Transfer Learning with Network Embeddings under Structured Missingness
Modern data-driven applications increasingly rely on large, heterogeneous datasets collected across multiple sites. Differences in data availability, feature representation, and underlying populations often induce structured missingness, complicating…
Authors: Mengyan Li, Xiaoou Li, Kenneth D M
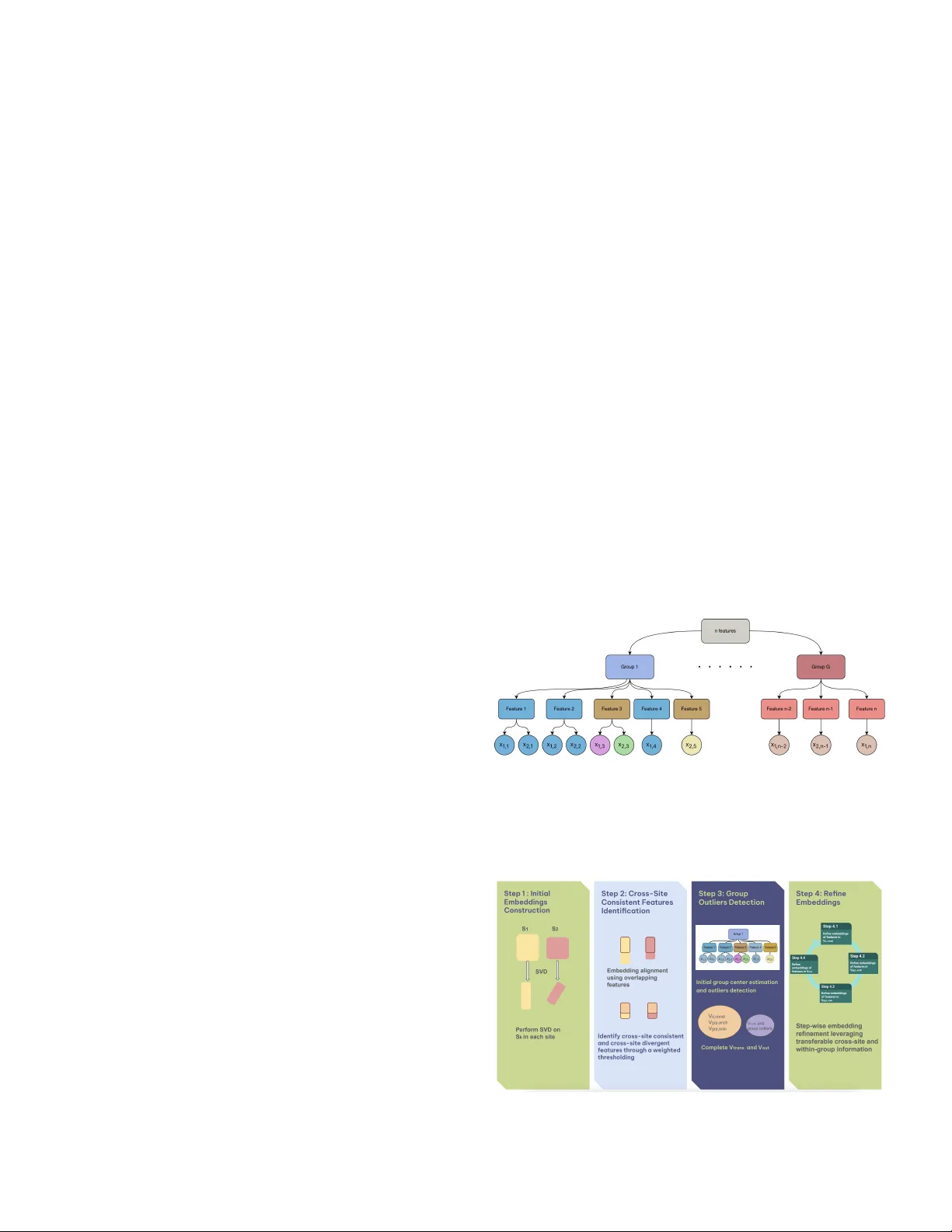
Submitted to Statistical Science T ransfer Learning with Netw ork Embeddings under Structured Missingness Mengy an Li 1 , Xiaoou Li 1 , Kenneth D Mandl, and Tianxi Cai * Abstract. Modern data-driv en applications increasingly rely on lar ge, het- erogeneous datasets collected across multiple sites. Dif ferences in data a vail- ability , feature representation, and underlying populations often induce struc- tured missingness, complicating efforts to transfer information from data-rich settings to those with limited data. Many transfer learning methods ov erlook this structure, limiting their ability to capture meaningful relationships across sites. W e propose T ransNEST ( T rans fer learning with N etwork E mbeddings under ST ructured missingness), a frame work that integrates graphical data from source and target sites with prior group structure to construct and re- fine netw ork embeddings. T ransNEST accommodates site-specific features, captures within-group heterogeneity and between-site dif ferences adapti vely , and improv es embedding estimation under partial feature overlap. W e es- tablish the con ver gence rate for the T ransNEST estimator and demonstrate strong finite-sample performance in simulations. W e apply TransNEST to a multi-site electronic health record study , transferring feature embeddings from a general hospital system to a pediatric hospital system. Using a hier- archical ontology structure, T ransNEST improv es pediatric embeddings and supports more accurate pediatric kno wledge extraction, achieving the best accuracy for identifying pediatric-specific relational feature pairs compared with benchmark methods. K ey wor ds and phr ases: T ransfer Learning, Network Embeddings, Structured Missing Data, Inner Product Model, Electronic Health Record Data. 1. INTRODUCTION 1.1 Backgr ound Modern data-driv en applications increasingly inv olve learning from multiple data sources or sites, where fea- ture sets only partially overlap, and data av ailability v aries across sources. Such settings arise in many domains, including multilingual representation learning, cross- Mengyan Li is Assistant Pr ofessor , Department of Mathematical Sciences, Bentle y University , W altham, USA (e-mail: mengyanli@bentle y .edu ). Xiaoou Li is Associate Pr ofessor , Sc hool of Statistics, University of Minnesota, Minneapolis, USA (e-mail: lixx1766@umn.edu ). K enneth D. Mandl is Pr ofessor , Chair in Biomedical Informatics and P opulation Health, Harvard Medical Sc hool, Boston, USA (e-mail: K enneth.Mandl@childr ens.harvar d.edu ). T ianxi Cai is John Roc k Pr ofessor , Harvar d T .H. Chan School of Public Health, Harvar d University , Boston, USA (e-mail: tcai@hsph.harvar d.edu ). * Tianxi Cai is the Corresponding author 1 Mengyan Li and Xiaoou Li are co-first authors and contributed equally to this work. platform recommendation systems, and multi-site clini- cal studies where measurement protocols and recorded cov ariates differ across centers. A fundamental challenge in these problems is structured missingness induced by heterogeneous feature spaces and population-le vel dif fer- ences across sites, which complicates transfer learning from information-rich sources to data-limited targets. In man y multi-site settings, a subset of features admits kno wn one-to-one correspondences across sites, while the remaining features are site-specific b ut or ganized through a shared group or hierarchical structure. For example, in multilingual representation learning, sites may corre- spond to corpora in different languages with partially ov erlapping vocabularies or kno wn cross-lingual align- ments. In cross-platform recommendation systems, sites may correspond to dif ferent platforms with o verlapping item sets or externally matched items. In both cases, fea- tures are connected through network-structured represen- tations deri ved from co-occurrence or similarity informa- tion, and group structure encodes shared semantic or tax- onomic relationships. 1 2 Another example is multi-site kno wledge extraction from electronic health record (EHR) data. Adv ances in EHR systems have enabled the use of large-scale real- world clinical data for data-driv en biomedical research, supporting tasks such as clinical knowledge extraction, risk prediction, and treatment response assessment [e.g. 10 , 15 , 23 , 29 ]. Despite this promise, the complex struc- ture of EHR data poses substantial modeling challenges. EHR features are high-dimensional and relational, com- prising diagnoses, medications, laboratory tests, and pro- cedures. Representation learning has therefore emerged as an effecti ve approach for modeling EHR data, en- abling low-dimensional embeddings that capture feature relationships and support do wnstream tasks [ 31 ]. Beyond high dimensionality , EHR data exhibit pronounced het- erogeneity across patient subpopulations and healthcare systems. In particular , representations learned from gen- eral or adult populations may transfer poorly to pedi- atric settings due to biological, pharmacological, and clin- ical dif ferences, as well as pediatric-specific v ocabularies [ 14 ]. Additionally , approximately half of pediatric health- care visits are pre venti ve well-checks [ 16 , 19 ], contribut- ing to the unique sparsity of pediatric EHR data. Although shared hierarchical ontology structures can potentially in- crease transferability between adult and pediatric popu- lations, these differences induce structured missingness and site-specific feature behavior , limiting the effecti ve- ness of standard transfer learning methods that assume shared feature spaces. These challenges moti vate transfer learning frame- works that can e xploit network structure, handle partial feature alignment, and incorporate shared group informa- tion across heterogeneous multi-site data. T o this end, we propose a T rans fer learning with N etwork E mbeddings under ST ructured missingness (TransNEST) method, by combining site-specific network embeddings with prior group structure to guide selectiv e information shar- ing. T ransNEST accommodates partially aligned feature spaces by allowing site-specific features, while adapti vely modeling heterogeneity both within groups and across sites. As a result, it enables more efficient and accurate kno wledge transfer to sparse, heterogeneous tar get popu- lations. 1.2 Related Literature and Our Contributions Learning from multi-site data with heterogeneous fea- ture spaces and structured missingness has been stud- ied across sev eral methodological lines, including matrix completion, multi-task learning, representation learning, and transfer learning. While these approaches offer partial solutions, they typically fall short in settings characterized by partial feature alignment, multi-block missingness pat- terns, and complex cross-site heterogeneity , which arise in many real-w orld applications. Classical matrix completion methods [e.g. 7 , 8 , 24 , 27 ] typically assume entries are missing completely at ran- dom, an assumption violated in multi-site settings where missingness is dri ven by systematic dif ferences in feature av ailability across sites. Structured matrix completion re- laxes this assumption for structured missingness, where an entire submatrix block is unobserved [ 6 ], but it does not directly accommodate more general multi-block miss- ingness patterns common in multi-source data. More re- cent work considers matrices with ov erlapping sub-blocks [ 36 ], yet enforces identical embeddings for overlapping features, which can be o verly restrictive when feature se- mantics or relationships dif fer across sites. Related challenges ha ve also been studied from a multi-task learning perspecti ve. For example, [ 32 ] pro- pose a two-step framew ork for heterogeneous multi- source block-wise missing data, addressing task-lev el heterogeneity but operating under a dif ferent modeling paradigm than the network-embedding–based transfer frame work de veloped here. More broadly , multi-task with shared-representation methods, including multi-task PCA and related approaches [e.g. 13 , 22 , 33 ], aim to borro w strength across tasks by learning shared latent structure. Ho wever , these methods generally assume a common fea- ture space and do not accommodate site-specific features or structured missingness induced by heterogeneous data collection processes. Se veral lines of statistical work incorporate external structural information to enhance embedding learning or facilitate kno wledge transfer . [ 5 ] proposes cov ariate- assisted spectral clustering that jointly lev erages graph structure and node cov ariates to recover latent commu- nities. [ 4 ] incorporates group structure among rows and columns, enabling matrix completion under missing not at random mechanisms. Ho wever , these two frameworks are de veloped for single network settings and do not con- sider block-wise missingness induced by heterogeneous feature spaces across multiple sites. [ 30 ] dev elops a spher - ical regression framew ork to learn a mapping between two embedding spaces under mismatches and can in- corporate group information, b ut it imposes strong cor - respondence constraints and does not address multi-site transfer with multi-block missingness. Complementing these statistical approaches, a large body of work studies knowledge transfer through learned representations in deep models. Deep representation learning and transfer learning methods[e.g. 3 , 17 , 18 ] learn transferable features from unlabeled data and adapt source-trained representations to target settings. While re- lated in spirit, these approaches typically do not e xplicitly model structured missingness, and often lack theoretical guarantees. More specialized deep transfer methods, such as distribution alignment [ 37 ] or similarity-induced em- beddings [ 25 ], alleviate some of these issues b ut still rely on restricti ve alignment assumptions. 3 T ransNEST explicitly accommodates site-specific fea- tures, partial feature alignment, and heterogeneous popu- lations by integrating network-structured representations observed at each site with shared group or hierarchical structure. The method adaptively determines which fea- tures should borrow strength across sites and/or from the group, and refines embeddings through a procedure that le verages cross-site and within-group homogeneity , and feature-le vel priors, such as feature-site weights. This se- lecti ve information sharing improv es embedding estima- tion ef ficiency while reducing bias induced by heteroge- neous feature behavior . Importantly , we also dev elop rigorous theoretical guar - antees for T ransNEST estimator under realistic multi-site structures. W e establish error bounds for embedding ac- curacy and pro ve that, under mild conditions, T ransNEST performs no worse than single-site singular v alue de- composition (SVD), ensuring robustness against neg ativ e transfer . With intermediate two-to-infinity norm bounds established, these results connect to and extend recent de- velopments in entrywise error bounds for low-rank mod- els [e.g. 1 , 9 , 11 , 12 ], b ut accommodate a substantially more complex, heterogeneous, and multi-block missing- ness structure. In summary , T ransNEST provides an adaptiv e, theoreti- cally grounded solution to transfer learning with network- structured data under structured missingness, with broad applicability across multi-site and multi-source learning problems. 2. METHOD 2.1 Notation For any positive integer n , let [ n ] = { 1 , . . . , n } . For any set S , let | S | denote its cardinality . For a matrix A = ( a ij ) ∈ R n × p , let A i · and A · j denote the i -th row and the j -th column, respectively , and let τ 1 ( A ) ≥ · · · ≥ τ min( n,p ) ( A ) denote its singular v alues. If A is positiv e semi-definite, then they are also the same as the eigen- v alues. Let ∥ A ∥ F and ∥ A ∥ 2 denote the Frobenius and spectral norm of a matrix A , respecti vely . In addition, ∥ A ∥ 2 →∞ = max i ∈ [ n ] ( P p j =1 a 2 ij ) 1 / 2 denotes the two-to- infity norm of A , which is the largest row norm of A . For any full rank matrix A , let U ( A ) = A ( A T A ) − 1 be its polar factor . Let O r denote the set of all r × r orthogonal matrices. W e further define the operator W ( X , ˜ X ) = arg min Q ∈ O r ∥ ˜ X − X Q ∥ F = U ( X T ˜ X ) , which is known as the orthogonal procrustes problem in the literature with a closed-form solution [ 28 ]. For two sequences a n and b n , we denote by a n = O ( b n ) if lim n →∞ | a n /b n | < ∞ , a n = o ( b n ) if lim n →∞ a n /b n = 0 , and a n = O p ( b n ) or a n = o p ( b n ) if a n = O ( b n ) or a n = o ( b n ) with a probability approaching 1 . W e use a n ≲ b n to denote a n ≤ C b n for some constant C > 0 , and use a n ∼ b n to denote C ≤ a n /b n ≤ C ′ for some constants C, C ′ > 0 . 2.2 Model Setup Let S = ( s k,i,j ) k =1 , 2; i,j ∈ [ n ] denote the underlying data tensor , where k index es the site and n is the total number of distinct features across sites. For each site k , the entry s k,i,j quantifies the relatedness between features i and j . Because the sites may not share identical feature sets, let V k ⊂ [ n ] denote the feature indices observed at each site, with |V k | = n k , k = 1 , 2 . The observed site-specific ma- trix is S k = ( s k,i,j ) i,j ∈ V k ∈ R n k × n k , and for simplicity we assume S k is symmetric. In practice, S k may be con- structed as a Shifted Positi ve Pointwise Mutual Informa- tion (SPPMI) matrix deri ved from feature co-occurrence data at site k [ 20 , 21 ], which provides a widely used rep- resentation of feature similarity based on co-occurrence patterns. W e model S k using a lo w-rank signal plus noise de- composition, S k = M k + E k , M k = X k X ⊤ k = U k D k U ⊤ k , for k = 1 , 2 , where M k ∈ R n k × n k is a symmetric rank- r signal matrix and E k ∈ R n k × n k is a mean-zero sym- metric noise matrix with independent upper-triangular entries. The diagonal matrix D k ∈ R r × r contains the nonzero eigen values of M k , and U k ∈ R n k × r contains the associated eigen vectors. W e define X k = U k D 1 / 2 k = ( x k, 1 , . . . x k,n k ) ⊤ ∈ R n k × r as the population-lev el em- bedding matrix for the features observed at site k , and x k,i ∈ R r denotes the true embedding of feature i at site k . This formulation is motiv ated by the fact that the SVD of SPPMI matrices recovers skip-gram-with- negati ve-sampling embeddings [ 20 ], providing a princi- pled interpretation of X k as the latent feature embeddings at site k . In addition to the site-specific matrices S 1 and S 2 , we lev erage an external grouping structure shared across sites. Let G denote the total number of groups, and let g i ∈ [ G ] indicate the group membership of feature i . This hierarchical structure provides v aluable prior information: features within the same group are expected to have sim- ilar embeddings, up to a small number of group-specific outliers whose representations de viate substantially from the group pattern. For example, in EHR data, Ibuprofen and Naproxen are both nonsteroidal anti-inflammatory drugs and thus play similar clinical roles, suggesting their embeddings should exhibit high similarity . By incorporat- ing this group information, we aim to impro ve embedding estimation ef ficiency , enhance cross-site alignment, and stabilize embedding recov ery in the presence of multi- block missingness. Let V = V 1 ∪ V 2 , V o = V 1 ∩ V 2 with | V o | = n o , and V no = V \ V o denote the index sets of all, over - lapping, and non-overlapping features, respecti vely . W e assume some ov erlapping features admit a shared pop- ulation embedding across the two systems. W e refer to 4 these as cr oss-site consistent features and denote the set by V o,cons . Others exhibit genuine cross-site heterogene- ity and admit distinct population embeddings across sites. W e refer to these as cr oss-site diver gent features and de- note the set by V o,div . For each group, we assume most features share a similar embedding (excluding cross-site di vergent features) and allow the existence of group out- liers whose embeddings de viate significantly from the majority in that group. W e denote by V out the set of fea- tures whose population embeddings do not conform to ei- ther cross-site or within-group homogeneity assumptions, including cross-site di ver gent features and group outliers. The remaining features form V trans = V o,cons ∪ V no,g rp , where V no,g rp contains non-ov erlapping features that are not group outliers. W e further divide V no,g rp into a set of features belonging to groups anchored by at least one cross-site consistent ov erlapping code, denoted by V g rp,anch , and a set of features in groups without an y cross-site consistent features, denoted by V g rp,sol o . These four sets, {V o,cons , V g rp,anch , V g rp,sol o , V out } , determine ho w information is shared or restricted across sites and groups. For an y feature i ∈ V \ V o,div , we introduce an indi- cator h i , where h i = 1 if the feature is not a group out- lier and h i = 0 otherwise. W e assume the existence of a population-level partition P = { P 1 , . . . , P L } of V o,cons such that features within the same block share a common population embedding. Each block corresponds to either a group shared across sites or a singleton feature whose embedding is consistent across sites. Formally , for any i, i ′ ∈ P ℓ with | P ℓ | > 1 , we ha ve g i = g i ′ , h i = h i ′ = 1 , and x 1 ,i = x 1 ,i ′ = x 2 ,i = x 2 ,i ′ . In addition, we impose population-le vel consistency for site-specific features through their group structure. For any site-specific feature l ∈ V g rp,anch ∩ V k , there exists at least one feature j ∈ V o,cons such that g l = g j and h l = h j = 1 . W e assume x k,l = x 1 ,j = x 2 ,j . Finally , for site- specific features i 1 ∈ V g rp,sol o ∩ V 1 and i 2 ∈ V g rp,sol o ∩ V 2 with g i 1 = g i 2 and h i 1 = h i 2 = 1 , belonging to the same group containing no cross-site consistent features, we as- sume x 1 ,i 1 = x 2 ,i 2 . Figure 1 illustrates the hierarchical feature structure and the corresponding population-lev el embedding relation- ships across sites. The estimation procedure described in the next section is designed to reco ver this hierarchical structure by identifying feature types and enforcing the corresponding embedding constraints. 2.3 Overview of the Estimation Method The goal of our estimation procedure is to co-train em- beddings across two sites while selectiv ely borrowing in- formation for features estimated to belong to V trans and av oiding negati ve transfer for those estimated to belong to V out . T o this end, the algorithm combines initial single- site embeddings with a hierarchical feature classification and then refines the embeddings according to the resulting feature types. As illustrated in Figure 2 , the algorithm proceeds in four main steps: 1. Obtain initial embeddings within each site via SVD of S k . 2. Identify cross-site consistent features via a weighted thresholding rule on aligned initial embeddings, thereby constructing estimates b V o,cons and the o ver- lapping portion of b V out . 3. Compute a provisional group center (excluding cross-site di ver gent features) for each group and use a second thresholding rule to detect group out- liers whose embeddings deviate significantly from this center , completing the construction of b V out , b V g rp,anch , and b V g rp,sol o . 4. Refine embeddings for features in b V o,cons , b V g rp,anch , b V g rp,sol o , and b V out in a stepwise f ashion, le veraging cross-site and within-group information where ap- propriate, and treating heterogeneous features sep- arately . F I G 1 . Illustration of the hierar chical structur e of n featur es: featur es 1 and 2 belong to V o,cons , featur es 3 and 5 belong to V out , featur e 4 belongs to V g rp,anch , and featur es n − 2 , n − 1 and n belong to V g rp,sol o . F I G 2 . Schematic of the T r ansNEST algorithm. 5 R E M A R K 1. The pr oposed fr amework is not limited to settings in which sites share identical or directly over- lapping featur e sets. More generally , it applies whenever a subset of features admits known one-to-one corr espon- dences acr oss sites, while the r emaining featur es ar e site- specific and organized by shar ed gr oup structure . F or ex- ample, in multilingual r epr esentation learning, site 1 and site 2 may corr espond to corpora in differ ent languages, such as English and Haitian, with partially overlapping vocabularies or known cr oss-lingual alignments. In this case, S k r epr esents language-specific SPPMI matrices, aligned words ar e linked thr ough bilingual dictionaries, and g i encodes shar ed linguistic cate gories such as part of speech or semantic class. T r ansNEST can then r efine Haitian wor d embeddings by transferring co-occurrence structur e learned fr om the information rich English cor- pus and the shar ed gr oup structur e. 2.4 Detailed Estimation Method Step 1: Initial Single-Site Embeddings via SVD. W e obtain initial embeddings by performing an SVD of S k and approximating it with the rank- r matrix ˆ S k = ˆ U k ˆ D k ˆ U ⊤ k , where ˆ D k ∈ R r × r contains the top r singular v alues (in descending order) and ˆ U k ∈ R n k × r contains the corresponding left singular vectors. The initial em- bedding matrix is then ˜ X k = ˆ U k ˆ D 1 / 2 k ∈ R n k × r , and we denote by ˜ x k,i the i th ro w of ˜ X k , which serves as the initial embedding for feature i at site k . R E M A R K 2 . Although we use single-site SVD embed- dings as the initial estimator in this study , other consis- tent embeddings, such as those derived fr om lar ge lan- guage models (LLMs) [e.g. 26 , 34 ], can also be inte- grated to initialize our algorithm. The err or rate of the r efined T r ansNEST estimator can be derived using our pr oof frame work, incorporating the initial estimation er- r or of the chosen method. Step 2: Embedding Alignment and Identification of Cr oss-Site Consistent F eatur es. Because embeddings are identifiable only up to rotation, we first align the initial single-site embeddings. Specifically , for an y orthogonal matrix Q ∈ O r , ˜ X k and ˜ X k Q represent the same solu- tion. T o make cross-site comparisons meaningful, we ro- tate the embeddings at site 2 to best match those at site 1. Let ˜ X k,o denote the submatrix of ˜ X k for ov erlapping features. W e compute the optimal rotation ˆ Q = W ( ˜ X 2 ,o , ˜ X 1 ,o ) = arg min Q ∈ O r ∥ ˜ X 1 ,o − ˜ X 2 ,o Q ∥ F , and define the aligned embeddings as ˜ X a 1 = ˜ X 1 , ˜ X a 2 = ˜ X 2 ˆ Q . Let ˜ x a k,i denote the i th row of ˜ X a k , k = 1 , 2 . Using the aligned embeddings, we classify each ov er- lapping feature as either cr oss-site consistent or cr oss-site diver gent . A feature is deemed cross-site consistent if its embeddings at the two sites are suf ficiently similar: b V o,cons = { i ∈ V o : ∥ ˜ x a 1 ,i − ˜ x a 2 ,i ∥ ≤ λ i } , and cross-site div ergent otherwise: b V o,div = V o \ b V o,cons , where λ i is a feature-specific threshold. R E M A R K 3 . T o simplify tuning, we incorporate site- specific feature weight w k,i , e.g., mar ginal fr equency of featur e i at site k . Define the weighted mean embedding ˜ x a i = w 1 ,i ˜ x a 1 ,i + w 2 ,i ˜ x a 2 ,i w 1 ,i + w 2 ,i , and classify i ∈ V o as cr oss-site consistent if max n w 1 ,i ( w 1 ,i + w 2 ,i ) ∥ ˜ x a 1 ,i − ˜ x a i ∥ 2 , w 2 ,i ( w 1 ,i + w 2 ,i ) ∥ ˜ x a 2 ,i − ˜ x a i ∥ 2 o ≤ λ. In this formulation, λ i r educes to a closed-form expr es- sion depending on ( w 1 ,i , w 2 ,i , λ ) , so only one hyperpa- rameter λ needs to be tuned. This design is naturally more tolerant of estimation variability for infrequent features, encoura ging knowledge transfer towar d them. Step 3: Initial Gr oup Center s Estimation and a Thr esh- olding Rule for Mer ging Blocks. W e obtain initial esti- mates for group centers and use a thresholding rule to de- cide whether each feature should be merged into its group center . W e propose to estimate each group center as the weighted mean of features in the group, excluding any cross-site di vergent features. W e use ˆ γ init g to denote the initial group center of group g ∈ [ G ] , and it is calculated as ˆ γ init g = P i P 2 k =1 w k,i ˜ x a k,i I { g i = g , i ∈ V k \ b V o,div } w g , where w g = P i P 2 k =1 w k,i I { g i = g , i ∈ V k \ b V o,div } . If the feature-site weights w k,i are unav ailable, we can set all of them to one. The thresholding rule is defined as follo ws. Feature i is merged into the center of group g i if and only if ˆ h i = 1 , where ˆ h i = I max k =1 , 2 c k,i ∥ ˜ x a k,i − ˆ γ init g i ∥ 2 I { i ∈ V k \ b V o,div } ≤ µ i . Here c k,i are pre-specified weights, µ i are tuning param- eters, and the indicator restricts the decision to observed, non-di vergent features. Thus, ˆ h i = 1 indicates that fea- ture i is treated as group-homogeneous and merged into its group center , while ˆ h i = 0 indicates a group outlier . 6 R E M A R K 4 . In practice , when feature-site weights ar e available, one possible choice of c k,i can be w k,i w g i . W e can further set µ i = µ , the only hyperparameter that needs to be tuned. W e adopt this configuration in the numerical studies. Similarly , this configuration is more tolerant of within-gr oup heter ogeneity for gr oups with smaller weights or that ar e less fr equent. Step 4: Embedding Refinement. After applying the two thresholding rules, we obtain the estimated four sets: b V o,cons , b V g rp,anch := { i ∈ V no : ˆ h i = 1 and ∃ i ′ ∈ b V o,cons such that ˆ h i ′ = 1 , g i = g i ′ } , b V g rp,sol o := { i ∈ V no : ˆ h i = 1 and ∄ i ′ ∈ b V o,cons such that ˆ h i ′ = 1 , g i = g i ′ } , and b V out := V no ∩ { i : ˆ h i = 0 } ∪ b V o,div . W e then refine the embeddings for features in each set us- ing update rules tailored to their estimated cross-site and within-group transferability . Step 4.1: Refine embeddings of features in b V o,cons . W e refine embeddings for features in b V o,cons by enforc- ing the block structure learned in steps 1-3, denoted by b P = { b P 1 , . . . , b P b L } . For any i and i ′ ∈ b P l , g i = g i ′ and ˆ h i = ˆ h i ′ = 1 . Let X k, b V o,cons = ( x k,i ) i ∈ b V o,cons ∈ R | b V o,cons |× r denote the population embedding matrix at site k , and let S k, b V o,cons be the corresponding submatrix of S k . W ithin each block b P ℓ , embeddings are constrained to be identical across sites. Hence, we define the constrained parameter space M ˆ h = { X 1 , b V o,cons , X 2 , b V o,cons : x 1 ,i = x 1 ,i ′ = x 2 ,i = x 2 ,i ′ , ∀ i, i ′ ∈ ˆ P l , l ∈ [ ˆ L ] } . The refined embeddings are obtained by solving ( ˆ X 1 , b V o,cons , ˆ X 2 , b V o,cons ) = arg min ( X 1 , b V o,cons , X 2 , b V o,cons ) ∈M ˆ h X k =1 , 2 ˜ w k S k, b V o,cons − X k, b V o,cons X ⊤ k, b V o,cons 2 F , (1) where ˜ w k is the weight of site k . For any feature i ∈ b V o,cons , we denote its refined embedding as ˆ x 1 ,i = ˆ x 2 ,i . R E M A R K 5 . The weight of site ˜ w k plays an important r ole in ( 1 ) and potentially impr oving the estimation effi- ciency . F or example, if site 1 has much less noise in the construction of S 1 (e.g ., hospital 1 has much mor e pa- tients using to construct the co-occurrence matrix of EHR featur es) compar ed with site 2 , then ˜ w 1 should be much lar ger than ˜ w 2 . In practice, these site weights can be ob- tained fr om data [ 35 ], e xternal information (e.g ., number of patients fr om each hospital), or thr ough a tuning pr o- cedur e. Unlike steps 2 and 3, wher e feature-site weights w k,i ar e used to enhance accurate featur e type detection, we employ the site weight ˜ w k her e to simplify the compu- tation. Step 4.2: Refine embeddings of features in b V g rp,anch . For i ∈ b V g rp,anch , there is a feature i ′ ∈ b V o,cons such that they belong to one group, i.e., g i ′ = g i , and both are sim- ilar to the group center , i.e., ˆ h i ′ = ˆ h i = 1 . W e directly set ˆ x k,i = ˆ x k,i ′ , where ˆ x k,i ′ is obtained in Step 4.1. Step 4.3: Refine embeddings of features in b V g rp,sol o . For non-ov erlapping features similar to the center of the group consisting of only non-ov erlapping features, we propose to refine their embeddings by conducting a weighted linear regression, treating refined ˆ X k, b V o,cons as the design matrices. For each g ∈ { g i : i ∈ b V g rp,sol o } , we estimate the shared group embedding as ˆ γ g = arg min γ X k =1 , 2 ˜ w k X i X j n ( s k,i,j − γ T ˆ x k,j ) 2 × I ( i ∈ V k ∩ b V g rp,sol o , g i = g , j ∈ b V o,cons ) o . W e apply the site weight ˜ w k again to potentially enhance estimation ef ficiency . F or an y feature i ∈ b V g rp,sol o , we de- note its refined embedding as ˆ x k,i = ˆ γ g i . Step 4.4: Refine embeddings of features in b V out . For features in b V out , where direct knowledge transfer is im- possible, we refine their embeddings by solving linear re- gressions at each site using the refined embeddings of fea- tures in b V trans = b V o,cons ∪ b V g rp,anch ∪ b V g rp,sol o , together with the initial single-site SVD embeddings of features in b V out , as the full design matrices. Before the regression, we first need to align the re- fined embeddings and the initial single-site embeddings at each site. Specifically , we estimate the orthogonal ma- trix W k,r at each site by aligning ˜ x k,i with ˆ x k,i for i ∈ b V trans , and W k,r = W { ˜ X k, b V trans , ˆ X k, b V trans } , where ˜ X k, b V trans = ( ˜ x k,i ) i ∈ b V trans and ˆ X k, b V trans = ( ˆ x k,i ) i ∈ b V trans . Let ˜ x a2 k,i = W T k,r ˜ x k,i , which is aligned with the refined embeddings at site k . W e then solve for ˆ x k,i , i ∈ b V out as follo ws ˆ x k,i = arg min x n X j ( s k,i,j − x T ˆ x k,j ) 2 I ( j ∈ V k \ b V out ) + X j ( s k,i,j − x T ˜ x a2 k,j ) 2 I ( j ∈ b V out ) o . (2) 7 R E M A R K 6 . Compar ed with the initial single-site em- bedding estimation, her e we use the r efined embeddings as part of the design matrix, which can be interpr eted as r educing measurement err or in covariates, r esulting in better empirical performance. The term P j ( s k,i,j − x T ˜ x a2 k,j ) 2 I ( j ∈ b V out ) in the loss function is used to guar- antee that our embeddings ar e no worse than the ini- tial single-site embeddings in the worst scenario wher e b V trans = ∅ . 3. THEORETICAL RESUL TS 3.1 Assumptions In this subsection, we introduce the most important As- sumption 1 on the relationship among the number of dif- ferent types of features and the order of the largest and the r -th lar gest singular v alues of the true embedding ma- trices, X k and X k, V o,cons , k = 1 , 2 . W e defer other more technical assumptions, Assumptions S1– S4, on the sub- Gaussian error E k with σ 2 k as the sub-Gaussian param- eter , the incoherence condition of U k , the lar ge enough gap between the two embeddings of any cross-site div er- gent features and between the outliers of a group and the group center for selection consistenc y , to Section A in the Supplementary Materials. T o simplify the error rates, we assume the embedding rank r is fixed. Detailed error rates as functions of r are provided in the Supplementary Ma- terials. A S S U M P T I O N 1 . W e assume the following conditions hold: (i) The number s of featur es at the two sites and in their overlap ar e of the same or der , that is, n 1 ∼ n 2 ∼ n o ∼ n . (ii) The leading and r -th singular values of the true em- bedding matrices at both sites, as well as those as- sociated with the cr oss-site consistent features, ar e of the same or der: τ 2 1 ( X 1 ) ∼ τ 2 1 ( X 2 ) ∼ τ r ( X T 2 , V o,cons X 1 , V o,cons ) ∼ τ 2 r ( X 1 , V o,cons ) ∼ τ 2 r ( X 2 , V o,cons ) . This implies that the number of population-level blocks satisfies |P | ≥ r . (iii) The signal str ength of the embeddings dominates the noise level, in the sense that τ 2 r ( X k ) ≳ n k σ k , for k = 1 , 2 . (iv) The effective complexity of the non-transferable featur e set is contr olled, with |V out | ≲ r − 1 n . R E M A R K 7 . Condition (i) of Assumption 1 r equir es a sufficient number of overlapping features to enable effec- tive information tr ansfer acr oss sites. Conditions (ii)–(iii) impose a signal strength r equir ement ensuring that the shar ed latent structur e is identifiable in the pr esence of noise. Condition (iv) contr ols the size of cr oss-site diver- gent and gr oup-outlying featur es, pr eventing them fr om overwhelming the shar ed structur e. 3.2 Selection Consistency In this subsection, we show model selection consis- tency in terms of identifying cross-site consistent features and mer ging blocks based on the prior group structures in the following two Lemmas. The detailed proofs are pro- vided in Sections E and G of the Supplement Materials. L E M M A 3.1. Under Assumptions 1 and S1 – S4, we can consistently identify cr oss-site consistent features via the pr oposed thr esholding pr ocedur e in step 2, i.e., (3) lim n →∞ P ( b V o,cons = V o,cons ) = 1 . L E M M A 3.2 . Under Assumptions 1 and S1 – S4, for all featur es e xcept cr oss-site diver gent features, we can consistently mer ge blocks via the pr oposed thr esholding pr ocedur e in step 3, i.e., (4) lim n →∞ P ˆ h i = h i , for all i ∈ V \ V o,div = 1 . 3.3 Error Bounds after Refinements W e establish error bounds for feature embedding refine- ments on the sets V o,cons , V g rp,anch , V g rp,sol o , and V out in Propositions S1 – S4, respectiv ely , in the Supplementary Materials. In the main paper, we present only the final error rate for min W X ∈O r ∥ ˆ X k − X k W X ∥ F , which is ob- tained by aggregating the results of the four propositions. T o provide a simplified and interpretable error bound, we introduce Assumption S5, under which we assume that the embedding rank r is fixed, population-lev el groups are not se verely underrepresented at any site, and site-specific features are balanced across sites. The full error bound without Assumption S5 is given in Theorem S1 in the Supplementary Materials. W e summarize the resulting simplified error rate of the T ransNEST estimator in the following theorem. T H E O R E M 3.3 . Under Assumptions 1 and S1 – S5, if ˜ w 1 / ˜ w 2 ∼ σ 2 2 /σ 2 1 , for k = 1 , 2 , we have min W X ∈O r ∥ ˆ X k − X k W X ∥ F = O p h log 1 / 2 ( n ) τ − 1 r ( X k ) × ( |P | 1 / 2 + | G solo | 1 / 2 )( σ 1 ∧ σ 2 ) + |V out | 1 / 2 σ k i , (5) wher e G solo = { g i : i ∈ V g rp,sol o } . 8 Here W X ∈ O r denotes an orthogonal matrix achiev- ing the optimal Procrustes alignment between ˆ X k and X k . The presence of group structure and cross-site kno wledge transfer reduces the effecti ve number of pa- rameters from ( n 1 + n 2 ) r to ( |P | + | G solo | + | V out | ) r . W ith appropriately chosen site weights, the effecti ve noise lev el for features in V trans is reduced from σ k to σ 1 ∧ σ 2 . Without loss of generality , we treat site 2 as the target site with σ 1 < σ 2 . Compared with a target- only SVD estimator , whose error rate is of the order of τ − 1 r ( X 2 ) n 1 / 2 2 σ 2 , the T ransNEST estimator achie ves a strictly better rate when ( |P | 1 / 2 + | G solo | 1 / 2 ) σ 1 ≪ n 1 / 2 2 σ 2 and |V out | ≪ n. 4. SIMULA TION STUDIES For data generation, we consider the following embed- ding structure: x k,i := β g i I ( | G i | > 1) + ζ i + δ k,i , where β g i is the embedding of the group effect, | G i | is the number of features in group g i , ζ i is the embedding of the feature effect shared between sites, and δ k,i is the embedding of the feature-site effect. If feature i belongs to V o,cons , δ 1 ,i = δ 2 ,i = 0 . If feature i at site k has the same embedding as its group center , then ζ i + δ k,i = 0 . W e set n 1 = n 2 = 2000 , n = 3000 , n o = 1000 , and r = 50 . Let B = ( β 1 , . . . , β G ) ⊤ ∈ R G × r . W e generate group ef fects via a multiv ariate normal distribution, i.e., B · ,j ∼ N (0 , Σ 1 ) , j ∈ [ r ] , where Σ 1 ∈ R G × G is block diag- onal. Specifically , Σ 1 consists of G/ 10 repeated non- ov erlapping blocks, denoted by D ∈ R 10 × 10 . D is also block diagonal, with two sub-blocks. The first sub- block is a 3 × 3 autoregressi ve cov ariance matrix of or- der 1 (AR1) with correlation parameter ρ β = 0 . 4 , i.e., D ij = 0 . 4 | i − j | for i, j ∈ [3] . This implies that the cor- responding groups are related. The second sub-block is a 7 × 7 identity matrix. Further , to mimic the hierar- chical ontology structure of EHR features in Section 5, 500 features from each site do not belong to any groups. Let Z = ( ζ 1 , . . . , ζ n ) ⊤ ∈ R n × r . W e first generate fea- ture ef fects via a multi variate normal distribution, i.e., Z · ,j ∼ N (0 , Σ 2 ) , j ∈ [ r ] , where Σ 2 ∈ R n × n is block diagonal, and each block is AR1 with size 6 and cor- relation parameter ρ ζ = 0 . 4 . W e then randomly select n c ζ ov erlapping features, and set their ζ i to zero. For these features, the group effect dominates. Regarding feature-site ef fects, we randomly select n δ ov erlapping features from each site, denoting the set of index es as { d k, 1 , . . . , d k,n δ } , k = 1 , 2 , and generate feature-site ef- fects for them. While for the remaining features, δ k,i = 0 . Let ∆ k = ( δ k,d k, 1 , . . . , δ k,d k,n δ ) ⊤ ∈ R n δ × r . W e gener- ate feature-site effects via a multi variate normal distri- bution, i.e., ∆ k, · ,j ∼ N (0 , Σ 3 ) , j ∈ [ r ] , k = 1 , 2 , where Σ 3 ∈ R n δ × n δ is block diagonal with block size 50 and each block is AR1 with correlation parameter ρ δ = 0 . 95 . W e then generate the data matrices S k via s k,i,j = x ⊤ k,i x k,i + ϵ k,i,j , ϵ ij ∼ N (0 , σ k,i σ k,j ) . W e treat site 1 as the source site with σ 1 ,i = 5 for any i ∈ [ n 1 ] . For the tar- get site 2, we further separate features into frequent fea- tures and rare features to mimic the real EHR features. Specifically , there are n f = 1300 frequent features with σ 2 ,f = 20 , and n r = 700 rare features with σ 2 ,r = 80 . W e set w k,i = 1 /σ k,i in the simulation, and similar to [ 36 ], we calculate the site weights as (6) e w k = n − 1 3 − k ∥ S 3 − k − ˜ X 3 − k ( ˜ X 3 − k ) T ∥ 2 2 n − 1 1 ∥ S 1 − ˜ X 1 ( ˜ X 1 ) T ∥ 2 2 + n − 1 2 ∥ S 2 − ˜ X 2 ( ˜ X 2 ) T ∥ 2 2 . W e compare our method with four benchmarks: (1) single-site SVD (SSVD), i.e., ˜ X 2 , (2) single-site with group structures (SSG) method, (3) plain data pooling (DP) method, and (4) BONMI method [ 36 ]. The SSVD method fails to le verage the group structures and the information-rich source population. F or the SSG method, we utilize the prior group structures and replace the fea- ture embedding with the mean of the SSVD embed- dings of all features belonging to the group. This single- site method ignores the heterogeneity within groups. In the DP method, we first calculate the weighted sum of the observed data matrices for the overlapping features, i.e., P k =1 , 2 e w k { S k,i,j } i,j ∈ V o , and obtain the embedding of overlapping features via SVD; we then solve an or- thogonal procrustes problem using the co-trained embed- dings and the SSVD embeddings of overlapping features at the target site; we finally rotate the SSVD embed- ding of tar get-specific features using the estimated or - thogonal matrix to get aligned DP embeddings. BONMI method completes the full matrix S full ∈ R n × n by recov- ering the missing block S miss ∈ R ( n 1 − n o ) × ( n 2 − n o ) , i.e., the relationship between site-1-specific features and site- 2-specific features, under the lo w-rank assumption, and then applies SVD to the completed b S full to obtain the em- beddings for all n features. Note that DP and BONMI both ignore the heterogeneity between sites and do not le verage the prior group structures. W e consider configurations where G = 250 , 400 , n c ζ = 0 , 1000 , and n δ = 100 , 200 to study the influence of group size, the le vel of group-heterogeneity , and the le vel of site-heterogeneity on the performance of different meth- ods. When G = 250 , the uniform group size is 8; when G = 400 , the uniform group size is 5. When n c ζ = 0 , then Assumption S4 on the exact group selection consistency does not hold. W e e xplore the performance of T ransNEST under this model misspecification. For ev aluation, we not only report the estimation ac- curacy in terms of estimating the true embedding X 2 , 9 but also the performance of identifying positiv e feature- feature pairs against random pairs, moti vated by EHR ap- plications. In practice, we hav e kno wledge of which two EHR features are similar and which two are related. For example, type 1 diabetes is similar to type 2 diabetes, and they belong to the same group, diabetes mellitus. T ype 2 diabetes is related to insulin, although they do not belong to the same ontology group. Further , the sparsity of the network allo ws us to assume that each random pair repre- sents a weak or no relationship. W e curate related feature- feature pairs based on the structure of Σ l , l = 1 , 2 , 3 , for e valuation. If two features belong to the same group, the y are similar and related. If Σ 1 ,i,j > 0 , then features in the i th group are related to the features in the j th group. If Σ 2 ,i,j > 0 , ζ i = 0 and ζ j = 0 , then feature i and feature j are related. W e can further expand the set of related pairs specific for the target site using the non-zero elements in Σ 3 paired with the set { d 2 , 1 , . . . , d 2 ,n δ } . These target- specific pairs mimic the EHR feature pairs that hold only for a specific patient sub-population. A random subset of the positi ve pairs was used for tuning, and the remaining set was used for e valuation. The results of the four configurations with G = 400 are summarized in T ables 1 and 2 . In T able 1 , we report area under the curve (A UC) for all a vailable feature pairs, fre- quent pairs, rare pairs, frequent and transfer-eligible pairs, frequent and transfer-ineligible pairs, rare and transfer- eligible pairs, and rare and transfer-ineligible pairs. If one of the features in a pair is frequent and allows kno wledge transfer , i.e., belonging to V trans , we classify the pair as frequent and transfer eligible. The same classification logic is applied to the other types of feature pairs. F or each positi ve or random feature pair in the ev aluation set, we compute the cosine similarity between the embeddings of the two features. A UCs are then calculated by com- paring the vector of cosine similarities to a binary v ec- tor of the same length, where 1 indicates a positi ve pair , and 0 indicates a random (neg ativ e) pair . Our method, T ransNEST , achiev es the highest overall A UC for distin- guishing positi ve pairs from random pairs across all four configurations. Relati ve to ( C 1) , configuration ( C 3) ex- hibits greater heterogeneity within groups, which impairs the performance of SSG and reduces the ef fectiv e sam- ple size av ailable to T ransNEST . Despite this, T ransNEST generally outperforms the other methods by adaptively capturing the degree of heterogeneity and borro wing use- ful information from the source. Compared with ( C 1) , ( C 2) has a lar ger n δ , indicating a higher lev el of het- erogeneity both between sites and within groups. This in- creased heterogeneity degrades the performance of SSG, DP , and BONMI; howe ver , SSVD and T ransNEST remain relati vely robust. For TransNEST , the observed decrease in A UCs for rare and transfer-ineligible features is primar - ily due to the smaller number of cross-site consistent fea- tures and the smaller effecti ve sample sizes when updat- ing their embeddings. As a result, the data matrix used for updating the embeddings of rare and transfer-ineligible features, via the regression framework, is of lower qual- ity , ne gativ ely af fecting performance. Configuration ( C 4) presents the most challenging scenario for knowledge transfer due to its highest lev el of heterogeneity . Even so, TransNEST slightly outperforms SSVD overall and sho ws a clear adv antage ov er SSVD for rare and transfer - eligible features, although such features are relati vely fe w in this configuration. In T able 2 , we not only focus on the angles be- tween two embeddings b ut also on the scales. W e report F .err , defined as n − 1 2 ∥ M 2 − b S 2 ∥ F , where b S 2 = c X 2 c X ⊤ 2 ; F .rare.err , defined as n − 1 r ∥ M 2 , rare , rare − b S 2 , rare , rare ∥ F , where M 2 , rare , rare and b S 2 , rare , rare are sub-matrices of M 2 and b S 2 with only the rows and columns corre- sponding to the rare features selected, respecti vely; and F .freq.err , defined as n − 1 f ∥ M 2 , freq , freq − b S 2 , freq , freq ∥ F , where M 2 , freq , freq and b S 2 , freq , freq are sub-matrices of M 2 and b S 2 with only the rows and columns corresponding to the frequent features selected, respectively . Generally speaking, T ransNEST has the best performance in ( C 1) and ( C 2) , and comparable performance with DP in ( C 3) and ( C 4) , outperforming the rest methods. TransNEST significantly outperforms SSVD in the four configura- tions in terms of F .err and F .rare.err . It also has a smaller F .freq.err in the more homogeneous settings ( C 1) and ( C 2) . TransNEST outperforms SSG in terms of F .err and F .freq.err , especially in ( C 3) and ( C 4) , where the within- group heterogeneity is larger . BONMI has the smallest F .freq.err but the second largest F .rare.err in the four con- figurations, potentially caused by the wrong assumption that all ov erlapping features are site-homogeneous, and the lack of utilization of prior feature-site weights. DP has a comparable performance to T ransNEST in ( C 3) and ( C 4) , potentially due to TransNEST’ s failure to ef- fecti vely utilize group structure to boost ef ficiency when the group heterogeneity is large. The results of the remaining four configurations with G = 250 can be found in Section J of the Supplementary Materials, which sho w similar patterns. 5. REAL D A T A ANAL YSIS The analysis included data from a total of 0.25 million patients at Boston Children’ s Hospital (BCH), a quater- nary referral center for pediatric care, and 2.5 million pa- tients at Mass General Brigham (MGB), a Boston-based non-profit hospital system serving primarily an adult pop- ulation, each of whom had at least one visit with codified medical records. W e gathered four domains of codified features from BCH and MGB, including PheCodes for diagnosis, RxNorm codes for medication, LOINC codes for lab measurements, and CCS codes for procedures. After frequency control, 3055 codes are shared between 10 T A B L E 1 The performances of the embeddings from five methods on identifying the positive featur e pairs a gainst random/ne gative pairs. Reported A UCs are computed using the same evaluation pr ocedur e, with the set of positive pairs r estricted to all labeled available pairs (A UC), fr equent labeled pairs (A UC.fr eq), rar e labeled pairs (A UC.rar e), fr equent transfer-eligible labeled pairs (A UC.fr eq.T r), fr equent transfer -ineligible labeled pairs (A UC.fr eq.NT r), rar e transfer-eligible labeled pair s (A UC.rar e.T r), and rar e transfer -ineligible labeled pairs (A UC.rar e.NT r). A UC A UC.freq A UC.rare A UC.freq.Tr A UC.freq.NT r A UC.rare.T r A UC.rare.NTr ( C 1) SSVD 0.72 0.78 0.66 0.83 0.82 0.66 0.66 G = 400 SSG 0.73 0.69 0.78 0.87 0.63 0.81 0.80 n c ζ = 1000 DP 0.69 0.66 0.77 0.88 0.58 0.82 0.79 n δ = 100 BONMI 0.70 0.69 0.74 0.87 0.62 0.76 0.75 T ransNEST 0.77 0.76 0.79 0.87 0.76 0.80 0.77 ( C 2) SSVD 0.72 0.75 0.69 0.83 0.77 0.66 0.71 G = 400 SSG 0.69 0.67 0.68 0.87 0.61 0.80 0.62 n c ζ = 1000 DP 0.65 0.64 0.64 0.88 0.57 0.82 0.57 n δ = 200 BONMI 0.67 0.67 0.66 0.88 0.61 0.77 0.60 T ransNEST 0.76 0.75 0.75 0.87 0.74 0.79 0.72 ( C 3) SSVD 0.72 0.73 0.67 0.73 0.77 0.68 0.68 G = 400 SSG 0.70 0.66 0.74 0.72 0.62 0.76 0.76 n c ζ = 0 DP 0.70 0.67 0.74 0.78 0.59 0.77 0.75 n δ = 100 BONMI 0.70 0.68 0.73 0.77 0.62 0.75 0.74 T ransNEST 0.75 0.73 0.75 0.74 0.74 0.77 0.75 ( C 4) SSVD 0.72 0.73 0.69 0.73 0.74 0.67 0.70 G = 400 SSG 0.67 0.65 0.67 0.71 0.61 0.76 0.61 n c ζ = 0 DP 0.66 0.65 0.64 0.78 0.58 0.76 0.58 n δ = 200 BONMI 0.68 0.67 0.66 0.77 0.61 0.75 0.61 T ransNEST 0.73 0.71 0.71 0.74 0.71 0.76 0.69 T A B L E 2 The performances of five methods on estimating the tar get err or-fr ee matrix M 2 , M 2 , rare , rare , and M 2 , freq , freq in terms of F r obenius err ors. F .err F .rare.err F .freq.err F .err F .rare.err F .freq.err ( C 1) ( C 2) SSVD 16.11 34.24 8.60 16.00 33.86 8.31 SSG 10.23 13.80 9.12 10.24 13.13 9.04 DP 11.73 19.61 8.28 11.64 18.46 8.54 BONMI 13.86 28.37 7.91 13.53 26.86 7.82 T ransNEST 9.65 13.06 8.23 9.84 13.55 8.23 ( C 3) ( C 4) SSVD 15.53 31.64 8.65 15.48 31.37 8.63 SSG 11.72 14.17 10.69 12.07 14.89 10.75 DP 11.13 17.29 8.42 11.58 17.57 8.75 BONMI 12.26 23.14 7.68 12.18 22.30 7.68 T ransNEST 11.35 14.29 9.91 11.81 15.26 10.14 11 the two hospital systems, while 1221 codes are unique to BCH, and 2350 codes are unique to MGB. W e lev er- age the top-do wn hierarchical structures of PheCodes, RxNorm codes, and LOINC codes as prior group knowl- edge. More details can be found in [ 21 ]. W e then gen- erated a summary-level co-occurrence matrix and corre- sponding SPPMI matrix of EHR codes at each site as de- scribed in [ 2 ]. Similar to the simulation studies, we compare our T ransNEST method with four benchmarks: SSVD, SSG, DP , and BONMI. For ev aluation, we curated labels from pediatric articles on reputable sources, cov ering both Phe- Code to PheCode and PheCode to RxNorm code pairs. W e randomly selected a subset of the labels for tuning and the remaining for ev aluation. Similar to the simula- tion studies, we also randomly selected the same number of code pairs and treated them as negati ve pairs, i.e., the two codes are unrelated, based on the f act that the network is v ery sparse. W e assessed the quality of the embeddings by measuring the accuracy of the cosine similarity be- tween two embeddings of each code pair, and reported A UCs for dif ferent types of codes. T o further delineate the adv antages of T ransNEST , we categorized BCH EHR codes into two groups based on their occurrence frequen- cies within patient records: rare codes, seen in less than 0.1% of patients, and frequent codes, observed in 0.1% or more of patients. The results are summarized in T able 3 . Among the two single-site methods, SSG outperforms SSVD, high- lighting the utility of incorporating group structures. DP , BONMI, and T ransNEST all outperform SSVD, suggest- ing a de gree of similarity between MGB and BCH that al- lo ws effecti ve knowledge transfer from MGB to enhance pediatric embeddings. Furthermore, T ransNEST outper - forms BONMI by effecti vely modeling site heterogene- ity in overlapping codes and le veraging group structures. It achie ves performance comparable to DP in identify- ing frequent PheCode–PheCode pairs b ut substantially higher A UCs in identifying PheCode–RxNorm pairs and rare PheCode–PheCode pairs. This suggests that frequent PheCodes are more similar between MGB and BCH than RxNorm codes. For rare BCH codes, the combined bene- fits of group structure, transferred kno wledge from MGB, and adaptive modeling of heterogeneity are critical, en- abling T ransNEST to excel. 6. DISCUSSION W e propose a no vel transfer learning under a structured missingness framework, T ransNEST , based on network embeddings. It le verages prior hierarchical structures to enhance kno wledge transferability across sites and adap- ti vely captures complex heterogeneity both within groups and between sites in a robust manner . TransNEST is also intrinsically federated, preserving data pri vac y by com- municating only summary-le vel information across sites, e.g., code co-occurrence matrices of patient cohorts, with- out sharing patient-le vel data. W ith the advancement of LLMs, high-quality source or general-purpose embed- dings hav e become widely accessible. Howe ver , these embeddings may be too generic for specific downstream tasks. T ransNEST of fers a principled way to fine-tune such embeddings, enabling them to better capture target- specific characteristics and statistical patterns. More im- portantly , we rigorously establish theoretical guarantees for the T ransNEST method, which also sheds light on the broader theoretical analysis of low-rank models, where obtaining a ∥ · ∥ 2 →∞ error bound remains a significant challenge. Although we focus on two sites in this paper, T ransNEST can be naturally extended to a multi-source setting. T o facilitate the theoretical deri vations, we adopt a hard- thresholding strategy for classifying code types. An in- teresting direction for future work is to explore a soft- thresholding approach, which may better capture the con- tinuously v arying levels of heterogeneity within groups and between sites. Another promising direction is to more ef fectively le verage prior multi-layer hierarchical struc- tures, potentially through the use of Graph Neural Net- works. SUPPLEMENT AR Y MA TERIAL Supplementary Materials to ‘T ransfer Lear ning with Network Embeddings under Structured Missing- ness’ T echnical assumptions, intermediate theoretical results, proofs, and additional simulation results are provided in the Supplementary Materials. REFERENCES [1] A B B E , E . , F A N , J . , W A N G , K . and Z H O N G , Y . (2020). Entry- wise eigen vector analysis of random matrices with lo w expected rank. Annals of statistics 48 1452. [2] B E A M , A . L . , K O M P A , B ., S CH M A L T Z , A ., F R I E D , I ., W E - B E R , G . , P A L ME R , N ., S HI , X ., C A I , T. and K O H A N E , I . S . (2020). Clinical concept embeddings learned from massive sources of multimodal medical data. In P acific Symposium on Biocomputing. P acific Symposium on Biocomputing 25 295. [3] B E N G IO , Y . (2012). Deep learning of representations for unsu- pervised and transfer learning. In Proceedings of ICML work- shop on unsupervised and tr ansfer learning 17–36. JMLR W ork- shop and Conference Proceedings. [4] B I , X . , Q U , A . , W A N G , J . and S H E N , X . (2017). A group- specific recommender system. J ournal of the American Statis- tical Association 112 1344–1353. [5] B I N K IE W I C Z , N ., V O GE L S TE I N , J . T. and R O HE , K . (2017). Cov ariate-assisted spectral clustering. Biometrika 104 361–377. [6] C A I , T . , C A I , T. T. and Z H A N G , A . (2016). Structured matrix completion with applications to genomic data integration. Jour - nal of the American Statistical Association 111 621–633. 12 T A B L E 3 A UCs in identifying differ ent types of positive pairs against r andom/negative pair s. P-P repr esents PheCode-PheCode pairs, P-R r epr esents PheCode-RxNorm pairs, P-P fr eq r epresents PheCode-PheCode pairs with at least one code being fr equent, P-R fr eq r epresents PheCode-RxNorm pairs with at least one code being fr equent, P-P rar e r epresents PheCode-PheCode pairs with at least one code being rare , and P-R rar e r epresents PheCode-PheCode pair s with at least one code being rar e. P-P P-R P-P freq P-R freq P-P rare P-R rare Number of pairs 3715 339 3649 319 797 138 SSVD 0.75 0.70 0.74 0.71 0.60 0.63 SSG 0.77 0.75 0.76 0.75 0.71 0.71 DP 0.81 0.77 0.79 0.77 0.76 0.74 BONMI 0.79 0.72 0.77 0.74 0.74 0.68 T ransNEST 0.80 0.81 0.78 0.80 0.80 0.80 [7] C A I , T. T. and Z H O U , W.- X . (2016). Matrix completion via max-norm constrained optimization. [8] C A N D ÈS , E . J . and T AO , T. (2010). The power of con vex re- laxation: Near-optimal matrix completion. IEEE transactions on information theory 56 2053–2080. [9] C A P E , J ., T A N G , M . and P R I E B E , C . E . (2019). The two-to- infinity norm and singular subspace geometry with applications to high-dimensional statistics. The Annals of Statistics 47 2405– 2439. [10] C H E N , I . Y ., A G R A W A L , M . , H O R NG , S . and S O N T AG , D . (2019). Robustly extracting medical knowledge from EHRs: a case study of learning a health knowledge graph. In P acific Sym- posium on Biocomputing 2020 19–30. W orld Scientific. [11] C H E N , Y . and L I , X . (2024). A note on entrywise consistency for mixed-data matrix completion. Journal of Machine Learning Resear ch 25 1–66. [12] C H E R NO Z H UK O V , V . , H A N S E N , C ., L I AO , Y . and Z H U , Y . (2023). Inference for lo w-rank models. The Annals of statistics 51 1309–1330. [13] F A N , J ., W A N G , D . , W A N G , K . and Z H U , Z . (2019). Distributed estimation of principal ei genspaces. Annals of statistics 47 3009. [14] F E R N A N D E Z , E ., P E R E Z , R ., H E R NA N D E Z , A ., T E JA D A , P ., A RT E TA , M . and R A M O S , J . T . (2011). Factors and mecha- nisms for pharmacokinetic differences between pediatric popu- lation and adults. Pharmaceutics 3 53–72. [15] G O L D ST E I N , B . A ., N A V A R , A . M . , P E N C I N A , M . J . and I OA N - N I D I S , J . P . (2016). Opportunities and challenges in developing risk prediction models with electronic health records data: a sys- tematic revie w . J ournal of the American Medical Informatics As- sociation: J AMIA 24 198. [16] G R AC Y , D ., W E I SM A N , J ., G R A N T , R ., P RU I T T , J . and B R I T O , A . (2012). Content barriers to pediatric uptake of elec- tronic health records. Advances in P ediatrics 59 159–181. [17] H I N T O N , G . E . , O S I N D E RO , S . and T E H , Y . - W . (2006). A fast learning algorithm for deep belief nets. Neural computation 18 1527–1554. [18] H I N T O N , G . E . and S A L A K H U TD I N OV , R . R . (2006). Reduc- ing the dimensionality of data with neural netw orks. science 313 504–507. [19] J O S E PH , P . D ., C R A I G , J . C . and C AL DW E L L , P . H . (2015). Clinical trials in children. British journal of clinical pharmacol- ogy 79 357–369. [20] L E V Y , O . and G O L DB E R G , Y . (2014). Neural word embedding as implicit matrix factorization. Advances in neural information pr ocessing systems 27 . [21] L I , M ., L I , X . , P AN , K ., G E V A , A . , Y A N G , D . , S W E ET , S . M ., B O N Z E L , C . - L ., A Y A K U L A N G A R A P A N I C K A N , V . , X I O N G , X . , M A N D L , K . et al. (2024). Multisource representation learn- ing for pediatric knowledge extraction from electronic health records. NPJ Digital Medicine 7 319. [22] L I , Z . , Q I N , K ., H E , Y ., Z H O U , W . and Z HA N G , X . (2024). Knowledge transfer across multiple principal component analy- sis studies. arXiv pr eprint arXiv:2403.07431 . [23] L I U , H . , B I E L I N S KI , S . J . , S O H N , S . , M U R P HY , S . , W AG H O - L I K A R , K . B ., J O N NA L AG A D DA , S . R ., R A V I K U M AR , K ., W U , S . T., K U L L O , I . J . and C H U T E , C . G . (2013). An in- formation extraction framew ork for cohort identification using electronic health records. AMIA Summits on T ranslational Sci- ence Pr oceedings 2013 149. [24] M A Z U MD E R , R . , H A S T I E , T. and T I B S H I R A NI , R . (2010). Spectral regularization algorithms for learning large incomplete matrices. The Journal of Machine Learning Research 11 2287– 2322. [25] P AS S A L I S , N . and T E FA S , A . (2018). Unsupervised knowledge transfer using similarity embeddings. IEEE tr ansactions on neu- ral networks and learning systems 30 946–950. [26] R A S M Y , L . , X I A N G , Y . , X IE , Z . , T AO , C . and Z H I , D . (2021). Med-BER T : pretrained contextualized embeddings on large- scale structured electronic health records for disease prediction. NPJ digital medicine 4 86. [27] R O H D E , A . and T S Y B A K OV , A . B . (2011). Estimation of high- dimensional low-rank matrices. [28] S C H Ö NE M A NN , P . H . (1966). A generalized solution of the or- thogonal procrustes problem. Psychometrika 31 1–10. [29] S H E U , Y . - H . , M A G D A MO , C . , M I L L E R , M ., D A S , S . , B L AC K E R , D . and S M O L L E R , J . W . (2023). AI-assisted pre- diction of differential response to antidepressant classes using electronic health records. NPJ Digital Medicine 6 73. [30] S H I , X . , L I , X . and C A I , T. (2021). Spherical regression under mismatch corruption with application to automated knowledge translation. Journal of the American Statistical Association 116 1953–1964. [31] S I , Y ., D U , J ., L I , Z . , J I A N G , X . , M I L L E R , T . , W A N G , F . , Z H E N G , W . J . and R O B E RT S , K . (2021). Deep representation learning of patient data from Electronic Health Records (EHR): A systematic revie w . J ournal of biomedical informatics 115 103671. [32] S U I , Y ., X U , Q . , B A I , Y . and Q U , A . (2025). Multi-task Learning for Heterogeneous Multi-source Block-Wise Missing Data. Journal of Computational and Graphical Statistics just- accepted 1–15. [33] Y A M A N E , I . , Y G E R , F., B E R A R , M . and S U G I Y A M A , M . (2016). Multitask principal component analysis. In Asian Con- fer ence on Machine Learning 302–317. PMLR. 13 [34] Y U A N , Z ., Z HA O , Z . , S U N , H ., L I , J ., W A N G , F. and Y U , S . (2022). CODER: knowledge-infused cross-lingual medical term embedding for term normalization. Journal of biomedical infor- matics 126 103983. [35] Z H O U , D ., C A I , T. and L U , J . (2021). Multi-source Learning via Completion of Block-wise Overlapping Noisy Matrices. arXiv pr eprint arXiv:2105.10360 . [36] Z H O U , D ., C A I , T . and L U , J . (2023). Multi-source learning via completion of block-wise o verlapping noisy matrices. J ournal of Machine Learning Resear ch 24 1–43. [37] Z H UA N G , F., C H E N G , X . , L U O , P . , P A N , S . J . and H E , Q . (2015). Supervised representation learning: Transfer learning with deep autoencoders. In IJCAI 15 4119–4125. Buenos Aires.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment