MeDUET: Disentangled Unified Pretraining for 3D Medical Image Synthesis and Analysis
Self-supervised learning (SSL) and diffusion models have advanced representation learning and image synthesis. However, in 3D medical imaging, they remain separate: diffusion for synthesis, SSL for analysis. Unifying 3D medical image synthesis and an…
Authors: Junkai Liu, Ling Shao, Le Zhang
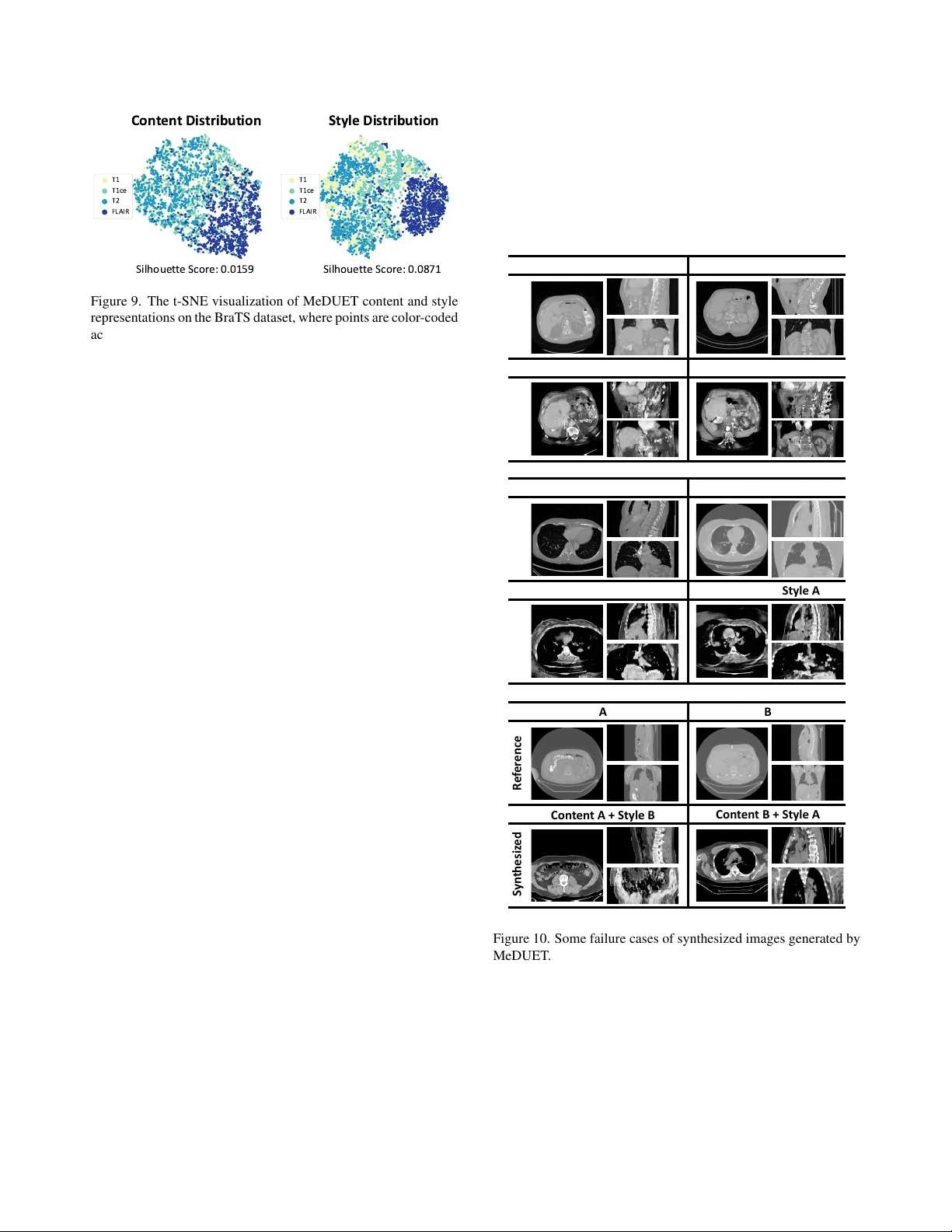
MeDUET : Disentangled Unified Pr etraining f or 3D Medical Image Synthesis and Analysis Junkai Liu 1 , Ling Shao 2 , Le Zhang 1 1 School of Engineering, Uni versity of Birmingham, UK 2 UCAS-T erminus AI Lab, Univ ersity of Chinese Academy of Sciences, Beijing, China jxl1920@student.bham.ac.uk; l.zhang.16@bham.ac.uk Abstract Self-supervised learning (SSL) and diffusion models have advanced r epr esentation learning and ima ge synthesis. However , in 3D medical imaging, they r emain separ ate: diffusion for synthesis, SSL for analysis. Unifying 3D medi- cal imag e synthesis and analysis is intuitive yet challenging, as multi-center datasets exhibit dominant style shifts, while downstr eam tasks r ely on anatomy , and site-specific style co-varies with anatomy acr oss slices, making factors unre- liable without e xplicit constraints. In this paper , we pr o- pose MeDUET , a 3D Me dical image D isentangled U nifi E d Pr e T raining framework that performs SSL in the V aria- tional Autoencoder (V AE) latent space which e xplicitly dis- entangles domain-invariant content fr om domain-specific style. The token demixing mechanism serves to turn disen- tanglement fr om a modeling assumption into an empirically identifiable pr operty . T wo novel pr oxy tasks, Mixed-F actor T oken Distillation (MFTD) and Swap-in variance Quadru- plet Contrast (SiQC), ar e devised to synergistically en- hance disentanglement. Once pretr ained, MeDUET is ca- pable of (i) delivering higher fidelity , faster conver gence, and impr oved contr ollability for synthesis, and (ii) demon- strating str ong domain generalization and notable label ef- ficiency for analysis across diverse medical benchmarks. In summary , MeDUET con verts multi-source heter ogeneity fr om an obstacle into a learning signal, enabling unified pr etraining for 3D medical image synthesis and analysis. The code is available at https : // github . com/ JK - Liu7/MeDUET . 1. Introduction Recently , self-supervised learning (SSL) following the pretraining-finetuning paradigm has emer ged as a power- ful approach, significantly improving performance across div erse downstream applications [ 7 , 28 , 29 , 53 ]. Mean- while, image generation has seen remarkable progress with ( a ) D om a i n S h i f ts ( b) F a ct or I d e n t i f i a b i l i t y Figure 1. The motivation of our proposed MeDUET . (a) La- tent similarity heatmap across domains ( S inter / S intra : inter- /intra-domain similarity). Compared with baseline medical SSL, which exhibits site-driven feature blocks, MeDUET isolates do- main shifts within the style map while maintaining a uniformly consistent content map across domains. (b) Latent t-SNE colored by domain. The baseline SSL clusters embeddings primarily by domain rather than anatomy , indicating style-dominated represen- tations, whereas MeDUET disentangles content and style in sepa- rate embedding spaces, enhancing factor identifiability . diffusion models [ 17 , 34 , 60 ]. Moti v ated by the intuition that generative modeling can foster a deeper understanding of visual data, pioneering studies [ 78 , 82 ] hav e shown that diffusion models pos sess outstanding representational ca- pability for perception tasks. In medical AI, both SSL and generativ e models have attracted considerable attention for 3D medical image synthesis [ 25 , 69 ] and analysis [ 4 , 79 ], respectiv ely . Nonetheless, the potential of diffusion mod- els in medical perception tasks, i.e., unified pretraining for both generation and understanding, remains largely unex- plored. Inspired by prior w ork [ 14 ], we ask the following research question: How can we establish a unified pr etrain- ing framework that benefits both 3D medical image synthe- sis and analysis tasks? Real-world 3D medical datasets aggreg ate heteroge- neous sites, scanners, field strengths, sequences, and co- horts [ 4 , 79 , 80 ], amplifying domain shifts mainly at the style lev el. F or example, a liv er CT model trained on Hos- pital A (Siemens, 120 kVp, v enous, 2.5 mm, soft kernel) de- grades on Hospital B (GE, 100 kVp, arterial, 1.0 mm, sharp) because HU histograms, noise, and edge profiles change while anatomy does not. In contrast, downstream tasks rely on content such as organ topology , lesion morphology , and anatomical continuity [ 44 , 54 ]. A na ¨ ıve unification of syn- thesis and analysis entangles style and anatomy , reducing generator controllability and causing perception models to ov erfit to style cues, which weakens generalization and la- bel efficienc y [ 52 , 67 ]. Thus, domain generalization is vital for unified pretraining but remains underexplored in current SSL framew orks (Fig. 1 (a)) [ 65 , 68 , 70 , 90 ]. In light of this challenge and opportunity , we formal- ize each volume as composed of two factors, a domain- in variant content factor that encodes intrinsic anatomy and captures stable semantics across domains, and a domain- specific style that reflects acquisition conditions and vi- sual appearance, accounts for peripheral, domain-varying changes [ 5 , 23 , 52 , 90 ]. Howe ver , clinical style differences are often subtle or ev en imperceptible, where the same structure can look different across centers while its geome- try is unchanged, making factor identifiability non-trivial for reliable encoding (Fig. 1 (b)) [ 44 , 85 ]. For instance, in contrast-enhanced CT , hyperattenuation may stem from true lesion enhancement (content) or from contrast timing, kVp, or reconstruction kernel (style). W ithout explicit dis- entanglement, multiple content and style e xplanations can account for the same image, rendering the factors unidenti- fiable. Motiv ated by the above analysis, we present MeDUET , a 3D Me dical image D isentangled U nifi E d Pre T raining model performed in the V AE latent space, facilitating both synthesis and analysis (Fig. 2 (a)). This latent space en- ables compact spatial tokenization of 3D volumes while naturally interfacing with latent diffusion. Our key in- sight is to reconcile the distinct goals of generation and perception by explicitly disentangling latent tokens into a domain-in variant content pathway and a domain-specific style pathw ay . Opposing task objectives encourage con- tent tokens to be domain-in variant while style tokens cap- ture acquisition-related v ariations. T o ensure practical iden- tifiability , a demixing module simulates “shared anatomy , different styles, ” yielding more rob ust features than masked reconstruction. T wo complementary pretext tasks reinforce the learning signals on the factorized tokens: Mixed Fac- tor T oken Distillation (MFTD) guides inference at mixed positions via teacher signals, and Swap-in variance Quadru- plet Contrast (SiQC) enforces within-factor consistency un- der content-style swaps while preserving inter-f actor dis- criminability . The pretrained MeDUET can be seamlessly adapted to both synthesis and analysis tasks, providing a C o nv e r g e nc e Co n t ro lla b ilit y D a t a E f f i c i e nc y D o m a i n Ge n e ra liz a t io n M e D U E T D is t illa t io n C o n t r a st C o n s t ra i n s D i s e nt a ng l e m e nt I d e n t if ia b le D em i xi n g E xis t in g S t r a t e g y Our S t r a t e g y Me di c a l I m a g e S y nt he s i s Me di c a l I m a g e A na l y s i s SSL O b j e c t i ve I n it S e g / Cls C o n d : M e t a d a t a S yn S e g / Cls S yn S y nt he s i s A na l y s i s I n it I n it Co n d A u g M eD UET O b j e c t iv e ( b ) ( a ) Figure 2. (a) Comparison between existing medical image syn- thesis/analysis paradigms and our unified strategy . (b) Overview of our proposed MeDUET . strong weight initialization for both applications (Fig. 2 (b)). Our contributions are summarized as follo ws: • W e propose MeDUET , which, to the best of our knowl- edge, is the first unified pretraining framew ork in medical imaging that supports both synthesis and analysis. • T o bridge the gap in medical imaging, we propose a nov el SSL paradigm within the V AE latent space, employing a disentanglement formulation that aligns the objectiv es of synthesis and analysis, reinforced by a demixing objec- tiv e that encourages factor separation identifiable. • W e design two factor -aw are pretext tasks, MFTD and SiQC, which target mixed-token inference and swap- in variant contrast within the factor spaces, respecti vely , jointly optimizing these complementary objectiv es. • Comprehensive e v aluation demonstrates the superior per- formance of MeDUET in both do wnstream synthesis and analysis tasks across 5 datasets, 4 tasks and 2 modali- ties. For synthesis, it yields substantial improv ements ov er baseline methods in generation fidelity , div ersity , con vergence speed, and controllability , while for analy- sis, it exhibits strong domain generalization and remark- able data efficienc y . 2. Related W ork 3D Medical Image Synthesis. 3D medical image synthe- sis aims to produce synthetic v olumetric data for various clinical applications [ 75 , 93 ]. Adv ances in natural-image diffusion have accelerated progress in this area [ 10 , 36 ], with multiple works sho wing high-fidelity 3D generations D o m a in C la s s if ie r GR L M L P M L P D o ma i n s S t ude n t B r a nc h T eac h er B r an c h ( ) ( ) ( ) ( ) S tude nt & ( ) ( ) ( ) ( ) T ea c h er & EM A EM A C o n t en t S p a c e S t y l e S p a c e S i Q C S t o p G ra d i e n t E MA U p d a t e L e a rn a b l e Fr o z e n T o k e n F o rw a rd F a c t o r F o rw a rd L eg en d T ok e n Mi x i ng T o ken Un m i x i n g D ua l R ec . VA E E nc o de r R a ndo m C r o p M i x i n g & D u a l R e c on s t r u ct i on F a c t o r D i s e n t a n gl e m e n t S i Q C : S w a p - I n v a r i a n ce Q u a d r u p l e t C on t r a s t i v e L e a r n i n g M F TD : M i xe d F a ct or T ok e n D i s t i l l a t i on C on v C on v T o k e n U n m ixin g S tude nt & T ea c h er & EM A S tude n t T ea c h er S tude n t T ea c h er S tude nt & T ea c h er & w / o M ix in g w / o M ix in g w / o M ix in g w / o M ix in g S t ude n t B r a nc h T eac h er B r an c h Mi x e d R e g i o n A lign m e n t M ixin g M a s k M ixin g M a s k M F TD , = { , , , } C o n t r a st i v e L o ss GR L Figure 3. The overall framework of our MeDUET : (a) Mixing & Dual Reconstruction aims to perform demixing between two mixed patch tokens. (b) F actor Disentanglement module explicitly decomposes encoded latent patches into content and style representations, using a domain classifier to empo wer them as domain-in variant and domain-identifiable, respectively . (c) MFTD performs knowledge distillation on the mixed regions within the factor space. (d) SiQC enforces contrastiv e consistency within the factor space, encouraging in v ariance to the swapped factor while preserving the discriminability of the retained one. capturing intricate structures and realistic details [ 24 , 56 ]. Diffusion models ha ve also been applied to di verse clin- ical tasks, including data augmentation [ 69 ], CT recon- struction [ 15 ], and counterfactual generation [ 88 ]. In this work, we propose initializing dif fusion models with pre- trained SSL models in the V AE latent space, which not only accelerates con vergence but also improves synthesis qual- ity . Moreov er , the content and style f actors encoded by the disentangled representation learning in MeDUET en- able promising controllability in generation. 3D Medical Image Analysis. Giv en the scarcity of labeled data in medical imaging, pretraining SSL on large unlabeled 3D data has become a strong paradigm for medical image analysis [ 13 , 39 ]. Most methods follo w contrasti ve learning (CL) [ 30 , 79 ] or masked image modeling (MIM) [ 72 , 96 ]. Previous CL-based approaches rely on strong volumetric augmentations [ 61 ], while mask or remove 3D patches and learn to reconstruct them [ 65 ]. Howe ver , existing methods tend to o veremphasize scanner appearance, entangling con- tent and style [ 2 , 57 , 65 ], which hinders their performance in cross-domain and low-data settings. In contrast, our method employs explicit factor disentanglement to learn ro- bust and domain-consistent semantics, thus improving out- of-distribution performance and data ef ficiency . Diffusion Models f or Repr esentation Learning. Giv en the competitive performance of dif fusion models in vari- ous generation tasks, recent studies ha ve emerged to explore their representational capabilities [ 14 , 35 , 78 ], demonstrat- ing that dif fusion models can serve as ef fecti ve visual learn- ers [ 51 , 71 ]. Other approaches modify dif fusion architec- tures to learn discriminativ e representations for visual per- ception tasks [ 12 , 58 , 82 ]. Howe ver , lar ge-scale applica- tions in medical imaging remain underexplored. Disentangled Representation Learning . Disentangled representation learning aims to train models that can sep- arate interpretable and informativ e factors hidden in ob- served data [ 32 , 45 , 73 ]. Prior works focus on augmenting generativ e models to enhance their disentanglement capa- bility [ 41 , 86 ]. In medical imaging, typical applications include disease decomposition [ 66 ], harmonization [ 81 ], controllable generation [ 8 , 50 , 52 ], and domain generaliza- tion [ 23 , 85 ]. In this paper , our MeDUET is inspired by recent trends of decomposing input volumes into domain- in variant content and domain-specific style [ 6 , 23 , 52 ]. 3. Method The overall framework of MeDUET is presented in Fig. 3 , which is based on the V AE latent space, disentangling content from style and using token demixing as supervi- sion. A student-teacher design supports two auxiliary proxy tasks, MFTD and SiQC, which pro vide targeted guidance to mixed tokens and enforce sw ap-in variant structure. 3.1. Demixing & Dual Reconstruction T o turn multi-center heterogeneity into an explicit super- visory signal and make factor disentanglement practically identifiable, we introduce a token demixing module that constructs and in verts controlled factor mixtures, which are consistent with real clinical shifts. Specifically , gi ven an input volume, we first obtain its latent representation via the frozen V AE tokenizer , and all subsequent operations are performed in latent space during pretraining. T wo sets of la- tent volume patches { z i , z j } are sampled from tw o random volumes within a batch. Follo wing the MIM paradigm, we then generate a mixed patch using a mixing function ϕ m that combines the corresponding visible cubic tokens from z i and z j according to a binary mask M [ 9 , 43 ]. Then the V iT encoder F enc takes the mixed tokens as input for rep- resentation learning. An unmixing function ϕ u restores to- kens to their original spatial positions based on the mixing notation M . The factor disentanglement module F d is then applied, while the lightweight decoder F dec reconstructs the original volumetric patches { ˆ z i , ˆ z j } from the visible tokens. Finally , the dual reconstruction loss is formulated as L r = ∥ ( ˆ z i − z i ) ⊙ M ∥ 2 2 + ∥ ( ˆ z j − z j ) ⊙ (1 − M ) ∥ 2 2 . (1) 3.2. F actor Disentanglement The factor disentanglement module F d decomposes the encoder output into two factors that serve complemen- tary purposes: a domain-in variant content representation c i ∈ R L × D c and a domain-specific style representation s i ∈ R L × D s , where L is the token length. W e implement F d with a lightweight con volutional layer . Domain Classifier . T o further reinforce disentanglement, we introduce domain classifiers to both factor learning pro- cess. The content branch is trained in an adversarial man- ner via a gradient-rev ersal layer (GRL) [ 19 ] so that the en- coder removes domain shift information from c i , whereas the style branch is trained normally to make s i distinguish- able of the domain. Specifically , each branch uses a three- layer MLP domain classifier with a cross-entropy loss: L d ( i ) = − log ([ σ ( G c ( R ( ¯ c i )))] y i ) − log [ σ ( G s ( ¯ s i ))] y ′ i , (2) where y i is the domain label, ¯ c i and ¯ s i denote the factor vec- tors after token-le vel pooling, G c and G s are domain classi- fiers for content and style disentanglement modules, respec- tiv ely , R represents the GRL, and σ denotes the softmax function. W e perform the same operation on z j , yielding L d ( j ) . The overall domain classifier loss is e xpressed as L d = L d ( i ) + L d ( j ) . (3) Then the two factor tokens are integrated via another con volutional layer to obtain aggregated tokens for subse- quent reconstruction process. 3.3. MFTD Intuitiv ely , token mixing introduces information loss, leav- ing mixed regions ambiguous and thereby weakening the constraints on the factor space. T o this end, we de- vise MFTD, which provides factor -specific teacher super- vision by distilling content from the content source and style from the style source to prev ent factor leakage, en- force in variance under incomplete contextual information, and strengthen disentanglement through fine-grained factor - lev el rather than coarse patch-level alignment. Concretely , given the recovered patch tokens ˆ z i and ˆ z j , we re-encode them by feeding them into F enc and F d again without mixing, yielding novel factor tokens { ˆ c i , ˆ s i } and { ˆ c j , ˆ s j } , which denote the full patch factors. Addition- ally , we introduce a teacher model, which is updated via the Exponential Moving A v erage (EMA). For the MFTD teacher network, we obtain { ˆ c T i , ˆ s T i } and { ˆ c T j , ˆ s T j } through an identical re-encoding and re-disentanglement process performed by the teacher model, which provides global con- textual priors to guide the student model. The objectiv e functions are designed to minimize the discrepancy between the teacher and student factor outputs in the mixed re gions: ∆ c i = 1 D c ∥ ˆ c i − sg[ ˆ c T i ] ∥ 2 2 , ∆ c j = 1 D c ∥ ˆ c j − sg[ ˆ c T j ] ∥ 2 2 , (4) L c MFTD = ∥ ∆ c i ⊙ M + ∆ c j ⊙ (1 − M ) ∥ 1 L , (5) where sg [ · ] denotes the stop gradient operation. Notably , the token distances are calculated by averaging across dimen- sions to mitigate the influence of high-dimensional factor representations. By combining it with the similarly com- puted loss of style tokens, the total token distillation loss is defined as the weighted sum: L MFTD = λ c L c MFTD + L s MFTD . (6) 3.4. SiQC Mixing yields paired views that share either content or style. T o make disentanglement effecti ve in practice, a special- ized objective is required to (i) pull together views shar- ing the same content while remaining sensitiv e to style, and (ii) symmetrically , pull together views sharing the same style while remaining sensitive to content, which generic contrasti ve objecti ves f ail to enforce. T o this end, SiQC employs a swap-in v ariant quadruplet objectiv e that pulls together same-content (different-style) and same-style (different-content) pairs while pushing apart cross-factor negati ves, thus explicitly structuring both factor spaces. Content-style swapping. Given the learned factors, we exchange factors between two input volumes, producing { c i , s j } and { c j , s i } , which are then fed into F dec to gener- ate recov ered patches z ij and z j i , respectiv ely . Next, F enc and F d are utilized again for re-encoding without mixing operation, yielding { c ij , s ij } and { c j i , s j i } . The former re- tains the anatomical structure information of z i and domain style characteristics of z j , while the latter does the opposite. Quadruplet construction. Let the two inputs be z i and z j , we form the quadruplet V ( i, j ) = { z i , z j , z ij , z j i } . T aking the content space contrasti ve learning as an example, q c ( a ) and k c ( b ) are defined as the pooled and L2-normalized stu- dent/teacher content features for view a, b ∈ V ( i, j ) . Contrastive loss. Drawing inspiration from [ 40 ], we define the binary positiv e mask as δ c ( z i ) = δ c ( z ij ) = i, δ c ( z j ) = δ c ( z j i ) = j, (7) P c ( a, b ) = 1 { δ c ( a ) = δ c ( b ) } · 1 { a = b } , (8) where δ c ( · ) is the content identity map, and 1 is the indica- tor function. Then the similarity and loss are defined as S c ( a, b ) = exp( α c ) q c ( a ) · sg[ k T c ( b )] , S c ( a, a ) = −∞ , (9) ℓ c ( a, b ) = S c ( a, b ) − log X b ′ ∈V ( i,j ) \{ a } exp( S c ( a, b ′ )) , (10) L c SiQC = E a ∈V ( i,j ) " − P b ∈V ( i,j ) P c ( a, b ) ℓ c ( a, b ) P b ∈V ( i,j ) P c ( a, b ) # , (11) L SiQC = L c SiQC + L s SiQC . (12) where α c is a learnable scale. This objectiv e pulls the pos- itiv e pairs { z i , z ij } and { z j , z j i } together in content space while pushing apart all other pairs. During SiQC, teacher features are stop-grad, while student features and the stu- dent path that produces swapped views remain fully dif fer- entiable, so that SiQC improves both factor encoders and the swap mechanism. The total loss of MeDUET pretraining is formulated as L = L r + λ 1 L d + λ 2 L MFTD + λ 3 L SiQC . (13) where λ i are coefficients to balance loss contrib ution. 3.5. Initialization f or Downstr eam A pplications For downstream generative tasks, we carefully modify the pretrained V iT model to adapt and initialize the DiT/SiT [ 49 , 55 ] models for diffusion-based generation. Follo wing [ 14 ], we reintroduce the shift ( β ) and scale ( γ ) parameters to implement AdaLN-Zero and layer normaliza- tion blocks as conditional input modules. For analysis tasks, we adopt UNETR [ 27 ] as the backbone, which seamlessly inherits the pretrained V iT weights from MeDUET . T o ad- dress inconsistencies in positional embeddings, trilinear in- terpolation is applied to upscale them, ensuring consistency between the patch sizes used during pretraining and those in the downstream phases. 4. Experiments 4.1. Experimental Setup Datasets. T o ensure a fair comparison with pre vious works, we employ the V oCo-10k dataset [ 79 , 80 ] for pretrain- ing, which includes 10 public CT datasets cov ering diverse sources and anatomical regions [ 1 , 16 , 22 , 42 , 48 , 59 , 62 , T able 1. Synthesis performance comparison. The best results are bolded , and the second best results are underlined. † : using pre-defined metadata vectors. ‡ : using learned content and style vectors. 100k/200k: pretrained steps of MeDUET . W e ev aluate SiT B/4 + MeDUET for subsequent experiments, using pretraining weights from 200k steps by default. Method FID ↓ MMD ↓ MS-SSIM ↓ Medical Image Synthesis Models WDM [ 18 ] 0.9668 0.6612 0.2284 MedSyn [ 84 ] 0.9873 0.6734 0.2325 MAISI [ 24 ] 0.9139 0.6292 0.2057 3D MedDiffusion [ 69 ] 0.9216 0.6327 0.2032 General Diffusion Models DiT -B/4 † 0.9207 0.6329 0.1957 DiT -B/4 ‡ 0.9074 0.6175 0.1906 MeDUET 100k/200k † 0.8763 / 0.8727 0.6097 / 0.6074 0.1892 / 0.1876 MeDUET 100k/200k ‡ 0.8642 / 0.8611 0.6028 / 0.6003 0.1824 / 0.1803 SiT -B/4 † 0.8670 0.6023 0.1834 SiT -B/4 ‡ 0.8533 0.6012 0.1798 MeDUET 100k/200k † 0.8039 / 0.8011 0.5806 / 0.5782 0.1712 / 0.1692 MeDUET 100k/200k ‡ 0.7908 / 0.7874 0.5627 / 0.5598 0.1677 / 0.1659 63 , 77 , 97 ], consisting of 10,500 CT scans in total. For downstream experiments, all diffusion models are trained using the V oCo-10k dataset. Additionally , we conduct analysis experiments on fi ve public datasets: BTCV [ 42 ], WORD [ 47 ], AMOS [ 38 ] and BraTS 21 [ 3 ] for segmen- tation tasks, and CC-CCII [ 91 ] for the CO VID-19 classifi- cation task. W e use consistent dataset settings as previous works [ 65 , 79 , 95 , 96 ]. Further details of the datasets can be found in the supplementary materials. Implementation Details. W e employ MAISI-V AE [ 24 ] as the tokenizer to generate latent representations of the in- put v olumes, which are cached to facilitate efficient train- ing and inference. The pretraining process consists of 200k steps, using the AdamW [ 46 ] optimizer with a co- sine learning rate schedule. The isotropic voxel spacing is set to 1.5 mm, and each subv olume patch has a size of 96 × 96 × 96 . W e introduce the MFTD module at 40k steps when the teacher model has been updated stably . W e treat each dataset as a domain because it typically reflects a distinct center/protocol/scanner and thus serves as a sta- ble proxy for style shifts. W e generate content and style vectors for entire volumes using the pretrained MeDUET weights. For synthesis, we condition DiT/SiT on the con- tent and style, replacing coarse metadata with fine-grained control. For analysis, we use the style vector to dri ve label- preserving style augmentation. Additional implementation details are provided in the supplementary materials. 4.2. Medical Image Synthesis W e compare MeDUET with SO T A 3D medical image syn- thesis methods [ 18 , 24 , 69 , 84 ]. Furthermore, we inte- grate our pretrained model into two diffusion transformers: DiT [ 55 ] and SiT [ 49 ], which we follo w the training settings by default. W e reimplemented all baseline methods from scratch using the pretraining dataset. Notably , we lev er - Me D UET ( O ur s ) 3D M e dD i f f us i o n M A I S I M e d S yn W DM Figure 4. Qualitative comparison of synthesized v olumes. 100k 300k 500k 800k 1M 1.5M 2M T raining Steps 0.85 0.90 0.95 1.00 FID 9.3 × Speedup Convergence Speed (DiT) D i T - B / 4 D i T - B / 4 M e D U E T M e D U E T 100k 300k 500k 800k 1M 1.5M 2M T raining Steps 0.78 0.80 0.85 0.90 0.95 FID 8.4 × Speedup Convergence Speed (SiT) S i T - B / 4 S i T - B / 4 M e D U E T M e D U E T Figure 5. Conver gence speed comparison. Left: Con ver gence acceleration for DiT . Right: Con ver gence acceleration for SiT . † : using pre-defined metadata. ‡ : using learned content and style. age the learned content and style vectors as condition with classifier-free guidance (CFG) [ 33 ] instead of pre-defined metadata v ectors (e.g. body region, vox el spacing). Evalua- tions are conducted on the generated 1k volumes. For ev al- uation metrics, following [ 69 ], Fr ´ echet Inception Distance (FID) [ 31 ] and Maximum Mean Discrepancy (MMD) [ 21 ] are used to assess the fidelity and realism of the synthetic images, while Multi-Scale Structural Similarity Index (MS- SSIM) [ 76 ] is employed to ev aluate the div ersity . For FID and MMD ev aluation, we use the MedicalNet 3D [ 11 ] as the feature extractor to compute whole-volume 3D features per scan. Synthesis Quality Comparison. W e exclude any im- prov ements to DiT and SiT , such as architectural mod- ifications [ 20 , 94 ] and acceleration techniques [ 87 , 89 ], which are not line with our core objecti ve. From T able 1 , MeDUET achie ves SO T A performance across all three met- rics, with FID scores of 0.7874 and 0.8611 for SiT and DiT , respecti vely , and achiev es lo wer MMD and MS-SSIM values than all baselines, indicating superior anatomical fi- delity and appearance div ersity . The learned disentangled factors consistently outperform the previously used meta- data v ectors, emphasizing the ef fecti veness of our disentan- gled factor learning. Fig. 4 presents a visual comparison of images synthesized by our method and baselines. Con vergence Acceleration. Fig. 5 displays the training progress for DiT , SiT , and MeDUET . With MeDUET ini- tialization, DiT achiev es a 9.3 × faster FID con ver gence speed and SiT achiev es an 8.4 × speedup. The gains grow 1 2 3 4 5 Figure 6. Performance comparison of data augmentation . † : us- ing pre-defined metadata vectors. ‡ : using learned content and style vectors. S 1 : 100% synthetic data. S 2 : 50% real data and 50% synthetic data. S 3 : 100% real data. S 4 : 100% real data and 50% synthetic data. S 5 : 100% real data and 100% synthetic data. c s D c s C Figure 7. Content and style consistency surface. w c , w s : CFG scales for content and style. R D : Dice ratio between the seg- mentations of the reference and the generated volumes; a higher value indicates greater anatomical similarity and thus higher con- tent consistency . R C : Hit rate of the style classifier on the gener- ated volumes; a higher value reflects greater style similarity , i.e., higher style consistency . with learned f actor and longer pretraining, confirming the effecti veness of our disentanglement and initialization. Conditional Generation f or Data A ugmentation. Fol- lowing [ 24 , 69 ], we integrate ControlNet [ 64 , 92 ] and con- dition the diffusion model on segmentation masks to im- prov e controllability . Subsequently , the synthesized vol- umes are employed as augmented data to enhance segmen- tation performance. W e train nnU-Net [ 37 ] on T otalSeg- mentator [ 77 ] under five training regimes. As illustrated in Fig. 6 , our MeDUET -guided, controllable synthesis con- sistently surpasses baselines across the five augmentation settings. Disentangled factors further boost segmentation, indicating anatomy-preserving content control and style- aware di versification, and validating our disentangled, con- trollable generation framew ork. Content and Style Consistency . T o assess control- lable generation without manual latent edits, we use dual- conditional CFG [ 33 ], independently scaling content w c and style w s during sampling. For content consistency , we fix T otalSe gmentator [ 77 ] content references, sample styles from other domains, sweep ( w c , w s ) , and compute the Dice ratio R D between segmentations of the references and the T able 2. Results on the segmentation task under different data proportions. The third row denotes the data ratios for training. W e report the mean Dice score (%). The best results are bolded , and the second best results are underlined. Method Dice Score (%) ↑ BTCV AMOS WORD BraTS 21 1-shot 10% 50% 100% A vg 1-shot 10% 50% 100% A vg 1-shot 10% 50% 100% A vg 1-shot 10% 50% 100% A vg F rom Scratch UNETR [ 27 ] 68.19 78.30 87.28 92.33 81.53 60.29 86.82 93.54 94.62 83.82 62.48 90.76 92.87 94.03 85.03 51.91 77.41 84.56 85.24 74.78 Swin UNETR [ 26 ] 71.28 78.54 88.06 92.65 82.63 63.18 87.26 93.80 95.29 84.88 64.39 91.05 92.93 94.01 85.59 55.82 79.56 85.42 86.15 76.74 General SSL MAE3D [ 13 ] 65.95 76.71 85.81 91.28 79.94 57.39 85.27 92.63 94.25 82.38 58.96 90.24 92.68 94.06 83.98 54.57 78.86 85.29 86.19 76.23 SimMIM [ 83 ] 65.42 76.08 84.93 90.89 79.33 57.82 85.06 92.48 93.94 82.33 57.75 90.05 92.51 93.81 83.53 52.39 77.63 85.12 85.54 75.17 MoCo v3 [ 28 ] 66.49 77.86 86.57 91.83 80.69 59.60 86.29 93.37 94.67 83.48 57.83 90.22 92.46 93.92 83.61 52.20 77.72 85.24 85.76 75.23 Medical Image SSL Swin UNETR [ 26 ] 72.87 80.42 89.93 92.96 84.05 63.01 89.75 93.52 94.69 85.24 66.85 91.13 93.26 94.03 86.32 56.93 79.70 85.58 86.62 77.21 SwinMM [ 74 ] 72.28 81.97 91.36 93.58 84.80 63.56 89.24 93.75 94.84 85.35 74.65 91.39 93.48 94.19 88.43 57.12 79.51 85.40 86.73 77.19 GVSL [ 30 ] 74.92 83.75 91.41 93.43 85.88 63.70 89.72 93.68 94.97 85.52 76.92 91.30 93.39 94.12 88.93 54.79 80.20 85.61 86.92 76.88 V oCo [ 79 ] 73.27 84.15 90.20 93.62 85.31 62.75 89.84 94.07 95.37 85.51 70.90 91.47 93.51 94.24 87.53 57.30 80.71 85.77 87.09 77.72 GL-MAE [ 95 ] 69.29 79.63 89.61 93.07 82.90 61.93 87.52 93.70 95.14 84.57 65.93 91.66 93.13 94.16 86.22 54.98 79.38 84.42 86.75 76.38 MIM [ 96 ] 71.86 81.05 90.23 93.14 84.07 61.84 88.68 94.05 95.03 84.90 71.49 91.91 93.30 94.02 87.68 56.36 79.06 84.82 86.84 76.77 Hi-End-MAE [ 65 ] 72.45 83.02 90.80 93.19 84.86 63.23 90.03 94.16 95.76 85.80 77.24 92.24 93.65 94.34 89.37 54.46 80.23 85.71 87.33 76.93 MeDUET 78.72 87.04 92.24 94.06 88.02 65.18 90.36 94.34 95.48 86.34 79.56 92.61 93.83 94.28 90.07 58.05 80.83 85.82 87.39 78.02 T able 3. Results on the classification task on the CC-CCII dataset under different data proportions. The second row denotes the data ratios for training. W e report the Accurac y (%). Method Accuracy (%) ↑ 10% 50% 100% A vg F rom Scratch UNETR [ 27 ] 73.80 82.40 88.65 81.62 Swin UNETR [ 26 ] 77.01 86.62 88.32 83.98 Medical Image SSL Swin-UNETR [ 26 ] 77.64 87.33 89.42 84.80 SwinMM [ 74 ] 81.49 91.27 92.23 88.33 GVSL [ 30 ] 86.16 91.08 93.26 90.17 V oCo [ 79 ] 86.85 91.86 93.45 90.72 GL-MAE [ 95 ] 80.63 88.62 92.13 87.13 MIM [ 96 ] 82.57 89.74 92.75 88.35 Hi-End-MAE [ 65 ] 78.76 88.15 92.59 86.50 MeDUET 88.68 91.79 93.59 91.35 generated volumes. For style consistency , we fix style ref- erences and report the style-classifier hit rate R C and its monotonicity with w s . W e use nnU-Net [ 37 ] as the seg- menter and MeDUET’ s domain classifier as the style classi- fier . As sho wn in Fig. 7 , R D increases with w c and quickly saturates near 1 across w s , while R C rises strictly with w s and is insensitiv e to w c . Thus, content tokens remain geometrically stable and style tokens control appearance, demonstrating MeDUET’ s impressiv e controllability . 4.3. Medical Image Analysis W e validate MeDUET with both General [ 13 , 28 , 29 , 83 ] and Medical SSL [ 26 , 27 , 30 , 65 , 74 , 79 , 95 , 96 ] on analy- sis tasks for a thorough comparison. The experimental re- sults are reported using official pretrained model weights for Medical SSL methods. Superior Performance for Cross-Domains and Data Ef- ficiency . W e fine-tune pretrained models for segmentation tasks on four datasets with v arying proportions of labeled data, and the results are summarized in T able 2 . AMOS and WORD correspond to unseen domains during pretraining, T able 4. Synthesis performance comparison with 3D medical image SSL in the latent space. † : using pre-defined metadata vectors. ‡ : using learned content and style vectors. Method FID ↓ MMD ↓ MS-SSIM ↓ SiT -B/4 + MAE3D † [ 13 ] 0.8416 0.5972 0.1871 SiT -B/4 + MAE3D ‡ [ 13 ] 0.8265 (-0.0151) 0.5816 (-0.0156) 0.1796 (-0.0075) SiT -B/4 + GL-MAE † [ 95 ] 0.8322 0.5947 0.1803 SiT -B/4 + GL-MAE ‡ [ 95 ] 0.8147 (-0.0175) 0.5803 (-0.0144) 0.1762 (-0.041) SiT -B/4 + Hi-End-MAE † [ 65 ] 0.8189 0.5846 0.1773 SiT -B/4 + Hi-End-MAE ‡ [ 65 ] 0.8094 (-0.0095) 0.5785 (-0.0061) 0.1736 (-0.0037) SiT -B/4 + MeDUET † 0.8011 0.5782 0.1692 SiT -B/4 + MeDUET ‡ 0.7874 (-0.0137) 0.5598 (-0.0184) 0.1659 (-0.0033) while BraTS 21 represents an unseen modality (from CT to MRI). MeDUET consistently impro v es Dice across do- mains and label ratios, while prior methods are less stable and underperform with unseen domains or limited labels. The robustness and data efficiency of MeDUET can be at- tributed to its strong capability to learn in variant anatomical representations that are robust to domain v ariations. Remarkable One-shot Segmentation Perf ormance. From T able 2 , under the one-shot segmentation setting, MeDUET surpasses the strongest SSL baseline by an av erage of 2.09% Dice Scores across four 3D medical benchmarks. These results suggest that the factor disen- tanglement and demixing mechanism in MeDUET enables more label-efficient representations. Strong Generalization Capability for MRI data. As shown in T able 2 , MeDUET outperforms existing SO T A methods on the BraTS 21 dataset under various data propor - tions, demonstrating the promising generalization ability of our MeDUET for unseen modalities. Classification T asks. From T able 3 , our MeDUET yields outperformance over SO T A baselines for the classification task on the CC-CCII dataset, which achieves a 0.63% im- prov ement of accuracy in av erage under various data ratios. T able 5. Ablation r esults on loss functions and model compo- nents. w/o Disent.: Factor disentanglement is remov ed by exclud- ing L d and L SiQC , and MFTD is modified to perform token-le vel distillation. w/o Demix.: The demixing module is disabled, result- ing in a vanilla mask ed reconstruction paradigm. Method Synthesis Analysis FID ↓ MMD ↓ MS-SSIM ↓ Dice Score (%) ↑ BTCV WORD 1-shot 100% 1-shot 100% Loss Functions w/o L d 0.8147 0.5836 0.1758 73.86 93.34 72.18 94.14 w/o L MFTD 0.7916 0.5623 0.1689 77.43 93.81 78.62 94.17 w/o L SIQC 0.7929 0.5619 0.1696 77.94 93.78 79.30 94.21 Components w/o Disent. 0.8460 0.5924 0.1802 71.63 93.28 70.41 94.06 w/o Demix. 0.7962 0.5677 0.1713 78.09 93.92 79.27 94.19 MeDUET 0.7874 0.5598 0.1659 78.72 94.06 79.56 94.28 T able 6. Linear probe classification results of learned content and style codes. W e report the Accurac y (%) and A UC (%). Method Content Style Accuracy (%) ↓ A UC (%) ↓ Accuracy (%) ↑ AUC (%) ↑ w/o L d 57.53 75.48 51.46 71.52 w/o L MFTD 24.20 63.21 96.07 93.11 w/o L SIQC 28.77 68.98 92.04 90.23 w/o Demix. 23.95 62.92 96.18 93.26 MeDUET 23.59 61.56 96.43 93.59 4.4. Medical SSL f or Generative T ransfer T o isolate the contribution of our disentangling objec- tiv es, we reimplement representative 3D V iT -based medi- cal SSL [ 13 , 65 , 95 ] within the same V AE latent space and transfer their encoders to initialize SiT . Under matched ex- perimental protocols, we compare their synthesis quality , as reported in T able 4 . As sho wn, MeDUET surpasses base- lines transferred within the identical latent space. Further- more, the introduction of our disentangled factors leads to varying degrees of improvement for latent medical SSL. Crucially , with setups all held fixed, the consistent gains across three metrics isolate the benefit of our disentangling components rather than model capacity or data advantages. T ak en together , these results establish MeDUET as a more transferable and ef fecti ve pretraining paradigm for 3D med- ical imaging than latent-space SSL alone. 4.5. Ablation Study Loss Functions and Components. From T able 5 , remov- ing any single loss consistently worsens synthesis, and dis- abling disentanglement leads to the largest few-shot Dice drop. W ith all components enabled, MeDUET achiev es the best synthesis and highest segmentation Dice, indicat- ing these components are complementary and essential for both generativ e quality and segmentation generalization. Factor Identifiability . In this section, we analyze the ef fec- tiv eness of disentanglement both quantitativ ely and qual- Figure 8. The t-SNE visualization of MeDUET content and style codes, where points are color-coded according to their domain la- bels. Left: Content distribution. Right: Style distribution. itativ ely by inv estigating factor identifiability empirically: the content vector should be domain-in v ariant and suffi- cient for downstream tasks, while the style vector should be domain-specific and task-independent. W e first conduct linear-probe classification experiments using the learned factors as input and domain categories as labels. As re- ported in T able 6 , the linear probe trained on style vectors achiev es high domain sensitivity than variants, confirming that the style branch effecti vely captures more protocol- specific cues. Con versely , the probe trained on content vectors yields lower domain discriminative that is approxi- mately random classification, indicating lower domain leak- age and more effecti ve factor disentanglement. In addition, we visualize the t-SNE of factor codes learned by MeDUET and its variant w/o Demix. As shown in Fig. 8 , the content vectors learned by MeDUET ex- hibit an entangled and interspersed distribution across do- main labels, with a lower silhouette score than the v ariant w/o Demix., v erifying its ability to extract domain-inv ariant knowledge. In contrast, the style representations display better domain alignment, with clear and distinct boundaries and a higher silhouette score, indicating their capability to capture domain-specific information. These results further justify the rationale behind our demixing module, which f a- cilitates more disentangled and identifiable f actor learning. 5. Conclusion In this work, we present MeDUET , a unified pretraining framew ork for 3D medical imaging that learns identifi- able content-style factors within a V AE latent space and transfers seamlessly to both generativ e and analytical tasks. Our design explicitly enforces disentanglement so that content governs anatomical structure while style captures domain attributes. Extensiv e experiments demon- strate that MeDUET achiev es superior performance for versatile synthesis and analysis applications. By un- leashing the potential of SSL for both medical image generation and understanding, we believe MeDUET provides valuable insights that can guide future re- search on pretraining strategies for 3D medical imaging. References [1] Samuel G. Armato III, Geoffrey McLennan, Luc Bidaut, Michael F . McNitt-Gray , Charles R. Meyer , Anthony P . Reev es, Binsheng Zhao, Denise R. Aberle, Claudia I. Hen- schke, Eric A. Hoffman, Ella A. Kazerooni, Heber MacMa- hon, Edwin J. R. van Beek, David Y ankelevitz, Alberto M. Biancardi, Peyton H. Bland, Matthe w S. Brown, Roger M. Engelmann, Gary E. Laderach, Daniel Max, Richard C. Pais, David P .-Y . Qing, Rachael Y . Roberts, Amanda R. Smith, Adam Starkey , Poonam Batra, Philip Caligiuri, Ali Farooqi, Gregory W . Gladish, C. Matilda Jude, Reginald F . Munden, Iva Petkovska, Leslie E. Quint, Lawrence H. Schwartz, Baskaran Sundaram, Lori E. Dodd, Charles Fenimore, David Gur , Nicholas Petrick, John Freymann, Justin Kirby , Brian Hughes, Alessi V ande Casteele, Sangeeta Gupte, Maha Sal- lam, Michael D. Heath, Michael H. Kuhn, Ekta Dharaiya, Richard Burns, David S. Fryd, Marcos Salganicoff, V ikram Anand, Uri Shreter, Stephen V astagh, Barbara Y . Croft, and Laurence P . Clarke. The lung image database consortium (lidc) and image database resource initiativ e (idri): A com- pleted reference database of lung nodules on ct scans. Medi- cal Physics , 38(2):915–931, 2011. 5 , 1 , 2 [2] Mahmoud Assran, Mathilde Caron, Ishan Misra, Piotr Bo- janowski, Florian Bordes, Pascal V incent, Armand Joulin, Mike Rabbat, and Nicolas Ballas. Masked siamese networks for label-ef ficient learning. In Eur opean conference on com- puter vision , pages 456–473, 2022. 3 [3] Ujjwal Baid, Satyam Ghodasara, Suyash Mohan, Michel Bilello, Evan Calabrese, Errol Colak, Keyv an Farahani, Jayashree Kalpathy-Cramer, Felipe C Kitamura, Sarthak Pati, et al. The rsna-asnr-miccai brats 2021 benchmark on brain tumor segmentation and radiogenomic classification. arXiv pr eprint arXiv:2107.02314 , 2021. 5 , 2 [4] Qi Bi, Jingjun Y i, Hao Zheng, W ei Ji, Y a wen Huang, Y uex- iang Li, and Y efeng Zheng. Learning generalized medical image representation by decoupled feature queries. IEEE T ransactions on P attern Analysis and Machine Intelligence , pages 1–18, 2025. 1 , 2 [5] Manh-Ha Bui, T oan Tran, Anh Tran, and Dinh Phung. Ex- ploiting domain-specific features to enhance domain gener- alization. In Advances in Neural Information Processing Sys- tems , pages 21189–21201, 2021. 2 [6] Zhuotong Cai, Jingmin Xin, Chenyu Y ou, Peiwen Shi, Siyuan Dong, Nicha C. Dvornek, Nanning Zheng, and James S. Duncan. Style mixup enhanced disentanglement learning for unsupervised domain adaptation in medical im- age segmentation. Medical Image Analysis , 101:103440, 2025. 3 [7] Mathilde Caron, Hugo T ouvron, Ishan Misra, Herv ´ e J ´ egou, Julien Mairal, Piotr Bojanowski, and Armand Joulin. Emerg- ing properties in self-supervised vision transformers. In Pr oceedings of the IEEE/CVF International Confer ence on Computer V ision (ICCV) , pages 9650–9660, 2021. 1 [8] Zengyang Che, Zheng Zhang, Y aping W u, and Meiyun W ang. Disentangle and then fuse: A cross-modal net- work for synthesizing gadolinium-enhanced brain mr im- ages. IEEE T ransactions on Circuits and Systems for V ideo T echnology , 35(6):6021–6033, 2025. 3 [9] Kai Chen, Zhili Liu, Lanqing Hong, Hang Xu, Zhen- guo Li, and Dit-Y an Y eung. Mixed autoencoder for self- supervised visual representation learning. In Proceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 22742–22751, 2023. 4 [10] Qi Chen, Xiaoxi Chen, Haorui Song, Zhiwei Xiong, Alan Y uille, Chen W ei, and Zongwei Zhou. T ow ards generalizable tumor synthesis. In Pr oceedings of the IEEE/CVF Confer- ence on Computer V ision and P attern Recognition (CVPR) , pages 11147–11158, 2024. 2 [11] Sihong Chen, Kai Ma, and Y efeng Zheng. Med3d: Trans- fer learning for 3d medical image analysis. arXiv preprint arXiv:1904.00625 , 2019. 6 [12] Xinlei Chen, Zhuang Liu, Saining Xie, and Kaiming He. De- constructing denoising diffusion models for self-supervised learning. In The Thirteenth International Confer ence on Learning Repr esentations , 2025. 3 [13] Zekai Chen, Dev ansh Agarwal, Kshitij Aggarwal, Wiem Safta, Mariann Micsinai Balan, and Ke vin Brown. Masked image modeling advances 3d medical image analysis. In Pro- ceedings of the IEEE/CVF W inter Confer ence on Applica- tions of Computer V ision (W ACV) , pages 1970–1980, 2023. 3 , 7 , 8 , 1 [14] Xiangxiang Chu, Renda Li, and Y ong W ang. Usp: Uni- fied self-supervised pretraining for image generation and understanding. In Pr oceedings of the IEEE/CVF Inter- national Confer ence on Computer V ision (ICCV) , pages 18475–18486, 2025. 1 , 3 , 5 [15] Hyungjin Chung, Suhyeon Lee, and Jong Chul Y e. De- composed diffusion sampler for accelerating large-scale in- verse problems. In The T welfth International Conference on Learning Repr esentations , 2024. 3 [16] Kenneth Clark, Bruce V endt, Kirk Smith, John Freymann, Justin Kirby , Paul K oppel, Stephen Moore, Stanley Phillips, David Maffitt, Michael Pringle, et al. The cancer imag- ing archiv e (tcia): maintaining and operating a public infor- mation repository . Journal of digital imaging , 26(6):1045– 1057, 2013. 5 , 1 , 2 [17] Prafulla Dhariwal and Alexander Nichol. Diffusion models beat gans on image synthesis. In Advances in Neural Infor- mation Pr ocessing Systems , pages 8780–8794, 2021. 1 [18] Paul Friedrich, Julia W olleb, Florentin Bieder, Alicia Durrer , and Philippe C Cattin. Wdm: 3d wa velet diffusion mod- els for high-resolution medical image synthesis. In MICCAI workshop on deep g enerative models , pages 11–21. Springer , 2024. 5 [19] Y arosla v Ganin, Evgeniya Ustinov a, Hana Ajakan, Pas- cal Germain, Hugo Larochelle, Franc ¸ ois La violette, Mario March, and V ictor Lempitsky . Domain-adversarial training of neural networks. Journal of Machine Learning Researc h , 17(59):1–35, 2016. 4 [20] Shanghua Gao, Pan Zhou, Ming-Ming Cheng, and Shuicheng Y an. Masked dif fusion transformer is a strong image synthesizer . In Pr oceedings of the IEEE/CVF In- ternational Conference on Computer V ision (ICCV) , pages 23164–23173, 2023. 6 [21] Arthur Gretton, Karsten M. Borgwardt, Malte J. Rasch, Bernhard Sch ¨ olkopf, and Alexander Smola. A kernel two- sample test. J. Mach. Learn. Res. , 13(null):723–773, 2012. 6 [22] Aaron J Grossberg, Abdallah SR Mohamed, Hesham Elha- lawani, William C Bennett, Kirk E Smith, T racy S Nolan, Bowman Williams, Sasikarn Chamchod, Jolien Heukelom, Michael E Kantor , et al. Imaging and clinical data archive for head and neck squamous cell carcinoma patients treated with radiotherapy . Scientific data , 5(1):1–10, 2018. 5 , 1 , 2 [23] Ran Gu, Guotai W ang, Jiangshan Lu, Jingyang Zhang, W en- hui Lei, Y inan Chen, W enjun Liao, Shichuan Zhang, Kang Li, Dimitris N. Metaxas, and Shaoting Zhang. Cddsa: Con- trastiv e domain disentanglement and style augmentation for generalizable medical image segmentation. Medical Image Analysis , 89:102904, 2023. 2 , 3 [24] Pengfei Guo, Can Zhao, Dong Y ang, Ziyue Xu, V ish- wesh Nath, Y ucheng T ang, Benjamin Simon, Mason Belue, Stephanie Harmon, Baris T urkbey , and Daguang Xu. Maisi: Medical ai for synthetic imaging. In 2025 IEEE/CVF W in- ter Confer ence on Applications of Computer V ision (W ACV) , pages 4430–4441, 2025. 3 , 5 , 6 , 1 , 2 , 4 [25] Ibrahim Ethem Hamamci, Sezgin Er, Anjan y Sekuboy- ina, Enis Simsar , Alperen T ezcan, A yse Gulnihan Sim- sek, Sevval Nil Esirgun, Furkan Almas, Irem Do ˘ gan, Muhammed Furkan Dasdelen, et al. Generatect: T ext- conditional generation of 3d chest ct volumes. In Eur opean Confer ence on Computer V ision , pages 126–143, 2024. 1 [26] Ali Hatamizadeh, V ishwesh Nath, Y ucheng T ang, Dong Y ang, Holger R Roth, and Daguang Xu. Swin unetr: Swin transformers for semantic se gmentation of brain tumors in mri images. In International MICCAI brainlesion workshop , pages 272–284. Springer , 2021. 7 [27] Ali Hatamizadeh, Y ucheng T ang, V ishwesh Nath, Dong Y ang, Andriy Myronenko, Bennett Landman, Holger R. Roth, and Daguang Xu. Unetr: Transformers for 3d medical image segmentation. In Pr oceedings of the IEEE/CVF W in- ter Confer ence on Applications of Computer V ision (W ACV) , pages 574–584, 2022. 5 , 7 , 4 [28] Kaiming He, Haoqi Fan, Y uxin W u, Saining Xie, and Ross Girshick. Momentum contrast for unsupervised visual repre- sentation learning. In Pr oceedings of the IEEE/CVF Confer- ence on Computer V ision and P attern Recognition (CVPR) , 2020. 1 , 7 [29] Kaiming He, Xinlei Chen, Saining Xie, Y anghao Li, Piotr Doll ´ ar , and Ross Girshick. Masked autoencoders are scalable vision learners. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 16000–16009, 2022. 1 , 7 [30] Y uting He, Guanyu Y ang, Rongjun Ge, Y ang Chen, Jean- Louis Coatrieux, Boyu W ang, and Shuo Li. Geometric vi- sual similarity learning in 3d medical image self-supervised pre-training. In Proceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 9538–9547, 2023. 3 , 7 , 2 [31] Martin Heusel, Hubert Ramsauer , Thomas Unterthiner , Bernhard Nessler, and Sepp Hochreiter . Gans trained by a two time-scale update rule conv erge to a local nash equilib- rium. In Advances in Neural Information Pr ocessing Sys- tems , 2017. 6 [32] Irina Higgins, David Amos, David Pfau, Sebastien Racaniere, Loic Matthey , Danilo Rezende, and Alexander Lerchner . T o wards a definition of disentangled representa- tions. arXiv preprint , 2018. 3 [33] Jonathan Ho and Tim Salimans. Classifier-free diffusion guidance. arXiv preprint , 2022. 6 , 2 [34] Jonathan Ho, Ajay Jain, and Pieter Abbeel. Denoising dif- fusion probabilistic models. In Advances in Neural Informa- tion Pr ocessing Systems , pages 6840–6851, 2020. 1 [35] Drew A. Hudson, Daniel Zoran, Mateusz Malinowski, An- drew K. Lampinen, Andre w Jae gle, James L. McClelland, Loic Matthey , Felix Hill, and Alexander Lerchner . Soda: Bottleneck diffusion models for representation learning. In Pr oceedings of the IEEE/CVF Conference on Computer V i- sion and P attern Recognition (CVPR) , pages 23115–23127, 2024. 3 [36] Mahmoud Ibrahim, Y asmina Al Khalil, Sina Amirrajab, Chang Sun, Marcel Breeuwer , Josien Pluim, Bart Elen, G ¨ okhan Ertaylan, and Michel Dumontier . Generative ai for synthetic data across multiple medical modalities: A system- atic re vie w of recent de velopments and challenges. Comput- ers in Biology and Medicine , 189:109834, 2025. 2 [37] Fabian Isensee, Jens Petersen, Andre Klein, David Zim- merer , P aul F Jae ger , Simon K ohl, Jakob W asserthal, Gregor K oehler , T obias Norajitra, Sebastian W irkert, et al. nnu-net: Self-adapting framework for u-net-based medical image seg- mentation. arXiv preprint , 2018. 6 , 7 [38] Y uanfeng Ji, Haotian Bai, Chongjian GE, Jie Y ang, Y e Zhu, Ruimao Zhang, Zhen Li, Lingyan Zhanng, W anling Ma, Xi- ang W an, and Ping Luo. Amos: A lar ge-scale abdominal multi-organ benchmark for versatile medical image se gmen- tation. In Advances in Neural Information Processing Sys- tems , pages 36722–36732, 2022. 5 , 2 [39] Y ankai Jiang, Mingze Sun, Heng Guo, Xiaoyu Bai, Ke Y an, Le Lu, and Minfeng Xu. Anatomical inv ariance modeling and semantic alignment for self-supervised learning in 3d medical image analysis. In Pr oceedings of the IEEE/CVF International Conference on Computer V ision (ICCV) , pages 15859–15869, 2023. 3 [40] Prannay Khosla, Piotr T eterwak, Chen W ang, Aaron Sarna, Y onglong Tian, Phillip Isola, Aaron Maschinot, Ce Liu, and Dilip Krishnan. Supervised contrastive learning. In Advances in Neural Information Pr ocessing Systems , pages 18661–18673, 2020. 5 [41] Hyunjik Kim and Andriy Mnih. Disentangling by factoris- ing. In Pr oceedings of the 35th International Conference on Machine Learning , pages 2649–2658. PMLR, 2018. 3 [42] Bennett Landman, Zhoubing Xu, Juan Igelsias, Martin Styner , Thomas Langerak, and Arno Klein. Miccai multi- atlas labeling beyond the cranial vault–workshop and chal- lenge. In Pr oc. MICCAI multi-atlas labeling beyond cra- nial vault—workshop c hallenge , page 12. Munich, Germany , 2015. 5 , 1 , 2 [43] Jihao Liu, Xin Huang, Jinliang Zheng, Y u Liu, and Hong- sheng Li. Mixmae: Mixed and masked autoencoder for effi- cient pretraining of hierarchical vision transformers. In Pr o- ceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 6252–6261, 2023. 4 [44] Xiao Liu, Pedro Sanchez, Spyridon Thermos, Alison Q. O’Neil, and Sotirios A. Tsaftaris. Learning disentangled rep- resentations in the imaging domain. Medical Image Analysis , 80:102516, 2022. 2 [45] Francesco Locatello, Stefan Bauer , Mario Lucic, Gunnar Raetsch, Sylvain Gelly , Bernhard Sch ¨ olkopf, and Olivier Bachem. Challenging common assumptions in the unsuper- vised learning of disentangled representations. In Pr oceed- ings of the 36th International Conference on Machine Learn- ing , pages 4114–4124. PMLR, 2019. 3 [46] Ilya Loshchilov and Frank Hutter . Decoupled weight de- cay regularization. In International Conference on Learning Repr esentations , 2019. 5 [47] Xiangde Luo, W enjun Liao, Jianghong Xiao, Jieneng Chen, T ao Song, Xiaofan Zhang, Kang Li, Dimitris N. Metaxas, Guotai W ang, and Shaoting Zhang. W ord: A large scale dataset, benchmark and clinical applicable study for abdom- inal organ segmentation from ct image. Medical Image Anal- ysis , 82:102642, 2022. 5 , 2 [48] Jun Ma, Y ao Zhang, Song Gu, Cheng Ge, Ershuai W ang, Qin Zhou, Ziyan Huang, Pengju L yu, Jian He, and Bo W ang. Au- tomatic organ and pan-cancer segmentation in abdomen ct: the flare 2023 challenge. arXiv pr eprint arXiv:2408.12534 , 2024. 5 , 1 , 2 [49] Nanye Ma, Mark Goldstein, Michael S Albergo, Nicholas M Boffi, Eric V anden-Eijnden, and Saining Xie. Sit: Explor- ing flow and dif fusion-based generati ve models with scalable interpolant transformers. In Eur opean Conference on Com- puter V ision , pages 23–40. Springer , 2024. 5 , 2 [50] Y e Mao, Lan Jiang, Xi Chen, and Chao Li. Disc-diff: Disentangled conditional diffusion model for multi-contrast mri super-resolution. In International Conference on Med- ical Image Computing and Computer-Assisted Intervention , pages 387–397. Springer , 2023. 3 [51] Soumik Mukhopadhyay , Matthe w Gwilliam, Y osuk e Y am- aguchi, V atsal Agarwal, Namitha Padmanabhan, Archana Swaminathan, Tianyi Zhou, Jun Ohya, and Abhinav Shri- vasta va. Do text-free dif fusion models learn discriminativ e visual representations? In Eur opean Conference on Com- puter V ision , pages 253–272, 2024. 3 [52] Sarah M ¨ uller , Lisa M. Koch, Hendrik P .A. Lensch, and Philipp Berens. Disentangling representations of retinal im- ages with generative models. Medical Image Analysis , 105: 103628, 2025. 2 , 3 [53] Maxime Oquab, Timoth ´ ee Darcet, Th ´ eo Moutakanni, Huy V . V o, Marc Szafraniec, V asil Khalidov , Pierre Fernandez, Daniel HAZIZA, Francisco Massa, Alaaeldin El-Nouby , Mido Assran, Nicolas Ballas, W ojciech Galuba, Russell Howes, Po-Y ao Huang, Shang-W en Li, Ishan Misra, Michael Rabbat, V asu Sharma, Gabriel Synnaev e, Hu Xu, Herve Je- gou, Julien Mairal, Patrick Labatut, Armand Joulin, and Pi- otr Bojanowski. DINOv2: Learning robust visual features without supervision. T ransactions on Mac hine Learning Re- sear ch , 2024. Featured Certification. 1 [54] Mathias ¨ Ottl, Frauke Wilm, Jana Steenpass, Jingna Qiu, Matthias R ¨ ubner , Arndt Hartmann, Matthias Beckmann, Pe- ter F asching, Andreas Maier , Ramona Erber , et al. Style- extracting dif fusion models for semi-supervised histopathol- ogy segmentation. In Eur opean Conference on Computer V ision , pages 236–252, 2024. 2 [55] William Peebles and Saining Xie. Scalable diffusion mod- els with transformers. In Proceedings of the IEEE/CVF In- ternational Conference on Computer V ision (ICCV) , pages 4195–4205, 2023. 5 , 2 , 3 [56] Kunpeng Qiu, Zhiqiang Gao, Zhiying Zhou, Mingjie Sun, and Y ongxin Guo. Noise-consistent siamese-diffusion for medical image synthesis and segmentation. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P at- tern Recognition (CVPR) , pages 15672–15681, 2025. 3 [57] Zelin Qiu, Xi W ang, Zhuoyao Xie, Juan Zhou, Y u W ang, Lingjie Y ang, Xinrui Jiang, Juyoung Bae, Moo Hyun Son, Qiang Y e, et al. Large-scale multi-sequence pretraining for generalizable mri analysis in versatile clinical applications. arXiv pr eprint arXiv:2508.07165 , 2025. 3 [58] Rahul Ravishankar , Zeeshan Patel, Jathushan Rajasegaran, and Jitendra Malik. Scaling properties of diffusion mod- els for perceptual tasks. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 12945–12954, 2025. 3 [59] Marie-Pierre Revel, Samia Boussouar , Constance de Margerie-Mellon, In ` es Saab, Thibaut Lapotre, Dominique Mompoint, Guillaume Chassagnon, Audrey Milon, Math- ieu Lederlin, Souhail Bennani, et al. Study of thoracic ct in covid-19: The stoic project. Radiology , 301(1):E361–E370, 2021. 5 , 1 , 2 [60] Robin Rombach, Andreas Blattmann, Dominik Lorenz, Patrick Esser , and Bj ¨ orn Ommer . High-resolution image synthesis with latent diffusion models. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 10684–10695, 2022. 1 [61] Shaohao Rui, Lingzhi Chen, Zhenyu T ang, Lilong W ang, Mianxin Liu, Shaoting Zhang, and Xiaosong W ang. Multi- modal vision pre-training for medical image analysis. In Pr o- ceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 5164–5174, 2025. 3 [62] Arnaud Arindra Adiyoso Setio, Alberto T raverso, Thomas de Bel, Moira S.N. Berens, Cas van den Bogaard, Pier- giorgio Cerello, Hao Chen, Qi Dou, Maria Evelina Fan- tacci, Bram Geurts, Robbert van der Gugten, Pheng Ann Heng, Bart Jansen, Michael M.J. de Kaste, V alentin K o- tov , Jack Y u-Hung Lin, Jeroen T .M.C. Manders, Alexan- der S ´ o ˜ nora-Mengana, Juan Carlos Garc ´ ıa-Naranjo, Evge- nia Papav asileiou, Mathias Prokop, Marco Saletta, Cor- nelia M Schaefer-Prokop, Ernst T . Scholten, Luuk Scholten, Miranda M. Snoeren, Ernesto Lopez T orres, Jef V ande- meulebroucke, Nicole W alasek, Guido C.A. Zuidhof, Bram van Ginneken, and Colin Jacobs. V alidation, comparison, and combination of algorithms for automatic detection of pulmonary nodules in computed tomography images: The luna16 challenge. Medical Image Analysis , 42:1–13, 2017. 5 , 1 , 2 [63] Amber L Simpson, Michela Antonelli, Spyridon Bakas, Michel Bilello, Ke yv an Farahani, Bram V an Ginneken, An- nette K opp-Schneider, Bennett A Landman, Geert Litjens, Bjoern Menze, et al. A large annotated medical image dataset for the development and ev aluation of segmentation algo- rithms. arXiv preprint , 2019. 5 , 1 , 2 [64] Zhenxiong T an, Songhua Liu, Xingyi Y ang, Qiaochu Xue, and Xinchao W ang. Ominicontrol: Minimal and univer - sal control for diffusion transformer . In Proceedings of the IEEE/CVF International Conference on Computer V ision (ICCV) , pages 14940–14950, 2025. 6 [65] Fenghe T ang, Qingsong Y ao, W enxin Ma, Chenxu Wu, Zi- hang Jiang, and S. Ke vin Zhou. Hi-end-mae: Hierarchi- cal encoder-dri ven masked autoencoders are stronger vision learners for medical image segmentation. Medical Image Analysis , 107:103770, 2026. 2 , 3 , 5 , 7 , 8 , 1 , 4 [66] Y oubao T ang, Y uxing T ang, Y ingying Zhu, Jing Xiao, and Ronald M. Summers. A disentangled generative model for disease decomposition in chest x-rays via normal image syn- thesis. Medical Image Analysis , 67:101839, 2021. 3 [67] Piyush Tiw ary , Kinjawl Bhattacharyya, and Prathosh AP . LangD Aug: Lange vin data augmentation for multi-source domain generalization in medical image se gmentation. In F orty-second International Conference on Mac hine Learn- ing , 2025. 2 [68] T assilo W ald, Constantin Ulrich, Stanisla v Luk yanenko, An- drei Goncharov , Alberto Paderno, Maximilian Miller , Lean- der Maerkisch, Paul Jaeger , and Klaus Maier-Hein. Revis- iting mae pre-training for 3d medical image segmentation. In Pr oceedings of the IEEE/CVF Conference on Computer V ision and P attern Recognition (CVPR) , pages 5186–5196, 2025. 2 [69] Haoshen W ang, Zhentao Liu, Kaicong Sun, Xiaodong W ang, Dinggang Shen, and Zhiming Cui. 3d meddiffusion: A 3d medical latent diffusion model for controllable and high- quality medical image generation. IEEE T ransactions on Medical Imaging , pages 1–1, 2025. 1 , 3 , 5 , 6 , 4 [70] Jindong W ang, Cuiling Lan, Chang Liu, Y idong Ouyang, T ao Qin, W ang Lu, Y iqiang Chen, W enjun Zeng, and Philip S. Y u. Generalizing to unseen domains: A survey on domain generalization. IEEE T ransactions on Knowledge and Data Engineering , 35(8):8052–8072, 2023. 2 [71] Jinglong W ang, Xiawei Li, Jing Zhang, Qingyuan Xu, Qin Zhou, Qian Y u, Lu Sheng, and Dong Xu. Diffusion model is secretly a training-free open vocabulary semantic seg- menter . IEEE T ransactions on Image Pr ocessing , 34:1895– 1907, 2025. 3 [72] Siwen W ang, Churan W ang, Fei Gao, Lixian Su, Fandong Zhang, Y izhou W ang, and Y izhou Y u. Autoregressi ve se- quence modeling for 3d medical image representation. Pro- ceedings of the AAAI Conference on Artificial Intelligence , 39(8):7871–7879, 2025. 3 [73] Xin W ang, Hong Chen, Si’ao T ang, Zihao W u, and W enwu Zhu. Disentangled representation learning. IEEE T ransac- tions on P attern Analysis and Machine Intelligence , 46(12): 9677–9696, 2024. 3 [74] Y iqing W ang, Zihan Li, Jieru Mei, Zihao W ei, Li Liu, Chen W ang, Shengtian Sang, Alan L Y uille, Cihang Xie, and Y uyin Zhou. Swinmm: masked multi-view with swin transformers for 3d medical image segmentation. In In- ternational conference on medical image computing and computer-assisted intervention , pages 486–496. Springer, 2023. 7 [75] Y uran W ang, Zhijing W an, Y ansheng Qiu, and Zheng W ang. Devil is in details: Locality-aware 3d abdominal ct volume generation for self-supervised organ segmentation. In Pr o- ceedings of the 32nd A CM International Confer ence on Mul- timedia , page 10640–10648, 2024. 2 [76] Z. W ang, E.P . Simoncelli, and A.C. Bo vik. Multiscale struc- tural similarity for image quality assessment. In The Thrity- Seventh Asilomar Conference on Signals, Systems & Com- puters, 2003 , pages 1398–1402 V ol.2, 2003. 6 [77] Jakob W asserthal, Hanns-Christian Breit, Manfred T Meyer , Maurice Pradella, Daniel Hinck, Alexander W Sauter, T obias Heye, Daniel T Boll, Joshy Cyriac, Shan Y ang, et al. T o- talsegmentator: robust segmentation of 104 anatomic struc- tures in ct images. Radiology: Artificial Intelligence , 5(5): e230024, 2023. 5 , 6 , 1 , 2 [78] Chen W ei, Karttikeya Mangalam, Po-Y ao Huang, Y anghao Li, Haoqi Fan, Hu Xu, Huiyu W ang, Cihang Xie, Alan Y uille, and Christoph Feichtenhofer . Diffusion models as masked autoencoders. In Pr oceedings of the IEEE/CVF In- ternational Conference on Computer V ision (ICCV) , pages 16284–16294, 2023. 1 , 3 [79] Linshan W u, Jiaxin Zhuang, and Hao Chen. V oco: A simple- yet-effecti ve volume contrastiv e learning framework for 3d medical image analysis. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 22873–22882, 2024. 1 , 2 , 3 , 5 , 7 , 4 [80] Linshan W u, Jiaxin Zhuang, and Hao Chen. Large-scale 3d medical image pre-training with geometric context priors. arXiv pr eprint arXiv:2410.09890 , 2024. 2 , 5 [81] Mengqi Wu, Lintao Zhang, Pew-Thian Y ap, Hongtu Zhu, and Mingxia Liu. Disentangled latent energy-based style translation: An image-lev el structural mri harmonization framew ork. Neural Networks , 184:107039, 2025. 3 [82] W eilai Xiang, Hongyu Y ang, Di Huang, and Y unhong W ang. Denoising diffusion autoencoders are unified self- supervised learners. In Proceedings of the IEEE/CVF In- ternational Conference on Computer V ision (ICCV) , pages 15802–15812, 2023. 1 , 3 [83] Zhenda Xie, Zheng Zhang, Y ue Cao, Y utong Lin, Jianmin Bao, Zhuliang Y ao, Qi Dai, and Han Hu. Simmim: A simple framew ork for masked image modeling. In Proceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 9653–9663, 2022. 7 [84] Y anwu Xu, Li Sun, W ei Peng, Shuyue Jia, Katelyn Morrison, Adam Perer, Afrooz Zandifar , Shyam V isweswaran, Motah- hare Eslami, and Kayhan Batmanghelich. Medsyn: T ext- guided anatomy-aware synthesis of high-fidelity 3-d ct im- ages. IEEE T ransactions on Medical Imaging , 43(10):3648– 3660, 2024. 5 [85] Siyuan Y an, Zhen Y u, Chi Liu, Lie Ju, Dwarikanath Ma- hapatra, Brigid Betz-Stablein, V ictoria Mar, Monika Janda, Peter Soyer , and Zongyuan Ge. Prompt-driven latent domain generalization for medical image classification. IEEE T rans- actions on Medical Imaging , 44(1):348–360, 2025. 2 , 3 [86] T ao Y ang, Cuiling Lan, Y an Lu, and Nanning Zheng. Diffu- sion model with cross attention as an inducti ve bias for disen- tanglement. In Advances in Neural Information Pr ocessing Systems , pages 82465–82492, 2024. 3 [87] Jingfeng Y ao, Bin Y ang, and Xinggang W ang. Reconstruc- tion vs. generation: T aming optimization dilemma in latent diffusion models. In Pr oceedings of the IEEE/CVF Confer- ence on Computer V ision and P attern Recognition (CVPR) , pages 15703–15712, 2025. 6 [88] Y ousef Y e ganeh, Azade Farshad, Ioannis Charisiadis, Marta Hasny , Martin Hartenberger , Bj ¨ orn Ommer , Nassir Na v ab, and Ehsan Adeli. Latent drifting in diffusion models for counterfactual medical image synthesis. In Pr oceedings of the IEEE/CVF Confer ence on Computer V ision and P attern Recognition (CVPR) , pages 7685–7695, 2025. 3 [89] Sihyun Y u, Sangk yung Kwak, Huiwon Jang, Jongheon Jeong, Jonathan Huang, Jinwoo Shin, and Saining Xie. Representation alignment for generation: T raining diffusion transformers is easier than you think. In The Thirteenth In- ternational Conference on Learning Representations , 2025. 6 [90] An Zhang, Han W ang, Xiang W ang, and T at-Seng Chua. Disentangling masked autoencoders for unsupervised do- main generalization. In Eur opean Conference on Computer V ision , pages 126–151, 2024. 2 [91] Kang Zhang, Xiaohong Liu, Jun Shen, Zhihuan Li, Y e Sang, Xingwang W u, Y unfei Zha, W enhua Liang, Chengdi W ang, Ke W ang, et al. Clinically applicable ai system for accu- rate diagnosis, quantitativ e measurements, and prognosis of covid-19 pneumonia using computed tomography . Cell , 181 (6):1423–1433, 2020. 5 , 2 [92] Lvmin Zhang, Anyi Rao, and Maneesh Agrawala. Adding conditional control to text-to-image diffusion models. In Pr oceedings of the IEEE/CVF International Confer ence on Computer V ision (ICCV) , pages 3836–3847, 2023. 6 [93] Qilong Zhao, Y ifei Zhang, Mengdan Zhu, Siyi Gu, Y uyang Gao, Xiaofeng Y ang, and Liang Zhao. Due: Dynamic uncertainty-aware explanation supervision via 3d imputa- tion. In Pr oceedings of the 30th ACM SIGKDD Con- fer ence on Knowledge Discovery and Data Mining , page 6335–6343, 2024. 2 [94] Hongkai Zheng, W eili Nie, Arash V ahdat, and Anima Anandkumar . Fast training of dif fusion models with mask ed transformers. T ransactions on Machine Learning Resear ch , 2024. 6 [95] Jiaxin Zhuang, Luyang Luo, Qiong W ang, Mingxiang W u, Lin Luo, and Hao Chen. Advancing volumetric medical image segmentation via global-local masked autoencoders. IEEE T ransactions on Medical Imaging , pages 1–1, 2025. 5 , 7 , 8 , 1 , 2 , 3 [96] Jiaxin Zhuang, Linshan W u, Qiong W ang, Peng Fei, V arut V ardhanabhuti, Lin Luo, and Hao Chen. Mim: Mask in mask self-supervised pre-training for 3d medical image analysis. IEEE T ransactions on Medical Imaging , 44(9):3727–3740, 2025. 3 , 5 , 7 , 1 , 2 , 4 [97] Xiahai Zhuang. Multiv ariate mixture model for myocardial segmentation combining multi-source images. IEEE T rans- actions on P attern Analysis and Machine Intelligence , 41 (12):2933–2946, 2019. 5 , 1 , 2 MeDUET : Disentangled Unified Pr etraining f or 3D Medical Image Synthesis and Analysis Supplementary Material This supplementary material is organized as follo ws: • Section A presents the full pretraining algorithm of MeDUET . • Section B describes the pretraining datasets and down- stream analysis datasets. • Section C provides additional training and implementa- tion details, including factor extraction for downstream tasks, dual classifier-free guidance for synthesis, and the style augmentation scheme for analysis. • Section D reports a computational analysis comparing FLOPs, wall-clock time, and overall efficiency with ex- isting medical SSL baselines. • Section E presents additional experiments, including ab- lations on the V AE tokenizer , factor effecti veness and generalization, and failure case analysis. • Section F provides discussions on domain configurations, empirical f actor identifiability , and remaining limitations. A. Algorithm Algorithm 1 illustrates the detailed pretraining procedure of MeDUET . B. Datasets B.1. Pretraining Datasets Details of the pretraining V oCo-10k dataset are provided in T able 7 . Specifically , V oCo-10k comprises 10 pub- lic datasets: BTCV [ 42 ], MM-WHS [ 97 ], Spleen [ 63 ], TCIA Covid19 [ 16 ], LUNA16 [ 62 ], STOIC 2021 [ 59 ], FLARE23 [ 48 ], LiDC [ 1 ], HNSCC [ 22 ], and T otalSegmen- tator [ 77 ]. W e follow the pre viously released open-source data splits [ 79 , 95 , 96 ] for the pretraining datasets to ensure consistency and fair comparison. B.2. Downstr eam Analysis T ask Datasets For downstream analysis tasks, we perform a comprehen- siv e ev aluation on four datasets cov ering both segmenta- tion and classification across two modalities. Details of the do wnstream datasets used in the analysis tasks are pro- vided in T able 8 . W e also follow the data splits used in prior work [ 65 , 79 ] to ensure fair comparison. C. Additional T raining Details C.1. Implementation Details All experiments are implemented using PyT orch and MON AI, and training is conducted on four NVIDIA A100 Algorithm 1 Pretraining procedure of MeDUET Require: Dataset with multi domains and domain labels; V AE encoder; MeDUET student model; EMA teacher model. Ensure: Model parameters of student network. Initialize model parameters. 1: for each training iteration do 2: Sample two batches { x i , x j } and encode into latent tokens { z i , z j } by the V AE. ▷ Demixing & Dual Reconstruction 3: Mix and encode tokens z mix ij = ϕ m ( z i , z j , M ) , z e i , z e j = ϕ u ( F enc ( z mix ij ) , M ) . 4: Apply F d and F dec to obtain factors ( c i , s i , c j , s j ) , and reconstructed tokens ( ˆ z i , ˆ z j ) . 5: Compute the dual reconstruction loss L r by Eq. ( 1 ). ▷ Factor Disentanglement 6: Obtain disentangled f actors by con volution ( c i , s i ) , ( c j , s j ) = con v ( z e i ) , conv ( z e j ) . 7: Compute domain classifier loss L d by Eq. ( 2 ). 8: Re-aggregate factors to tokens ˆ z e i = con v ( c i , s i ) , ˆ z e j = con v ( c j , s j ) . ▷ MFTD 9: Re-encode ( ˆ z i , ˆ z j ) with student and teacher branches (teacher stop-gradient) to obtain student factors ( ˆ c i , ˆ s i , ˆ c j , ˆ s j ) and teacher factors ( ˆ c T i , ˆ s T i , ˆ c T j , ˆ s T j ) . 10: Distill tokens of mixed regions using L MFTD by Eq. ( 6 ). ▷ SiQC 11: Generate swapped tokens z ij = F dec ( con v ( c i , s j )) , z j i = F dec ( con v ( c j , s i )) . 12: Re-encode { z i , z j , z ij , z j i } to obtain pooled con- tent/style factors { q c , k c , q s , k s } and construct the quadruplet V ( i, j ) . 13: Build positi ve masks in content/style spaces and compute L SiQC by Eq. ( 12 ). 14: Compute total loss L by Eq. ( 13 ). 15: Update the student model with L and update the teacher model by EMA. 16: end for GPUs. Following previous works [ 13 , 65 , 95 ], we adopt V iT -B as the backbone encoder of MeDUET , which is an appropriate choice gi ven the dataset scale. The input and output channels are modified to 4 to match the latent di- mensionality of our V AE tokenizer [ 24 ]. The class tokens are discarded in all downstream applications. The remain- T able 7. The details of pretraining datasets. Dataset Modality Region of Inter est #V olumes BTCV [ 42 ] CT Abdomen 24 MM-WHS [ 97 ] CT Chest 16 Spleen [ 63 ] CT Spleen 32 TCIA Covid19 [ 16 ] CT Chest 722 LUN A16 [ 62 ] CT Chest 843 STOIC 2021 [ 59 ] CT Chest 2000 FLARE23 [ 48 ] CT Abdomen 4000 LiDC [ 1 ] CT Chest 589 HNSCC [ 22 ] CT Head/Neck 1071 T otalSegmentator [ 77 ] CT Head/Neck/Chest/leg/ Abdomen/pelvis/feet 1203 T able 8. The details of downstream analysis datasets. Dataset Modality #T rain #V alid Full 1-shot 10% 50% 100% AMOS [ 38 ] CT 240 1 24 120 240 120 WORD [ 47 ] CT 100 1 10 50 100 20 BraTS21 [ 3 ] MRI 1000 1 100 500 1000 251 CC-CCII [ 91 ] CT 2514 1 251 1257 2514 1664 ing experimental settings follow prior methods [ 30 , 65 , 79 , 95 , 96 ] for fair comparison. The dimensions of the content and style representations, D c , D s , are set to 768 and 192, respectiv ely . The hyper - parameters of the loss functions are set as follows: λ 1 = 0 . 2 , λ 2 = 0 . 5 , λ 3 = 0 . 3 , and λ c = 0 . 5 . The GRL strength of the domain classifier is set to 1.0. For the EMA teacher, the decay rate is initialized at 0.997 and gradually increased to 0.9997 during pretraining using a cosine schedule. De- tails of the pre-processing and pretraining settings are pro- vided in T able 9 . C.2. Factor Generation for Downstr eam Applica- tions W e extract the content and style vectors for each volume for use in do wnstream tasks. Specifically , each v olume is cropped into fix ed-size sub-v olumes and fed into the frozen MeDUET model, where the factor disentanglement mod- ule produces content and style representations c ∈ R L × D c and s ∈ R L × D s . The sub-volume size is kept identical to the pre-processing configuration (i.e., 96 × 96 × 96 ) to ensure consistency . The resulting content and style repre- sentations from all sub-volumes are then aggregated at the volume lev el. With the sub-volumes covering the full vol- ume, we apply patch-level and token-lev el mean pooling to the aggregated representations, yielding the final content and style vectors c 0 ∈ R D c and s 0 ∈ R D s , which capture the domain-in variant anatomical information and domain- discriminativ e style characteristics of the entire volume, re- spectiv ely . T able 9. The overvie w of pre-processing and pretraining settings in the experiments. Pre-pr ocessing Settings Spacing 1 . 5 × 1 . 5 × 1 . 5 Intensity [ − 175 , 250] Norm [0 , 1] Sub-volume size 96 × 96 × 96 Latent size 4 × 24 × 24 × 24 Augmentation Random Rotate and Flip Pretraining Settings Pretraining steps 200k Optimizer AdamW W eight decay 1e-2 Optimizer momentum β 1 , β 2 = 0 . 9 , 0 . 95 Optimizer lr 1e-4 Batch size 64 × 4 = 256 Lr schedule W armup cosine W armup steps 2k Factor dimension D c , D s = 768 , 192 Loss coefficients λ 1 , λ 2 , λ 3 , λ c = 0 . 2 , 0 . 5 , 0 . 3 , 0 . 5 EMA decay 0.997, 0.9997 GRL coefficient 1.0 C.3. Downstr eam Synthesis Hyperparameters. For the diffusion-based synthesis task, we build our model upon the original DiT and SiT imple- mentations and adopt the same hyperparameter configura- tions [ 49 , 55 ]. The latent representations and factor codes are precomputed using MAISI-V AE [ 24 ] and MeDUET , re- spectiv ely . The hyperparameter settings of DiT/SiT used in our generation experiments are summarized in T able 10 . During both training and sampling, we apply dual classifier- free dropout and dual CFG conditioning [ 33 ] based on the learned disentangled content and style codes. T raining: Dual Classifier -Free Dropout. Gi ven a 3D volume, our frozen MeDUET produces a content vector c 0 ∈ R D c and a style v ector s 0 ∈ R D s . W e standardize each vector with per-sample L 2 normalization and temper- ature scaling as norm τ ( v ) = v ∥ v ∥ 2 + ε · 1 τ , (14) ˆ c = norm τ c ( c 0 ) , ˆ s = norm τ s ( s 0 ) , (15) where τ c , τ s > 0 are learnable scalars. W e maintain learn- able null embeddings c ∅ ∈ R D c , s ∅ ∈ R D s , which pass through the same normalization path: ˆ c ∅ = norm τ c ( c ∅ ) , ˆ s ∅ = norm τ s ( s ∅ ) . At each step we draw independent Bernoulli masks for content and style: m c ∼ Bernoulli(1 − p c ) , m s ∼ Bernoulli(1 − p s ) , (16) with drop probabilities p c , p s ∈ [0 , 1] , which are set to 0 . 2 in our model. The conditional vectors used by the dif fusion T able 10. The overvie w of downstream synthesis settings in the experiments. DiT -B/4 SiT -B/4 Pre-pr ocessing Spacing 1 . 5 × 1 . 5 × 1 . 5 1 . 5 × 1 . 5 × 1 . 5 Intensity [ − 175 , 250] [ − 175 , 250] Norm [0 , 1] [0 , 1] Sub-volume size 256 × 256 × 128 256 × 256 × 128 Latent size 4 × 64 × 64 × 32 4 × 64 × 64 × 32 Architectur e Input dim. 4 × 64 × 64 × 32 4 × 64 × 64 × 32 Hidden dim. 768 768 Num. blocks 12 12 Num. heads 12 12 Patch size 4 4 Optimization Batch size 16 × 4 = 64 16 × 4 = 64 Optimizer AdamW AdamW Lr 5e − 5 5e − 5 ( β 1 , β 2 ) (0 . 9 , 0 . 999) (0 . 9 , 0 . 999) Lr schedule W armup cosine W armup cosine T raining steps 2M 2M EMA decay 0.9999 0.9999 Interpolants α t 1 − t – σ t t – ω t σ t – T – 1000 T raining objecti ve v-prediction noise-prediction Sampler Euler -Maruyama DDPM Sampling steps 250 250 Dropout prob p c , p s = 0 . 2 , 0 . 2 p c , p s = 0 . 2 , 0 . 2 Guidance w c , w s = 3 . 0 , 3 . 0 w c , w s = 3 . 0 , 3 . 0 transformer (DiT/SiT) are ˜ c = m c ˆ c + (1 − m c ) ˆ c ∅ , ˜ s = m s ˆ s + (1 − m s ) ˆ s ∅ . (17) W e inject (˜ c, ˜ s ) via dual-branch AdaLN-Zero [ 55 ]. For each block we predict af fine/gate parameters from content/style and combine them: ( γ c , β c ) = W c ˜ c , ( γ s , β s ) = W s ˜ s , (18) γ = γ c + γ s , β = β c + β s , (19) Giv en pre-LN activ ations h , AdaLN-Zero modulation is AdaLN( h ) = (1 + γ ) ⊙ LN( h ) + β . (20) Sampling: Dual CFG. For a target pair ( c i , s j ) (e.g., con- tent and style from reference cases), we form normalized ˆ c i , ˆ s j and their null counterparts ˆ c ∅ , ˆ s ∅ . At each time step we compute three model ev aluations: e cs = ϵ θ ( z t , t ; ˆ c i , ˆ s j ) , (21) e ns = ϵ θ ( z t , t ; ˆ c ∅ , ˆ s j ) , (22) e cn = ϵ θ ( z t , t ; ˆ c i , ˆ s ∅ ) , (23) T able 11. The ov ervie w of downstream analysis settings in the experiments. Pre-pr ocessing Settings Spacing 1 . 5 × 1 . 5 × 1 . 5 Intensity [ − 175 , 250] Norm [0 , 1] Sub-volume size 96 × 96 × 96 Latent size 4 × 24 × 24 × 24 Num. sub-crops 4 Augmentation Random Rotate, Flip, Scale, Shift Fine-tuning Settings Optimizer AdamW W eight decay 1e-5 Optimizer momentum β 1 , β 2 = 0 . 9 , 0 . 999 Optimizer lr 2e-4 Batch size 1 × 4 = 4 Inference Sliding W indo w Inference Inference ov erlap 0.5 Sw batch size 32 T able 12. The hyperparameter settings of style augmentation un- der different data ratios in the do wnstream analysis tasks. Config Data Ratios 1-shot 10% 50% 100% p aug 0.9 0.8 0.6 0.3 α min , α max 0.4, 0.8 0.4, 0.8 0.3, 0.6 0.2, 0.5 λ aug 0.4 0.4 0.4 0.4 λ cons 0.3 0.3 0.3 0.3 EMA decay 0.90 0.90 0.90 0.90 where z t denotes the input of diffusion transformers, and t is the timestep. W e then apply dual guidance with indepen- dent strengths w c , w s ≥ 0 : ϵ CFG = e cs + w c ( e cs − e ns ) + w s ( e cs − e cn ) . (24) Setting w c = w s = 0 yields near unconditional sampling. In our experiment, we set w c = 3 . 0 and w s = 3 . 0 by de- fault. T o sum up, our dual CFG design lets us (i) train with missing-condition robustness through the dual CF-dropout, and (ii) perform finely controllable generation at inference time with independent knobs for content consistency and style strength, all while keeping the content/style semantics disentangled. C.4. Downstr eam Analysis Hyperparameters. Follo wing prior work [ 65 , 79 , 95 , 96 ], the fine-tuning setups for the analysis tasks are kept largely consistent with the pretraining configurations, as summa- rized in T able 11 . For segmentation tasks, the latent-space outputs of UNETR are passed through the V AE decoder to T able 13. The computational cost and wall-clock time comparison of medical SSL methods. Method Backbone Param. (M) Flops (G) Per Epoch Time (s) Pretrain T ime (GPU hours) Dice Score (%) ↑ Accuracy (%) ↑ BTCV AMOS WORD BraTS21 CC-CCII V oCo [ 79 ] Swin-B 127.44 1264.9 1867.3 331.9 85.31 85.51 87.53 77.72 90.72 MIM [ 96 ] V iT -B 71.0 1002.5 615.4 683.8 84.07 84.90 87.68 76.77 88.35 Hi-End-MAE [ 65 ] V iT -B 98.9 133.9 164.6 731.6 84.86 85.80 89.37 76.93 86.50 MeDUET V iT -B 105.1 50.1 72.9 405.2 88.02 86.34 90.07 78.02 91.35 reconstruct the predicted segmentation masks. For classifi- cation tasks, we remove the V AE decoder and modify the final output layer of UNETR [ 27 ] to directly produce the predictiv e logits. Moreover , we further ev aluate our disen- tangled latent space on downstream analysis by introduc- ing a style augmentation scheme for both segmentation and classification tasks. Style A ugmentation. With the learned style representa- tions from pretrained MeDUET , for cross-domain adapta- tion, we first build a style prototype from K unlabeled target-domain v olumes: s pro = 1 K K X i =1 s ( i ) . (25) T o reduce sensitivity to the particular choice of K volumes and to track the empirical tar get distrib ution, we further up- date the prototype online using an EMA o ver target-domain mini-batches. During training, for each labeled source sam- ple ( z , y ) , we stochastically generate a style-augmented volume in the MeDUET latent space. W e keep the con- tent tokens fixed and linearly mix the style tokens to ward the target prototype with a random strength α : s ′ = (1 − α ) s + α s pro , (26) z aug = F dec (con v( c , s ′ )) . (27) where F dec is the decoder of MeDUET that reconstruct the patch tokens from factors, and con v denotes the con volu- tional layer used to aggregate factors. Let f θ be the segmen- tation/classification network (UNETR in our experiments), for each mini-batch we optimize a combined supervised and consistency objecti ve: L sup = L task ( f θ ( z ) , y ) + λ aug L task ( f θ ( z aug ) , y ) , (28) L cons = D KL ( σ ( f θ ( z aug )) ∥ σ ( f θ ( z ))) , (29) L = L sup + λ cons L cons . (30) where L task is task loss for corresponding perception tasks, D KL represents the KL div ergence, σ is the softmax func- tion, λ aug and λ cons are coefficients for balancing the loss term contributions. W e apply latent style augmentation to a fraction p aug of training samples per batch, drawing α ∼ U [ α min , α max ] . T able 12 illustrates the hyperparam- eter settings of our style augmentation strategy under dif- ferent label proportions. D. Computational Analysis Considering that the ov erall pretraining pipeline of our MeDUET is relativ ely complex, containing multiple com- ponents and loss terms, we provide a computational analysis to illustrate the practicality of our method. Specifically , we implement a compute cost and wall-clock time comparison with sev eral Medical SSL methods [ 65 , 79 , 96 ] to ensure a fair comparison, as demonstrated in T able 13 . Notably , all comparisons are conducted on identical pre-processing settings and the pretrain time is measured on four NVIDIA A100 GPUs. It can be observed that MeDUET achie ves the best Dice/Accuracy while also enjoying the lowest Flops and per-epoch wall-clock time among all compared meth- ods. This efficiency mainly comes from performing SSL in the V AE latent space with a 4 × spatial compression ratio, which greatly reduces the number of tokens processed by the V iT backbone. Hence, the overall pretraining time re- mains comparable to V oCo [ 79 ] and is substantially lower than MIM [ 96 ] and Hi-End-MAE [ 65 ], despite our more elaborate disentanglement objectiv es. E. Experiments E.1. More Ablation Study Role of the V AE T okenizer . T o assess the sensitivity of MeDUET to the choice of V AE tokenizer , we further ev al- uate our framew ork using alternati ve V AE models. Giv en that tailored 3D V AEs for medical imaging remain rela- tiv ely undere xplored, we adopt the P atch-V olume Autoen- coder from 3D MedDiffusion [ 69 ], a 3D VQGAN-based tokenizer pretrained on large-scale CT and MRI datasets. As shown in T able 14 , replacing MAISI-V AE [ 24 ] with the Patch-V olume Autoencoder leads to only minor fluctu- ations in both synthesis and analysis performance. While MAISI-V AE achiev es slightly higher averages, both tok- enizers maintain consistently strong results, suggesting that the improv ements of MeDUET arise primarily from the dis- entangled pretraining scheme rather than from any specific V AE design. These findings indicate that our framework is relativ ely insensitive to the tok enizer choice. Factor Impact f or Medical Synthesis Baselines. T o fur- ther assess the usefulness of the learned factors, we replace the original metadata condition vectors in medical synthe- T able 14. Ablation experiments on the role of V AE. Method Synthesis Analysis FID ↓ MMD ↓ MS-SSIM ↓ Dice Score (%) ↑ BTCV WORD 1-shot 100% 1-shot 100% Patch-V olume Autoencoder [ 69 ] 0.7948 0.5630 0.1686 78.55 94.13 79.29 94.22 MAISI-V AE [ 24 ] 0.7874 0.5598 0.1659 78.72 94.06 79.56 94.28 T able 15. Ablation experiments on the impact of our learned factor codes on baseline medical synthesis methods. Method FID ↓ MMD ↓ MS-SSIM ↓ MAISI [ 24 ] 0.9139 0.6292 0.2057 + Factor Conditions (Ours) 0.9046 (-0.093) 0.6159 (-0.133) 0.1918 (-0.139) 3D MedDiffusion [ 69 ] 0.9216 0.6327 0.2032 + Factor Conditions (Ours) 0.9081 (-0.135) 0.6198 (-0.129) 0.1931 (-0.101) sis baselines [ 24 , 69 ] with our pretrained content and style embeddings. As sho wn in T able 15 , plugging in our fac- tors yields consistent improvements on all three metrics for both baselines, e ven though the generativ e architectures and training setups are unchanged. This indicates that MeDUET provides richer and more informati ve conditioning signals than handcrafted metadata, capturing continuous, image- deriv ed anatomy and appearance cues that are not fully de- scribed by discrete site labels, and can thus benefit a broad class of medical synthesis models. E.2. F actor Generalization f or MRI T o further demonstrate the generalization capability of our disentanglement scheme, we perform t-SNE visualization on the content and style factors extracted from the BraTS dataset [ 3 ], as shown in Fig. 9 . On this out-of-distribution MRI dataset, the t-SNE plots reveal that the style embed- dings form distinct clusters corresponding to the four MRI modalities (T1, T1ce, T2, FLAIR), whereas the content em- beddings remain mixed across modalities. Notably , these BraTS modalities are nev er used as domain labels dur- ing pretraining. This emergent clustering indicates that our style branch automatically adapts to unseen modality- specific appearance factors, while the content branch re- mains modalit y-in variant. T ogether , these observ ations pro- vide empirical evidence that the proposed disentanglement scheme generalizes beyond the supervised domains. E.3. F ailure Cases Analysis Sev eral failure cases of factor-guided synthesis with MeDUET are shown in Fig. 10 . T ypical qualitati ve failures include anatomically implausible shapes, missing or inac- curate pathology representation, and incomplete content- style disentanglement. Below , we discuss sev eral plausi- ble causes and outline potential mitigation strategies: (i) In multi-center datasets, anatomical and pathological pat- terns often correlate with specific scanners or sites, caus- ing the style branch to inadvertently learn anatomical cues. This issue may be alleviated by improving cross-center data balancing and introducing independence regulariza- tion between content factors and domain labels. (ii) Be- cause content and style factors originate from a shared en- coder and must satisfy strong reconstruction constraints, the model may redundantly store anatomical information in both branches. Stronger architectural separation, such as decorrelation constraints between branches, may help reduce this leakage. (iii) Joint optimization across mul- tiple objectives can prioritize reconstruction fidelity over perfectly disentangled factor separation. In addition, ex- treme factor guidance during inference may push sampling into underexplored regions where residual entanglement be- comes more apparent. Future w ork will in vestigate harder factor -swap sampling, as well as calibrated or re gularized guidance ranges, to mitigate these ef fects. F. Discussions F .1. Exploration for Domain Setups It is worth noting that not all datasets provide reliable style metadata (e.g., scanner vendor , field strength, kVp, recon- struction kernel). Consequently , we do not exploit such fine-grained style annotations; instead, we use dataset-lev el domain IDs as domain labels to ensure consistenc y across datasets. This choice is admittedly coarse, where each dataset may still contain multiple protocols and gradual shifts, b ut it roughly captures the dominant cross-dataset ap- pearance differences that the style branch is meant to model. Because the style encoder learns a continuous style space under weak domain supervision, MeDUET does not rely on perfectly clean domains, and our experiments show that this simple labeling already yields stable disentanglement and strong performance, with finer-grained domain definitions left for future work. Figure 9. The t-SNE visualization of MeDUET content and style representations on the BraTS dataset, where points are color-coded according to their modality labels. Left: Content code distribution. Right: Style code distribution. F .2. Empirical Factor Identifiability W e emphasize that our use of the term identifiability is purely empirical rather than theoretical. MeDUET does not provide formal guarantees that content and style f actors are uniquely recov erable. Instead, we demonstrate approx- imate disentanglement through diagnostics such as domain linear probes, t-SNE visualizations, and factor -controlled generation. These results suggest that the learned domain- in variant content and domain-specific style spaces behav e in a factorized manner under our training setup. F .3. Limitations While MeDUET demonstrates strong performance across div erse medical imaging tasks through a unified pretrain- ing framework, se veral limitations remain. First, MeDUET introduces multiple components, increasing implementa- tion complexity compared with simpler SSL baselines. Al- though operating in a compressed latent space keeps the computational cost and wall-clock time manageable, the ov erall pretraining pipeline remains resource-intensi ve. A promising future direction is to design lighter-weight objec- tiv es or distillation strategies that retain the learned factors while reducing architectural and training o verhead. Addi- tionally , our generativ e ev aluation is primarily conducted on datasets sharing similar anatomical distributions as those used for pretraining. Cross-dataset or out-of-distribution synthesis is not extensiv ely explored and warrants further in vestigation. Moreov er , we currently focus on segmenta- tion and classification as downstream analysis tasks. Ex- tending MeDUET to other tasks such as detection, re gistra- tion, or report generation, as well as to multi-institutional real-world data with heterogeneous protocols, is an impor- tant future direction. R e f er en c e S y n th es i z ed A B C o nt e nt A + S t y l e B C o n t en t B + S ty l e A R e f er en c e S y n th es i z ed A B C o nt e nt A + S t y l e B C o n t en t B + S ty l e A R e f er en c e S y n th es i z ed A B C o nt e nt A + S t y l e B C o n t en t B + S ty l e A Figure 10. Some failure cases of synthesized images generated by MeDUET .
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment