MolHIT: Advancing Molecular-Graph Generation with Hierarchical Discrete Diffusion Models
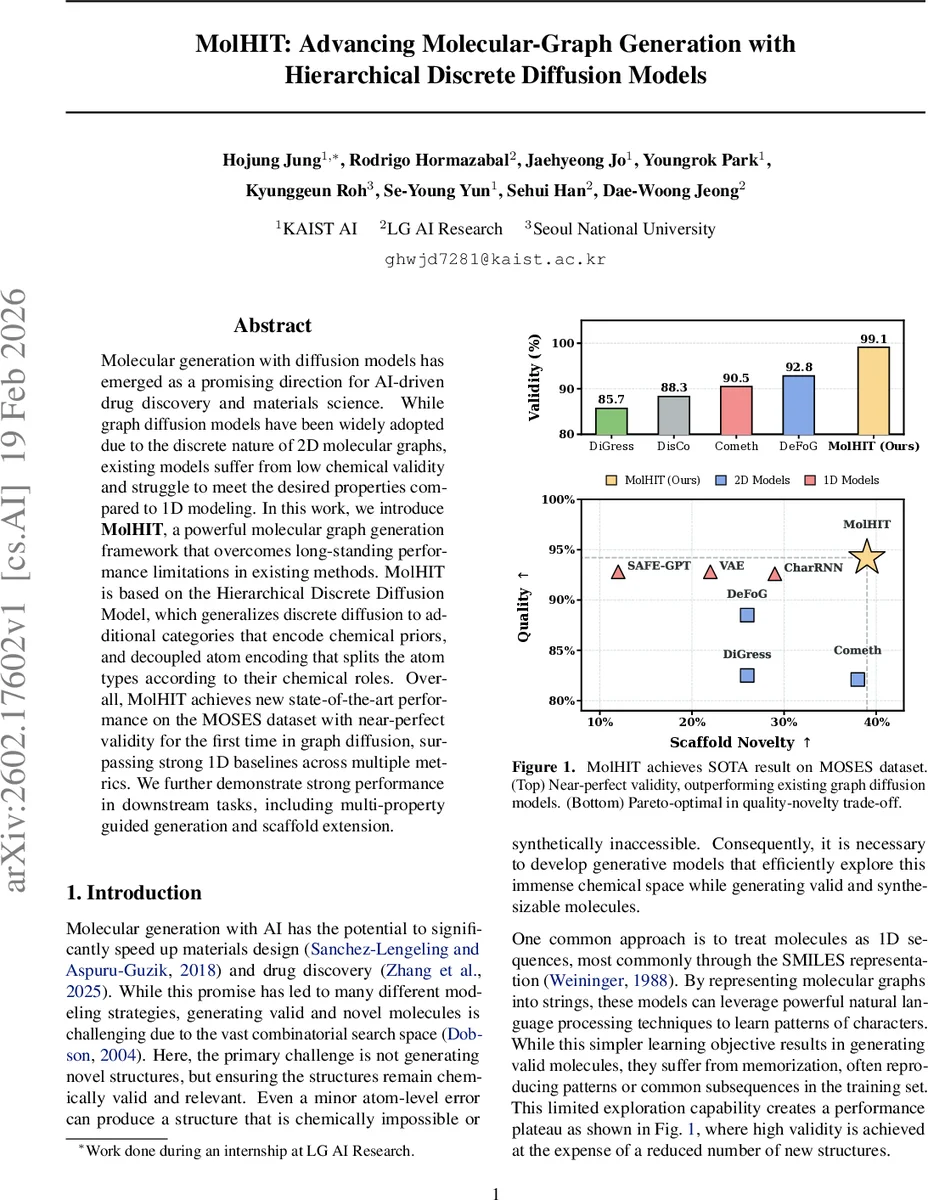
Molecular generation with diffusion models has emerged as a promising direction for AI-driven drug discovery and materials science. While graph diffusion models have been widely adopted due to the discrete nature of 2D molecular graphs, existing models suffer from low chemical validity and struggle to meet the desired properties compared to 1D modeling. In this work, we introduce MolHIT, a powerful molecular graph generation framework that overcomes long-standing performance limitations in existing methods. MolHIT is based on the Hierarchical Discrete Diffusion Model, which generalizes discrete diffusion to additional categories that encode chemical priors, and decoupled atom encoding that splits the atom types according to their chemical roles. Overall, MolHIT achieves new state-of-the-art performance on the MOSES dataset with near-perfect validity for the first time in graph diffusion, surpassing strong 1D baselines across multiple metrics. We further demonstrate strong performance in downstream tasks, including multi-property guided generation and scaffold extension.
💡 Research Summary
MolHIT introduces a novel framework for molecular graph generation that bridges the performance gap between graph‑based diffusion models and sequence‑based (SMILES) models. The method builds upon two key innovations: a Hierarchical Discrete Diffusion Model (HDDM) and Decoupled Atom Encoding (DAE).
HDDM extends the conventional discrete diffusion process by adding an intermediate state space (S₁) between the clean atom categories (S₀) and the absorbing masked state (S₂). A deterministic projection matrix Φ clusters the K atom types into G chemically meaningful groups. The forward transition at each timestep is a convex combination of three kernels: identity (I), the S₀→S₁ transition Q(1) defined by Φ, and the masking transition Q(2). By choosing monotonic schedules αₜ and βₜ (αₜ≤βₜ) the authors prove that the Chapman‑Kolmogorov equation holds, yielding a closed‑form expression for the cumulative transition matrix. This enables a principled derivation of the continuous‑time negative ELBO, which reduces to the standard masked diffusion loss when the intermediate states are omitted. In practice, atoms are first denoised from the masked prior to a coarse group label (S₁) and then refined to the exact atom token (S₀), providing a coarse‑to‑fine generation trajectory that respects chemical relationships among atom types.
DAE addresses the ill‑posed nature of previous atom encodings that relied solely on atomic numbers. By separating formal charge and aromaticity into distinct attributes, the token space expands to represent each chemically distinct variant (e.g., N,
Comments & Academic Discussion
Loading comments...
Leave a Comment