LGD-Net: Latent-Guided Dual-Stream Network for HER2 Scoring with Task-Specific Domain Knowledge
It is a critical task to evalaute HER2 expression level accurately for breast cancer evaluation and targeted treatment therapy selection. However, the standard multi-step Immunohistochemistry (IHC) staining is resource-intensive, expensive, and time-…
Authors: Peide Zhu, Linbin Lu, Zhiqin Chen
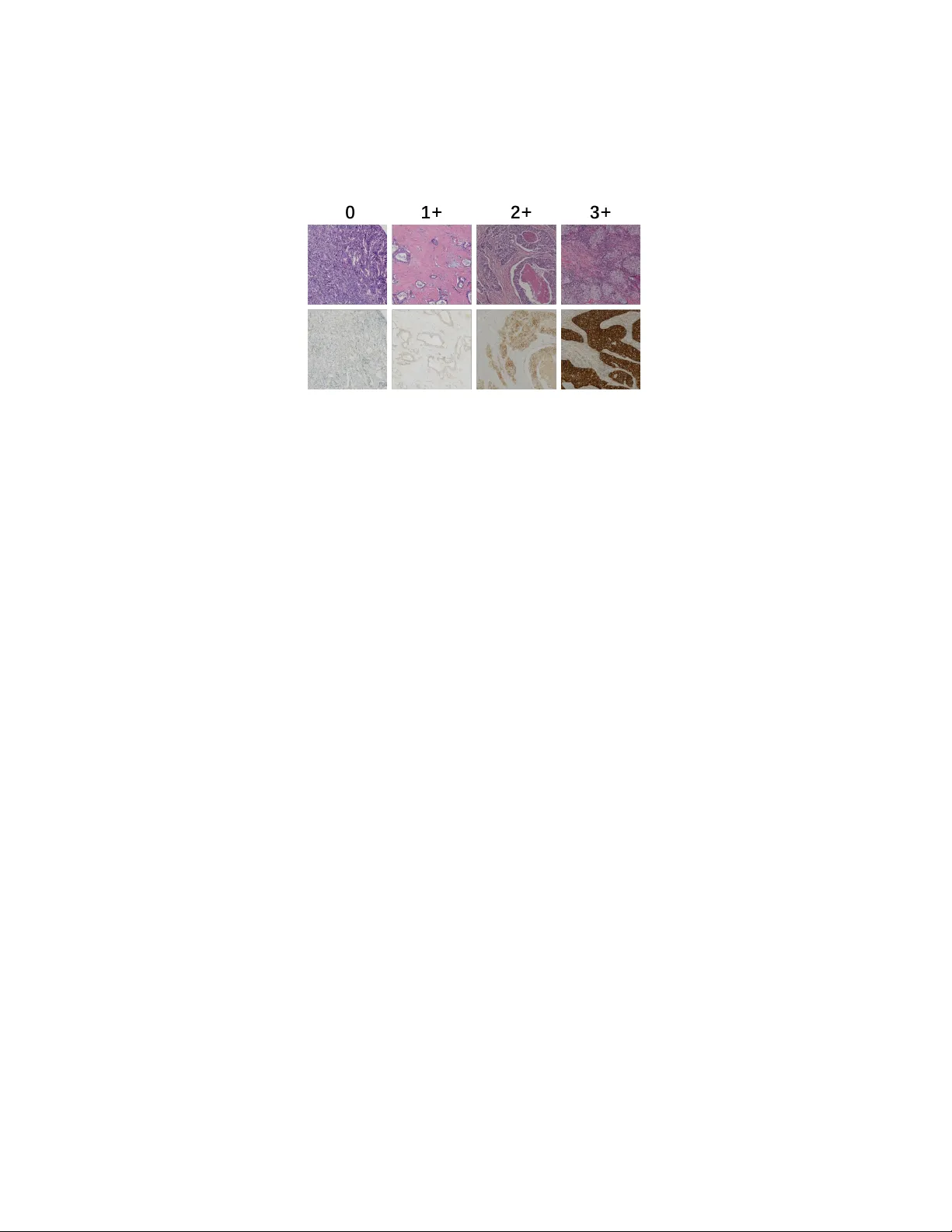
LGD-Net: Laten t-Guided Dual-Stream Net w ork for HER2 Scoring with T ask-Sp ecific Domain Kno wledge P eide Zh u a,1 , Lin bin Lu b,1 , Zhiqin Chen a , Xiong Chen b, ∗ a Scho ol of Mathematics and Statistics, F ujian Normal University, F uzhou, 350117, China b Dep artment of Onc olo gy, Mengchao Hep atobiliary Hospital of F ujian Me dic al University, F uzhou, 350117, China Abstract It is a critical task to ev alaute HER2 expression lev el accurately for breast cancer ev aluation and targeted treatmen t therapy selection. Ho wev er, the standard m ulti-step Imm unohisto c hemistry (IHC) staining is resource- in tensive, exp ensiv e, and time-consuming, whic h is also often una v ailable in man y areas. Consequently , predicting HER2 levels directly from H&E slides has emerged as a potential alternativ e solution. It has b een sho wn to b e effectiv e to use virtual IHC images from H&E images for automatic HER2 scoring. Ho wev er, the pixel-level virtual staining metho ds are computation- ally exp ensiv e and prone to reconstruction artifacts that can propagate diag- nostic errors. T o address these limitations, we prop ose the Laten t-Guided Dual-Stream Net work (LGD-Net) , a no v el framew ork that employ es cross-mo dal feature hallucination instead of explicit pixel-lev el image gener- ation. LGD-Net learns to map morphological H&E features directly to the molecular latent space, guided by a teac her IHC enco der during training. T o ensure the hallucinated features capture clinically relev an t phenot yp es, w e explicitly regularize the mo del training with task-sp ecific domain kno wl- edge, sp ecifically nuclei distribution and mem brane staining in tensity , via ligh tw eight auxiliary regularization tasks. Extensiv e exp erimen ts on the pub- lic BCI dataset demonstrate that LGD-Net achiev es state-of-the-art p erfor- mance, significan tly outp erforming baseline metho ds while enabling efficien t inference using single-mo dalit y H&E inputs. Keywor ds: HER2 Scoring, F eature Hallucination, Domain Kn o wledge ∗ Corresp onding Author 1 Equal contribution Regularization 1. In tro duction Breast cancer is one of the most frequently diagnosed malignancy , imp os- ing a significan t burden on public health systems [1]. HER2 (Human Epi- dermal Gro wth F actor Receptor 2) has been widely adopted as a fundamen- tal biomarker for guiding therap eutic strategies. The accurate assessmen t of HER2 expression lev el (categorized as 0, 1+, 2+, or 3+) is required to iden tify patien ts eligible for targeted therapies[2, 3]. The standard clinical workflo w t ypically consists of an initial morphological examination using H&E (Hema- to xylin and Eosin) staining, follo wed b y sp ecific molecular testing via IHC (Imm unohisto c hemistry) [4]. This multi-step pro cess is resource-in tensiv e, exp ensiv e, and time-consuming. F urthermore, in many developing regions, IHC infrastructure is often inaccessible. In addition, the sub jectiv e in terpre- tation of staining in tensity can lead to substantial inter-rator v ariabilit y [5]. Consequen tly , developing to ols that are capable of HER2 scoring automati- cally directly from H&E slides has b ecome a critical researc h ob jective. The rapid adv ancemen t of deep learning-based metho ds has enabled the extraction of complex patterns from histopathological images. Early studies on HER2 scoring directly from H&E images by training deep neural net- w orks lik e Conv olutional Neural Net works (CNNs) to identify morphological correlates of protein expression [6, 7]. Although these unimo dal approaches established a baseline, they often struggle to distinguish b et w een in termedi- ate HER2 scores (e.g., distinguishing 1+ from 2+), as H&E staining primar- ily visualizes tissue structure rather than sp ecific molecular signals [8]. T o bridge this information gap, the field has increasingly turned tow ard virtual staining techniques. These metho ds typically employ generative mo dels such as Generative A dversarial Net works (GANs) to translate H&E images into syn thetic IHC images, effectiv ely creating a pro xy mo dalit y for do wnstream classification [9]. F or instance, Liu et al. [10] introduced the BCI dataset and prop osed a p yramid-based Pix2Pix mo del to generate registered IHC pairs. More recently , Qin et al. [11] developed a dynamic reconstruction framew ork that synthesizes missing mo dalities to improv e classification robustness. Although these generativ e approac hes ha ve demonstrated the superier p erformance ov er unimo dal approac hes, they rely on pixel-lev el virtual stain- ing and thus are p osed with tw o ma jor challenges. First, the ob jective of im- age translation often diverges from the ob jective of diagnosis. The generation 2 mo dels trained to minimize pixel-wise reconstruction error or adv ersarial loss ha ve to learn ho w to reconstruct details like the high-frequency patterns that are irrelev an t to HER2 scoring, such as stromal texture, red blo o d cells, and bac kground noise. This results in computational inefficiency and instabilit y during b oth training and inference. Second, the virtual staining mo dels are prone to generating artifacts or hallucinations in the image space when faced with domain shifts [12, 13]. If a model generates a false p ositiv e membrane staining pattern to satisfy the discriminator, the downstream classifier can pro duce an incorrect diagnosis due to it. T o address these c hallenges, w e in tro duce a no vel metho d for HER2 ex- pression scoring that a v oids explicit image generation in this w ork. W e hy- p othesize that it is not the image of the IHC stain that is required for clas- sification, b ut rather the mole cular semantics enco ded within it. Therefore, w e prop ose the Laten t-Guided Dual-Stream Net work (LGD-Net) , a framew ork that replaces pixel-level translation with fe atur e hal lucination . In- stead of reconstructing a visually interpretable IHC image, LGD-Net learns a mapping from morphological H&E features to a molecular latent space, guided b y a teacher netw ork tht enco des the IHC image to the latent space during training. This approac h elliminates the need for the deco der for vir- tual IHC image generation, fo cusing the mo del’s capacit y on discriminativ e biomark er features rather than textural reconstruction. T o ensure the hallucinated features remain biologically meaningful, w e regualize the the latent space feature generation with the domain knowledge. Sp ecifically , we design light w eight auxiliary tasks that force the latent fea- tures to b e predictive of nuclei dis tribution and membrane staining intensit y . This ensures that the learned represen tation is not just a statistical approx- imation of the teac her net work, but explicitly enco des the structural and molecular cues essen tial for HER2 scoring. The primary con tributions of this pap er are as follo ws: • W e prop ose a cross-mo dal feature hallucination framew ork that effec- tiv ely bridges the gap b et w een H&E morphology and IHC molecular information, eliminating the computational ov erhead and artifact risks asso ciated with pixel-level image generation. • W e introduce a domain-kno wledge regualization mec hanism that en- forces ph ysical constrain ts, specifically , the n uclei densit y and mem- brane in tensity , on the latent features, impro ving the mo del’s sensitiv- it y to subtle HER2 expression differences. 3 Student Encoder T eacher Encoder Attention Fusion Nuclei Decoder Membrane Decoder Feature Hallucinator Distillation Loss Classifier Nuclei Loss Membrane Loss T raining Phase Inference Phase Classification Loss Student Encoder Feature Hallucinator Attention Fusion Classifier Figure 1: Overview of the prop osed LGD-Net framework. • W e conduct extensiv e exp erimen ts on the public BCI dataset, demon- strating that LGD-Net outp erforms b oth unimo dal baselines and re- cen t state-of-the-art generativ e metho ds [10, 11], while requiring only standard H&E images during inference. 2. Metho dology W e prop ose the Latent-Guided Dual-Stream Net w ork (LGD-Net) , a framework designed for HER2 expression level scoring from H&E images. Unlik e prev ailing virtual staining approac hes that dep end on computation- in tensive pixel-level translation [10, 11], LGD-Net employs a fe atur e hal lu- cination strategy . This approach enables the inference of missing IHC se- man tics directly within the laten t space without generating the actual visual images, constrained b y task-sp ecific pathological priors. 2.1. Pr oblem F ormulation and F r amework Overview F ormally , let D = { ( x i H E , x i I H C , y i ) } N i =1 represen t a set of registered H&E and IHC paired patc hs from the BCI cohort [10], where x H E , x I H C ∈ R H × W × 3 are 3-c hannel R GB images with the heigh t H and width W pixels, and y ∈ { 0 , 1+ , 2+ , 3+ } denotes the HER2 expression score. Our ob jective is to learn 4 a predictiv e mo del that, during infer enc e , accepts only a single H&E input x H E while lev eraging the latent molecular information of the absen t IHC mo dalit y to predict y . As depicted in Fig. 1, LGD-Net comprises three ma jor comp onen ts: (1) a Dual-Stream Enco ding Mo dule structured in a T eacher-Studen t config- uration; (2) a Latent F eature Hallucinator that maps morphological fea- tures to pseudo-molecular representations; and (3) a Domain-Kno wledge REgularization module that enforces biological consistency through auxil- iary regularization tasks. 2.2. Cr oss-Mo dal F e atur e Hal lucination Div erging from con ven tional image-to-image translation, w e perform cross- mo dal synthesis exclusiv ely within the high-level feature space. In this w ay , in stead of generating high-frequency artifacts, we fo cus on the mo del’s ca- pacit y on discriminative biomark ers. 2.2.1. Dual-Str e am Enc o ders W e adopt a parallel enco der architecture: a Studen t Enco der E S pro cess- ing H&E inputs and a T eacher Enco der E T pro cessing IHC inputs. Lev eraging the feature extraction proto col from [11], w e obtain the latent representa- tions: z H E = E S ( x H E ) , z I H C = E T ( x I H C ) (1) where z ∈ R C × h × w denotes the spatial feature maps. During training, E T pro vides the target distribution for molecular feature learning; critically , it is discarded during the inference phase, requiring only H&E input. 2.2.2. L atent Mapping T o bridge the domain gap b et ween the H&E and IHC modalities, w e in tro duce the F eature Hallucinator M . This mo dule learns a non-linear transformation to pro ject H&E features in to a “hallucinated” IHC laten t space: ˆ z I H C = M ( z H E ) (2) The primary ob jectiv e is to align ˆ z I H C structurally and seman tically with the authentic IHC features z I H C without pixel reconstruction. 5 2.3. Domain Know le dge-c onstr aine d T r aining via Auxiliary T asks Minimizing statistical discrepancy b et ween latent vectors is insufficient to guaran tee that the hallucinated features capture clinically relev an t phe- not yp es. The c haracterization of nuclei distribution and membr ane staining intensity is critical for accurate HER2 assessment. Therefore, w e prop ose to in ternalize these biological constraints into the latent representation via tw o ligh tw eight auxiliary deco ders. 2.3.1. Nuclei Density R e gularization T o ensure ˆ z I H C retains accurate structural information, w e incorp orate a Nuclei Deco der D nuc . Ground truth density maps K g t are generated from the Hemato xylin c hannel of the training images using color decon volution and Gaussian filtering [14]. The deco der estimates this distribution as: ˆ K = D nuc ( ˆ z I H C ) (3) By minimizing the reconstruction error b et ween ˆ K and K g t , we constrain the laten t features to enco de precise cellular lo calization. 2.3.2. Membr ane Intensity R e gularization Giv en that HER2 scoring is principally defined by the completeness of mem brane staining (D AB signal), we employ a Mem brane Deco der D mem to predict membrane activ ation. The ground truth mask M g t is derived via adaptiv e thresholding of the DAB channel in the HED color space [14]. The predicted mask is formulated as: ˆ M = D mem ( ˆ z I H C ) (4) These auxiliary tasks ensure that ˆ z I H C is not merely an abstract v ector but explicitly enco des the biological semantics required for the diagnostic task. 2.4. Dynamic F usion and Classific ation F or the final prediction, we integrate the morphological features z H E with the hallucinated molecular features ˆ z I H C . T o account for p otential v ariations in the reliabilit y of the hallucinated features, w e implemen t the mo dalit y- sensitiv e feature attention-based fusion mechanism. W e concatenate the feature maps and compute a spatial-channel attention map A via shared MLP lay ers: z f used = Concat ( z H E , ˆ z I H C ) · A ( z H E , ˆ z I H C ) (5) 6 The fused representation is then fed in to a classifier C to compute the prob- abilit y distribution ov er HER2 expression levels: ˆ y = C ( z f used ) (6) 2.5. L oss F unctions and Optimization T o train the prop osed mo del, w e design a comp osite function to collab o- rativ ely optimize the classification p erformance while enforcing cross-mo dal alignmen t and biological consistency , sp ecifically with the following training ob jectives: 1. Classification Loss ( L cls ): W e utilize a Cross-Entrop y loss: L cls = − X c y c log( ˆ y c ) (7) 2. F eature Distillation Loss ( L dist ): T o align the hallucinated and real IHC features, w e minimize the cosine distance, prioritizing semantic direction ov er magnitude: L dist = 1 − ˆ z I H C · z I H C ∥ ˆ z I H C ∥ 2 ∥ z I H C ∥ 2 (8) 3. Auxiliary Biological Losses ( L bio ): These losses enforce the domain kno wledge constraints. W e apply Mean Squared Error (MSE) for nuclei densit y regression and Dice Loss for membrane segmen tation: L nuc = ∥ ˆ K − K g t ∥ 2 2 , L mem = 1 − Dice ( ˆ M , M g t ) (9) The total optimization ob jectiv e is then formulated as: L total = L cls + λ d L dist + λ n L nuc + λ m L mem (10) where λ d , λ n , λ m are h yp erparameters balancing feature alignmen t and bio- logical regularization. Up on completion of training, the auxiliary deco ders are remov ed, resulting in a streamlined inference arc hitecture that requires only H&E input. 7 Figure 2: Examples of paired H&E-IHC images with different HER2 expression levels (0, 1+, 2+, 3+ from left to right). 3. Exp erimen tal Setup 3.1. Dataset W e conducted exp erimen ts to v alidate the LGD-Net on the BCI Dataset [10], a large publicly av ailable cohort for HER2 assessment. The dataset comprises 4,873 pairs of H&E and IHC patches ( 1024 × 1024 pixels) deriv ed from strictly spatially registered whole slide images with the HER2 expression lev el annotations ranging from negativ e (0) to strongly p ositiv e (3+). W e use the official training and test sets (3896:977 images) for training and testing. T o generate ground truth sup ervision for the auxiliary tasks, we emplo yed the deterministic image pro cessing pip eline describ ed in [14]. Sp ecifically , n uclei density maps ( K g t ) w ere generated by applying color deconv olution to extract the hematoxylin channel, follow ed by Gaussian filtering. Membrane staining masks ( M g t ) were deriv ed via adaptiv e thresholding of the DAB c hannel in the Haemato xylin-Eosin-DAB (HED) color space. 3.2. Implementation Details W e implemen ted the propso ed framework in PyT orch. The model was trained on a server with 256GB memory and tw o NVIDIA R TX 3090 GPU. W e utilized ResNet-50 [15], initialized with ImageNet w eigh ts, as the bac k- b one for b oth the Student and T eac her enco ders. Input images were resized to 512 × 512 pixels to optimize the trade-off betw een computational effi- ciency and fine-grained feature retention. T o generate the fak e IHC images for baselines, w e trained the generator with an H800 GPU. W e conduct mo del training was p erformed in an end-to-end manner using the A dam W optimizer ( β 1 = 0 . 9 , β 2 = 0 . 999 , w eight deca y= 1 e − 4 ). W e 8 initialized the learning rate at 1 e − 4 and applied a cosine annealing schedule o ver 50 ep o c hs. The loss w eigh ting factors w ere empirically set to λ d = 10 . 0 , λ n = 5 . 0 , and λ m = 5 . 0 to prioritize feature alignment and biological regularization. 3.3. Evaluation Metrics W e report the A c cur acy (A c c) and Macr o-aver age d F1-sc or e (F1) to assess classification p erformance, with the latter b eing particularly critical giv en the inheren t class im balance. A dditionally , w e also rep ort the Cohen ’s Kapp a ( κ ) to quan tify the agreement b et w een model predictions and ground-truth annotations. 4. Results and Analysis 4.1. Comp arison with State-of-the-Art Metho ds T o v alidate the efficacy of LGD-Net, we compared it against the following categories of baseline approaches that cov er most HER2 prediction metho ds: 1. Unimo dal Baseline: A standard ResNet-50 trained exclusiv ely on single mo dalit y ( H&E images or IHC images only). 2. Dual-mo dal Baselines: F or this category of baselines, w e consider three settings. The first is image c onc at , which takes b oth the paired H&E image and IHC image as the input to the con v olutional la yer. The second is fe atur e level c onc at with attention whic h encoder the H&E image and the IHC image separately , and apply atten tion on the concated features. The third is the F e atur e-L evel F usion , where w e use the Bidirectional Cross-Mo dal Reconstruction framework [11], which represen ts the current state-of-the-art. T able 1: Quantitativ e comparison on the BCI testing set.. Metho d A cc(%, ↑ ) F1 ( ↑ ) κ ( ↑ ) H&E Unimo dal 82.29 0.7935 0.7543 IHC Unimo dal 89.46 0.8906 0.8642 ImageConcat ( H&E + real IHC ) 90.99 0.8929 0.8724 F eatureConcat ( H&E + real IHC) 93.76 0.9429 0.9208 F eatureF usion (H&E + real IHC) 94.37 0.9375 0.9190 LGD-Net (Ours) (H&E) 95.60 0.9644 0.9453 9 As reported in T able 1, the H&E-Only baseline yields sub optimal p er- formance (82.29%), confirming that morphological features alone are insuffi- cien t to resolv e ambiguous HER2 classes. The IHC-only baseline outperforms H&E baseline as expected since the IHC images directly con tain the visual features of the HER2 expression. The dual-mo dal metho ds improv e accuracy b y synthesizing the H&E and IHC information. W e can also observe that the feature-based metho ds including the atten tion on the concatenated features and the adv anced feature fusion can further improv e the p erformance. No- tably , LGD-Net surpasses the previous state-of-the-art method b y 1.23% in accuracy , 0.0269 in Macro-F1 and 0.0263 in Kappa. This p erformance gain corrob orates our h yp othesis that fe atur e hal lucination , constrained by in- ternalized domain kno wledge, captures diagnostic seman tics more effectively than pixel-level features. 4.2. A blation Study W e conducted a comprehensiv e ablation study to in vestigate the con- tribution of each comp onen t within LGD-Net. The results are detailed in T able 2. T able 2: Ablation study of key comp onen ts: F eature Hallucination (Halluc.), Atten tion F usion (Attn.), and Domain Kno wledge Internalization (Bio-Reg). V ariant Halluc. A ttn. Bio-Reg Acc (%) F1 κ A (Baseline) ✗ ✗ ✗ 82.29 0.7935 0.7543 B (F eat. Align) ✓ Concat ✗ 92.53 0.9120 0.8914 C (Dynamic) ✓ ✓ ✗ 93.35 0.9344 0.8884 D (w/ Nuclei) ✓ ✓ Nuclei 94.27 0.9498 0.9239 E (w/ Mem b.) ✓ ✓ Mem b. 94.78 0.9522 0.9281 F (F ull) ✓ ✓ Both 95.60 0.9644 0.9453 Efficacy of F eature Hallucination: Simply in tro ducing the feature hallucination mo dule (V arian t B) results in a substan tial p erformance im- pro vemen t ov er the baseline (+10.24% in accuracy , 0.1371 in κ ). This in- dicates that the student enco der and the feature hallucinator successfully learns to appro ximate the molecular feature distribution of the T eacher. Impact of Domain Kno wledge: The in tegration of biological con- strain ts further enhances robustness. Sp ecifically , membrane intensit y reg- ularization (V ariant E) yields sligh tly heigher improv emen t than nuclei reg- ularization (V ariant D). This aligns with clinical guidelines, whic h identify 10 mem brane staining completeness as the primary determinant for distinguish- ing HER2 scores [14]. The combination of both constraints (V arian t F) ac hieves the highest p erformance, demonstrating that structural (nuclei) and molecular (membrane) cues pro vide complementary diagnostic information. 5. Conclusion In this pap er, w e presen ted the Laten t-Guided Dual-Stream Net- w ork (LGD-Net) , a nov el framework for accurate HER2 scoring directly from H&E stained images. T o address the inherent limitation of H&E images of the lac k of molecular information required to distinguish HER2 represen- tation levels, we prop osed to in tegrate feature-lev el hallucination instead of pixel-lev el virtual staining. The prop osed approac h demonstrates that b y hallucinating molecular features in the laten t space and constraining them with task-sp ecific domain knowledge, including the nuclei density and mem- brane staining in tensity , LGD-Net effectively recov ers diagnostic signals lost in standard morphological stains. Comprehensiv e experiments on the BCI dataset v alidate the effectiv e- ness of our metho d. LGD-Net achiev ed a significant p erformance gain o ver the unimodal H&E baseline (impro ving accuracy from 82.29% to 95.60% ) and outp erformed state-of-the-art dual-mo dal fusion methods. F urther- more, by eliminating the computationally exp ensiv e image deco der during inference, LGD-Net offers a highly efficient solution suitable for large-scale clinical screening. References [1] H. Sung, J. F erla y , R. L. Siegel, M. Lav ersanne, I. So erjomataram, A. Je- mal, F. Bra y , Global cancer statistics 2020: Glob o can estimates of in- cidence and mortality w orldwide for 36 cancers in 185 countries, CA: a cancer journal for clinicians 71 (3) (2021) 209–249. [2] A. G. W aks, E. P . Winer, Breast cancer treatmen t: a review, Jama 321 (3) (2019) 288–300. [3] A. C. W olff, M. E. H. Hammond, K. H. Allison, B. E. Harvey , P . B. Mangu, J. M. Bartlett, M. Bilous, I. O. Ellis, P . Fitzgibb ons, W. Hanna, 11 et al., Human epidermal gro wth factor receptor 2 testing in breast can- cer: American so ciet y of clinical oncology/college of american patholo- gists clinical practice guideline fo cused up date, Archiv es of pathology & lab oratory medicine 142 (11) (2018) 1364–1382. [4] S. Ahn, J. W. W o o, K. Lee, S. Y. P ark, Her2 status in breast can- cer: c hanges in guidelines and complicating factors for interpretation, Journal of pathology and translational medicine 54 (1) (2020) 34–44. [5] C. Karak as, H. T yburski, B. M. T urner, X. W ang, L. M. Sc hiffhauer, H. Katerji, D. G. Hicks, H. Zhang, In terobserv er and in terantibo dy repro ducibilit y of her2 immunohistochemical scoring in an enriched her2-lo w–expressing breast cancer cohort, American Journal of Clini- cal Pathology 159 (5) (2023) 484–491. [6] S. F arahmand, A. I. F ernandez, F. S. Ahmed, D. L. Rimm, J. H. Ch uang, E. Reisenbic hler, K. Zarringhalam, Deep learning trained on hemato xylin and eosin tumor region of interest predicts her2 status and trastuzumab treatment resp onse in her2+ breast cancer, Modern P athol- ogy 35 (1) (2022) 44–51. [7] S. A. Rasm ussen, V. J. T a ylor, A. P . Surette, P . J. Barnes, G. C. Beth une, Using deep learning to predict final her2 status in inv asive breast cancers that are equivocal (2+) b y immunohistochemistry , Ap- plied Imm unohisto c hemistry & Molecular Morphology 30 (10) (2022) 668–673. [8] N. Brieu, N. T riltsc h, P . W ortmann, D. Winter, S. Saran, M. Reb elatto, G. Schmidt, Auxiliary cyclegan-guided task-a w are domain translation from duplex to monoplex ihc images, in: 2025 IEEE 22nd International Symp osium on Biomedical Imaging (ISBI), IEEE, 2025, pp. 1–5. [9] B. Bai, X. Y ang, Y. Li, Y. Zhang, N. Pillar, A. Ozcan, Deep learning- enabled virtual histological staining of biological samples, Light: Science & Applications 12 (1) (2023) 57. [10] S. Liu, C. Zhu, F. Xu, X. Jia, Z. Shi, M. Jin, Bci: Breast cancer im- m unohisto c hemical image generation through pyramid pix2pix, in: Pro- ceedings of the IEEE/CVF conference on computer vision and pattern recognition, 2022, pp. 1815–1824. 12 [11] J. Qin, W. Y ang, Y. Su, Y. Zh u, W. Li, Y. Pan, C. Pan, H. Qi, Her2 expression prediction with flexible multi-modal inputs via dynamic bidi- rectional reconstruction, in: Pro ceedings of the 33rd ACM International Conference on Multimedia, 2025, pp. 2036–2043. [12] J. P . Cohen, M. Luck, S. Honari, Distribution matching losses can hal- lucinate features in medical image translation, in: In ternational confer- ence on medical image computing and computer-assisted in terven tion, Springer, 2018, pp. 529–536. [13] J. V asiljević, Z. Nisar, F. F euerhak e, C. W emmert, T. Lamp ert, Cy- clegan for virtual stain transfer: Is seeing really b elieving?, Artificial In telligence in Medicine 133 (2022) 102420. [14] Q. Peng, W. Lin, Y. Hu, A. Bao, C. Lian, W. W ei, M. Y ue, J. Liu, L. Y u, L. W ang, Adv ancing h&e-to-ihc virtual staining with task-sp ecific do- main knowledge for her2 scoring, in: In ternational Conference on Med- ical Image Computing and Computer-Assisted In terv ention, Springer, 2024, pp. 3–13. [15] K. He, X. Zhang, S. Ren, J. Sun, Deep residual learning for image recognition, in: Proceedings of the IEEE conference on computer vision and pattern recognition, 2016, pp. 770–778. 13
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment