StrokeNeXt: A Siamese-encoder Approach for Brain Stroke Classification in Computed Tomography Imagery
We present StrokeNeXt, a model for stroke classification in 2D Computed Tomography (CT) images. StrokeNeXt employs a dual-branch design with two ConvNeXt encoders, whose features are fused through a lightweight convolutional decoder based on stacked …
Authors: ** *제공된 원문에 저자 정보가 명시되지 않았습니다.* (논문 본문이나 메타데이터에서 확인 필요) **
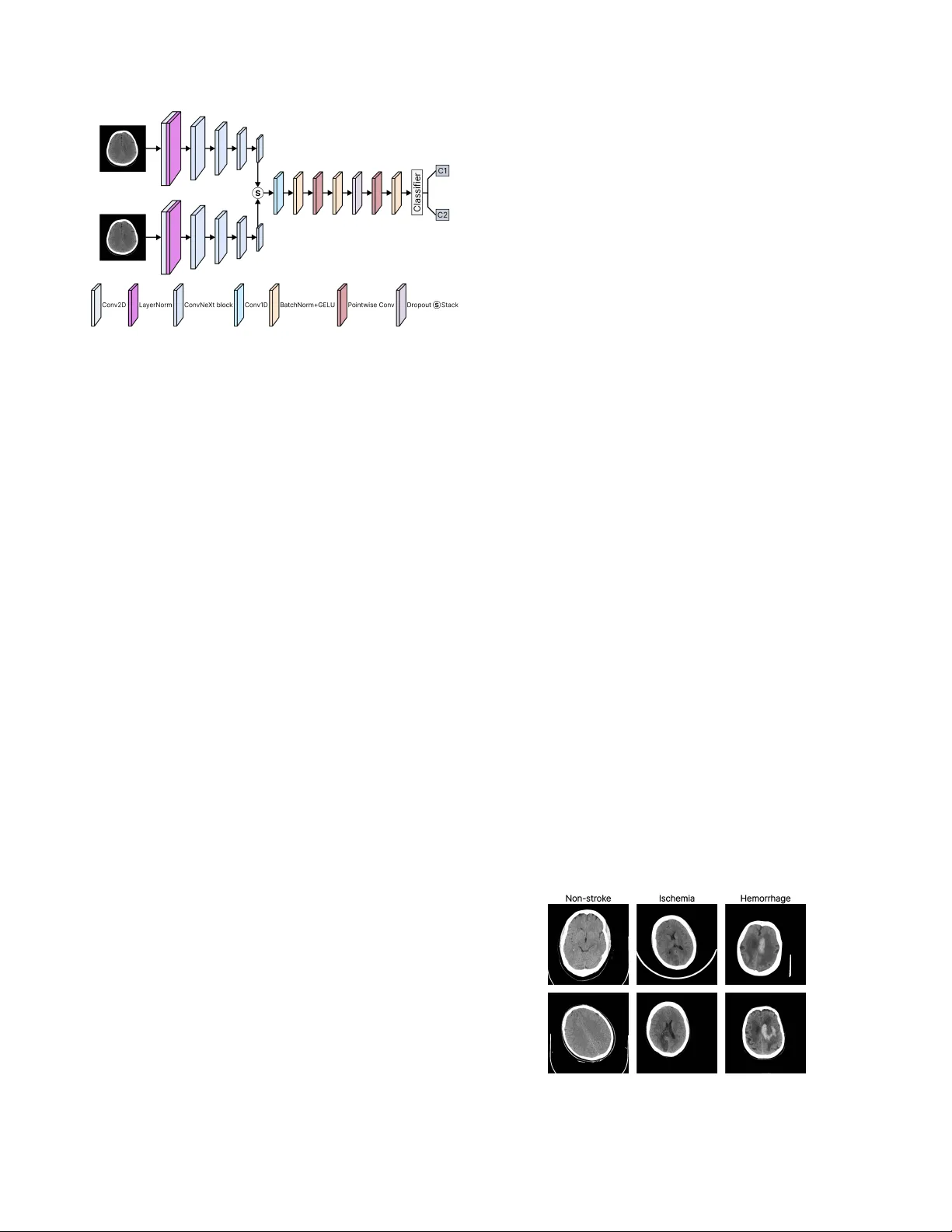
Str okeNeXt: A Siamese-encoder A ppr oach f or Brain Str ok e Classification in Computed T omography Imagery Leo Thomas Ramos 1,2 Angel D. Sappa 1,2,3 1 Computer V ision Center 2 Uni versitat Aut ` onoma de Barcelona 3 ESPOL Polytechnic Uni versity Abstract W e present Strok eNeXt, a model for str ok e classification in 2D Computed T omo graphy (CT) ima ges. Str ok eNeXt em- ploys a dual-branc h design with two Con vNeXt encoders, whose featur es ar e fused thr ough a lightweight convolu- tional decoder based on stack ed 1D operations, including a bottlenec k pr ojection and transformation layers, and a compact classification head. The model is evaluated on a curated dataset of 6,774 CT ima ges, addr essing both str oke detection and subtype classification between ischemic and hemorrhage cases. Str ok eNeXt consistently outperforms con volutional and T r ansformer-based baselines, r eac hing accuracies and F1-scores of up to 0.988. P air ed statis- tical tests confirm that the performance gains ar e statisti- cally significant, while class-wise sensitivity and specificity demonstrate rob ust behavior across diagnostic cate gories. Calibration analysis shows r educed pr ediction err or com- par ed to competing methods, and confusion matrix results indicate low misclassification rates. In addition, the model exhibits low inference time and fast con ver gence . Code is available at: www.hidden_for_review.com 1. Introduction Brain stroke is a life-threatening medical condition and a leading cause of adult mortality and long-term disability worldwide, af fecting millions of indi viduals each year [ 30 ]. It occurs when cerebral blood flow is disrupted due to v as- cular blockage or rupture [ 13 , 25 ], depriving brain tissue of oxygen and nutrients and resulting in cellular injury or death [ 13 ]. Owing to its abrupt onset and sev ere clinical consequences, stroke remains a major challenge in emer- gency medicine and neurology [ 11 ]. Clinically , strokes are broadly categorized into ischemic and hemorrhagic types, depending on whether they are caused by vascular blockage or rupture [ 13 ]. T ypically , initial assessment relies on neuroimaging techniques such as Magnetic Resonance Imaging (MRI) and Computed T o- mography (CT) [ 1 , 5 ], which enable visualization of af- fected brain re gions and discrimination between stroke sub- types, thereby supporting timely therapeutic decisions [ 1 ]. Howe ver , image interpretation remains complex and highly time-sensiti ve [ 7 ]. Manual analysis demands expert radiologists and often in volv es labor-intensi ve procedures to identify subtle lesions or distinguish pathological pat- terns [ 1 ], making it susceptible to human error and inter- observer variability [ 3 ]. Also, high operational costs and the need for specialized personnel can restrict timely diag- nosis in resource-limited environments [ 6 , 11 ]. These chal- lenges ha ve driv en increasing interest in computational ap- proaches, particularly Artificial Intelligence (AI), to support and enhance clinical decision-making in strok e assessment. AI methods, particularly those based on computer vision, enable automated analysis of medical images by learning discriminativ e visual patterns from lar ge annotated datasets [ 9 , 22 ]. In stroke diagnosis, such models can identify sub- tle abnormalities in MRI or CT scans with minimal human intervention, supporting more consistent and efficient as- sessment. Consequently , AI-based solutions ha ve gained increasing adoption in clinical research and practice [ 10 ]. Giv en the critical nature of stroke assessment, effecti ve methods must combine high diagnostic accuracy with com- putational ef ficiency [ 29 ]. Misclassification of stroke sub- types can lead to inappropriate treatment decisions, increas- ing the risk of irreversible brain damage or death [ 6 ], while excessi v e computational cost may limit deployment in time- critical clinical settings [ 2 ]. Accordingly , balancing reliable predictions with fast inference is essential. Based on the aforementioned, we present StrokeNeXt (Fig. 1 ), a Deep Learning (DL) approach for brain stroke classification based on a dual-branch feature extraction ar - chitecture. The two branches process the same input image independently , enabling the capture of complementary rep- resentations and mitigating information loss typically asso- ciated with single-path pipelines. The resulting features are fused through a lightweight decoding module designed to enhance feature integration and generate the final classifi- cation. StrokeNeXt is ev aluated on a real-world CT dataset, where it demonstrates competitiv e performance. The main contributions of this w ork are summarized as follo ws: • W e present StrokeNeXt, an approach for brain stroke Figure 1. Ov ervie w of the proposed StrokeNeXt architecture. The model processes the input CT image through two parallel Con- vNeXt encoders. Their outputs are fused via a custom con volu- tional decoder composed of a stack operation, a 1D conv olutional fusion path with transformation layers, and a classification head. classification that integrates dual-branch feature extrac- tion and a lightweight decoder . • W e demonstrate that StrokeNeXt achiev es high accuracy in distinguishing between stroke and normal cases using real-world CT scan data. • W e show that the model also performs effecti vely in dif- ferentiating among various strok e subtypes. • W e show that StrokeNeXt performs notably when com- pared to other state-of-the-art models from the literature. • W e establish a baseline architecture and performance benchmark that can serve as reference for future research. 2. Related work Stroke identification has been widely explored using div erse paradigms. Early approaches relied on traditional classi- fiers such as Random Forest [ 4 ] and ElasticNet [ 8 ], which achiev ed moderate success in stroke detection b ut were lim- ited by their dependence on handcrafted features and lack of spatial context. Subsequent efforts shifted toward DL, where CNN-based models such as MobileNetV2 [ 29 ] im- prov ed automation and spatial feature extraction but still exhibited limited representational capacity and rob ustness. Other architectures ha ve been proposed to enhance fea- ture representation. Models such as P-CNN [ 11 ], D-UNet [ 29 ], and OzNet [ 20 ] improve lesion localization through multi-scale encoding or task-specific optimizations. In par- allel, 3D approaches, including 3D-CNN [ 19 ] and CAD systems [ 28 ], exploit volumetric information to capture richer spatial context. Ho we ver , these methods typically rely on computationally expensi ve 3D pipelines, increasing complexity and o verfitting risk, limiting their suitability for real-time clinical deployment. Recent work has explored hybrid architectures to model long-range dependencies. For example, StrokeV iT [ 21 ] combines conv olutional networks with V ision T ransform- ers to enhance feature representation and prediction accu- racy . Ho we v er , T ransformer-based models generally de- mand large datasets, high computational cost, and long training times, limiting their practicality in resource- and time-constrained clinical settings. Overall, existing methods exhibit a trade-off between accuracy , ef ficiency , and reliability . While some achie v e strong classification performance, they often incur high computational cost, whereas others lack proper calibration for time-sensiti v e clinical deployment. Addressing these limitations, StrokeNeXt proposes a dual-branch con v olu- tional frame work that strengthens feature di versity through parallel processing while maintaining efficienc y . Its design focuses on achieving rob ust diagnostic performance without relying on large data v olumes or computationally expensi v e inference pipelines, positioning it as a balanced alternati ve within the current landscape of stroke analysis methods. 3. Methodology 3.1. Dataset and prepr ocessing W e use the Stroke Dataset released as part of the Artifi- cial Intelligence in Healthcare Competition (TEKNOFEST 2021) [ 15 ]. It consists of 6,774 brain CT cross-sectional images in PNG format, annotated and curated by a team of sev en radiologis ts. The dataset is di vided into three classes: 4,551 non-stroke, 1,093 hemorrhagic stroke, and 1,130 is- chemic stroke cases. The dataset is publicly available, and representativ e samples are sho wn in Fig. 2 . T o structure the experiments, the dataset was reorganized into two settings. The first addresses stroke detection by grouping ischemic and hemorrhagic cases into a single pos- itiv e class and using non-stroke samples as ne gati ves. The second focuses on stroke subtype classification, considering only ischemic and hemorrhagic cases and excluding non- stroke images. In both settings, the data were randomly split into training, v alidation, and test sets following an 80-10-10 ratio. Data augmentation was used, including random hor- izontal flips, rotations up to 10°, and brightness, contrast, Figure 2. Sample images from the used dataset in this work. and saturation jittering. Images were resized to 224 × 224 and normalized using standard ImageNet statistics. 3.2. Model design 3.2.1. Siamese encoder StrokeNeXt adopts a dual-branch feature extraction mod- ule in which two identical encoders process the same in- put image in parallel. The core hypothesis is that indepen- dent branches can explore dif ferent abstraction paths de- spite sharing the same input, since their parameters are not shared and optimization driv es them toward distinct repre- sentations. As each branch follo ws its o wn stochastic train- ing trajectory , the encoders naturally specialize in dif fer - ent visual cues, increasing representational di v ersity . This results in complementary features being passed to the de- coder , improving feature quality compared to single-branch extractors. Each encoder branch is based on Con vNeXt [ 18 ], a CNN architecture inspired by Transformer design princi- ples. ConvNeXt is built on the hypothesis that fully con v o- lutional models can match or surpass Transformer perfor- mance without incurring the high computational and data demands typically associated with it. Con vNeXt has been validated across multiple benchmarks, moti v ating its use as the backbone of the proposed dual-branch encoder , where it provides strong representational capacity while preserving computational efficienc y for stroke classification. Con vNeXt builds upon a ResNet50 through ke y modi- fications, including larger kernel sizes (7 × 7), GELU acti- vations, layer normalization, and a stage-wise design, en- abling effecti v e hierarchical representation learning with controlled computational cost. Architecturally , it comprises four sequential stages of conv olutional blocks followed by pooling and a classification head, as illustrated in Fig. 3 . In our implementation, the final pooling and classifier layers are removed, and only the feature extraction stages are re- tained to serve as the encoders. Con vNeXt is av ailable in tiny , small, base, and lar ge variants, which dif fer primarily in network depth and width, namely the number of channels and blocks per stage, as summarized in T able 1 . Figure 3. Con vNeXt architecture (tin y version). Source: [ 23 ] 3.2.2. Fusion decoder A custom fusion module (Fig. 4 ) is designed to combine the outputs of the two encoder branches into a unified repre- sentation for classification. The module applies a sequence T able 1. Details of Con vNeXt v ariants. T aken from: [ 24 ] Model Channels (per stage) Depths (per stage) Params (M) FLOPs (G) Con vNeXt-tiny [96, 192, 384, 768] [3, 3, 9, 3] 28 4.5 Con vNeXt-small [96, 192, 384, 768] [3, 3, 27, 3] 50 8.7 Con vNeXt-base [128, 256, 512, 1024] [3, 3, 27, 3] 89 15.4 Con vNeXt-large [192, 384, 768, 1536] [3, 3, 27, 3] 198 34.4 of 1D con v olutional layers along a synthetic sequence di- mension of length two, obtained by stacking the encoder outputs. An initial con v olution merges the paired feature vectors by learning local interactions between them, fol- lowed by a bottleneck formed by two pointwise (kernel size 1) con v olutions. The first projects the features to a config- urable hidden dimension, while the second restores them to the original size. More in depth, fusion is performed over a synthetic sequence obtained by stacking the encoder embeddings, [ B , C ] → [ B , C, 2] . A single 1D con v olution with k ernel size k = 2 collapses this axis to length one. This operation implements a learned, channel-mixing combination of the two branch embeddings, capturing cross-branch and cross- channel interactions rather than a simple per -channel sum. BatchNorm and GELU stabilize and gate this mer ge, yield- ing a fused vector in [ B , C ] . Figure 4. Structure of fusion decoder de veloped in this study . After fusion, a bottleneck transformation refines the rep- resentation using two pointwise conv olutions with an op- tional hidden width H ( C → H → C ) , each followed by normalization and GELU, with dropout applied between layers for regularization. These pointwise operations act as linear projections on the fused vector , where the bottleneck width H provides a direct mechanism to control model ca- pacity and FLOPs without modifying the encoders. Since the initial k = 2 merge already collapses the paired features [ f 1 , f 2 ] back to width C , all subsequent transformations op- erate at C or at most H , preserving low latency . This design offers concrete adv antages over common alternati ves. • Concatenation + MLP: Concatenating [ f 1 , f 2 ] doubles the input width to 2 C ; any do wnstream MLP must first process that larger vector . By merging with a k = 2 conv , we reduce to C before the non-linear block, so compara- ble expressi ve po wer is achieved with fewer parameters and lower memory traf fic. • Element-wise sum: Summation has zero parameters but cannot learn cross-channel reweighting or asymmetric branch contributions. The k = 2 conv learns both pos- itiv e/ne gati ve mixing and cross-channel couplings, so it can emphasize complementary cues from each branch rather than av eraging them aw ay . • Attention/gated fusion: W ith only two ‘tokens, ’ self- attention de generates to learning a 2 × 2 mixing plus pro- jections, yet still incurs Q/K/V projections, softmax, and extra acti v ations. The proposed k = 2 con v realizes an equiv alent linear mixing of the two vectors, and the sub- sequent pointwise + GELU stack supplies non-linearity , achieving similar effect at a fraction of the ov erhead and with deterministic latency . T wo additional properties are purposeful. First, the fusion block is order-a ware, as a fixed stacking order is preserved, allowing the model to treat the two encoder branches differently if they specialize during training. Sec- ond, the bottleneck/dropout/BN sequence improves opti- mization stability and calibration without incurring a no- ticeable speed penalty . 3.3. T raining and implementation details The models were implemented in PyT orch and trained with mixed-precision using Automatic Mixed Precision (AMP). Optimization was performed with AdamW (learning rate 1 × 10 − 4 , weight decay 1 × 10 − 5 ) and a ReduceLR On- Plateau scheduler . All models were trained for 20 epochs with a batch size of 80 using CrossEntropy loss with label smoothing (0.1). T o ensure fair comparison, StrokeNeXt and all baselines follo wed the same training protocol, and all Con vNeXt v ariants used their def ault torchvision hy- perparameters 1 . CT images were processed as 3-channel inputs, and all experiments were conducted on a single NVIDIA A100 GPU with 40 GB. Model ev aluation used a comprehensi ve set of com- plementary metrics. Classification performance was as- sessed using Accuracy , Precision, Recall, F1-score, A U- R OC, A UPRC, and Balanced Accuracy . Reliability and error structure were analyzed through Matthe ws Correla- tion Coefficient (MCC), Brier Score, and Expected Calibra- tion Error (ECE). Computational efficiency was e valuated via training time, inference latency , throughput, peak GPU memory usage, and FLOPs. In addition, per-class sensi- tivity , specificity , and support were reported, and McNe- mar’ s paired tests were applied to assess the statistical sig- nificance of performance dif ferences. Implementation of all other metrics employed follo w standard formulations. 1 https : / / docs . pytorch . org / vision / main / models / convnext.html 4. Results and Discussion 4.1. Classification of strok e presence T able 2 sho ws the performance of StrokeNeXt v ariants on stroke presence classification. All configurations achiev e consistently strong results, with accuracy and F1-scores e x- ceeding 97%. Increasing encoder size leads to systematic improv ements across all metrics, indicating that lar ger Con- vNeXt backbones yield more robust feature representations. This trend is further reflected in A UR OC and A UPRC v al- ues approaching perfect discrimination ( > 0 . 99 ) for the base and large v ariants, demonstrating reliable ranking per - formance under class imbalance. Balanced accuracy closely matches ov erall accuracy , and the MCC reaches 0.97 for StrokeNeXt-lar ge, confirming strong predictiv e agreement in imbalanced settings. Calibration quality remains stable across v ariants, with low Brier scores and small ECE val- ues, indicating well-aligned probabilistic outputs. Regarding efficiency , StrokeNeXt-tin y exhibits the low- est computational cost, with < 9 GFLOPs, 58M parameters, and a peak memory usage of 1.4 GB, resulting in a latency of 2 ms per image and a throughput of 570 images per sec- ond. As encoder size gro ws, computational and memory demands increase. StrokeNeXt-large reaches nearly 400M parameters and 69 GFLOPs, with latency rising to 9 ms and throughput dropping to 113 images per second. Despite this, training time remains modest across all v ariants, with the largest model completing within 15 minutes on the tar- get hardware. This shows that while larger backbones pro- vide incremental accurac y gains, the smaller StrokeNeXt variants already achieve a fav orable balance between per- formance and efficienc y . T able 3 compares StrokeNeXt with representati ve CNN and T ransformer baselines, considering both the best- performing and the lightest variant. Con ventional CNNs, including MobileNetV2, VGG16, and ResNet v ariants, reach accuracies in the 86-89% range with MCC values of 0.65-0.75, indicating frequent misclassifications. Swin T ransformer sho ws improv ed calibration and discrimination but remains below 90% accuracy . In contrast, StrokeNeXt- tiny achie v es 97.8% accurac y and an MCC of 0.95, while StrokeNeXt-lar ge reaches 98.7% accuracy and 0.97 MCC. Both variants attain near -perfect A UROC and A UPRC ( > 0 . 98 ), Brier scores < 0 . 03 , and ECE values ≈ 0.05, demon- strating not only superior classification performance but also better-calibrated probability estimates. In terms of efficienc y , lightweight CNNs such as Mo- bileNetV2 and ResNet50 achie ve the highest throughput ( > 2000 img/s) with sub-GB memory usage, b ut their lo wer accuracy limits competitiveness. VGG16 and ResNet152 retain fast inference but require substantially higher mem- ory ( > 4 GB and ∼ 1.2 GB, respectiv ely), reducing their suit- ability in constrained environments. StrokeNeXt-tin y , de- T able 2. Performance of the dif ferent StrokeNeXt v ariants on stroke presence classification (non-strok e vs. stroke). Method Accuracy Precision Recall F1-score AUR OC AUPRC Balanced Acc. MCC Brier score ECE Latency (s) Throughput (img/s) Peak GPU (GB) FLOPs (G) Params (M) T rain. time (h) StrokeNeXt-tiny 0.978 0.978 0.978 0.978 0.986 0.987 0.971 0.950 0.021 0.053 0.002 570 1.417 8.977 57.6 0.087 StrokeNeXt-small 0.980 0.981 0.981 0.980 0.976 0.981 0.972 0.957 0.019 0.047 0.003 349 1.747 17.474 100.8 0.126 StrokeNeXt-base 0.982 0.983 0.982 0.982 0.990 0.990 0.977 0.960 0.017 0.051 0.004 222 2.641 30.853 178.5 0.161 StrokeNeXt-large 0.987 0.987 0.987 0.987 0.995 0.995 0.983 0.970 0.013 0.059 0.009 113 4.950 68.940 399.9 0.245 T able 3. Performance of StrokeNeXt on strok e presence classification (non-stroke vs. stroke) compared with other methods. Method Accuracy Precision Recall F1-score A UROC AUPRC Balanced Acc. MCC Brier score ECE Latency (s) Throughput (img/s) Peak GPU (GB) FLOPs (G) Params (M) T rain. time (h) MobileNetv2 [ 26 ] 0.865 0.863 0.865 0.862 0.938 0.894 0.828 0.685 0.121 0.154 0.001 3374 0.868 0.319 2.2 0.040 VGG16 [ 16 ] 0.894 0.893 0.894 0.893 0.918 0.858 0.822 0.657 0.142 0.180 0.001 1750 4.020 15.519 134.2 0.041 ResNet50 [ 12 ] 0.875 0.879 0.875 0.870 0.926 0.893 0.827 0.711 0.125 0.164 0.001 2057 1.034 4.130 23.5 0.040 ResNet152 [ 12 ] 0.892 0.894 0.892 0.889 0.937 0.908 0.855 0.752 0.108 0.152 0.001 1213 1.298 11.601 58.2 0.046 Swin T ransformer [ 17 ] 0.893 0.893 0.892 0.890 0.939 0.899 0.859 0.751 0.094 0.099 0.002 363 2.196 10.550 87.1 0.067 ConvNeXt-base [ 18 ] 0.879 0.878 0.879 0.879 0.937 0.891 0.857 0.723 0.123 0.174 0.002 444 1.947 15.425 87.5 0.049 StrokeNeXt-tiny 0.978 0.978 0.978 0.978 0.986 0.987 0.971 0.950 0.021 0.053 0.002 570 1.417 8.977 57.6 0.087 StrokeNeXt-large 0.987 0.987 0.987 0.987 0.995 0.995 0.983 0.970 0.013 0.059 0.009 113 4.950 68.940 399.9 0.245 The best r esults ar e in bold, the second best ar e underlined. Note that we have selected the best and lightest Str okeNeXt model for comparison. spite its dual-branch design, requires only 57.6M parame- ters and 8.98 GFLOPs, with 1.4 GB peak memory and 570 img/s throughput. Compared to Con vNeXt-base, VGG16, and Swin, StrokeNeXt-tiny consistently uses fewer param- eters, less memory , and achiev es higher throughput at com- parable latency . Although training time increases slightly , it remains practical ( < 0 . 1 h). Con versely , StrokeNeXt-large maximizes accuracy at a substantially higher cost (400M parameters, ∼ 5 GB memory , 113 img/s). Overall, the Stro- keNeXt f amily of fers a tunable trade-of f, with the tin y v ari- ant enabling real-time deployment and the large variant fa- voring peak performance when resources permit. T able 4 reports McNemar test results comparing StrokeNeXt-lar ge (our best model) with the baselines. In all cases, p-v alues fall well below the α = 0 . 05 threshold, indicating that the observed performance gains are statis- tically significant rather than random. StrokeNeXt consis- tently corrects a substantially larger number of cases mis- classified by competing models, with at least 67 additional correct predictions across comparisons, while the in verse scenario occurs only rarely . The corresponding χ 2 values further show that these improvements are systematic rather than marginal. T able 4. Results of the McNemar test comparing our best model with other methods on stroke presence classification (non-stroke vs. stroke). Method A Method B A ✓ B ✗ B ✓ A ✗ χ 2 p-value StrokeNeXt-large VGG16 [ 16 ] 94 2 82.260 < 0.0001 StrokeNeXt-large MobileNetv2 [ 26 ] 87 4 73.890 < 0.0001 StrokeNeXt-large ResNet50 [ 12 ] 79 3 68.597 < 0.0001 StrokeNeXt-large ResNet152 [ 12 ] 67 3 56.700 < 0.0001 StrokeNeXt-large ConvNeXt-base [ 18 ] 76 3 65.620 < 0.0001 StrokeNeXt-large Swin Transformer [ 17 ] 69 5 53.635 < 0.0001 A ✓ B ✗ indicates the number of samples correctly classified by method A but misclassi- fied by method B, while B ✓ A ✗ indicates the opposite. Statistical significance is assessed at α = 0 . 05 . T able 5 reports class-wise sensitivity and specificity . Baseline methods e xhibit clear imbalances, since ResNet50 and ResNet152 achiev e high non-stroke sensitivity ( > 0 . 96 ) at the expense of low specificity ( < 0 . 75 ), reflecting a ten- dency to ov er -predict the non-stroke class. MobileNetV2 and VGG16 show weaker discrimination, with sensitivity and specificity falling below 0.94 in one or both classes. In contrast, StrokeNeXt maintains consistently balanced performance across categories. StrokeNeXt-large achiev es the highest ov erall values, with sensitivity and specificity > 0 . 99 for both classes, indicating near-perfect separation, while StrokeNeXt-tiny sustains values around 0.95, out- performing heavier baselines such as Con vNeXt-base and Swin. Unlike models that fav or one metric at the expense of the other , Strok eNeXt preserves symmetry , avoiding the trade-offs observ ed in ResNet and V GG architectures. T able 5. Per-class performance of StrokeNeXt on stroke presence classification (non-stroke vs. stroke) compared to other methods. Method Non-stroke Stroke Sensitivity Specificity Sensitivity Specificity MobileNetv2 [ 26 ] 0.934 0.722 0.722 0.934 VGG16 [ 16 ] 0.908 0.735 0.735 0.908 ResNet50 [ 12 ] 0.967 0.686 0.686 0.967 ResNet152 [ 12 ] 0.965 0.744 0.744 0.965 Swin T ransformer [ 17 ] 0.956 0.762 0.762 0.956 ConvNeXt-base [ 18 ] 0.921 0.794 0.794 0.921 StrokeNeXt-tiny 0.991 0.951 0.951 0.991 StrokeNeXt-small 0.998 0.946 0.946 0.998 StrokeNeXt-base 0.993 0.960 0.960 0.993 StrokeNeXt-large 0.993 0.973 0.973 0.993 Fig. 5 presents the confusion matrices for StrokeNeXt and the baseline models. StrokeNeXt exhibits near-perfect class separation, with only a small number of misclassifi- cations in either category , consistent with its high MCC. In contrast, baselines such as ResNet50, VGG16, and Mo- bileNetV2 sho w substantially higher false ne gati ves in the stroke class, a particularly critical failure mode in clinical settings. Even stronger backbones, including ConvNeXt- base, Swin T ransformer , and ResNet152, display greater imbalance between false positi ves and false negati v es than Figure 5. Confusion matrices comparison between StrokeNeXt and other models on strok e presence classification (non-stroke vs. stroke). StrokeNeXt. T able 6 compares StrokeNeXt against prior methods for stroke presence classification. StrokeNeXt-large achiev es the best overall performance, reaching an F1-score of 0.987, slightly surpassing OzNet (0.984) and clearly outperform- ing earlier CNN approaches (0.944), StrokeV iT (0.870), and classical methods such as ElasticNet (0.750) and CAD systems (0.730). The lightweight StrokeNeXt-tin y also at- tains an F1-score of 0.978, exceeding MobileNetV2 (0.974) and approaching the strongest models. This indicates that StrokeNeXt scales effecti v ely , with lar ger variants advanc- ing the state-of-the-art while smaller v ariants retain compet- itiv e accurac y at substantially lo wer computational cost. T able 6. Comparison of StrokeNeXt on stroke presence classifica- tion (non-stroke vs. stroke) with other methods from the literature. Method Accuracy Precision Recall F1-score StrokeV iT [ 21 ] 0.870 0.870 0.870 0.870 OzNet [ 20 ] 0.984 0.983 0.986 0.984 CNN [ 14 ] 0.951 0.953 0.936 0.944 ElasticNet [ 8 ] 0.760 0.790 0.710 0.750 CAD [ 28 ] 0.810 0.760 0.820 0.730 MobileNetv2 [ 29 ] 0.975 0.977 0.972 0.974 StrokeNeXt-tiny 0.978 0.978 0.978 0.978 StrokeNeXt-large 0.987 0.987 0.987 0.987 The best r esults ar e in bold, the second best ar e underlined. 4.2. Classification of strok e subtypes T urning to stroke subtype classification, T able 7 reports the performance of the StrokeNeXt variants. All models op- erate close to ceiling, with accuracy and F1-scores in the 0.986-0.988 range and A UROC/A UPRC values between 0.999 and 1.000. Balanced accurac y closely matches over - all accurac y , and MCC remains stable at ≈ 0.973, show- ing consistent behavior across stroke types. Calibration metrics remain lo w , with small Brier scores and ECE v al- ues, indicating well-calibrated probability estimates rather than ov erconfident predictions. Recall values around 0.985- 0.988 imply very few false negati ves for either subtype, which is clinically relev ant gi ven the div ergence in treat- ment strategies. Regarding ef ficiency , StrokeNeXt-tin y remains the most efficient configuration, with 0.002 s latency , 571 img/s throughput, ≈ 58M parameters, and < 1.5 GB peak GPU memory . In contrast, StrokeNeXt-large incurs higher cost, reaching 0.009 s latency , 114 img/s throughput, ≈ 400M pa- rameters, and a ∼ 5 GB memory footprint. T raining time in- creases with model capacity but remains < 0.15 h per epoch across variants. Gi v en the marginal performance differ- ences, smaller configurations such as tiny and small offer particularly f a vorable trade-offs, deli vering reliable subtype classification while remaining suitable for deployment. T able 8 compares StrokeNeXt with other models for stroke subtype classification. Conv entional single-branch architectures, (MobileNetV2, ResNet50, and Swin Trans- former) achie v e accurac y < 0 . 86 with MCC v alues < 0 . 75 , indicating limited reliability for clinical use. In contrast, StrokeNeXt-tiny and StrokeNeXt-base reach near-ceiling performance, with A UROC/A UPRC of 1.0, balanced ac- curacy of 0.986, and MCC of 0.973, reflecting rob ust dis- crimination across stroke subtypes. Calibration quality is also improv ed, with a low Brier score (0.011) and ECE < 0 . 06 . Giv en the importance of correctly distinguishing stroke subtypes for treatment selection, these gains are clin- ically meaningful. Notably , StrokeNeXt-tiny matches the diagnostic reliability of StrokeNeXt-base while requiring fewer computational resources. In terms of ef ficiency , lightweight baselines such as Mo- bileNetV2 and ResNet50 achie ve the lo west FLOPs, param- eter counts, and memory usage, yielding high throughput (e.g., > 1900 img/s for MobileNetV2), b ut with reduced predictiv e reliability . At higher capacity , StrokeNeXt-base requires 178.5M parameters and 30.8 GFLOPs, yet main- T able 7. Performance of the dif ferent StrokeNeXt v ariants on stroke type classification (ischemia vs. hemorrhage). Method Accuracy Precision Recall F1-score AUR OC AUPRC Balanced Acc. MCC Brier score ECE Latency (s) Throughput (img/s) Peak GPU (GB) FLOPs (G) Params (M) T rain. time (h) StrokeNeXt-tiny 0.986 0.985 0.986 0.986 1.000 1.000 0.986 0.973 0.011 0.058 0.002 571 1.417 8.977 57.6 0.041 StrokeNeXt-small 0.987 0.987 0.987 0.987 0.999 0.999 0.986 0.973 0.013 0.049 0.003 349 1.747 17.474 100.8 0.096 StrokeNeXt-base 0.988 0.988 0.988 0.988 1.000 1.000 0.986 0.973 0.011 0.054 0.005 219 2.641 30.853 178.5 0.101 StrokeNeXt-large 0.987 0.987 0.985 0.986 0.999 0.999 0.986 0.973 0.015 0.061 0.009 114 4.950 68.940 399.9 0.122 T able 8. Performance of StrokeNeXt on strok e type classification (ischemia vs. hemorrhage) compared with other methods. Method Accuracy Precision Recall F1-score A UROC AUPRC Balanced Acc. MCC Brier score ECE Latency (s) Throughput (img/s) Peak GPU (GB) FLOPs (G) Params (M) T rain. time (h) MobileNetv2 [ 26 ] 0.812 0.813 0.811 0.811 0.919 0.919 0.811 0.624 0.122 0.083 0.001 2341 0.868 0.319 2.2 0.020 VGG16 [ 16 ] 0.879 0.879 0.879 0.879 0.951 0.956 0.879 0.758 0.104 0.124 0.001 1434 4.020 15.519 134.3 0.020 ResNet50 [ 12 ] 0.852 0.854 0.852 0.852 0.918 0.924 0.851 0.706 0.133 0.136 0.001 1900 1.034 4.130 23.5 0.024 ResNet152 [ 12 ] 0.865 0.873 0.865 0.865 0.949 0.936 0.864 0.738 0.101 0.130 0.001 1217 1.298 11.601 58.2 0.025 Swin T ransformer [ 17 ] 0.825 0.825 0.825 0.825 0.905 0.911 0.825 0.650 0.132 0.084 0.003 335 2.196 10.550 87.1 0.029 ConvNeXt-base [ 18 ] 0.848 0.848 0.848 0.848 0.926 0.930 0.848 0.695 0.130 0.142 0.002 441 1.947 15.425 87.6 0.026 StrokeNeXt-tiny 0.986 0.985 0.986 0.986 1.000 1.000 0.986 0.973 0.011 0.058 0.002 571 1.417 8.977 57.6 0.041 StrokeNeXt-base 0.988 0.988 0.988 0.988 1.000 1.000 0.986 0.973 0.011 0.054 0.005 219 2.641 30.853 178.5 0.101 The best r esults ar e in bold, the second best ar e underlined. Note that we have selected the best and lightest Str okeNeXt model for comparison. tains competitiv e latency (0.005 s) and throughput (219 img/s). StrokeNeXt-tin y represents an effecti v e middle ground, with 57.6M parameters and 8.98 GFLOPs, sustain- ing 571 img/s and 0.002 s latency while outperforming all baselines in accuracy . T able 9 reports the McNemar test results comparing StrokeNeXt with baseline methods for stroke subtype clas- sification. In all comparisons, p-v alues fall well below the 0.05 threshold, confirming that the observed gains are statis- tically significant. StrokeNeXt consistently corrects a larger number of errors made by competing models, while the op- posite cases remain minimal. This indicates not only higher ov erall accurac y , but also more reliable discrimination be- tween ischemia and hemorrhage, a distinction with direct clinical impact due to differing treatment strate gies. T able 9. Results of the McNemar test comparing our best model with other methods on stroke type classification (ischemia vs. hemorrhage). Method A Method B A ✓ B ✗ B ✓ A ✗ χ 2 p-value StrokeNeXt-base V GG16 [ 16 ] 25 1 20.346 < 0.0001 StrokeNeXt-base MobileNetv2 [ 26 ] 40 1 35.219 < 0.0001 StrokeNeXt-base ResNet50 [ 12 ] 31 1 26.281 < 0.0001 StrokeNeXt-base ResNet152 [ 12 ] 29 2 21.807 < 0.0003 StrokeNeXt-base Con vNeXt-base [ 18 ] 32 1 27.272 < 0.0001 StrokeNeXt-base Swin Transformer [ 17 ] 37 1 32.236 < 0.0001 A ✓ B ✗ indicates the number of samples correctly classified by method A but misclassi- fied by method B, while B ✓ A ✗ indicates the opposite. Statistical significance is assessed at α = 0 . 05 . T able 10 reports per-class performance for ischemic and hemorrhagic stroke classification. Conv entional models show moderate results, with ischemia sensitivity often be- low 0.90, a concerning limitation giv en the clinical risk of missed ischemic cases. In contrast, StrokeNeXt vari- ants achieve higher sensitivity ( > 0 . 97 ) while maintaining strong specificity , indicating balanced discrimination across both subtypes. The high ischemia sensitivity reflects im- prov ed reliability in identifying the more frequent and diag- nostically challenging class, while the comparable perfor- mance for hemorrhage confirms cross-category rob ustness. T able 10. Per-class performance of StrokeNeXt on troke type clas- sification (ischemia vs. hemorrhage) compared to other methods. Method Ischemia Hemorrhage Sensitivity Specificity Sensitivity Specificity MobileNetv2 [ 26 ] 0.773 0.850 0.850 0.772 VGG16 [ 16 ] 0.873 0.885 0.885 0.873 ResNet50 [ 12 ] 0.809 0.894 0.894 0.809 ResNet152 [ 12 ] 0.790 0.938 0.938 0.790 Swin T ransformer [ 17 ] 0.818 0.832 0.832 0.818 ConvNeXt-base [ 18 ] 0.846 0.850 0.850 0.846 StrokeNeXt-tiny 0.972 0.999 0.999 0.972 StrokeNeXt-small 0.982 0.991 0.991 0.982 StrokeNeXt-base 0.973 0.999 0.999 0.973 StrokeNeXt-large 0.982 0.991 0.991 0.982 Fig. 6 presents the confusion matrices for stroke sub- type classification. StrokeNeXt-base shows near-perfect separation between subtypes, with only a fe w misclassi- fications, in contrast to the higher error rates observed in ResNet, Swin T ransformer , and MobileNetV2. Notably , StrokeNeXt is the only model correctly identifying more than 100 ischemic cases, while all others fall belo w this threshold. Concurrently , it achiev es perfect hemorrhage recognition, a voiding false negati v es that could critically af- fect clinical decisions. This shows that StrokeNeXt not only improv es overall accurac y but also minimizes subtype con- fusion, a key requirement in settings where treatment de- pends on precise stroke categorization. Higher MCC values further support its robustness under balanced e v aluation. T able 11 compares StrokeNeXt against prior methods for ischemic vs. hemorrhagic classification. StrokeNeXt-base achiev es the highest numbers, with an F1-score of 0.988, outperforming CNN approaches such as D-UNet (0.985) and P-CNN (0.983). Notably , StrokeNeXt-tiny matches or exceeds these baselines with an F1-score of 0.986 at a lower computational cost. In contrast, Random F orest and Enhanced-CNN remain below 0.96, and the 3D-CNN reaches only 0.880 despite using volumetric information. Figure 6. Confusion matrices comparison between StrokeNeXt and other models on strok e type classification (ischemia vs. hemorrhage). T able 11. Comparison of StrokeNeXt on stroke type classification (ischemia vs. hemorrhage) with other methods from the literature. Method Accuracy Precision Recall F1-score P-CNN-W [ 11 ] 0.975 0.975 0.975 0.975 Random Forest [ 4 ] 0.959 0.944 0.961 0.954 Enhanced-CNN [ 27 ] 0.952 0.949 0.972 0.960 P-CNN [ 11 ] 0.983 0.983 0.983 0.983 3D-CNN [ 19 ] 0.920 0.940 0.840 0.880 D-UNet [ 29 ] 0.985 0.986 0.985 0.985 StrokeNeXt-tiny 0.986 0.985 0.986 0.986 StrokeNeXt-base 0.988 0.988 0.988 0.988 The best r esults ar e in bold, the second best ar e underlined. Overall, the results sho w that the proposed StrokeNeXt achiev es higher accuracy and statistical reliability than competing approaches, with improvements consistently supported by significance testing. It maintains strong cal- ibration alongside high predictive performance, ensuring outputs that are both discriminativ e and trustworthy for clin- ical decision support. These properties hold across both stroke presence detection and subtype classification, indi- cating that the architecture generalizes ef fecti vely to dif- ferent diagnostic tasks. Importantly , the ef ficiency analysis demonstrates that these gains do not compromise usability , as even lightweight StrokeNeXt variants outperform con- ventional baselines while offering fa vorable trade-of fs be- tween accuracy and computational cost. This flexibility en- ables deployment across a wide range of resource settings, positioning StrokeNeXt as a reliable and efficient alterna- tiv e to e xisting methods for CT -based stroke classification. 5. Limitations This study has limitations that should be acknowledged. First, the dataset does not include patient-lev el information such as demographics or clinical outcomes. As StrokeNeXt is purely image-based, the absence of these variables lim- its the analysis of how model predictions may generalize across different patient populations or interact with clinical variables. In addition, the ev aluation is restricted to CT im- agery , and the reported efficiency results are specific to the hardware configuration used in this work. While these re- sults provide a realistic estimate, validation on larger and more di verse cohorts and on different hardware would be necessary to fully assess robustness and generalizability . in real-world clinical settings. 6. Conclusions W e present StrokeNeXt for brain stroke classification from CT imagery . It is a dual-branch architecture with two identi- cal Con vNeXt encoders operating in parallel on the same in- put. The extracted features are fused through a lightweight con v olutional decoder and processed by a compact classi- fier , aiming to enhance representational capacity while pre- serving computational efficiency . The approach is ev aluated on a curated dataset of 6,774 annotated CT images under two classification scenarios: stroke presence detection and stroke subtype classification (ischemic vs. hemorrhagic). Across both tasks, StrokeNeXt consistently outper- forms conv olutional, T ransformer -based, and state-of-the- art methods. StrokeNeXt achie ves F1-scores of up to 0.988, while maintaining consistently high class-wise sensitivity and specificity , low calibration error , and strong Matthews correlation coefficients. Statistical significance testing fur- ther indicates that the observed improvements are not due to random variation. Confusion matrix analysis also demon- strates reliable discrimination of challenging cases. The model maintains fast inference and practical training times, demonstrating that the approach design effecti vely balances diagnostic performance and practical deployability . References [1] Hossein Abbasi, Maysam Orouskhani, Samaneh Asgari, and Sara Shomal Zadeh. Automatic brain ischemic stroke seg- mentation with deep learning: A revie w . Neur osci. Inform. , 3(4):100145, 2023. 1 [2] Mohamed A. Abdou. Literature revie w: efficient deep neu- ral networks techniques for medical image analysis. Neural Comput. Appl. , 34:5791–5812, 2022. 1 [3] Priya Aggarwal, Narendra Kumar Mishra, Binish Fatimah, Pushpendra Singh, Anubha Gupta, and Shiv Dutt Joshi. Covid-19 image classification using deep learning: Ad- vances, challenges and opportunities. Comput. Biol. Med. , 144:105350, 2022. 1 [4] T essy Badriyah, Nur Sakinah, Iwan Syarif, and Daisy Rah- mania Syarif. Machine learning algorithm for stroke disease classification. In ICECCE , 2020. 2 , 8 [5] Heitor Cabral Frade, Susan E. W ilson, Anne Beckwith, and W illiam J. Powers. Comparison of outcomes of ischemic stroke initially imaged with cranial computed tomography alone vs computed tomography plus magnetic resonance imaging. J AMA Network Open , 5(7):e2219416, 2022. 1 [6] T ongan Cai, Haomiao Ni, Mingli Y u, Xiaolei Huang, Kelvin W ong, John V olpi, James Z. W ang, and Stephen T .C. W ong. Deepstroke: An efficient stroke screening framework for emergenc y rooms with multimodal adversarial deep learn- ing. Med. Imag e Anal. , 80:102522, 2022. 1 [7] Aditi Deshpande, Nima Jamilpour , Bin Jiang, Patrik Michel, Ashraf Eskandari, Chelsea Kidwell, Max Wintermark, and Kav eh Laksari. Automatic segmentation, feature extraction and comparison of healthy and stroke cerebral vasculature. Neur oImag e: Clin. , 30:102573, 2021. 1 [8] Soumyabrata Dev , He wei W ang, Chidozie Shamrock Nwosu, Nishtha Jain, Bharadwaj V eerav alli, and Deepu John. A predictive analytics approach for stroke prediction using machine learning and neural netw orks. Healthc. Anal. , 2:100032, 2022. 2 , 6 [9] Omar Elharrouss, Y ounes Akbari, Noor Almadeed, and So- maya Al-Maadeed. Backbones-revie w: Feature extractor networks for deep learning and deep reinforcement learning approaches in computer vision. Computer Science Review , 53:100645, 2024. 1 [10] Andre Este va, Katherine Chou, Serena Y eung, Nikhil Naik, Ali Madani, Ali Mottaghi, Y un Liu, Eric T opol, Jeff Dean, and Richard Socher. Deep learning-enabled medical com- puter vision. npj Digit. Med. , 4(1), 2021. 1 [11] Anjali Gautam and Balasubramanian Raman. T owards effec- tiv e classification of brain hemorrhagic and ischemic stroke using cnn. Biomed. Signal Pr ocess. Control. , 63:102178, 2021. 1 , 2 , 8 [12] Kaiming He, Xiangyu Zhang, Shaoqing Ren, and Jian Sun. Deep residual learning for image recognition. In CVPR , 2015. 5 , 7 [13] Md. Maruf Hossain, Md. Mahfuz Ahmed, Abdullah Al No- maan Nafi, Md. Rakibul Islam, Md. Shahin Ali, Jahu- rul Haque, Md Sipon Miah, Md Mahbubur Rahman, and Md Khairul Islam. A nov el hybrid vit-lstm model with ex- plainable ai for brain stroke detection and classification in ct images: A case study of rajshahi region. Comput. Biol. Med. , 186:109711, 2025. 1 [14] Buket Kaya and Muhammed Onal. A cnn transfer learning- based approach for segmentationand classification of brain stroke from noncontrast ctimages. Int. J. Imaging Syst. T ech- nol. , 33(4):1335–1352, 2023. 6 [15] Ural K oc, Ebru Akcapinar Sezer , Y asar Alper Ozkaya, Y asin Y arbay , Onur T aydas, V eysel Atilla A yyildiz, Huseyin Alper Kiziloglu, Ugur Kesimal, Imran Cankaya, Muhammed Said Besler , Emrah Karakas, F atih Karademir , Nihat Baris Se- bik, Murat Bahadir, Ozgur Sezer , Batuhan Y esilyurt, Songul V arli, Erhan Akdogan, Mustafa Mahir Ulgu, and Suayip Birinci. Artificial intelligence in healthcare competition (teknofest-2021): Stroke data set. Eurasian J. Med. , 54: 248–258, 2022. 2 [16] Shuying Liu and W eihong Deng. V ery deep con v olutional neural network based image classification using small train- ing sample size. In A CPR , 2015. 5 , 7 [17] Ze Liu, Y utong Lin, Y ue Cao, Han Hu, Y ixuan W ei, Zheng Zhang, Stephen Lin, and Baining Guo. Swin transformer: Hierarchical vision transformer using shifted windows. In ICCV , 2021. 5 , 7 [18] Zhuang Liu, Hanzi Mao, Chao-Y uan W u, Christoph Feicht- enhofer , Tre vor Darrell, and Saining Xie. A convnet for the 2020s. In CVPR , 2022. 3 , 5 , 7 [19] A.S. Neethi, S. Niyas, Santhosh Kumar Kannath, Jimson Mathew , Ajimi Mol Anzar , and Jeny Rajan. Stroke classi- fication from computed tomography scans using 3d con volu- tional neural netw ork. Biomed. Signal Pr ocess. Contr ol. , 76: 103720, 2022. 2 , 8 [20] Oznur Ozaltin, Orhan Coskun, Ozgur Y eniay , and Abdul- hamit Subasi. A deep learning approach for detecting stroke from brain ct images using oznet. Bioengineering , 9(12): 1–16, 2022. 2 , 6 [21] Rishi Raj, Jimson Mathew , Santhosh Kumar Kannath, and Jeny Rajan. Strokevit with automl for brain stroke classifica- tion. Eng. Appl. Artif . Intell. , 119:105772, 2023. 2 , 6 [22] Leo Thomas Ramos and Angel D. Sappa. Multispectral semantic segmentation for land cover classification: An ov erview . IEEE JST ARS , 17:14295–14336, 2024. 1 [23] Leo Thomas Ramos and Angel D. Sappa. Dual-branch con vnext-based network with attentional fusion decoding for land cover classification using multispectral imagery . In SoutheastCon , 2025. 3 [24] Leo Thomas Ramos and Angel D. Sappa. Multi-encoder con vnext network with smooth attentional feature fusion for multispectral semantic segmentation. arXiv preprint arXiv:2602.10137 , 2026. 3 [25] Archana Saini, Kalpna Guleria, and Shagun Sharma. Perfor- mance analysis of machine learning approaches for stroke prediction in healthcare. In INDIA Com , 2023. 1 [26] Mark Sandler , Andre w Howard, Menglong Zhu, Andre y Zh- moginov , and Liang-Chieh Chen. Mobilenetv2: In verted residuals and linear bottlenecks. In CVPR , 2019. 5 , 7 [27] M. Shakunthala and K. HelenPrabha. Classification of is- chemic and hemorrhagic stroke using enhanced-cnn deep learning technique. J. Intell. Fuzzy Syst. , 45(4):6323–6338, 2023. 8 [28] Azhar T ursyno va, Batyrkhan Omarov , Natalya T uk enov a, In- dira Salgozha, Oner gul Khaav al, Rinat Ramazanov , and Bag- dat Ospanov . Deep learning-enabled brain stroke classifica- tion on computed tomography images. Comput. Mater . Con- tin. , 75(1):1431–1446, 2023. 2 , 6 [29] Sercan Y alcın and Huseyin V ural. Brain stroke classification and segmentation using encoder-decoder based deep conv o- lutional neural networks. Comput. Biol. Med. , 149:105941, 2022. 1 , 2 , 6 , 8 [30] Guangming Zhu, Hui Chen, Bin Jiang, Fei Chen, Y uan Xie, and Max W intermark. Application of deep learning to is- chemic and hemorrhagic stroke computed tomography and magnetic resonance imaging. Semin. Ultrasound CT MRI , 43(2):147–152, 2022. 1
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment