Multimodal MRI Report Findings Supervised Brain Lesion Segmentation with Substructures
Report-supervised (RSuper) learning seeks to alleviate the need for dense tumor voxel labels with constraints derived from radiology reports (e.g., volumes, counts, sizes, locations). In MRI studies of brain tumors, however, we often involve multi-pa…
Authors: Yubin Ge, Yongsong Huang, Xiaofeng Liu
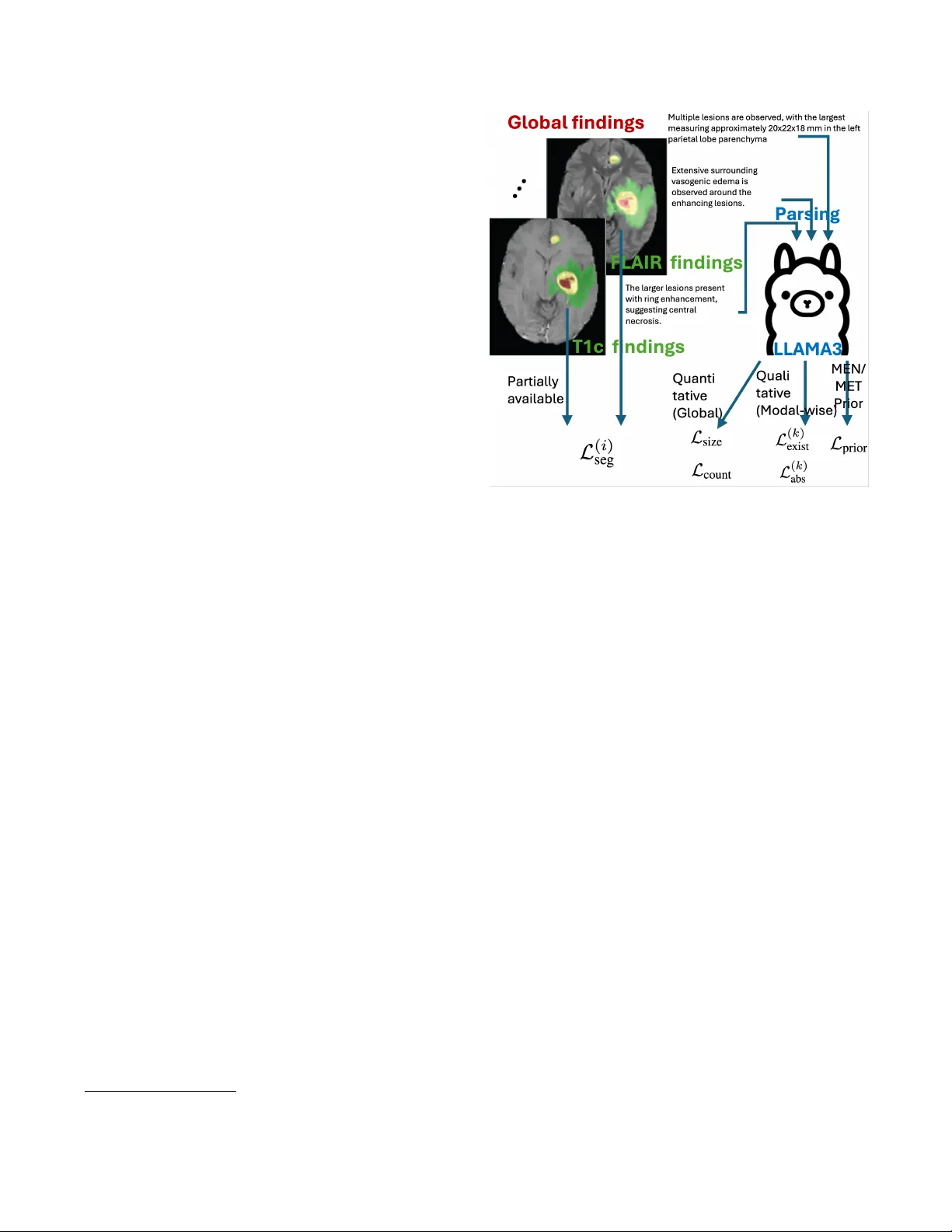
MUL TIMOD AL MRI REPOR T FINDINGS SUPER VISED BRAIN LESION SEGMENT A TION WITH SUBSTR UCTURES Y ubin Ge 1 † , Y ongsong Huang 2 , 3 † , Xiaofeng Liu 2 ∗ 1 Amazon A WS, 2 Y ale Uni versity , 3 T ohoku Univ ersity ABSTRA CT Report-supervised (RSuper) learning seeks to alleviate the need for dense tumor vox el labels with constraints deriv ed from radiology reports (e.g., volumes, counts, sizes, loca- tions). In MRI studies of brain tumors, ho wev er , we often in volv e multi-parametric scans and substructures. Here, fine- grained modality/parameter-wise reports are usually pro vided along with global findings and are correlated with different substructures. Moreov er , the reports often describe only the largest lesion and provide qualitativ e or uncertain cues (“mild, ” “possible”). Classical RSuper losses (e.g., sum volume consistency) can ov er-constrain or hallucinate unre- ported findings under such incompleteness, and are unable to utilize these hierarchical findings or exploit the priors of v ar- ied lesion types in a mer ged dataset. W e explicitly parse the global quantitati ve and modality-wise qualitati ve findings and introduce a unified, one-sided, uncertainty-aware formulation (MS-RSuper) that: (i) aligns modality-specific qualitativ e cues (e.g., T1c enhancement, FLAIR edema) with their cor- responding substructures using existence and absence losses; (ii) enforces one-sided lower -bounds for partial quantitative cues (e.g., largest lesion size, minimal multiplicity); and (iii) adds e xtra- vs. intra-axial anatomical priors to respect co- hort differences. Certainty tokens scale penalties; missing cues are do wn-weighted. On 1238 report-labeled BraTS- MET/MEN scans, our MS-RSuper largely outperforms both a sparsely-supervised baseline and a naiv e RSuper method. Index T erms — Report supervision, multimodal MRI, meningioma, brain metastases, segmentation. 1. INTRODUCTION Accurate delineation of lesion structures is a fundamental step in clinical diagnosis, intervention, and treatment planning [1]. Howe ver , vox el-wise annotation for 3D multimodal MRI is costly and subjectiv e, especially when substructures of tu- mor core (TC), enhancing tumor (ET), and edema (ED) must be delineated across sequences with different contrast mech- anisms [1]. How to utilize e xisting, routinely summarized radiology reports to aid (or assist) segmentation model train- † contribute equally . ∗ corresponding author: xiaofeng.liu@yale.edu ing is of great importance for more practical utilization of big medical data. T o exploit the v aluable and relatively large-scale text in- formation, early attempts usually form a multi-task learning with an auxiliary task in addition to the se gmentation with either the extracted tumor present/absent label for classifi- cation [2], or a contrastive language-image pre-training ob- jectiv e [3]. While the benefits of auxiliary tasks to segmen- tation are indirect and occasionally minor, the recently de- veloped Report-supervised learning (RSuper) [4] promises to directly le verage the detailed volumes, counts, and sizes e x- tracted from abundant abdominal CT radiology reports and enforces the corresponding loss functions alongside the con- ventional segmentation loss for only a small portion of seg- mentation labeled samples (e.g., 50 scans). This approach therefore largely reduces the b urden of manual labeling. Howe ver , in MRI studies of brain tumors, we often in- volv e multi-parametric scans of T1, T1c, T2, and FLAIR as well as substructures of TC, ET , and ED. In which the fine- grained modality/ parameter-wise finding reports are usually provided along with the global finding reports, and correlated to dif ferent substructures. Ho w to systematically exploit both the global descriptors (falx-/skull-base adjacency vs. deep parenchyma, approximate size, multiplicity , edema/midline shift) and modality-wise descriptors (T1c enhancement pat- tern; FLAIR hyperintensity) is largely undere xplored. Moreov er , the reports often describe only the largest le- sion, provide qualitativ e or uncertain cues (“mild, ” “possi- ble”). Specifically , in brain lesions, reports are often par- tially specified : many cases provide only the lar gest lesion size d max , omit axes for diameters, and/or use certainty quali- fiers ( mild , possible , equivocal ); and the counts are sometimes qualitativ e (“multiple”). Simply adopt the classical RSuper volume loss, which penalizes the difference of predicted and labeled sum volume of all tumors, may (a) learn to suppress small lesions that the report does not enumerate, or (b) unduly shrink tumors to match a partial volume hint. Finally , when combining image-report records from mul- tiple diseases, cohort-specific priors are lost. For e xample, BraTS-MET (metastases) typically includes multiple intra- axial parenchymal lesions, often with ring enhancement. In contrast, BraTS-MEN (meningioma) is typically e xtra-axial and dural-based (e.g., falx, skull base) with solid enhance- ment. A report-deri ved supervision should transfer across cohorts without imposing contradictory biases. A nai ve RSu- per loss cannot lev erage these strong anatomical priors. T o address these limitations, we propose a no vel multi- modality with substructure RSuper framew ork for brain le- sion segmentation. Our main contributions are: • Modality-Substructure Alignment: W e introduce a loss that links modality-specific report findings (e.g., T1c enhancement, FLAIR edema) directly to their corresponding segmentation substructures (ET and ED, respecti vely). • One-Sided Partial-Report Loss: W e propose a ”lo wer- bound” size loss and ”minimal-multiplicity” count loss to handle incomplete reports that only describe the largest le- sion or use qualitative counts, av oiding penalty for valid, unreported lesions. • Cohort-Specific Priors: W e integrate an anatomical prior loss that penalizes intra-axial predictions for MEN and extra-axial predictions for MET , guided by cohort-lev el cues from the reports. W e v alidated its ef fecti veness on the combined BraTS- MET and BraTS-MEN with segmentation and report 1 . 2. METHODOLOGY Our frame work trains a 3D se gmentation network using a par- tially segmented dataset, i.e., a lar ge set of image-report pairs ( D R ) and a small set of fully-masked data ( D M ). The model is trained with a composite loss. F or data in D M , we use a standard supervised segmentation loss, L seg (e.g., a combina- tion of Dice and Cross-Entropy loss). For data in D R , we introduce a nov el report-supervised loss, L report , designed to handle the hierarchical, qualitati ve, and partial nature of mul- timodal MRI reports. 2.1. Hierarchical Report Parsing and Mapping W e first employ a Large Language Model (LLM) with domain-specific prompts to parse each free-text radiology report. Critically , we categorize the extracted cues into two distinct types based on their nature and scope: (A) Quantitative Global Cues: These are specific measure- ments, typically found in the ”global findings” section, that apply to the entir e lesion or provide a total count. These are often partial (e.g., “largest lesion measuring 45x39x47 mm, ” ”multiple punctate... lesions”). (B) Qualitative Modality-Specific Cues: These are usually descriptiv e, non-numeric findings tied to a specific MRI sequence, which inherently map to tumor substructures. Though sometimes the tumor size is provided, it is the same as the global finding, which does not provide incremental information. For example, T1c often includes “obvious en- hancement, ” “ring enhancement, ” “no enhancement, ” while 1 ht tp s: // hu gg in g fa ce .c o/ da ta se ts/ Ji ay uL ei /R ad Ge nome- Brain_MRI/tree/main Fig. 1 . Overvie w of our proposed report-supervised frame- work. An LLM parses hierarchical findings from reports. Our losses align modality-specific findings, handle partial cues, and enforce anatomical location priors. FLAIR includes “surrounding extensiv e edema, ” “mild h y- perintense signal. ” Based on this parsing, we propose to establish a Modality- Substructure Alignment Principle . This is not a loss func- tion itself, b ut a crucial mapping rule that directs ho w con- straints are applied: • T1c findings (enhancement) constrain the Enhancing T umor ( P E T ) probability map. • FLAIR findings (edema, non-specific hyperintensity) constrain the Edema ( P E D ) probability map. • T1 or T2 findings (e.g., ”hypointense core”) constrain the T umor Core ( P T C ) map. • Global cues (e.g., total size, count) constrain the Whole T umor ( P W T ) map, where P W T = P E T + P E D + P T C . Uncertainty cues (e.g., ”possible, ” ”mild”) are parsed into a scaling weight λ ∈ [0 , 1] for the corresponding loss term. 2.2. Unified Report Constraint Loss ( L report ) Our primary report loss, L report , combines constraints from both qualitativ e and quantitative cues, applying them to the aligned substructure maps identified in § 2 . 1 . 2.2.1. Substructure Qualitative Existence and Absence Loss As identified, most modality-specific cues are qualitativ e (e.g., ”edema is present”) and lack quantitativ e volumes. W e cannot use a symmetric L1/L2 volume loss [4]. In- stead, we formulate a loss based on the existence or ab- sence of a finding. F or a gi ven substructure class k (e.g., k ∈ { ET , ED, TC } ), let V k be the predicted volume of P k ( x ) ≥ 0 . 5 for that substructure. If the report confirms the pr esence of substructure k (e.g., ”surrounding edema” → k = ED) with confidence λ k, pos , we apply an ”Existence Loss. ” This loss penalizes the model only if it fails to predict any presence (volume < 1 ) of that substructure L ( k ) exist = max(0 , 1 − V k ) , which encourages the model to segment at least 1 v oxel for class k , without halluci- nating a specific target v olume. Con versely , if the report explicitly confirms the absence of a substructure k (e.g., ”no enhancement” → k = ET), we apply an loss that penalizes any prediction for that class L ( k ) exist = V k . Therefore, we hav e L ( k ) exist = max(0 , 1 − V k ) if presence confirmed, V k if absence confirmed, 0 otherwise. (1) 2.2.2. Global One-Sided P artial Cue Loss (Size and Count) W e handle the quantitati ve b ut partial nature of global cues: • Size Loss: Reports often provide only the 3D-dim or diameter(s) of the larg est lesion, d max . Let C pred be the set of predicted connected components for the whole tu- mor ( P W T ≥ 0 . 5 ). Let d c be the volume of a component c ∈ C pred . The loss is: L size = | d max − max c ∈ C pred d c | . This loss used mean absolute error , which is more robust to small inaccurate measure of d max . • Count Loss: Reports often use qualitativ e counts like ”multiple” or ”a fe w . ” W e parse this to a minimal inte ger N qual (e.g., ”multiple” → N qual = 2 ). W e apply a one-sided count loss: L count = max(0 , N qual − | C pred | ) . It penalizes the model if it predicts fewer than N qual lesions. Therefore, we hav e L global = w size L size + w count L count . 2.3. Cohort-Specific Anatomical Prior Loss ( L prior ) Finally , we lev erage global location cues (e.g., “falx, ” “parenchy- mal”) to identify the cohort (MEN or MET) and apply a strong anatomical prior . W e use pre-defined binary masks for the dura/extra-axial space ( M dural ) and the brain parenchyma/ intra-axial space ( M parench ). • If the report suggests Meningioma (MEN) , which is extra-axial, we penalize an y intra-axial predictions: L prior = X x ( P W T ( x ) · M parench ( x )) . • If the report suggests Metastases (MET) , which are intra-axial, we penalize extra-axial predictions: L prior = X x ( P W T ( x ) · M dural ( x )) . This loss ef fectiv ely guides the model to search in the correct anatomical compartment, resolving ambiguity and reducing false positi ves. 2.4. T otal Loss Function The model is first pre-trained on D M using L seg . It is then fine-tuned on the combined dataset D M ∪ D R . For a mixed batch B = B M ∪ B R , the total loss is: L total = 1 | B M | X i ∈ B M L ( i ) seg + w r | B R | X j ∈ B R L ( j ) report , where w r is a balancing weight for the report-based supervi- sion, and L report is the sum of our proposed constraint losses: L report = 3 X k L ( k ) exist + w size L size + w count L count + w prior L prior , where w size , w count , and w prior are weights for each component of the report-supervised loss. 3. EXPERIMENTS AND RESUL TS W e used two large-scale, multi-modal MRI segmentation datasets. Their associated radiology reports are manually generated in RadGenome-Brain MRI dataset [5] 1 . Each sub- ject has a global finding and four modality-wise findings. • BraTS-MEN (Meningioma): A collection of 1000 subjects (4000 3D mpMRI scans) with meningioma. Reports frequently describe extra-axial, dural-based lesions (e.g., ”falx cerebri, ” ”skull base, ” ”cerebellopontine angle”) with ”marked, ” ”uniform” enhancement on T1c. • BraTS-MET (Metastases): A collection of 238 sub- jects (952 3D mpMRI scans) with brain metastases. Reports describe ”multiple, ” ”parenchymal” (intra-axial) lesions, of- ten with ”ring enhancement” and ”extensi ve surrounding edema” on FLAIR. W e held out 50 MEN and 50 MET for testing, and re- maining subjects for training (all with reports, while 50 MEN and 50 MET has segmentation masks). The LLM parser was implemented using Llama 3.1 70B as in [4] with prompts engineered to e xtract the hierarchical attributes, uncertainty weights, and cohort priors. W e used the 3D nnU-Net framework as our base segmen- tation architecture due to its strong performance. The model was pre-trained on BraTS2018 for Glioblastoma (dif ferent tu- mor from MEN or MET) with the supervised CE loss for labeled substructures of TC, ET , and ED [1]. No report is T able 1 . Dice Score on the held-out test sets. Method T est Set WT (DSC) TC (DSC) ET (DSC) Masks-Only MEN 0.481 0.323 0.370 R-Super [4] MEN 0.452 0.301 0.353 MS-RSuper(Ours) MEN 0.554 0.428 0.489 Masks-Only MET 0.420 0.385 0.321 RSuper [4] MET 0.443 0.391 0.333 MS-RSuper(Ours) MET 0.529 0.494 0.452 av ailable. Loss weights were set empirically as w r = 0 . 2 , w size = 1 . 0 , w count = 0 . 5 , and w prior = 0 . 2 . W e compare three methods: (1) Masks-Only (Base- line): fine-tuned only on the 100 labeled scans ( D M ). (2) R-Super [4] finetuned on D M ∪ D R , using the summed volume and count applied to the ”Whole T umor” (WT = ET+ED+TC) prediction. (3) Ours MS-RSuper: multimodal with substructure supervised by L report . As shown in T able 1, our method largely outperforms both baselines. The Masks-Only model suf fers from poor general- ization, as expected from only 50 labels in each disease. The RSuper [4] baseline dose not provides improv ement, since its symmetric “summed volume” loss is confused by the par- tial reports (e.g., only the largest lesion), leading to subopti- mal performance. Our MS-RSuper achiev es the highest Dice scores across all substructures and both cohorts. The gains are promising in the MET dataset, where our L count (handling “multiple” lesions) and our qualitative losses ( L exist ) (which align ’edema’ to P E D and ‘enhancement’ to P E T ) are critical. For the MEN dataset, L prior (enforcing extra-axial location) was ke y to reducing false positi ves in the brain parenchyma. An ablation study (T able 2) on the MET dataset confirms that each of our proposed loss components contributes to the final performance. The quantitativ e, partial-cue losses ( L exist ) provide the first major boost by handling size and count. Adding the qualitati ve modality-aligned losses ( L global ) fur- ther improves performance by correctly using the T1c and FLAIR cues. Finally , the cohort prior ( L prior ) provides an additional gain by penalizing anatomically implausible pre- dictions. 4. CONCLUSION W e introduced a novel report-supervised learning framework tailored for the complexities of multi-parametric brain MRI and substructure segmentation. Unlike prior work on CT , our method addresses three ke y challenges: (1) it aligns qualita- tiv e modality-specific findings with their corresponding seg- mentation substructures using novel existence and absence losses, (2) it uses one-sided, uncertainty-aware losses to ro- bustly handle partial quantitative reports (e.g., “largest lesion only , ” “multiple”), and (3) it integrates cohort-lev el anatom- ical priors (intra- vs. extra-axial) deri ved from report key- words. W e ev aluate on a large dataset of 1238 meningioma and metastases scans, our approach largely outperformed both T able 2 . Ablation study on the BraTS-MET T est set (WT Dice). Each component of our proposed loss ( L exist , L global , L prior ) provides a cumulati ve benefit. Method WT (DSC) Masks-Only (Baseline) 0.420 + L exist (Partial size/count) 0.475 + L exist + L global (Adds qualitativ e) 0.513 + L exist + L global + L prior (Full MS-RSuper) 0.529 a sparsely-supervised baseline and a nai ve application of ex- isting RSuper methods. This work demonstrates that by de- signing losses that faithfully reflect the hierarchical and often- incomplete nature of radiology reports, we can ef fectiv ely lev erage large-scale text data to improve multi-class segmen- tation in multimodal imaging. 5. COMPLIANCE WITH ETHICAL ST AND ARDS This retrospectiv e study used open-access human subject data; no additional ethical approv al was required. 6. A CKNO WLEDGMENTS Supported in part by NIH R21EB034911 and NVIDIA Aca- demic Grant Program. 7. REFERENCES [1] Xiaofeng Liu, Helen A Shih, Fangxu Xing, Emiliano Santarnecchi, Georges El F akhri, and Jonghye W oo, “In- cremental learning for heterogeneous structure segmenta- tion in brain tumor mri, ” in MICCAI . Springer , 2023. [2] Y ongtao Zhang, Haimei Li, Jie Du, Jing Qin, Tianfu W ang, Y ue Chen, Bing Liu, W enwen Gao, Guolin Ma, and Baiying Lei, “3d multi-attention guided multi-task learning network for automatic gastric tumor segmenta- tion and lymph node classification, ” IEEE TMI , 2021. [3] Louis Blankemeier , Joseph Paul Cohen, Ashwin Ku- mar , Dave V an V een, Syed Jamal Safdar Gardezi, Mag- dalini P aschali, Zhihong Chen, Jean-Benoit Delbrouck, Eduardo Reis, Cesar T ruyts, et al., “Merlin: A vision lan- guage foundation model for 3d computed tomography , ” Resear ch Squar e , pp. rs–3, 2024. [4] Pedro RAS Bassi, W enxuan Li, Jieneng Chen, Zheren Zhu, Tian yu Lin, Sergio Decherchi, Andrea Cav alli, Kang W ang, Y ang Y ang, Alan L Y uille, et al., “Learn- ing segmentation from radiology reports, ” in MICCAI . Springer , 2025. [5] Jiayu Lei, Xiaoman Zhang, Chaoyi W u, Lisong Dai, Y a Zhang, Y anyong Zhang, Y anfeng W ang, W eidi Xie, and Y uehua Li, “ Autorg-brain: Grounded report genera- tion for brain mri, ” , 2024.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment