Protein Language Models Diverge from Natural Language: Comparative Analysis and Improved Inference
Modern Protein Language Models (PLMs) apply transformer-based model architectures from natural language processing to biological sequences, predicting a variety of protein functions and properties. However, protein language has key differences from n…
Authors: Anna Hart, Chi Han, Jeonghwan Kim
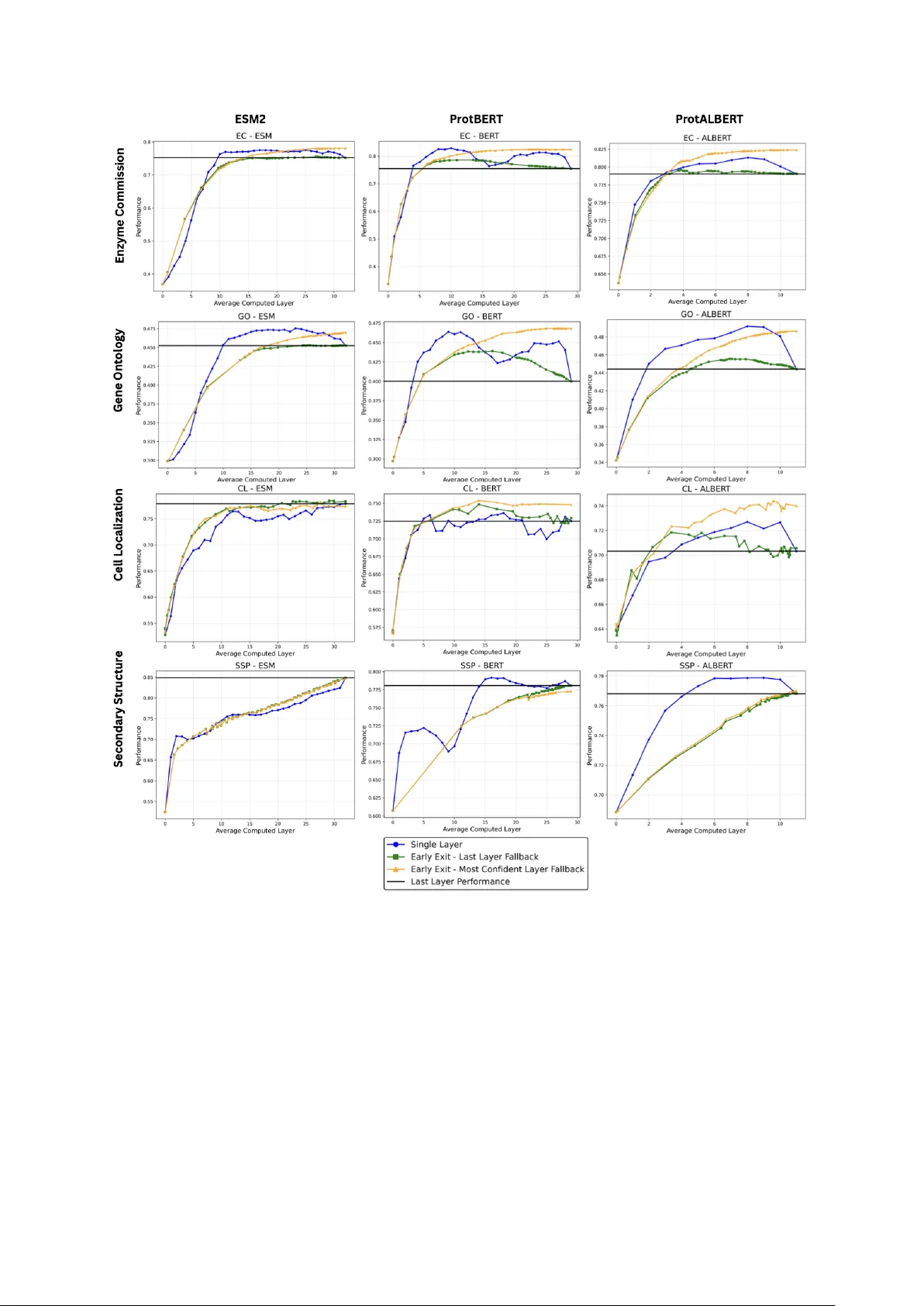
Protein Language Mo dels Div erge from Natural Language: Comparativ e Analysis and Impro v ed Inference Anna Hart 1, 2 , Chi Han 1 , Jeongh w an Kim 1, 2 , Huimin Zhao 2, 3 , and Heng Ji 1, 2, * 1 Sieb el Scho ol of Data Scienc e and Computing, University of Il linois Urb ana-Champ aign, 201 N Go o dwin Ave, Urb ana, IL 61801, USA 2 DOE Center for A dvanc e d Bio ener gy and Biopr o ducts Innovation, University of Il linois Urb ana-Champ aign, 1206 W Gr e gory Drive, Urb ana, IL 61801, USA 3 Chemic al and Biomole cular Engine ering, University of Il linois Urb ana-Champ aign, 600 S Matthews Ave, Urb ana, IL 61801, USA * Corr esp onding author: hengji@illinois.edu Abstract 0.1 Motiv ation Mo dern Protein Language Models (PLMs) apply transformer-based model architectures from natural language pro cessing to biological sequences, predicting a v ariety of protein functions and prop erties. How ever, protein language has k ey differences from natural lan- guage, such as a ric h functional space despite a v o cabulary of only 20 amino acids. These differences motiv ate research in to how transformer-based arc hitectures op erate differently in the protein domain and ho w w e can b etter leverage PLMs to solve protein-related tasks. 0.2 Results In this work, w e b egin by directly comparing how the distribution of information stored across la yers of attention heads differs b et ween the protein and natural language domain. F urthermore, w e adapt a simple early-exit technique—originally used in the natural language domain to improv e efficiency at the cost of p erformance—to achiev e b oth increased accuracy and substantial efficiency gains in protein non-structural prop erty prediction b y allowing the mo del to automatically select protein representations from the in termediate lay ers of the PLMs for the sp ecific task and protein at hand. W e achiev e p erformance gains ranging from 0.4 to 7.01 p ercen tage p oin ts while simultaneously improving efficiency by ov er 10% across mo dels and non-structural prediction tasks. Our work op ens up an area of researc h directly comparing how language mo dels change b eha vior when mo v ed in to the protein domain and adv ances language modeling in biological domains. 0.3 A v ailabilit y and Implementation Co de is av ailable at https://gith ub.com/ahart34/protein with instructions on downloading data. 0.4 Con tact Corresp onding author Heng Ji, hengji@cs.illinois.edu Keyw ords: Bioinformatics, Natural language pro cessing, Protein function prediction, Pro- tein prop ert y prediction, T ransformers 1 1 In tro duction Proteins encode v ast amoun ts of biological information in their sequences. Inspired b y the success of natural language models (NLMs)(V aswani et al., 2017; Peters et al., 2018; Bro wn et al., 2020; Clark et al., 2020; Devlin et al., 2019; Zhuang et al., 2021; Dai et al., 2019; Y ang et al., 2019; Lan et al., 2020; Raffel et al., 2020), protein language mo dels (PLMs) harness the information found in v ast databases of protein sequences to predict protein function and prop erties. PLMs ha ve a v ariety of applications imp ortan t to protein science and engineering suc h as generating proteins with desired prop erties (Madani et al., 2023; Liu et al., 2025; T ruong Jr and Bepler, 2023; Lv et al., 2020), predicting mutational effects (Brandes et al., 2023; T ruong Jr and Bepler, 2023), predicting protein prop erties (Xu et al., 2022; Brandes et al., 2023; Elnaggar et al., 2022; Lin et al., 2023; Rao et al., 2019), and predicting metab olic-engineering constan ts (Boorla and Maranas, 2025). Man y curren t PLMs apply the same arc hitectures and techniques that w orked in natural language (NL) to proteins (Shuai et al., 2023; Elnaggar et al., 2022; Lin et al., 2023). Sev eral new er mo dels (Heinzinger et al., 2024; Lup o et al., 2022; Ha yes et al., 2025) primarily address the a v ailabilit y of additional data types (e.g., incorp orating structural data or m ultiple sequence alignmen ts). Ho wev er, this do es not address the fundamen tal differences in how transformer arc hitectures enco de protein sequences compared to natural language sequences. In this w ork, w e sp ecifically lo ok at the differences in how enco der-based PLMs (Elnaggar et al., 2022; Lin et al., 2023) build a representation of the protein in the mo del differen tly than how enco der-based NLMs do for natural language. Suc h behavioral differences are lik ely giv en the fundamen tal differences b et ween NL and protein language. F or example, NL text often has a long length and a large v ariet y of tok ens (which are analogous to words): for example, GPT-4 handles an input of up to 32,768 tokens (Op enAI et al., 2023) and Llama (T ouvron et al., 2023) understands 32,000 differen t tokens. By contrast, most PLMs consider each amino acid as a tok en, resulting in sligh tly o ver 20 tokens, and the a verage protein input is only ab out 300 amino acids long (Alb erts et al., 2002). F urthermore, while the meaning of NL comes from h uman-made conv en tions, protein function is determined b y complex ph ysical and c hemical interactions within the context of a biological system. Sev eral previous approaches seek to understand how PLMs op erate. In Vig et al. (2021), atten tion within PLMs is linked to biological features within the protein, such as binding sites, to demonstrate whic h protein features affect the protein’s represen tation within the PLM. Si- mon and Zou (2024) study the biological concepts enco ded in PLM neurons. Li et al. (2024) in vestigates transfer learning in PLMs and demonstrates that ESM2 (Lin et al., 2023) mo del p erformance saturates in the middle lay ers for non-structural tasks. While yielding v aluable insigh ts, these studies do not directly compare internal mechanisms of PLMs and NLMs, leaving a gap in our understanding of ho w LM b eha vior c hanges when moving from the NL to protein domain. By finding wa ys in which LM b eha vior is differen t in the protein domain, we can unlo c k new opp ortunities for the researc h communit y to develop domain-sp ecific, biologically grounded language mo dels for protein data. The innate differences b etw een proteins and natural language inspire us to in vestigate the implications of these differences in the mo dels. T o understand these differences, we address t wo k ey questions: (i) How do the in ternal mec hanisms of NLMs and PLMs differ when pro cessing NL and protein input sequences? (ii) Ho w do we better leverage the laten t information embedded in the intermediate la yer representations of PLMs? T o b egin addressing these broader questions, w e conduct tw o targeted inv estigations: examining how the distribution of information stored in atten tion mechanisms differs b et w een NLMs and PLMs through a direct comparison and demonstrating ho w an inference-time early-exit technique (Sc h wartz et al., 2020) lev erages the information stored in PLM lay er represen tations, offering p erformance b enefits not seen with the tec hnique in NLMs. T o our kno wledge, this is the first w ork to directly compare internal 2 atten tion represen tations betw een NLMs and PLMs and to emplo y early-exit in PLMs, opening up new av en ues for translating adv ances in NLP in to the biological domain. 2 Metho ds 2.1 A tten tion Analysis 2.1.1 Preliminaries In this w ork, w e explore differences b et ween the enco ders of PLMs and NLMs b y analyzing infor- mation the attention heads fo cus on. Encoder mo dels, such as BER T (Devlin et al., 2019), follow the transformer enco der arc hitecture (V asw ani et al., 2017). The input is first tok enized into pieces (analogous to words), and these tok ens are passed together through lay ers of feed-forward net works and m ulti-head self-atten tion mec hanisms. In each lay er, feed-forward netw orks incor- p orate representations of individual tok ens while self-attention mechanisms learn asso ciations b et w een tokens and pass information betw een related tok ens. The self-atten tion mechanism computes attention weigh ts using A ttention( Q, K, V ) = softmax QK T √ d k V where QK ⊤ pro vides the relatedness b et w een the query , Q (representation of the token seeking context) and the keys, K, (representation of the surrounding tok ens providing context). The softmax normalizes the re- latedness v alues into attention w eights, and the v alue v ectors (V) pass the information from the con textual tok ens to influence the representation of the query token. The attention mec hanisms capture the con text of eac h token: ho w eac h tok en’s meaning is influenced b y the surrounding tok ens. A ttention mec hanisms assign w eight to surrounding tok ens by their relev ance to the k ey token. Each la yer contains m ultiple attention heads (atten tion computations o ccurring in parallel) which represen t different relationship patterns. The goal of enco der-based mo dels is to dev elop a goo d represen tation of the input sequence, which enco des laten t features that are conduciv e to enabling b etter prediction in downstream tasks with fine-tuning. F or further ex- planation of NLMs and attention mechanisms, we refer readers to the original transformer pap er (V aswani et al., 2017) and survey papers suc h as (Minaee et al., 2024). PLMs (Elnaggar et al., 2022; Lin et al., 2023; Xiao et al., 2025) often follow similar architectures to NLMs; how ever, the tokens in PLMs are typically individual amino acids. In this study , w e explore differences in the b eha vior of attention mechanisms betw een NLMs and PLMs; sp ecifically , we compute the imp ortance of p ositional and seman tic information, explained next, in each atten tion head. 2.1.2 Approac h T o examine whether PLM atten tion heads determine relationships b etw een amino acids differ- en tly than NLM atten tion heads do for words, we first seek a method for directly analyzing and comparing the information stored in atten tion heads. In order to perform a one-to-one compar- ison, we need to detect concepts within attention heads that are applicable to b oth proteins and NL. Atten tion in NLMs can b e brok en down in to focusing on semantic and p ositional informa- tion: where p osition is the lo cation of tokens in the sequence and semantics is the contextualized meaning of the tokens (Han and Heng, 2025). These concepts can b e directly translated into protein language, where p ositional information refers to the distance b et ween the target token and con textual amino acids in the primary protein sequence, and semantics relates to the rep- resen tation of the amino acid, incorp orating the amino acid’s identit y and surrounding con text of the amino acid in the sequence. W e adapt the method by Han and Heng (2025) to disentan- gle the matrix of attention logits into positional and semantic comp onen ts. Sp ecifically , logits w ( i − j, q i , k j ) are found to be appro ximated by the follo wing form: w ( i − j, q i , k j ) ≈ a ( i − j ) + b ( k j ) + c ( q i ) 3 where a represents the con tribution of the relative p osition of the key (k) at position j and query (q) at p osition i , b represents the con tribution of the contextualized information of the k ey , and c represen ts contribution of the current query tok en. The original metho d (Han and Heng, 2025) w as designed for generativ e autoregressive mo dels, where eac h token only attends to previous tokens ( j ≤ i ). W e relax this constrain t to bidirectional PLMs. The approximation is solved by linear regression. As a result, the v ariance of sequence a = [ a ( i − j )] L i − j =0 w ould indicate the imp ortance of p ositional patterns, while the v ariance of b = [ b ( k j )] L j =0 indicates the con tribution of semantic comp onen t. W e compute the ratio of p ositional to semantic information as v ar( positional comp onent ) v ar( seman tic comp onen t ) . The ratio of p ositional to semantic information con veys ho w m uc h the tok en (e.g., amino acid)’s position in the sequence influences its w eigh ting in the atten tion head v ersus how m uch that tok en’s identit y and context influences the weigh ting. 2.2 Early-Exit 2.2.1 Preliminaries Since the internal atten tion mechanisms pro cess protein sequences differently than natural lan- guage sequences, we not only analyze these mec hanistic differences but also inv estigate how w e can better leverage the mo del’s in termediate represen tations of proteins for do wnstream tasks. As findings (Li et al., 2024) hav e demonstrated that performance saturates in middle la yers for non-structural tasks in ESM2, w e explore an early-exit metho d (Sch w artz et al., 2020) as a viable tec hnique for improving m ultiple PLMs - ESM2 (Lin et al., 2023), ProtBER T, (Elnaggar et al., 2022) and ProtAlBER T (Elnaggar et al., 2022). T ypically , a do wnstream task uses the last lay er of the pre-trained mo del to make its prediction. How ev er, some inputs are simpler than others and ma y not need the full last-lay er representation; as such, early-exit detects when the mo del is confident enough to mak e a prediction from an earlier la yer and therefore allows "easier" inputs to exit the pretrained mo del so oner, as surv eyed in (Rahmath P et al., 2024). In NLMs, simple early-exit metho ds typically struggle to match final-lay er p erformance in NL tasks, with approaches lik e the metho d of Sch w artz et al. showing efficiency gains at the cost of reduced accuracy (Sc hw artz et al., 2020). W e adapt this straightforw ard early-exit approac h across PLM mo dels to test whether simple early-exit can ac hieve b etter p erformance and efficiency on protein tasks b y leveraging the in termediate la y er representations of PLMs. 2.2.2 Approac h First, w e attac h a m ulti-la yer p erceptron (MLP) with a single hidden lay er on top of eac h PLM la yer, following the approac h in Zhang et al. (Zhang et al., 2024) and Xu et al.(Xu et al., 2022), whic h lev erages the T orchDrug framew ork (Zhu et al., 2022). Each MLP is trained to predict the task lab el from the protein representation at its corresp onding PLM lay er. Adapted from the natural language metho d in Sch w artz et al. (Sc hw artz et al., 2020), early-exit inference pro ceeds as follows. Beginning at la yer l = 0 , we pass the protein representation at la yer l through the MLP attached to that lay er (denoted as MLP l ). W e then use the maximum predicted class probabilit y from MLP l as the confidence score. If the confidence score at lay er l exceeds a predefined threshold t the output of lay er l ’s MLP is used as the prediction and execution is ceased. If the confidence score at la y er l do es not exceed threshold t , computation pro ceeds to lay er l + 1 , and the pro cedure rep eats. This pro cess con tinues until either the confidence threshold is met or the final PLM lay er is computed. F or our analysis w e iterate o v er a range of thresholds, though in practice a single threshold could be c hosen from a v alidation set. If no lay er exceeds the confidence threshold, w e consider t w o fallbac k strategies. In standard NLP settings including (Sc h wartz et al., 2020) the output of the final la yer is t ypically used as fallbac k, we refer to this as L ast L ayer F al lb ack . Ho wev er, prior w ork on PLMs (Li et al., 2024) 4 has observ ed that in termediate lay ers can yield stronger performance on certain protein-related tasks. T o account for this, we create a Most Confident L ayer F al lb ack , in which confidence scores are recorded across all lay ers and the prediction from the lay er with the highest confidence is selected. A schematic of early-exit is shown in Figure 1. Pseudo code for the mo dified early-exit is shown in Algorithm 1. Algorithm 1 A dapted Early-Exit Algorithm Giv en: Protein sequence x , Confidence Threshold t , F allback Strategy S Output: T ask prediction 1: M axC onf ← 0 2: M ostC onf identP r ed ← None 3: h ← Embed ( x ) 4: for lay er l = 0 to L do 5: h ← PLM l ( h ) 6: log its l ← MLP l ( h ) 7: conf ← max( Sigmoid ( l og its l )) 8: pr ed ← Predict ( log its l ) 9: if conf > t then 10: return pr ed ▷ Threshold met: Exit 11: end if 12: if conf > M axC onf then 13: M axC onf ← conf 14: M ostC onf identP r ed ← pr ed 15: end if 16: end for ▷ F allback Strategies 17: if S = L ast L ayer F al lb ack then 18: return pr ed 19: end if 20: if S = Most Confident L ayer F al lb ack then 21: return M ostC onf identP r ed 22: end if 3 Results and Discussion 3.1 A tten tion Analysis 3.1.1 Exp erimen ts W e collect 1,000 random proteins from UniProtKB/SwissProt, a high-quality subset of UniProt (The UniProt Consortium et al., 2023) and 1,000 random text excerpts from a random subset (Dongk eyun Y o on, 2023) of SlimPa jama (Sob olev a et al., 2023) describ ed in (Shen et al., 2024), whic h is a div erse NL corpus spanning w eb text, b o oks, Github, ArxiV, Wikip edia, and Stack- Exc hange. F or four encoder architectures (BER T (Devlin et al., 2019), AlBER T (Lan et al., 2020), T5 encoder (Raffel et al., 2020), and XLNet (Y ang et al., 2019)), we compute attention heads across all la yers using b oth the ab o ve pretrained NLMs for all NL sequences and their cor- resp onding PLMs (ProtBER T, ProtAlBER T, ProtT5, and ProtXLNet (Elnaggar et al., 2022)) for all protein sequences. Our mo dification of (Han and Heng, 2025) is used to decomp ose each attention head into p ositional, semantic, and residual comp onen ts, as discussed in Section 2.1.2. In T able 2 we confirm that the decomp osed components can b e reconstructed into a matrix with sufficien t 5 Figure 1: The sc heme for early-exit, based on Sc hw artz et al. (Sch w artz et al., 2020). The input protein sequence is fed in to the PLM. A t each la y er, an MLP mak es a prediction for the downstream task and the confidence of this prediction is calculated. When the confidence reac hes a predetermined threshold, the model will output the result from the current la yer and cease further execution. 6 similarit y to the original atten tion matrix, indicating that the attention decomp osition explain a ma jor p ortion of what the self-atten tion mechanism enco des. The ratio of p ositional-to-seman tic information was calculated as in Section 2.1.2. T o statistically analyze the input-dep enden t, head-dependent, and la yer-dependent v ariance of the atten tion fo cus, w e estimate the p opulation v ariance for eac h v ariable across 10 disjoin t subsets of 100 inputs eac h, and we provide the mean and standard deviation of these v ariance estimations. 3.1.2 Findings Our analysis reveals that BER T, AlBER T, and T5 con tain a greater input-dep enden t v ariance in the ratio of p ositional:seman tic atten tion fo cus in the PLM model versus the corresponding NLM. W e visualize this v ariation with a heatmap, sho wn in Figure 2, whic h bins attention heads across 1,000 inputs by their p ositional:seman tic attention information ratio. Qualitativ ely , we observ e a wider distribution of p ositional:seman tic atten tion in the PLMs than the NLMs for the BER T, AlBER T, and T5 architectures - b oth within and b et ween lay ers. T o in v estigate what v ariables this v ariation comes from, we run statistical analysis to quan tify the amount of v ariation in the attention ratio on an input-level, head-level, and la yer-lev el basis, as described in T able 3.1.1. W e find that the PLMs for arc hitectures BER T, AlBER T, and T5 indeed hav e a higher v ariability in atten tion ratio with respect to all three v ariables: the protein input, the atten tion head, and the mo del la y er, as shown in T able 1. These findings indicate that protein language mo dels exhibit a greater degree of v ariability in ho w attention heads contain p ositional v ersus seman tic information. While XLNet does not exhibit the same pattern, its p erm utation of training inputs is exp ected to alter p ositional information and render it an outlier in our analysis. One p ossible explanation for this finding is that the protein language has a v ery small v o cabu- lary: approximately 20 tokens - one for each amino acid - compared to the h undreds of thousands of tokens in NL. Despite this limited vocabulary , proteins still cov er a large functional space: eac h sequence enco des complex information that determines the protein’s structure, function, and prop erties. Muc h of the information in proteins comes from ph ysical and c hemical interac- tions b et ween amino acids in the sequence. As such, it is not surprising that a PLM would need increased flexibility in its atten tion mec hanisms to prop erly enco de the complex relationships b et w een amino acids in the protein. More broadly , patterns of interaction among amino acids ma y b e more complex than patterns of interaction among words in natural language, which are often guided b y grammar and sentence structure, leading to greater v ariation in attention mec hanisms in PLMs. Because it is not feasible to artificially construct protein languages with differen t prop erties while preserving v alid protein sequences, w e leav e testing these hypotheses to future work. A dditionally , the increased v ariabilit y in attention mechanisms across lay ers and inputs in PLMs suggests that early-exit mechanisms could b e esp ecially beneficial by allo wing the mo del to select different la yers for differen t inputs, a method that w e study next. 3.2 Early-Exit 3.2.1 Exp erimen ts W e p erform early-exit in multiple PLMs - ESM2 (Lin et al., 2023), ProtBER T (Elnaggar et al., 2022), and ProtAlBER T (Elnaggar et al., 2022)- for three non-structural classification tasks: gene on tology-biological pro cess, enzyme commission, and subcellular lo calization, and one struc- tural classification task: secondary structure prediction. Gene on tology - biological pro cess (GO) (Ash burner et al., 2000) iden tifies the biological pro cess that the protein plays a role in, and Enzyme Commission (EC) (Bairo c h, 2000) identifies the types of c hemical reactions that an 7 Figure 2: Many PLMs displa y more v ariabilit y in their atten tion fo cus than the cor- resp onding NLM. The heat map displa ys ho w attention heads distribute their fo cus b et w een p ositional and seman tic information across 1,000 inputs, plotting eac h head for eac h input by ratio of p ositional to semantic information fo cus. These plots are generated for NLMs BER T (Devlin et al., 2019), AlBER T (Lan et al., 2020), T5 enco der (Raffel et al., 2020), and XLNet (Y ang et al., 2019) and their corresp onding PLMs (Elnaggar et al., 2022). The y axis represen ts the ratio of p ositional:seman tic information captured by the attention heads, and the color rep- resen ts the num b er of atten tion heads in that lay er p er ratio bin. All attention heads, for each of 1,000 inputs, are accounted for in eac h la y er. As sho wn in the figure, more v ariability in the atten tion fo cus is display ed in the protein v ersions of BER T, ALBER T, and T5, with XLNet as an exception. enzyme can catalyze. Subcellular lo calization (CL) denotes the organelle in a euk ary otic cell where the protein is found and structural classification (SSP) giv es the t yp e of secondary struc- ture that each amino acid is found in. W e c hose GO to test PLM p erformance on learning the functions of diverse proteins, whereas EC illustrates the PLM’s ability to learn to predict the functions within a sp ecific type of proteins. The EC and GO sets are sourced from (Gligorijević et al., 2021), and we use the split with a maxim um of 95% sequence similarit y betw een the training and testing set as giv en b y (Zhang et al., 2024). W e use the PEER b enc hmark (Xu et al., 2022) for the CL and SSP datasets, with the CL dataset containing a maximum of 30% sequence similarit y betw een the training and testing set, sourced from (Almagro Armen teros et al., 2017) and the SSP dataset con taining a testing set sourced from (Klausen et al., 2019) and a training set sourced from (Cuff and Barton, 1999) with a maximum of 25% sequence similarity b et w een the training and testing set. In the ev en t no la yers meet the confidence threshold, early-exit assumes that the final la y er t ypically mak es the b est predictions, and th us selects the final lay er if no earlier lay ers meet the confidence threshold - denoted L ast L ayer F al lb ack (Sc hw artz et al., 2020). How ev er, we find that in the non-structural classification tasks of GO, EC, and CL, p erformance in the middle la yers can outp erform the last la yer by several percentage points across ESM2, ProtBER T, and ProtAlBER T, consistent with observ ations regarding ESM2 non-structural tasks in (Li et al., 2024). As suc h, we provide a simple yet effective mo dification to the early-exit method: in cases where no la yer meets the confidence threshold, we select the most confiden t la y er, an ywhere in the mo del Most Confident L ayer F al lb ack . F or each dataset, we p erform early-exit with multiple confidence thresholds and calculate the performance and a v erage num b er of computed lay ers for eac h. W e directly use the predicted probabilities as the confidence metric to reduce the need for training of an additional parameter, as has b een used in NLP metho ds suc h as (Berestinzshevsky and Ev en, 2019). The mo del predicted probability is calculated as the maxim um logit probability 8 Model Input-Dependent V ariance Lay er-Dependent V ariance Head-Dependent V ariance NLM PLM NLM PLM NLM PLM NLM PLM BER T ProtBER T 0.493 ( ± 0.040) 1.262 ( ± 0.095) 2.973 ( ± 0.034) 7.317( ± 0.167) 2.412 ( ± 0.041) 4.620( ± 0.099) ALBER T ProtALBER T 0.288 ( ± 0.021) 0.752 ( ± 0.075) 2.040 ( ± 0.010) 2.986( ± 0.038) 2.056 ( ± 0.019) 3.851 ( ± 0.044) T5 ProtT5-UniProt 0.440 ( ± 0.021) 0.878 ( ± 0.042) 1.456( ± 0.010) 2.567 ( ± 0.033) 2.658 ( ± 0.015) 3.438 ( ± 0.023) XLNet ProtXLNet 0.828 ( ± 0.068) 0.451 ( ± 0.033) 3.459 ( ± 0.079) 2.390 ( ± 0.025) 2.464 ( ± 0.062) 1.732 ( ± 0.017) T able 1: Man y PLMs sho w greater input, lay er, and attention-head dep enden t v ari- abilit y in attention fo cus than the corresp onding NLM. This table reports the mean and standard deviation of the estimated v ariance in attention ratio (semantic vs. p ositional) from 10 disjoin t samples of 100 inputs each. These statistics are computed across NLMs BER T (Devlin et al., 2019), AlBER T (Lan et al., 2020), T5 enco der (Raffel et al., 2020), and XLNet (Y ang et al., 2019) and their corresp onding PLMs (Elnaggar et al., 2022). PLMs ProtBER T, ProtAlBER T, and ProtT5 hav e higher input, lay er, and attention-head dep endent v ariance than the correp onding NLMs—BER T, ALBER T, and T5, with XLNet as an exception NLP Mo del Correlation Protein Mo del Correlation Mo del Co efficien t Mo del Co efficien t BER T 0.770 ( ± 0.029) ProtBER T 0.733 ( ± 0.050) ALBER T 0.903 ( ± 0.011) ProtALBER T 0.769 ( ± 0.034) T5 0.601 ( ± 0.032) ProtT5-UniProt 0.708 ( ± 0.037) XLNet 0.638 ( ± 0.053) ProtXLNet 0.7047( ± 0.025) T able 2: Correlation of original attention matrix with of attention matrix reconstructed from 3-comp onen t decomp osition for NLMs BER T (Devlin et al., 2019), AlBER T (Lan et al., 2020), T5 enco der (Raffel et al., 2020), and XLNet (Y ang et al., 2019) and their corresponding PLMs (Elnaggar et al., 2022). The results indicate that the decomposed comp onen ts explain a ma jor p ortion of the information enco ded in the self-atten tion. from the MLP for EC, GO, and CL and as the maximum logit probability av eraged across amino acids for SSP . The total n um b er of computed la y ers is used as an indicator of efficiency due to its repro ducibilit y , as is done in Xin et al. (Xin et al., 2021) whic h v alidates that this metric has a linear correlation with w all-time. W e provide the plot b et ween the total num b er of computed la yers and wall-time for ESM2 in Figure 4, v alidating the exp ected linear correlation in the protein domain. W e p erform early-exit for the aforementioned t wo settings: L ast L ayer F al lb ack and Most Confident L ayer F al lb ack . W e compute tw o baselines: single-lay er p erformance, which is the performance of each individual lay er for the dataset, and last-la yer performance whic h is the p erformance of the last lay er. F urthermore, we calculate the calibration of the confidence metric using the Excess AUR C (Geifman et al., 2019), with a binary cross en trop y loss used in the calculation for EC and GO and correctness used in the calculation for CL (Figure 5). 3.2.2 Findings Through our early-exit analysis, we find k ey observ ations in PLM b eha vior that contrast with the observ ations of NLMs described in (Sc hw artz et al., 2020). Notably , w e find that early-exit in PLMs not only greatly improv es efficiency , but also offers p erformance gains across mo dels and non-structural tasks. First, w e demonstrate that the early p erformance saturation observed in ESM2 Li et al. (2024) generalizes to ProtBER T and ProtAlBER T, with middle-lay er p erformance surpassing last-la yer p erformance across mo dels for non-structural tasks. The high middle-lay er perfor- mance allo ws us to refine the early-exit metho d b y designating the most-confident la y er as fallbac k, as describ ed in section 3.2.1. W e compute early-exit performance across multiple pre- defined thresholds and for eac h threshold, w e plot the p erformance versus the av erage computed la yer in Figure 3. Unlik e in the natural language results rep orted in Sc hw artz et al. (Sch w artz 9 et al., 2020), we see p erformance improv emen ts using b oth the L ast L ayer F al lb ack and Most Confident L ayer F al lb ack , with the b est results seen with Most Confident L ayer F al lb ack . Using the most confident lay er as fallback, EC prediction in ESM2 achiev ed last la y er p erformance with a 52.38% efficiency improv emen t and gained 2.85 p ercen tage points in F1 max with a 12.53% efficiency impro v ement. GO prediction in ESM2 ac hiev ed last lay er performance with a 43.94% efficiency impro vemen t and improv ed 1.55 p ercen tage p oin ts in F1 max with a 10.37% efficiency improv emen t. CL prediction in ESM2 ac hiev ed last la yer p erformance with a 16.57% efficiency improv emen t and improv ed accuracy by 0.4 p ercen tage p oin ts with a 16.57% efficiency impro vemen t. F ull results are sho wn in Figure 3. These results demonstrate that early-exit is a viable approac h for not only impro ving algorithm efficiency but also improving p erformance on non-structural tasks across mo dels and applications. Second, in PLMs, the early-exit method outp erforms the performance of the last lay er while not consisten tly outp erforming single-lay er p erformance. This is in con trast to NLMs, where early-exit significan tly outp erforms single-la y er p erformance but fails to outp erform the last-la yer p erformance (Sch w artz et al., 2020). When analyzing the confidence metric using excess A UR C (Geifman et al., 2019), we find that the confidence metric for PLM early-exit is generally w ell- calibrated for EC, impro ves in calibration in later lay ers for CL, and remains po orly calibrated for the GO task. Giv en that this simple early-exit tec hnique already improv es b oth p erformance and efficiency in PLMs, these results demonstrate that further developmen t of early-exit with protein-sp ecific confidence metrics is a promising area of researc h. F urthermore, early-exit do es offer distinct adv antages o v er selecting a single exit lay er: it eliminates the need for lay er selection on a v alidation set and pro vides greater robustness for inference on diverse protein sets, as the early-exit mechanism adapts on a p er-protein basis. Third, while early-exit improv es inference in non-structural tasks (GO, EC, CL), w e find that early-exit do es not make meaningful gains for structural tasks (SSP). This is consistent with observ ations of ESM2 in Li et al. (Li et al., 2024) , where performance of non-structural tasks saturated early but structural tasks did not. Thus, while w e agree with Li et al. that pre-training b etter aligned with non-structural tasks may improv e task p erformance, we additionally sho w that inference-time methods suc h as early exit can impro v e both the efficiency and accuracy of PLMs on non-structural tasks b y leveraging the stronger p erformance of intermediate la y ers. A promising direction for future w ork would be to discov er the primary v ariables leading to exit-decisions, which would lend more in terpretabilit y to mo del decisions and confidence. As w e did not see a meaningful relationship b et w een the ratio of semantic:positional attention in the atten tion heads of a la yer and exit decisions, it is likely that many other v ariables ha ve a larger impact on early-exit decisions. 4 F uture W ork This work fo cuses on enco der-only protein sequence mo dels, understanding ho w proteins are represen ted within a transformer arc hitecture. F uture work could study deco der mo dels such as ProGen (Madani et al., 2023) or multimodal mo dels such as (Hay es et al., 2025) to b etter understand ho w mo dels generate nov el proteins or ho w v arious types of data, such as protein se- quence and structural data, are handled by a mo del. This work fo cuses on analyzing as directly as p ossible differences b et w een NLMs and PLMs by finding concepts - p osition, semantics, and a logit-based static early-exit - that can b e compared across domains. Prior work on interpretabil- it y in PLMs (Vig et al., 2021; Simon and Zou, 2024) fo cuses on protein-sp ecific concepts, suc h as binding sites. A promising future direction of research would b e to unite domain-agnostic comparisons b et ween PLMs and NLMs to domain-sp ecific concepts and b eha viors of the mo dels; for example, b y connecting p ositional and semantic information to biological structures in the proteins. F urthermore, we hope that our analysis of the differences b etw een PLMs and NLMs stresses the imp ortance of not simply transferring NLMs into the protein domain, but instead 10 Figure 3: Early-Exit Impro v es b oth P erformance and Efficiency in Non-Structural T asks across Multiple PLMs. The total n umber of computed lay ers is used as a proxy for efficiency . The trade-offs b et w een model p erformance and efficiency are calculated for: (1) Indi- vidual La yer Performance, (2) Early-Exit L ast L ayer F al lb ack , and (3) Early-exit Most Confident L ayer F al lb ack . The baseline p erformance of the last lay er is drawn across with a blac k line. Computations are done for ESM2 (Lin et al., 2023), ProtBER T, and ProtALBER T (Elnaggar et al., 2022). Early-exit Most Confident L ayer F al lb ack outp erforms b oth the last-lay er p erfor- mance baseline and early-exit L ast L ayer F al lb ack regarding b oth performance and efficiency in non-structural tasks. F or the secondary structure prediction, early-exit allows efficiency gains but harms p erformance. 11 Figure 4: W alltimes v ersus total n um b er of computed la y ers The w alltime for the testing set on 1 V100 GPU v ersus the av erage n umber of computed lay ers is plotted across all models and tasks. Early-exit Most Confident L ayer F al lb ack is plotted. A diamond mark er at the final la yer denotes the baseline walltime. W e see that w alltime corresponds linearly with the num ber of computed lay ers inno v ating new arc hitectures and metho ds to better learn biological knowledge. F or example, a new attention mechanism ma y be needed to better capture the v aried structures and functions of proteins enco ded by a small sequence vocabulary . Ov erall, we b eliev e that b etter understand- ing the differences b et ween in machine learning algorithms in their original domain and their new biological domain will unlo c k promising research directions into adapting and mo difying mac hine learning methods for biology . Comp eting in terests No comp eting in terest is declared. A c kno wledgmen ts This w ork was funded b y the DOE Center for A dv anced Bio energy and Biopro ducts Innov ation (U.S. Department of Energy , Office of Science, Biological and Environmen tal Research Program under A ward Num b er DE-SC0018420). Any opinions, findings, and conclusions or recommen- dations expressed in this publication are those of the author(s) and do not necessarily reflect the views of the U.S. Department of Energy . W e sincerely thank Professor Ge Liu for helpful and v aluable discussions. 12 Figure 5: Confidence calibration . A lo w er excess AUR C score (Geifman et al., 2019) denotes a b etter calibrated confidence metric. W e see that, for all mo dels, confidence is w ell calibrated across la yers for EC, is well calibrated in middle and later la yers for CL, and is p o orly calibrated for GO tasks. References B. Alb erts, A. Johnson, and J. Lewis. The shap e and structure of proteins. In Mole cular Biolo gy of the Cel l. 4th e dition. Garland Science, 2002. J. J. Almagro Armenteros, C. K. Sønderby , S. K. Sønderby , et al. DeepLo c: prediction of protein sub cellular lo calization using deep learning. Bioinformatics , 33(21):3387–3395, 2017. ISSN 1367-4803, 1367-4811. M. Ash burner, C. A. Ball, J. A. Blak e, et al. Gene ontology: to ol for the unification of biology . Natur e Genetics , 25(1):25–29, 2000. ISSN 1061-4036, 1546-1718. doi: 10.1038/75556. A. Bairoch. The ENZYME database in 2000. Nucleic A cids R ese ar ch , 28(1):304–305, 2000. ISSN 13624962. K. Berestinzshevsky and G. Even. Dynamically sacrificing accuracy for reduced computation: Cascaded inference based on softmax confidence. Artificial Neur al Networks and Machine L e arning - ICANN 2019: De ep L e arning: 28th International Confer enc e on Artificial Neur al Networks , 2019. V. S. Bo orla and C. D. Maranas. CatPred: a comprehensive framew ork for deep learning in vitro enzyme kinetic parameters. Natur e Communic ations , 16(1):2072, 2025. ISSN 2041-1723. 13 N. Brandes, G. Goldman, C. H. W ang, et al. Genome-wide prediction of disease v arian t effects with a deep protein language mo del. Natur e Genetics , 55(9):1512–1522, 2023. ISSN 1061-4036, 1546-1718. T. B. Bro wn, B. Mann, N. Ryder, et al. Language mo dels are few-shot learners. In A dvanc es in Neur al Information Pr o c essing Systems 33: Annual Confer enc e on Neur al Information Pr o c essing Systems , 2020. K. Clark, M.-T. Luong, Q. V. Le, et al. ELECTRA: Pre-training text enco ders as discriminators rather than generators. In 8th International Confer enc e on L e arning R epr esentations, ICLR 2020, A ddis Ab ab a, Ethiopia, April 26-30, 2020 . Op enReview.net, 2020. J. A. Cuff and G. J. Barton. Ev aluation and impro vemen t of m ultiple sequence metho ds for protein secondary structure prediction. Pr oteins: Structur e, F unction, and Genetics , 34(4): 508–519, 1999. ISSN 0887-3585, 1097-0134. Z. Dai, Z. Y ang, Y. Y ang, et al. T ransformer-XL: Atten tiv e language mo dels b ey ond a fixed- length con text. In Pr o c e e dings of the 57th Annual Me eting of the Asso ciation for Compu- tational Linguistics , pages 2978–2988. Asso ciation for Computational Linguistics, 2019. doi: 10.18653/v1/P19- 1285. J. Devlin, M.-W. Chang, K. Lee, et al. BER T: Pre-training of deep bidirectional transformers for language understanding. In Pr o c e e dings of the 2019 Confer enc e of the North Americ an Chapter of the Asso ciation for Computational Linguistics: Human L anguage T e chnolo gies, V olume 1 (L ong and Short Pap ers) , pages 4171–4186. Asso ciation for Computational Linguistics, 2019. Dongk eyun Y o on. SlimPa jama-6b. https://huggingface.co/datasets/DKYoon/ SlimPajama- 6B , 2023. A. Elnaggar, M. Heinzinger, C. Dallago, et al. ProtT rans: T ow ard understanding the language of life through self-supervised learning. IEEE T r ansactions on Pattern A nalysis and Machine Intel ligenc e , 44(10):7112–7127, 2022. ISSN 0162-8828, 2160-9292, 1939-3539. Y. Geifman, G. Uziel, and R. El-Y aniv. Bias-reduced uncertaint y estimation for deep neural classifiers. International Confer enc e on L e arning R epr esentations , 2019. V. Gligorijević, P . D. Renfrew, T. K osciolek, et al. Structure-b ased protein function prediction using graph con volutional net w orks. Natur e Communic ations , 12(1):3168, 2021. ISSN 2041- 1723. C. Han and J. Heng. Computation mec hanism b ehind LLM p osition generalization. In Pr o c e e d- ings of the 63r d A nnual Me eting of the Asso ciation for Computational Linguistics (V olume 1: L ong Pap ers) , 2025. T. Hay es, R. Rao, H. Akin, et al. Sim ulating 500 million years of ev olution with a language mo del. Scienc e , 2025. M. Heinzinger, K. W eisseno w, J. G. Sanchez, et al. Bilingual language mo del for protein sequence and structure. NAR Genomics and Bioinformatics , 6(4):lqae150, 2024. ISSN 2631-9268. M. S. Klausen, M. C. Jesp ersen, H. Nielsen, et al. NetSurfP-2.0: Impro v ed prediction of protein structural features b y integrated deep learning. Pr oteins: Structur e, F unction, and Bioinfor- matics , 87(6):520–527, 2019. ISSN 0887-3585, 1097-0134. Z. Lan, M. Chen, S. Go o dman, et al. ALBER T: A lite BER T for self-sup ervised learning of language represen tations. In 8th International Confer enc e on L e arning R epr esentations, ICLR 2020, A ddis Ab ab a, Ethiopia, April 26-30, 2020 . Op enReview.net, 2020. 14 F.-Z. Li, A. P . Amini, Y. Y ue, et al. F eature reuse and scaling: Understanding transfer learning with protein language mo dels. In F orty-first International Confer enc e on Machine L e arning, ICML 2024, Vienna, Austria, July 21-27, 2024 . Op enReview.net, 2024. Z. Lin, H. Akin, R. Rao, et al. Evolutionary-scale prediction of atomic-level protein structure with a language mo del. Scienc e , 379(6637):1123–1130, 2023. ISSN 0036-8075, 1095-9203. S. Liu, Y. Li, Z. Li, et al. A text-guided protein design framew ork. Natur e Machine Intel ligenc e , 2025. U. Lup o, D. Sgarb ossa, and A.-F. Bitb ol. Protein language mo dels trained on multiple sequence alignmen ts learn phylogenetic relationships. Natur e Communic ations , 13(1):6298, 2022. ISSN 2041-1723. L. Lv, Z. Lin, H. Li, et al. Prollama: A protein large language mo del for multi-task protein language pro cessing. Journal of IEEE T r ansactions on Artificial Intel ligenc e , 2020. A. Madani, B. Krause, E. R. Greene, et al. Large language mo dels generate functional protein sequences across div erse families. Natur e Biote chnolo gy , 41(8):1099–1106, 2023. ISSN 1087- 0156, 1546-1696. doi: 10.1038/s41587- 022- 01618- 2. S. Minaee, T. Mikolo v, N. Nikzad, et al. Large language mo dels: A surv ey . ArXiv pr eprint , abs/2402.06196, 2024. Op enAI, J. A c hiam, S. Adler, S. Agarwal, et al. GPT-4 tec hnical rep ort. ArXiv pr eprint , abs/2303.08774, 2023. URL . M. E. Peters, M. Neumann, M. Iyyer, et al. Deep con textualized word represen tations. In Pr o c e e dings of the 2018 Confer enc e of the North A meric an Chapter of the Asso ciation for Computational Linguistics: Human L anguage T e chnolo gies, V olume 1 (L ong Pap ers) , pages 2227–2237. Asso ciation for Computational Linguistics, 2018. C. Raffel, N. Shazeer, A. Rob erts, et al. Exploring the limits of transfer learning with a unified text-to-text transformer. J. Mach. L e arn. R es. , 21:140:1–140:67, 2020. H. Rahmath P , V. Sriv asta v a, K. Chaurasia, et al. Early-exit deep neural netw ork - a compre- hensiv e survey . A CM Computing Surveys, V olume 57, Issue 3 , 2024. R. Rao, N. Bhattachary a, N. Thomas, et al. Ev aluating protein transfer learning with tap e. Neur al Information Pr o c essing Systems , 2019. R. Sch w artz, G. Stanovsky , S. Sw ay amdipta, et al. The right to ol for the job: Matching mo del and instance complexities. In Pr o c e e dings of the 58th Annual Me eting of the Asso ciation for Computational Linguistics , pages 6640–6651. Asso ciation for Computational Linguistics, 2020. Z. Shen, T. T ao, L. Ma, et al. Slimpa jama-dc: Understanding data combinations for llm training. A rxiV pr eprint , 2024. R. W. Sh uai, J. A. Ruffolo, and J. J. Gra y . IgLM: Infilling language mo deling for antibo dy sequence design. Cel l Systems , 14(11):979–989.e4, 2023. ISSN 24054712. E. Simon and J. Zou. In terPLM: Discov ering in terpretable features in protein language mo dels via sparse auto enco ders. ArXiV pr eprint , 2024. D. Sob olev a, F. Al-Khateeb, R. Myers, et al. SlimPa jama: A 627B tok en cleaned and dedu- plicated version of RedP a jama, 2023. URL https://huggingface.co/datasets/cerebras/ SlimPajama- 627B . 15 The UniProt Consortium, A. Bateman, M.-J. Martin, S. Orc hard, et al. UniProt: the univ ersal protein knowledgebase in 2023. Nucleic A cids R ese ar ch , 51:D523–D531, 2023. H. T ouvron, T. La vril, G. Izacard, et al. LLaMA: Op en and efficient foundation language mo dels. A rXiv pr eprint , abs/2302.13971, 2023. T. F. T ruong Jr and T. Bepler. Poet: A generative mo del of protein families as sequences-of- sequences. 37th Confer enc e on Neur al Information Pr o c essing Systems , 2023. A. V asw ani, N. Shazeer, N. P armar, et al. A tten tion is all y ou need. Pr o c e e dings fo the 31st International Confer enc e on Neur al Information Pr o c essing Systems , 2017. J. Vig, A. Madani, L. R. V arshney , et al. BER T ology meets biology: Interpreting attention in protein language mo dels. In 9th International Confer enc e on L e arning R epr esentations, ICLR 2021, Virtual Event, Austria, May 3-7, 2021 . Op enReview.net, 2021. Y. Xiao, W. Zhao, J. Zhang, et al. Protein large language models; a comprehensive survey . A rXiv pr eprint , 2025. J. Xin, R. T ang, Y. Y u, et al. BERxiT: Early exiting for BER T with b etter fine-tuning and extension to regression. In Pr o c e e dings of the 16th Confer enc e of the Eur op e an Chapter of the Asso ciation for Computational Linguistics: Main V olume , pages 91–104. Asso ciation for Computational Linguistics, 2021. doi: 10.18653/v1/2021.eacl- main.8. M. Xu, Z. Zhang, J. Lu, et al. PEER: A comprehensiv e and m ulti-task b enc hmark for protein sequence understanding. In A dvanc es in Neur al Information Pr o c essing Systems 35: A nnual Confer enc e on Neur al Information Pr o c essing Systems 2022, NeurIPS 2022, New Orle ans, LA, USA, Novemb er 28 - De c emb er 9, 2022 , 2022. Z. Y ang, Z. Dai, Y. Y ang, et al. XLNet: Generalized autoregressiv e pretraining for language understanding. In A dvanc es in Neur al Information Pr o c essing Systems 32: A nnual Confer- enc e on Neur al Information Pr o c essing Systems 2019, NeurIPS 2019, De c emb er 8-14, 2019, V anc ouver, BC, Canada , pages 5754–5764, 2019. Z. Zhang, J. Lu, V. Chenthamarakshan, et al. Structure-informed protein language mo del. GEM W orkshop, ICLR , 2024. Z. Zhu, C. Shi, Z. Zhang, et al. T orc hdrug: A p o werful and flexible machine learning platform for drug discov ery . ArxiV pr eprint , 2022. L. Zhuang, L. W a yne, S. Y a, et al. A robustly optimized BER T pre-training approac h with p ost-training. In Pr o c e e dings of the 20th Chinese National Confer enc e on Computational Linguistics , pages 1218–1227. Chinese Information Pro cessing Society of China, 2021. 16
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment