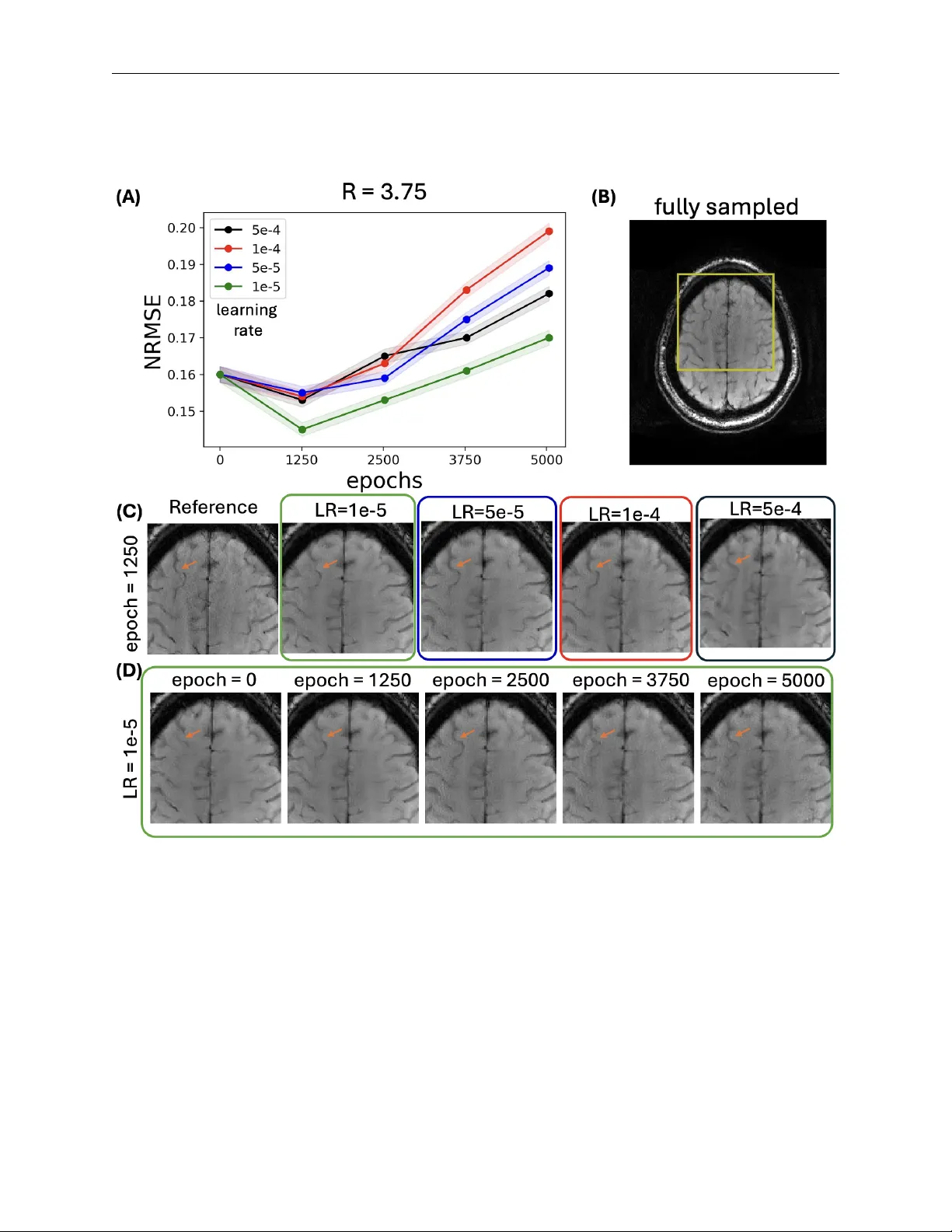
A ccelerating Strok e MRI with Diffusion Probabilistic Mo d- els through Large-Scale Pre-training and T arget-Sp ecific Fine-T uning Y amin Arefeen 1,2, † , Sidharth Kumar 1, † , Steven W arach 3 , Hamidreza Sab er 3,4 , Jonathan I. T amir 1,5,6 † These authors con tributed equally to this w ork. 1 Chandra F amily Departmen t of Electrical and Computer Engineering The Univ ersity of T exas at Austin Austin, TX, USA 2 Departmen t of Imaging Ph ysics MD Anderson Cancer Center Houston, TX, USA 3 Dell Medical School Department of Neurology The Universit y of T exas at Austin Austin, TX, USA 4 Dell Medical School Department of Neurosurgery The Universit y of T exas at Austin Austin, TX, USA 5 Dell Medical School Department of Diagnostic Medicine The Universit y of T exas at Austin Austin, TX, USA 6 Oden Institute for Computational Engineering and Sciences The Universit y of T exas at Austin Austin, TX, USA * Corresp onding author: Submited to Magnetic Resonance in Medicine Name Y amin Arefeen Departmen t Chandra F amily Department of Electrical Engineering and Computer Science Institute The Universit y of T exas at Austin; Austin, TX, United States E-mail y aminarefeen@gmail.com W ord Coun t: 250 (Abstract) ∼ 5000 (b ody) Figure/T able Coun t: 10 2 / 28 Submited to Magnetic Resonance in Medicine Abstract Purp ose: T o develop a data-efficient strategy for accelerated MRI reconstruction with Diffu- sion Probabilistic Generativ e Mo dels (DPMs) that enables faster scan times in clinical strok e MRI when only limited fully-sampled data samples are av ailable. Metho ds: Our simple training strategy , inspired b y the foundation model paradigm, first trains a DPM on a large, diverse collection of publicly av ailable brain MRI data in fastMRI and then fine-tunes on a small dataset from the target application using carefully selected learning rates and fine-tuning durations. The approach is ev aluated on controlled fastMRI experiments and on clinical stroke MRI data with a blinded clinical reader study . Results: DPMs pre-trained on approximately 4000 sub jects with non-FLAIR contrasts and fine-tuned on FLAIR data from only 20 target sub jects ac hiev e reconstruction p erformance com- parable to mo dels trained with substantially more target-domain FLAIR data across m ultiple acceleration factors. Exp erimen ts rev eal that mo derate fine-tuning with a reduced learning rate yields impro ved performance, while insufficient or excessive fine-tuning degrades reconstruction qualit y . When applied to clinical stroke MRI, a blinded reader study in volving tw o neuroradi- ologists indicates that images reconstructed using the prop osed approach from 2 × accelerated data are non-inferior to standard-of-care in terms of image qualit y and structural delineation. Conclusion: Large-scale pre-training combined with targeted fine-tuning enables DPM-based MRI reconstruction in data-constrained, accelerated clinical strok e MRI. The proposed approac h substan tially reduces the need for large application-sp ecific datasets while maintaining clinically acceptable image quality , supp orting the use of foundation-inspired diffusion mo dels for accel- erated MRI in targeted applications. Keyw ords : Accelerated MRI; Diffusion probabilistic mo dels; F oundation mo dels; Strok e MRI; Clinical T ranslation 1 / 28 Submited to Magnetic Resonance in Medicine 1 In tro duction F oundation models (1) represent a p o werful paradigm in mac hine learning where models pre-trained on large, div erse datasets adapt effectiv ely to target applications in a zero-shot fashion or with fine-tuning (2). As a non-exhaustiv e list, this approach successfully applies across man y domains, including natural language pro cessing (3), vision (4), segmen tation (5), fluorescence microscopy (6), and v arious healthcare applications (7–9). Mac hine learning currently ac hieves state-of-the-art p erformance in reconstructing an MRI image from under-sampled data for man y anatom y , contrasts, and datasets (10–13). Ho wev er, these models are t ypically trained on datasets sp ecific to the application of interest and are applied without mo dification for inference. While effective given sufficient data, this approac h degrades in settings with limited data to train the mo dels for the desired application (10). As a motiv ating example, medical imaging is critical to diagnosis of ischemic stroke in patients who come to the emergency ro om presenting symptoms; many sites use computed tomograph y (CT) due to its effectiveness, widespread a v ailability , and relativ ely short scan times (14). How ever, MRI offers improv ed sensitivity when detecting isc hemic stroke and more precise lo calization of the ischemic lesion (15). Despite these adv an tages, MRI typically requires longer scan times and is more susceptible to degradation from patient motion, whic h can delay image acquisition and, consequen tly , time to treatmen t. Machine learning–based acceleration metho ds could mitigate these limitations by reducing scan duration and motion sensitivity , but limited stroke sp ecific training data precludes their application. Sev eral tec hniques ha ve b een prop osed to train mac hine learning reconstruction models in set- tings with limited, fully-sampled data. Some methods train reconstruction models on just the under-sampled scan itself by lev eraging calibration regions (16, 17), implicit regularization imp osed b y neural netw ork structure (18, 19), and self-sup ervised learning through k-space splitting (20). Ho wev er, these methods fall short of state-of-the-art reconstruction quality offered b y methods trained on large amounts of high qualit y data (10–12). Other self-sup ervised training strategies instead utilize under-sampled or corrupt images for training (21–25) but still require a large corpus of data. In addition, these metho ds implemen t end-to-end reconstructions mapping under-sampled data to images given a sp ecific acquisition mo del. Motiv ated b y data limitations in our strok e application and inspired b y the foundation mo del paradigm (26), we prop ose a metho d to train diffusion probabilistic generative mo dels (DPMs) 2 / 28 Submited to Magnetic Resonance in Medicine for accelerated MRI reconstruction given a small dataset for the contrast of in terest. Inspired by similar approaches for large language mo dels (27), our simple but effective strategy first trains a DPM on a large, public dataset of div erse MR images and then fine-tunes the DPM on the smaller, fully-sampled dataset from the target application. During fine-tuning, we select h yp erparameters to adapt the mo del effectiv ely without ov erfitting and retaining the b enefits of pre-training on a large, diverse dataset (28). W e select DPMs for the reconstruction mo del b ecause they are agnostic to the acquisition forw ard mo del (29–38). Thus unlik e end-to-end metho ds which require the larger external dataset and the smaller target dataset to be acquired with the same measuremen t mo del (sampling pattern, coil geometry , etc. . . ) (10), a DPM pre-trained on the larger dataset can be adapted to smaller target datasets with different acquisition mo dels. First, w e comprehensively ev aluate the prop osed approac h on fastMRI (11) b y initially pre- training a DPM on T 1 , T 2 , and T 1 -p ost brain scans from 4000 sub jects. T o sim ulate access to limited data on a target application, w e then finetune the DPM on FLAIR data from just 20 sub jects of fastMRI. A cross a range of acceleration factors, DPMs trained with our metho d achiev e comparable p erformance to DPMs trained with 344 target FLAIR sub jects. Our exp erimen ts also highligh t b est practices when selecting inference hyperparameters for DPM-based MRI reconstruction and sho wcase application on prosp ectiv ely under-sampled data. Next, w e apply our method to the motiv ating setting of strok e MRI. W e initially pre-train a DPM on T 1 , T 2 , T 1 -p ost, and FLAIR brain scans from approximately 4000 fastMRI sub jects and then fine-tune the DPM on data from 25 stroke patien ts from our lo cal hospital. T esting on 5 additional patien ts demonstrates that the proposed metho d enables reconstruction of SWI, MPRAGE, DWI, and FLAIR images from 2 × further under-sampled stroke data. Finally , w e conduct a clinical v alidation study in which t wo radiologists experienced in strok e MRI compared SWI, MPRA GE, DWI, and FLAIR images reconstructed from retrosp ectiv e, 2 × under-sampled data using our metho d to standard-of-care images. The reader study indicates that images reconstructed b y the DPM pre-trained on fastMRI and fine-tuned on lo cal stroke data ac hieves non-inferior image qualit y and structural delineation of anatom y relative to the standard- of-care. 3 / 28 Submited to Magnetic Resonance in Medicine 2 Metho ds 2.1 Diffusion Probabilistic Generative Mo dels for MRI Reconstruction W e mo del the multi-coil MRI acquisition as y = P F S x + n where y are the acquired measuremen ts, P is the under-sampling op erator, F is the F ourier T ransform, S is the m ulti-coil operator, n is additiv e gaussian noise, and x is the image to reconstruct. DPMs reconstruct accelerated MRI acquisitions by first learning the underlying probability distribution of the target MR contrast and anatom y , and then guiding the reconstruction tow ards solutions that b oth matc h the data and are statistically likely . Sp ecifically , w e use score-based diffusion mo dels where a neural netw ork G θ appro ximates the score function ∇ x t log p t ( x ) , where p t ( x ) denotes image distribution perturb ed b y gaussian noise with standard deviation σ t (29, 34, 35, 39). Equipp ed with the score function appro ximation, images can b e reconstructed from under-sampled data b y appro ximately sampling from the p osterior p ( x | y ) with p osterior sampling algorithms (36). In this w ork, we combine the prior sampling form ulation from Ref. (39) with Diffusion Posterior Sampling (31) to appro ximately sample from p ( x | y ) in accelerated MRI. Sp ecifically , w e solv e the follo wing ordinary differential equation (ODE): d x = ˙ s ( t ) s ( t ) x − s ( t ) 2 ˙ σ ( t ) σ ( t ) ∇ x log p x s ( t ) | y ; σ ( t ) dt. [1] Setting s ( t ) = 1 and σ ( t ) = t based on Ref. (39), applying Bay e’s rule to separate the p osterior score in to the sum of the log-likelihoo d and prior score, and inserting the DPM that approximates the prior score function and the closed-form expression for the lik eliho o d score (31, 32) results in the following ODE, d x = h − t ∇ x ∥ PFS˜ x ( x ) − y ∥ 2 2 + D θ ( x , t i dt. [2] W e apply the approximation ˜ x ( x ) = E [ x 0 | x ] since the log-likelihoo d score is only kno wn at t = 0 (31). 2.2 DPM Details All DPMs in this work use the U-Net st yle architecture from Ref. (29), as implemented b y Ref. (39), and we adopt a mo dified version of the architecture that supp orts inputs with v arying matrix sizes (40). Initial training on the external dataset and fine-tuning on the target dataset applied the “EDM” training loss, data augmentation, optimizer c hoice, and noise level scheduling. 4 / 28 Submited to Magnetic Resonance in Medicine T o accommodate datasets with heterogeneous con trasts, we simultaneously train on all a v ailable data b y conditioning the mo del on con trast type (40). Each image contrast t yp e in the training set is assigned a one-hot vector, which is passed through a small fully-connected neural netw ork to pro duce an em b edding. Eac h U-Net blo c k in the DPM netw ork then takes this em b edding vector as an input, allowing the model to adapt its b ehaviour based on the con trast. The DPM and em b edding netw ork are trained jointly . 2.3 P osterior Sampling Details Algorithm 1 outlines the p osterior sampling pro cedure utilized in this w ork for accelerated MRI reconstruction adapted from the ODE prior sampling pro cedure used in Ref. (39) and the log- lik eliho o d approximation prop osed by Ref. (31). The noise lev els, ˆ σ i follo w the recommended sc hedule from Ref. (39), and N denotes the n umber of sampling steps. Since this posterior sampling is approximate, ζ i is introduced as a tunable hyperparameter that balances data consistency and learned prior score at eac h iteration. Algorithm 1 Diffusion Posterior Sampling (DPS) Require: G θ ( x ; σ ) , N , { ζ N − 1 i =0 } , { ˆ σ i } N − 1 i =0 sample x t 0 ∼ N (0 , I ) for i ∈ { 0 , ..., N − 1 } do ˆ s = G θ ( x ; σ ) − x i σ ˆ x 0 = 1 √ α i ( x i + (1 − ¯ α i ˆ s ) n ∼ N (0 , I ) ˜ x p = √ α i (1 − ¯ α i − 1 ) 1 − ¯ α i x i + √ ¯ α i − 1 β i 1 − ¯ α i − 1 ˆ x 0 ˜ x DC = ∇ x i || y − A ( ˆ x 0 ) || 2 2 x i +1 = ˜ x p − ζ i ˜ x DC + σ i n end for 2.4 Prop osed T raining Strategy Inspired b y stra tegies for foundation and large language mo dels (27), w e propose a simple and effectiv e training approach to train DPMs for accelerated MRI when only limited target-domain data is a v ailable. Our strategy inv olv es tw o stages: (i) pre-training on a large, div erse external dataset, like fastMRI (11), cov ering multiple MRI con trasts; and (ii) fine-tuning on the smaller, 5 / 28 Submited to Magnetic Resonance in Medicine target dataset, up dating all mo del w eights. T o mitigate ov erfitting during fine-tuning, we reduce the learning rate b y an order of magnitude and train for a small num b er of ep ochs, corresponding to roughly 2% of pre-training time. In addition, we assign the target contrast its o wn one-hot enco ded v ector and up date the weigh ts of the embedding netw ork during fine-tuning. Figure 1 compares (A) standard and (B) the prop osed training strategies for DPMs. Figure 1: (A) The standard DPM training pip eline trains exclusively on the target-domain dataset, but if the target dataset is small, this can lead to degraded p erformance. (B) The proposed training pip eline mitigates data scarcit y by first pre-training the DPM on a large, div erse external MRI dataset, follow ed by fine-tuning on a small target-domain dataset using a reduced learning rate and limited num b er of ep o c hs. The fine-tuned mo del is then used for p osterior sampling–based reconstruction from under-sampled k-space. 2.5 Con trolled Exp erimen ts with F astMRI T o v alidate the prop osed training strategy , w e use the m ulti-coil brain dataset from F astMRI (11). Our dataset included Axial T 2 from 2678 sub jects, Axial T 1 from 498 sub jects, p ost-con trast Axial T 1 from 949 sub jects, and Axial FLAIR from 344 sub jects. W e fo cused on reconstructing Axial FLAIR images, sim ulating a lo w-data regime b y limiting the FLAIR training set to only 20 sub jects. All k-space data w ere pre-whitened and re-sized to 320 × 320 matrix size, and training w as p erformed 6 / 28 Submited to Magnetic Resonance in Medicine on fully-sampled, coil-combined images normalized to the 99th p ercen tile of the maximum absolute v alue. F or testing, 500 random Axial FLAIR slices w ere selected from held-out sub jects not used during training. W e first inv estigated the effect of learning rate and training time during fine-tuning. A DPM w as pre-trained on the larger external dataset with a learning rate of 10 − 4 for 625 , 000 ep o c hs, and then fine-tuned on the 20-sub ject FLAIR dataset with learning rates of { 5 × 10 − 4 , 1 × 10 − 4 , 5 × 10 − 5 , 5 × 10 − 5 } and ep o c hs of { 0 , 625 , 1250 , 1875 , 2500 } . In addition, we compared our prop osed training paradigm to alternativ e metho ds. • Metho d 1: T rained a DPM on the full dataset, using all 2678 Axial T 2 , 498 Axial T 1 , 949 P ost-contrast Axial T 1 , and 344 Axial FLAIR sub jects. • Metho d 2: Pre-trained on T 1 , T 2 and P ost-contrast T 1 data and fine-tuned on all 344 FLAIR sub jects. • Metho d 3: T rained only on the 344-sub ject FLAIR dataset. • Metho d 4 (Prop osed): Pre-trained on all T 2 , T 1 , and T 1 p ost-con trast data and then fine-tuned on 20 FLAIR sub jects with reduced learning rate and shorter training time. • Metho d 5: T rained solely on the 20-sub ject FLAIR dataset. • Metho d 6: T rained join tly on the full external dataset and the 20-sub ject FLAIR dataset without fine-tuning. Metho ds 1-3 serve as p erformance upp er b ounds, as they use extensive target con trast (FLAIR) data. Metho ds 4-6 represent the constrained data regime, relying on large external datasets and limited target-domain data. All metho ds were ev aluated on the 500 v alidation FLAIR slices using under-sampling rates of R = { 3 , 4 , 5 , 6 , 7 , 8 } with no auto-calibration (ACS) data. 2.6 Ev aluating P osterior Sampling Hyp erparameters W e ev aluated the effect of h yp erparameter selection in Algorithm 1 for approximate p osterior sam- pling. In particular, the data consistency weigh ting ζ is often set to b e inv ersely prop ortional to the norm of the data consistency error (31), while the num b er of steps N trades off b et ween qualit y and computation. Giv en a DPM trained for FLAIR reconstruction with the proposed metho d, we 7 / 28 Submited to Magnetic Resonance in Medicine compared reconstruction performance for different choices of ζ b et ween 1 / 2 and 5 and N b et ween 200 and 300 when reconstructing data at R = { 3 , 4 , 5 , 6 , 7 , 8 } . 2.7 Application to Clinical Stroke MRI In collaboration with our lo cal hospital’s medical sc ho ol, under IRB approv al and informed consen t, w e collected a small dataset of clinical stroke MRI from 30 patients on a 3T Siemens Vida scanner. Eac h sub ject underwen t a standard-of-care strok e imaging protocol following triage, including T2- FLAIR, MPRAGE, SWI, and DWI. The standard-of-care data were reconstructed with SENSE- based parallel imaging using coil maps calibrated with BAR T (41). The list b elo w summarizes the acquisition parameters: • T2-FLAIR: 0 . 5 × 0 . 5 mm in-plane resolution, 5 mm slice thickness, 9000 ms TR. • MPRA GE: 1 . 5 × 1 . 5 × 1 . 5 mm resolution, 240 × 240 × 168 mm F O V, 2500 ms TR. • SWI: 1 . 0 × 1 . 0 mm resolution, 250 × 211 mm FO V, 2 . 5 mm slice thickness. • D WI: 0 . 8 × 0 . 8 mm resolution, 250 × 250 mm FO V, 30 slices, 5 . 0 mm slice thickness, b-v alues: 0 , 1200 s/mm 2 . T o ev aluate our metho d in this clinical setting, w e first pre-trained a DPM on all Axial FLAIR, Axial T 2 , Axial T 1 , and Axial T 1 p ost con trast data a v ailable from the 4125 sub jects in fastMRI. Then, we fine-tuned the DPM sep erately for eac h clinical sequence (T2-FLAIR, MPRAGE, SWI, D WI) using 20 strok e patien ts from our lo cal dataset. Fine-tuning w as performed with a reduced learning rate and limited training duration as describ ed in Section 2.4. An additional 5 strok e sub jects were used for v alidation to select h yp er-parameters for fine-tuning and p osterior sampling. Fine-tuned DPMs then reconstructed retrosp ectiv ely under-sampled data (accelerations around R ≈ 4 ) on the remaining 5 test sub jects. 2.8 Clinical Strok e Reader Study W e acquired additional clinical stroke MRI data from 80 sub jects at our lo cal hospital with IRB ap- pro v al and informed consent to conduct a blinded reader study . Eac h sub ject was scanned using the standard-of-care strok e proto col, including T2-FLAIR, MPRA GE, SWI, and DWI acquisitions ob- tained at an acceleration factor of approximately R ≈ 2 and reconstructed using SENSE-based par- allel imaging. T o ev aluate accelerated imaging, the same data were retrosp ectiv ely under-sampled 8 / 28 Submited to Magnetic Resonance in Medicine b y an additional factor of tw o, resulting in a total effective acceleration of approximately R ≈ 4 , and reconstructed using diffusion probabilistic models (DPMs) fine-tuned with the prop osed training strategy . F or each of the 80 sub jects, the reconstructed images corresp onding to the standard-of-care and prop osed accelerated proto cols w ere group ed by sub ject, blinded, and randomized, yielding a total of 160 image sets for ev aluation. Imp ortantly , each image set comprised contrasts from the stroke proto col, and readers assessed the image set as a whole rather than individual sequences in isolation. Reader H.S. ev aluated all 160 image sets, while reader S.W. ev aluated 42 image sets (corresp onding to 21 sub jects) due to time constraints; H.S. and S.W. hav e 11 and 30 years of clinical exp erience, resp ectiv ely . Eac h image set w as scored for structural delineation of white matter, gra y matter, and ven tri- cles, as well as for image qualit y metrics including signal-to-noise ratio (SNR), contrast, sharpness, artifacts, and ov erall image quality . All ev aluations were p erformed using a fiv e-p oin t ordinal scale where [1 ← repeat scan needed , 2 ← limited , 3 ← diagnostic , 4 ← go od , 5 ← outstanding ] . Statis- tically significan t differences in metrics w as assessed using paired t wo-sided Wilcoxon signed-rank tests and inter-rater reliabilit y was measured with the Cohen’s k appa co efficien t. 2.9 Exp erimen ts on Prosp ectiv ely Under-sampled A cquisitions Noting that our clinical ev aluation inv olved retrosp ectiv e subsampling, we further ev aluated our metho d with prosp ectiv e under-sampling. W e acquired SWI and FLAIR data from a health y sub ject under IRB approv al and informed consen t using the parameters of the t ypical stroke proto col at our lo cal hospital. T w o acquisitions were p erformed: one at the standard-of-care acceleration rate, R = 1 . 7 , and another at a prospective acceleration rate of R = 3 . 75 . W e applied the prop osed metho d, pre-trained on all T 2 , T 1 , and p ost-con trast T 1 from fastMRI and fine-tuned on our FLAIR and SWI strok e data, to reconstruct the data prosp ectiv ely under-sampled to R = 3 . 41 and R = 3 . 75 resp ectiv ely and to reconstruct the R = 1 . 75 under-sampled data after it was further retrospectively under-sampled b y R = 3 . 52 and R = 3 . 75 resp ectiv ely . Reconstructions on b oth datasets w ere compared to a L1-wa v elet baseline implemented with BAR T (41). 9 / 28 Submited to Magnetic Resonance in Medicine Figure 2: Quan titative results from the controlled fastMRI exp erimen ts where DPMs w ere fine-tuned on 20 sub jects to reconstruct fastMRI flair data. NRMSE on 500 v alidation slices as a function of fine-tuning duration (ep ochs) is plotted for different learning rates and acceleration factors. A cross all acceleration factors, mo derate fine-tuning impro ves reconstruction qualit y relative to no fine- tuning, while excessiv e fine-tuning leads to degraded p erformance, particularly at higher learning rates. 3 Results 3.1 Con trolled Exp erimen ts with F astMRI Figure 2 quantitativ ely ev aluates the prop osed fine-tuning method across different learning rates and training ep ochs, measured with normalized ro ot mean squared error (NRMSE) at v arious acceleration factors on the 500 fastMRI FLAIR v alidation slices. Across all acceleration rates and learning rates, fine-tuning for 650 ep o c hs yields the low est NRMSE, outp erforming b oth no fine- tuning (0 ep o c hs) and longer (1250, 1875, 2500 ep ochs) fine-tuning durations. Similarly , a learning rate of 1 × 10 − 5 ac hieves the b est p erformance in comparison to higher learning rates. Figure 3 illustrates a represen tative example from the quantitativ e FLAIR exp erimen t at R = 4 . As shown in (A), combining a learning rate of 1 × 10 − 5 and 650 ep ochs giv es the b est quan ti- tativ e p erformance. (B) shows the fully-sampled reference and (C) zo oms into the reference and reconstructions under v arious learning rates and ep ochs. Qualitativ e improv ements are observed, 10 / 28 Submited to Magnetic Resonance in Medicine highligh ted by the orange arrow, when the appropriate fine-tuning hyperparameters are used. Figure 4 compares DPMs fine-tuned using our metho d (650 ep o c hs and 1 × 10 − 5 learning rate) on FLAIR data from 20 sub jects to metho ds (1-3) which ha ve access to FLAIR data from 344 sub jects and metho ds (5,6) with access to the same 20 FLAIR sub jects and large external dataset. (A) plots quantitativ e reconstruction p erformance for acceleration rates R = { 4 , 5 , 6 } . Our metho d ac hieves comparable p erformance to methods 1-3 whic h ha ve access to m uch more FLAIR data and impro ved performance in comparison to methods 5,6 with access to similar amoun ts of data. (B) sho ws example reconstructions from R = 4 under-sampled data. Quan titatively and qualitativ ely , as highlighted b y the orange arrow, our metho d ac hieves comparable quality to metho ds 1-3 and outp erforms metho ds 5 and 6. Using our b est fine-tuned DPM, Figure 5 plots the a v erage NRMSE across the v alidation samples for v arying acceleration rates as a function of the data consistency step-size ζ in Algorithm 1. Green dots highligh t the typical default setting of ζ = 1 and the red dots mark the v alue of ζ that achiev es the lo west reconstruction error at eac h acceleration rate. The optimal choice of ζ increases with higher acceleration rate. Supp orting T able S1 summarizes the b est p erforming of ζ and num b er of p osterior sampling steps N for each acceleration rate. 3.2 Application to Clinical Stroke MRI Figure 6 ev aluates DPMs fine-tuned using the proposed technique with limited strok e data to reconstruct R ≈ 3 . 75 SWI data from clinical strok e acquisitions. Similar to FLAIR in the fastMRI setting, the NRMSE plots in (A) av eraged o ver the 5 v alidation sub jects suggest that fine-tuning for to o few or to o man y epo c hs results in w orse quan titativ e performance. In addition, higher learning rates yield w orse p erformance for all training ep o c hs. These results suggest that 1250 ep ochs and a learning rate of 1 × 10 − 5 yields impro ved performance for fine-tuning a DPM to strok e SWI. (B) shows an example standard-of-care SWI image and (C) compares zo omed in views of this reference with reconstruction using DPMs fine-tuned with differen t h yp erparameters. Analysis on the 5 sub ject v alidation dataset found ζ = 2 . 1 as the optimal data consistency w eighting and w as used for all test reconstructions. Figure 7 presents fine-tuning a DPM to reconstruct DWI acquired with b-v alue 1200 s/mm 2 . (A) Fine-tuning with 252 ep ochs and a learning rate of 1 × 10 − 5 yields the b est p erformance. (B) sho ws an example standard-of-care DWI image and (C) compares zo omed in views of the reference and reconstructions using our fine-tuned DPM with v arious hyperparameters. 11 / 28 Submited to Magnetic Resonance in Medicine Supp orting Figure S1 analyzes the influence of fine-tuning epo c hs and learning rate for the additional FLAIR, MPRA GE, and D WI (b-v alue = 0) acquisitions in the test strok e data. Similar to SWI and DWI b-v alue 1200 s/mm 2 , appropriate selection of fine-tuning hyperparameters improv es reconstruction performance on R ≈ 4 under-sampled data. Supp orting Figure S2 presen ts an example MPRAGE slice from the quantitativ e analysis of Supp orting Figure S1. Figure 8 presen ts example stroke proto col images from the standard-of-care and images from the same proto col retrosp ectiv ely under-sampled b y an additional factor of 2 and reconstructed with DPMs fine-tuned with the prop osed strategy . F or DWI, w e display deriv ed trace and apparent diffusion co efficien t (ADC) maps, commonly used in clinical strok e assessmen t, instead of displaying the raw DWI images directly . 3.3 Clinical Reader Study Figure 9 summarizes the results of the blinded clinical reader study , in which t wo board-certified neuroradiologists ev aluated full strok e-proto col image sets reconstructed from standard-of-care data and from retrosp ectively accelerated data, b y a factor of 2, reconstructed using DPMs trained with the prop osed approach. The first reader ev aluated image sets from all 80 sub jects. F or structural delineation, the mean scores [standard, accelerated] for gray matter, white matter, and ven tricles were [4.8, 4.9], [4.8, 4.9], and [4.8, 4.9], resp ectiv ely . No statistically significan t difference w as observed for gray matter delin- eation, while the proposed accelerated reconstruction ac hieved statistically significan t improv ements for white matter and v entricular delineation. F or image quality metrics, mean scores [standard, accelerated] for SNR, contrast, sharpness, artifacts, and o verall image quality were [4.2, 4.7], [4.7, 4.8], [4.7, 4.8], [4.1, 4.3], and [4.5, 4.8], respectively . The accelerated reconstruction demonstrated statistically significant impro vemen ts in SNR, sharpness, artifact level, and ov erall image quality , while no statistically significant difference was observed for contrast. The second reader ev aluated image sets from 21 sub jects. F or structural delineation, mean scores [standard, accelerated] for gray matter, white matter, and ven tricles were [4.6, 4.1], [4.6, 4.1], and [5.0, 4.9], resp ectively , with no statistically significant differences observ ed. F or image quality metrics, mean scores [standard, accelerated] for SNR, con trast, sharpness, artifacts, and ov erall image quality w ere [4.9, 4.4], [4.5, 4.1], [4.4, 4.0], [4.0, 3.9], and [4.4, 4.0], resp ectiv ely . In this cohort, the standard-of-care reconstruction ac hiev ed a statistically significan tly higher SNR score, while no statistically significant differences were observed for the remaining image qualit y metrics. 12 / 28 Submited to Magnetic Resonance in Medicine In ter-rater reliability , as measured b y Cohen’s k appa co efficien t, was fair agreemen t ( 0 . 21 − 0 . 40 ) for contrast, sharpness, artifacts, and ov erall image quality , slight agreemen t ( 0 . 00 − 0 . 20 ) for SNR, white matter, and grey matter, and p o or agreemen t ( < 0 . 00 ) for ven tricles 3.4 Exp erimen ts on Prosp ectiv ely Under-sampled Data Figure 10 displays the mo del pre-trained with T 1 , T 2 , and p ost-con trast T 1 fastMRI and fine-tuned on FLAIR or SWI data from 20 sub jects applied to reconstruct retrosp ectiv ely and prosp ectiv ely under-sampled FLAIR and SWI data. Comparison to an L1-wa v elet baseline shows that the mo del qualitativ ely reduces reconstruction artifacts. In addition, the reconstructions from the retrosp ec- tiv ely and prosp ectiv ely under-sampled acquisitions show similar image qualit y . 4 Discussion W e prop osed a simple and effective training strategy for diffusion probabilistic mo dels (DPMs) that enables high-qualit y MRI reconstruction in data-constrained settings. Dra wing inspiration from the foundation mo del paradigm, the prop osed approac h com bines large-scale pre-training on diverse external MRI datasets follow ed by targeted fine-tuning on small, fully-sampled datasets from the application of interest, with an astute selection of hyperparameters. Controlled exp erimen ts on F astMRI demonstrate that DPMs trained with our strategy achiev ed reconstruction quality com- parable to mo dels trained on substantial ly larger target sp ecific datasets. W e further v alidated the approac h on our clinically motiv ated strok e MRI application, where the prop osed metho d gener- alized across multiple con trasts within the standard strok e imaging proto col. In a blinded clinical reader study , images reconstructed from data retrosp ectiv ely accelerated by an additional factor of t wo and fine-tuned using only 20 fully sampled sub jects were rated as non-inferior to standard-of- care reconstructions for b oth structural delineation and image quality metrics, In the controlled fastMRI FLAIR exp erimen ts, we observ ed a clear trade-off b et ween insufficient adaptation and ov erfitting during fine-tuning. As shown in Figures 2 and 3, omitting fine-tuning resulted in reconstructions that w ere p oorly adapted to the target contrast, leading to suboptimal image quality . In con trast, excessiv e fine-tuning, either through an increased num b er of epo c hs or an ov erly aggressive learning rate, degraded p erformance, consisten t with o verfitting to the limited size of the target dataset. Similar trends w ere observed when fine-tuning to differen t contrasts in the clinical strok e proto col, where the optimal fine-tuning configuration v aried across target datasets. 13 / 28 Submited to Magnetic Resonance in Medicine T ogether, these results highlight that effective transfer of diffusion models to data-limited MRI reconstruction tasks requires careful selection of fine-tuning h yp erparameters to balance contrast adaptation against ov erfitting or catastrophic forgetting (42). The fastMRI setting further enabled a direct comparison of the proposed training strategy against alternativ e approac hes with access to substantially larger amoun ts of target-domain FLAIR data (Figure 4). Metho ds 1–3 incorp orated FLAIR data from all 344 sub jects during training and therefore represent upp er-bound baselines that rely on extensive target-sp ecific sup ervision. Despite b eing fine-tuned using FLAIR data from only 20 sub jects, the prop osed approac h (Metho d 4) ac hieved reconstruction qualit y comparable to these upp er-bound baselines. In contrast, Metho ds 5 and 6, whic h either trained exclusively on the limited 20-sub ject FLAIR dataset or join tly on the large external dataset and the 20 sub ject FLAIR dataset, exhibited degraded p erformance relativ e to the prop osed metho d. These results indicate that large-scale pre-training on external data is b eneficial, and that sequential fine-tuning on the target domain is more effective than join t training when target-domain data are scarce. Since Algorithm 1 represen ts an appro ximate p osterior sampling pro cedure, fastMRI exp erimen ts in Figure 5 also c haracterized the effect of the data consistency w eighting ζ on reconstruction p erformance. Higher acceleration rates corresp onded to a larger optimal ζ . W e susp ect that less total data in the data consistency term results in a low er ov erall v alue relative to the prior term, so a larger weigh t needs to b e placed on the consistency term with less a v ailable k-space data. T raining DPMs for our limited stroke dataset motiv ated the prop osed approach. Figures 6, 7, and 8 demonstrate that the prop osed training strategy effectiv ely fine-tunes DPMs, with just data from 20 patients, to the v arious con trasts used in the stroke MRI proto col, including FLAIR, MPRA GE, SWI, and D WI. Results from the blinded clinical reader study (Figure 9) demonstrate that images reconstructed using the prop osed DPM-based approach from 2 × less data are non-inferior to the standard-of-care reconstructions for b oth structural delineation and image quality metrics. F or the first reader, who ev aluated all 80 sub jects, the prop osed approach achiev ed statistically significan t, sligh tly higher scores than the standard-of-care in several categories, including SNR, artifact level, and o verall image qualit y . In contrast, the second reader, who ev aluated a smaller subset of 21 sub jects, tended to rate the standard-of-care images slightly higher across most metrics. Ho wev er, no statistically significan t differences were observ ed for this reader except for SNR, where the standard-of-care ac hieved higher scores. 14 / 28 Submited to Magnetic Resonance in Medicine Imp ortan tly , despite these inter-reader differences, mean scores for b oth reconstruction ap- proac hes remained consistently high for b oth readers, with most metrics av eraging near or ab o v e a score of 4, corresp onding to go od image quality . This suggests that, while individual reader preferences and scoring tendencies may differ, both the prop osed and standard-of-care reconstruc- tions pro duced clinically acceptable images across the ev aluated stroke proto cols. The observed impro vemen t in SNR for the proposed approac h in the first reader ma y be attributable to the de- noising effect of posterior sampling guided by a learned score function, while the reduced artifact scores may reflect the exclusion of motion-corrupted k-space lines when reconstructing from few er measuremen ts. Most exp erimen ts in this work relied on retrosp ectiv e under-sampling, while the prosp ectiv e ev aluation was limited to data acquired from a health y volun teer. F uture work will therefore fo- cus on v alidating the prop osed approach using prosp ectiv ely under-sampled clinical stroke MRI acquisitions to more fully assess performance. In addition, future reader studies will ev aluate the impact of the prop osed reconstructions on do wnstream clinical interpretation and decision-making relativ e to standard-of-care images. Finally , the current p osterior sampling implemen tation incurs substan tially longer reconstruction times p er slice compared to conv entional parallel imaging and end-to-end learning–based metho ds. F uture work will in vestigate more efficient implementations and improv ed p osterior sampling strategies to reduce reconstruction time and facilitate practical clinical deploymen t. 5 A c kno wledgment This w ork was supp orted in part b y NSF CCF-2239687 (CAREER), NSF IFML 2019844, and JCCO fello wship. References 1 Rishi Bommasani, Kyle Hudson, Ehsan Adeli, Russ Altman, Simran Arora, Sydney von Arx, Mic hael S Bernstein, Jeannette Bohg, An toine Bosselut, Emma Brunskill, et al. On the opp or- tunities and risks of foundation mo dels. arXiv , 2021. 2 Ce Zhou, Qian Li, Chen Li, Jun Y u, Yixin Liu, Guang jing W ang, Kai Zhang, Cheng Ji, Qiben Y an, Lifang He, Hao Peng, Jianxin Li, Jia W u, Ziwei Liu, Pengtao Xie, Caiming Xiong, Jian 15 / 28 Submited to Magnetic Resonance in Medicine P ei, Philip S. Y u, and Lichao Sun. A comprehensive survey on pretrained foundation mo dels: a history from b ert to chatgpt. International Journal of Machine L e arning and Cyb ernetics , 2024. 3 W ayne Xin Zhao, Kun Zhou, Junyi Li, Tianyi T ang, Xiaolei W ang, Y up eng Hou, Yingqian Min, Beic hen Zhang, Junjie Zhang, Zican Dong, Yifan Du, Chen Y ang, Y ushuo Chen, Zhip eng Chen, Jinhao Jiang, Ruiyang Ren, Yifan Li, Xinyu T ang, Zik ang Liu, P eiyu Liu, Jian-Y un Nie, and Ji-Rong W en. A survey of large language mo dels. arXiv , 2023. 4 Alec Radford, Jong W o ok Kim, Chris Hallacy , Adit y a Ramesh, Gabriel Goh, Sandhini Agar- w al, Girish Sastry , Amanda Ask ell, Pamela Mishkin, Jac k Clark, Gretc hen Krueger, and Ilya Sutsk ever. Learning transferable visual models from natural language sup ervision. Pr o c e e dings of Machine L e arning R ese ar ch , 2021. 5 Alexander Kirillov, Eric M in tun, Nikhila Ra vi, Hanzi Mao, Chlo e Rolland, Laura Gustafson, T ete Xiao, Sp encer Whitehead, Alexander C. Berg, W an-Y en Lo, Piotr Dollár, and Ross Girshick. Segmen t anything. IEEE/CVF International Confer enc e on Computer Vision (ICCV) , 2023. 6 Chenxi Ma, W eimin T an, Ruian He, and Bo Y an. Pretraining a foundation model for general- izable fluorescence microscopy-based image restoration. Natur e Metho ds , 2024. 7 Zhi Huang, F ederico Bianchi, Mert Y uksekgonul, Thomas J. Montine, and James Zou. A vi- sual–language foundation model for pathology image analysis using medical t witter. Natur e Me dicine , 2023. 8 Y ukun Zhou, Mark A. Chia, Siegfried K. W agner, Murat S. A yhan, Dominic J. Williamson, Robb ert R. Struyv en, Timing Liu, Mouc heng Xu, Mateo G. Lozano, P eter W oo dw ard-Court, Y uk a Kihara, UK Biobank Ey e, Vision Consortium, Andre Altmann, Aaron Y. Lee, Eric J. T op ol, Alastair K. Denniston, Daniel C. Alexander, and Pearse A. Keane. A visual–language foundation mo del for pathology image analysis using medical twitter. Natur e , 2023. 9 Mic hael Moor, Oishi Banerjee, Zahra Shak eri Hossein Abad, Harlan M. Krumholz, Jure Lesk ov ec, Eric J. T op ol, and Pranav Ra jpurk ar. F oundation mo dels for generalist medical artificial intel- ligence. Natur e , 2023. 10 Reinhard Heck el, Mathews Jacob, Akshay Chaudhari, Or Perlman, and Efrat Shimron. Deep learning for accelerated and robust mri reconstruction. Magnetic R esonanc e Materials in Physics, Biolo gy and Me dicine , 2024. 16 / 28 Submited to Magnetic Resonance in Medicine 11 Matthew Muc kley , Bruno Riemensc hneider, Alireza Radmanesh, Sun W oo Kim, Geun u Jeong, Jingyu Ko, Y ohan Jun, Hyungseob Shin, Dosik Hwang, Mahmoud Mostapha, Simon Arb eret, Dominik Nick el, Zaccharie Ramzi, Philipp e Ciuciu, Jean-Luc Starck, Jonas T euw en, Dim- itrios Kark alousos, Chaoping Zhang, Anuroop Sriram, Zhengnan Huang, Nafissa Y akub o v a, Y vonne W. Lui, and Florian Knoll. Results of the 2020 fastmri c hallenge for machine learning mr image reconstruction. IEEE T r ansactions on Me dic al Imaging , 2021. 12 Jun Lyu, Chen Qin, Sh uo W ang, F an w en W ang, Y an Li, Zi W ang, Kun yuan Guo, Cheng Ouy ang, Mic hael Tänzer, Meng Liu, Longyu Sun, Mengting Sun, Qing Li, Zhang Shi, Sha Hua, Hao Li, Zhensen Chen, Zhenlin Zhang, Bingyu Xin, Dimitris N. Metaxas, George Yiasemis, Jonas T euw en, Liping Zhang, W eitian Chen, Yidong Zhao, Qian T ao, Y an w ei P ang, Xiaohan Liu, Artem Razumo v, Dmitry V. Dylo v, Quan Dou, Kang Y an, Y uy ang Xue, Y uning Du, Julia Dietlmeier, Carles Garcia-Cabrera, Ziad Al-Ha j Hemidi, Nora V ogt, Ziqiang Xu, Y a jing Zhang, Ying-Hua Ch u, W eib o Chen, W enjia Bai, Xiahai Zhuang, Jing Qin, Lianming W u, Guang Y ang, Xiaob o Qu, He W ang, and Chengy an W ang. Results of the 2020 fastmri c hallenge for machine learning mr image reconstruction. Me dic al Image A nalysis , 2023. 13 Arjun D. Desai, Andrew M. Sc hmidt, Elk a B. Rubin, Christopher M. Sandino, Marianne S. Blac k, V alentina Mazzoli, Kathryn J. Stev ens, Rob ert Boutin, Christopher Ré, Garry E. Gold, Brian A. Hargrea ves, and Aksha y S. Chaudhari. Skm-tea: A dataset for accelerated mri reconstruction with dense image lab els for quan titativ e clinical ev aluation. NeurIPs , 2024. 14 Dale Birenbaum, Laura Bancroft, and Gary F elsb erg. Imaging in acute stroke. W estern Journal of Emer gency Me dicine , 12:67–76, 2011. 15 Costanza Maria Rapillo, Vincent Dunet, Silvia Pisto cc hi, Vincent Darioli, Bruno Bartolini, Stev en Da vid Ha jdu, P atrik Michel, and Da vide Stram b o. Mo ving from ct to mri paradigm in acute ischemic strok e: F easibilit y , effects on stroke diagnosis and long-term outcomes. Str oke , 2024. 16 Mehmet Akçak ay a, Steen Mo eller, Sebastian W eingärtner, and Kâmil Uğurbil. Scan-sp ecific robust artificial-neural-net works for k-space interpolation (raki) reconstruction: Database-free deep learning for fast imaging. Magnetic R esonanc e in Me dicine , 2019. 17 Y amin Arefeen, On ur Beker, Jaejin Cho, Elfar A dalsteinsson, and Berkin Bilgic. Scan-sp ecific 17 / 28 Submited to Magnetic Resonance in Medicine artifact reduction in k-space (spark) neural net works synergize with physics-based reconstruction to accelerate mri. Magnetic R esonanc e in Me dicine , 2022. 18 Ruimin F eng, Qing W u, Jie F eng, Hua jun She, Ch unlei Liu, Y uyao Zhang, and Hong jiang W ei. Imjense: Scan-specific implicit representation for joint coil sensitivity and image estimation in parallel mri. IEEE T r ansactions on Me dic al Imaging , 2024. 19 Mohammed Darestani and Reinhard Hec k el. Accelerated mri with un-trained neural net works. IEEE T r ansactions on Computational Imaging , 2021. 20 Burhaneddin Y aman, Sey ed Amir Hossein Hosseini, and Mehmet Ak çak a ya. Zero-shot self- sup ervised learning for mri reconstruction. International Confer enc e on L e arning R epr esenta- tions , 2024. 21 Burhaneddin Y aman, Sey ed Amir, Hossein Hosseini, Steen Mo eller, Jutta Ellermann, Kâmil Uğurbil, and Mehmet Akçak a y a. Self-sup ervised learning of ph ysics-guided reconstruction neural net works without fully sampled reference data. Magnetic R esonanc e in Me dicine , 2020. 22 Charles Millard and Mark Chiew. A theoretical framew ork for self-sup ervised mr image recon- struction using sub-sampling via v ariable densit y noisier2noise. IEEE T r ansactions on Compu- tational Imaging , 2023. 23 Arjun D. Desai, Batu M. Ozturkler, Christopher M. Sandino, Rob ert Boutin, Marc Willis, Shrey as V asana w ala, Brian A. Hargrea ves, Christopher Ré, John M. Pauly , and Aksha y S. Chaudhari. Noise2recon: Enabling snr-robust mri reconstruction with semi-sup ervised and self- sup ervised learning. Magnetic R esonanc e in Me dicine , 2023. 24 Andrew W ang and Mik e Da vies. Benc hmarking self-supervised metho ds for accelerated mri reconstruction. arXiv , 2025. 25 Mohammad Zalbagi Darestani, Jia yu Liu, and Reinhard Hec k el. T est-time training can close the natural distribution shift p erformance gap in deep learning based compressed sensing. ICML , 2022. 26 Matthieu T erris, Sam uel Hurault, Maxime Song, and Julian T ac hella. Reconstruct an ything mo del a light weigh t foundation mo del for computational imaging. ICLR , 2026. 18 / 28 Submited to Magnetic Resonance in Medicine 27 Jupinder P armar, Sanjev Satheesh, Mostofa P atw ary , Mohammad Sho eybi, and Bryan Catan- zaro. Reuse, don’t retrain: A recip e for contin ued pretraining of language mo dels. arXiv , 2024. 28 Reinhard Heck el Kang Lin. Robustness of deep learning for accelerated mri: Benefits of div erse training data. International Confer enc e on Machine L e arning , 2024. 29 Y ang Song, Jasc ha Sohl-Dickstein, Diederik P . Kingma, Abhishek Kumar, Stefano Ermon, and Ben Poole. Score-based generative mo deling through sto c hastic differen tial equations, 2021. 30 Hyung jin Chung and Jong Chul Y e. Score-based diffusion mo dels for accelerated mri. Me dic al Image Analysis , 2022. 31 Hyung jin Chung, Jeongsol Kim, Michael T. Mccann, Marc L. Klasky , and Jong Chul Y e. Diffu- sion p osterior sampling for general noisy in verse problems, 2023. 32 Ajil Jalal, Marius Arvinte, Giannis Daras, Eric Price, Alexandros G Dimakis, and Jonathan I T amir. Robust compressed sensing mri with deep generative priors. A dvanc es in Neur al Infor- mation Pr o c essing Systems , 2021. 33 Guanxiong Luo, Moritz Blumenthal, Martin Heide, and Martin Uec ker. Ba yesian mri reconstruc- tion with joint uncertain ty estimation using diffusion mo dels. Magnetic R esonanc e in Me dicine , 2023. 34 Ling Y ang, Zhilong Zhang, Y ang Song, Shenda Hong, Runsheng Xu, Y ue Zhao, W entao Zhang, Bin Cui, and Ming-Hsuan Y ang. Diffusion mo dels: A comprehensive surv ey of metho ds and applications. ACM Computing Surveys , 2023. 35 Jonathan Ho, Aja y Jain, and Pieter Abb eel. Denoising diffusion probabilistic mo dels. A dvanc es in Neur al Information Pr o c essing Systems , 2020. 36 Hongk ai Zheng, W enda Chu, Bingliang Zhang, Zihui W u, Austin W ang, Berthy T. F eng, Caifeng Zou, Y u Sun andNikola K ov achki, Zac hary E. Ross, Katherine L. Bouman, and Yisong Y ue1. In versebench: Benc hmarking plug-and-pla y diffusion priors for inv erse problems in physical sci- ences. International Confer enc e on L e arning R epr esentations , 2025. 37 Guanxiong Luo, Na Zhao, W enhao Jiang, Edw ard S Hui, and P eng Cao. Mri reconstruction using deep bay esian estimation. Magnetic R esonanc e in Me dicine , 2020. 19 / 28 Submited to Magnetic Resonance in Medicine 38 Kerem T ezcan, Christian Baumgartner, Roger Luechinger, Klaas Pruessmann, and Ender K onuk oglu. Mr image reconstruction using deep densit y priors. IEEE T r ansactions on Me d- ic al Imaging , 2019. 39 T ero Karras, Miik a Aittala, Timo Aila, and Sam uli Laine. Elucidating the design space of diffusion-based generativ e models. In A dvanc es in Neur al Information Pr o c essing Systems , vol- ume 35, pages 26565–26577, 2022. 40 Y amin Arefeen, Brett Lev ac, Bhairav Patel, Chang Ho, and Jonathan I T amir. Diffusion prob- abilistic generative models for accelerated, in-nicu p ermanen t magnet neonatal mri. Magnetic R esonanc e in Me dicine , 2025. 41 Martin Ueck er, F rank Ong, Jonathan I T amir, Dara Bahri, P atrick Virtue, Joseph Y Cheng, T ao Zhang, and Mic hael Lustig. Berkeley adv anced reconstruction to olbox. Pr o c e e dings of the International So ciety of Magnetic R esonanc e in Me dicine , 2015. 42 James Kirkpatric k, Razv an Pascan u, Neil Rabino witz, Joel V eness, Guillaume Desjardins, An- drei A. Rusu, Kieran Milan, John Quan, Tiago Ramalho, Agnieszk a Grabsk a-Barwińsk a, et al. Ov ercoming catastrophic forgetting in neural netw orks. Pr o c e e dings of the National A c ademy of Scienc es , 114(13):3521–3526, 2017. 20 / 28 Submited to Magnetic Resonance in Medicine Figure 3: A represen tative example from the fastMRI FLAIR exp eriment. (A) Quan titative NRMSE as a function of fine-tuning duration for differen t learning rates, highligh ting that a comparatively smaller learning rate and fewer fine-tuning ep ochs yields the b est p erformance. (B) F ully sampled reference FLAIR image. (C) Zo omed reconstructions obtained using different learning rates with 650 fine-tuning ep o c hs, compared against the fully sampled reference. (D) Reconstructions obtained using a fixed learning rate of 1 × 10 − 5 across different fine-tuning durations. Appropriate fine-tuning h yp erparameters lead to visibly improv ed structural fidelity and reduced artifacts, as highlighted b y the orange arro ws. 21 / 28 Submited to Magnetic Resonance in Medicine Figure 4: (A) DPMs are compared across metho ds with access to differen t amoun ts of target- domain FLAIR data. Metho ds 1–3 are trained using FLAIR data from all 344 sub jects, while Metho ds 5 and 6 are trained using 20 FLAIR sub jects and the external dataset. The prop osed approac h (Metho d 4), pre-trained on a large external dataset and fine-tuned on 20 FLAIR sub jects, ac hieves comparable NRMSE to Metho ds 1-3, despite using substantially less target domain data and consisten tly outp erforms Metho ds 5,6 trained on the same limited data. (B) displa ys example reconstructions from this exp erimen t. 22 / 28 Submited to Magnetic Resonance in Medicine Figure 5: Using the b est fine-tuned DPM, av erage NRMSE on fastMRI FLAIR v alidation data is plotted as a function of the data-consistency step size ζ for multiple acceleration factors. Red mark ers denote the default c hoice ζ = 1 , while green mark ers indicate the v alue of ζ that minimizes reconstruction error at each acceleration. The optimal ζ increases with acceleration. 23 / 28 Submited to Magnetic Resonance in Medicine Figure 6: (A) NRMSE of SWI reconstructions av eraged ov er five v alidation sub jects from data ret- rosp ectiv ely accelerated by R ≈ 3 . 75 . (B) F ully-sampled reference. (C) Zo omed-in reconstructions obtained with fixed ep o c hs and v arying learning rate and (D) fixed learning rate and v arying ep o c hs. Similar to the fastMRI FLAIR exp erimen ts, insufficient or excessive fine-tuning duration and higher learning rates consisten tly result in worse reconstructions, demonstrating that the prop osed training strategy is effective for clinical stroke SWI data. 24 / 28 Submited to Magnetic Resonance in Medicine Figure 7: (A) NRMSE of DWI reconstructions av eraged ov er five v alidation sub jects from data ret- rosp ectiv ely accelerated by R ≈ 3 . 75 . (B) F ully-sampled reference. (C) Zo omed-in reconstructions obtained with fixed ep o c hs and v arying learning rate and (D) fixed learning rate and v arying ep o c hs. Similar to the fastMRI FLAIR exp erimen ts, insufficient or excessive fine-tuning duration and higher learning rates consisten tly result in worse reconstructions, demonstrating that the prop osed training strategy is effective for clinical stroke DWI data. 25 / 28 Submited to Magnetic Resonance in Medicine Figure 8: Represen tative images from a clinical stroke protocol are shown for a single sub ject, com- paring standard-of-care reconstructions (top row) with images retrosp ectiv ely under-sampled by an additional factor of tw o and reconstructed using the prop osed DPM-based approach (b ottom row). The proto col includes diffusion-w eighted imaging (shown as deriv ed trace and apparent diffusion co efficien t (ADC) maps), susceptibility-w eighted imaging (SWI), FLAIR, and MPRA GE. Across all con trasts, the prop osed accelerated reconstructions preserve structural detail and contrast compa- rable to the standard-of-care. 26 / 28 Submited to Magnetic Resonance in Medicine Figure 9: Mean reader scores for structural delineation (left) and image quality metrics (right) are shown for tw o b oard-certified neuroradiologists ev aluating full stroke-protocol image sets recon- structed from standard-of-care data (GT) and from retrosp ectiv ely accelerated data reconstructed using the prop osed DPM-based approach (Recon). The top row rep orts results from Reader 1, who ev aluated all 80 sub jects, while the b ottom row rep orts results from Reader 2, who ev aluated 21 sub jects. F or Reader 1, the prop osed accelerated reconstructions ac hieved statistically significant higher scores for several image qualit y metrics and for white matter and ven tricular delineation. F or Reader 2, no significan t differences were observ ed except for SNR, where the standard-of-care reconstruction scored higher; how ev er, mean scores for b oth metho ds remained high across all cat- egories. 27 / 28 Submited to Magnetic Resonance in Medicine Figure 10: The prop osed DPM, pre-trained on fastMRI data and fine-tuned on FLAIR or SWI data from 20 sub jects, is applied to reconstruct b oth retrosp ectiv ely and prosp ectiv ely under-sampled clinical stroke data. F or eac h contrast, the fully sampled reference, retrosp ectiv e reconstructions, and prosp ectiv e reconstructions are display ed along with their effective acceleration factors. A cross b oth contrasts, the proposed metho d qualitativ ely reduces reconstruction artifacts relativ e to the ℓ 1 baseline, and ac hieves comparable image quality b et ween retrosp ectiv ely and prosp ectiv ely under- sampled acquisitions 28 / 28
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment