IBISAgent: Reinforcing Pixel-Level Visual Reasoning in MLLMs for Universal Biomedical Object Referring and Segmentation
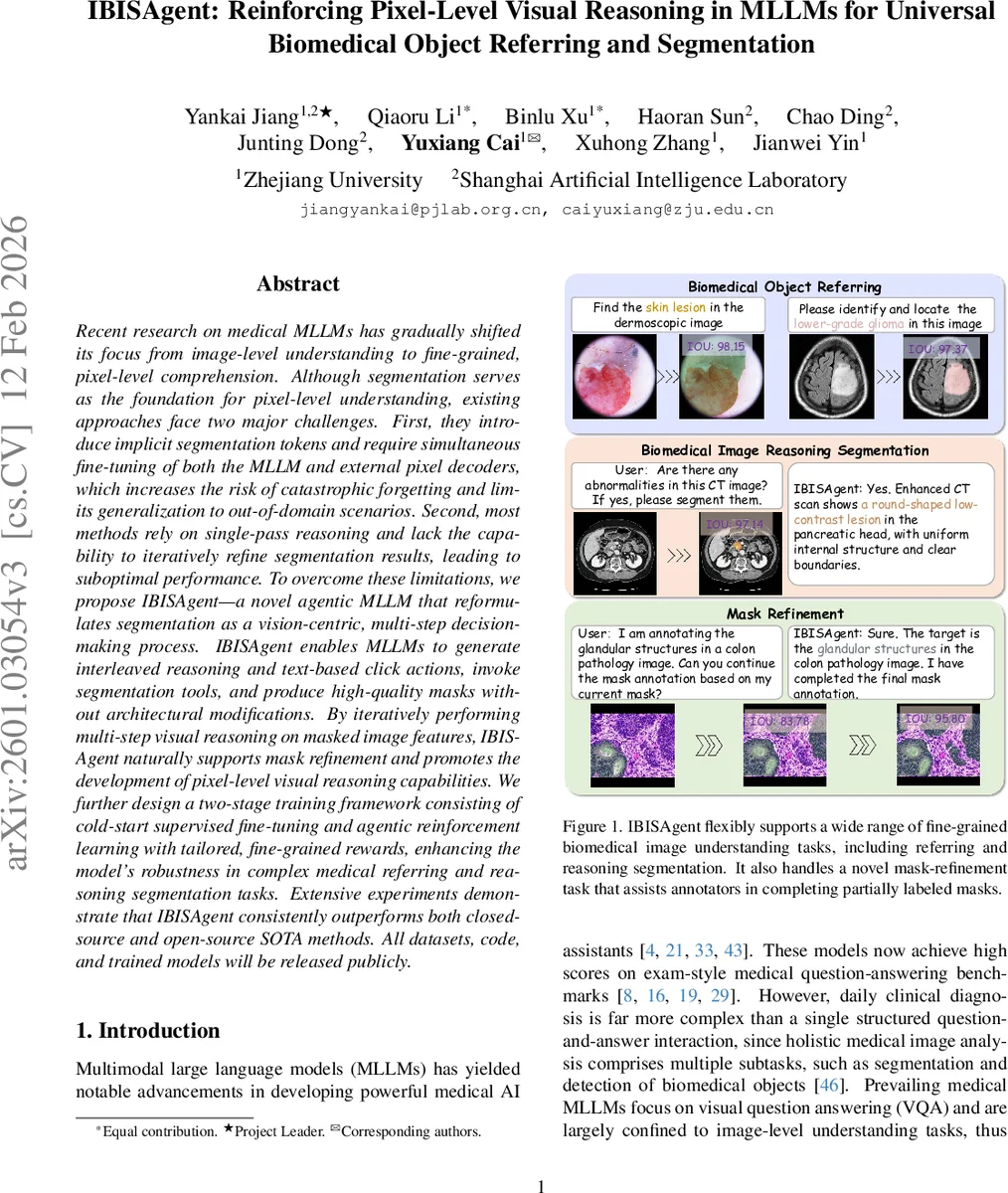
Recent research on medical MLLMs has gradually shifted its focus from image-level understanding to fine-grained, pixel-level comprehension. Although segmentation serves as the foundation for pixel-level understanding, existing approaches face two major challenges. First, they introduce implicit segmentation tokens and require simultaneous fine-tuning of both the MLLM and external pixel decoders, which increases the risk of catastrophic forgetting and limits generalization to out-of-domain scenarios. Second, most methods rely on single-pass reasoning and lack the capability to iteratively refine segmentation results, leading to suboptimal performance. To overcome these limitations, we propose a novel agentic MLLM, named IBISAgent, that reformulates segmentation as a vision-centric, multi-step decision-making process. IBISAgent enables MLLMs to generate interleaved reasoning and text-based click actions, invoke segmentation tools, and produce high-quality masks without architectural modifications. By iteratively performing multi-step visual reasoning on masked image features, IBISAgent naturally supports mask refinement and promotes the development of pixel-level visual reasoning capabilities. We further design a two-stage training framework consisting of cold-start supervised fine-tuning and agentic reinforcement learning with tailored, fine-grained rewards, enhancing the model’s robustness in complex medical referring and reasoning segmentation tasks. Extensive experiments demonstrate that IBISAgent consistently outperforms both closed-source and open-source SOTA methods. All datasets, code, and trained models will be released publicly.
💡 Research Summary
The paper introduces IBISAgent, an agentic multimodal large language model (MLLM) designed to perform fine‑grained, pixel‑level biomedical object referring and segmentation without architectural modifications or the use of implicit segmentation tokens. Existing medical MLLMs typically rely on special tokens (e.g.,
IBISAgent reframes segmentation as a vision‑centric, multi‑step Markov Decision Process (MDP). Given a user query Q and an image I, the agent iteratively generates a reasoning‑action‑observation tuple (rₜ, aₜ, oₜ) for each step t. The textual thinking rₜ is expressed between
Training proceeds in two stages. First, a cold‑start supervised fine‑tuning (SFT) phase uses a large synthetic dataset derived from BiomedParseData (3.4 M image‑mask‑label triples covering 82 object types across nine modalities). An automated click‑simulation algorithm (F_cs) converts each ground‑truth mask into a sequence of positive and negative clicks, yielding over 400 K high‑quality reasoning and action trajectories. The SFT stage teaches the model the correct format of interleaved reasoning, click generation, and mask observation. Second, an agentic reinforcement learning (RL) phase optimizes a policy π_θ with fine‑grained rewards: (1) a region‑based click placement reward that measures proximity of clicks to error regions, (2) a progressive IoU improvement reward that encourages each step to increase overlap with the ground truth, and (3) a trajectory‑length penalty that discourages unnecessary steps. These rewards guide the model to discover efficient, self‑reflective strategies rather than merely mimicking the SFT trajectories.
Extensive experiments on multiple biomedical segmentation benchmarks demonstrate that IBISAgent consistently outperforms both closed‑source and open‑source state‑of‑the‑art (SOTA) methods. In‑domain results show higher IoU scores across tasks such as organ detection, lesion localization, and tumor segmentation. Zero‑shot evaluations on unseen modalities confirm strong generalization. A novel mask‑refinement scenario, where the model receives a partially annotated mask and completes it, yields IoU improvements of up to 95 %, highlighting practical utility for annotation assistance. Ablation studies verify the importance of multi‑step reasoning, the RL fine‑grained rewards, and the removal of implicit segmentation tokens.
The key contributions are: (1) a novel agentic framework that equips MLLMs with pixel‑level visual reasoning without altering the underlying language architecture; (2) a large synthetic trajectory dataset and a two‑stage training pipeline (cold‑start SFT + RL) that together foster robust decision‑making and action planning; (3) comprehensive evaluation showing superior performance on biomedical object referring, segmentation, and mask‑refinement tasks. By treating segmentation tools as plug‑and‑play language‑controlled modules, IBISAgent offers a flexible, extensible solution that bridges the gap between human‑like interactive annotation and automated AI analysis, with potential implications beyond the medical domain.
Comments & Academic Discussion
Loading comments...
Leave a Comment