El Agente Gráfico: Structured Execution Graphs for Scientific Agents
Large language models (LLMs) are increasingly used to automate scientific workflows, yet their integration with heterogeneous computational tools remains ad hoc and fragile. Current agentic approaches often rely on unstructured text to manage context…
Authors: Jiaru Bai, Abdulrahman Aldossary, Thomas Swanick
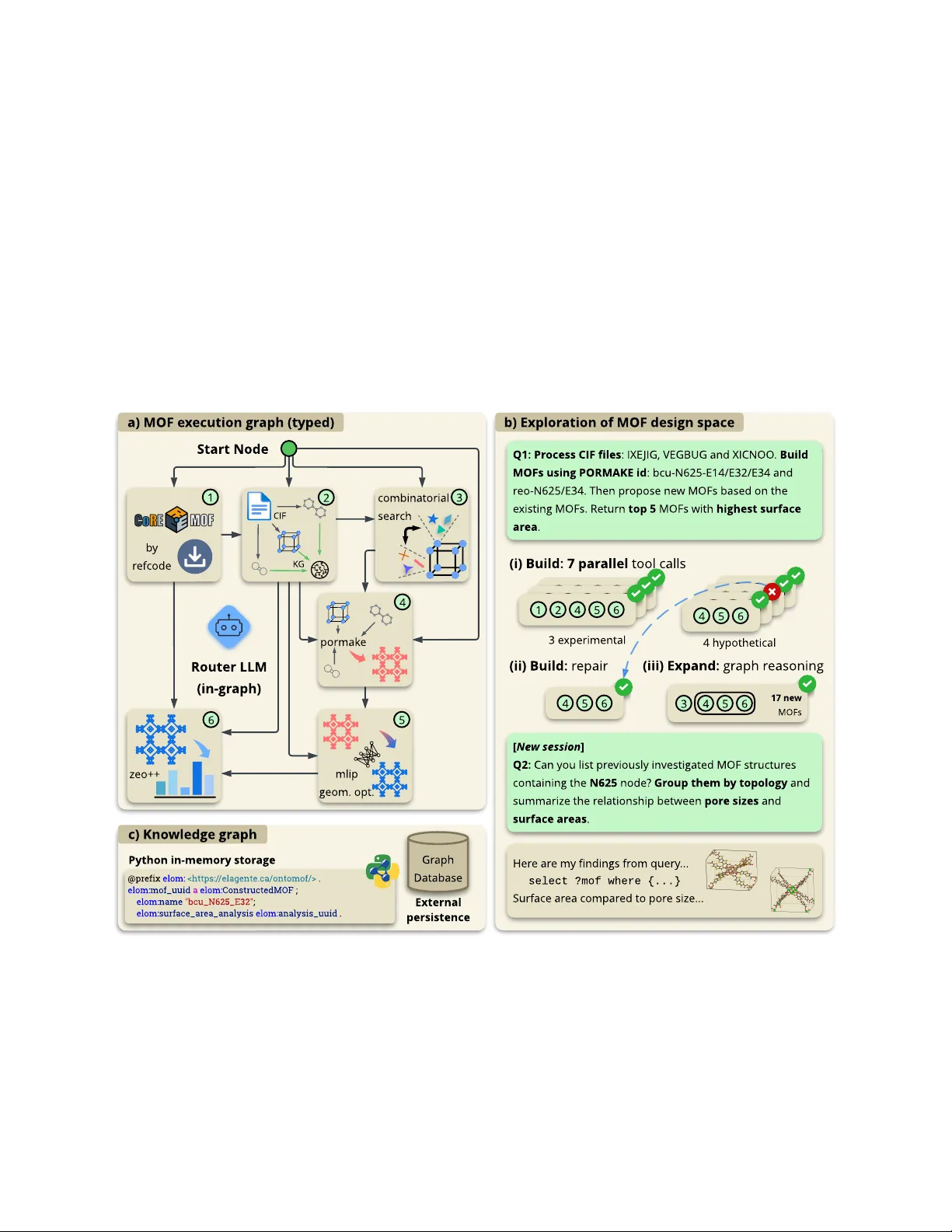
El Ag ente Gráfico: S tructured Ex ecution Graphs for Scientific Ag ents Jiaru Bai 1 , 2 , † , , Abdulrahman Aldossary 1 , 2 , † , , Thomas Sw anick 3 , 4 , , Marcel Müller 1 , 2 , , Y eonghun Kang 1 , 2 , , Zijian Zhang 2 , 3 , , Jin W on L ee 5 , , T sz W ai K o 1 , 2 , , Mohammad Ghazi V akili 1 , 2 , 3 , , V arinia Bernales 1 , 3 , 9 , ∗ , , Alán Aspuru-Guzik 1 , 2 , 3 , 6 , 7 , 8 , 9 , 10 , 11 , ∗ , 1 Departmen t of Chemistry , Universit y of T oron to, 80 St. George St., T oron to, ON M5S 3H6, Canada 2 V ector Institute for Artificial In telligence, W1140-108 College St., Sc hw artz Reisman Inno v ation Campus, T oron to, ON M5G 0C6, Canada 3 Departmen t of Computer Science, Universit y of T oron to, 40 St George St., T oron to, ON M5S 2E4, Canada 4 Departmen t of Mathematics, Univ ersity of T oron to, 40 St George St., T oronto, ON M5S 2E4, Canada 5 Sc ho ol of Computer Science, McGill Universit y , 3480 Univ ersit y St., Montréal, QC H3A 0E9, Canada 6 Departmen t of Materials Science & Engineering, Univ ersit y of T oronto, 184 College St., T oronto, ON M5S 3E4, Canada 7 Departmen t of Chemical Engineering & Applied Chemistry , Univ ersity of T oronto, 200 College St., T oronto, ON M5S 3E5, Canada 8 Institute of Medical Science, 1 King’s College Circle, Medical Sciences Building, Ro om 2374, T oronto, ON M5S 1A8, Canada 9 A cceleration Consortium, 700 Universit y A ve., T oronto, ON M7A 2S4, Canada 10 Canadian Institute for A dv anced Research (CIF AR), 661 Universit y A ve., T oron to, ON M5G 1M1, Canada 11 NVIDIA, 431 King St W #6th, T oronto, ON M5V 1K4, Canada † Con tributed equally to this work. Large language mo dels (LLMs) are increasingly used to automate scien tific workflo ws, y et their in tegration with heterogeneous computational to ols remains ad ho c and fragile. Curren t agentic approac hes often rely on unstructured text to manage con text and co ordinate execution, generating often ov erwhelming volumes of information that may obscure decision pro venance and hinder au- ditabilit y . In this work, we present El Ag ente Gráfico , a single-agent framework that embeds LLM-driv en decision-making within a type-safe execution environmen t and dynamic knowledge graphs for external p ersistence. Cen tral to our approach is a structured abstraction of scientific concepts and an ob ject-graph mapp er that represents computational state as typed Python ob jec ts, stored either in memory or p ersisted in an external knowledge graph. This design enables con text management through typed symbolic identifiers rather than raw text, thereby ensuring consistency , supp orting pro venance tracking, and enabling efficient to ol orchestration. W e ev aluate the system b y developing an automated b enchmarking framework across a suite of universit y-level quan tum chemistry tasks previously ev aluated on a multi-agen t system, demonstrating that a single agent, when coupled to a reliable execution engine, can robustly p erform complex, multi-step, and parallel computations. W e further extend this paradigm to tw o other large classes of applications: conformer ensemble generation and metal-organic framework design, where knowledge graphs serve as b oth memory and reasoning substrates. T ogether, these results illustrate how abstraction and type safety can provide a scalable foundation for agentic scientific automation b eyond prompt-centric designs. Date: F ebruary 23, 2026 Correspondence: v arinia@b ernales.org and alan@aspuru.com 1 1 Introduction The integration of large language mo dels ( LLM s) into scientific workflo ws enables agen ts to inv ok e external to ols and orchestrate m ulti-step pro cedures ( 1 ; 2 ; 3 ; 4 ; 5 ; 6 ; 7 ; 8 ; 9 ; 10 ; 11 ; 12 ; 13 ). These agents can parameterize function calls, reason ov er quantitativ e outputs, and synthesize results conv ersationally . In our previous work, El A gente Q ( 5 ) introduced a computational chemistry agent, rep orting > 87 % accuracy using a multi-agen t architecture. A new family of agentic frameworks based on this arc hitecture is expanding in to a wide v ariet y of applications ( 14 ; 15 ; 16 ; 17 ). T o mitigate LLM con text limits, a common approach is multi-agen t decomp osition to reduce p er-agen t context load ( 18 ), but this introduces co ordination and v erification failures typical of multi-agen t systems ( 19 ; 20 ). Consisten t with this view, recen t work ( 21 ) rep orts diminishing or negativ e returns from co ordination once a single agent exceeds a mo derate capability threshold. These limitations are acute in scien tific computing, where numerical correctness and state fidelity dominate o ver conv ersational coherence. Numerical sim ulation pipeline s generate large volumes of structured data and binary artifacts ( 22 ; 23 ) that are impractical to transmit through LLM con text windo ws. Sev eral systems, therefore, externalize execution metadata; for example, DREAMS ( 10 ) uses a centralized “can v as” to track calculation status and file paths. How ev er, rep eated disk-based (de-)serialization of large ob jects can still imp ose substantial ov erhead, esp ecially in GPU-accelerated settings ( 24 ; 25 ). A further barrier is soft ware heterogeneity: w orkflows span diverse molecular formats [ e.g. , xyz , SELFIES ( 26 ), InChI ( 27 )] and large configuration spaces, making misconfiguration b oth common and costly . Hard-coded conv erters and probabilistic LLM -based glue remain brittle at scale and under comp osition. W e argue that these issues share a ro ot cause: the execution con text is treated as unstructured and ephemeral, rather than a first-class represen tation of the scientific state. Scalable agen ts require execution state to b e explicitly t yp ed, v alidated, and decoupled from transient textual con text ( 28 ). A t yp ed abstraction la yer enables safe state transmission across heterogeneous to ols, supp orts automated trace v alidation and b enc hmarking, and preserv es prov enance of intermediate results. In this w ork, we present El Agente Gráfico , a single-agen t framework with a type-safe execution en vironment for scientific w orkflows. Scien tific state is represen ted as structured Python ob ject graphs through a typed abstraction lay er and is p ersisted in an external knowledge graph ( K G ) via an ob ject graph mapp er ( OGM ). W e ev aluate Gráfico on GPU-accelerated quantum c hemistry tasks using GPU4PySCF ( 24 ) and demonstrate its extensibility through case studies in conformer search for Boltzmann-weigh ted sp ectroscop y in solution and metal-organic framework ( MOF ) design. An ov erview of the main comp onen ts of Gráfico can b e found in Figure 1 . 2 R esults W e first discuss the arc hitecture of Gráfico , the underlying technology , and the to ols used. W e then discuss the results of the quantum chemistry exercises, comparing different LLM s with resp ect to p erformance and cost. Finally , we presen t tw o case studies that demonstrate applicability to other to ols, namely conformer searc h and MOF design. 2.1 S ystem design Figure 2 presen ts an o verview of the Gráfico arc hitecture, highlighting its core design principles and mo dular comp onen ts. W e describ e the key elements b elo w to provide context for how the s ystem enables structured, extensib le scientific workflo ws. Execution Graph A t its core, Gráfico exp oses typed execution graphs as to ols, reducing context pressure b y treating workflo ws as v alidated state transformations rather than free-form text. In the GPU4PySCF w orkflow, for example, the key challenge is managing complex intermediate ob jects ( e.g. , mean-field state) that are exp ensiv e to recompute and impractical to serialize through LLM con text windo ws. W e enco de single-p oin t, geometry optimization, frequency , and time-dep endent density functional theory ( TDDFT ) 2 Her e ar e 20 c o n f o r m er s o f t h e syst em ! select ed : O1 L i ve S o f t w a r e & T o o l s A sk o r ch at w it h E l A g ent e G r áf i c o ! Wh at is t h e DM S O sol vat i o n ef f ect ? M C P c o d e exec u t i o n W eb sear c h K n o w l ed ge gr ap h q u er y E x e cu t i o n G r a p h s G r a p h C h a t F u n ct i o n al S o l vat i o n DF T C al cu lat io n DF T R esu lt E n e r gy Ob j e ct G r a p h M a p p e r M O F G P U 4Py S C F R o u t e r Age n t X Y Z Un i t C el l F r eq u en c y C o n cep t u al A t o m s Bas i s S e t PO R M AKE Cr y st alN et s G e n e r a l p u r p o se t o o l s Ot h e r t o o l s M OFid Figure 1 High-lev el illustration showing the main comp onen ts of El A gente Gráfico : (i) GraphChat as the user in terface, displa ying real-time even ts such as geometry optimization; (ii) the ob ject graph mapp er (a customized v ersion of Ref. ( 29 )) is used to (de)serialize Python ob jects into a kno wledge graph; (iii) execution graphs for the main w orkflows with router agen ts; and (iv) additional tools, including web search, sandboxed code execution, and knowledge graph interaction. The GraphCh at user interface can b e viewed from the Supplementary Video h ttps://www .y outube.com/pla ylist?list=PLaUD8plXw _ecR7A1EwVAKL3pzIZqjvuVU . calculations as no des connected by directed edges that define admissible data flow, including conditional and cyclic transitions ( e.g. , imaginary-frequency chec ks). Routing Ag ent When the graph to ol is in vok ed, the agent supplies a summary of the user’s request and initial configurations to the start no de. When multiple successor no des are admissible, a fo cused routing con troller (an LLM call) selects and inv okes the next no de via schema-conditioned structured output that b oth constrains transitions and instantiates v alid inputs. The controller can also terminate early once the request is satisfied. Abstraction Layer Across heterogeneous pack ages, the ConceptualA toms class provides a unified in-memory in terface for molecular and perio dic systems, including c harge/multiplicit y verification. P ac k age-sp ecific input/output classes comp ose ConceptualA toms for state representation, enabling on-the-fly transformations ( e.g. , conformer sampling follow ed b y electronic-structure calculations) while maintaining direct Python ob ject references for zero-cop y state transfer. 3 Memory Management Python ob jects created by the agent, either directly or through the execution graph, are p ersisted to a graph database ( K G ). The OGM serializes the Python instances in to K G en tries with unique identifiers (internationalized resource identifier ( IRI )) for subsequent retriev al. This enables retriev al of prior results and allows “heavy” scien tific data to propagate across computation stages without rep eated serialization or re-initialization, while also supp orting relation-aw are queries ov er p ersisted state. T ools and Implementation W e implement the system using p ydantic-ai , leveraging type hints and p ydantic v alidation for runtime type safety and sc hema enforcement across all graph transitions. Domain-sp ecific to ol c hains used in this w ork (molecular generation, quantum c hemistry , mac hine-learned force fields, conformer searc h, and MOF analysis) are detailed in section 5.2 . Additionally , the agent has general-purp ose to ols for w eb search, sandb o xed co de execution, and KG retriev al. Execution Journey The execution journey begins when the agen t in v okes the graph to ol p er molecule, instan tiating multiple graphs in parallel. F or each execution graph, the agent initializes the appropriate ConceptualA toms sub class, a summarized user request, and the configurations needed for the entry no de (t ypically a single-p oin t calculation). If initial configurations are inv alid ( e.g. , incorrect charge/m ultiplicit y), the LLM is automatically reprompted for correction. Each graph initializes with isolated GPU execution con texts to maximize parallelism, typically with three concurrent calculations p er GPU. The routing controller then selects subsequent no des, parameterizing any intermediate settings until the graph exits. The agent retriev es the results, each con taining an instance of ConceptualA toms , executes any subsequent graph/to ol calls as needed, and finally resp onds to the user’s request. Use r G r áf i co a ) S y st e m ov er v i ew t o o l call d ) D y n a m i c r o u t i n g o r e xi t ( E n d N o d e ) - C ol le c t ou t p u t s ( as t o o l r esul t s) - U p d a t e k n o w l e d ge gr a p h ( if co n f igu r ed ) d ) c) S h a r e d r u n t i m e st a t e G r ap h De p e n d e n c i e s - F r on t en d ac cess p or t - Grap h d at ab ase en d p o in t Wo r k f l o w S t a t e (t y p ed ) - C onc ept ua lA t oms - Lar ge nu m er ical ar r ays - N o d e s p ecif i c in p u t s - N o d e lo cal o u t p u t s Wo r k i n g M e m o r y (t ex t u a l ) - S u m m ar ised u ser in t en t - N o d e e xecu t i o n s u m m ar ies Ob j e ct s shi p p ed in P y t hon memory G e o m et r y Op t i m i sat i o n F r e q u en cy C a l cu l at i o n I m a gi n a r y F r eq . R em ov al M o l ec u l ar An a l y si s S t a r t N o d e : S i n gle P o in t C al c u l at i o n b ) G PU-a cce l e r a t e d P y S C F ex e cu t i o n gr a p h (t y p e d ) 1 sup p o r t s di re c ted c yc li c g raph 2 3 d ) ev e r y n o d e can t er m i n at e c) c) TD-DF T Wo r k i n g S u m m a r y No d e A: [ sm il es: ... No d e B: [ coo r d : . .. a d m i ssi b l e n e x t n o d es 3 2 I n p u t sch e m a { "key " : ...} 1 - R e as o n in g - C on f i d en ce - Gener at e d in p u t s { "key " : ...} 3 sel ect n e x t n o d e R o u t er LLM (i n -gr a p h ) u ser pr o m p t s an d in t er act io n st r eam ed even t s t r i g ger f u r t h e r r o u t i n g n o d e ex e cu t i o n Figure 2 Ov erall structure of El A gente Gráfi c o : (a) Users can interact with Gráfico through prompts or by uploading xyz files, while Gráfico pro vides real-time up dates of calculations and molecular tra jectories; (b) Typed execution graph for GPU4PySCF showing the execution no des with m ultiple admissible states that are controlled by the router agent; (c) Generated runtime (typed) states are kept in the knowledge graph for later retriev al and allow real-time user insp ection; (d) Routing agent allows for schema-complian t decision making while instantiating typed argumen ts to the next no de. 4 2.2 Benchmark for performance e v aluation University-le vel quantum chemistry exercises W e ev aluate El Agente Gráfi c o on six quan tum chemistry exercises from El Agente Q ( 5 ), each with tw o difficulty levels. T asks include analysis of organic/inorganic molecules, hydrogen-abstraction energetics, cycloalk ane ring strain, pKa prediction for halogenated acetic acids, and TDDFT excited-state energies. Prompts and rubrics are provided in Supplementary Sec. B.2 and B.3 . Each task was rep eated ten times, for a total of 120 runs p er ev aluated LLM. Models W e selected eigh t LLM s for comparison, including b oth op en-source and proprietary fron tier mo dels. F or b enc hmarking, the agent was restricted to the PySCF workflo w, co de execution, and unit conv ersion to ols to av oid unnecessary degrees of freedom in the ev aluation setting, and the routing controller was fixed as gpt-4o-mini . Within this controlled execution framework, each ev aluated LLM w as resp onsible solely for planning and to ol parameterization, thereb y enabling direct comparison of agent reliability under identical constrain ts. 2.2.1 Automated e valuation framework Ev aluation Strategy Ev aluating LLM -driv en scientific agents p oses unique challenges due to their sto c hastic decision-making, multi-step to ol use, and the absence of a single ground truth in text-based outputs. T o enable scalable b enchmarking, we implemented a fully automated ev aluation framework using p ydantic-ev als that op erates directly on execution traces and structured outputs. In line with the recommended best practices ( 30 ), w e adopt a dual-ev aluator design that indep endently assesses: (i) computational correctness, and (ii) semantic task adherence. The computational ev aluator is a deterministic numerical chec k er that verifies the v alidit y of Python ob jects pro duced by the agent’s execution graph ( e.g. , energies and geometries; details are provided in section 5.1 ). The semantic ev aluator uses an LLM -as-a-judge to assess task completeness, reasoning, and rep orting quality based on the full agent trace, including all to ol calls and final text outputs. Metrics F or each mo del, we rep ort the numerical and LLM judge ev aluation scores, a veraged across all runs. Ov er the agentic trace, we also trac k total tok ens, monetary cost, and the num b er of application programming in terface ( API ) requests. During an agentic trace, all previous messages are resent to the LLM pro vider, creating a “snowball effect” that typically leads to accumulating tok en costs, particularly when accounting for tok ens induced by re-prompting mo dels in the even t of errors. T o further understand the impact of v arying LLM b eha viours, we introduce additional metrics: (i) c ontext window satur ation , whic h measures the ratio of the total tokens in the final LLM API request to its maximum context window allow ed by the provider; (ii) err or r e c overy c ost , which is asso ciated with the exceptions and tracebacks provided to the mo del during error handling; and (iii) c arryover tokens , defined as the ratio of accum ulated cacheable tokens to the total tokens consumed throughout the agentic trace. This metric highligh ts the imp ortance of careful cache management for efficient agentic systems ov er long-horizon tasks. While many providers offer reduced rates for cached tok ens, implementation v aries across pro viders. F or simplicity , our calculations rely on raw token counts and original pricing rather than cached rates. The monetary cost could b e further reduced by employing additional caching strategies 1 , 2 or deploying lo cal mo dels. In T able 1 , we show a summary of the b enc hmarking exp erimen t. Compared to El Agente Q ( 5 ), which adopts a multi-agen t arc hitecture driven by sonnet-3.7 , Gráfico ac hieves substantially higher efficiency in b oth monetary cost and wall-clock time. Op erational cost drops from $4 . 67 to as low as $0 . 17 with gpt-5 ( ≈ 96% reduction), while wall-clock time decreases from 1 , 827 s to 200 – 300 s ( ≥ 6 x sp eedup), along with a reduction in trace token consumption from ∼ 1 . 6 M to ∼ 100 k . These gains arise primarily from the single-agent execution-graph design, which eliminates inter-agen t comm unication ov erhead that would otherwise amplify LLM API requests in multi-agen t systems. The reduction in execution time further demonstrates Gráfico ’s abilit y to exploit parallelized, GPU-accelerated PySCF workflo ws without incurring co ordination latency . F or computational chemistry workflo ws c haracterized by sequential reasoning and strict numerical v alidation, these results imply that once a single agent reac hes a high baseline accuracy , additional multi-agen t co ordination is unlikely to yield prop ortional p erformance gains. Instead, a single agent can pro vide optimal accuracy 1 https://platform.claude.com/docs/en/build- with- claude/prompt- caching 2 https://openrouter.ai/docs/guides/best- practices/prompt- caching 5 T able 1 Performance b enchmarking of LLM driv ers within the Gráfico arc hitecture, av eraged o ver 6 exercises with 2 difficulty levels rep eated 10 times, for a total of 120. The token cost assumes no cached tokens. Context window saturation is the num ber of tokens used in the last LLM API request of the trace divided by the maximum context windo w. Retry cost captures error-reco very ov erhead. Carry ov er tokens capture rep eated pro cessing of cacheable tok ens across successive calls. All calculations were performed on a compute no de with 4 H100 GPUs, a maximum concurrency of 3 calculations p er GPU, and 4 agents running in parallel. The sp ecification of each LLM is provided in Supplemen tary Sec. B.1 . LLM model Numerical ev al. (%) LLM Judge ev al. (%) T race tokens LLM API requests T ok en cost (USD) T ask duration (s) Context window saturation (%) Error recovery cost (%) Carryo ver tokens (%) El Agente Q ( 5 ) 88.25 with human eval 1,649,616 168.43 4.67 1,827 n/a n/a n/a gpt-4.1 93.71 96.52 113,175 5.11 0.25 208 3.59 8.17 66.79 gpt-5 98.88 98.50 83,613 3.32 0.17 228 9.78 9.59 53.21 gpt-5.1 98.37 96.61 99,775 3.65 0.17 211 10.43 8.69 58.19 gpt-5.2 98.69 96.19 95,153 4.40 0.22 180 8.42 8.27 64.59 minimax-m2 89.66 87.61 210,018 6.40 0.05 315 21.93 16.38 78.60 qwen3-max 93.54 90.33 178,671 3.84 0.24 296 26.27 11.82 61.45 sonnet-3.7 93.69 90.94 284,036 9.14 0.92 404 19.46 5.07 86.30 sonnet-4.5 96.07 95.67 320,397 6.58 1.09 273 26.79 9.02 83.28 and cost, particularly for sequential reasoning tasks, in which multi-agen t systems can exhibit up to 39 – 70% p erformance degradation ( 21 ). Bey ond aggregate efficiency W e observ e a consistent pattern in p erformance reflected in reasoning b eha viour across mo del generations within the GPT family . In the ring strain level 2 task, gpt-4.1 frequen tly generates cycloalk enes instead of cycloalk anes, which is resolved in later generations of gpt-5.x or in level 1, which includes more hints. The scores for eac h exercise can b e found in Supplementary Figure S1 and S2 . Under a fixed reasoning budget (configured by the same reasoning_effort parameter), three gpt-5.x mo dels achiev e comparable accuracy , while their reasoning token consumption decreases monotonically across generations (see Supplementary Figure S10 ), consistent with prior observ ations in pure LLM b enc hmarks ( 31 ; 32 ). As ra w accuracy approaches saturation, gains across mo del generations increasingly manifest as reductions in tok en usage rather than further accuracy improv emen ts, aligning with indep endent analyses rep orting a sharp decline in the “cost of intelligence” ( 33 ). W e also observe systematic differences in interaction patterns b et w een GPT and Claude mo dels that affect workflo w efficiency: GPT tends to batch and parallelize to ol calls, while Claude more often interlea v es incremental reasoning with serial to ol use. In our setting, sonnet-3.7 is costlier due to limited parallelization of GPU4PySCF workflo ws, whereas sonnet-4.5 incurs ov erhead from frequent co de-execution calls, suggesting that b etter-aligned (or more sp ecialized) to ols could reduce unnecessary in vocations. This div ergent interaction b ehaviour further reveals a critical engineering challenge: the need to reconcile agentic systems with the temp oral constraints of provider-side prompt caching (typically a 5-minute time-to-liv e ( TTL )). F or high-concurrency mo dels, this results in an “all-or-nothing” cost profile, where the dela y of a single to ol to return within the TTL windo w can trigger a costly re-computation of the en tire con text. F ull discussions can b e found in Supplementary Sec. B.5 . This dual-ev aluator design prov ed essential in practice; across repeated runs of the same task, agents frequen tly exhibited v ariabilit y in workflo w structure and rep orting style despite executing similar underlying computations. The n umerical ev aluator remained stable across such v ariations, while the LLM judge captured differences in task completeness and interpretabilit y . T o rigorously ev aluate agen t p erformance b ey ond statistical means, we adopt tw o complementary metrics that balance capability and reliability: pass@k ( 34 ) and passˆk ( 30 ). W e define pass as numerical score equals to 1.00 and LLM-as-a-judge score greater than 0.90 and achiev e pass@3 of 0.99 and passˆ3 of 0.51 with gpt-5 . Details can b e found in Supplementary Sec. B.4 . 6 A lightweight “bare” LLM agent An agent equipp ed only with w eb search and co de execution was tested on tw o El Agente Q tasks: inorganic comp ounds (L1) and pKa prediction (L2) using gpt-5 . These t wo exercises examine the LLM ’s ability to p erform multi-step tasks that cannot b e p erformed with a single command in PySCF. While it could script molecule setup and PySCF runs through trial and error, it failed key scientific c hecks and was inefficien t compared to Gráfico . F or the inorganic (L1) task, the bare agent built molecules with Op enBabel and ran GPU4PySCF geometry optimization follo wed by single-p oin t calculations. How ever, it made errors in p oint-group detection and p opulation analysis, omitted frequency/imaginary-mo de handling, and pro duced the wrong ClF 3 geometry (trigonal planar instead of T-shap ed). F or pKa (L2), it p erformed geometry optimization and Gibbs free energies but handled solv ation incorrectly , by mixing up COSMO solv ation vs. deprotonation energy , and did not chec k for imaginary frequencies, yielding pKa ≈ − 5 . 0 , which is outside the acceptable range in the rubric. The runs consumed ∼ 40 min/650k tokens (inorganic) and ∼ 16 min/450k tok ens (pKa), roughly an order of magnitude more tokens than Gráfico when using gpt-5 ( ∼ 3 min/25k tokens for inorganic L1 and ∼ 5 min/122k tokens for pKa L2). Due to the sto c hastic nature of these runs, it was infeasible to ev aluate them automatically with rep etition. The light weigh t agent’s capability for on-the-fly web search and iterative co de execution nonetheless demonstrates its p otential for b ootstrapping to ol generation. F ull discussion of these results, including session transcripts, can b e found in Supplementary Sec. B.6 . 2.3 Extending the abstraction W e now demonstrate the extensibility of the Gráfico arc hitecture by introducing additional sp ecialized to ols, show casing use cases in tw o distinct fields that leverage adv anced workflo w orchestration capabilities. These examples illustrate how Gráfico can accommo date complex, multi-step computational chemistry tasks that require dynamic deci sion-making, parallel execution, and robust error handling. More imp ortan tly , they demonstrate the practical utility of the OGM-back ed ConceptualA toms abstraction for seamless data exc hange across different pac k ages and workflo w stages, while supp orting p ersistent storage and retriev al through a s hared KG. 2.3.1 Boltzmann-weighted spectroscopic properties in solution Chemical pro cesses are rarely gov erned by a single static molecular structure; instead, thermodynamic ensem bles usually comprise man y distinct conformations that con tribute to observ able properties ( 35 ). A ccurate prediction of sp ectroscopic prop erties, therefore, requires explicit conformer sampling and Boltzmann- w eighted ensemble av eraging, motiv ating the use of mo dern conformer-search algorithms b ey ond brute-force molecular dynamics ( 35 ; 36 ; 37 ; 38 ). T o demonstrate agentic atomistic simulation workflo ws that co ordinate conformer sampling, solv ation mo delling, and spectral analysis, we assessed solven t effects on electronic absorption sp ectra under b oth implicit and explicit solv ation. In b oth cases, Gráfico enables repro ducible across-to ol handoff via p ersisten t ConceptualA toms iden tifiers. F urther discussions and metho dological details on conformer ensemble generation and Boltzmann-weigh ted sp ectral construction are pro vided in Supplemen tary Sec. C . A summary of the case studies is presented in Figure 3 , which shows the tasks and plots generated b y Gráfico . Implicit solvation T o inv estigate the effect of different implicit solven ts on the absorption sp ectra of a mero- cy anine comp ound (see Supplementary Sec. C.1 ), Gráfico p erformed indep endent and concurrent CREST conformer searches at the GFN2-xTB/ALPB level ( 39 ; 40 ) for water and n -heptane. The ConceptualA toms iden tifiers served as light weigh t handles for dispatching do wnstream PySCF workflo ws in parallel, resulting in five and four density functional theory ( DFT ) calculations, resp ectively , for water and n -heptane at the ω B97X-D4 (SMD)/def2-TZVP level ( 41 ; 42 ; 43 ; 44 ). In the water pip eline, one conformer exhibited an imagi- nary frequency after optimization; consequently , the router returned con trol to the main agent, prompting it to reuse the intermediate geometry in a targeted repair lo op b efore pro ceeding to TDDFT . After deduplication of conformers using an energy-based filter with additional ro ot mean square deviation ( RMSD ) v alidation, and recomputation of Boltzmann weigh ts from DFT energies, ensemble absorption sp ectra w ere generated via w eighted Gaussian broadening. Under the chosen mo del and broadening parameters, the ensemble maxima for w ater and n -heptane ov erlapp ed closely , indicating minimal sp ectral shifts induced by the implicit solv ation. 7 Figure 3 Generation of Boltzmann-w eigh ted absorption spectra using time-dep endent density functional theory ( TDDFT ), showing Gráfico generating (solv ated) conformers and passing their internationalized resource identifiers ( IRI s) to (GPU4)PySCF. Absorption sp ectra plots were generated with Gráfico . In the top plot, the geometry was pro vided in the prompt (not shown here). Prompts were summarized for brevity; complete prompts and transcripts are pr o vided in Supplemen tary Sec. C . This query was completed in 35 minutes with 440k tokens costing $1.11 with gpt-5.2 (medium reasoning effort). Effect of e xplicit solvation T o capture explicit solute–solven t interactions, Gráfico compared absorption sp ectra of 2,3-ep oxybutanol ( 45 ) in the gas phase and in explicit solv ation within a single execution (see Supplemen tary Sec. C.2 ). Gas-phase conformers were generated using CREST, while, in parallel, an explicit solv ation shell of 15 w ater molecules was constructed around the solute using quantum cluster growth ( QCG ) ( 46 ) and further sampled with CREST in its non-co v alent interaction ( --nci ) mo de. Conformers con tributing up to 95 % cum ulativ e Boltzmann weigh t were refined at the ω B97X-D4 (SMD)/def2-SVP , and TDDFT calculations were executed concurrently to exploit conformer-level parallelism. Sp ectra were subsequen tly broadened and combined for a direct visual comparison. The explicitly solv ated ensemble exhibits a pronounced solven t-induc ed red shift of the dominan t absorption band relative to the gas-phase sp ectrum, consisten t with stabilization of the excited state by the surrounding water cluster. T ogether, 8 this demonstrates an agentic system that p erforms high-level scientific planning ov er a structured scientific state while orchestrating sto c hastic sampling, explicit solv ation, and electronic-structure calculations within a unified workflo w. This query was completed in 30 minutes with 185k tokens costing $0.44 with gpt-5.2 (medium reasoning effort). 2.3.2 Interactive design for metal-organic frameworks Reticular chemistry reframes crystalline materials as assemblies of mo dular building blo c ks constrained by top ological frameworks, generating an immense combinatorial space that is ideal for graph-based reasoning ( 47 ; 48 ; 49 ). Na vigating this v ast landscap e ( 50 ; 51 ) requires the orchestration of heterogeneous data sources ( 52 ; 53 ) and sp ecialized to ols ranging from structure generation ( 54 ; 55 ; 56 ) to high-throughput GPU-accelerated mac hine-learned interatomic p oten tials ( MLIP s) ( 57 ; 58 ; 59 ). Connecting these isolated computational steps in to a coherent discov ery pro cess requires a stateful representation that preserves context across to ols. T o demonstrate this agentic reticular design paradigm, we tasked the agent with incrementally constructing and exploring a K G of exp erimen tally rep orted, hypothetical, and graph-inferred MOF s. F urther discussions and metho dological details are provided in Supplementary Sec. D . Figure 4 The MOF w orkflow includes: (1) structure acquisition by CCDC refco de ( 60 ) from CoRE-MOF database ( 52 ), (2) semantic decomp osition of CIF files into top ology , metal no des, and organic linkers, (3) combinatorial searc h within K G to prop ose new hypothetical MOF s, (4) MOF construction with PORMAKE ( 56 ), (5) geometry optimization using GPU-accelerated MLIP s, and (6) p orosity analysis using Zeo++ ( 61 ). These to ols allow Gráfico to pro cess, prop ose and analyze new MOF s, as well as p erform the necessary queries for b oth in-memory and external graph. Prompts w ere summarized for brevity; complete prompts and transcripts are provided in Supplementary Sec. D . Build and e xplore W e tasked Gráfico with pro cessing CIF files and h yp othetical MOF s based on the building blo c ks, and prop ose new MOF s accordingly (see Figure 4 (b) Q1). T o build the requested MOF s, Gráfico 9 dispatc hed seven parallel workflo ws (see Supplementary Sec. D.2.1 ). Across concurrently executed tasks, a deduplication strategy managed by the OGM ensured that top ology and building blo ck IRI s remained unique and consisten t. This was achiev ed via a “retrieve-or-create” strategy across b oth in-memory canonicalization of Python ob jects and K G p ersistence. T o prop ose new MOF s, Gráfico p erformed a combinatorial se arc h based on K G , en umerating feasible (top ology , metal no de, and organic linker) combinations deriv ed from the accum ulated graph entities, and the workflo w subsequently built and analyzed seven teen hypothetical MOF s. Candidates were rank ed by surface area, identifying the highest-p erforming structures among b oth exp erimen tally sourced and newly prop osed MOF s. Agent analysis of the generated structures concluded that altering top ology while keeping the comp onen ts fixed could strongly decouple p ore size from surface area, offering a distinct design lever relative to linker substitution. This query was completed in 10 minutes, consuming 124k tok ens and costing $0.16 with gpt-5.2 (medium reasoning effort). Cross-session persistence W e asked the agent in a new session to query the K G for all previously inv estigated MOF structures con taining a specific metal no de, group them b y top ology , and summarize the observ ed trade-offs b et w een p ore size and surface area ( Figure 4 (b) Q2, see Supplemen tary Sec. D.2.2 ). The agent insp ected the ontology to identify relev ant classes and prop erties, comp osed targeted graph queries to retrieve the p ersisted structures along with their p orosity descriptors, wrote Python co de to compute p er-top ology statistics and rank structures, and synthesized a concise takea wa y . It concluded that for a given top ology , increasing p ore size, esp ecially p ore limiting diameter, was asso ciated with a higher accessible surface area, whereas across top ologies they did not correlate. F ramework density and surface-to-volume effects can cause v ery large-p ore v arian ts to underp erform smaller but more accessible ones. This query was completed in 2 min utes, consuming 136k tokens and costing $0.20 using gpt-5.2 (medium reasoning effort). 3 Discussion The developmen t and ev aluation of Gráfico yield several critical insights into the design of robust LLM - driv en scien tific systems. Our w ork sho ws that scaling scien tific agen ts requires shifting from prompt engineering to context engineering 3 b y externalizing and typing scientific state (via OGM / ConceptualA toms ) suc h that control remains light weigh t, token-efficien t, and robustly parallelizable. Herein, w e discuss the lessons learned and the limitations of our design, along with an outlook to ward future work. These are summarized in the roadmap in Figure 5 . Code generation agents F rom the “bare” LLM exercise, we found that the outputs are only trusted after domain exp erts manually audit b oth the generated co de and the scientific results. This shifts the cognitiv e burden from to ol design to results ev aluation p ost-query , consistent with analyses of AI-assisted engineering ( 62 ). These unconstrained systems (bare agen ts equipp ed with co de execution and web search) can function as dev elopmental scaffolding, rather than as reliable scientific copilots. In domains with real-world consequences, suc h as self-driving labs ( 63 ; 64 ; 65 ; 66 ), strict and v alidated exp erimental pro cedures prior to execution necessitate structured framew orks ( 67 ). T rust through structured ex ecution A scientific copilot is only as useful as the trust it engenders in its outputs. W e therefore em brace engineering techniques that mak e LLM -driv en systems more reliable and scalable ( 68 ). Gráfico enforces schema v alidation with p ydantic b efor e to ols run and execute co de in con trolled environmen ts, aligning with emerging practices in autonomous lab oratory systems ( 67 ). T o av oid the information ov erload often caused by ov erly verbose LLM outputs ( 69 ), Gráfico exp oses a curated, state-driv en interface that makes the internal states co-observ able to users and agen ts alike in real time. F or example, the user can insp ect three-dimensional molecular structures in real time during geometry optimization and review explicit execution traces and logs. Bridging the engineering gap Building on this, we envision a shift in scientific softw are developmen t fo cused on runtime-generated to ol rep ositories ( 70 ; 71 ). While light w eight agents incur high tok en costs and require the sup ervision noted ab o ve, their capacity to rapidly generate functional prototypes marks a new frontier, 3 https://www.anthropic.com/engineering/ef fectiv e- context- engineering - f or- ai- agents 10 reducing hours of manual developmen t to moments. W e argue that the primary challenge is no longer LLM co de generation capability , but rather the transition from agent-generated prototypes to robust systems. By automating ground-truth generation and verification lo ops, one can harness the sp eed of rapid prototyping while maintaining the scientific rigour essential to the field. 0 T ext -c en t r i c agen t i c s yst em Ag e n t i c Ar ch i t e ct u r e C h e m i st r y C a p a b i l i t y S t r u c t u r e d ex e cu t i o n - T yp ed execu t i o n grap h - K G f o r p er s i st en ce 1 Di st r i b u t ed & l o n g-h o r i zo n a ge n t s - S el f -d r i vi n g lab s - M u l t i -m o d al m o n it o r i n g - Web o f agent s - S el f -evol vi n g agen t s 4 S e m a n t i c b o u n d a r y ev o l u t i o n - R eact i o n n et w o r k s - H yp o t h es i s gen er at io n - On t o lo gy evol u t io n - V er s i o n ed t o o l gen er at i o n 3 Asy n ch r o n o u s & r e so u r ce -a w a r e - F u ll P yS C F f u n ct io n a li t i es - M L IP and M D - P ar t ial r esu lt s r et u r n - Do cker an d san d b o x 2 Figure 5 Curren t stage and roadmap of future work showing: (1) structured execution of typed execution graphs and to ols (as implemented in the current Gráfico system); (2) asynchronous and resource-a ware execution to enable proactiv e agents; (3) semantic b oundary evolution allowing rapid extension of to ols and ontologies; (4) long-horizon agen ts tow ards a distributed netw ork of AI scientists. Asynchronous and resource-aware environment The curren t single-run time design enables lo w-ov erhead, in-memory state sharing but implicitly couples workflo ws through global GPU con texts and dep endencies, requiring careful parallelism configuration ( e.g. , multi-threading or by the launc h of computational subpro cesses) and making hardware resource p olicies implicit. Moreov er, maintaining heterogeneous to ols via custom forks is brittle and do es not scale. F uture work will encapsulate domain-sp ecific workflo ws as p ortable agent skills 4 , exp ose resource and environmen t primitives as first-class concepts ( e.g. , device selection, memory/concurrency limits), and adopt hybrid, selectively isolated containerization and/or sandb oxing ( 72 ) for high-risk to ols to balance fault con tainment and repro ducibilit y with low-latency comp osabilit y . Semantic boundary ev olution Gráfico seman tic b oundaries are reflected in b oth the design of the ontologies underlying the execution graphs and the automatic ev aluation. Nonetheless, the b enc hmark construction and the underlying ontological definitions remain time-consuming and lab orious. Because agen ts are inheren tly probabilistic, they often discov er solution pathw ays that fall outside manually-specified rubrics, making it increasingly difficult to ev aluate accurately ( 73 ; 74 ). These observ ations allude to a cen tral challenge for future researc h, which is the co-evolution of semantic representation and the ev aluation framework, in direct analogy to how unit tests co-evolv e with softw are. F uture work will expand the semantic space through con trolled ontology refinement to guide adaptive to ol generation while simultaneously up dating the ev aluation criteria. Progress along these directions would transform b enc hmarking from a man ual, p ost-ho c pro cess in to an integral comp onent of agentic system design and optimization. Distributed and long-horizon tasks Distributed and long-horizon tasks require more adv anced context man- agemen t strategies. While Gráfico effectiv ely externalizes state to minimize token usage, the curren t design assumes a single-agen t, single-session con text operating o ver a persistent K G . Extending this to 4 https://claude.com/blog/equipping- agen ts- f or- the- real- world- w ith- agent- skills 11 m ulti-agent collab oration or long-term op eration introduces not only c hallenges in synchronizing shared state, managing divergen t K G s, and ensuring consistency across sessions, but also the need for richer context graphs that capture evolving decision traces across agents and time 5 . F uture work could explore distributed K G arc hitectures, versioned ontology expansion sync hronization with git 6 , and inter-agen t communication proto cols to supp ort collab orative scientific workflo ws. Additionally , mechanisms for temp oral reasoning and memory consolidation w ould enable agents to build up on prior knowledge ov er extended time horizons. 4 Conclusions In this work, we introduce El A gente Gráfi co , a t yp e-safe agen tic framework that em b eds LLM reasoning within explicit execution graphs and structured computational state, enabling robust orchestration of complex scien tific workflo ws. By representing scientific concepts, intermediate results, and execution con text as typed ob jects (rather than unstructured text), the system decouples high-level decision making from low-lev el execution while preserving prov enance and auditability . Across quantum chemistry and materials design tasks, including conformer search and MOF design space exploration, this architecture supp orts parallel execution and failure recov ery , with efficient con text management enabling these tasks to run within a single agent. Benc hmarking against a previous multi-agen t system ( 5 ) demonstrated ov er a 14-fold reduction in token usage with ov erall improv ed accuracy . Our results suggest that as agentic systems mov e tow ard p ersistent, stateful op eration, their reliability is increasingly gov erned b y execution semantics rather than by prompt engineering alone: sp ecifically , by how to ols and concepts are represen ted and constrained. By foregrounding t yp e-safe abstractions and explicit execution structure, this work reframes agen tic scientific w orkflows as a systems engineering problem and provides a concrete foundation for scalable and extensible AI-driv en scientific disco very . Ultimately , we envision deploying Gráfico as part of the El Agente family ( 5 ; 14 ; 15 ; 16 ; 17 ) on the cloud, facilitating the global demo cratization of science. 5 Methods 5.1 Gráfico infrastructure The ob ject graph mapp er (OGM) in this work is a customized version of the W orld A v atar ( 75 ; 76 ; 77 ) Python pac k age t wa ( 29 ), refactored to provide a fully Python-native interface and to align more closely with Pydantic data structures and usage patterns. This OGM provides a bidirectional serialization lay er in which Python classes are mapp ed to classes and relationships in on tologies, with strict type enforcement. Key custom features include a lazy-loading mechanism that efficiently retrieves subgraph data on demand when pro cessing large-scale ontologies, and sp ecialized supp ort for serializing NumPy arrays using n umpydan tic . By integrating with rdflib and p ydantic-ai , the OGM ensures that complex chemical data structures are represented with formal semantic v alidation while maintaining idiomatic Python interoperability for downstream in tegration with LLM-based agen ts. The computational chemistry agent was instantiated using the p ydantic-ai framew ork, configured with a system prompt that injects the workflo w constrain ts and domain-sp ecific instructions. T o ols are strictly t yp ed using Pydantic mo dels and registered via a to ol decorator or through the to ols list, enabling the LLM to inv oke complex functions with v alidated schemas. The framework’s dep endency injection system is critical for our architecture, passing a GraficoDeps ob ject to to ols that provide access to shared resources lik e the frontend and K G endp oin ts. W e also leverage pydan tic-ai’s mo del context proto col (MCP) supp ort to in tegrate external execution environmen ts ( e.g. , a sandb o xed Python runner) and its Mo delRetry mec hanism to handle to ol execution failures by providing feedback lo ops back to the mo del. The LLM s w ere configured with temp eratures of 0 for non-reasoning mo dels and 1 for mo dels using explicit reasoning mo des. Custom GPU scheduling was developed for GPU4PySCF to preven t CUDA context initialization conflicts and illegal memory access patterns common in multi-threaded GPU4PySCF jobs. The concurrency levels w ere managed by a thread-safe tok en queue, defaulting to three execution slots p er GPU to optimize device 5 https://neo4j.com/blog/agen tic- ai/hands- on- with- context- graphs- and- neo4j/ 6 Analogous to agent teams collectively w orking on a co debase: https://www.anthropic.com/engineering/building- c- compiler 12 utilization while preven ting memory exhaustion. Upon task submission, a dedicated scheduler assigns an a v ailable GPU identifier and execution slot, filtering visible devices through environmen t v ariables. Each w orkflow executes in a separate OS-level child pro cess while sharing the same CUDA driver context. In ter- pro cess communication is handled via pickled execution pa yloads. In tegrated distributed tracing (via logfire) pro vides real-time observ abilit y into slot o ccupancy . The ev aluation b enc hmark was developed using p ydantic-ev als . The system assesses the agent’s p erformance b y extracting structured workflo w results from to ol outputs and comparing them against the ground truth. The ev aluation metho dology employs a multi-dimensional scoring metric that v erifies correctness across several c hemical prop erties: (i) parameter integrit y , to ensure that the selected functional, basis set, charge, and m ultiplicity match the intended calculation; (ii) energetic accuracy , which v alidates total energies against reference v alues within a threshold of 0.01 Ha; (iii) structural fidelity , quantified by the RMSD of atomic p ositions relative to reference geometries within a threshold of 0.15 Å, utilizing sp yrmsd to account for symmetry and atom indexing p ermutations; and (iv) prop ert y analysis, which chec ks derived observ ables suc h as dip ole moments, HOMO-LUMO gaps (threshold of 0.1 Ha), p oint group symmetry , and vibrational frequencies (ensuring no imaginary mo des for stable minima). Scores are aggregated and normalized across these dimensions to yield a composite accuracy metric for each test case, with distributed tracing (via logfire ) used to monitor agent reasoning and to ol-in v o cation patterns during execution. T oken counts and cost information w ere tak en from the logfire records. F or ev aluations p erformed prior to the release of p ydantic-ev als==v1.23.0 (2025-11-25), token coun ts recorded by logfire w ere corrected to accoun t for a double-coun ting bug ( h ttps ://github.c om/pydantic /p yd an tic- ai/issues/ 3529 ), which was resolved in the v1.23.0 release. 5.2 Computational tools The pro ject employ ed uv for a virtual environmen t using Python 3.12.12. All computational pack ages w ere installed in the same en vironment to enable single-agent op eration using a shared Python runtime. T o resolv e dep endency conflicts and enable custom functions, some pack ages were forked from their original rep ositories. Sp ecific mo difications are noted for each pack age. Where feasible and once the interfaces stabilize, we intend to upstream generalizable improv ements to the original rep ositories via pull requests. 1. py dantic and numpy dantic : The customized ob ject graph mapp er is implemented using p ydantic (v2.11.7) for basic t yp ed classes and nump ydan tic (v1.6.11) for serialisation of NumPy arrays. 2. py dantic-ai family : The agentic framew ork and workflo w execution are implemented using p ydantic-ai (v1.44.0), which provides type-safe agent definitions, to ol registration, and v alidated data exc hange. Structured workflo w graphs are managed via the p ydantic-graph comp onen t, with customized base no de classes for dynamic routing ov er t yp ed no de definitions. The framework was iteratively up dated during developmen t to incorp orate new features and was stabilized at version v1.44.0. Execution traces, to ol in vocations, and structured state transitions were instrumented using logfire (accessed via p ydantic- ai ) to enable fine-grained observ ability , debugging, and post-ho c analysis of agen t b eha viour. F or ligh tw eigh t external information retriev al, the agent utilised the DuckDuc kGo search to ol via p ydantic_- ai.common_to ols.duc kduc kgo , allowing web search without direct dep endency on proprietary search APIs or in ternal web search to ols that are only av ailable for proprietary LLM mo dels. 3. PySCF family : Density functional theory calculations were p erformed using the p yscf (v2.10.0), with gpu4p yscf-cuda12x (v1.5.0) for GPU acceleration on CUDA 12.x hardware and p yscf-disp ersion (v1.3.0) for semiclassical disp ersion corrections ( 78 ; 79 ; 80 ; 42 ; 81 ). 4. PubChemPy : The molecular generation from names or SMILES was implemen ted using pub c hemp y (v1.0.4) ( 82 ; 83 ). 5. RDKit : The molecular generation from SMILES was implemented using rdkit (v2025.3.5) ( 84 ). 6. OpenBabel : In cases where RDKit failed, op en bab el-wheel (v3.1.1.22) (a prebuilt wheel through PyPI) w as used ( 85 ; 86 ). 7. QCElemental : Elemental data, physical constants, and unit conv ersions w ere standardized using qcele- men tal (v0.29.0), ensuring consisten t representation of molecular quantities across electronic-structure 13 calculations and w orkflow comp onen ts. 8. QCG and CREST : The conformer and rotamer ensem ble sampling to ol CREST generates rank ed en- sem bles of lo w-energy conformers for a given three-dimensional molecular structure ( 36 ; 37 ). Using iterativ e metadynamics simulations ( 87 ) in combination with conv en tional molecular dynamics, CREST systematically samples the p otential energy surface and explores the accessible conformational space. In this work, we implemented its GPU-accelerated v arian t describ ed in Ref. ( 25 ) (commit 75135dd ). F or the generation of explicitly solv ated molecular structures, w e use the Quantum Cluster Growth (QCG) algorithm ( 46 ) imple men ted in crest-3.0.2 together with xtb-6.7.1 . 9. PORMAKE : PORMAKE ( 56 ) enables top ology-aw are assembly of hypothetical MOF s from predefined building blo c ks (metal no des and organic link ers). Building blo cks are assigned to top ology connection sites using RMSD-based matching of lo cal co ordination environmen ts (RMSD ≤ 0.3 Å). PORMAKE’s exp erimen tal decomp osition routines are used to extract provisional building blo cks from existing CIF files. W e use a GitHub snapshot of PORMAKE pinned to commit 639caad ( h ttps://github.com/San gwon91/PORMAKE ), ensuring repro ducible b eha viour. 10. CoRE-MOF- T ools : CoRE-MOF-T o ols (pack age: coremof-to ols , v0.3.1) ( 52 ) is used for prepro cessing exp erimen tal CIF files, including solv en t remov al and structural normalization, and for in terfacing with Zeo++ to compute p orosit y-related descriptors. W e use a fork of CoRE-MOF-T o ols ( ht t p s : //github.com/jb2197/CoRE- MOF- To ols ) in whic h mo difications are confined to setup.p y to resolv e dep endency incompatibilities with the broader softw are environmen t. Sp ecifically , strict v ersion pins are relaxed to compatible low er b ound constraints ( e.g. , scikit-learn==1.3.2 to scikit-learn>=1.3.2 ). The fork also redirects the MOF Classifier ( 88 ) dep endency to a compatible fork ( http s: // gi thu b. c om/jb2197/MOF Classif ier ), where analogous changes are applied in setup.p y : strict version pins for n umpy , torc h , p ymatgen , scikit-learn , tqdm , and pandas are relaxed to lo wer-bounded constraints, the requests pac k age is added to install_requires , and an unused requests imp ort is remov ed from the setup script. 11. MOFid : MOFid (upstream v1.1.0; forked and released as v1.1.1 with mofid-v2 utilities) provides the con vert_ase_p ymat function for ASE-to-pymatgen con v ersion, used during building blo c k deduplication. This conv ersion enables structural comparison via pymatgen’s StructureMatc her , as part of a second- stage verification when PORMAKE’s exp erimental hash-based decomp osition yields multiple candidate metal nodes or organic linkers. W e use a fork of MOFid ( ht t p s :/ / g i t h u b .c o m / s wa n i ck t /m o fid ) that corrects the upstream pac k age distribution to make the mofid-v2 analysis co de programmatically accessible. Sp ecifically , the fork up dates setup.p y to include the mofid_v2 mo dules in the installable pac k age set, adds missing __init__.p y files, refactors the no de-grouping logic into an imp ortable library mo dule ( group_no de_b y_structure_library .py ), and updates dep endency declarations to supp ort reliable installation in Python environmen ts. These changes remov e script-style side effects presen t in the upstream implementation and enable direct programmatic use within Python workflo ws. 12. pymatg en : p ymatgen (v2024.8.9) ( 89 ) pro vides structure-level equiv alence c hecking throughout the w orkflow. F or building blo c k deduplication, StructureMatc her compares candidate structures conv erted from ASE via MOFid’s con vert_ase_p ymat utilit y . F or general structure v alidation and matching in the ConceptualA toms data mo del, pymatgen’s native AseA tomsAdaptor p erforms ASE-to-pymatgen con version, with StructureMatc her used for p erio dic structures and MoleculeMatc her for non-p eriodic molecules. T ogether, these capabilities supp ort deduplication, K G queries for equiv alen t comp onen ts, and structure v alidation. 13. CrystalNets.jl : CrystalNets.jl (v1.1.0) ( 90 ), accessed via juliacall (v0.9.29), is used for top ological classification of MOF structures. The pack age determines the underlying net top ology , dimensionalit y , and catenation from input CIF files. W e emplo y the SingleNo des clustering mode, in which each iden tified building blo c k is mapp ed to a single vertex. 14. Z eo++ : Zeo++ (v0.3) ( 61 ) is used for p orosity analysis, including largest cavit y diameter (LCD), p ore limiting diameter (PLD), largest free sphere path diameter (LFPD), accessible surface area, p ore volume, and framework dimensionality . Zeo++ is accessed through the CoRE-MOF-T o ols in terface and is obtained from the zeo++/0.3 mo dule directly loaded on the trillium-gpu HPC compute no de. 14 15. MA CE : MACE-torc h (v0.3.14) ( 58 ) provides machine-learned in teratomic p oten tials for geometry op- timization. W e employ MACE-MP-MOF0 ( mof-omat-0-v2 ) to p erform structure relaxation prior to p orosit y analysis. W e use a fork ( http s:/ /gi th ub. com /j b21 97/ ma ce ) that relaxes the strict dep en- dency constrain t e3nn==0.4.4 to an unpinned version ( e3nn ), and further in our environmen t, we pin e3nn==0.5.0 to ensure compatibility with other pack ages (MatterSim). 16. MatterSim : MatterSim (v1.2.0) ( 91 ) is supp orted as an alternative machine-learned force-field back end for geometry optimization within the workflo w. 17. Orb models : The Orb mo dels (v0.5.4) ( 92 ) pro vide an additional machine-learned interatomic p otential option for geometry relaxation. 18. ASE : The Atomic Simulation En vironment (v3.26.0) ( 93 ) serves as the common data structure for atomic configurations throughout the workflo w. Building blo cks from the PORMAKE decomp oser, structures for MLIP optimization, and final MOF geometries are all represented as ASE Atoms ob jects. 19. RDFLib and SP ARQL Wrapper : RDFLib (v7.1.4) is used for RDF serialization and lo cal graph op erations, while SP ARQL W rapp er (v2.0.0) handles remote SP ARQL queries against the K G . T ogether, they enable com binatorial search algorithms, equiv alence queries for comp onen t deduplication, and p ersistence of w orkflow results. 20. Blazegraph : Blazegraph (engine v2.1.6) serves as the triplestore back end, hosting the p ersistent K G and exposing a SP ARQL endpoint for querying MOF comp onen ts, top ologies, and compatibility relationships. The K G is deplo yed using the Cambridge CARES Do ck er image ghcr.io/cam bridge- cares/blazegraph:1.2.0 (con tainer registry: h ttps://gi thub.com/orgs/c am bridge- cares/pack ages/co ntainer/ package/ blaze gra ph ), which provides a standardized containerized runtime consistent with the W orld A v atar KG infrastructure. 21. mcp-run-python : Sandb oxed Python co de execution mcp-run-p ython w as used as a mo del context proto col server, where a sp ecific forked version ( h ttps://github.com/jb2197/mcp- run- python/releas es/tag/0.0.22.2- f ile ) was used to enable output files. 22. PythonREPL : The Python co de execution to ol from langc hain-exp erimen tal (v0.4.0) used for the “bare LLM ” agen t b enc hmark. This to ol should b e used with caution as it can execute arbitrary co de on the host machine ( e.g. , delete files, make netw ork requests). 5.3 Graphical user interface The Graphical User Interface (GUI) is implemented using the React framew ork. Comm unication b et ween the frontend and bac kend is facilitated by the GraphChat SDK, a custom Python library that utilizes the W ebSo ck et proto col for efficien t data exchange. The k ey features of the c hat interface include real-time streaming of agent resp onses, visibility into to ol execution logs, and display of the agent’s reasoning steps, all implemen ted using the Pydantic AI framework and rendered via p ycrdt for synchronization and real-time up dates. F urthermore, the interface dynamically renders a molecular viewer up on agent activ ation. When the agent generates molecules, the molecule viewer uses the corresp onding conceptual atom IRI as a unique iden tifier. The molecule viewer supp orts b oth molecular and p erio dic systems, with in teractive 3D rendering, optional atom selection, and the ability to send snapshots (captured images) to the LLM . The molecule viewer visualizes the molecular tra jectory in real-time and allows users to download structures (and tra jectories) in standard formats suc h as (extended) xyz and CIF. Additionally , plots generated by calls to the co de execution to ol are display ed directly in the chat interface for immediate in terpretation. The frontend also supp orts dynamic LLM mo del switching, allowing users to exp erimen t with different mo dels on the fly without in terrupting the ongoing session. A cknowledgments The authors thank Yiqun Bian for the helpful discussions in data analysis and Dr. Pit Steinbac h for the helpful suggestions in setting up the GPU-accelerated implementation of CREST. J.B. ackno wledges funding from the Eric and W endy Schmidt AI in Science Postdoctoral F ellowship Program, a program by Sc hmidt 15 F utures. A.A. ackno wledges supp ort from King Ab dullah Universit y of Science and T echnology (KAUST), Kingdom of Saudi Arabia, for the KAUST Ibn Rushd P ostdo ctoral F ellowship. Y.K. ackno wledges supp ort from the CIF AR AI Safety Catalyst A ward (Catalyst F und Pro ject #CF26-AI-001). T.W.K. ackno wledges the supp ort of the V ector Distinguished Postdoctoral F ellowship. A.A.-G. thanks Anders G. F røseth for his generous supp ort. A.A.-G. also ackno wledges the generous supp ort of Natural Resources Canada and the Canada 150 Research Chairs program. This work was supp orted by the AI2050 program of Schmidt Sciences. This work was supp orted by the Defense Adv anced Research Pro jects Agency (DARP A) under Agreement No. HR0011262E022.This research is part of the Univ ersity of T oronto’s Acceleration Consortium, which receiv es funding from the CFREF-2022-00042 Canada First Research Excellence F und. This research was enabled in part b y supp ort provided by SciNet HPC Consortium ( https://sci nethpc. ca/ ) and the Digital Researc h Alliance of Canada ( h ttps://www.alliancecan.ca ). Computations were p erformed on the T rillium sup ercomputer at the SciNet HPC Consortium. SciNet is funded b y: the Canada F oundation for Innov ation; the Gov ernment of Ontario; On tario Research F und - Researc h Excellence; and the Universit y of T oronto. R eferences [1] Andres M. Bran, Sam Cox, Oliver Schilter, Carlo Baldassari, Andrew D. White, and Philipp e Sch waller. Augment- ing Large Language Mo dels with Chemistry T o ols. Nat. Mach. Intel l. , 6(5):525–535, 2024. doi:10.1038/s42256- 024-00832-8 . [2] Daniil A. Boiko, Rob ert MacKnight, Ben Kline, and Gab e Gomes. Autonomous Chemical Research with Large Language Mo dels. Natur e , 624(7992):570–578, 2023. doi:10.1038/s41586-023-06792-0 . [3] Jura j Gottw eis, W ei-Hung W eng, Alexander Daryin, T ao T u, Anil Palepu, Petar Sirko vic, Artiom Myask ovsky , F elix W eissenberger, Keran Rong, Ryutaro T anno, Khaled Saab, Dan Popovici, Jacob Blum, F an Zhang, Katherine Chou, A vinatan Hassidim, Burak Gokturk, Amin V ahdat, Pushmeet Kohli, Y ossi Matias, Andrew Carroll, Kavita Kulk arni, Nenad T omasev, Y uan Guan, Vikram Dhillon, Eeshit Dhav al V aishnav, Byron Lee, Tiago R D Costa, José R Penadés, Gary Peltz, Y unhan Xu, Annalisa Pa wlosky , Alan Karthikesalingam, and Vivek Natara jan. T ow ards an AI Co-Scientist. arXiv Preprint , 2025. doi:10.48550/arxiv.2502.18864 . [4] Ludo vico Mitchener, Angela Yiu, Benjamin Chang, Mathieu Bourdenx, Tyler Nadolski, Arvis Sulov ari, Eric C. Landsness, Daniel L. Barabasi, Siddharth Naray anan, Nicky Ev ans, Shriya Reddy , Martha F oiani, Aizad Kamal, Leah P . Shriver, F ang Cao, Asmamaw T. W assie, Jon M. Laurent, Edwin Melville-Green, Mayk Caldas, Alb ert Bou, Kaleigh F. Rob erts, Sladjana Zagorac, Timothy C. Orr, Miranda E. Orr, Kevin J. Zwezdaryk, Ali E. Ghareeb, Laurie McCo y , Bruna Gomes, Euan A. Ashley , Karen E. Duff, T onio Buonassisi, T om Rainforth, Randall J. Bateman, Michael Sk arlinski, Samuel G. Ro driques, Michaela M. Hinks, and Andrew D. White. K osmos: An AI Scien tist for Autonomous Discov ery. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2511.02824 . [5] Y unheng Zou, Austin H. Cheng, Ab dulrahman Aldossary , Jiaru Bai, Shi Xuan Leong, Jorge Arturo Camp os- Gonzalez-Angulo, Changhy eok Choi, Cher Tian Ser, Gary T om, Andrew W ang, Zijian Zhang, Ilya Y ak av ets, Han Hao, Chris Creb older, V arinia Bernales, and Alán Aspuru-Guzik. El Agente: An autonomous agent for quantum c hemistry. Matter , 8(7):102263, 2025. doi:10.1016/j.matt.2025.102263 . [6] Key an Ding, Jing Y u, Junjie Huang, Y uchen Y ang, Qiang Zhang, and Hua jun Chen. SciT o olAgen t: A knowledge- graph-driv en scientific agent for multitool integration. Nat. Comp. Sci. , 5(10):962–972, 2025. doi:10.1038/s43588- 025-00849-y . [7] Y eonghun Kang and Jihan Kim. ChatMOF: An artificial intelligence system for predicting and generating metal- organic frameworks using large language mo dels. Nat. Commun. , 15(1), 2024. doi:10.1038/s41467-024-48998-4 . [8] Sh uxiang Cao, Zijian Zhang, Mohammed Alghadeer, Simone D F asciati, Michele Piscitelli, Mustafa Bakr, Peter Leek, and Alán Aspuru-Guzik. Automating quantum computing lab oratory exp eriments with an agent-based AI framew ork. Patterns , 6(10), 2025. doi:10.1016/j.patter.2025.101372 . [9] Thang D. Pham, Adit ya T anik an ti, and Murat Keçeli. ChemGraph as an agentic framework for computational c hemistry workflo ws. Commun. Chem. , 9(1), 2026. doi:10.1038/s42004-025-01776-9 . [10] Ziqi W ang, Hongshuo Huang, Hancheng Zhao, Changwen Xu, Shang Zhu, Jan Janssen, and V enk atasubramanian Visw anathan. D REAMS: Density F unctional Theory Based Research Engine for Agentic Materials Simulation. arXiv Preprint , 2025. doi:10.48550/arxiv.2507.14267 . 16 [11] Theo Jaffrelot Inizan, Sherry Y ang, Aaron Kaplan, Y en-hsu Lin, Jian Yin, Sab er Mirzaei, Mona Ab delgaid, Ali H. Ala wadhi, Kw angHw an Cho, Zhiling Zheng, Ekin Dogus Cubuk, Christian Borgs, Jennifer T. Chay es, Kristin A. P ersson, and Omar M. Y aghi. System of Agentic AI for the Discov ery of Metal-Organic F rameworks. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2504.14110 . [12] Jinming Hu, Hassan Na waz, Y uting Rui, Lijie Chi, Arif Ullah, and Pa vlo O. Dral. Aitomia: Y our intelligen t assistan t for ai-driven atomistic and quantum chemical simulations. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2505.08195 . [13] Carles Nav arro, Mariona T orrens, Philipp Thölke, Stefan Do err, and Gianni De F abritiis. Speak to a Protein: An In teractive Multimo dal Co-S cien tist for Protein Analysis. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2510.17826 . [14] Ignacio Gustin, Luis Mantilla Calderón, Juan B. Pérez-Sánc hez, Jérôme F. Gonthier, Y uma Nak amura, Karthik P anick er, Manav Ramprasad, Zijian Zhang, Y unheng Zou, V arinia Bernales, and Alán Aspuru-Guzik. El Agente Cuán tico: Automating Quantum Simulations. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2512.18847 . [15] Juan B. Pérez-Sánc hez, Y unheng Zou, Jorge A. Camp os-Gonzalez-Angulo, Marcel Müller, Ignacio Gustin, Andrew W ang, Han Hao, T sz W ai K o, Changhy eok Choi, Eric S. Isbrandt, Mohammad Ghazi V akili, Hany ong Xu, Chris Creb older, V arinia Bernales, and Alán Aspuru-Guzik. El agente quntur: A research collab orator agent for quantum c hemistry . arXiv Pr eprint , 2026. doi:10.48550/arxiv.2602.04850 . [16] Changh yeok Choi, Y unheng Zou, Marcel Müller, Han Hao, Y eonghun Kang, Juan B. Pérez-Sánc hez, Ignacio Gustin, Han yong Xu, Mohammad Ghazi V akili, Chris Creb older, Alán Aspuru-Guzik, and V arinia Bernales. El Agente Estructural: An Artificially Intelligen t Molecular Editor. arXiv Pr eprint , 2026. doi:10.48550/arxiv.2602.04849 . [17] Sai Govind Hari Kumar, Y unheng Zou, Andrew W ang, T sz W ai Ko, Jesus V aldez Hernandez, Nathan Y u, V arinia Bernales, and Alán Aspuru-Guzik. El Agente Solido: A New Agent for Solid State Simulations. 2026. [18] Jiaqi W ei, Y uejin Y ang, Xiang Zhang, Y uhan Chen, Xiang Zhuang, Zhangyang Gao, Dongzhan Zhou, Guangshuai W ang, Zhiqiang Gao, Juntai Cao, Zijie Qiu, Ming Hu, Chenglong Ma, Shixiang T ang, Junjun He, Chunfeng Song, Xuming He, Qiang Zhang, Chenyu Y ou, Sh uang jia Zheng, Ning Ding, W anli Ouyang, Nanqing Dong, Y u Cheng, Siqi Sun, Lei Bai, and Bow en Zhou. F rom AI for Science to Agentic Science: A Survey on Autonomous Scientific Disco very. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2508.14111 . [19] Mert Cemri, Melissa Z. Pan, Shuyi Y ang, Lakshy a A. Agraw al, Bhavy a Chopra, Rishabh Tiwari, Kurt Keutzer, A dity a Paramesw aran, Dan Klein, Kannan Ramchandran, Matei Zaharia, Joseph E. Gonzalez, and Ion Stoica. Wh y Do Multi-Agent LLM Systems F ail? arXiv Pr eprint , 2025. doi:10.48550/arxiv.2503.13657 . [20] Arpandeep Khatua, Hao Zhu, Peter T ran, Arya Prabhudesai, F rederic Sadrieh, Johann K. Lieb erwirth, Xink ai Y u, Yic heng F u, Michael J. Ryan, Jiaxin Pei, and Diyi Y ang. Co op erBenc h: Why Co ding Agents Cannot b e Y our T eammates Y et. arXiv Preprint , 2026. doi:10.48550/arxiv.2601.13295 . [21] Y ubin Kim, Ken Gu, Chanw oo Park, Chunjong Park, Sam uel Schmidgall, A. Ali Heydari, Y ao Y an, Zhihan Zhang, Y uchen Zhuang, Mark Malhotra, Paul Pu Liang, Hae W on Park, Y uzhe Y ang, Xuhai Xu, Yilun Du, Shw etak P atel, Tim Althoff, Daniel McDuff, and Xin Liu. T ow ards a Science of Scaling Agent Systems. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2512.08296 . [22] Ew a Deelman, Gurmeet Singh, Mei-Hui Su, James Blythe, Y olanda Gil, Carl Kesselman, Gaurang Mehta, Karan V ahi, G Bruce Berriman, John Go o d, Anastasia Laity , Joseph C. Jacob, and Daniel S. Katz. Pegasus: A F ramework for Mapping Complex Scientific W orkflows onto Distributed Systems. Sci. Pr o gr am. , 13(3):219–237, 2005. doi:10.1155/2005/128026 . [23] Ry an Chard, Jim Pruyne, Kurt McKee, Josh Bryan, Brigitte Raumann, Rachana Ananthakrishnan, Kyle Chard, and Ian T. F oster. Globus Automation Services: Research Pro cess Automation across the Space–Time Contin uum. F utur e Gener. Comput. Syst. , 142:393–409, 2023. doi:10.1016/j.future.2023.01.010 . [24] Xiao jie W u, Qiming Sun, Zhichen Pu, Tianze Zheng, W enzhi Ma, W en Y an, Y u Xia, Zhengxiao W u, Mian Huo, Xiang Li, W eiluo Ren, Sheng Gong, Y umin Zhang, and W eihao Gao. Enhancing GPU-Acceleration in the Python- Based Si m ulations of Chemistry F rameworks. WIREs Comput. Mol. Sci. , 15(2), 2025. doi:10.1002/wcms.70008 . [25] Pit Steinbac h and Christoph Bannw arth. A cceleration of Semiempirical Electronic Structure Theory Calculations on Consumer-Grade GPUs Using Mixed-Precision Density Matrix Purification. J. Chem. The ory Comput. , 21(15): 7335–7351, 2025. doi:10.1021/acs.jctc.5c00262 . [26] Mario Krenn, Florian Häse, AkshatKumar Nigam, Pascal F riederich, and Alan Aspuru-Guzik. Self-Referencing Em b edded Strings (SELFIES): A 100% Robust Molecular String Representation. Mach. L e arn.: Sci. T e chnol. , 1 (4):045024, 2020. doi:10.1088/2632-2153/aba947 . 17 [27] Stephen R Heller, Alan McNaught, Igor Pletnev, Stephen Stein, and Dmitrii T chekho vskoi. InChI, the IUP AC In ternational Chemical Identifier. J. Cheminf. , 7:23, 2015. doi:10.1186/s13321-015-0068-4 . [28] Matthew Muhob erac, Atharv a Parikh, Nirvi V akharia, Saniya Virani, Aco Radujevic, Sav annah W o od, Meghav V erma, Dimitri Metaxotos, Jeyaraman Soundarara jan, Thierry Masquelin, Alexander G. Go dfrey , Sean Gardner, Dobrila Rudnicki, Sam Michael, and Gaurav Chopra. State and Memory is All Y ou Need for Robust and Reliable AI Agen ts. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2507.00081 . [29] Jiaru Bai, Simon D. Rihm, Aleksandar Kondinski, F abio Saluz, Xinhong Deng, George Brown bridge, Sebastian Mosbac h, Jethro Akroyd, and Markus Kraft. tw a: The w orld av atar python pack age for dynamic knowledge graphs and its application in reticu lar chemistry . Digital Disc overy , 4(8):2123–2135, 2025. doi:10.1039/d5dd00069f . [30] Engineering at Anthropic. Dem ystifying ev als for AI agents, 2026. h ttps://ww w.anthropic.com/engineering/de m ystif ying - ev als- f or- ai- agents . [31] Y uyang W u, Yifei W ang, Ziyu Y e, Tianqi Du, Stefanie Jegelk a, and Yisen W ang. When More is Less: Understanding Chain-of-Though t Length in LLMs. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2502.07266 . [32] Zheng Du, Hao Kang, Song Han, T ushar Krishna, and Ligeng Zhu. OckBenc h: Measuring the Efficiency of LLM Reasoning. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2511.05722 . [33] Artificial Analysis. Artificial Analysis Intelligence Index, 2026. h ttps://artificialanalysis.ai/ev aluations/artif icia l- analysis- in telligence- index . [34] Mark Chen, Jerry T worek, Heewoo Jun, Qiming Y uan, Henrique Ponde de Oliveira Pinto, Jared Kaplan, Harri Edw ards, Y uri Burda, Nic holas Joseph, Greg Bro c kman, Alex Ra y , Raul Puri, Gretchen Krueger, Michael P etrov, Heidy Khlaaf, Girish Sastry , Pamela Mishkin, Bro ok e Chan, Scott Gray , Nick Ryder, Mikhail Pa vlov, Alethea Po wer, Luk asz Kaiser, Mohammad Bav arian, Clemens Winter, Philipp e Tillet, F elip e Petroski Such, Da ve Cummings, Matthias Plapp ert, F otios Chantzis, Elizab eth Barnes, Ariel Herb ert-V oss, William Hebgen Guss, Alex Nichol, Alex Paino, Nikolas T ezak, Jie T ang, Igor Babuschkin, Suchir Bala ji, Shantan u Jain, William Saunders, Christopher Hesse, Andrew N. Carr, Jan Leike, Josh Ac hiam, V edant Misra, Ev an Morik aw a, Alec Radford, Matthew Knight, Miles Brundage, Mira Murati, Katie May er, Peter W elinder, Bob McGrew, Dario Amo dei, Sam McCandlish, Ilya Sutskev er, and W o jciech Zaremba. Ev aluating large language mo dels trained on co de. arXiv Pr eprint , 2021. doi:10.48550/arxiv.2107.03374 . [35] Bernardo de Souza. GOA T: A Global Optimization Algorithm for Molecules and Atomic Clusters. Angew. Chem. Int. Ed. , 64(18):e202500393, 2025. doi:10.1002/anie.202500393 . [36] Philipp Prach t, F abian Bohle, and Stefan Grimme. Automated Exploration of the Low-Energy Chemical Space with F ast Quantum Chemical Metho ds. Phys. Chem. Chem. Phys. , 22(14):7169–7192, 2020. doi:10.1039/c9cp06869d . [37] Philipp Prac ht, Stefan Grimme, Christoph Bannw arth, F abian Bohle, Sebastian Ehlert, Gereon F eldmann, Johannes Gorges, Marcel Müller, Tim Neudeck er, Christoph Plett, Sebastian Spicher, Pit Steinbac h, Patryk A. W esołowski, and F elix Zeller. CREST – A program for the exploration of low-energy molecular chemical space. J. Chem. Phys. , 160(11):114110, 2024. doi:10.1063/5.0197592 . [38] RDKit Contributors. RDKit: Open-source cheminformatics. Zeno do, 2026. 10.5281/zenodo.591637 . [39] Christoph Bannw arth, Sebastian Ehlert, and Stefan Grimme. GFN2-xTB - An Accurate and Broadly P arametrized Self-Consisten t Tight-Binding Quantum Chemical Metho d with Multip ole Electrostatics and Density-Dependent Disp ersion Contributions. J. Chem. Theory Comput. , 15(3):1652–1671, 2019. doi:10.1021/acs.jctc.8b01176 . [40] Sebastian Ehlert, Marcel Stahn, Sebastian Spicher, and Stefan Grimme. Robust and Efficient Implicit Solv ation Mo del for F ast Semiempirical Methods. J. Chem. Theory Comput. , 17(7):4250–4261, 2021. doi:10.1021/acs.jctc.1c00471 . [41] Narb e Mardirossian and Martin Head-Gordon. ω b97X-V: A 10-parameter, range-separated hybrid, generalized gradien t approximation density functional with nonlo cal correlation, designed by a surviv al-of-the-fittest strategy . Phys. Chem. Chem. Phys. , 16(21):9904–9924, 2014. doi:10.1039/c3cp54374a . [42] Eik e Caldeweyher, Sebastian Ehlert, Andreas Hansen, Hagen Neugebauer, Sebastian Spicher, Christoph Bannw arth, and Stefan Grimme. A generally applicable atomic-charge dep enden t London disp ersion correction. J. Chem. Phys. , 150(15):154122, 2019. doi:10.1063/1.5090222 . [43] Florian W eigend and Reinhart Ahlrichs. Balanced basis sets of split v alence, triple zeta v alence and quadruple zeta v alence qualit y for H to Rn: Design and assessment of accuracy . Phys. Chem. Chem. Phys. , 7(18):3297–3305, 2005. doi:10.1039/b508541a . 18 [44] Aleksandr V. Marenich, Christopher J. Cramer, and Donald G. T ruhlar. Univ ersal Solv ation Mo del Based on Solute Electron Density and on a Contin uum Mo del of the Solv ent Defined by the Bulk Dielectric Constant and A tomic Surface T ensions. J. Phys. Chem. B , 113(18):6378–6396, 2009. doi:10.1021/jp810292n . [45] Ian T. Beck, Erica C. Mitchell, Annab elle W ebb Hill, Justin M. T urney , Brandon Rotav era, and Henry F. I I I Sc haefer. Ev aluating the Imp ortance of Conformers for Understanding the V acuum-Ultraviolet Sp ectra of Oxiranes: Exp erimen t and Theory. J. Phys. Chem. A , 128(50):10906–10920, 2024. doi:10.1021/acs.jpca.4c04391 . [46] Sebastian Spicher, Christoph Plett, Philipp Prach t, Andreas Hansen, and Stefan Grimme. Automated molecular cluster growing for explicit solv ation by efficient force field and tight binding metho ds. J. Chem. The ory Comput. , 18(5):3174–3189, 2022. doi:10.1021/acs.jctc.2c00239 . [47] P eter G. Boyd and T om K. W o o. A generalized metho d for constructing hypothetical nanop orous materials of an y net top ology from graph theory . CrystEngComm , 18(21):3777–3792, 2016. doi:10.1039/c6ce00407e . [48] Aleksandar Kondinski, Angiras Menon, Daniel Nurko wski, F eroz F arazi, Sebastian Mosbach, Jethro Akroyd, and Markus Kraft. Automated Rational Design of Metal–Organic Polyhedra. J. Am. Chem. So c. , 144(26):11713–11728, 2022. doi:10.1021/jacs.2c03402 . [49] Jiy eon Kim, Dongsik Nam, Hye Jin Cho, Eunchan Cho, Dharmalingam Siv anesan, Changhy eon Cho, Jaewoong Lee, Jihan Kim, and W ony oung Cho e. Up–down approach for expanding the chemical space of metal–organic framew orks. Nat. Synth. , 3(12):1518–1528, 2024. doi:10.1038/s44160-024-00638-x . [50] Hao Lyu, Z he Ji, Stefan W uttke, and Omar M. Y aghi. Digital reticular chemistry . Chem , 6(9):2219–2241, 2020. doi:10.1016/j.c hempr.2020.08.008 . [51] Zhiling Zheng, Nakul Rampal, Theo Jaffrelot Inizan, Christian Borgs, Jennifer T. Chay es, and Omar M. Y aghi. Large language mo dels for reticular chemistry. Nat. R ev. Mater. , 10(5):369–381, 2025. doi:10.1038/s41578-025- 00772-8 . [52] Guobin Zhao, Logan M. Brabson, Saumil Chheda, Ju Huang, Haewon Kim, Kunhuan Liu, Kenji Mo chida, Thang D. Pham, Prerna, Gianmarco G. T errones, Sunghyun Y o on, Lionel Zoubritzky , F rançois-Xavier Coud- ert, Maciej Haranczyk, Heather J. Kulik, Sey ed Mohamad Mo osavi, David S. Sholl, J. Ilja Siepmann, Ran- dall.Q. Snurr, and Y ongch ul G. Chung. CoRE MOF DB: A curated exp erimen tal metal-organic framework database with machine-learned prop erties for integrated material-pro cess screening. Matter , 8(6):102140, 2025. doi:10.1016/j.matt.2025.102140 . [53] Marco Gibaldi, Anna Kap eliukha, Andrew White, Jun Luo, Rob ert Alex May o, Jak e Burner, and T om K. W o o. Mosaec-db: a comprehensiv e database of exp erimental metal–organic frameworks with verified chemical accuracy suitable for molecular simulations. Chem. Sci. , 16(9):4085–4100, 2025. doi:10.1039/d4sc07438f . [54] Benjamin J. Bucior, Andrew S. Rosen, Maciej Haranczyk, Zhenp eng Y ao, Michael E. Zieb el, Omar K. F arha, Joseph T. Hupp, J. Ilja Siepmann, Alán Aspuru-Guzik, and Randall Q. Sn urr. Iden tification Sc hemes for Metal–Organic F rameworks T o Enable Rapid Search and Cheminformatics Analysis. Cryst. Gr owth Des. , 19(11): 6682–6697, 2019. doi:10.1021/acs.cgd.9b01050 . [55] Zhenp eng Y ao, Benjamín Sánchez-Lengeling, N. Scott Bobbitt, Benjamin J. Bucior, Sai Govind Hari Kumar, Sean P . Collins, Thomas Burns, T om K. W o o, Omar K. F arha, Randall Q. Sn urr, and Alán Aspuru-Guzik. Inv erse design of nanop orous crystalline reticular materials with deep generative mo dels. Nat. Mach. Intel l. , 3(1):76–86, 2021. doi:10.1038/s42256-020-00271-1 . [56] Sangw on Lee, Baekjun Kim, Hyun Cho, Hooseung Lee, Sarah Y unmi Lee, Eun Seon Cho, and Jihan Kim. Computational Screening of T rillions of Metal–Organic F rameworks for High-P erformance Methane Storage. ACS Appl. Mater. Interfac es , 13(20):23647–23654, 2021. doi:10.1021/acsami.1c02471 . [57] Matthew A. Addicoat, Nina V anko v a, Ismot F arjana Akter, and Thomas Heine. Extension of the Universal F orce Field to Metal–Organic F rameworks. J. Chem. The ory Comput. , 10(2):880–891, 2014. doi:10.1021/ct400952t . [58] Ily es Batatia, Da vid P K ov acs, Gregor Simm, Christoph Ortner, and Gáb or Csányi. MA CE: Higher order equiv ariant message passing neural netw orks for fast and accurate force fields. A dv. Neur al Inf. Pr o c ess. Syst. , 35: 11423–11436, 2022. doi:10.48550/arxiv.2206.07697 . [59] Hendrik Kraß, Ju Huang, and Seyed Mohamad Mo osa vi. MOFSimBenc h: ev aluating universal machine learn- ing interatomic p oten tials in metal-organic framework molecular mo deling. npj Comput. Mater. , 12(1), 2025. doi:10.1038/s41524-025-01872-3 . 19 [60] P eyman Z. Moghadam, Aurelia Li, Seth B. Wiggin, Andi T ao, Andrew G. P . Maloney , Peter A. W o od, Suzanna C. W ard, and David F airen-Jimenez. Developmen t of a cambridge structural database subset: A collection of metal–organic frameworks for past, present, and future. Chem. Mater. , 29(7):2618–2625, 2017. doi:10.1021/acs.c hemmater.7b00441 . [61] Thomas F. Willems, Chris H. Rycroft, Michaeel Kazi, Juan C. Meza, and Maciej Haranczyk. Algorithms and to ols for high-throughput geometry-based analysis of crystalline p orous materials. Micr op or. Mesop or. Mat. , 149 (1):134–141, 2012. doi:10.1016/j.micromeso.2011.08.020 . [62] Saffron Huang, Bryan Seethor, Esin Durmus, Kunal Handa, Miles McCain, Michael Stern, and Deep Ganguli. Ho w AI Is T ransforming W ork at Anthropic, 2025. h ttps://anthropic.com/research/ho w- ai- is- transforming- w or k- at- anthropic/ . [63] Gary T om, Stefan P . Sc hmid, Sterling G. Baird, Y ang Cao, K ourosh Darvish, Han Hao, Stanley Lo, Sergio P ablo-García, Ella M. Ra jaonson, Marta Skreta, Naruki Y oshik aw a, Samantha Corapi, Gun Deniz Akkoc, F elix Strieth-Kalthoff, Martin Seifrid, and Alán Aspuru-Guzik. Self-Driving Lab oratories for Chemistry and Materials Science. Chem. R ev. , 124(16):9633–9732, 2024. doi:10.1021/acs.chemrev.4c00055 . [64] Shi Xuan Leong, Caleb E Griesbach, Rui Zhang, Kourosh Darvish, Y uchi Zhao, Abhijoy Mandal, Y unheng Zou, Han Hao, V arinia Bernales, and Alán Aspuru-Guzik. Steering tow ards safe self-driving lab oratories. Nat. R ev. Chem. , 9(10):707–722, 2025. doi:10.1038/s41570-025-00747-x . [65] F elix Strieth-Kalthoff, Han Hao, V andana Rathore, Joshua Derasp, Théophile Gaudin, Nicholas H. Angello, Martin Seifrid, Ek aterina T rushina, Mason Guy , Junliang Liu, Xun T ang, Masashi Mamada, W esley W ang, T uul T sagaantsoo j, Cyrille Lavigne, Rob ert Pollice, T ony C. W u, Kazuhiro Hotta, Leticia Bo do, Shangyu Li, Mohammad Haddadnia, Agnieszk a W ołos, Rafał Roszak, Cher Tian Ser, Carlota Bozal-Ginesta, Riley J. Hickman, Jeny a V estfrid, Andrés Aguilar-Granda, Elena L. Klimarev a, Ralph C. Sigerson, W enduan Hou, Daniel Gahler, Slaw omir Lac h, Adrian W arzyb ok, Oleg Boro din, Simon Rohrbach, Benjamin Sanchez-Lengeling, Chihay a Adac hi, Bartosz A. Grzyb o wski, Leroy Cronin, Jason E. Hein, Martin D. Burke, and Alán Aspuru-Guzik. Delo calized, asynchronous, closed-lo op discov ery of organic laser emitters. Scienc e , 384(6697), 2024. doi:10.1126/science.adk9227 . [66] Jiaru Bai, Sebastian Mosbach, Connor J. T aylor, Dogancan Karan, Kok F o ong Lee, Simon D. Rihm, Jethro Akro yd, Alexei A. Lapkin, and Markus Kraft. A Dynamic Knowledge Graph Approach to Distributed Self-Driving Lab oratories. Nat. Commun. , 15:462, 2024. doi:10.1038/s41467-023-44599-9 . [67] Alexus A. Smith, Edmund L. W ong, Ronan C. Donov an, Brad A. Chapman, Ryan Harry , Pooy an Tirandazi, P aulina Kanigowsk a, Elizab eth A. Gendreau, Rob ert H. Dahl, Michal Jastrzebski, Jose E. Cortez, Christopher J. Bremner, José C. Morales Hemuda, James Do oner, Ian Grav es, Rahul Karandik ar, Christopher Lionetti, Kevin Christopher, Andrew L. Consiglio, Alyssa T ran, William McCusker, Duy X. Nguyen, Isis Botelho Nunes da Silv a, Alv aro R. Bautista-A y ala, Monica P . McNerney , Sean Atkins, Michael McDuffie, Will Serb er, Bradley P . Barb er, T rinh Thanongsinh, Andrew Nesson, Bib ek Lama, Brandon Nichols, Cameron LaF rance, T enzing Nyima, Alicia Byrn, Rashard Thornhill, Bryan Cai, Lizvette A yala-V aldez, Alycia W ong, Austin J. Che, W alter Thav ara jah, Daniel Smith, Jr. Thomas F. Knight, Da vid W. Borhani, Jerry T worek, Mostafa Rohaninejad, Ahmed El-Kishky , Nathan C. T edford, T ejal Pat wardhan, Y unxin Joy Jiao, and Reshma P . Shetty . Using a gpt-5-driven autonomous lab to optimize the cost and titer of cell-free protein syn thesis, 2026. h ttps://cdn.openai.com/pdf /5a12a3bc- 9 6b7- 4e07- 9386- db6ee5bb2ed9/u sing- a- g pt- 5- d riv en- autono mous- lab- to- optimize- the- cost- and- titer- of- cell- f re e- protein- synthesis.pdf . [68] Human Lay er. 12-F actor Agents - Principles for building reliable LLM applications, 2025. h ttps://github.com/h umanlay er/12- f actor- agen ts . [69] V adim Borisov, Michael Gröger, Mina Mikhael, and Richard H. Schreiber. Do Chatb ot LLMs T alk T o o Much? The Y apBench Benchmark. arXiv Pr eprint , 2026. doi:10.48550/arxiv.2601.00624 . [70] Zijian Zhang, Aiwei Yin, Amaan Baw eja, Jiaru Bai, Ignacio Gustin, V arinia Bernales, and Alán Aspuru-Guzik. El Agen te F orjador: T ask-driven Agent Generation for Quantum Simulation. Manuscript in preparation, 2026. [71] Xu Huang, Junwu Chen, Y uxing F ei, Zhuohan Li, Philipp e Sch waller, and Gerbrand Ceder. Cascade: Cum ulative agen tic skill creation through autonomous dev elopment and ev olution. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2512.23880 . [72] Alibaba Op en Source. A universal sandb o x platform for AI application scenarios, providing multi-language SDKs, unified sandb ox proto cols, and sandb ox runtimes for LLM-related capabilities., 2025. h ttps://github.com/aliba ba/Op enSandbo x . 20 [73] Miles W ang, Joy Jiao, Neil Chowdh ury , Ethan Chang, and T ejal Pat w ardhan. F rontierScience: Ev aluating AI’s Abilit y to Perform Exp ert-Level Scientific T asks, 2025. h ttps://cdn.openai.com/pdf /2fcd284c- b468- 4c21- 8ee0- 7 a783933efcc/frontierscience- paper.pdf . [74] Zhangde Song, Jieyu Lu, Y uanqi Du, Botao Y u, Thomas M. Pruyn, Y ue Huang, Kehan Guo, Xiuzhe Luo, Y uanhao Qu, Yi Qu, Yink ai W ang, Haorui W ang, Jeff Guo, Jingru Gan, Parshin Sho jaee, Di Luo, Andres M Bran, Gen Li, Qiyuan Zhao, Shao-Xiong Lennon Luo, Y uxuan Zhang, Xiang Zou, W anru Zhao, Yifan F. Zhang, W ucheng Zhang, Sh unan Zheng, Saiyang Zhang, Sartaa j T akrim Khan, Mahy ar Ra jabi-Kochi, Samantha Paradi-Maropakis, T ony Baltoiu, F engyu Xie, Tiany ang Chen, Kexin Huang, W eiliang Luo, Meijing F ang, Xin Y ang, Lixue Cheng, Jia jun He, Soha Hassoun, Xiangliang Zhang, W ei W ang, Chandan K. Reddy , Chao Zhang, Zhiling Zheng, Mengdi W ang, Le Cong, Carla P . Gomes, Chang-Y u Hsieh, Adit ya Nandy , Philipp e Sch waller, Heather J. Kulik, Hao jun Jia, Huan Sun, Seyed Mohamad Mo osavi, and Chenru Duan. Ev aluating Large Language Mo dels in Scientific Disco very. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2512.15567 . [75] Nenad Krdzav ac, Sebastian Mosbach, Daniel Nurko wski, Philipp Buerger, Jethro Akroyd, Jacob Martin, Angiras Menon, and Markus Kraft. An Ontology and Seman tic W eb Service for Quantum Chemistry Calculations. J. Chem. Inf. Mo del. , 59(7):3154–3165, 2019. doi:10.1021/acs.jcim.9b00227 . [76] Laura Pascazio, Simon Rihm, Ali Naseri, Sebastian Mosbach, Jethro Akroyd, and Markus Kraft. Chemical Sp ecies Ontology for Data Integration and Knowledge Discov ery. J. Chem. Inf. Mo del. , 63(21):6569–6586, 2023. doi:10.1021/acs.jcim.3c00820 . [77] Xiao c hi Zhou, Patric k Bulter, Changxuan Y ang, Simon D. Rihm, Thitik arn Angk anaporn, Jethro Akro yd, Sebastian Mosbac h, and Markus Kraft. Ontology-to-tools compilation for executable semantic constrain t enforcement in LLM agen ts. arXiv Pr eprint , 2026. doi:10.48550/arxiv.2602.03439 . [78] Stefan Grimme, Jens An tony , Stephan Ehrlich, and Helge Krieg. A consisten t and accurate ab initio parametrization of density functional disp ersion correction (DFT-D) for the 94 elements H-Pu. J. Chem. Phys. , 132(15):154104, 2010. doi:10.1063/1.3382344 . [79] Stefan Grimme, Stephan Ehrlich, and Lars Go erigk. Effect of the damping function in disp ersion corrected densit y functional theory . J. Comput. Chem. , 32(7):1456–1465, 2011. doi:10.1002/jcc.21759 . [80] Eik e Caldeweyher, Christoph Bannw arth, and Stefan Grimme. Extension of the D3 disp ersion co efficien t mo del. J. Chem. Phys. , 147(3):034112, 2017. doi:10.1063/1.4993215 . [81] Luk as Wittmann, Igor Gordiy , Marvin F riede, Benjamin Helmich-P aris, Stefan Grimme, Andreas Hansen, and Markus Bursch. Extension of the D3 and D4 London disp ersion corrections to the full actinides series. Phys. Chem. Chem. Phys. , 26(32):21379–21394, 2024. doi:10.1039/D4CP01514B . [82] Sungh wan Kim, Paul A Thiessen, Ev an E Bolton, Jie Chen, Gang F u, Asta Gindulyte, Lianyi Han, Jane He, Siqian He, Benjamin A Sho emaker, Jiy ao W ang, Bo Y u, Jian Zhang, and Stephen H Bryan t. PubChem Substance and Comp ound Databases. Nucleic A cids R es. , 44(D1):D1202–D1213, 2016. doi:10.1093/nar/gkv951 . [83] Matthew Swain. PubChemPy, 2017. h ttps://pub chempy .org . [84] Greg Landrum, Paolo T osco, Brian Kelley , Ricardo Ro driguez, et al. rdkit/rdkit: 2025_03_1 (q1 2025) release, Marc h 2025. h ttps://doi.org . [85] Noel M O’Boyle, Michael Banck, Craig A James, Chris Morley , Tim V andermeersch, and Geoffrey R Hutchison. Op en Bab el: An Op en Chemical T o olbox. J. Cheminf. , 3:33, 2011. doi:10.1186/1758-2946-3-33 . [86] Jinzhe Zeng. op en bab el-wheel, 2026. h ttps://github.com/njzjz/op en babel- w heel . Accessed 9 F eb 2026. [87] Stefan Grimme. Exploration of Chemical Comp ound, Conformer, and Reaction Space with Meta-Dynamics Sim ulations Based on Tight-Binding Quantum Chemical Calculations. J. Chem. Theory Comput. , 15(5):2847–2862, 2019. doi:10.1021/acs.jctc.9b00143 . [88] Guobin Zhao, P engyu Zhao, and Y ongch ul G. Chung. Mofclassifier: A machine learning approach for v alidating computation-ready metal–organic framew orks. J. A m. Chem. So c. , 147(37):33343–33349, 2025. doi:10.1021/jacs.5c10126 . http://dx.doi.org/10.1021/jacs.5c10126 . [89] Sh yue Ping Ong, William Davidson Richards, Anubha v Jain, Geoffroy Hautier, Michael Kocher, Shreyas Cholia, Dan Gunter, Vincent L Chevrier, Kristin A Persson, and Gerbrand Ceder. Python Materials Genomics (pymat- gen): A Robust, Op en-Source Python Library for Materials Analysis. Comput. Mater. Sci. , 68:314–319, 2013. doi:10.1016/j.commatsci.2012.10.028 . 21 [90] Lionel Zoubritzky and F rançois-Xavier Coudert. CrystalNets.jl: Iden tification of Crystal T op ologies. SciPost Chem. , 1:005, 2022. doi:10.21468/SciPostChem.1.2.005 . [91] Han Y ang, Chenxi Hu, Yichi Zhou, Xixian Liu, Y u Shi, Jielan Li, Guanzhi Li, Zekun Chen, Shuizhou Chen, Claudio Zeni, et al. MatterSim: A Deep Learning Atomistic Mo del Across Elements, T emp eratures and Pressures. arXiv Preprint , 2024. doi:10.48550/arxiv.2405.04967 . [92] Benjamin Rho des, Sander V andenhaute, V aidotas Šimkus, James Gin, Jonathan Go dwin, Tim Duignan, and Mark Neumann. Orb-v3: atomistic simulation at scale. arXiv Pr eprint , 2025. doi:10.48550/arxiv.2504.06231 . [93] Ask Hjorth Larsen, Jens Jørgen Mortensen, Jakob Blomqvist, Iv ano E Castelli, Rune Christensen, Marcin Dułak, Jesp er F riis, Michael N Grov es, Bjørk Hammer, Cory Hargus, Eric D Hermes, P aul C Jennings, Peter Bjerre Jensen, James Kermo de, John R Kitchin, Esb en Leonhard Kolsb jerg, Joseph Kubal, Kristen Kaasb jerg, Steen Lysgaard, Jón Bergmann Maronsson, T ristan Maxson, Thomas Olsen, Lars Pastewk a, Andrew Peterson, Carsten Rostgaard, Jak ob Schiøtz, Ole Sch ütt, Mikkel Strange, Kristian S Thygesen, T ejs V egge, Lasse Vilhelmsen, Michael W alter, Zhenh ua Zeng, and Karsten W Jacobsen. The atomic simulation environmen t—a Python library for working with atoms. J. Phys. Condens. Matter. , 29(27):273002, 2017. ISSN 1361-648X. doi:10.1088/1361-648x/aa680e . [94] Stefan Grimme, F abian Bohle, Andreas Hansen, Philipp Prach t, Sebastian Spicher, and Marcel Stahn. Efficien t Quan tum Chemical Calculation of Structure Ensembles and F ree Energies for Nonrigid Molecules. J. Phys. Chem. A , 125(19):4039–4054, 2021. doi:10.1021/acs.jp ca.1c00971 . 22 Supporting Information Contents A List of to ols 24 B Universit y-lev el quantum chemistry b enc hmark 30 B.1 LLM mo del sp ecifications . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 30 B.2 User prompts . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 31 B.3 Rubrics for LLM judge . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 34 B.4 P ass k analysis . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 36 B.5 Statistics plots . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 37 B.6 Bare LLM agent . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 46 C Use case extension 1: Boltzmann-weigh ted sp ectroscopic prop erties 48 C.1 Boltzmann-w eigh ted sp ectroscopic prop erties in implicit solutions . . . . . . . . . . . . . . . . 49 C.2 Boltzmann-w eigh ted sp ectroscopic prop erties with explicit solv ation . . . . . . . . . . . . . . 50 D Use case extension 2: Exploration of metal-organic framew orks design space 50 D.1 MOF execution graph no de implemen tations . . . . . . . . . . . . . . . . . . . . . . . . . . . 50 D.2 Chat transcripts . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 58 23 A List of tools T ool Name T ool description received b y agent/LLMs run_mof_w orkflow Run the MOF wor kflow graph for building, optimizing, and/or analyzing MOF s. This w orkflow uses dynamic AI-based routing to chain together multiple stages based on the user’s inten t. The workflo w is NOT limited to a single op eration— it can automatically pro ceed through multiple stages (e.g., download -> decom- p ose -> build -> optimize -> analyze). W ORKFLOW ST AGES (can b e combined automatically): • Do wnload: F etch CIF from CSD/CoRE-MOF database • Decomp ose: Extract top ology and building blo cks from CIF • Searc h: Query knowledge graph for compatible MOF combinations • Build: Construct MOF structure from comp onen ts • Optimize: Geometry relaxation via MLFF (MACE-MOF, etc.) • Analyze: Zeo++ p orosit y analysis SAMPLE WORKFLO WS: Build directly from PORMAKE comp onents Required: – top ology_pormake_id (e.g., "p cu", "dia) – no de_pormake_id (e.g., "N409", "N111") – link er_p ormak e_id (e.g., "E1", "E42") Optional: – mof_name Example user in tents: – "Build a MOF with top ology Z using metal no de X and organic linker Y." – "Construct this sp ecific MOF (I know the building blo c ks)." CSD/CoRE-MOF Refco de -> Download, then automatically routed based on user inten t (summarised_user_query) Required: – csd_refco de Optional: – mof_name Example user in tents: – "What is the top ology of MOF ABA VIJ?" – "What is the surface area of this CSD structure?" – "Optimize the MOF with refco de ABA VIJ." Pro cess a lo cal CIF file Required: – input_cif_filename Optional: – mof_name T ypical user inten ts: – "Here is a CIF. What’s its top ology?" – "Optimize this MOF." – "Run Zeo++ analysis on this structure." Kno wledge Graph Search -> Find compatible com binations, then optionally build Optional: – top ology_iris : list of top ology instance IRIs to restrict – metal_no de_iris : list of metal-no de instance IRIs to restrict – organic_link er_iris : list of linker instance IRIs to restrict Parameters schema 24 {"additionalProp erties":false,"properties":{"summarised_user_query":{"type":"string"}," top ology_pormake_id":{"default":n ull,"an yOf":[{"type":"string"},{"type":"null"}]}," no de_pormake_id":{"default":n ull,"an yOf":[{"type":"string"},{"type":"null"}]}," link er_p ormak e_id":{"default":null,"an yOf":[{"t yp e":"string"},{"t yp e":"n ull"}]},"mof_name ":{"default":n ull,"anyOf":[{"t ype":"string"},{"type":"null"}]},"csd_refcode":{"default":null," an yOf":[{"type":"string"},{"type":"null"}]},"input_cif_filename":{"default":n ull,"anyOf":[{" t yp e":"string"},{"t yp e":"n ull"}]},"top ology_iris":{"default":n ull,"anyOf":[{"items":{"t ype":" string"},"t yp e":"arra y"},{"type":"null"}]},"metal_node_iris":{"default":null,"an yOf":[{"items ":{"t yp e":"string"},"t yp e":"arra y"},{"type":"null"}]},"organic_link er_iris":{"default":n ull," an yOf":[{"items":{"type":"string"},"type":"array"},{"t ype":"null"}]},"update_graph":{" default":false,"t yp e":"b oolean"}},"required":["summarised_user_query"],"type":"ob ject"} T ool Name T ool description received b y agent/LLMs duc kduckgo_w eb_- searc h Searc hes Duc kDuckGo for the giv en query and returns the re- sults. The searc h results. Parameters schema {"additionalProp erties":false,"properties":{"query":{"description":"The query t o searc h for.","t yp e ":"string"}},"required":["query"],"t yp e":"ob ject"} T ool Name T ool description received b y agent/LLMs get_con version_fac- tor Con vert ph ysical quan tities from a list of UnitCon vPair each con taining a n umerical v alue along with source and target units, and the function returns a list of conv erted v alues. list[float] Conv erted v alues in the target unit. str: Error message if conv ersion is not supp orted or input is inv alid. Parameters schema {"additionalProp erties":false,"properties":{"UnitConv ersionP airs":{"description":"List of UnitCon vPair ob jects containing v alue, from_unit, and to_unit.","items":{"$ref":"#/$defs/ UnitCon vPair"},"t ype":"array"}},"required":["UnitCon v ersionPairs"],"t yp e":"ob ject","$defs ":{"UnitCon vPair":{"properties":{"v alue":{"t yp e":"n um b er"},"from_unit":{"en um":["hartree ","e V","kJ/mol","kcal/mol","cm^-1","Hz"],"t ype":"string"},"to_unit":{"enum":["hartree","e V ","kJ/mol","k cal/mol","cm^-1","Hz"],"type":"string"}},"required":["v alue","from_unit"," to_unit"],"t yp e":"ob ject","additionalProp erties":false}}} T ool Name T ool description received b y agent/LLMs capture_view er_- screenshot Capture a screenshot of the current Molecule Viewer and provide it to the mo del. List of str with BinaryCon- ten t image included in the conten t so the LLM can analyze it. Parameters schema 25 {"additionalProp erties":false,"properties":{"molecule_id":{"default":null,"description":"Optional molecule id to target; defaults to \"current\" if omitted.","anyOf":[{"t yp e":"string"},{"t yp e":" n ull"}]}},"type":"ob ject"} T ool Name T ool description received b y agent/LLMs run_crest_con- former_searc h Run a CREST conformer search for a giv en molecule iden ti- fier. A JSON-serializable dictionary describing the CREST ensemble. Parameters schema {"additionalProp erties":false,"properties":{"identifier":{"description":"The molecule identifier ( SMILES string, name, or XYZ file path).","type":"string"},"identifier_t yp e":{"description":" The type of the identifier ( ' smiles ' , ' name ' , or ' xyz ' ).","enum":["smiles","name","xyz"],"t yp e":" string"},"c harge":{"default":0,"description":"The molecular charge (default: 0).","type":" n umber"},"spin_multiplicit y":{"default":1,"description":"The spin multiplicit y (default: 1)."," t yp e":"in teger"},"implicit_solven t":{"default":n ull,"description":"The implicit solv ent mo del to use (default: None), example: \"w ater\", \"acetonitrile\".","an yOf":[{"type":"string"},{" t yp e":"n ull"}]},"calculation_level_method":{"default":"gfn2","description":"The metho d to use for the CREST calculation (default: ' gfn2 ' ).","enum":["tblite","gfn2","gfn1","gfnff","gfn0 ","gfn0*","xtb","gfn","gfn-xtb","orca","generic"],"t yp e":"string"},"crest_run type":{"default ":"im td-gc","description":"CREST sampling mo de ( ' imtd-gc ' for standard sampling, ' nci ' for non-co v alen t complexes and aggregates sampling whic h should b e used for explicitly solv ated structures).","en um":["imtd-gc","nci"],"t ype":"string"},"run_on_gpu":{"default":false," description":"Whether to run the calculation on GPU (default: F alse).","type":"b o olean"}}," required":["iden tifier","identifier_t ype"],"type":"ob ject"} T ool Name T ool description received b y agent/LLMs run_qcg_cluster Run a QCG grow calculation and return the merged cluster as XYZ. Parameters schema {"additionalProp erties":false,"properties":{"solute_identifier":{"description":"Solute identifier ( SMILES, name, or XYZ string).","type":"string"},"solven t_iden tifier":{"description":"Solv ent iden tifier (SMILES, name, or XYZ string).","type":"string"},"solute_identifier_t ype":{" description":"T yp e of solute identifier.","en um":["smiles","name","xyz"],"t yp e":"string"}," solv ent_iden tifier_t yp e":{"description":"T yp e of solven t identifier.","en um":["smiles","name ","xyz"],"t yp e":"string"},"c harge":{"default":0,"description":"T otal molecular c harge.","type ":"n umber"},"spin_multiplicit y":{"default":1,"description":"Spin multiplicit y (e.g., 1 for singlet, 2 for doublet).","minimum":1,"t yp e":"in teger"},"nsolv":{"default":15,"description":" Num b er of solven t molecules to add.","exclusiveMinim um":0,"t yp e":"in teger"},"threads":{" default":12,"description":"Threads passed to CREST via --T.","exclusiv eMinimum":0,"t ype":" in teger"},"alpb_solven t":{"default":"w ater","description":"ALPB solven t name (case- insensitiv e). A v ailable: acetone, acetonitrile, aniline, b enzaldeh yde, b enzene, c h2cl2, c hcl3, cs2, dio xane, dmf, dmso, ethanol, ether, ethylacetate, furane, hexadecane, hexane, methanol, nitromethane, o ctanol, o ctanol (w et), phenol, toluene, thf, water","an yOf":[{"t yp e":"string "},{"t yp e":"n ull"}]}},"required":["solute_identifier","solv en t_iden tifier"," solute_iden tifier_type","solven t_iden tifier_t yp e"],"t yp e":"ob ject"} 26 T ool Name T ool description received b y agent/LLMs sho w_molecule_in_- view er Instruct the Molecule Viewer to display a ConceptualAtoms b y its IRI. A confirmation message indicating the viewer has b een up dated. Parameters schema {"additionalProp erties":false,"properties":{"conceptual_atoms_iri":{"description":"The IRI of the ConceptualA toms instance to displa y.","type":"string"}},"required":["conceptual_atoms_iri "],"t yp e":"ob ject"} T ool Name T ool description received b y agent/LLMs run_p yscf_workflo w Run PySCF calculations using the workflo w graph with dynamic routing capabilities. list[PyscfOutput] PySCF calculation outputs gathered from the w orkflo w run. Parameters schema {"additionalProp erties":false,"properties":{"summerised_user_query":{"description":"Description of the calculations the user w ants to run, this will b e used b y the workflo w routing agen t to determine which next no des to execute.","type":"string"},"identifier_t yp e":{"description":" Kind of identifier supplied to describ e the molecule.","enum":["name","smiles","xyz"," xyz_filename","conceptual_atoms_iri"],"t yp e":"string"},"iden tifier":{"description":"Identifier con tent (name, SMILES, or XYZ string) describing the conceptual atom; for XYZ, provide the literal file text such as `` 3\\nCOMMENT\\nO 0.000 0.000 0.000\\nH 0.000 0.757 0.586\\nH 0.000 -0.757 0.586 `` .","type":"string"},"charge":{"default":0,"description":"Ov erall molecular c harge; defaults to 0.","t yp e":"in teger"},"spin_multiplicit y":{"default":1,"description":"Spin m ultiplicity of the molecule; defaults to 1.","type":"integer"},"basis_set":{"default":n ull," description":"Basis set name for the calculation; falls back to workflo w defaults when omitted .","an yOf":[{"type":"string"},{"type":"null"}]},"restricted":{"default":true,"description":" Whether to use a restricted reference (RHF/ROHF/RDFT) instead of unrestricted; defaults to T rue.","type":"b o olean"},"xc_functional":{"default":n ull,"description":"Exc hange-correlation functional to activ ate DFT; when omitted, the workflo w uses Hartree-F o ck.","an yOf":[{"t yp e":" string"},{"t yp e":"n ull"}]},"solv ation_model":{"default":null,"description":"Implicit solv ation mo del lab el; uses v acuum when not provided.","an yOf":[{"en um":["CPCM","COSMO","IEF- PCM","SS(V)PE","SMD"],"t yp e":"string"},{"t yp e":"n ull"}]},"implicit_solven t":{"default": n ull,"description":"Solven t environmen t name for the chosen solv ation mo del; defaults to w ater when paired with an implicit mo del.","an yOf":[{"en um":["W ater","Dimethylsulfo xide"," Nitromethane","A cetonitrile","Methanol","Ethanol","Acetone","Meth ylenec hloride"," T etrahydrofurane","Aniline","Chlorobenzene","Chloroform","T oluene","Benzene"," Cyclohexane","N-heptane"],"t yp e":"string"},{"t yp e":"n ull"}]},"exit_no de":{"default":n ull," description":"Iden tifier of a w orkflow no de to terminate at, enabling partial runs.","anyOf":[{" t yp e":"string"},{"t yp e":"n ull"}]}},"required":["summerised_user_query","identifier_t ype"," iden tifier"],"type":"ob ject"} 27 T ool Name T ool description received b y agent/LLMs run_sparql_query Execute a read-only SP AR QL query against the knowledge graph. Supp orts SELECT, CONSTRUCT, ASK, and DESCRIBE queries. UPDA TE op erations are rejected. Queries are sub ject to: 45 second timeout Complexity limits (max 50 triple patterns, OPTIONAL depth 5, UNION branc hes 10) Injection pattern detection and basic syntax v alidation Query results as a string. Results are truncated if to o large. Parameters schema {"additionalProp erties":false,"properties":{"sparql":{"description":"The SP ARQL query string. Must b e a v alid read-only query.","t yp e":"string"}},"required":["sparql"],"t yp e":"ob ject"} T ool Name T ool description received b y agent/LLMs get_on tology_snap- shot Retriev e complete sc hema snapshot for an ontology . Returns all classes with their data prop erties and ob ject prop erties in a single call, giving context for reasoning ab out the knowledge graph structure. "ontology_iri": str, "ontology_description": str, "class_coun t": int, "classes": class_iri: "py_class": "mo dule.ClassName", "description": "Class do cstring summary", "data_prop erties": prop ert y_- iri: "field": "field_name", "t yp e": "python.t yp e.P ath", "description": "Field description" , "ob ject_properties": ... Parameters schema {"additionalProp erties":false,"properties":{"ontology_iri":{"description":"The IRI of the on tology to insp ect.\nExample: \"https://elagen te.ca/on tomof\"","type":"string"},"class_iris":{"default ":n ull,"description":"Optional filter - only return these sp ecific classes.\nIf None, returns all classes in the ontology.","an yOf":[{"items":{"t yp e":"string"},"t yp e":"arra y"},{"type":"null "}]}},"required":["on tology_iri"],"type":"ob ject"} T ool Name T ool description received b y agent/LLMs get_instance_from_- kno wledge_graph Retriev e a kno wledge graph ob ject b y its IRI. Large data fields (ND Arrays, CIF text, etc.) are stripp ed and replaced with metadata. Use get_- cif_con tent to retrieve full CIF text. "instance_iri": str, "class_t yp e": "ClassName", "data": ... fields with large data stripp ed ... , "large_fields_av ailable": ["cif_text", "p ositions", ...] Parameters schema {"additionalProp erties":false,"properties":{"instance_iri":{"description":"The IRI of the ob ject to retriev e.\nExample: \"h ttps://elagente.ca/on tomof/ConstructedMOF_ab c123\"","type":" string"},"recursiv e_depth":{"default":3,"description":"How deep to recursiv ely pull related ob jects.\nDefault is 3 (direct prop erties and some related context). Increase for more context. F or pulling ConceptualAtoms or related structures, use -1.","type":"integer"}},"required":[" instance_iri"],"t yp e":"ob ject"} 28 T ool Name T ool description received b y agent/LLMs get_cif_con tent Retriev e the CIF conten t for a ConstructedMOF. CIF (Crystallo- graphic Information File) contains the full atomic structure for visualization or analysis. "mof_iri": str, "mof_name": str or None, "cif_text": "full CIF conten t...", "n_atoms": int Parameters schema {"additionalProp erties":false,"properties":{"mof_iri":{"description":"The IRI of the ConstructedMOF instance.","type":"string"}},"required":["mof_iri"],"type":"ob ject"} T ool Name T ool description received b y agent/LLMs run_p ython_co de (ex- p osed via MCP server) T o ol to execute Python co de and return stdout, stderr, and return v alue. Guidelines The c ode may b e async, and the v alue on the last line will b e returned as the return v alue. The co de will b e executed with Python 3.13 using p yodide - so adapt your co de if needed. Y ou co de must b e executed within a timeout. Y ou hav e 60 seconds b efore the run is canceled. Y ou hav e these python pack ages installed: n umpy ,pandas,matplotlib T o output files or images, sav e them in the "/output_files" folder. Parameters schema {"t yp e": "ob ject", "prop erties": {"python_code": {"type": "string", "description": "Python co de to run"}, "global_v ariables": {"type": "ob ject", "additionalProp erties": {}, "default": {}, " description": "Map of global v ariables in context when the co de is executed", "prop erties": {}}}, "required": ["python_code"], "additionalProp erties": false} 29 B Univ ersity-lev el quantum chemistry benchmark B.1 LLM model specifications T able S1 Pricing (p er million tokens) and maximum context window size for b enchmark ed LLM s. Data were sourced from official provider websites for proprietary mo dels and Op enRouter for op en mo dels. Pricing for cached tokens was not considered since cashing p olicies v ary across providers. LLM model Input tok ens ($/M) Output tok ens ($/M) Maximum context window gpt-4.1 2 8 1,047,576 gpt-5 1.25 10 400,000 gpt-5.1 1.25 10 400,000 gpt-5.2 1.75 14 400,000 minimax-m2 0.2 1 205,000 qw en3-max 1.2 6 262,144 claude-sonnet-3.7 3 15 200,000 claude-sonnet-4.5 3 15 200,000 "gpt-4.1-2025-04-14": {"temp erature": 0.0, "provider_name": "op enai", "provider_url": "https:// api.op enai.com/v1/"} "gpt-5-2025-08-07": {"temp erature": 1.0, "op enai_reasoning_effort": "low", "provider_name": " op enai", "provider_url": "https://api.openai.com/v1/"} "gpt-5.1-2025-11-13": {"temp erature": 1.0, "op enai_reasoning_effort": "low", "provider_name": " op enai", "provider_url": "https://api.openai.com/v1/"} "gpt-5.2-2025-12-11": {"temp erature": 1.0, "op enai_reasoning_effort": "low", "provider_name": " op enai", "provider_url": "https://api.openai.com/v1/"} "op enrouter:minimax/minimax-m2": {"temp erature": 1.0, "pro vider_name": "op enrouter", " pro vider_url": "h ttps://op enrouter.ai/api/v1"} "op enrouter:qw en/qw en3-max": {"temp erature": 0.0, "provider_name": "op enrouter", " pro vider_url": "h ttps://op enrouter.ai/api/v1"} "claude-3-7-sonnet-20250219": {"temp erature": 1.0, "anthropic_thinking": {"type": "enabled", " budget_tok ens": 1024}, "extra_headers": {"an thropic-b eta": "token-efficien t-to ols-2025-02-19", "pro vider_name": "an thropic", "provider_url": "https://api.an thropic.com"}} "claude-sonnet-4-5-20250929": {"temp erature": 0.0, "kind": "resp onse", "provider_name": " an thropic", "pro vider_url": "https://api.an thropic.com"} 30 B.2 User prompts Prompts were minimally revised to remov e OR CA-sp ecific data and ensure an unbiased comparison with the original El Agente ( 5 ) b enc hmark questions; no additional instructions w ere given. F unctionals and basis sets w ere adjusted sp ecifically for PySCF compatibility . Prompt Organic compounds level 1 Perform in parallel geometry optimization of the [comp ounds below] with the Hartree-F o ck (HF) metho d and def2-SVP basis set in the gas phase. Once the calculations hav e b een successfully completed, please generate individual rep orts for eac h of the molecules listed below, one at a time. Each rep ort should include the final Cartesian coordinates (in Å), total energy (in Hartrees), p oint group symmetry , dip ole moment (in Deby e), molecular orbital analysis (including an MO energy table and the HOMO-LUMO gap), atomic charge analysis (Mulliken, Löwdin, and IAO). Organic Compounds: 1. caffeine (SMILES: CN1C=NC2=C1C(=O)N(C(=O)N2C)C) 2. theobromine (SMILES: CN1C=NC2=C1C(=O)NC(=O)N2C) 3. acetylsalicylic_acid (SMILES: CC(=O)OC1=CC=CC=C1C(=O)O) Prompt Organic compounds level 2 Perform in parallel geometry optimization using the xyz files listed b elo w from the default working directory with the Hartree-F o c k (HF) metho d and def2-SVP basis set in the gas phase. Once the calculations hav e been successfully completed, please generate individual rep orts for ea c h of the molecules listed b elo w, one at a time. Each rep ort should include the final Cartesian co ordinates (in Å), total energy (in Hartrees), p oint group symmetry , dip ole moment (in Debye), molecular orbital analysis (including an MO energy table and the HOMO-LUMO gap), atomic charge analysis (Mulliken, Löwdin, and Hirshfeld). Organic Compounds: 1. caffeine_op enbabel.xyz (charge = 0; multiplicit y = 1) 2. theobromine_op enbabel.xyz (charge = 0; multiplicit y = 1) 3. aspirin_op enbabel.xyz (charge = 0; multiplicit y = 1) 4. methyl_salicylate_openbabel.xyz (charge = 0; multiplicity = 1) 5. acetaminophen_op enbabel.xyz (charge = 0; multiplicit y = 1) 6. triazaadamantane_openbabel.xyz (charge = 0; multiplicity = 1) 7. limonene_op enbabel.xyz (charge = 0; multiplicit y = 1) 8. D-glucose_op enbabel.xyz (charge = 0; multiplicit y = 1) 9. creatinine_amine_tautomer_op enbabel.xyz (charge = 0; multiplicit y = 1) 10. creatinine_imine_tautomer_op enbabel.xyz (charge = 0; multiplicit y = 1) 11. L-phenylalanine_zwitterion_openbabel.xyz (charge = 0; multiplicity = 1) 12. 2-chloronitrobenzene_op enbabel.xyz (charge = 0; multiplicit y = 1) 13. cis-1_2-cyclohexanediol_op enbabel.xyz (charge = 0; multiplicit y = 1) 14. L-histidine_non_zwitterion_op enbabel.xyz (charge = 0; multiplicit y = 1) 15. 2_2-biphenol_op enbabel.xyz (charge = 0; multiplicit y = 1) 16. S-2-ethyl-2-fluoropentan-1-ol_openbabel.xyz (charge = 0; multiplicity = 1) 17. R-3-hydro xycyclop en tan-1-one_op en bab el.xyz (charge = 0; multiplicit y = 1) 18. 3-methylbutanoate_anion_openbabel.xyz (charge = -1; multiplicity = 1) 19. diisopropylamide_anion_openbabel.xyz (charge = -1; multiplicity = 1) 20. diisopropylammonium_cation_openbabel.xyz (charge = +1; multiplicity = 1) Prompt Inorganic compounds level 1 Plan and act directly . Do not ask for my confirmation this time. Complete the following request: Perform in parallel geometry optimization of the [compounds b elow] with the Hartree-F o ck (HF) metho d and def2-SVP basis set in the gas phase. Once the calculations hav e b een successfully completed, please generate individual rep orts for each of the molecules listed below, one at a time. Each report should include the final Cartesian co ordinates (in Å), total energy (in Hartrees), point group symmetry , dip ole moment (in Deby e), molecular orbital analysis (including an MO energy table and the HOMO-LUMO gap), atomic charge analysis (Mulliken, Löwdin, and IAO). Inorganic Compounds: 1. Chromium(0) hexacarb onyl (low spin) – SMILES: [Cr](=C=O)(=C=O)(=C=O)(=C=O)(=C=O)(=C=O) 2. Chlorine trifluoride – SMILES: FCl(F)F 3. Fluorophosphoric acid (singly deprotonated form) – SMILES: [O-]P(F)(O)=O 31 Prompt Inorganic compounds level 2 Plan and act directly . Do not ask my confirmation this time. Complete the following request: Perform in parallel geometry optimization using the xyz files listed b elo w with the Hartree-F o c k (HF) metho d and def2-SVP basis set in the gas phase. Once the calculations hav e b een successfully completed, please generate individual rep orts for eac h of the molecules listed below, one at a time. Each rep ort should include the final Cartesian coordinates (in Å), total energy (in Hartrees), p oint group symmetry , dip ole moment (in Deby e), molecular orbital analysis (including an MO energy table and the HOMO-LUMO gap), atomic charge analysis (Mulliken, Löwdin, and Hirshfeld). List of Inorganic Comp ounds: 1. chromium_hexacarbonyl.xyz (charge = 0; multiplicit y = 1) 2. chlorine_trifluoride.xyz (charge = 0; multiplicit y = 1) 3. fluorophosphoric_acid_singly_deprotonated_form.xyz (charge = -1; multiplicity = 1) 4. trifluoromethane_sulfonate.xyz (charge = -1; multiplicity = 1) 5. cyclohexyldimethylphosphine.xyz (charge = 0; multiplicit y = 1) 6. t-butylisothiocyanate.xyz (charge = 0; multiplicit y = 1) 7. chromic_acid.xyz (charge = 0; multiplicit y = 1) 8. p ermanganic_acid.xyz (charge = 0; multiplicit y = 1) 9. p erchlorate.xyz (charge = -1; multiplicit y = 1) 10. hexafluorophosphate.xyz (charge = -1; multiplicity = 1) 11. tetrafluorob orate.xyz (charge = -1; multiplicit y = 1) 12. dicyanoaurate.xyz (charge = -1; multiplicit y = 1) 13. nitrogen_trifluoride.xyz (charge = 0; multiplicity = 1) 14. sulfur_hexafluoride.xyz (charge = 0; multiplicity = 1) 15. sulfur_tetrafluoride.xyz (charge = 0; multiplicity = 1) 16. xenon_tetrafluoride.xyz (charge = 0; multiplicity = 1) Prompt Carbocations compounds level 1 A carb o cation formation reaction is given by R-H -> R+ + H-. Y our task is to calculate the carb ocation formation enthalpies and Gibbs free energies for R-H = methane, ethane, propane, 2-methylpropane, toluene, benzene, dimethyl ether, trimethylamine, and propene. In your working directory , you can find: carbo_ch4.xyz, carbo_C2H6.xyz, carb o_C3H8.xyz, carbo_2-methylpropane.xyz, carb o_toluene.xyz, carb o_benzene.xyz, carb o_et2o.xyz, carb o_et3n.xyz, carbo_prop ene.xyz, and carbo_h-.xyz. Y o u will also find carb o_c h3+.xyz, carb o_C2H5+.xyz, carb o_C3H7+.xyz, carb o_2-meth ylpropyl+.xyz, carbo_toluene+.xyz, carb o_benzene+.xyz, carb o_et2o+.xyz, carb o_et3n+.xyz, and carb o_prop ene+.xyz for the cations. Please optimize these structures (except the hydride) using DFT with the B3L YP functional and def2-SVP basis set, and from the outputs, extract the relevan t information to calculate the carb ocation formation enthalpies and Gibbs free energies of each R-H. Rep ort the results (in kcal/mol) in a table. F or charge and multiplicit y , for molecules charge 0, multiplicit y 1; carbo cations charge 1, multiplicit y 1; hydride charge -1, multiplicit y 1. Prompt Carbocations compounds level 2 A carb o cation formation reaction is given by R-H -> R+ + H-. Y our task is to calculate the carb ocation formation enthalpies and Gibbs free energies for R-H = methane, ethane, propane, 2-methylpropane, toluene, b enzene, dimethyl ether, trimethylamine, and prop ene. The SMILES of each R-H are as follows: C, CC, CCC, CC(C)C, Cc1ccccc1, c1ccccc1, COC, CN(C)C, C=CC. The SMILES of each R+ is given b y [CH3+], [CH2+]C, C[CH+]C, C[C+](C)C, [CH2+]c1c(cccc1), c1[c+]cccc1, CO[CH2+], CN(C)[CH2+], [CH2+]C=C. Please use the SMILES strings in the table to generate the appropriate geometries, optimize them using DFT with the B3L YP functional and def2-SVP basis set, and from the outputs, extract the relev an t information to calculate the carbo cation formation enthalpies and Gibbs free energies of each R-H. Rep ort the results (in kcal/mol) in a table. Prompt Ring Strain compounds le vel 1 Compute the v alues of ∆ H and ∆ G for the following reactions: cyclo(CnH2n) → cyclo(Cn-1H2n-3)-CH3 Perform these calculations using B3L YP/def2-svp for v alues of n from 4 to 8 and use them to appro ximate the relativ e ring strain energies of cycloalk anes of size 3 to 8. Hint: The first reaction (n = 4) is cyclobutane (SMILES string C1CCC1) conv erting into methylcyclopropane (SMILES string CC1CC1). Each structure needs to b e optimized and frequencies must b e calculated to get the enthalp y and Gibbs free energies. Y ou will need to pick a reference p oin t to use as the “zero ring strain” p oint and compare the others relative to that. Rep ort a table of ring size vs. ring strain enthalp y and free energy . T o calculate the ring strain energy , start by assuming cyclo octane (n = 8) is the reference p oin t, and that its ring strain is zero. Then, the ring strain of cycloheptane (n = 7) is determined by the enthalp y or Gibbs free energy of the reaction cyclo octane → methylcycloheptane (n = 8 to n = 7), and the ring strain energy of cyclo o ctane (n = 8). Obtain this for n = 8 to n = 3. Finally , use cyclohexane (n = 6) as the reference p oin t of zero ring strain. 32 Prompt Ring Strain compounds le vel 2 Compute the v alues of ∆ H and ∆ G for the following reactions: cyclo(CnH2n) → cyclo(Cn-1H2n-3)-CH3 Perform these calculations using B3L YP/def2-svp for v alues of n from 4 to 8 and use them to appro ximate the relativ e ring strain energies of cycloalk anes of size 3 to 8. Hint: Y ou will need to pick a reference p oin t to use as the “zero ring strain” p oint and compare the others relative to th at. Report a table of ring size vs. ring strain enthalp y and free energy . The ring strain of cyclo(CnH2n) is determined by the reaction energy of cyclo(CnH2n) → cyclo(Cn-1H2n-3)-CH3 and the ring strain energy of cyclo(CnH2n). Prompt pKa Prediction compounds le vel 1 Plan and act directly . Do not ask for my confirmation this time. Complete the following request: Calculate the pKa of acetic acid in water using tw o calculations at the B3L YP/def2-SVP level of theory with the CPCM implicit solvation model. Prompt pKa Prediction compounds le vel 2 Plan and act directly . Do not ask my confirmation this time. Complete the following request: Calculate the pKa of chlorofluoroacetic acid using B3L YP def2-SVP . T o do so, first calibrate the free energy of solv ation of the proton based on the known literature v alues of some re- lated carboxylic acids: 1. Acetic acid; pKa = 4.76 2. Fluoroacetic acid; pKa = 2.586 3. Chloroacetic acid; pKa = 2.86 Prompt TDDFT compounds level 1 Compute the energy level of S1, the energy difference b etw een S1 and T1, and the oscillator strength to the S1 state for the following structures from the default working directory: tddft_2.xyz, tddft_3.xyz, tddft_5.xyz. Perform a single-p oin t TDDFT (after geometry optimization and chec king for geometric stability) calculation with B3L YP/def2-SVP . Prompt TDDFT compounds level 2 Compute the energy level of S1, the energy difference b etw een S1 and T1, and the oscillator strength to the S1 state for the following structures from the default working directory: tddft_2.xyz, tddft_3.xyz, tddft_5.xyz. Perform a single-p oin t TDDFT calculation with B3L YP/def2-SVP . 33 B.3 R ubrics for LLM judg e W e used gpt-4o as an indep endent LLM judge to ev aluate results on a scale from 0 to 1 based on the following rubric. The judge is required to return a single final score, whic h totals 100% (1.0) if all task-sp ecific criteria are satisfied. T able S2 The rubric used by the LLM judge when grading the b enchmark questions. Question L ev el W eight T ask Organic comp ounds 1 20% Correct input file, i.e., level of theory , required keyw ords, charge, and multiplicit y; 20% Con vergence of calculation, i.e., SCF, geometry optimization, and absence of imaginary frequency; 20% Rep ort generation; 20% Successful extraction and do cumentation of all rep ort v alues; 20% Successful generation of XYZ from SMILES. 2 20% Correct input file, i.e., level of theory , required keyw ords, charge, and multiplicit y; 20% Con vergence of calculation, i.e., SCF, geometry optimization, and absence of imaginary frequency; 20% Rep ort generation; 20% Successful extraction and do cumentation of all rep ort v alues; 20% Successful pro cessing of all input XYZ. Inorganic comp ounds 1 20% If correct input file, i.e., lev el of theory , required keyw ords, charge, and multiplicit y; 20% If con vergence of calculation, i.e., SCF, geometry optimization, and absence of imaginary frequency; 20% If generated a rep ort; 20% If suc cessful extraction and do cumen tation of all rep ort v alues; 20% If generated of XYZ from SMILES. 2 20% If correct input file, i.e., lev el of theory , required keyw ords, charge, and multiplicit y; 20% If con vergence of calculation, i.e., SCF, geometry optimization, and absence of imaginary frequency; 20% If generated a rep ort; 20% If suc cessful extraction and do cumen tation of all rep ort v alues; 20% If suc cessful pro cessing of all input XYZ. Carb ocations 1 40% Molecules: correct input file, output geometry , completed calculation, and data extraction; 40% Carb ocations: correct input file, output geometry , completed calcu- lation, and d ata extraction; 20% Results: correct ∆ H and ∆ G trends, and match chemical intuition. 2 40% Molecules: correct input file, output geometry , completed calculation, and data extraction; 40% Carb ocations: correct input file, output geometry , completed calcu- lation, and d ata extraction; 20% Results: correct ∆ H and ∆ G trends, and match chemical intuition. 34 T able S2 – contin ued from previous page Question L ev el W eight T ask Ring Strain 1 10% Correct struc tures from the formula; 10% Reasonable energy scale (not Hartree); 10% No imaginary frequencies; 10% P erforms just the right num ber of calculations (no strange single p oin t); 10% Consisten t level of theory used (DFT, basis set, solv ent mo del if an y); 10% Correctly extracted en thalpy and Gibbs free energies; 10% Correct referenc e energy (cyclohexane); 10% Correct ring strain magnitude (from extracted v alues); 10% Correct sign for ring strain; 10% All v alues rep orted (cyclopropane, Gibbs, and Enthalp y). 2 10% Correct struc tures from the formula; 10% Reasonable energy scale (not Hartree); 10% No imaginary frequencies; 10% P erforms just the right num ber of calculations (no strange single p oin t); 10% Consisten t level of theory used (DFT, basis set, solv ent mo del if an y); 10% Correctly extracted en thalpy and Gibbs free energies; 10% Correct referenc e energy (cyclohexane); 10% Correct ring strain magnitude (from extracted v alues); 10% Correct sign for ring strain; 10% All v alues rep orted (cyclopropane, Gibbs, and Enthalp y). pKa Prediction 1 33% Correct input files, i.e., level of theory , required keyw ords, charge, and multiplicit y; 33% Con vergence of calculation, i.e., SCF, geometry optimization, and absence of imaginary frequency; 33% Computed correct pK a ∼ 22 . 05 , anything roughly ab ov e 19 or b elo w 25 is c orrect. 2 25% Correct input files, i.e., level of theory , required keyw ords, charge, and multiplicit y; 25% Con vergence of calculation, i.e., SCF, geometry optimization, and absence of imaginary frequency; 25% Calibrated proton solv ation energy using linear regression or a verag- ing; 25% Computed reasonable v alue for pK a (e.g., a v alue b et ween -2.70 and 1.50). TDDFT 1 40% Correct inpu t file (level of theory as requested, tddft blo c k); 40% Calculation c ompleted normally; 20% Extract and rep ort the v alues correctly , and the num ber matched. 2 40% Correct inpu t file (level of theory as requested, tddft blo c k); 40% Calculation c ompleted normally; 20% Extract and rep ort the v alues correctly , and the num ber matched. 35 B.4 Pass k analy sis T o rigorously ev aluate agent p erformance b eyond simple accuracy , we adopt a tiered threshold approach with tw o complementary metrics as a balance b et ween capability and reliability: pass@k ( 34 ) to measure the probabilit y of obtaining at least one successful run in k attempts, and passˆk ( 30 ) to measure the probability that al l k attempts succeed as a metric for the robustness of the system. W e define a pass as a numerical score equal to 1.00 and an LLM-as-a-judge score greater than 0.90. This ensures b oth strict numerical v alidit y and high-quality p ost-pro cessing and report correctness as ev aluated by the LLM judge ( e.g. , rep orting the pKa correctly after a linear regression fit of the resulting energies). The results are summarized in Supplemen tary T able S5 . W e achiev e pass@3 of 0.99 and passˆ3 of 0.54 with gpt-5 . This demonstrates strong p oten tial for further system refinement tow ard pro duction deploymen t. T able S3 Pass@k and Passˆk rates with criteria: τ numerical ≥ 0 . 90 and τ llm_judge ≥ 0 . 90 . pass@k passˆk LLM Model @1 @3 @5 @10 ˆ1 ˆ3 ˆ5 ˆ10 gpt-4.1 0.81 0.99 1.00 1.00 0.81 0.52 0.34 0.11 gpt-5 0.94 1.00 1.00 1.00 0.94 0.83 0.74 0.53 gpt-5.1 0.87 1.00 1.00 1.00 0.87 0.65 0.48 0.22 gpt-5.2 0.84 1.00 1.00 1.00 0.84 0.59 0.42 0.17 minimax-m2 0.53 0.90 0.98 1.00 0.53 0.15 0.04 0.00 qw en3-max 0.53 0.90 0.98 1.00 0.53 0.15 0.04 0.00 sonnet-3.7 0.57 0.93 0.99 1.00 0.57 0.19 0.06 0.00 sonnet-4.5 0.78 0.99 1.00 1.00 0.78 0.48 0.29 0.08 T able S4 Pass@k and Passˆk rates with criteria: τ numerical = 1 . pass@k passˆk LLM Model @1 @3 @5 @10 ˆ1 ˆ3 ˆ5 ˆ10 gpt-4.1 0.68 0.97 1.00 1.00 0.68 0.30 0.13 0.02 gpt-5 0.82 1.00 1.00 1.00 0.82 0.56 0.38 0.13 gpt-5.1 0.76 0.99 1.00 1.00 0.76 0.43 0.24 0.06 gpt-5.2 0.82 1.00 1.00 1.00 0.82 0.56 0.38 0.13 minimax-m2 0.55 0.91 0.98 1.00 0.55 0.16 0.05 0.00 qw en3-max 0.55 0.91 0.98 1.00 0.55 0.16 0.05 0.00 sonnet-3.7 0.54 0.91 0.98 1.00 0.54 0.16 0.04 0.00 sonnet-4.5 0.68 0.97 1.00 1.00 0.68 0.30 0.13 0.02 T able S5 Pass@k and Passˆk rates with criteria: τ numerical = 1 and τ llm_judge ≥ 0 . 90 . pass@k passˆk LLM Model @1 @3 @5 @10 ˆ1 ˆ3 ˆ5 ˆ10 gpt-4.1 0.63 0.95 0.99 1.00 0.63 0.25 0.10 0.01 gpt-5 0.82 0.99 1.00 1.00 0.82 0.54 0.36 0.12 gpt-5.1 0.70 0.97 1.00 1.00 0.70 0.34 0.16 0.02 gpt-5.2 0.75 0.99 1.00 1.00 0.75 0.42 0.23 0.05 minimax-m2 0.43 0.81 0.94 1.00 0.42 0.07 0.01 0.00 qw en3-max 0.39 0.78 0.92 0.99 0.39 0.06 0.01 0.00 sonnet-3.7 0.43 0.82 0.95 1.00 0.43 0.08 0.01 0.00 sonnet-4.5 0.62 0.95 0.99 1.00 0.62 0.24 0.09 0.01 36 B.5 Statistics plots All b enc hmarks were executed on a high-p erformance computing no de equipp ed with four NVIDIA H100 (80 GB) GPUs. W e chose a maximum concurrency of four agen t runs. GPU4PySCF jobs were parallelized with up to three concurrent jobs p er GPU to saturate device utilization, allowing up to tw elv e molecules to b e pro cessed simultaneously . W e identify systematic differences in interaction patterns b et ween the GPT and Claude mo del families that ha ve direct implications for workflo w efficiency . GPT mo dels tend to batch to ol in vocations, dispatc hing m ultiple calls concurrently and returning to reason ov er aggregated results, whereas Claude mo dels more frequen tly interlea v e to ol calls with incremen tal reasoning. Within the Claude family , the elev ated cost of sonnet-3.7 is largely driv en by its limited ability to parallelize to ol calls, a kno wn constraint that p ersists even under recommended system prompt configurations. 7 Sonnet-4.5 impro ves in its ability to inv ok e GPU4PySCF w orkflows in parallel but incurs high cost due to frequent calls to a general-purp ose run_p ython_co de to ol for rep ort generation (see Figs. S11 and S12 ). Notably , sonnet-3.7 exhibits a recurring execution pattern in geometry optimization tasks, progressively reducing parallelism by issuing to ol calls in batches of five, then three, tw o, and finally one, while sonnet-4.5 rep eatedly inv okes excessive generic co de execution calls for p ost-processing. These b eha viours highlight that mo del capability alone do es not determine system-lev el efficiency; rather, the alignment b et ween a mo del’s interaction style and the av ailable to ol abstractions plays a critical role. In suc h cases, encapsulating recurrent p ost-pro cessing steps in to dedicated to ols could further mitigate unneces sary to ol in vocation ov erhead. F or the b o x plots (Figs. S7 , S8 , S9 , and S10 ), eac h b o x spans from quartile 1 (Q1) to quartile 3 (Q3). The second quartile (Q2) is marked by a line inside the b ox. By default, the whiskers extend to Q1 - 1.5 × IQR and Q3 + 1.5 × IQR, where IQR = Q3 - Q1. Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0.5 0.6 0.7 0.8 0.9 1 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0.5 0.6 0.7 0.8 0.9 1 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Lev el 1 Lev el 2 Figure S1 Radar plots of n umerical-ev aluator scores a veraged ov er 10 runs by task and mo del for Level 1 and Level 2. Eac h p olygon sho ws a mo del’s mean score across tasks. 7 https://platform.claude.com/docs/en/agents- and- to ols/tool- use/implemen t- to ol- use#maximizing- parallel- tool- use 37 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0.5 0.6 0.7 0.8 0.9 1 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0.5 0.6 0.7 0.8 0.9 1 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Lev el 1 Lev el 2 Figure S2 Radar plots of LLM-as-a-judge-ev aluator scores av eraged ov er 10 runs by task and mo del for Level 1 and Lev el 2. Eac h p olygon shows a mo del’s mean score across tasks. 38 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 100k 200k 300k 400k 500k 600k gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 0.2M 0.4M 0.6M 0.8M Base Input (Static) T ool R esults Input R easoning Output Action/T ext Output Lev el 1 Lev el 2 Figure S3 Mean token comp osition by generation source across tasks, mo dels, and levels. Stack ed bars categorize input tok ens into base prompts (system prompts, to ol schemas, and user task prompts) and to ol-result inputs, and output tok ens into reasoning tokens, action/to ol-call outputs, and text outputs. 39 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 0.5 1 1.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 0.5 1 1.5 2 2.5 3 Base Input (Static) T ool R esults Input R easoning Output Action/T ext Output Lev el 1 Lev el 2 Figure S4 Same split as Fig. S3 , but measured in USD cost calculated from pricing provided by LLM providers. 40 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 100k 200k 300k 400k 500k 600k gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 0.2M 0.4M 0.6M 0.8M Base Input (Static) Normal Oper ation (In/Out) Immediate R etry Carry o v er Burden Lev el 1 Lev el 2 Figure S5 Mean token comp osition by role across tasks, mo dels, and levels. Stack ed bars partition total tokens into base input, normal op eration (in/out), immediate retry , and carryo v er burden across agentic traces; rows separate Lev el 1 and Level 2. 41 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 0.5 1 1.5 2 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T 0 0.5 1 1.5 2 2.5 3 Base Input (Static) Normal Oper ation (In/Out) Immediate R etry Carry o v er Burden Lev el 1 Lev el 2 Figure S6 Same split as Fig. S5 , but measured in USD cost calculated from pricing provided by LLM providers. 42 5 10 15 20 25 30 5 10 15 20 25 30 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Lev el 1 Lev el 2 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T Figure S7 Distribution of LLM requests by mo del across tasks and levels. Boxes show p er-model v ariabilit y in LLM requests, with ov erlaid p oin ts for individual runs; facets separate tasks (columns) and levels (rows), and colours denote mo dels. 0 50k 100k 150k 200k 0 50k 100k 150k 200k gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Lev el 1 Lev el 2 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T Figure S8 Distribution of final call tokens by mo del across tasks and levels. Bo xes show p er-model v ariability in final call total tokens, with ov erlaid p oin ts sized by n umber of LLM requests; facets separate tasks (columns) and levels (ro ws), and colours denote mo dels. 43 0% 20% 40% 60% 0% 20% 40% 60% gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 Lev el 1 Lev el 2 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T Figure S9 Distribution of final call context windo w pressure (final call tokens divided by the maximum context window) b y mo del across tasks and levels. Boxes show p er-model v ariability , with ov erlaid p oints representing individual runs scaled by the n umber of LLM requests; facets split tasks (columns) and levels (rows), with colours denoting mo dels. 0 2k 4k 6k 8k 10k 0 2k 4k 6k 8k 10k gpt -5 gpt -5.1 gpt -5.2 minimax -m2 Lev el 1 Lev el 2 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T Figure S10 Distribution of reasoning token usage by mo del across tasks and levels. Boxes show p er-model v ariabilit y in reasoning tokens (filtered to LLM mo dels with reasoning capabilities; sonnet-3.7 has a consistent reasoning budget of 2024), with ov erlaid p oints for individual runs; facets split tasks (columns) and levels (rows), and colours denote mo dels. One run from minimax-m2 for Ring Strain (Level 2) consumed 102.91k reasoning tokens and is omitted from the plot due to the y-axis limit. 44 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 0 2 4 6 8 10 12 14 R un PySCF (T ool) R un PySCF (LLM) R un Python (T ool) R un Python (LLM) Unit Con v ersion (T ool) Unit Con v ersion (LLM) Figure S11 A verage num ber of to ol calls versus LLM API requests p er mo del for the b enchmark exercise. F or each mo del, stack ed bars compare three to ol types ( run_p yscf_workflo w , run_p ython_co de , get_con version_factor ), sho wn separately for actual to ol calls and corresp onding LLM requests (side-by-side groups). gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 0 5 10 15 20 25 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 sonnet -4.5 gpt -4.1 gpt -5 gpt -5.1 gpt -5.2 minimax -m2 qwen3-max sonnet -3.7 0 10 20 30 R un PySCF (T ool) R un PySCF (LLM) R un Python (T ool) R un Python (LLM) Unit Con v ersion (T ool) Unit Con v ersion (LLM) Lev el 1 Lev el 2 Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T Carbocations Inorganic Organic pK a Prediction Ring Str ain TDDF T Figure S12 T o ol usage v ersus LLM API requests by task and lev el for the b enc hmark exercise. F or eac h task (group ed blo c ks) and mo del (ticks), stack ed bars compare av erage to ol calls (left bar) with corresp onding LLM requests (right bar) for the run_p yscf_workflo w , run_p ython_co de , and get_con version_factor to ols; rows separate levels and colours en code to ol typ e. 45 B.6 Bare LLM ag ent B.6.1 Summary of results Informed by prior work on agentic systems that integrate co de execution and web search, where LLM agen ts dynamically construct quantum sim ulation scripts using these to ols, we examine whether a similar approach generalizes to computational chemistry b enchmarks. W e set up a light w eight LLM agen t with a web search to ol and co de execution capabilities and tested it on tw o represen tative El Agente tasks: inorganic comp ounds lev el 1 and pKa prediction level 2. The first exercise examines the LLM ’s ability to write scripts to optimize the geometry , remov e imaginary frequencies, obtain the p oin t group symmetry , and obtain the IAO p opulations, none of which are readily accessible from PySCF with one-liners. The latter exercise examines the LLM ’s abilit y to extract the Gibbs free energy and fit the results against the provided exp erimen tal v alues. F or the inorganic comp ounds exercise, the agent sets up the molecules from SMILES using the Op enBab el pac k age, p erforms geometry optimization with PySCF, follow ed by a tighter conv ergence single-p oint calcu- lation. How ev er, it was not able to extract all prop erties correctly . In particular, it made mistakes in: (1) extracting the p oin t-group symmetry , (2) running Lowdin analysis correctly , (3) omitting frequency analysis and imaginary frequency remov al, and (4) creating the correct geometry of chlorine trifluoride (pro ducing a trigonal planar structure instead of the correct T-shap ed structure). The full transcript can be found in Sec. B.6.2 . F or the pKa prediction exercise level 2, the agent was also able to run chloro-, fluoro-, and chlorofluoro-acetic acid, as well as acetic acid, successfully . Molecular geometries w ere first generated from SMILES and then optimized with PySCF, follow ed b y frequency analysis to obtain the Gibbs free energy . Ho wev er, to estimate the free energy of solv ation, the LLM obtained energies using the COSMO implicit solven t mo del and subtracted them from the Gibbs free energy of the gas-phase molecules. With this inaccurate proto col, it obtained a pKa of -5.0, well outside the range considered correct in our rubric (-2.70 to 1.50). The LLM also omitted chec king for imaginary frequencies after geometry optimization. The full transcript can b e found in Sec. B.6.3 . These tw o exp erimen ts were conducted with gpt-5 . The inorganic comp ounds and pk a prediction exercises to ok 40 min utes and 16 minutes and cost 650k and 450k tokens (total), resp ectiv ely . This is an order of magnitude more tok ens than were needed to run Gráfico , demonstrating the adv an tage of careful agentic to ol design. Moreo ver, the sto chastic nature of LLM s is evident in their approaches: the agent decided to search the web for the pKa prediction task and insp ected the PySCF source co de for the inorganic compounds exercise. While these b eha viours are interesting to study , they underline the unreliability of LLM b eha viours across rep eated runs, rendering them unreliable rather than dep endable co-pilots. Nonetheless, the LLM -generated functions could b e used for automatic to ol generation to extend agentic system capabilities. System prompt pro vided to this light w eigh t agent is (no additional context or instructions provided): Y ou ' re a computational chemistry agent that interprets user in tent and executes quan tum c hemistry calculations by writing your own python functions using the follo wing p ython pac k ages: gpu4pyscf, rdkit, nump y, and op en bab el. Y ou can execute your Python co de via the ` Python_REPL ` to ol. Y ou can use DuckDuc kGo to searc h for co de examples of gpu4p yscf on https://gith ub.com/p yscf/ gpu4p yscf/tree/master/examples. Y ou ha ve access to GPU, so AL W A YS use imp orts from gpu4pyscf to accelerate the calculation, or use ` to_gpu() ` metho d from p yscf to make sure it ' s accelerated on GPU. When running to ols in parallel, fix error for an y of the failed to ols b efore reply bac k. 46 B.6.2 Bare agent transcript: inorganic compound: lev el 1 Gráfico was configured with gpt-5 (temp erature = 1, reasoning effort set to lo w ) for this run, with access to the to ols Python_REPL for co de execution and duc kduckgo_searc h for web search. F or the complete chat transcript, please refer to GitHub rep o: h ttps://github.com/jb2197/ElAgenteGraf ic o- ChatTranscript . B.6.3 Bare agent transcript: pka prediction: lev el 2 Gráfico was configured with gpt-5 (temp erature = 1, reasoning effort set to lo w ) for this run, with access to the to ols Python_REPL for co de execution and duc kduckgo_searc h for web search. F or the complete chat transcript, please refer to GitHub rep o: h ttps://github.com/jb2197/ElAgenteGraf ic o- ChatTranscript . 47 C Use case e xtension 1: Boltzmann-w eighted spectroscopic properties Chemical pro cesses rarely o ccur on the basis of a single, static molecular structure. Instead, a thermo dynamic ensem ble, e.g., one mole of molecules, comprises a large num ber of distinct conformations, often num bering in the millions or more ( 35 ). This configurational diversit y go es b ey ond finite-temp erature effects arising from the o ccupation of vibrational energy levels, which are typically accoun ted for via thermostatistical corrections ( 94 ). A ccurately predicting sp ectroscopic prop erties in solution, therefore, requires explicit sampling of relev ant molecular conformations and Boltzmann-w eigh ted av eraging ov er the resulting ensem ble. Bey ond brute- force sampling via long and/or high-temp erature molecular dynamics ( MD ) simulations, a range of efficient algorithms for c onformer search and ensemble generation has emerged in recent y ears ( 35 ; 37 ; 38 ). T o assess solven t effects on electronic absorption sp ectra under b oth implicit and explicit solv ation mo dels, we execute tw o user queries to demonstrate agentic quantum-c hemistry workflo ws that co ordinate conformer sampling, solv ation mo delling, geometry refinement using DFT, and sp ectral analysis. A cross b oth use cases, the agent orc hestrated sp ecialized to ols, including CREST ( 25 ) for conformer discov ery , QCG for explicit solven t cluster construction ( 46 ), and PySCF for electronic structure calculations, such as geometry optimizations, vibrational analysis, and TDDFT , while managing parallel execution and enforcing metho dological constraints. In termediate molecular structures were exc hanged using knowledge-graph-bac k ed ConceptualA toms iden tifiers, enabling repro ducible to ol-to-tool handoff without rep eated serialization of large co ordinate payloads through the context window of LLMs. Effect of implicit solvents T o inv estigate the effect of different implicit solven ts on the absorption sp ectra of a mero cy anine comp ound (see Supplementary Sec. C.1 for Cartesian co ordinates), the agent decomp osed the task in to tw o fully indep enden t solven t-specific pip elines for w ater and n -heptane, which were executed concurrently . CREST conformer searches at the GFN2-xTB/ALPB ( 39 ; 40 ) level of theory were launched in parallel for each solv ent, yielding ensembles of low-energy conformers represented by p ersisten t ConceptualA toms iden tifiers. These identifiers served as light weigh t handles for dispatc hing downstream PySCF workflo ws in parallel, resulting in five and four DFT calculations, resp ectiv ely , for water and n -heptane at the ω B97X-D4/def2-TZVP lev el of theory ( 41 ; 42 ; 43 ). Within the PySCF workflo w, dynamic routing enforced correct metho dological sequencing: geometry optimization and frequency analysis were mandatory gates b efore TDDFT . In the w ater pip eline, one conformer exhibited an imaginary frequency after optimization, prompting an attempted imaginary-mo de remov al follow ed by re-optimization. The routing logic terminated the workflo w after re- optimization without pro ceeding to TDDFT . Although this outcome was not flagged as an exception that w ould trigger an automatic retry , the agent initiated a targeted remediation step that reused the intermediate geometry and re-ran optimization and frequency analysis until a true minimum was confirmed. This “repair” lo op ensured that TDDFT w as p erformed only on v alidated minima, in accordance with the user’s inten t, thereb y preven ting in v alid sp ectra and unnecessary computation. After DFT refinemen t, Gráfico follow ed the user request and deduplicated the conformers in to unique minima using an energy-based filter with additional RMSD v alidation. Boltzmann weigh ts were recomputed from DFT energies, and TDDFT sp ectra w ere combined via weigh ted Gaussian broadening to pro duce solven t-sp ecific ensemble sp ectra. Under the c hosen mo del and broadening parameters, the ensemble maxima for water and n -heptane ov e rlapped closely , indicating minimal sp ectral shifts induced by the implicit solv ation in this case. This query was completed in 35 minutes with 440k tokens, costing $1.11 with gpt-5.2 (medium reasoning effort). Effect of explicit solvation F or explicit solv ation, the agen t orchestrated a multi-stage w orkflow to compare absorption sp ectra of 2,3-ep o xybutanol in the gas phase and in explicit solv ation within a single execution plan (see Supplementary Sec. C.1 ). Gas-phase conformer ensembles were generated using CREST (GFN2-xTB), while, in parallel, an explicitly solv ated solute with a cluster of 15 w ater molecules was constructed using QCG ( 46 ) and further sampled with CREST in its non-co v alen t interaction ( --nci ) mo de. F or the subsequen t DFT calculations, lo w-energy conformers from b oth environmen ts were selected up to a cumulativ e Boltzmann w eight of 95 %. F our gas-phase conformers and six conformers from the explicitly solv ated pip eline were refined using a PySCF w orkflow ( ω B97X-D4/def2-TZVP) analogous to the implicit-solv ation case study , but with vibrational analysis omitted to reduce computational cost. Geometry optimization and TDDFT excited-state calculations were executed concurrently across multiple conformers to exploit conformer-level parallelism. Sp ectra w ere subsequen tly broadened and com bined in to a comparative visualization. The 48 explicitly solv ated ensem ble exhibits a pronounced solven t-induced red shift of the dominant absorption band relativ e to the gas-phase sp ectrum, consistent with stabilization of the excited state by the surrounding w ater cluster. As in the implicit-solven t case, ConceptualA toms iden tifiers enabled p ersisten t, repro ducible referencing of molecular structures across to ols. T ogether, this workflo w demonstrates an agentic system that p erforms high-level planning and reasoning ov er a structured scientific state, while orchestrating sto c hastic sampling, explicit solv ation mo delling, and electronic-structure calculations as a compiled workflo w. This query was completed in 30 minutes with 185k tokens costing $0.44 with gpt-5.2 (medium reasoning efforts). C.1 Boltzmann-weighted spectroscopic properties in implicit solutions Gráfico w as configured with gpt-5.2 (temp erature = 1, reasoning effort set to lo w ) for this run. The chat- completion API endp oin t was used b ecause, at the time of execution, the resp onses API did not supp ort SV G inputs (accepting only image/jp eg , image/png , image/gif , and image/w ebp formats). As a result, the detailed reasoning summary was not exp osed. The routing agent was configured with gpt-4.1 (temp erature = 0.1 using the chat-completion API endp oint). F or the complete chat transcript, please refer to GitHub rep o: h ttps://github.com/jb2197/ElAgenteGraf ic o- ChatTranscript . 49 C.2 Boltzmann-weighted spectroscopic properties with e xplicit solvation Gráfico was configured with gpt-5.2 (temp erature of 1, ‘medium’ reasoning effort, and ‘detailed’ reasoning summary) for this run. The resp onses API endp oin t w as used here. The routing agent was configured with gpt-4.1 (temp erature = 0.1 using the chat-completion API endp oin t). F or the complete chat transcript, please refer to GitHub rep o: h ttps://github.com/jb2197/ElAgenteGraf ic o- ChatTranscript . D Use case extension 2: Exploration of metal-organic frameworks design space D .1 MOF e xecution graph node implementations This section describ es the in ternal data flow within the MOF execution graph, detailing how structures are decomp osed, represented, translated b etw een softw are libraries, and passed through each computational stage. Throughout, the execution graph op erates ov er typed state ob jects and optionally p ersists intermediate and final artifacts to the knowledge graph via a custom OGM lay er (based on t wa ( 29 ); see Sec. 5.1 ). High-lev el node chains. Based on the connections (type hin ts) of each no de, we exp ect the MOF execution graph to supp ort three main entry routes. It should b e noted, ho wev er, that the routing b et ween these no des is con trolled by an LLM API call that decides the next no de based on b oth the current state of the execution graph and the summarized user inten t. It is therefore p ossible that unexp ected b eha viour may emerge from an agent op erating ov er the MOF execution graph, e.g. , as observed in Supplementary Sec. D.2.3 . • CIF route (local or downloaded): StartW orkflow → (DownloadF romCSD) → Pro cessCIF → Build- MOFBasic → MLFFGeomOpt → ZeoppAnalysis → MOF GraphEnd . [ Note: Downlo adF r omCSD is use d only when a CSD r efc o de is pr ovide d; when a CIF p ath is pr ovide d, the exe cution gr aph enters dir e ctly at Pr o c essCIF . ] • K G -driven route: StartW orkflow → CombinatorialSearc h → BuildMOFBasic → MLFF GeomOpt → ZeoppAnalysis → MOFGraphEnd . [ Note: it has b e en observe d that sometimes the main agent only wants to insp e ct the p ossible new MOF s without c onstructing them, ther efor e it instructs the r outing agent to exit after the c ombinatorial se ar ch. ] • PORMAKE-component route (direct building blocks): StartW orkflow → BuildMOFBasic → MLFFGeo- mOpt → ZeoppAnalysis → MOFGraphEnd . D .1.1 CIF decomposition to OGM building blocks When an exp erimental CIF enters the execution graph via the ProcessCIF no de, it undergo es a three-step decomp osition pip eline implemented in on tomofs_from_cif.py . Step 1: T opology classification. The CIF is passed to CrystalNets.jl ( 90 ) (via juliacall ) to determine its underlying top ological net. The classification returns a top ology lab el ( e.g. , p cu , m tn ), a dimensionality , and a catenation num b er. The no de currently restricts its scop e to catenation = 1 with a single top ology lab el; CIF s yielding multiple lab els are rejected. Step 2: Structure decomposition. PORMAKE’s ( 56 ) exp erimen tal MOFDecomp oser is inv oked on the CIF to extract provisional building blo cks as ASE ( 93 ) A toms ob jects. These raw fragments include b oth real atoms and PORMAKE “do c king p oints” (pseudo-atoms with atomic num ber Z = 0 ) that mark connection sites. Step 3: Building block deduplication. Because the decomp oser pro duces one fragment p er crystallographic site (whic h may include symmetry-equiv alen t copies), a tw o-stage deduplication strategy reduces the set to unique comp onen ts: • Stag e 1 (fast grouping): apply PORMAKE’s hash_atoms (complexit y parameter 7 ) to structurally hash and group fragments. 50 • Stag e 2 (structural verification): only when multiple candidate no des/linkers remain after hashing, p erform pairwise structural comparison using mofid ’s ( 54 ) interface to pymatgen’s StructureMatc her ( 89 ) with tolerances ltol=0.3 , stol=2.0 , and angle_tol=5.0 . The pip eline then enforces the current scop e constraint: exactly one unique metal no de and one unique organic link er p er CIF. OGM instantiation and in-process canonicalization. Eac h deduplicated building block is instantiated as an OGM class: MetalNo de or OrganicLink er , b oth sub classes of BuildingBlo c k . These OGM classes wrap the underlying PORMAKE BuildingBlo c k ob jects and enforce seman tic constraints: MetalNo de v alidates has_metal=T rue (via PORMAKE metal detection), while OrganicLink er v alidates has_metal=F alse . The top ology is similarly wrapp ed as a T op ology OGM instance. T o preven t d uplicate Python ob jects when parallel execution graph inv o cations pro cess the same comp onen t concurren tly , T op ology , MetalNo de , and OrganicLink er instances are registered in an in-pro cess canoni- calization registry guarded b y p er-k ey threading.RLo c k instances. This provides within-pr o c ess identit y stabilit y under concurrency . When K G in tegration is enabled, a second deduplication lay er queries the external graph for pre-existing equiv alen ts. These functions implement a t wo-stage strategy: a SP ARQL query first filters candidates b y atom count (for building blo c ks) or identifier (for top ologies), then p erforms structural comparison using pymatgen’s StructureMatc her . If an equiv alent is found in the K G , the existing instance is reused rather than creating a duplicate. D .1.2 OGM instances and ConceptualAtoms The ontomofs OGM classes store atomic geometry through ConceptualA toms , a Pydantic mo del that serves as the framework’s canonical v alidated geometry container. When a PORMAKE building blo c k is wrapp ed, its ASE A toms ob ject is split into tw o parts: • BuildingBlo c k.atoms ( ConceptualA toms ): contains only real atoms ( Z = 0 ), with p ositions in Å, the unit cell (if p erio dic), p eriodic boundary conditions, and electronic state (c harge, spin multiplicit y). A tomic num bers and p ositions are stored in an XYZ sub-mo del as v alidated NumPydantic arrays. • BuildingBlo c k.do c king_p oin ts ( XYZ ): con tains the pseudo-atoms ( Z = 0 ) that mark PORMAKE connection sites . Round-tripping across libraries. ConceptualA toms provides bidirectional conv ersion utilities: • from_ase_atoms(atoms) / to_ase_atoms(...) : round-trip to and from ASE A toms , preserving charge and spin m ultiplicity via atoms.info . • from_qcelemen tal_molecule(mol) / to_qcelemen tal_molecule() : round-trip to and from QCElemen- tal Molecule instances. Identity across in-memory state and K G persistence. ConceptualA toms is integrated with the OGM la yer (via GraphBaseMo del ), meaning each instance carries an instance_iri that uniquely iden tifies it across in-memory Python state and the external knowledge graph. When desired, this IRI can b e preserved through ASE round-trips by embedding it in atoms.info["instance_iri"] using to_ase_atoms(preserv e_iri=T rue) . When from_ase_atoms encoun ters an IRI in the info dict, it can retrieve the existing ConceptualA toms from the OGM lo okup table (when a v ailable) rather than creating a new instance, ensuring ob ject iden tity is main tained across serialization b oundaries. D .1.3 MOF combinatorial search algorithms Query S1 presen ts a SP ARQL query to prop ose candidate MOF s from historical syn thesis data, inspired by the similar algorithm implemented for metal-organic p olyhedra ( 48 ; 29 ). The underlying hypothesis is that c hemical building blo c ks (metal no des and organic linkers) that hav e successfully crystallized into a sp ecific top ological net in prior exp erimen ts p ossess inherent chemical or steric compatibilities with that top ology . F or a target topology T , the algorithm retrieves the set of metal no des M prov en = { m | ∃ ( m, T ) ∈ K obs } and organic linker L prov en = { l | ∃ ( l, T ) ∈ K obs } , where K obs represen ts the set of exp erimentally observed MOF s instantiated in the graph. A Cartesian pro duct P = M prov en × L prov en is computed and subsequently 51 filtered for pairs ( m, l ) where the metal no de and linker represent different lo cal structural roles to satisfy top ological constraints. The p ossible com binations are then filtered by negation to remov e any pair ( m, l , T ) that already exists as a mofs:ConstructedMOF in the graph. The query can b e configured with V ALUES clause to restrict the prop osed candidate to only include certain top ologies or building blo c ks. Query S1 In tra top ology search # Algorithm 1: find new MOF combinations using top ologies that have already # succeeded with each comp onen t (metal no de and organic linker) individually. # # Placeholder markers ( ` # {{V ALUES_*}} ` ) are replaced programmatically by # ` sparql_alg_with_v alues ` in ` sparql_utilities.py ` to inject optional V ALUES clauses. PREFIX mofs : PREFIX rdfs : PREFIX grafico : PREFIX rdfs : PREFIX grafico :
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment