Foundation Models for Medical Imaging: Status, Challenges, and Directions
Foundation models (FMs) are rapidly reshaping medical imaging, shifting the field from narrowly trained, task-specific networks toward large, general-purpose models that can be adapted across modalities, anatomies, and clinical tasks. In this review,…
Authors: Chuang Niu, Pengwei Wu, Bruno De Man
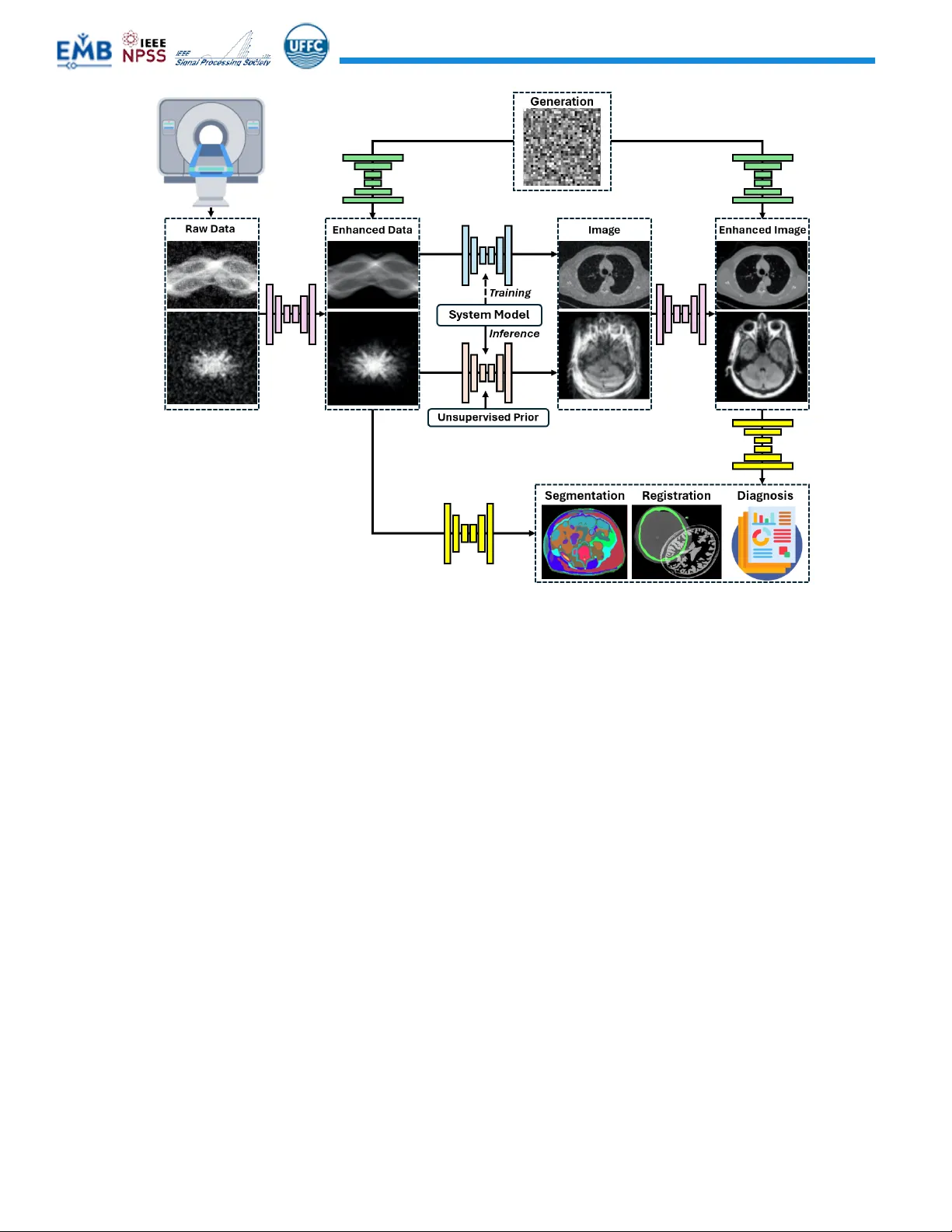
IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 1 Foundation Models for Medical Imaging: Status, Challenges, and Directions Chuang Niu , Member , IEEE , Pengwei Wu , Member , IEEE , Bruno De Man , Fell ow , IEEE , and Ge Wang, Fellow , IEEE Abstract — Foundation models (FMs) are rapidly reshaping medical imaging, s hifting the field from narrowly trained, task - specific networks toward large, general - purpose models that can be adapted across modalities, anatomies, and clinical tasks. I n this revi ew, we synthesize the emerging lands c ape of medical imaging FMs along three major axes: principles of FM design, applicatio ns of FMs, and forwar d - looking ch allenges and opportunities . Taken together, this revi ew provides a technically grounded, clinically aware, and future - facing roadmap for developing FMs that are not only powerful and versatile but also trustworthy and r eady for responsible translation into clinical practice. Index Terms — Foundation models , m edical imaging , i mage rec onstru ction , i mage anal ysis , m ultimodal lea rning I. INTRODUCT ION rtificial intelligence (AI) for medical imaging is experiencing a transformative shift from task - specific models towar d f oundation mo dels (FMs), wh ich are lar ge artificial neural networks pre - trained on vast, diverse datasets and adapted efficiently to a variety of downstream tasks. In medical imaging, where lab els are scarc e, heterogeneous, and expensive, FMs show a strong promise for rapid adaptation with minima l annotation, improved generalization across sites, scanners, and populations, and a plausible route to “generalist” medical i maging as sistants that r eason acr oss cont exts. Recent overviews from both the radiology and computer vision communities document a surge of FM research, spanning 2D/3D segmentation, image – text representation learning through vision – language fusion, and generative models. Together, these developments mo tivate a new synthesis of principles, capabilities, and translational considerations tailored to the healthcare ecosystem [1] . To c ontextualize foundation models, we begin by explor ing their relationship with the broader AI landscape, coupled with Figure 1.1 il lustrating the relative ti melines of the related ar eas along with s ome seminal publications . AI r efers to non - human systems performing tasks that mimic human perception and reasoning, such as language understanding and image analy sis. Machine learnin g, a s ubset o f AI, trains models to det ect C. Niu and P. Wu are co - first authors with equal contributio ns, B. De Man a nd G. Wang a re co - corresponding authors. C. Niu, and G. Wang are with Biomedical Imaging Center, C enter for Computational Innovations , Center for Biotechnology & Interdisciplinary Studies, Department of Biomedic al Engineeri ng, School of Engineering, Rensselaer Polytechnic Institute, Tr oy, New York, USA (e - mail: niuc@rpi.edu ; wangg6@rpi.edu ). P. Wu and B. De Man are with GE HealthCare Technology & Innovati on Center, Niskayuna, NY, USA (e - mail: pengwei.wu@gehealthcare.com ; deman@gehealthcare.com ). patterns in data, evolving from simple statistical methods to more sophisticat ed tools like random forests and s upport v ector m achine s. Deep learning uses multi - layer artificial neural networks to represent dat a in a data - driven fashion, leading to advanced architectures, like Convolutional Neural Networks (CNNs) , Recurrent Neural Ne tworks (RNNs) , Graph Neural Networks (GNNs ) , and Transformers. The term found ation mod el was coined by th e Center for Research on Foundation Models at the Stanf ord Insti tute for Human - Centered Ar tificial Intelligence in August 2 021 [19] . Foundation models are a class of deep learning models that ar e initially trained based on a diverse dataset for broad applications, and that can then be fine - tuned for spec ific down - stream applications. Typically, they are initially trained in a self - super vised fashi on. These pre - trained FMs then serve as the basis for developing task - specific models through transfer learning. The term fo undation model is sometimes used lo osely: a critical examination of the criteria for a model to qualify as a foundation model is give n in [20] . Foundation model s are charac terized by enormous training data and par ameter counts, which lead to emergent capabilities that do not present in smaller mo dels. In other words, a foundation model serves as a general - purpose platform that, with minimal task -s pecific training, can achieve strong performance across a variety of tasks. Another hallmark of foundation models is scalability. Their performance improves predictably as model size, trai ning dat a, and amount of compute increase, following empirical scali ng laws. This scaling yields surprising capabilities, e.g., GPT - 3 demonstrated in - context learning to solve tasks it was not explicitly trained for. Foundation models also e xhibit strong generalization and transferability, mean ing that the knowledge captur ed during pretraining on broad data can be transferred to unseen tasks. A single pretrained model can be fine - tuned to excel in applications ranging from natural language processing (NLP) to computer vision and robotics. This versatility has incentivized h omogenization of AI research around a few architectures, especially the Transformer. However, this also means any defects or biases i n a foundat ion model might propagate to its downstream uses. We fi rst introduc e s everal pr evious re view papers related to foundation m odels. A comprehensive survey of s elf - supervised A IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 2 Turing [2] , Deep Blue [3] , MYCIN [4] , SHR DLU [5] IBM [6] , Linear Regression [7] , S VMs [8] , Random Forests [9] LeCun [10] , AlexNet [11] , ResNet [12] , LST M [13] , GAN [14] , U - Net [15] Deep M ind AlphaFold [16] , Google M ed - PaLM [17] , Bommasani [1 8] . Fig . 1.1: Publication trends from 2000 to 2025 (Source: Scopus, Sep’ 2025) highlight the relative growth of Artificial Intelligenc e (AI), Machine Learning (ML), Deep Learning (DL), and Foundation Models (FM), alongside key milestones that shaped each fiel d. l earning (SSL) is provided by Liu et al in 2021 [21] . Two 2023 survey s by Kaz erooni et al. and by Yang et al [22], [23] provide in - depth overview s of t he rapidly evolving field of diff usion models, which are increasingly being integrated into foundation models. A more recent review of generative models is provided by Hein et al [24] . Lo ngpre et al. [25] pres ent a practical guide to suppo rt responsible and transparent development of FMs across text, vision, and speech modalities . L arge l anguage models (LLMs) are the most popular type of FMs . Zhou et al. [26] trace the evolution from BERT to ChatGPT, emphasizing key advancements in ar chitecture, training methods, and model capabilities . Zhao e t al. [27] summarize LLMs and emerging trends like multi - agent collaboration and chain - of - th ought reason ing. Ian A. Scott [28] introduces physicians to FMs and LLMs , explaining how they can perform diverse tasks across modalities (text, audio, images, video), with potential applications in medicine . Yang et al. [29] offer an overview of ChatGPT, BERT, and other LLMs , detailing their underlying architectures, trainin g strategies, and b road applications . Truhn et al. [30] explore how LLMs and multimodal f oundation models are transforming precision oncology . Some excellent reviews focused on medical imaging and image analysis . Azad et al. [31] explore how FMs are reshaping the field of medical imaging, including a structured taxonomy of FMs in m edical imaging and clinical applications, challenges, and directions. The vision - language mo dels (VLMs) are covered in several reviews, including Hua ng et al. [32] , Ryu et al [33] , and Sun et al [34] . They analyze how multimodal FMs are reshaping clinical AI by integrating visual data (e.g., X - rays, MRIs) with textual information (e.g., radiology reports, clinical notes). Zhang et a l [35] , Huix et al [36] , and Veldhui zen et al [37] survey FMs for me dical image analysis and outlin e unique challenges of applying them in radiology, pathology, and ophthalmology. Khan et al [38] present an analysis of subgroup fairness in medical imaging FMs, indicating that improved overall accuracy may come at the expense of reduced s ubgroup fairness. The latest milestone in this emerg ing field is the IEEE Transactions on Medical Imaging Special Issue on Advancements i n Foundation Models for Medical Imaging (2025), w hich assem bled 18 papers spanning se gmentation, multimodal integrat ion, arch itectura l innovat ions, benchmarking, ethics, and generative synthesis. Collect ively, these contributions underscore both the breadth and depth of current progress: from SAM - inspired segmentation frameworks and Mamba - based backbones to multimodal vision – language adaptations and large - scale echocardiography models; f rom topol ogy - guided generative pathology models to benchmark and et hical analyses that foreground fairness, interpretability, and governance. The Special Issue illustrates not only rapid technical advances bu t also the broader community recognition that foundation models for medical imaging and beyo nd mus t be judg ed by accu racy , equity, transparency, and clinical utility. This collection thus provides a valuable snapshot of the state of the art, while also mot ivating the n eed fo r integrativ e up - to - date reviews like the present article. This review advances t he current literature in three distinct ways. First, it adopts a broad coverage of FMs in medical imaging, especially incorporating the und errepresented do main of i mage reconstr uction for C T, SPECT, PET, MRI, ultrasound, and optical imaging . Second, b y integrati ng the most recent developments in this rapidly evolving field, our review addresses temporal gaps in prior surveys , such as generative AI, reinforcement learning, and modern reasoning methods for medical imaging researchers and practitioners. Final ly, we offer an extensive perspective as the last part that reflects our current vision to promote further advancement. The remainde r of this revi ew is organized as follows. In the next section, w e distill the principles behind FMs that are most relevant to imaging, ranging from major model architectures, common trai ning strategies, to the key components of development and deployment of FMs . In the third section, we survey applications across medical imaging modaliti es (CT, MR, PET, US, X - ra y, ophthalm ology , pathology) and tasks (segmentation, detection, diagnosis, triage, report generation, reconstruction), highlighting str engths, caveats, and challenges . In the final section, we identify future directions in terms of four pillars supporting medical imaging FMs , which are data/knowledge, model/optimization, computing power, and regulatory science. Overall, w e hope to provide a unify ing view that is technica lly grou nded , clinically actionable , and f orward - looking . IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 3 Fig. 2.1 . Principl es of Foundation Models. Overview of medi cal foundat ion models, illustr ating how large - scale heterogeneous clinical data, including medical images with asso ciated re ports, lab tests, genetics, and electronic health records, are use d to t rain scalable, genera lizable, and transferable foundation models that can be adapted to diverse downstream ap plications such as radiology, oncology, pathology, neurology, and cardiology. The lowe r panel summarizes the major technical components , including model architect ures, mode ling parad igms and t raining wor kflows, and efficien cy techniq ues for op timization and deploy ment. II. P RINCIPLES OF F OUNDATION M ODELS In medical imaging, FMs learn from large - scale, image - centric multimodal datasets along with associated radiology reports, laboratory results, genetic profiles, and electronic health record (EHR) data. These models support a wide range of clinical s pecialties, including radiology, oncology, pathology, neurology, cardiology, and so on. This section outlines the core principl es behind medical imaging FMs, covering model archit ectures, training strategies, and efficiency techniques, as summ arized in Figure 2.1. A. Model Arc hitectur es Several neur al network architectur es serve as the building blocks for foundat ion models. Bri efly speaking, Transformers have taken the lead in many domains, especially in NLP and image analysis. CNNs, however, often outperform Transformers on smaller - scale vision tasks w hen data is scarce, due to their built - in locality bias. For some tasks w ith very long sequential data, state space models (SSMs) like Mamba now show promising results, even surpassing Transformers of similar or l arger sizes. In this subsect ion, we review the major architectures, their variants, highlighting st rengths and limitations. 1) Transformer The Transfo rmer [39] has become the de facto core of most FMs in language and incr easingly in vision and medical imaging. Transformers dispense with recurrence and convolutions in favor of self - attention mechanisms that explicitly model long - range dependen cies. A Transf ormer block is typically composed of mult i - head self - attention layers and feed - forward layers, enab ling it to attend to a ll positions of an input sequence in parallel. Vision Tra nsformers (ViTs) [40], [ 41] token ize images and analyze them based on sufficient traini ng data. ViTs can achieve excellent per formance across vari ous vision tasks. ViTs’ strengths lie in their ability to capture global context easily via self - attention and their scalability with gradually refined attention coverage . However, since Transformers lack inductive biases, a ViT trained from scratch on limited data may underperform a CNN [41]. The Swin Transf ormer [42] was designed to address the ViT’s i ssues by computing self - attention in non - overlapping windows and shif ting the window positions between layers to allow cross - window connections to reduce the computation and improve generalization on smaller datasets. Another strategy of reducing the compu tation for high - resolution/dimension images is to interleave global and local window atten tion acros s layers [43]. Decoder - only Transformers [44] are simplified Trans formers where only the decoder stack is used. Through causally masked self - attention, each t oken can attend only to preceding tokens, making t hem inher ently autoregressive. Import antly, causal self - attenti on is powerful in deployment due to the ability to perform key - value caching, which brings several tangible advantages, such as huge speedup in inference and stable latency. Thus, it has be come foundation al for modern language models. The decod er only arch itecture is also critical in multimodal mode ling by dec oding vision and la nguage tokens for various tasks, such as image/video captio ning [45] a nd medical r eport ge neration [46]. The Mixture of Experts (MoE) architectur e [47] is an enhancement to Transformer architectures and replaces the … … Tr a i n i n g Applicati on • Sca lability • Gene ralizability • Tra n sf er ab i l i t y Medical Images Report s Lab Test Ge n e t i c EH R Radiolog y Oncology Pathology Ne u r o l o g y Ca r d i o l o g y Large - scale Data Diverse Applications Generative Foundation Models Ar chi t ect ur es Tr a i n i n g Efficiency Diff usion GAN VAE AR Discri minat ive SSVRL CLIP Masked JEPA ViT Decoder - onl y MoE ResNet CNN - Attent ion Mamba H3 RWKV Quantization Effici ent At tention Disti llat ion FSDP / ZeRO Mixed Precision Checkpoi nting FlashAttent ion RetNet Pre - Traini ng Generative Discri minat ive Training Workflow Modeling Paradigms Transformer Convolut ional Model s State Space Model s Effici ent O ptimiz ation Effici ent Depl oyment UNet Kernel Fusi on Gradient Accu mulation vLLM / SGLang / Oll ama RL PPO GRPO DPO Post - Traini ng SFT RL LoRA 4 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 standard feed forward layer in each Transformer block with a set of parallel sub networks called experts and a learnable gating network that routes each input token to a small subset of experts based on token - specific features, dramatically inc reasing mode l capacity by only activating a fraction of the total parameters per token. Each gate typically employs a sparsely gated softmax function along with auxiliary load balancing losses and bias terms to distribute tokens evenly across experts [48]. This architecture enables training models with trilli ons of parameters, where only a few billion of t hose parameters are used during inference for any given token, making MoE an enabler of scalability in modern FMs. Efficient attenti on mechan isms address the quadratic cost of standard Transformers by reducing computation and memory while preserving ess ential context. Spa rse attention limit s ea ch token’s receptive field through structured or adaptive sparsity [49]. Lin ear attention replaces softmax with kernelized or alternative formulations so attention scales linearly in sequence length [50], [51]. Low - rank and factorized approaches compress keys, values, or the attention matrix itself [52]. Meanwhile , multi - query and group - query attention [53], [54] share key – value projections across heads or head groups, boosting memory efficiency and throughput wit h minimal loss in expressiven ess. Together, these methods form a toolkit that enables scalable Trans formers for l ong - con text, high - resolution, and multimodal tasks. 2) Convolution - based Models Convolutional Neural Networ ks (CNNs) [55] dominated computer vision for years, and they remain highly relevant in the era of FMs. CNNs like ResNet [56] and UNet [15] are g ood at learning local pat terns and are tr anslation - invariant. These inductive biases al low CNNs to generalize well with relatively small training datasets and excel on medium - scale tasks with strong local features. However, CNNs have a restricted receptive field, so localized convolutions migh t miss global context. Attention - Convolut ional Models [57] aim to ge t t he best of both CNNs and Transformers, such as by augmenting CNN backbones with attention blocks [58], or by augment ing Transformers with convolutional token embeddings [59]. However, they could also inhe rit some limitation s of b oth paradigms. Nonetheless, these hybrid models form an important class of FM architecture. 3) State - Space Models Recurrent Neural Networks (RNNs) [60] we re the wor khorse for sequence modeling. However, RNN and its variants like Long Short - Term Memory (LSTMs) and Gated Recurrent Unit (GRUs) [61] faced challenges in capturing long - range dependencies. In particular, they could not be parallelized across sequence positions. To this end, State space models (SSMs) provide a powerful framework for sequence modeling by representing how hidden states evolve over time in response to inputs, offering a fundamentally recurrent alternative to attention - based architectures. Instead of computing pairwise interactions across all tokens, SSMs propagate information through a structured state update that can be computed efficiently in linear time, making t hem well - suited for v ery l ong sequences and streaming scenarios. The modern resurgence of SSMs bega n with the Structured State Space sequence model (S4) [62], which introduced stable diagonal - plus - low - rank parameterizations enabling long - range memory and efficient convolutional impleme nt ations. This foundati on has since driven the development of highly expressive, scalable architectures such as Selecti ve SSMs (Mamba) [63], RWKV [64], H3 [65], and RetNet [66], which have evolved into competitive sequence learners capable of matching or sur passing t ransformer performance in long - context tasks while offering significant advantages i n scalability and memory efficiency. SSMs have been successf ully adopted in medical imaging [67], [68]. B. Modeling and Trai ning 1) Modeling Paradigms We can d ivide the v arious modeling methods for FMs into generative and discriminative/ contrastive paradigms. Generative models provi de a full under standing of data and produce new examples, whereas discriminative/contrast ive models excel at p roducing gene ralizable representations and making de cisions. Genera tive mo dels of fer t ools f or data generation, uncertainty quantification, and discovering underlying data structure, which can be invaluable in improving medical imaging qua lities. Contra stive, as a t ype of discriminative representation learning approach, currentl y dominate t he pret raining for image analysis tasks such as classification, segmentation, detection, and regression, underlining predictive accuracy. Generative Mo deling Variational Autoencoders (VAE) [69] exe mplifies an e arly latent - variable generative approach that marries probabili stic models with deep learnin g. A VAE consists of an encoder network that m aps input data to a latent distribution and a decoder network that reconstructs the data from a latent sam ple, trained jointly by maximizing a variational lower bound on data likelihood. This framework enables learning a deep latent representation while permitting effective and efficient inference. VAEs have been pivotal as a principled method for learning unsupervised generative models of images, offering stable training and explicit probability density estimation. Furthermore, some extensions of VAEs were proposed; e.g. β - VAE [70] for disentangled facto rs, VQ - VAE [71] f or discrete latent spaces. VAEs generate new samples rapidly but often at a relatively compromised quality in comparison with more recent models introduced below. In generative foundation models, VAEs serves as an important method for vision tokenization/c ompression [72]. Generative Adversarial Networks (GANs) [73] are a major step forward for generative modeling, form ulated as a minimax game between a generator that synthesizes data and a discriminator that distinguishes real from fake data. Successive innovations such as DCGAN [7 4], Progress ive GAN [75], StyleGAN [76], and BigGAN [77] improved training st ability, scale, and controllability. In medical imaging, GANs have been widely adopted fo r cross - modalit y transla tion [78], super - 5 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 resolution [79], denoising [80], inpainting [81], and simulation [82], serving as both a critical modeling paradigm in the evolution of FMs and a practical tool in medical imaging. While GANs remain attractive for their efficiency at inference, they often suffer from training difficulties and mode collapses. Diffusion models [8 3] are a r ecent class of generat ive models that achieve state - of - the - art results. These models define a forward process that gradually adds noise to every image or sample in a training dataset until it becomes pure noise. Then, a learned reverse process will remove noise gradually to synthesize a new image or sample. The seminal Deno ising Diffusion Proba bilistic Model (DDPM) [83 ] d emonstrated that diffusion models can produce excellent images, typically outperforming GAN results while off ering advantages like stable training and distribution coverage. A main drawback of DDPM is the compu tational c ost: genera ting an imag e requires many iterative denoising steps, making them slower than one - shot generators like VAEs and GANs. Recent research addresses this drawback with optimized samplers which allow fewer or even just one step, such as latent diffusion techniques [84] which diffuse in a lower - dimensional latent space to speed up generation and consist ency models [85] which remove noise in on e or few steps. Diffusion models have rapidly been adopted in medical imaging for various tasks, such as image reconstruction and enhancement [86]. These diffusion models are based on thermodynamics and have been extended in reference to electrodynamics an d other mechanisms [86]. Autoregress ive (AR) [87] generativ e mod els treat data synthesis as a sequential prediction problem, modeling the joint distribution of high - dimensional data as a product of conditionals. In natural language processing, this framework underlies the next - tok en prediction mechanism in LLMs, where each token is generated by conditioning on all previously generated tokens [44]. In computer vision, early examples include PixelRNN [88] and PixelCNN [89], which demonstrated that images can be generated pixel - by - pix el by scanning an image field and predicting the next pixel intensity conditioned on the context. However, they are notoriously slow since all output elements are produced sequentially. Recent advances demonstrate that images can be first compressed into d iscrete latent codes and then modele d with a Transform er as an autoregressive sequence of tokens, as illustrated by DALL·E [90] for text - to - image generation. This paradigm has proven successful in natural language processing first and more recently for ima ge generati on [91], and even i n multi - modal tasks [92]. Discriminati ve Modeling Self - supervised visual representation learning (SSVRL) exploits large - scale unlabeled images to learn features. Discriminati ve self - supervised methods are essential for SSVRL. These methods do not attempt to model the input distribution fully; instead, they train neural networks on pretext tasks such that solving these tasks requires extracting high - level semantic features. One prominent class of methods is contrastive learning, exemplified by methods like CPC [93], SimCLR [94], MoCo [95], and PI RL [96]. Fu rthermore, the teacher - student learning paradigm, such as in BYO L [97], SimSiam [ 98], and DINO [99], took a surprisi ng s tep by removing explicit negative pairs in contrastive learning. Despite the absence of con trasting against nega tives, these methods avoid collapse thr ough their asymmetric teacher - versus - student architecture. Clustering - based methods [100], [101], [102] leverage self - supervised visual representation learning, achieving excellent results. Recently, inform ation maximizati on methods emerged as a promising direction for self - supervised learning due to their simplicity without contrastive negative examples nor asymmetric design [103], [104]. Self - supervised discriminative learning enables rich feature learning from unlabeled datasets, greatly r educing the need for costly annotations in medical imaging [105], [106]. Vision - Language Contras tive Learning lea rns joint representations from paired images and text. These approaches, such as CLIP [107] and ALIGN [108], extend contrastive learning to multimodal data: an image and its accompanying caption form a positive pair, and mismatched image – caption combinations f orm negatives. By leveraging extremely large datasets of image – text pairs, these models learn remarkably general and transferable visual features. After CLIP - based multimodal pretraining, the image e ncoder may us e zero - shot training for classification. By training on noisy but abundant web dat a, t hese models e ncode a rich associ ation between visual concepts and natural language, enabling important downstream applications [107]. In the medical imaging field, analog ous approaches have been extensivel y explored, e.g. aligning radiology images w ith report text, to bring the benefits of multimodal pretraining to speci alized domains [109]. Generative - Disc riminative Modeling Masked autoenco ders reconstr uct missing or corrupte d portions of t he input. A prime example in NLP is BERT [110], which learns a deep bidirectional Transformer by mas king out random words in a sentence and training the model to predict missing tokens. In c omputer vi sion, the Masked Autoencoder (MAE) [111] blocks a fraction of image patches and then reconstruct them, even outperforming supervised pre - training on the same architect ure for downstream t asks. The MAEs are optimized vi a reconst ruction error rathe r than likelihood. As such, they straddle generative and discriminative paradigms: the training objective is generative, but the resulting encoder is typically used for discrimin ative tasks. Joint - Embedding Predictive Architecture (JEPA) [112] builds on the idea of a “world model”, learning by predicting future or missing higher - level representations. A recent instantiation is I - JEPA [113].Another example is data2vec [114]. These approaches define an appealing middle ground that combines embedding , alignment, and prediction, learning both generative and discriminative features across modalities to support intelligent behavior. This paradigm resonates strongly with the Bayesian br ain hypothesis [115 ] and the minimum free - energy principle [116], which similarly view intelligence as predictive modeling of latent structure in the world. Reinforcement Le arning Reinforcement learning (RL) has be come critica l in trai ning 6 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 FMs, providing a principled mechanism to optimize model performance beyond supervised learning. While large - scale pretraining equips models with broad linguistic and world knowledge, RL enables them to incorporate explicit evaluative signals — ranging from h uman f eedback to ver ifiable task - based rewards — into their policy updates, thus facilitating goal - directed refinement of model capabilities. This transition from next - token prediction to preference - aligned optimization reflects a most important trend in fou ndation model research, where performance reliability, controllability, ge neralizati on, and interpretability incr easingly depend on an iterative feedback loop [117], [118]. RL - based methods help ensure that model outputs adhere to human - preferred behaviors, s afety norms, and i nteraction standards. The canonical RLHF pipeline centers on Proximal Policy Optimization (PPO) [119], paired with learned reward models de rived f rom human prefere nce dat a [120]. This KL - regularized objective enables stable policy updates while preventing di vergence from a reference policy. More recently, Direct Preference Opt imization (DPO) [121] has emerge d as a compelling alternative, reformulating the KL - constrained preference - alignment objective into a tractable supervised - learning – style loss that eliminates the need for value estimation and on - policy sampling. Complementary formulations, such as ORPO [122], fur ther strea mline prefere nce optimizat ion by merging likelihoo d trai ning wi th pref erence modeling. In medical imaging, t hese methods were applied to radiolo gy report generation [123] and radiology question answering systems [124]. For reasoning - intensive domains, mod els are optimized for verifiable correctness and multi - step reasoning quality. This shift has catalyzed the adoption of Group Relative Policy Optimization (GRPO) [125 ] and its variants. These met hods compute relative advantages among multiple sampled trajectories for each prompt, thereby avoiding explicit critics and improving stability in domains where correctness signals are sparse but reliable. Enhancements such as DAPO [126] and GSPO [127] furt her refine gr oup - based policy - gradient dynamics for large - scale reasoning optimization. Beyond GRPO, a growing l iterature explores RL wit h verifiabl e rewards [128], tree - search – augmented RL [129] , self - play – driven reasoning improvement [130], and of fline RL for complex reasoning trajecto ries [131]. Collectively, these methods position RL as a corner stone for advancing FMs from superficially coherent reasoning to demonstrably correct, logically structured problem solving. RL - based reasoning methods have been expl ored for medical i m aging applications, such as medical image question answering [132] and personalized lung cancer risk predict ion [133]. C. Training Workfl ow A typical traini ng workfl ow of FMs involve s a l arge - scale pre - training stage for learning generalizable representations, followed by an iterative post - training process including supervised fine - tuning (SFT) and reinforcemen t learning for alignment and/or reasoning. Although different types of FMs may adopt varie d strategie s, this two - stage paradigm remains the dominant develop mental framework. Pre - training on Broad Data : The pre - training stage typically leverages large - scale, heterogeneous datasets t o learn robust representations and capture complex data distributions. For text and vision transformers, this involves billions of text tokens or millions of images, optimize d through self - supervised objectives such as masked language modeling (BERT), next - token prediction (GPT), or contrastive alignment (CLIP). In biomedical domains, “broad data’’ further includes large collections of clinical text, medi cal images, and multimoda l corpora, enabling the development of domain - specialized models such as Bi oLMs [ 134], MedSAM [ 135], and M3FM [136]. In many cases, domain - adaptive pretraining is performed beforehand where models are furt her pretrained on lar ge in - domain corpora (e.g., Bi oBERT [134] trai ned on PubMed) to i mprove handling of domain - specific terminology and semantics. The pre - training of generative models such a s diffusion models [84] and GANs [137] learns to approximate the underlying data distribution closely e nough to generate realistic samples or reconstruct missing or corrupted information. This generativ e pre - training enables models to internalize fine - grained structural and semantic patter ns, while providing additional benefits unique to synthesis - based objectives. In medical imaging, s uch pre - trained generative models have been s uccessful ly applied to me dical image reconstruction [138], super - resolution [139], denoising [140], and robust artifact correction [141]. Despite its computationa l complexity, pretraining is essential to extract implicit, rich, and transferable features a nd knowledge for downstream adaptation [136]. Supervised Fine - Tuning : After large - scale pre - training, models are special ized for target doma ins or d ownstream task s through fine - tuning on smaller, high - quality labeled datas ets, typically with reduced learning rates to preserve general features. A central challenge is balanc ing specialization with the preservatio n of g eneral knowled ge. Risks of overfitting and catastrophic forgetting are address ed through sever al effective techniques, such as layer freezing, adaptiv e optimization, and parameter - efficient adaptation methods (e.g., LoRA [142]). Reinforcement Learning for Alignment and Rea soning : A cr itical aspect in the deve lopment of FMs is alignment, which ensures that outputs are not only plausible and fluent but also accurate, reliable, and consistent with domain standards and human values . A widely adopted paradigm is Reinforcement Learning from Human Feedback (RLHF), wherein human evaluators provide preference rankings of model outputs that are distilled into a reward model, subsequently optimized via reinforcement learning algorithms [119]. More recently, reinforcement learning has been ren ovated to improve reasoning quality. Reasoning - oriented models such as DeepSeek - R1[143] embody this shift of reinforcement learning [125] to encourage multi - step problem solving, self - consistency, and robu stness in complex decision - making tasks. This shift underscores the versati lity of r einforcement le arning both as an alignment mechanism and as a r oute to strengthen reasoning performance, an essential property for high - stakes 7 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 medical imaging appli cations. In some t raining pipel ines, data generated after RL is fed back into the SFT dataset, creating an iterative loop that progressively improves real - world performance [143]. D. Computational Efficiency The rapid scaling of FMs has intensified the need for computationally efficient training and cost - effective deployment, especially for medical imaging applications where the data are of high dimensions. Efficiency is now a foundational design goal rather t han a secondary consideration. Although some archit ectural effici ency designs such as MoE and efficient attention mechanisms have been introduced, this section reviews critical and popular techniques used to optimize training and inference pha ses of large - scale models. 1) Efficient Opt imization Training FMs i s computation ally challen ging. Effici ent optimization techniques address this challenge by reducing memory overhe ad, improv ing paral lelism, a nd levera ging hardware capabilities, with some important techni ques reviewed below. Sharded Training and Memory Partitioning : Fully Sharded Data Parallelism (FSDP) [144] and ZeRO - based [145] optimization represent two st ate - of - the - art strategies for memory - efficient distri buted training. These methods partiti on model par ameters, gradie nts, an d optimi zer sta tes ac ross devices, enabling training of models that exceed the memory limits of individual GPUs. The ZeRO family (ZeR O - 1/2/3) further allows flexible control over the trade - off between memory savi ngs and i nter - device communication. Parameter - Efficient Fin e - Tuning : Low - Rank Adaptation (LoRA) [146] and its variants [147] reduce the computational and storage cost of fine - tuning by introducing lightweight low - rank matrices into Transformer modules. Only these small sets of parameters are updated during training, allow ing adaptation to new tasks with orders - of - magnitude fewer trainabl e weights. Efficient Attention and Kernel Imple mentations : FlashAttention [148] provides an optimized, memory - efficient attention implementation that minimizes redundant data movement, direct ly improving traini ng speed. Similarl y, kernel fusion techniques [149] combine multiple GPU operations into a single execut ion step, reducing kernel - laun ch overhead and improving hardware u tilization. Mixed Precisi on a nd Act ivati on Che ckpoin ting : Mixed - precision trai ning [150] with FP16 or BF16 formats reduces memory foot print a nd incre ases ari thmetic throughput, particularly on modern Tensor Core devices. Activation checkpointing [151] further reduces memory use by storing only a subset of inter mediate a ctivati ons and rec omputing others during backpropagation. Together these techniques allow larger batch sizes and deeper models to be trained with the same hardware resou rces. Gradient Accumulation : When batch sizes exceed GPU memory capaci ty, gradi ent accumul ation simu lates lar ge - batch training by aggregating gradients across several forward passes [152]. This enables stable optimization behavior without requiring large accelerator clusters. 2) Efficient De ployment Once training i s complete, mode l deployment mus t balance performance, latency requirements, and hardware constraints . Efficient deployment focuses on compressing models, reducing precision, and leveragi ng specialized inference engines to achieve high - throu ghput, low - cost inference. Model Compr essio n via Dis tillat ion : Knowledge distillation [153] transfers the behavior of a large “teacher” model to a small er “student” model. The resulti ng s tudent model ret ains much of the t eacher’s predict ive capab ility wh ile requiring substantially fewer parameters and reduced comput ing. Distillation is especially effectiv e for edge d evices, mobile pl atforms, and lat ency - sensitive applications. Quantizati on for Reduced Precision Inference : Quantization converts model weights and acti vations from high - precision formats ( e.g., FP32) t o lower - precision representations such as INT8 or INT4 [154]. Modern quantization - aware and post - training quantization technique s maintain accuracy while d ramat ically improving inference throughput and lowering memory usage. These methods have become standard in production - scale model serving. Specialized Inference Runtimes : Recent advancements in inference system design have led to optimized runtimes tailored specifically for LLMs and multimodal LLMs. • vLLM [ 155] introduces PagedAttention, an effi cient memory management mechanism t hat impro ves multi - request batching and maximizes GPU utilization. • SGLang [156] ext ends these ideas by incorporat ing structured caching and partial - decoding reuse, enabling hi gh - throughput, multi - tenant d eployment scenarios. • Olla ma focus es on ease of dep loyment f or loc al environments, particularly for quantized models running on consumer hardware. These syste ms provide substantial impr ovements over traditional d eep learning servers, achieving significantly higher throughput for practical ap plications. III. M EDICAL I MAGING A PPLICATIONS In this section, we survey successes and challenges of FMs in medical imaging for various modalities and tasks as shown in Fig. 3.1, and we list the major benchm arks and medical imaging data platforms for development of FMs. Med ical imaging methods often f ace the long - tail data scenario, caused by heavily imbalanced datasets in which many common disease cases coexist with fewer rare disease cases. Consequently, the scarcity of data for training models to accurately identify these rare cases leads to perform ance degradation. The few - shot setting of FMs aligns perfectly with this long - tailed scenario, serving as a versatile base for a wide range of i maging modalitie s, anato mies, and downstre am tasks. A. Image Reconstructio n and Enhanceme nt Medical image reconst ructio n involves solvin g i nverse problems to recover high - quality images from incomplete or corrupted data. For example, reconstructing images from undersampled k - space signals or noisy/incomplete sinograms 8 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 [157], [158], [159], [160]. FMs are increasingly researched for this purpose to improve reconstruction quality under challenging conditions for various imaging modalities. Ins tead of t raining a separate network for every scanner or protocol, a single large model can be pretrai ned on div erse imag e data an d then adap ted to specific reconstruction tasks. This section focuses on three important types of foundatio ns models for image reconstruction & enhancement tasks as shown in Fig. 3.2. Fig . 3.1. Example applications of foundation models in di fferent aspects of medical imaging, across dif ferent modalities, anatomies, and tasks. 1) Image Enhancem ent These types of networks dir ectly improve the data qual ity, either in a measurement or image domain, or in bot h domains as shown in pink in Fig. 3.2. Common examples include sinogram completion and inpainting, K - space completion, denoising [80], [161], supe r- resolution [79], artifacts correction, inhomogeneity correction, and harmonization networks [162]. They do not require explicitly def ined forward models, s ince the y are not solving an inve rse probl em. Early attempts have demonstrated that deep learning can greatly enhance image quality in specific scenarios. However, these models typically are limited to pre - defined tasks or selected anatomic reg ions, and they often suffer from poor generalizability for out - of - domain tasks. Recently, there has been an increasing interest in FMs for multiple image enhancement tasks. TAMP for example leverages a physics - drive pre - training and parameter - efficient adaptation for universal CT image quality improvement in both sinogram and image domains [163 ]. 2) Direct Reconst ruction Researchers began leveragi ng la rge, paired datasets to directly learn mappings from measurements (e.g., sinograms in CT, k - space in MRI) to the final clean, high - quality images in an end - to - end fashion [ 164]. These approaches focus on minimizing recons truc tion and other losses (e.g., L1, L2, perceptual [165], adversarial [165]) during the training procedure. They are typically not i terative in nature and can use imaging system mo dels to enforce physical cons traints. Compared to image enhancement networks , r econstructi on networks can incorporate the measurement operator and observed data. For example, the “Reconstruct Anything” model proposes a universal direct inversion model by introducing a new conditioning mechanis m tha t in tegrates the imagi ng physics through multigri d Krylov iterations [166]. This single backbone model performs multiple image reconstruction and enhancement tasks. Another prominent example is the unrolled network framework type of approach, which mi mics itera tive optimization algorithms (e.g., ISTA, ADMM) by embedding data consistency and learned regularization into a trainable architecture [157], [ 158], [159], [160]. Finally, another line of work aims to directly learn the inverse model with dee p le arning (i.e., not explicitly providing the imaging physics / forward model to the network), for example, iRadonMap [167], [168], hierarchical DL reconstruction [169], and AUTOMAP [170], [171]. However, making the network learn an inverse models can be chal lenging due to their dime nsionality. We distinguish these methods from image enhancement methods since they involve a domain change between input and output (e.g., from K - space to image domain). This is still an emerging application for foundation models. 3) Prior Modeli ng The fina l cate gory of networks focuses on prior dist ribution modeling. S uch a model c an help th e inversi on/recons tructio n process significantly. Instead of relyi ng on paired data, they can naturally be applied to different inverse pr oblems without fine - tuning. Most of these methods ne ed the forward model to enforce various conditions such as tomographic data to finish tomographic reconstruction. While most of these methods are iterative, they are not the defining characteristics of the proposed method; for example, a conditional diffusion model trained to correct for different typ es of artifacts would still classify as an image enhancement task, despite being iterative in nature. One popular line of work utilizes score - based models to estimate the unconditional score function of the prior distribution. Then, during inference, some f orm of measurement matching technique i s used to condition t he reverse diffusion with a closed - form, approximated measurement matching scor e. Some example s are DPS, Scor e ALD, Score - SDE, Repaint (for inpainting only), BlindDPS, DDRM, PFGMs [24], DDS [172], Bl aze3DM [17 3], and FORCE [174]. Another line of work uses plug - and - play networks [175], [176], which use a pre - trained denoiser to regularize an iterative reconstruction process of solving inverse problems, as opposed to using a hand - crafted regularizer such as total variation [177] or wavelet spar sity [178]. While all the above - mentioned methods can potentia lly b e used for various applications (thus qualif ying as FMs), the first two categories often require retraining for new applications. In contrast, the third category offers a general framework for arbitrar y inverse problems without ret raining nor fine - tuning, IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 9 Fig . 3.2. Illustration of foundation models for different tasks in medical imaging. The center portion shows three types of founda tion models for image reconstruction & enhancement. Pink: image (or data) enhan cement model. Blue: direct reconstruction model. Or ange: prior distribution model. The other portions show data generation models (green) and data analysis models (yellow). likely at a cost of slow inference speed. Overall, while the use of FMs for tomographic reconstruct ion is still emerging, the trend is clear: a single large model (often generative) can flexibly handle multiple inverse problems by virtue of comprehensive prior knowledge. In major clinical areas, this means fa ster scans (by reconstructing from sparse data), lower radiation and contrast doses, higher image quality, and better diagnostic performance. B. Image Analysis In me dical imag e analysis, innovative FM techniques are continually emerging in various clinical tasks. A few representative applications are summariz ed here. 1) Classificat ion and Regres sion FMs have dr iven advances in medical image class ification and r egression, enabling diagnostic prediction and feature discovery with minimal supervision. Early deep learning models alre ady matched exp ert perfor mance in task s like disease detection from i mage s, but they required l arge, labeled datasets. FMs address this challenge by leveraging self - supervised pretraining on vast unlabeled datasets, often coupled with text, to reduc e labeling efforts in variou s domains. In radiology, the CheXzero model [179] was trained on hundreds of thousands of chest X - rays and their clinical reports using contrastive vision - language learning. CheXzero achieves zero - shot pathology classification, i.e., it can detect diseases that were not annotated, reaching area - under - curve (AUC) values around 0.95 f or several findings on external X - ray datasets. In oncology, FMs are used to discover imaging biomarkers from radiological scans. Pai et al. trained a self - supervised encoder on 11,467 dive rse tum or images [180], yielding a model that outperformed conventional supervised methods in pr edicting clinical biomarkers , especia lly in low - data regimes. In pathology, FMs are al so pivotal for classification like cancer subtypi ng. For example, Prov - Gig aPath [181] achieved state - of - the - art accuracy on 25 of 26 tasks in a pathology benchmark (covering cancer subtypes and “pathomics” predict ive t asks), significantly outperfor ming prior methods on the majori ty of t hose t asks. Finally, regression - based appro aches remain central to cancer risk assessment and survi val prediction, as indicated by studies such as DeepSurv [182] and TabSurv [183]. 2) Segmentation and Detection Early attempts at deep learning for segmentation has relied on task - specific models, each requiring laborious annotations [184]. FMs are redefining this area by learning generalized segmentation cap abilities across diverse organs and m odalities. Several st udies explored how large - scale pretrained models can generalize across segmentation tasks in medical imaging. Ma et al. [185], and Noh et al [186] provide over views of FMs f or segmentation tasks, including tumor detection, organ delineation, cell segmentat ion, and anomaly identification. The Segment Anything Model (SAM) [187] is a prompta ble 10 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 paradigm and t he first FM for general - purpose image segmentation. However, applying it naively showed limited accuracy on many medical images [188]. To bridge this gap, researchers developed medical - domain variants like MedSAM [189], a FM for “universal” m edical image segmentation, which was pretrained on 1.5 7 mi llion image – mask pairs spanning 10 imaging modalities and over 30 disease types. It was evaluated favorably on 86 internal tasks and 60 external test tasks. Other attempts include MedLSAM [190], 3DS AM - Adapter [191], SAM - Med2D [192], SAMed - 2 [193], and SAM - U [194], enhancing the sensitivity and specificity of medical image segmentation tasks. However, prompt able segmenta tion can be t ime - consuming in 3D cases fo r large cohort data analysis. Given that b oth SAM and SAM2 (video variant of SAM) handle either single or a stream or 2D images, their performa nce is typically worse than specially traine d 3D models. To address this issue, various 3D medical segment ation models (with and without prompts) were proposed. Promptless models t ypically support automated segmentation of most human anatomies and major pathologies directly [195]. For example, Brain SegFounder uses SwinUNETR for 3D neuroimage segmentation tasks [196]. VISTA3D i s a 3D segmentat ion mo del, whose performanc e was bo osted by d istilling sta te - of - the - art 2D i mage segmentation with supervoxels [196]. 3) Registration Conventional regi stration al gorithms (e.g. de formable registration) are generic but slow [197], while recent learning - based methods are fast but tend to overfit [198], [199], [200], [201]. Emerging FMs deliver state - of - the - art results. UniGradICON[202] , [2 03] is an early FM for medical image registration, trained on a dozen public d atasets covering various anatomies. This model achieved high accuracy across multiple registration tasks (e.g., aligning brain MRI scans as well as thoracic C T scans). Similarly, Hu et al enhanced the robustness and generalizability of registration using a FM after sharpnes s - aware minimization [204]. Another l ine of work focuses on perf orming ze ro - or few - shot transfer learning using pretrained vision models. For example, DINO - REG uses the feature maps from DINO (a natural image foundation model) to compute the registration loss [205]. MultiCo3D levera ged anatomical information from the SAM model to guide registration (via aligning semantics) [206]. FoundationM orph utilizes a pretrained vision - language model to guide registr ation and a multi - dimensional attention module to fuse vis ion - language represent ations [205]. C. Image Generation Image generation especially relevant in me dical im aging due to data scarc ity including data im balances (rarity of certain medical conditions and/or populations) , high human - annotation cost, and patient privacy concerns [207]. By creating artif icial yet rea listic medical images with generative AI, one can greatly reduce the dependency on real patient data for training powerful deep learni ng models [208], [209], virtual clinical trials [210], [211], and training medical professionals [212]. FMs have revolutio nized image generation by enabling scalable, high - fidelity synthesis across diverse modalities. Recent efforts demonstrated that generative AI can synthesize high - quality chest X - ray images, 3D MR and CT images, 2D pathology images, and so on [213], [214], [215], [216], [217], [218]. We highlight two rese arch aspects of image generation: Model Arc hitect ure & Con dition ing: Ea rly att empts a t medical image generati on typically rely on generative adversarial networks (GANs), which – while powerful - suffered from issues like mode collapse, difficulty in training, and limited sample diversity [ 219]. Transformers have then been used (e.g., Med - Art [220], Trans Med [221] , MedFormer [222]) to offer global context modeling and scalability. Recently, diffus ion - type models h ave emerged for high - quality image synthesis [213], [214], [215], [216], [217], [218]. For example, MINIM has presented a unified text - conditioned latent diffusion model for different domains, including OCT, chest X - ray, and CT [218]. In terms of new conditioning a pproaches (i.e., controlled generation): one line of work uses text - to - image diffusion models adapted to medicine. For example, RoentGen [223] fine - tunes a popular latent - space vision – langu age diffusio n model (Stable Diffusi on , trained on natural images and captions) using tens of thousands of chest X - rays paired with radiology report sentences. With DiffTumor [224], CT scans can be synthesized with a liver tumor of a specified size and location. Beyond text prompts, FMs can ge nerate i mages conditioned on other inputs, such as segmentation maps or existing images (image - to - image translation). Diffusion models are also used to i npaint or modify medical images in a controlled way [225]. Scalable Generation: Realistic high - resolution 3D/4D volume generation is rather challenging due to the high memory footprint required by a unified 3D framework. This is further complicated by the inhomogeneities of medical images in terms of volume dimens ions and pixel sizes. GenerateCT [226] addresses this challenge by decomposing the 3D generation process into a sequential generation of indi vidual sl ices. While a volum e can be generated to arbitrar y sizes with this approach, there are lingering concerns regarding the 3D structural inconsistencies across slices. O thers attem pt at direct 3D image generation: for example, MAISI achieves 5123 real istic CT image generation via latent space diffusion and Ten sor splitting parallelism (TSP) [227], [228]. D. Report Generati on and Visio n Question - Answer ing FMs have substant ially advanced automated r adiology report generation. For example, FMs can generate human - readable reports from multi - modal data in structur ed or unstructure d formats in professional and plain languages [229], [230]. Compared with earlier encoder – decoder systems, modern multimodal Transformer s pretrai ned on dataset s such as MIMIC - CXR can produce more coherent , stru ctured, and clinically aligned reports, often with explicit sections for findings, impressio ns, and comparisons [231]. Recent work further applies prefe rence - based optimization such as Direct Preference Optimizat ion (DPO) t o suppress ha llucinated prior examinations and better align generated text with r adiologist IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 11 T ABLE 1. M AJOR DATASETS AND PLATFORMS FOR MEDICAL IMAGING . Name / Plat form Modalit y / T ype Scope (Anatomy / T ask) Annotation / Data T ype Scale ChestX - ray14 [39] X- ray Chest disease s Image - level labels 100k+ CheXpert [40] X- ray Chest (in/ou tpatient) Uncertain labels 220k+ MIMIC - CXR [41] X- ray + reports Chest (ICU) Full radiology re ports; labels 370k+ DeepLesion [42] CT Whole - body lesions Bounding boxes 30k+ slices RadImageNet [43] CT/MRI/US Multi - or gan, multimodal Diagnostic labels 1.3M+ BraTS [44] MRI (3D) Brain tumors Segmentation masks ~2k cases EchoNet - Dynamic [45] Ultrasound (video) Cardiac EF values; masks 10k+ videos TCGA [46] WSI + CT /MRI/PET Multica ncer Dx labels; genomics; ROIs 20k – 30k WSIs + radiology PA N D A [47] Pathology (WSI) Prostate Gleason gr ading 11 k ROCO [48] /MedICaT [49] Multi - modal ity + tex t Scientific figur es Captions; ar ticle text 80k – 200k MedMNIST v2 [50] Multi - modal ity 18 organ/task datasets Class labels ~700k MSD [51] CT/MRI 10 organs Pixel masks 2.6k volumes fastMRI [52] MRI Knee, brai n Raw k - space; fully & undersampled Millio ns of s lices Calgary - Campinas [53] MRI Brain Fully sampled k - space ~370 volumes MRiLab s yntheti c MRI [54] MRI (si mulated) Brain Synthetic k - space Millio ns Mayo LDCT Chal lenge [55] CT Chest/abdomen Full - dose/low - dose paired CT Hundreds of volumes LIDC - IDRI [56] CT Lung Nodule lab els; full CT vo lumes 1,018 cases TCIA [57] Platform (multi - modal ity) Multica ncer , multi - organ Curated datas ets; Dx; segment ation; genomic s 50k+ studies MIDRC [58] Platform (X - ray , CT) COVID - 19, thoracic Standardized imaging + metadata 500k+ images UK Biobank Ima ging [59] Platform (MRI, X - ray , fundus) Population cohort Structural/functi onal MRI; clinical data 100k+ participants NLST [60] CT , Clinical Data Lung cancer scre ening Nodule anno tations; CT ser ies Te n s o f t h o u s a n d s OASIS [61] / HCP [62] MRI Neuroimagin g Structural + funct ional MRI Thousands AAPM Challenges [63] Multi - modal ity CXR, CT , colonoscopy , fractures Ta s k - specific annotations Va r i e s PhysioNet Imaging [64] Platform (X - ray , US, CT) ICU and clinical cohorts Images linked to EHR/waveform s 100k+ expectations [232]. Vision ques tion - answering (VQA) provides a novel interaction mode in which FMs answer targeted queries such as “Is there cardiomegaly?” or “What is the size of the liver lesion?” rather than producing a full report. Leveraging large - scale image – text pretraining and attention - based localization, contemporary medical VQA systems achieve strong performance across CXR, CT, MRI, ul trasound, and pathology, and can often operate in zero - shot or few - shot regime s [231]. Alignment methods like DPO have also be en adap ted to radiology VQA (RadQA - DPO) [233]. E. Other Task s Finally, we describe how FMs may solve some appli cations peripherally related to medical imaging. FMs can automate patient follow - up. Using imaging data and electronic patient records, FMs can personalize messages with recommendations and appoi ntments. FMs can identify public - health relevant disease patt erns or biomarkers. For example, FMs were suggested for detecting quantitative cancer biomarkers and predicting public disease progression [234]. FMs can also be used for workflow opt imization, such as image quality monitoring in a hospital, which can t hen alert technologists and/or physicians to i mage quality issues and/or incorrect protocol selection [235], [236], [237]. Other usages include automatic protocol selection and recommendation based on scout ima ges and patient information, and aut omatic data management (e.g., similar case retrieval, [238]). This list seems endless, only limited by our imagination. F. Medical Imaging Dat asets an d Benchmark s Foundation models in medical i maging rely on large, divers e, and multimodal datasets that span radiography, CT, MRI, ultrasound, nucl ear medicine, digital pat hology, and image – text corpora, as well as highly specialized datasets for tasks like reconstruction, segm entation, and report generation. Public platforms such as The Cancer Imaging Archive (TCIA) [239] and the Medical Imaging and Data Resource Center (MIDRC) [240] have become central hubs for standardized, curated collections acros s cancer imaging, C OVID - 19 imaging, lung screening, and multi - organ cohorts, enabling reproducible benchmarks and large - scale pretraining. In parallel, dedicated CT and MRI reconstru ction datasets — such as fastMRI [241], MRiLab dat a [242] , the May o Clini c Low - Dose CT Chal leng e datasets [243], and AAPM Grand Challenge collections [244] — provide high - quality raw k - space or projection data needed to support physics - informed foundation models. Table 1 summarizes representative dataset s and platforms that underpin the development and evaluation of generalist and multimodal foundati on models in medic al imagin g. IV. P ERSPECTIVES The successes of AI have been commonly attributed t o the three pillars: data, models, and computing power [265]. Large, diverse, and mult imodal data f uel the lear ning process; advanced architectures and optimization techniques extract information from data to empower generalizable models; and computing infrastructure enables the training and deployment of AI systems. Together, these three pill ars have driven much of the rapid evolut ion of foundation models in medical imaging. However, medicine is a mission - critical domain, and medical imaging serves as the ey es of modern medicine, w here errors IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 12 Fig . 4.1. F our pillars of foundation models for medical imaging. While three pillars for AI progress have been widely recognized, including data, mo dels, and computing, in medical imaging the stakes demand the fourth pillar: regulatory science. have life - altering consequences. Unlike other fields where innovation could temporarily outpace regulation, healthcare demands not only technical sophistication but also trust, safety, and accountability. This reality calls for the fourth pillar: regulator y sci ence. Robust evaluation frameworks, fairness auditing, clinical trials, and alignment with ethics ensure that foundation models for medical imaging meet the highest standards before entering real - world clinical workflows. By explicitly adding regulato ry science to the traditional triad as shown in Figure 4.1, we underline that AI in medical imaging demands more than just bigger datasets, smarter models, and faster GPUs – it requires a disciplined pathway from technical breakthroughs to clinical int egra tion. A. Data/Knowledge An unpre cedented scale of datas ets h as bee n used in pretraining large AI models i ncluding foundation models. However, it is increasingly recognized that bigger is not always better, particularly in the medical domain where image subtleties , clinical representative ness, and biological complexities direct ly affect model outcomes [65] . While the size of a dataset is a good indicator of information content, it has become the consensus that data quality, diversity, and multimodal ity are equal ly importan t for develo pment of trustworthy AI models and their clinical performance. Indeed, data quality ensures that foundation models learn meaningful patterns rather than noise or arti facts. Medical images vary widely in type, quality, style , and annotation accuracy. Low - quality or inconsistently labeled data can propagate errors through downstre am tasks. Also, data diversity underpins generalizability across pat ient populat ions, scanner s, and clinical settings . Many existing datasets overrepresent specific demographics or disease types. Finally, integrating imaging with clini cal text , genomics , a nd longitudinal health records, multimodal ity unlocks represent ations that reflec t the fu ll complexity of patient care. Despite their value , datasets remain largely fragmented across hospitals, vendors, and juri sdictions due to privacy regul ations (e.g., HIPAA, GDPR), intellectual property concerns, and institutional policies. Federated learning has already served as an alternative [66] . To further improve privacy, secure computation techniques can be combined with feder ated learning to ensure that intermediate model updates remain encrypted [67] . Moreover, synthetic data synthesis using generative AI can fill gaps where real data are scarce, sensitive, or inaccessible. When integrated with privacy - preserving pipelines, synt hetic data can be shared without compromising patient confidentiality [68] . Additionall y, we e nvision a le gally mand ated fra mework under which medical datasets would be preserved securely during patients’ lifetimes and be declassified a few decades later for research use. Such a mechanism, analogous to historical archives in other domains, would balance privacy and scientific value. This will allow future researchers to inherit a comprehensive, ethically sourced repository for studying disease evolution, h ealth trends, and long - ran ge biomedical questions. This means that data governance must not only protect individuals now but al so enable t ransformative discoveries for generations to come. Beyond sheer data volume, incorporating medical knowledge is emerging as a critical enabler for medical foundation models [69] . Knowledge graphs capturing biomedical ontologies, disease – symptom – treatment relationships, and imaging – genomics linkages can guide representation lear ning toward clinically meaningful concept s. Likewise, r etrieval - augmented generation (RAG) allows models to query curated clinical databases, imaging atlases, and knowledge bases duri ng Foundation Model Downstr eam Applica tion A Downstr eam Applica tion B Computing Power – GPUs and ecos yste ms that en able scalable , energy - efficient model training and deployment, with pathways toward ne xt - generation computing pl atfor ms. Data/Knowledge – Diverse datasets and kno wledge bases that in tegr ate multimo dal and multitask information, enabling secure data sharing thr ough fede rate d le ar nin g wi th in t he healthcare meta verse . Models/Optimization – Innovative architectures tr ained with self - learning and rein forc eme nt l ea rn ing , integrating multimodal data into a worl d model t o suppor t causal rea son ing , co nt in ual l ea rn ing , and inter pretabl e multitask ing. Regulatory Science – Robust evaluation frameworks and re gulato ry pathways that ensure saf e transla tion and rigorou s validation of foun dat io n mod el s for me di ca l imaging. 13 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 inference, providing verifiable yet up - to - date context for predictions [70] . B. Models/Opt imizati on Currently, autor egressive, and diff usion - type models represent two major paradigms. Large language models (LLMs) work via autoregression and are very successful. On the other hand , diffusion - type models have demonstrated exceptional perfor mance in image, video, and multimodal generation. These paradigms can be in contrast: tokenization versus transformation, next - token prediction versus field - based generation, symbolic reasoning versus perceptual changing , and semantic analysis versus manifold learning. Aut oregressive models ma y suff er fro m exposu re bia s / error accumulation, but diffusion - type models fa ce comput ational costs due to iterati ve sampling, though new solvers and latent - space parameterizations mitigate this issue. No w, a growing trend is to unify both paradigms [71], [72] . Up to tod ay t he Transformer architecture remains the mainstrea m, but breakth roughs are nee ded fo r per formance boost. Emerging ar chitectures such as Mamba models show the ability to capture long - range context with sub - quadratic complexity [73] . Further improvements like MambaExtend enhance its long - context capabilities via a training - free scaling calibration, enabling up to 32 times longer context windows with minimal computational overhead [74 ] . Also, differentiable reasoning engines and hybrid neuro - symbolic architecture are emerging to integrate symbolic knowledge with deep learning [75] . Efforts are further made toward brain - inspired architectures that work with novel links, loops, and emerging behaviors [76] . Inspiration from the human brain suggests future artificial neural network architectures with modularity, recurrent connectivity, memory, predictive coding, and cross - modal integrat ion, percepti on, and reasoning in fast and slow modes. While data - driven techniques offer immense value, they cannot fully replace the role of physics - based models. When t he underlying physics is well understood and models accur ately approximate r eality in a generalizabl e way, physics - based approaches will con sistently outperform purely data - driven ones. Therefor e, hybr id architectures that combine physics modeling with deep learning — such as physics - informed foundation models (FMs) — are likely to deliver the most robust and powerful solutions. When grounded in f irst principles, these FMs become e ven more foundational. Since a neural architecture is nothing but a computational prototype, we must train it to optimize its parameters and performance, including pre - training, training, and post - trainin g (during test - time/inferenc e) [77] . This is critical but high ly nontrivial, as it demands non - convex optimization. Techniques such as reinforcement learning with human or tool feedback and fine - turning in various forms can guide models toward desirable outcomes and meaningful decision - maki ng [78] . Dynamic optimiza tion strat egies, incl uding test - time training and adaptation, will be essential to m aintain performance under distribution shifts [79] . As medi cal imaging foundation models evolve, a natural tension lies between generalist models and specialist models . Generalist mode ls benefit fro m large - scale, multi - modal pretraining, offering transferable representations and a unified inference pipeline across diverse task s [80] . On the other hand, specialist models, often built through parameter - efficient fine - tuning of a generalist backbone, can achieve higher accuracy, regulatory clarity, and workflow integration for targeted use cases such as lung nodule tracking or cardiac f unction assessment [81], [82] . Future ecosystems will likely adopt a combined paradigm, where a robust gener alist foundation provides shared represent ations, while speci alist derivatives deliver precisi on, interpret ability, and regul atory compliance for mission - critical clinical endpoints. We believe that interactions between speci alist and generalist models exemplify bottom - up and top - down methods, defi ning the dynamics of medical AI . C. Computing Power The rapid advancemen t of medical imaging foundati on models depends cr iticall y on high - performance computing resources, with GPU being the main work horse. The pace of computational innovati on, f rom NVIDIA’s cutting - edge architectures to emerging specialize d AI accelerators, has enabled ever - larger models, shorter training cycles, and faster inferences, but it also demands investment to stay competitive. Initiatives such as New York’s EmpireAI exemplify forward - looking efforts to enable and democratize acces s to powerful computing resources, fostering partner ships across the state. As the first state - initiated AI - oriented computational infrastructure, Empire AI [83] brings together nine universities in New York to operate a shared high - performance computing resources. Designed to becom e a unique academic computing platform, Empire AI has demonstrated strong resul ts with its first - generation Alpha version. The upcoming Beta version represents a major boost, with 7X speedup and 20X acceleration in inference, capable of training multi - trillion - parameter models, Beyond the Beta version, the Buffalo supercomputing facility is expected in 2027 to deliver orders of magnitude greater computational power. Meanwhile, tech giants have active projects like Stargate, highlighting the global best - in - class resources. For m edical imaging, forging synergies between universities, hospitals, industry stakehol ders, and federate initiative s like EmpireAI will be key to ensuring t hat cutting - edge computational capacity translates into real - world clinical innovations. In addition to GPUs, emerging computing paradigms hold promises for development of medical i maging foundation models. Quantum computing has the po tential to revolution ize foundation models by accelerating large - scale training, enabling quantum - inspired architectures, and unlocking generative and/or discriminative capabilities for complex, high - dimensional probl ems. As quantum hardware matures, hybrid quantum - classical FMs could become ess ential , paving the way for breakthroughs in medical imaging and beyond . Neuromorphic computing, inspired by spiki ng neural networks and event - driven architectures, off ers ultra - low - power inference and real - time edge intelligence. Optical computing harnesses photonic interconnects and analog l ight - based operations to achi eve ma ssively parallel, energy - efficient matrix computations beyond the scaling limits of electronic chips. At another front ier, synthetic biological intelligence do es IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. X , NOVEMB ER 2020 14 Fig . 4.2. High - level roadmap for development of medical imaging foundation models underpinned by the four pillars, highlighting how technica l, re gulatory, and clinical advances converge to enable trustworthy, high - performance AI systems, while ensuring their s afe, ethical, and viable deployment for healthcare benefits. computation using engineered cellular or molecular systems, opening a radically new substrate for learning and decision - making under biochemic al constrain ts. As t hese technologi es mature, hybrid computi ng ecos ystems that integrat e ele ctronic, photonic, neuromorphic, and biological processors could deliver orders - of - magnitude improvements in speed, energy efficiency, and adaptability. D. Regulatory Sc ience The r apid development of medical imagi ng foundation models (FMs) has outpaced ex isting regulatory frameworks, underscoring the urgent need to develop new rules tailored to address their unique challenges. While the U.S. FDA has begun adapting its existing AI/ML - based soft ware regulati ons, s uch as predetermined change cont rol plans and Good Machine Learning Practice (GMLP), these ge neral frameworks do not fully account for the uniqu e features of medical found ation models. Unlike traditi onal task - specific AI systems, foundation models are pre - trained on vast, heterogeneous datasets and can be fine - tuned or prompted for a range of downstream tasks. This introduces new regulatory challenges. Given the mission - critical nature of medical applications, it is essent ial to develop a dedicated regulatory science strategy for foundation models to address generalizability, explainability , monitoring, and so on. To embed explainability in foundation models, a promising approach is to synergistically combine Chain - of - Thought (CoT) reasoning [84] and causal analysis [85] . CoT reasoning offers a narrative, step - by - step explanation of how a mod el arrives at a decision, helping users interpret its internal logi c. However, t hese re asoning traces c an be post hoc rationalizations rather than genuine causal mechanisms. On the other hand, causal analysis aims to uncover the underlying cause - and - effect relationships driving a model’s predictions . While more rigo rous, causal models alone may lack the intuitive transparency needed for clinicians to trust and adopt the system. We advo cate that integrating these two complementary approaches by ali gning the CoT explanat ion with the model’s learned or infe rred cau sal str ucture [86] . In principle, this coupling can produce a dual - layered explainability framework that is both human - interpretable and epistemically s ound. Such an approach not only enhances the trustworthiness of medical AI systems but also provides regulators with a pri ncipled method to evaluate explainability claims. In the context of foundati on models where tasks and data may shift signif icantly over time, t his approach offers a scalable and rigorous way to monitor model reasoning across clinical scenarios. A corners tone of regulator y oversi ght is ensuring generalizability under dist ribution s hifts. Prospective multi - institutional benchmarks should stress - test models across raw acquisition, reconstruction, enhancement, and diagnostic tasks to quantify variability introduced by hardware differences, imaging physics, and clinical protocols [87] [88] . Techniques such as domain adaptation, test - time training, and federated or cross - site training reduce sensitivity to site - or devi ce - specific artifacts. Subgroup performance disparit ies can undermine clinical trust, and can be measured with disaggregated met rics, dataset design through balanced cohort selecti on, and augmentation for under represented groups. Equally important is that AI models should produce well - calibrated confidence estimates [89] and abstain from pr edictions when uncertainty is high , providing an indicator for potential deterioration of m odel performance over time . Also, retrieval - augmented generation and tool grounding help ensure that diagnostic claims remain linked to verifiable evidence rather than hallucinated content [90] . Present (2025 - 2027) Near Future (until 2030) Long T erm (beyond 2030) Te c h n i c a l Innovation Regulatory Evolution Clinical Integration Healthcare Systems Closed - Loop F eedback Multimodal Data Tra n s f o r m e r s Fede rated L ea rnin g Genera tive AI Mamba / Mamba V ariants Te s t - Time Adaptation Neuro - Symbolic Reasoning Br ain - Inspired Architectures Neuromo rphic / Optical Compu ting Automated Mul ti - Modal Pipelines FDA PCCP s EU AI Act Regulatory Sandboxes Global AI Safety Standards & Guide lines Continuou s P ost - Mark et Surveilla nce Retrospective Va l i d a t i o n Single - site Pilots Prospe ct ive Multi - Center T rials Explainability (Co T + Causal) Real - Time Auditing, Adapting, & Optimization Hybrid & Other Architectures Digital T wins Healthcare Metaverses Evolvin g Eco - syste m o f AI Mo del s Quantum C omputing ? 15 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 Regulatory science treats model deployment as the beginning of oversight. Clinical int egration requires continuous monitoring for d ata drift, adversari al inputs, and performance regressions, coup led with shadow mode evaluations before live deployment. The FDA’s Predetermined Change Control Plans (PCCPs), the EU AI Act, the UK’s MHRA SaMD framework, and the NIST AI Risk Management Framework converge on principles for pre - authorized updates, locked evaluation datasets, r ollback procedures, and post - market sur veillance. In this mon itoring proce ss, transparent governance underpins all regulatory stages. Model cards, data sheets, and stratified performance specifications by sit e, modality, and patient subgroup create an auditable evidence base. Documenting traini ng datasets and steps, algorithmic changes, calibration methods, a nd real - wor ld perfor mance metrics supports reproducibility and facilitates global regulatory harm onization. E. Concluding Remar ks The convergence of data/knowledge, models/optimization, computing power, and regulatory science is redefining medical imaging foundation models. Further success demands high - quality, multimodal data, innovative ar chitectures, sust ainable computing ecosyste ms, and contemporary regulatory frameworks. Future foundation models will likely combine generalist representations across modalities and tasks with specialist derivatives fine - tuned for h igh - stakes clinical endpoints, supported by advances in feder ated learning, privacy - preserving syntheti c augmentation, and retrieval - augmented reasoning to break down dat a sil os. At the same time, brain - inspired architectures, physics - informed generative models, and emerging computing paradigms promise major gains in effic iency and capability. Yet, clinical translation will ultimately hinge on rigorous and transparent governance and continuous post - deployment oversight. By uniting technical breakthroughs with ethical and regulatory rigor, medical imaging foundation models c an evolve into future healt hcare systems , be ing powerful, trustworthy, and impactful. R EFERENCES [1] M. Awai s et al . , “Founda tional Models Defining a New Era in Vision: A Survey and Outl ook,” 2023, arXiv . doi: 10.48550/ARXIV.2307.13721. [2] A. M. Turing, “Computin g Machinery an d Intellige nce,” in Parsing the Turing Test , R. Epstein, G. Roberts, and G. Bebe r, Ed s., Dordrecht: Spr inger Netherlands, 2009, pp. 23 – 65. doi: 10.1007/978 -1- 4020 - 6710 - 5_3. [3] M. Campbell, A. J. Hoane J r, and F. Hsu , “Deep blue,” Artif. Intel l. , vol. 134, no. 1 – 2, pp. 57 – 83, 2002. [4] E. Shortliffe, Computer - based medical consultations: MYCIN , vol. 2. Elsevier, 2012. Accessed: Oct. 17, 2025. [Online]. Available: https://books.google.com/books?hl=zh - CN&lr=&id=i9QXugPQw6oC&oi=f nd&pg=PP1&dq= Computer - based+medical+consultations:+MYCIN%5BM%5D.+El sevier,+2012.&ots=Ib7h - 09wQG&sig=KVtV3cRHEDRj9xzLLY3k PDZeu5Y [5] T. Winograd, “Procedur es as a r epresentation for dat a in a computer program for understanding natural language,” 1971. Accessed: Oct. 17, 2025. [Onli ne]. Available: https://apps.dtic.mil/s ti/html/tr/AD0721399/ [6] D. A. Ferrucci, “In troduction to ‘this is watson,’ ” IBM J. Res. Dev. , vol. 56, no. 3.4, pp. 1 – 1, 2012. [7] S. M . Stigler, “Gauss and the invention of least squares,” Ann. Stat. , pp. 465 – 474, 1981. [8] C. Cortes and V. Vapnik, “Support - vector networks,” Mach. Lear n. , vol. 20, no. 3, pp. 273 – 297, Sept. 1995, doi: 10.1007/BF00994018. [9] L. Breiman, “Random Forest s,” Mach. Learn. , vol. 45, no. 1, pp. 5 – 32, Oct. 2001, doi: 10.1023/A:1010933404324. [10] Y. Le Cun, L. Bot tou, Y. Bengio, and P. Ha ffner, “Gradient - based learning applied to document recognition,” Proc. I EEE , vol. 86, no. 11, pp. 2278 – 2324, 2002. [11] A. Krizhevsky, I. Sutskever, and G. E. Hi nton, “Imagenet classification with deep convolutional neur al networks,” Adv. Ne ural Inf. Process. Syst. , vol. 25, 2012, Accessed: Oct. 17, 2025. [Online]. Avail able: https://proceedings.neurips.cc/ paper/2012/hash/c399862 d3b9d6b76c8436e924a68c45b - Abstract.ht ml [12] K. He, X. Zhang, S. Ren, and J. Sun, “Deep residual learning for image recognition,” in Proceedings of the IEEE conference on computer vision and pattern recognition , 2016, pp. 770 – 778. Accessed: Oct. 17, 2025. [Online]. Available: http://openaccess.thecvf.com/content_cvpr_2016/ html/ He_Deep_Residua l_Learning_ CVPR_2016_paper. html [13] S. Hochreite r and J. Schmidhuber, “Long short - term memory,” Neural Comput. , vol. 9, no. 8, pp. 1735 – 1780, 1997. [14] I. Goodfellow et al. , “Generative adversarial network s,” Commun. ACM , vol. 63, no. 11, pp. 1 39 – 144, Oct. 2020, doi: 10.1145/3422622. [15] O. Ronn eberger, P. Fischer, and T. Brox, “U - Net: Convolutional Netwo rks for Biomedical Image Segmentation,” May 18, 2015, arXiv : doi: 10.48550/arXiv.1505.04597. [16] J. Jumper et al. , “Highly accurate protein structure prediction with AlphaFold, ” nature , vol. 596, no. 7873, pp. 583 – 589, 2021. [17] K. Singhal et al. , “Large language models encode clinical knowledge,” Nature , vol. 620, no. 7972, pp. 172 – 180, 2023. [18] R. Bommasani, “On the oppor tunities and ri sks of foundation models,” ArXiv Pr epr. ArXiv210807 258 , 2021. [19] “Introducing the Center for Research on Foundation Models (CRFM) | Stanford HAI.” Accessed: Oct. 17, 2025. [Online]. Available: https://hai.stanford.edu/news/ introducing - center - research - foundation - models - crfm [20] S. Alfasly et al. , “When is a Foundation Model a Foundation Model,” Sept. 14, 2023, arXiv : arXiv:2309.11510. doi: 10.48550/arXiv.2309.11510. 16 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 [21] X. Liu et al. , “Self - supervised Learning: Generative or Contrastive,” IEEE Trans. Knowl. Data Eng. , pp. 1 – 1, 2021, doi: 10.1109/TKDE.2021.3090866. [22] L. Yang et al. , “Diffusion Mod els: A Comprehensive Survey of Methods and Applications,” Sept. 27, 2025, arXiv : arXiv:2209.00796. doi: 10.48550/arXiv.2209.00796. [23] A. Kazerouni et al. , “Diffusion Mo dels for Medical Image Analysis: A Comprehensiv e Survey,” June 03, 2023, arXiv : arXiv:2211.07804 . doi: 10.48550/arXiv.2211.07804. [24] D. Hein, A. Bozorgpour, D. Merhof, and G. Wang, “Physics - Inspired Generative Models in Medical Imaging: A Review,” Aug. 23, 2024, arXiv : arXiv:2407.10856. doi: 10.48550/arXiv.2407.10856. [25] S. Longpre et al. , “The Responsible Foundation Model Development Che atsheet: A Re view of Tool s & Resources,” Feb. 17, 2025, arXiv : doi: 10.48550/arXiv.2406.16746. [26] W. Zhou et al. , “The security of using large language models: A survey with emphasis on ChatGPT,” IEEECAA J. Autom. Sin. , 2024, Accesse d: O ct. 17, 2025. [Online]. Available: https://ieeexplore.ieee.org/ abstract/document/10751746/ [27] W. X . Zhao et al. , “A survey of large language models,” ArXiv Prepr. ArXiv230318223 , vol. 1, no. 2, 2023, Accessed: Oct . 17, 202 5. [Online ]. Availabl e: https://www.researchgate.net/profile/ Tang - Tianyi - 3/publication/369740832_A_Survey_of_Large_Langua ge_Models/links/665fd2e3637e4448a37dd281/A - Survey - of - Large - Language - Models .pdf [28] I. A. Scott and G. Zuccon, “The new paradigm in machine learn ing – foundation models, large language models and beyond: a primer for physicians,” Intern. Med. J. , vol. 54, no. 5, pp. 705 – 715, May 2024, doi: 10.1111/imj.16393. [29] J. Yang et al. , “Harnessing the Power of LLM s in Practice: A Survey on ChatGPT and Beyond,” ACM Trans. Knowl. Discov. Data , vol. 18, no. 6, pp. 1 – 32, July 2024, doi: 10.1145/3649506. [30] D. Tru hn, J. - N. Eckardt, D. Ferber, and J. N. Kather, “Large language models and multimodal foundati on models for precisi on oncology,” NPJ Precis. Oncol. , vol. 8, no. 1, p. 72, 2024. [31] B. Azad et al. , “Foundational Models in Medical Imaging: A Comprehensive Survey and Future Vision,” Oct. 28, 20 23, arXiv : arXiv:2310.18689. doi: 10.48550/arXiv.2310.18689. [32] S. - C. Huang, M. Jensen , S. Yeung - Levy, M. P. Lungren, H. P oon, and A. S. Chaudhari, “Multimodal Foundation Models for Med ical Imaging - A Sys tematic Review and Implementation Guidelines,” medRx iv , pp. 2024 – 10, 2024. [33] J. S. Ryu, H. Kang, Y . Chu, and S. Yang, “V ision - language foundation models for medical imaging: a review of current practices and innovations,” Biomed. Eng. Lett. , vol. 15, no. 5, pp. 809 – 830, Sept. 2025, doi: 10.1007/s13534 - 025 - 00484 - 6. [34] K. Sun et al. , “Medical Multim odal Foundation Models in Clinical Diagnosis and Treatment: Applications, Challenges, and Future Directions,” Dec. 03, 2024, arXiv : arXiv:2412.02621. doi: 10.48550/arXiv.2412.02621. [35] S. Zhang and D. Metaxas, “On the Challenges and Perspectives of Foundation Models for Medical Image Analysis,” Nov. 21, 2023, arXiv : arXiv:23 06.05705. doi: 10.48550/arXiv.2306.05705. [36] J. P. Huix, A. R. Ganeshan, J. F. Haslum, M. Söderberg, C. Matsoukas, and K. Smith, “Are Natural Domain Foundation Models Useful f or Medical Image Classificat ion?,” in 2024 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV) , Waikoloa, HI, USA: IEEE, J an. 2024, pp. 7619 – 7628. doi: 10.1109/WACV57701.2024.00746. [37] V. van Vel dhuizen et al. , “F oundation Models in Medical Imaging -- A Review and Outlook, ” June 16, 2025, arXiv : arXiv:2506.09095. doi: 10.48550/arXiv.2506.09095. [38] M. O. Khan, M. M. Afz al, S. Mirz a, and Y. Fang, “How Fair are Medical Imaging Foundation Models ?,” in Proceedings of the 3rd Machine Learning for Health Symposium , PMLR, Dec. 2023, pp. 217 – 231. Accessed: Oct. 17, 2025. [Online]. Avail able: https://proceedings.mlr.press/ v225/khan23a.html [39] A. Vaswani et al. , “Attention is All you N eed”. [40] C. Niu and G. Wang, “Unsupervised contrast ive learning based transformer for lung nodule detection,” Phys. Med. Biol. , vol. 67, no. 20, p. 204001, Oct. 2022, doi: 10.1088/1361 - 6560/ac92ba. [41] A. Dosovitskiy et al. , “An Image is Worth 16x16 Words: Transformers for Image Recogni tion at Scal e,” June 03, 2021, arXiv : arXiv:2010.11929 . doi: 10.48550/arXiv.2010.11929. [42] Z. Liu et al. , “Swin Transformer: Hierarchical Vision Transformer using Shi fted Windows,” Aug. 17, 2021, arXiv : arXiv:2103.14030. doi: 10.48550/arXiv.2103.14030. [43] Y. Li, H. Mao, R. Girs hick, and K. He, “Explori ng Plai n Vision Transfor mer Backbones for Object Detect ion,” in Computer Vision – ECCV 2022 , vol. 13669, S. Avidan, G. Brostow, M. Cissé, G. M. Fari nella, and T. Hassner , Eds., in Lect ure Notes in Computer Science, vol. 13669. , Cham: Springer Nature Switzerlan d, 2022, pp. 280 – 296. doi: 10.1007/978 -3- 031 - 20077 - 9_17. [44] A. Radf ord, K. Narasimhan, T. Salimans, and I. Sutskever, “Improving Language Understanding by Generative Pre - Training”. [45] S. Bai et al. , “Qwen2.5 - VL Technical Report,” Feb. 19, 2025, arXiv : arXiv:2502.13923 . doi: 10.48550/arXiv.2502.13923. [46] L. Blankemeier et al. , “Merlin: A Vision Language Foundation Model for 3D Computed Tomography,” June 10, 2024, arXiv : arXiv:2406.06512. doi: 10.48550/arXiv.2406.06512. [47] N. Shazeer et al. , “Outrageously Large Neural Networks: The Spar sely - Gated Mixture - of - Expert s Layer ,” J an. 23 , 17 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 2017, arXiv : arXiv:1701.06538 . doi: 10.48550/arXiv.1701.06538. [48] D. Dai et al. , “DeepSeekMoE : Towards Ultimate Expert Specialization in Mixture - of - Experts Language Models,” Jan. 11, 2024, arXiv : arXiv:2401.06066 . doi: 10.48550/arXiv.2401.06066. [49] R. Child, S. Gr ay, A. Ra dford, and I. Sutskever, “Generating Long Sequences with Sparse Transformers,” Apr. 23, 2019, arXiv : arXiv:1904.10 509. doi: 10.48550/arXiv.1904.10509. [50] K. Cho romanski et al. , “Rethinking Attention with Performers,” Nov. 19, 2022, arXiv : doi: 10.48550/arXiv.2009.14794. [51] A. Kathar opoulos, A. Vyas, N. Papp as, and F. Fleu ret, “Transformers are RNNs: Fast Autoregressive Transformers with Lin ear Attention,” Aug. 31, 2020, arXiv : arXiv:2006.16236. doi: 10.48550/arXiv.2006.16236. [52] S. Wang, B. Z. Li, M. Khabsa, H. Fang, and H. Ma, “Linformer: Self - Attenti on wit h Linea r Complex ity,” June 14, 2020, arXiv : arX iv:2006.04768. doi: 10.48550/arXiv.2006.04768. [53] J. Ainslie, J. Lee - Thorp, M. de Jong, Y. Zemlyanski y, F. Lebrón, and S. Sanghai, “ GQA: Training Gener alized Multi - Query Transformer Models from Mult i - Head Checkpoints,” De c. 23, 2023, arXiv : arXiv:2305 .13245. doi: 10.48550/arXiv.2305.13245. [54] N. Shazeer, “Fast Transformer Decoding: One Write - Head i s Al l You Need,” Nov. 06, 2019, arXiv : arXiv:1911.02150. doi: 10.48550/arXiv.1911.02150. [55] A. Krizhevsk y, I. Suts kever, and G. E. Hinto n, “ImageNet classif ication with deep convoluti onal neur al networks,” Commun. ACM , vol. 60, no. 6, pp. 84 – 90, May 2017, doi: 10.1145 /306538 6. [56] K. He, X. Zhang, S. Ren, and J. Sun, “Deep Residual Learning for Image Recognition,” Dec. 10, 2015, arXiv : arXiv:1512.03385. doi: 10.48550/arXiv.1512.03385. [57] S. Woo, J. Pa rk, J. - Y. Lee, and I. S. Kweon, “CBAM: Convolutional Block Attention Module,” in Computer Vision – ECCV 2018 , vol. 11211, V. Ferrari, M. Hebert, C. Sminchisescu, and Y. Weis s, Eds., in Lecture Notes i n Computer Science, vol. 11211. , Cham: Springer International Publishing, 2018 , p p. 3 – 19. doi: 10.1007/978 -3- 030 - 01234 - 2_1. [58] I. Bello , B. Zoph, Q. Le, A. Vaswani, and J. Shlen s, “Attention Augmented Convolutional Networks,” in 2019 IEEE/CVF International Conference on Computer Vision (ICCV) , Seoul, Korea (South): IEEE , Oct. 2019, pp. 3285 – 3294. doi: 10.1109/ICCV.2019.00338. [59] H. Wu et al. , “CvT: Introducing Convolutions to Vision Transformers,” in 2021 IEEE/CVF I nternational Conference on Comp uter Vision ( ICCV) , Montreal, QC, Canada: IEEE, Oct . 2021, pp. 22 – 31. doi : 10.1109/ICCV48922.2021.00009. [60] A. S herstinsky , “ Fundamentals of Recurr ent Neural Network (RNN) and Long Short - Term Memory (LSTM) Network,” Phys. Nonlinear Phenom. , vol. 404, p. 132306, Mar. 2020, doi: 10.1016/j.phys d.2019.132306. [61] J. Chung, C. Gulcehre, K. Cho, and Y. Bengio, “Empirical Evaluation of Gated Recurrent Neural Networks on Sequence M odeling,” Dec. 11, 2014, arXiv : arXiv:1412.3555. doi: 10.48550/arXiv.1412.3555. [62] A. Gu, K. Goel, and C. Ré, “Efficie ntly Modeling Long Sequences with Structured State Spaces,” Aug. 05, 2022, arXiv : arXiv:2111.00396. doi: 10.48550/arXiv.2111.00396. [63] A. Gu and T. Dao, “Mamba: Linear - Time Sequence Modeling with Select ive St ate S paces,” May 3 1, 202 4, arXiv : arXiv:2312.00752. doi: 10.48550/arXiv.2312.00752. [64] B. Peng et al. , “RWKV: Reinventing RNNs for the Transformer Era,” Dec. 11, 2023, arXiv : arXiv:2305.13048. doi: 10.48550/arXiv.2305.13048. [65] D. Y. Fu, T. Dao, K. K. Sa ab, A. W. Thomas, A. Ru dra, and C. Ré, “Hungry Hungry Hippos: Towards Language Modeling with State Space Model s,” Apr. 29 , 2023, arXiv : arXiv:2212.14052. doi: 10.48550/arXiv.2212.14052. [66] Y. Sun et al. , “Retentive Network: A Successor to Transformer for Large Language Models, ” Aug. 09 , 2023, arXiv : arXiv:2307.08621 . doi: 10.48550/arXiv.2307.08621. [67] Y. Yue a nd Z. Li, “MedMamba: Vision Mamba for Medical Image Classificati on,” Sept. 29, 2024, arXiv : arXiv:2403.03849. doi: 10.48550/arXiv.2403.03849. [68] Z. Chen et al. , “LangMamba: A Language - driven Mamba Framework for Low - dose CT Denoising with Vision - language Models,” July 08, 2025, arXiv : arXiv:2507.06140. doi: 10.48550/arXiv.2507.06140. [69] D. P. Kingma and M. Welling, “Auto - Encoding Variational Bayes,” Dec. 10, 2022, arXiv : arXiv:1312.6114. doi: 10.48550/arXiv.1312.6114. [70] I. Higgins et al. , “β - VAE: LEARNING BASIC VISUAL CONCEPTS WITH A CONSTRAINED VARIATIONAL FRAMEWORK,” 20 17. [71] A. van den Oord, O. Vinyals , and K. Kavukcuoglu , “Neural Discrete Representation Learning,” May 30, 2018, arXiv : arXiv:1711.00937 . doi: 10.48550/arXiv.1711.00937. [72] Z. Zheng et al. , “Open - Sora: Democratizing Efficient Video Produ ction for All,” Dec. 29, 202 4, arXiv : arXiv:2412.20404. doi: 10.48550/arXiv.2412.20404. [73] I. J. Goodfellow et al. , “Generative Adversarial Networks,” June 1 0, 2014, arXiv : 10.48550/arXiv.1406.2661. [74] A. Radford, L. Metz, and S. Chintala, “Unsupervised Representatio n Learning with Deep Convolutional Generative Adve rsarial Networ ks,” Jan. 07, 2016, arXiv : arXiv:1511.06434. doi: 10.48550/arXiv.1511.06434. [75] T. Karras , T. Aila, S. Laine, and J. Lehtinen, “Progressive Growing of GANs for Improved Quality, Stability, and Variation,” Feb. 26, 2018, arXiv : arX iv:1710.10196. doi: 10.48550/arXiv.1710.10196. [76] T. Karras, S. Laine, and T. Aila, “A St yle - Based Generator Archit ecture for Generati ve Adversarial Networks,” Ma r. 29, 20 19, arXiv : doi: 10.48550/arXiv.1812.04948. 18 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 [77] A. Brock, J . Donahue, and K. Simonyan , “Large Sc ale GAN Training for High Fidelity Natural Imag e Synthesis,” Feb. 25, 2019, arXiv : arXiv:1809.11096. doi: 10.48550/arXiv.1809.11096. [78] S. Dayarathna, K. T. Islam, S. Uribe, G. Yang, M. Hayat, and Z. C hen, “Deep learning based synthesis of M RI, C T and PET: Review and anal ysis,” Med . Image An al. , vol. 92, p. 103046, 2024, doi: https://doi.org/10.1016/j. media.2023.103046. [79] C. You et al. , “CT Super - resolution GAN Constrained by the Identical, Residual, and Cycle Lea rning Ensemble(GAN - CIRCLE),” IEEE Trans. Med. Imaging , vol. 39, no. 1, pp. 188 – 203, Jan. 2020, doi: 10.1109/TMI.2019.2922960. [80] Q. Yang et al. , “Low Dose CT Im age Denoising Using a Generative Adversarial Net work with Wasserstein Distance a nd Percept ual Loss, ” IEEE Trans. Med. Imaging , vol. 37, no. 6, pp. 1348 – 1357, June 2018, doi: 10.1109/TMI.2018.2827462. [81] C. Niu et al. , “Low - Dimensional Manifold - Constrain ed Disentangleme nt Network for Met al Artifact Reduction,” IEEE Trans. Radiat. Plasma Med. Sci. , vol. 6, no. 6, pp. 656 – 666, July 2022, doi: 10.1109/TRPMS.2021.3122071. [82] C. Ni u et al. , “Noise Entangled GAN For Low - Dose CT Simulation,” Feb. 18, 2021, arXiv : arXiv:2102.09615 . doi: 10.48550/arXiv.2102.09615. [83] J. Ho, A. Jain, and P. Abbeel, “Denoising Diffusion Probabilistic Models,” Dec. 16, 2020, arXiv : arXiv:2006.11239. doi: 10.48550/arXiv.2006.11239. [84] L. Yang et al. , “Diffusion Mod els: A Comprehensive Survey of Methods and Applications,” Dec. 02, 2024, arXiv : arXiv:2209.00796. doi: 10.48550/arXiv.2209.00796. [85] Y. So ng, P. Dhariwa l, M. Chen, and I. Suts kever, “Consistency Models,” May 31, 2023, arXiv : arXiv:2303.01469. doi: 10.48550/arXiv.2303.01469. [86] D. Hein, A. Bozorgpour, D. Merhof, and G. Wang, “Physics - Inspired Generative Models in Medical Imaging: A Review,” Aug. 23, 2024, arXiv : arXiv:2407.10856. doi: 10.48550/arXiv.2407.10856. [87] T. B. Brown et al. , “La nguage Models are Few - Shot Learners,” July 22, 2020, arXiv : arXiv:2005.14165. doi: 10.48550/arXiv.2005.14165. [88] A. van den Oord, N. Kalchbrenner, and K. Kavukcuoglu, “Pixel Recurrent Neural Networks,” Aug. 19, 2016, arXiv : arXiv:1601.06759. doi: 10.48550/arXiv.1601.06759. [89] A. va n den Oord, N. Kalchbr enner, O. Vi nyals, L. Espeholt, A. Graves, a nd K. Kavukcuoglu, “Conditional Image Generation with PixelCNN Decoders,” June 18, 2016, arXiv : arXiv:1606.05328 . doi: 10.48550/arXiv.1606.05328. [90] A. Ramesh et al. , “Zero - Shot Text - to - Image Generation, ” Feb. 26, 2021, arXiv : doi: 10.48550/arXiv.2102.12092. [91] P. Sun et al. , “Autoregressive Model Beats Diffusion: Llama for Scalable Image Generation,” June 1 0, 2024, arXiv : arXiv:2406.06525. doi: 10.48550/arXiv.2406.06525. [92] X. Wang et al. , “Emu3: Next - Token Prediction is All You Need ,” Sep t. 2 7, 20 24, arXiv : arXiv:24 09.18869. doi: 10.48550/arXiv.2409.18869. [93] A. van den Oord, Y. Li, and O. Vinya ls, “Represe ntation Learning with Cont rastive Predic tive Coding,” Jan. 22, 2019, arXiv : arXiv:1807.03748 . doi: 10.48550/arXiv.1807.03748. [94] T. Chen, S. Kornblith, M. Norouzi, and G. Hinton, “A Simple Framework for Contrastive Lear ning of Visual Representatio ns,” July 01, 2020, arXiv : arXiv:2002.05709. doi: 10.48550/arXiv.2002.05709. [95] K. He, H. Fan, Y. Wu, S. Xie, an d R. Girsh ick, “Momentum Contrast for Unsupervised Visual Representatio n Lear ning,” Mar. 23, 2020, arXiv : arXiv:1911.05722. doi: 10.48550/arXiv.1911.05722. [96] I. Misra and L. van der Maaten, “Self - Supervised Learning of Pretext - Invariant Representations,” Dec. 04, 2019, arXiv : arXiv:1912.01991 . doi: 10.48550/arXiv.1912.01991. [97] J. - B. Grill et al. , “Bootstrap your own laten t: A new approach to self - supervised Learning,” Sept. 10, 2020, arXiv : arXiv:2006.07733. doi: 10.48550/arXiv.2006.07733. [98] X. Chen and K. He, “Exploring Simpl e Siame se Representatio n Learning,” Nov. 20, 2020, arXi v : arXiv:2011.10566. doi: 10.48550/arXiv.2011.10566. [99] M. Caron et al. , “Emer ging Properties in S elf - Supervised Vision Transformers,” May 24, 2021, arXiv : arXiv:2104.14294. doi: 10.48550/arXiv.2104.14294. [100] M. Ca ron, I. Misra, J. Mair al, P. Goyal, P. Boja nowski, and A. Jouli n, “Unsupervised Learning of Visual Features by Contrast ing Cluster Assignments, ” Jan. 08, 2021, arXiv : arXiv:2006.09882 . doi: 10.48550/arXiv.2006.09882. [101] M. Car on, P. Bo janowski , A. Joul in, and M. Douze, “Deep Clustering for Unsupervised Learning of Visual Features,” Mar. 18, 2019, arXiv 10.48550/arXiv.1807.05520. [102] Y. M. Asano, C. Rupp recht, a nd A. Ved aldi, “Se lf - labelling via simultaneous clustering and representation learning,” Feb. 19, 2020, arXiv : arXiv:1911.0 5371. doi: 10.48550/arXiv.1911.05371. [103] A. Bard es, J. Pon ce, and Y. LeCun, “VICReg: Variance - Invariance - Covariance Regularization for Self - Supervised Learning,” Jan. 28, 2022, arXiv : arXiv:2105.04906. doi: 10.48550/arXiv.2105.04906. [104] C. Niu, W. Xia, H. Shan, and G. Wang, “ Information - Maximized Soft Variable Discret izati on for Self - Supervised Image Representation Learning,” Jan. 07, 2025, arXiv : arXiv:2501.03469 . doi: 10.48550/arXiv.2501.03469. [105] C. Niu et al. , “Noise suppression with similarity - based self - supervised deep learning,” IEEE Trans. Med . Imaging , vol. 42, no. 6, pp . 1590 – 1602, 2022. [106] S. - C. Huang, A. Paree k, M. Jensen, M. P. Lungren , S. Yeung, and A. S. Chaudhari, “Self - supervised learning for medical image classification: a system atic review and 19 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 implementation guidelines,” Npj Digit . Me d. , vol. 6, no. 1, p. 74, Apr. 2023, doi: 10.1038/s41746 - 023 - 00811 - 0. [107] A. Radf ord et al. , “Learning Transferab le Visual Models From Natural Language Supervision,” Feb. 26, 2021, arXiv : arXiv:2103.00020. doi: 10.48550/arXiv.2103.00020. [108] C. Jia et al. , “Scaling Up Visual and Vision - Language Representatio n Learning Wit h Noisy Text Supervision,” June 11, 2021, arXiv : arXiv:2102.05918. doi: 10.48550/arXiv.2102.05918. [109] Z. Zhao et al. , “CLIP in medical imaging: A survey,” Med. Ima ge Anal . , vol. 102, p. 103551, May 2025, doi: 10.1016/j.media.2025.103551. [110] J. Devlin, M. - W. Chang, K. Lee , and K. Toutano va, “BERT: Pre - training of Dee p Bidirectional Transformers for Language Understanding,” May 24, 2019, arXiv : arXiv:1810.04805. doi: 10.48550/arXiv.1810.04805. [111] K. He, X. Chen, S. Xie, Y. Li, P. Dollár, and R. Girshick, “Masked Autoencoders Are Scalable Vision Learners ,” Dec. 19, 2021, arXiv : arXiv:2111.06377. doi: 10.48550/arXiv.2111.06377. [112] Y. LeCun, “A Path Towards Autonomous Machine Intelligence Version 0.9.2, 2022 - 06 - 27”. [113] M. Assran et al. , “Self - Supervised Learning from Images with a Joi nt - Embedding Predictive Architectur e,” Apr. 13, 2023, arXiv : arXiv:2301.08243. doi: 10.48550/arXiv.2301.08243. [114] A. Baevs ki, W. - N. Hsu, Q. Xu , A. Babu, J. Gu, and M. Auli, “data2 vec: A General Fr amework for Sel f - supervised Learning in Speech, Vision and Language,” Oct. 25, 20 22, arXiv : arXiv:2202.03555. doi: 10.48550/arXiv.2202.03555. [115] D. C. Knill and A. Pouget, “The Bayesi an brain: the r ole of uncertainty in neural coding and computation,” TRENDS Neurosci. , vol. 27, no. 12, pp. 712 – 719, 2004. [116] K. Frist on, “The free - energy principle: a unified brain theory?,” Nat. Rev. Neurosci . , vol. 11, no. 2, pp. 127 – 138, 2010. [117] L. Ouyang et al. , “Training language models to follow instructions with human feedback,” Mar. 04, 2022, arXiv : arXiv:2203.02155. doi: 10.48550/arXiv.2203.02155. [118] P. Christiano, J. Leike, T. B. Brown, M. Martic, S. Legg, and D. Amodei, “Deep reinforcement learning from human preferences,” Feb. 17, 2023, arXiv : arXiv:1706.03741. doi: 10.48550/arXiv.1706.03741. [119] J. Schulman, F. Wolski, P. Dhariwal, A. Radford, and O. Klimov, “Proximal Policy Opt imization Algorithms,” Aug. 28, 2017, arXiv : arXiv:1707.06347. doi: 10.48550/arXiv.1707.06347. [120] N. Stiennon et al. , “Learning to summarize from human feedback,” Feb. 15, 2022, arXiv : arXiv:2009.01325. do i: 10.48550/arXiv.2009.01325. [121] R. Rafailov, A. Sharma, E. Mitchel l, S. Er mon, C. D. Manning, a nd C. Finn, “ Direct Pr eferenc e Optimiza tion: Your Language Model is Secretly a Reward Model,” July 29, 2024, arXiv : arXiv:2305.18290. doi: 10.48550/arXiv.2305.18290. [122] J. Hong, N. Le e, and J. Thorne, “O RPO: M onolithic Preference Optimization without Reference Model,” Mar. 14 , 202 4, arXiv : arXiv:2403.07691. doi: 10.48550/arXiv.2403.07691. [123] O. Banerjee, H. - Y. Zhou, S. Adi than, S . Kwak, K. Wu, and P. Rajpurkar, “Direct Preference Optimization for Suppressing Hallucinated Prior Exams in Radiology Report Generation, ” June 14, 2024, arXiv : arXiv:2406.06496. doi: 10.48550/arXiv.2406.06496. [124] S. A. Nahian and R. Kavul uru, “RadQA - DPO: A Radiology Qu estion Ans wering Sys tem with Encoder - Decoder Model s Enhanced by Direct Preferen ce Optimization” . [125] Z. Shao et al. , “DeepSeekMath: Pushing the Limits of Mathemati cal Reasonin g in Open Language Models,” Apr. 27, 2024, arXiv : arXiv:2402.03300. doi: 10.48550/arXiv.2402.03300. [126] Q. Yu et al. , “DAPO: An Open - Source LLM Reinforcement Learning System at Scale,” May 20, 2025, arXiv : arXiv:2503.14476 . doi: 10.48550/arXiv.2503.14476. [127] C. Zheng et al. , “Group Sequence Policy Optimization,” July 28, 2025, arXiv : arXiv:2507.18071. doi: 10.48550/arXiv.2507.18071. [128] H. Lightman et al. , “L et’s Verify Step by Step,” M ay 31, 2023, arXiv : arXiv:2305.20050 . doi: 10.48550/arXiv.2305.20050. [129] S. Yao et al. , “Tree of Thoughts: Deliberate Problem Solving with Large Language Models,” Dec. 03, 2023, arXiv : arXiv:2305.10601. doi: 10.48550/arXiv.2305.10601. [130] Z. Chen, Y. Deng, H. Yuan, K. Ji, and Q. Gu, “S elf - Play Fine - Tuning Converts Weak Language Models to Strong Language Model s,” June 14, 2024, arXiv : arXiv:2401.01335. doi: 10.48550/arXiv.2401.01335. [131] H. Wang et al. , “Offline Reinf orcement Learning for LLM Multi - Step Reasoni ng,” Dec. 25, 2024, arXiv : arXiv:2412.16145. doi: 10.48550/arXiv.2412.16145. [132] Y. Lai et al. , “Med - R1: Reinf orcement Learning for Generalizabl e Medical Reasoning in Vision - Language Models,” 2025, arXiv . doi: 10.48550/ARXIV.2503.13939. [133] C. Niu and G. Wang, “Reasoning Language Model for Personalized Lung Cancer Screening,” Sept. 07, 2025, arXiv : arXiv:2509.06169. doi: 10.48550/arXiv.2509.06169. [134] J. Lee et al. , “BioBERT: a pre - trained biomedical language representation model for biomedical text mining,” Bi oinformatics , vol. 36, no. 4, pp. 1234 – 1240, Feb. 2020, doi: 10.1093/bioinf ormatics/btz682. [135] J. Ma, Y. He, F. Li, L. Ha n, C. You, and B. Wang, “Segment anything in medical images,” Na t. Commun . , vol. 15, no. 1, p. 654, Jan. 2024, doi: 10.1038/s41467 - 024 - 44824 - z. [136] C. Niu et al. , “Medical multimodal multitask foundation model f or lu ng can cer s creening, ” Nat. Commun. , vol. 16, no. 1, p. 1523, Feb. 2025, doi: 10.1038/s41467 - 025 - 56822 - w. 20 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 [137] J. Gui, Z. Sun, Y . Wen, D. Tao, and J. Ye, “A Review on Generative Adver sarial Network s: Algorithms, Theory, and Applications,” Jan. 20, 2020, arXiv : arXiv:2001.06937. doi: 10.48550/arXiv.2001.06937. [138] W. Xia , Y. Shi, C. Niu, W. Cong, and G. Wang, “Diffusion Prior Regularized Iterative Reconstruction for Low - dose CT,” Oct. 10, 2023, arXiv : arXiv :2310.06949. doi: 10.48550/arXiv.2310.06949. [139] C. Niu, C. Wiedeman, M. Li, J. S. Maltz, and G. Wang, “3D Photon Counting CT Image Super - Resol ution Using Conditional Diffusion Model,” Aug. 22, 2024, arXiv : arXiv:2408.15283. doi: 10.48550/arXiv.2408.15283. [140] X. Li u, Y. Xie , J. Ch eng, S. Diao, S. Tan, and X. Liang, “Diffusion Probabilistic Priors for Zero - Shot Low - Dose CT Image Denoising,” July 14, 2023, arXi v : arXiv:2305.15887. doi: 10.48550/arXiv.2305.15887. [141] W. Xia et al. , “Dual - Domain Denoising Dif fusion Probabilistic Model for Metal Artifact Reduction,” IEEE Trans. Radiat. Plasma Med. Sci. , pp. 1 – 1, 2025, doi: 10.1109/TRPMS.2025.3582528. [142] E. J . Hu et al. , “LoRA : Low - Rank Adaptation of Larg e Language Models,” Oct. 16, 2021, arXiv : arXiv:2106.09685. doi: 10.48550/arXiv.2106.09685. [143] “DeepSeek_R1.” [144] Y. Z hao et al. , “PyTor ch FSDP: Experiences on Scaling Fully Sharded Data Pa rallel,” Sept. 12, 2023, arXiv : arXiv:2304.11277. doi: 10.48550/arXiv.2304.11277. [145] R. Y. Aminabadi et al. , “DeepSpe ed Inference: Enabling Efficient Inference of Tra nsformer Models at Unprecedented Scale, ” Jun e 30, 2022, arXiv : arXiv:2207.00032. doi: 10.48550/arXiv.2207.00032. [146] E. J . Hu et al. , “LoRA : Low - Rank Adaptation of Larg e Language Models,” Oct. 16, 2021, arXiv : arXiv:2106.09685. doi: 10.48550/arXiv.2106.09685. [147] Y. Mao et al. , “A Survey on LoRA of Large Language Models,” Front. Comput. Sci. , vol. 19, no. 7, p. 197605, July 2025, doi: 10.1007/s11704 - 024 - 40663 - 9. [148] T. Dao, “FlashAtt ention - 2: Faster Attention with Better Parallelism and Work Partitioning,” July 17, 2023, arXiv : arXiv:2307.08691. doi: 10.48550/arXiv.2307.08691. [149] P. - L. Hsu et al. , “Liger Kernel: Efficient Triton Kernels for LLM Training,” Jan. 24, 2025, arXiv : arXiv:2410.10989. doi: 10.48550/arXiv.2410.10989. [150] P. Micikevicius et al. , “Mixed Precision Training,” Feb. 15, 2018, arXiv : arXiv:1710.03740. doi: 10.48550/arXiv.1710.03740. [151] V. Korthikanti et al. , “REDUCING ACTIVATION RECOMPUTATION IN LARGE TRANSFORMER MODELS”. [152] J. Lamy - Poirier, “Layered gradient accumulation and modular pipeli ne paralleli sm: fast and effici ent training of large language models,” June 04, 2021, arXiv : arXiv:2106.02679. doi: 10.48550/arXiv.2106.02679. [153] G. Hinton, O. Vinyals , and J. Dean, “Distilling the Knowledge in a Neural Network,” Mar. 09 , 2015, arXiv : arXiv:1503.02531. doi: 10.48550/arXiv.1503.02531. [154] S. Li et al. , “Eva luating Quantized La rge Language Models,” June 06, 2024, arXiv : arXiv:2402.18158. doi: 10.48550/arXiv.2402.18158. [155] W. Kwon et al. , “Efficient Memory Manag ement for Large Language Model Serving wi th PagedAt tention,” Sept. 12, 2023, arXiv : arXiv:2309.06180. doi: 10.48550/arXiv.2309.06180. [156] L. Zheng et al. , “SGLang: Efficient Execution of Structured Language Model Programs,” June 06, 2024, arXiv : arXiv:2312.07104. doi: 10.48550/arXiv.2312.07104. [157] Z. Ramzi, G. R. Chaithya, J . - L. Starck, and P. Ciuciu, “NC - PDNet: A density - compensated unrolled network for 2D and 3D non - Cartesian M RI reconstruction,” IEEE Trans. Med. Imaging , v ol. 41, no. 7, pp. 16 25 – 1638, 2022. [158] K. Hammernik et al. , “Learning a varia tional network for reconstruction of accelerated MRI data,” Magn . Reson. Med. , vol. 79, no. 6, pp. 3055 – 3071, June 2018, doi: 10.1002/mrm.26977. [159] J. Adler and O. Öktem, “Learned primal - dual reconstruction,” IEEE Trans. Med. Imaging , vol. 37, no. 6, pp. 1322 – 1332, 2018. [160] “Pruning Unrolled Networks (PUN) at Initialization for MRI Reconst ructio n Improve s General izatio n.” Accessed: Oct . 17, 202 5. [Online]. Available: https://arxiv.org/html/2412. 18668v1 [161] K. Zhang et al. , “Practical Blind Image Denoising via Swin - Conv - UNet and Data Synthes is,” Mach . Int ell. Res. , vol. 20, no. 6, pp. 822 – 836, Dec. 2023, doi: 10.1007/s11633 - 023 - 1466 - 0. [162] L. Zuo et al. , “Unsupervised MR harmonization by learning disentang led representations using information bottleneck theory,” NeuroImage , vol. 243, p. 11 8569, 2021. [163] Y. Li u et al. , “Imaging foundation model for univers al enhancement of non - ideal measurement CT,” Feb. 25, 2025, arXiv : arXiv:2410.01591 . doi: 10.48550/arXiv.2410.01591. [164] S. Arridge , P. Maass, O. Ökt em, and C. - B. Schönl ieb, “Solving inverse problems using data - driven models,” Acta Numer. , vol. 28, pp. 1 – 174, 2019. [165] J. Johnson, A. Alahi, and L. Fei - Fei, “Perceptual Losses for Real - Time Style Transfer and Super - Resol ution,” Mar. 27 , 201 6, arXiv : arXiv:1603.08155. doi: 10.48550/arXiv.1603.08155. [166] M. Terris, S. Hur ault, M. So ng, and J. Tachella, “Reconstruct Anything Model: a lightweight foundation model for c omputation al imaging ,” Sept. 2 7, 2025, arXiv : arXiv:2503.08915. doi: 10.48550/arXiv.2503.08915. [167] H. Wan g et al. , “Towards Unified CT Reconstruction: Federated Metadata Learning Wit h Personalized Condition - Modul ated iRa donMAP,” IEEE Trans. Radiat. Pl asma Med. Sci. , 2025, Accessed: Oct. 17, 2025. [Online]. Available: https://ieeexplore.ieee.org/ abstract/document/11017337/ [168] J. He, Y. Wang, and J. Ma, “Radon inversion via deep learning,” IEEE Trans. Med. Imaging , vol. 39, no. 6, pp. 2076 – 2087, 2020. [169] L. Fu and B. De Man, “A hierarchical approach to deep learning and its application to tom ographic 21 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 reconstruction,” in 15th international meeting on fully three - dimensional image reconst ruction in radiology and nuclear medicine , SPIE, 201 9, p. 1107202. Accessed: Nov. 22, 2025. [Online ]. Available: https://www.spiedigitallibrary. org/conference - proceedings - of - spie/11072/1107202/A - hierarchical - approach - to - deep - learning - and - its - application - to/10.1117/12.2534 615.short [170] B. Zhu, J. Z. Li u, S. F. Cauley, B. R. Rosen, and M. S. Rosen, “I mage rec onstruction by domain - transform manifold learni ng,” Nat ure , vol. 555, no. 7697, pp. 487 – 492, 2018. [171] P. Jain, P. Bv, and N. Sinha, “Novel Complex AUTOMAP for Ac celerated MRI,” in Proceedings of the Thirteenth Indian Conference on Computer Vision, Graphics and Image Processing , Gandh inagar India: ACM, Dec. 2022, pp. 1 – 9. doi: 10.1145/3571600.3571636. [172] H. Chung, S. Lee, a nd J. C. Ye, “Decomposed Diffus ion Sampler for Accelerat ing Large - Scale Inverse Problems,” Feb. 19, 2024, arXiv : arXiv:2303.0 5754. doi: 10.48550/arXiv.2303.05754. [173] J. He, B. Li, G. Yang, and Z. Liu, “Blaze3DM: Marry Triplane Representation with Di ffusion for 3D Medical Inverse Problem Solving,” May 24, 2024, arXiv : arXiv:2405.15241. doi: 10.48550/arXiv.2405.15241. [174] W. Xia, C. Niu, and G. Wan g, “Tomographic Foundat ion Model -- FORCE: Flow - Ori ented Reconstruction Conditioning Engine,” June 02, 2025, arXiv : arXiv:2506.02149. doi: 10.48550/arXiv.2506.02149. [175] S. V. Venkatakrishnan, C. A. Boum an, and B. Wohlberg, “Plug - and - play priors for model based reconstruction,” in 2013 IEEE global conference on signal and information processing , IEEE, 2013, pp. 945 – 948. Accessed: Oct . 17, 202 5. [Online ]. Availabl e: https://ieeexplore.ieee.org/ abstract/document/6737048/ [176] J. - C. Pes quet, A. Repetti, M. Terris, and Y. Wiaux, “Learning Maximally Monotone Operators for Image Recovery,” SIAM J. Imaging Sci. , vol. 14, n o. 3, pp. 1206 – 1237, Jan. 2021, doi: 10.1137/20M1387961. [177] L. I. Rudin, S. Osher , and E. Fatemi, “Nonlinear total variation based noise removal algorithms,” Phys. Nonlinear Phenom. , vol. 60, no. 1 – 4, pp. 259 – 268, 1992. [178] M. Stepha ne, “A wa velet tour of signa l p rocessi ng.” Elsevier, 1 999. Accessed: Oct. 17, 2025. [Online]. Available: https://www.sciencedirect.com/book/9780123743701/a - wavelet - tour - of - signal - processing [179] E. Tiu, E. Tal ius, P . Patel , C. P. Langlotz, A. Y. Ng, and P. Rajpurka r, “Exper t - level detection of pathologies from unannotated chest X - ray images via self - supervised learning,” Nat. Biomed. Eng. , vol. 6, no. 12, pp. 1399 – 1406, 2022. [180] S. Pai et al. , “Foundation model for cancer imaging biomarkers,” Nat. Mach. Intell. , vol. 6, no. 3, pp. 354 – 367, 2024. [181] H. Xu et al. , “A whole - slide foundation model for digital pathology from real - wor ld data,” Nature , vol. 630, no. 8015, pp. 181 – 188, 2024. [182] H. Yu, Q. Wang, X. Zhou, L. Gu, and Z. Zh ao, “Deep weighted sur vival neura l networks t o survival risk prediction,” Complex Intell. Syst. , vol. 11, no. 1, p. 41, Jan. 2025, doi: 10.1007/s40747 - 024 - 01670 - 2. [183] T. Vu et al. , “Tabular Foundation Model for Breast Cancer Prognosis using Gene Expression Data,” Oct. 05, 2025, medRxiv . doi: 10.1101/2025.10.03.25337 265. [184] J. Long, E. Shelhamer, and T. Darrell, “Fully convolutional networks for semantic segmentation,” in Proceedings of the IEEE conference on c omputer visi on and pattern recogniti on , 2015, pp. 3431 – 3440. Accessed: Nov. 22, 2025. [Online ]. Available: http://openaccess.thecvf.com/content_cvpr_2015/ html/L ong_Fully_Convolutional_Networks_2015_CVPR_pap er.html [185] J. Ma, Y. He, F. Li, L. Ha n, C. You, and B. Wang, “Segment anything in medical images,” Na t. Commun . , vol. 15, no. 1, p. 654, Jan. 2024, doi: 10.1038/s41467 - 024 - 44824 - z. [186] S. Noh and B. - D. Lee, “A narrative review of foundation models for medical image segmen tation: zero - shot performance evaluation on diverse modalities,” Quant. Imaging Med. Surg. , vol. 15, no. 6, pp. 5825 – 5858, June 2025, doi: 10.21037/qims - 2024 - 2826. [187] A. Kiril lov et al. , “Segment Any thing,” Apr. 05, 2023, arXiv : arXiv:2304.02643. doi: 10.48550/arXiv.2304.02643. [188] K. Fan, L. Liang, H. Li, W. Situ, W. Zhao , and G. Li , “Research on Medical Image Segmentation Based on SAM and Its Future Prospects,” Bio engineering , vol. 12, no. 6, p. 608, 2025. [189] J. Ma, Y. He, F. Li, L. Ha n, C. You, and B. Wang, “Segment anything in medical images,” Na t. Commun . , vol. 15, no. 1, p. 654, 2024. [190] W. Le i, W. Xu , K. Li, X. Zha ng, and S. Zhan g, “MedLSAM: Localize and segment anything model for 3D CT images,” Med. Image An al. , vol. 99, p. 103370, 2025. [191] S. Gong et al. , “3dsam - adapter: Holistic adaptation of sam from 2d to 3d for promptable medical image segmentation,” ArXiv E - Prints , p. arXiv - 2306, 2023. [192] J. Sun et al. , “Medical image analysis using improved SAM - Med2D: segmentation and classif icatio n perspectives,” BMC Med. Imaging , vol. 24, no. 1, p . 241 , Sept. 2024, doi: 10.1186/s12880 - 024 - 01401 - 6. [193] Z. Yan et al. , “SAMed - 2: Selective Memory Enhanced Medical Seg ment Anythi ng Model,” i n Medical Imag e Computing and Compu ter Assisted Intervention – MICCAI 2025 , vol. 15 972, J. C. G ee, D. C. Alexander, J. Hong, J. E. Ig lesias, C. H. Sudre, A. Venkat araman, P. Golland, J. H. Kim, and J. Park, Eds., in Lecture Notes in Computer Scie nce, vol. 15972. , Cham: Springer Nature Switzerland, 2026, pp. 540 – 550. doi: 10.1007/978 -3- 032 - 05169 - 1_52. [194] G. Deng et al. , “SAM - U: Multi - box Prompts Triggered Uncertainty Esti mation for Reliable SAM in Medical Image,” in Medical Ima ge Compu ting a nd Compu ter Assisted Int ervention – MICC AI 2023 Worksh ops , vol. 14394, J. Woo, A. Hering, W. Silva, X. Li, H. Fu, X. Liu, 22 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 F. Xing, S. Purushotham, T. S. Mathai, P. Mukherjee, M. De Grauw, R. Beets Tan, V. Corbetta, E. Kotter, M. Reyes, C. F. Ba umgartner, Q. Li, R. Leahy, B. Dong, H. Chen, Y. Huo , J . Lv, X. Xu, X. Li, D. Mahapatra, L. Cheng, C. Petitjean, and B. Presles, Eds., in Lecture Notes in Computer Scie nce, vol. 1439 4. , Cham: Spri nger Nature Switzerland, 2023, pp. 368 – 377. doi: 10.1007/978 -3- 031 - 47425 - 5_33. [195] Y. Tian, M. Shi , X. Zhang, B. Zhang, M. Wang, and Y. Shi, “Assist ing embodied AI: a survey of 3D segmentation models for medical CT images: Y. Tian et al.,” CCF Trans . Pervasi ve Comput. Interact. , pp. 1 – 22, 2025. [196] J. Cox et al. , “BrainSegFoun der: Towards 3D foundation models for ne uroimage segmenta tion,” Med. Image Anal. , vol. 97, p. 103301, 2024. [197] B. B. Avants, C. L. Epstein, M. Grossman, and J. C. Gee, “Symmetric diffeomorphic image registration with cross - correlation: evaluating automated labeling of elderly and neurodegenerative brain,” Med. Image Anal. , vol. 12, no. 1, pp. 26 – 41, 2008. [198] G. Balakrish nan, A. Zhao, M. R. Sabuncu, J. Guttag, and A. V. Dalca, “Voxel Morph: A Learning Fr amework for Deformable Medi cal Image Reg istration ,” IEEE Trans. Med. Imaging , vol. 38, n o. 8, pp. 1788 – 1800, Aug. 2019, doi: 10.1109/TMI.2019.2897538. [199] J. Chen, E. C. Frey, Y. He, W. P. Segars, Y. Li, and Y. Du, “TransMor ph: Transf ormer for unsupervised medical image re gistrati on,” Med. Image Anal. , vol. 82, p. 102615, Nov. 2022, doi: 10.1016/j.media.2022.102615. [200] M. Hoffmann, B. Billot, D. N. Gr eve, J. E. Iglesias, B. Fischl, and A. V. Dalca, “SynthMorph: Learning Contrast - Invariant Registration Without Acquired Images,” IEEE Trans. Med. Imaging , vol. 41, no. 3, pp. 543 – 558, Mar. 2022, doi: 10.1109/TMI.2021.3116879. [201] M. P. He inrich and L. Hansen, “Voxelm orph++ Going beyond the cranial vault with keypoint super vision and multi - channel ins tance optimisation,” Feb. 28, 2022, arXiv : arXiv:2203.00046. doi: 10.48550/arXiv.2203.00046. [202] L. Tian et al. , “uniGradICON: A Foundatio n Model for Medical Image Registr ation,” in Medical Image Computing and Compu ter Assisted Intervention – MICCAI 2024 , vol. 15002, M. G. Linguraru, Q. Dou, A. Feragen, S. Giannarou, B. Glocker, K. Lekadir, and J. A. Schnabel, Eds., in Lecture Notes in Computer Science, vol. 15002. , Cham: Springer Nature Swit zerland, 2024, pp. 749 – 760. doi: 10.1007/978 -3- 031 - 72069 - 7_70. [203] B. De mir et al. , “MultiG radICON: A Foundation Model for Multimodal Medical Image Registration,” in Biomedical Image Registration , vol. 15249, M. Modat, I. Simpson, Ž. Špiclin, W. Bastiaansen, A. Hering, and T. C. W. Mok, Eds., in Lecture Notes in Computer Science, vol. 15249. , Cham: Springer Nature Swit zerland, 2024, pp. 3 – 18. doi: 10.1007/978 -3- 031 - 73480 - 9_1. [204] J. Hu, K. Yu, H. Xian, S. Hu, and X. Wang, “Improving Generalizati on of Medi cal Image Registration Foundation Model,” May 10, 2025, arXiv : arXiv:2505.06527. doi: 10.48550/arXiv.2505.06527. [205] X. Song , X. Xu, an d P. Yan, “Ge neral Purpos e Image Encoder DINOv2 for Medical Image Registratio n,” Feb. 24, 2024, arXiv : arXiv:2402.15687. doi: 10.48550/arXiv.2402.15687. [206] H. Xu et al. , “MultiCo3D: Multi - Label Voxel Co ntrast for One - Shot Incremental Segmentati on of 3D Neuroimages,” Mar. 09, 2025, arXiv : doi: 10.48550/arXiv.2503.06598. [207] H. Sin gh, “Generat ive AI for Synthetic Data Creation: Solving Dat a Scar city in Machine Lea rning,” Av ailable SSRN 5267914 , 2025, Accessed: Nov. 22, 2025. [Online]. Available: https://papers.ssrn.com/sol3/ papers.cfm?abstract_id=52 67914 [208] V. Thambawita et al. , “SinGAN - Seg: Synthetic tra ining data generation for medical image segmentation, ” PloS One , vol. 17, no. 5, p. e026797 6, 2022. [209] L. R. Koetzier et al. , “Generating Synthetic Data for Medical Imagi ng,” Radiology , vol. 312, no. 3, p. e232471, Sept. 2024, doi: 10.1148/radiol.232471. [210] C. - Y. Ung et al. , “Artificial Clinic Intelligence (ACI): A Generative AI - Powered Modeling Platform to Optimize Patient Cohort Enri chment and Clinical Trial Optimization, ” Cancers , vol. 17, no. 21, p. 3543, 2025. [211] B. D. Killeen et al. , “Towards Virtual Clinical Trials of Radiology AI with Conditional Generative Modeling, ” Feb. 13, 2025, arXiv : arXiv:2502.09688. doi: 10.48550/arXiv.2502.09688. [212] S. S. Bhuyan et al. , “Generative Artificial Intelligence Use in He althcare: Opportunit ies for Clinical Excellenc e and Administrative Efficiency, ” J. Med. Syst. , vol. 49, no. 1, p. 10, Jan. 2025, doi: 10.1007/s10916 - 024 - 02136 - 1. [213] J. S. Yoon, C. Zhang, H . - I. S uk, J. Guo, and X. Li, “SADM: Sequence - Aware Diffus ion Model for Longitudinal Medical Image Generation,” in Information Processing in Medical Imagi ng , vol. 13939, A. Frangi, M. De Bruijne, D. Wassermann , and N. Navab, Eds., in Lecture Notes in Computer Scie nce, vol. 13939. , Cham: Springer Nature Switzerland, 2023, pp. 388 – 400. doi: 10.1007/978 -3- 031 - 34048 - 2_30. [214] L. X. Nguyen, P. S. Aung, H. Q. Le, S. - B. Park, and C. S. Hong, “A new chapter for medica l i mage gener ation: The stable dif fusion method,” in 2023 International Conference on I nformation Net working (ICOIN) , IEEE, 2023, pp. 483 – 486. Accessed: Nov. 22, 2025. [Online]. Available: https://ieeexplore.ieee.org/ abstract/document/10049010/ [215] N. Konz, Y. Chen, H. Dong, and M. A. Mazurowski , “Anatomically - Controllable Medi cal Image Genera tion with Segmen tation - Guided Diffusi on Model s,” in Medical Image Comput ing a nd Co mputer Assis ted Intervention – MIC CAI 2024 , vol. 15007, M. G. Linguraru, Q. Dou, A. Feragen, S. Gi annarou, B. Glocker, K. Lekadir, and J. A. Schnabel, Eds., in Lecture Notes in Computer Scie nce, vol. 1500 7. , Cham: Spri nger Nature Switzerland, 2024, pp. 88 – 98. doi: 10.1007/978 - 3- 031 - 72104 - 5_9. 23 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 [216] F. Khader et al. , “Denoising diffusion probabilistic models for 3D medical image generati on,” Sci. Rep. , vol. 13, no. 1, p. 7303, 2023. [217] A. L. Y. Hung et al. , “Med - cdiff: Conditional medical image generation with diffusion models,” Bioengineering , vol. 10, no. 11, p. 1258, 2023. [218] J. Wang et al. , “Self - improving generative foun dation model for synthetic med ical ima ge generat ion and clinical applications,” Nat. Med. , vol. 31, no. 2, pp. 6 09 – 617, Feb. 2025, doi: 10.1038/s41591 - 024 - 03359 - y. [219] M. Cobbin ah et al. , “Diversity in Stable GANs: A Systematic Review of Mode Collapse Mitigation Strategies,” Eng. Rep. , vol. 7, no. 6, p. e70209, June 2025, doi: 10.1002/eng2.70209. [220] C. Guo, A. N. Christensen, and M. R. Hannemose , “Med - Art: Diffusion Transformer for 2D Medica l Text - to - Image Generation,” June 25, 2025, arXiv : arXiv:2506.20449. doi: 10.48550/arXiv.2506.20449. [221] Y. Dai, Y. Gao, and F. Liu, “Transmed : Transf ormers advance multi - modal medical ima ge classif ication ,” Diagnostics , vol. 11, no. 8, p. 1384, 2021. [222] Z. Xia, H. Li, and L. Lan, “MedFormer: Hierarchical Medical Visi on Tr ansform er wi th Content - Aware Dual Sparse Selection Attention,” Aug. 05, 2025, arXi v : arXiv:2507.02488. doi: 10.48550/arXiv.2507.02488. [223] P. Chambon et al. , “RoentGen : Vision - Language Foundation Model for Chest X - ray Generation,” Nov. 23, 2022, arXiv : arXiv:2211.12737 . doi: 10.48550/arXiv.2211.12737. [224] Q. Chen et al. , “ Towards generalizable tumor synthesis,” in Proceedi ngs of the IEEE/CVF conference on c omputer vision and pattern recognition , 2024, pp. 11147 – 11158. Accessed: Oct . 17, 202 5. [Online ]. Availabl e: http://openaccess.thecvf.com/content/ CVPR2024/html/ Chen_Towards_Genera lizable_Tumor_Syn thesis_CVP R_2024_paper.html [225] L. Zhang, A. Rao, and M. Agrawala, “Addi ng condit ional control to text - to - image diffusion models,” in Proceedings of the IEEE/CVF international conference on computer vision , 2023, pp. 3836 – 3847. Accessed: Nov. 22, 2025. [Online ]. Available: http://openaccess.thecvf.com/content/ ICCV2023/html/Z hang_Adding_Conditional_Control_to_Text - to - Image_Diffusion_Models_IC CV_2023_paper.html [226] I. E. Hamamci et al. , “GenerateCT: Text - Conditional Generation of 3 D Chest CT Vo lumes,” in Computer Vision – ECCV 2024 , vol. 1513 7, A. Leonardis, E. Ricci, S. Roth, O. Russakovsky, T. Sattler, and G. Varol, Eds., in Lectu re Notes in Computer S cience, vol. 15137. , Cham: Spr inger Nature Switzerland, 2025, pp. 126 – 143. doi: 10.1007/978 -3- 031 - 72986 - 7_8. [227] C. Zhao et al. , “MAISI - v2: Accelerated 3D High - Resolution M edical Image Synthesi s with Rectified Flow and Region - specific Contrastive Loss,” Aug. 07, 2025, arXiv : arXiv:2508.05772. doi: 10.48550/arXiv.2508.05772. [228] P. Guo et al. , “Maisi: Medical ai for synthetic imaging,” in 2025 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV) , IEEE, 2025, pp. 4430 – 4441. Accessed: Oct . 17, 202 5. [Online ]. Availabl e: https://ieeexplore.ieee.org/ abstract/document/10943915/ [229] S. - C. Huang, M. Jensen, S. Yeun g - Levy, M. P. Lungren, H. P oon, and A. S. Chaudhari, “Multimodal Foundation Models for Med ical Imaging - A Sys tematic Review and Implementation Guidelines,” medRx iv , pp. 2024 – 10, 2024. [230] C. Wu, X. Zhang, Y. Zhang, H. Hui, Y. Wang, and W. Xie, “Towards generalis t foundation m odel for radiology by leveraging web - scale 2d&3d medical data,” Nat. Commun. , vol. 16, no. 1, p. 7866, 2025 . [231] S. - C. Huang, M. Jensen, S. Yeun g - Levy, M. P. Lungren, H. P oon, and A. S. Chaudhari, “Multimodal Foundation Models for Med ical Imaging - A Sys tematic Review and Implementation Guidelines,” medRx iv , pp. 2024 – 10, 2024. [232] O. Banerjee, H. - Y. Zhou, S. Adi than, S . Kwak, K. Wu, and P. Rajpurkar, “Direct preference optimization for suppressing hallucinated prior exams in radiology report generation,” ArXiv Pre pr. ArXiv24060649 6 , 2024, Accessed: Nov. 24, 2025. [Onli ne]. Availabl e: https://arxiv.org/abs/2406.06496 [233] E. - D. M. E. by Direct, “Ra dQA - DPO: A Radiology Question Ans wering Syst em with Encod er - Decoder Models Enhance d by Dir ect Preferen ce Op timizat ion”, Accessed: Nov. 24, 2025. [Onli ne]. Availabl e: https://aclanthology.org/2025.bi onlp - 1.10v2.pdf [234] S. Pai et al. , “Foundation models for quantitative biomarker discovery in cancer imaging,” MedRxiv , 2023, Accessed: Oct . 17, 202 5. [Online ]. Availabl e: https://pmc.ncbi.nlm.nih.gov/art icles/PMC10508804/ [235] A. S. Tejani , T. S. Cook, M. Hussain, T. Sippel Schmidt, and K. P. O’Donnell, “Int egrating and Adopting AI in the Radiology Workflow: A Primer for Standards and Integrating the Healthcare Enterprise (IHE ) Profiles,” Radiology , vol. 311, no. 3, p. e23265 3, June 2024, doi: 10.1148/radiol.232653. [236] A. Vian, D. A. Eif er, M. Anes, G. R. Garcia , and M. Recamonde - Mendoza , “Explor ing the F easibi lity of AI - Assisted Spine MRI Protocol Optimizati on Us ing DICOM Image Metadata,” Feb. 04, 2 025, arXiv : arXiv:2502.02351. doi: 10.48550/arXiv.2502.02351. [237] L. Mel azzini et al . , “AI for image quality and patient safety in CT and MRI,” Eur. Radio l. Exp. , vol. 9, no. 1, p. 28, Feb. 2025, doi: 10.1186/ s41747 - 025 - 00562 - 5. [238] H. R. Tizhoosh, “Foundation models and informatio n retrieval in digital pathology,” in Art ificial Intelligence i n Pathology , Elsevier, 2025, pp. 211 – 232. Accessed: Oct. 17, 2025. [Online]. Available: https://www.sciencedirect.com/science/art icle/pii/B9780 32395359700011X [239] K. Clark et al. , “The Canc er Imaging Archive (TCIA): Maintain ing and Operat ing a Public In formati on Repository,” J. Digit. I maging , vol. 26, no. 6, pp. 1045 – 1057, Dec. 2013, doi: 10.1007/s10278 - 013 - 9622 - 7. [240] “MIDRC,” MIDRC. Accessed: Nov. 27, 2025. [Online]. Available: https://www. midrc.org [241] J. Zbontar et al. , “fastMRI: An Open Dataset and Benchmarks for Acce lerated MRI,” Dec . 11, 2019, 24 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 arXiv : arXiv:1811.08839. doi: 10.48550/arXiv.1811.08839. [242] F. Liu, J. V. Velikina, W. F. Block, R. Kijowski, and A. A. Samsonov, “Fast Realistic MRI Simulations Based on Generalized Multi - Pool Exchange Tissue Model,” IEEE Trans. Med. Imaging , vol. 36, no. 2, pp. 527 – 537, Feb. 2017, doi: 10.1109/TMI.2016.2620961. [243] C. H. McCollough et al. , “Low‐dose CT for the detection and classification of metastatic liver lesions: Results of the 2016 Low Dose CT Grand Challenge,” Med . Phy s. , vol. 44, no. 10, Oct. 2017, doi: 10.1002/mp.12345. [244] “Grand Challenge.” Accessed: Nov. 27, 2025. [Online]. Available: https://www.aapm.org/GrandChallenge/Default.asp [245] X. Wang, Y. Peng, L. Lu, Z. Lu, M. Bagheri, and R. M. Summers, “ChestX - ray8: Hospital - Scale Chest X - Ray Database and Benchmarks on Weakly - Supervised Classificat ion and Localizati on of Common Thorax Diseases”. [246] J. Irvin et al. , “CheXpert: A Larg e Chest Radiogra ph Dataset with Uncertainty Labels and Expert Comparison,” Proc. AAAI Conf. Artif. Intell. , v ol. 33, no. 01, pp. 590 – 597, July 2019, doi: 10.1609/aaai.v33i01.3301590. [247] A. E. W. Johnson et al. , “MIMIC - CXR, a de - identified publicly available database of chest radiographs with free - text reports,” Sci. Data , vol. 6, no. 1, p. 317, Dec. 2019, doi: 10.1038/s41597 - 019 - 0322 - 0. [248] K. Yan, X. Wang, L. Lu, and R. M. Summers, “DeepLesion: automated mining of large - scale lesion annotations and universal lesion detection with deep learning,” J. Med. Imaging , vol. 5, no. 03, p. 1, July 2018, doi: 10.1117/1.JMI.5.3.036501. [249] X. Mei et al. , “RadImageNet: An Open Radiolo gic Deep Learning Res earch Datas et for Effective Transfer Learning,” Radiol. Artif. Intell. , vol. 4, no . 5, p. e210315, Sept. 2022, doi: 10.1148/ryai .210315. [250] B. H. Menz e et al. , “The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS),” IEEE Trans. Med. Imaging , vol. 34, no. 10, pp. 1993 – 2024, Oct. 2015, doi: 10.1109/TMI.2014.2377694. [251] D. Ouyang et al. , “Video - based AI for beat - to - beat assessment of cardiac function,” Nat ure , vol. 580, no. 7802, pp. 252 – 256, Apr. 2020, doi: 10.1038/s41586 - 020 - 2145 - 8. [252] F. Prior et al. , “T he public ca ncer radiology imaging collections of The Cancer Imaging Archive,” Sci. Data , vol. 4, no. 1, p. 170124, Sept. 2017, doi: 10.1038/sdata.2017.124. [253] W. Bulten et al. , “Artificia l intelligence for diagnosis and Gleason gra ding of prostate can cer: the PANDA challenge,” Nat. Med. , vol. 28, no. 1, pp. 154 – 163, Jan. 2022, doi: 10.1038/s41591 - 021 - 01620 - 2. [254] O. Pelka, S. Koitka, J . Rückert, F. Nensa, and C. M. Friedrich, “Radiology Objects i n COntext (ROCO): A Multimod al Image Dataset,” in Intravascular Imaging and Computer Assisted Stenti ng and Large - Scale Annotation of Biomedical Data and Expert Label Synthesis , D. Stoyanov, Z. Taylor, S. Balocco, R. Sznitman, A. Martel, L. Maier - Hein, L . Duong, G. Zahnd, S. Demirci, S. Albarqouni, S. - L. Lee, S. Moriconi , V. Cheplygin a, D. Mateus, E. Trucc o, E. Granger, and P. Jann in, Eds., Cha m: Spri nger International Publishing, 2018, pp. 180 – 189. [255] S. Subramanian et al. , “MedICaT: A Dataset of Medical Images, Captions, and Textual References,” in Findings of the Association for Computational Linguistics: EMNLP 2020 , Online: Association fo r Computational Linguistics, 2020, pp. 2112 – 2120. doi: 10.18653/v1/2020.findings - emnlp.191. [256] J. Yang et al. , “MedMNIST v2 - A large - scale lightweight benchm ark for 2D and 3D bio medical image classification,” Sci. Data , vol. 10, no . 1, p. 41, Ja n. 2023, doi: 10.1038/s41597 - 022 - 01721 - 8. [257] M. Antonelli et al. , “The Medical Segmentation Decathlon,” Nat. Commun. , vol. 13, no. 1, p. 4128, July 2022, doi: 10.1038/s41467 - 022 - 30695 - 9. [258] R. Souza et al. , “An open, m ulti - vendor, multi - field - strength brain MR dataset and analysis of publicly available skul l stri pping methods agreement.,” NeuroImage , vol. 170, pp. 482 – 494, Apr. 2018, doi: 10.1016/j.neuroimage.2017.08.021. [259] S. G. Armato et al. , “The Lung Image Database Consortium (LIDC) and Image Dat abase Resource Initiative (IDRI): A Completed Reference Database of Lung Nod ules on CT Sca ns,” Med. Phys . , vol. 38, no. 2, pp. 915 – 931, Feb. 2011, doi: 10.1118/1.3528204. [260] T. J. Littl ejohns et al. , “The UK Biobank imaging enhancement of 100,000 participants: rationale, data collection, management and future directions,” Na t. Commun. , vol. 11, no. 1, p. 2624, May 2020, doi: 10.1038/s41467 - 020 - 15948 - 9. [261] The Nat ional Lung Screening Tria l Research Team, “Reduced Lung - Cancer Mort ality with Low - Dose Computed To mographic Screening,” N. Engl. J . Med. , vol. 365, no. 5, pp. 395 – 409, Aug. 2011, doi: 10.1056/NEJMoa1102873. [262] D. S. Marcus, T. H. Wang, J. Parker, J. G. Csernansky, J. C. Morris, and R. L. Buc kner, “Open Acces s Series of Imaging Studies (OASIS): Cross - sectional MRI Data in Young, Middle Aged, Nondemented, a nd Demented Older Adul ts,” J. Cogn. Neurosci. , vol. 19, no. 9, pp. 1498 – 1507, Sept. 2007, doi: 10.1162/jocn.2007.19.9.1498. [263] D. C. Van Essen, S. M. Smith, D. M. Ba rch, T. E. J. Behrens, E. Ya coub, an d K. Ugurbil, “The WU - Minn Human Connec tome Proj ect: An overvie w,” Mapp. Connect. , vol. 80, pp. 62 – 79, Oct. 2013, doi: 10.1016/j.neuroimage.2013.05.041. [264] A. L. Goldberger et al. , “PhysioBank, PhysioToolkit, and PhysioNet: Components of a New Research Resource for Complex Physiologic Signals,” Circulation , vo l. 101, no. 23, June 2000, doi: 10.1161/01.CIR.101. 23.e215. [265] “Alexandr Wang on AI’s Potential and Its ‘Deficiencies’ | TIME.” Accessed: Aug. 31, 2025. [Online]. Available: https://time.com/7296215/alexandr - wang - interview/?utm_source =chatgpt.com 25 IEEE TRANSACTIONS O N MEDICAL IMAGING, VO L. xx, NO. x , 2020 [266] W. Yang, H. Z hang, W. Tan, Y. Su n, and B. Yan, “A Self - Supervised Paradigm for Data - Efficient Medical Foundation Model Pr e - training: V - information Optimization Framework, ” Apr. 06, 2025, arXiv : arXiv:2408.07107. doi: 10.48550/arXiv.2408.07107. [267] I. Dayan et al. , “Federated learning for predicting clinical outcomes in patients with COVID - 19,” Nat. Med. , vol. 27, no. 10, pp. 1735 – 1743, Oct. 2021, doi: 10.1038/s41591 - 021 - 01506 - 3. [268] X. Zha ng, H. Den g, R. Wu, J. Re n, and Y. Ren, “PQSF: post - quantum secure privacy - preserving federated learning,” Sci. Rep. , vol. 14, no. 1, p. 23553, Oct. 2024, doi: 10.1038/s41598 - 024 - 74377 - 6. [269] Y. Shi, W. Xia, C. Niu , C. Wie deman, and G. Wang, “Enabling Com petitive Performance of Medical Im aging with Diffusi on Model - generated Images without Privacy Leakage,” Feb. 15, 2024, arXiv : 10.48550/arXiv.2301.06604. [270] K. Huang et al. , “A foundation model for clinician - centered drug repurposing,” Nat. Med. , vol. 30, no. 12, pp. 3601 – 3613, Dec. 2024, doi: 10.1038/s41591 - 024 - 03233 - x. [271] Z. Wang, E. Khatibi, and A. M. Rahmani, “MedCoT - RAG: Causal Chain - of - Thought RAG for Medical Question Answe ring,” Aug. 20, 2025, ar Xiv : arXiv:2508.15849. doi: 10.48550/arXiv.2508.15849. [272] N. Fa thi, T. Scholak, and P. - A. Noël, “Unifying Autoregress ive and Dif fusion - Based Seq uence Generation, ” Apr. 08, 2025, arXi v : doi: 10.48550/arXiv.2504.06416. [273] J. Hu et al. , “ACDiT: Interpolating Autoregressive Conditional Modeling and Diffusion Transfor mer,” Mar. 13, 2025, arXiv : arXiv:2412.07720. doi: 10.48550/arXiv.2412.07720. [274] S. Azizi, S. Kundu, M. E. Sadeghi, and M. Pedram, “MambaExtend: A Training - Free Approa ch to Improve Long Co ntext Extension of Mamba,” prese nted at the The Thirteenth International Conference on Learning Representatio ns, Oct. 2024. Acce ssed: Sept. 01, 2025. [Online]. Available: https://openreview.net/forum?id=LgzRo1RpLS&utm_s ource=chatgpt.com [275] “On the Promise for Assurance of Differentiable Neurosymbolic Reasoni ng Par adigms.” Accessed: Sept. 01, 2025. [Online]. Available: https://arxiv.org/html/2502. 08932v1?utm_source=chatg pt.com [276] G. Wang and F. - L. Fan, “Dimensionality and dynamics for next - generation artifi cial neural networks,” Patterns , vol. 6, no. 8, Aug. 2025, doi: 10.1016/j.patter.2025.101231. [277] S. Sinha, P. Gehler, F. Locate llo, and B. Schiele, “TeST: Test - time Self - Trainin g under Distrib ution Shift,” Sept. 23, 2022, arXiv : arXiv:2209.11459. doi: 10.48550/arXiv.2209.11459. [278] DeepSeek - AI et al. , “DeepSeek - V3 Technical Report,” Feb. 18, 2025, arXiv : arXiv:2412.19437. doi: 10.48550/arXiv.2412.19437. [279] J. Liang, R. He, and T. Tan, “A Comprehensive Survey on Test - Time Adaptation under Distributi on Shif ts,” Int. J. Comput. Vis. , vol. 133, no. 1, pp. 31 – 64, Jan. 2025, doi: 10.1007/s11263 - 024 - 02181 - w. [280] M. Moor et al. , “Foundation models for generalist medical artificial intelligen ce,” Nat ure , vol. 616, no. 7956, pp. 259 – 265, Apr. 2023, doi: 10.1038/s41586 - 023 - 05881 - 4. [281] H. Chao et al. , “Deep learning predicts cardiovascular disease risks from lung cancer screening l ow dose computed tomography,” Nat. Commun. , vol. 12, no . 1, p. 2963, May 2021, doi: 10.1038/s41467 - 021 - 23235 - 4. [282] C. Niu et al. , “Medical multimodal multitask foundation model f or lu ng can cer s creening, ” Nat. Commun. , vol. 16, no. 1, p. 1523, Feb. 2025, doi: 10.1038/s41467 - 025 - 56822 - w. [283] R. Harrison, “Alpha + Beta har dware (fall 2025),” Empire AI. Accessed: Sept. 01, 2025. [Online]. Available: https://empireai.freshdesk.com/support /solutions/articles /157000363466 - alpha - beta - hardware - fa ll - 2025 - [284] “A Survey of Chain of Thought Reasoning: Advances, Frontiers and Future,” ar5iv. Accessed: Sept. 01, 2025. [Online]. Available: https://ar5iv.labs.arxiv. org/html/2309.15402 [285] L. Jiao et al. , “C ausal Infere nce Mee ts Deep Learning: A Comprehensive Survey,” Res. Wash. DC , vol. 7, p. 0467, 2024, doi: 10.34133/research.0467. [286] X. Yu et al. , “Causal Sufficiency and Necessity Improves Chain - of - Thought Reasoning,” July 26, 2025, arXiv : arXiv:2506.09853. doi: 10.48550/arXiv.2506.09853. [287] K. Al hamoud et al. , “FedMedICL: Towards Holistic Evaluation of Distributi on Shifts in Federated Medical Imaging,” July 11, 2024, arXiv : arXiv:2407.08822. doi: 10.48550/arXiv.2407.08822. [288] H. Guan, P. - T. Yap, A. Bozoki, and M. Liu, “Federated learning for medical image analysis: A survey,” Pattern Recognit. , v ol. 151, p. 1 10424, July 2024, doi: 10.1016/j.patcog.2024.110424. [289] G. Pap adopoulos, P. J. Edwards, a nd A. F. Mu rray, “Confidence est imation methods for neural networks: a practical comparison,” IEEE Trans. Neural Netw. , vol. 12, no. 6, pp. 1278 – 1287, Nov. 2001, doi: 10.1109/72.963764. [290] Z. Shi et al. , “Generate - then - Ground in Retri eval - Augmented Generation for Mul ti - hop Question Answering,” in Proceedings of the 62nd Annual Meeting of the Association for Computational Lingui stics (Volume 1: Long Papers) , Bangkok, Thailand: Association for Co mputational Linguistics, 2024, pp. 7339 – 7353. doi: 10.18653/v1/2024.acl - long.397.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment