UniCrop: A Universal, Multi-Source Data Engineering Pipeline for Scalable Crop Yield Prediction
Accurate crop yield prediction increasingly relies on diverse data streams, including satellite observations, meteorological reanalysis, soil composition, and topographic information. However, despite rapid advances in machine learning, most existing…
Authors: Emiliya Khidirova, Oktay Karakuş
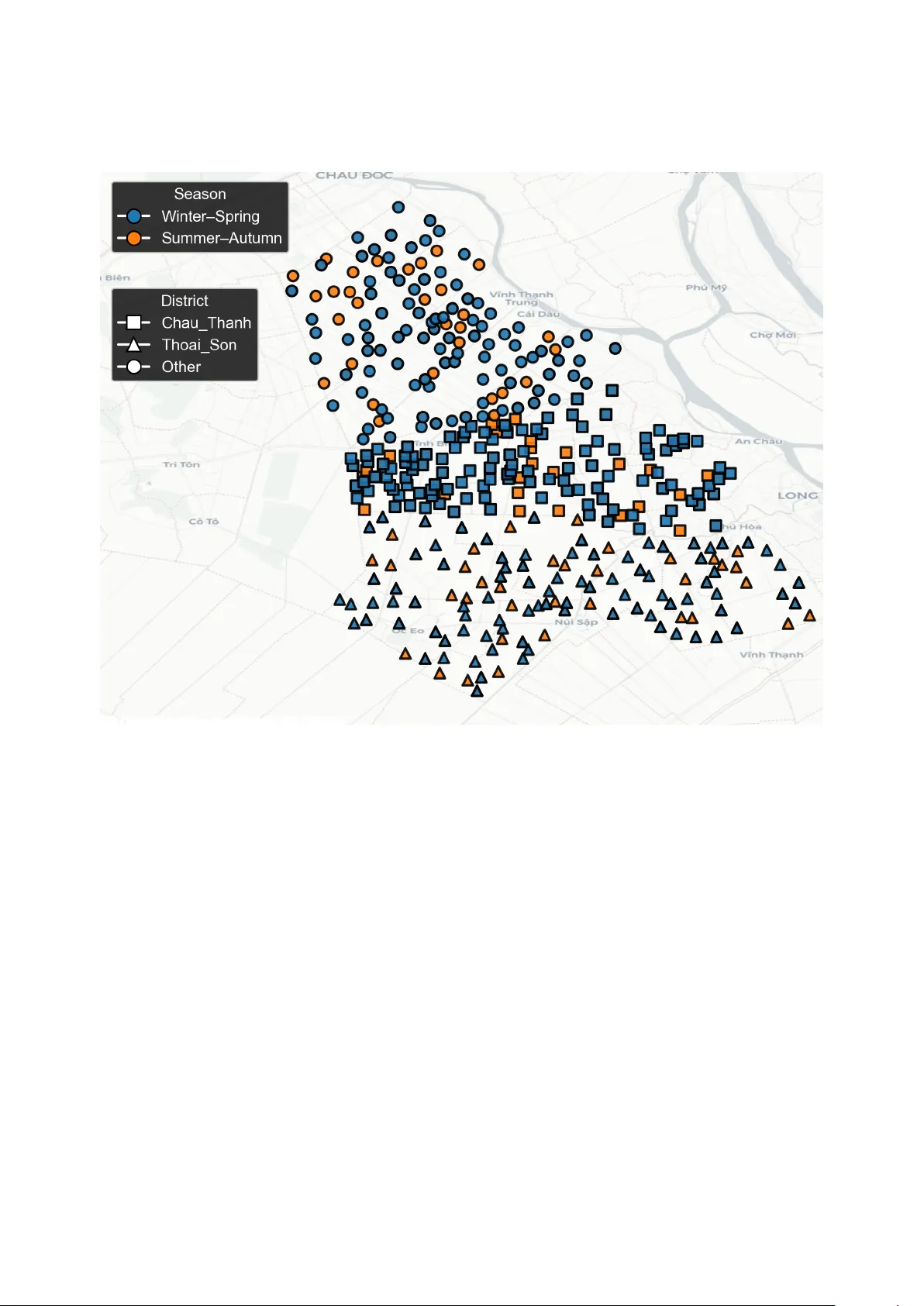
UniCrop: A Uni versal, Multi-Source Data Engineering Pipeline for Scalable Crop Y ield Prediction Emiliya Khidirov a a , Oktay Karaku ¸ s b, ∗ a Car di ff University , School of Mathematics, CF24 4A G UK. b Car di ff University , School of Computer Science & Informatics, CF24 4A G UK Abstract Accurate crop yield prediction increasingly relies on di verse data streams, including satellite observations, meteo- rological reanalysis, soil composition, and topographic information. Howe v er , despite rapid advances in machine learning, most e xisting approaches remain crop- or region-specific and require substantial bespoke data engineering e ff orts. This limits scalability , reproducibility , and operational deployment. This study introduces UniCrop , a uni ver - sal and reusable data pipeline designed to automate the acquisition, cleaning, harmonisation, and feature engineering of multi-source en vironmental data for crop yield prediction. For any gi v en location, crop type, and temporal windo w , UniCrop automatically retrie ves, harmonises, and engineers over 200 en vironmental variables from heterogeneous satellite, climate, soil, and topographic sources (Sentinel-1 / 2, MODIS, ERA5-Land, N ASA PO WER, SoilGrids, and SR TM), reducing them to a compact, analysis-ready feature set utilising a structured feature reduction workflow with minimum redundancy maximum relev ance (mRMR). T o v alidate the pipeline, UniCrop was applied to a rice yield dataset comprising 557 field observations. Using only the selected 15 features, four baseline machine-learning models (LightGBM, Random Forest, Support V ector Regression, and ElasticNet) were trained using rigorous cross- validation. LightGBM achie ved the best single-model performance (RMSE = 465.1 kg / ha, R 2 = 0 . 6576), while a constrained ensemble of all baselines further impro ved accurac y (RMSE = 463.2 kg / ha, R 2 = 0 . 6604). SHAP analysis confirmed agronomically plausible relationships and demonstrated how UniCrop lev erages multi-modal predictors. UniCrop contributes a scalable and transparent data-engineering framew ork that addresses the primary bottleneck in operational crop yield modelling: the preparation of consistent and harmonised multi-source data. By decoupling data specification from implementation and supporting any crop, re gion, and time frame through simple configuration updates, UniCrop provides a practical foundation for transferable, high-quality agricultural analytics at scale. The code and implementation documentation are shared in https://github.com/CoDIS- Lab/UniCrop . K e ywor ds: Crop yield prediction, remote sensing, data harmonisation, agricultural machine learning, feature engineering, multi-source integration, Sentinel, ERA5, SoilGrids, data pipelines. ∗ Corresponding author Email addr esses: emiliyakhidirova@gmail.com (Emiliya Khidirova), karakuso@cardiff.ac.uk (Oktay Karaku ¸ s) 1. Introduction Agriculture plays a central role in global food security and economic stability , yet it faces intensifying pressures from climate change, population growth, and increasingly volatile weather patterns [1, 2]. Accurate crop yield pre- diction is essential for informing government policies, stabilising supply chains, guiding agrib usiness logistics, and supporting farmers’ decisions on irrigation, fertiliser application, and harvest scheduling. These needs ha ve grown more urgent as production shocks in one re gion can propagate rapidly across international markets [3]. Significant adv ances in Earth Observ ation (EO), agro-meteorology , and machine learning (ML) have expanded the potential for data-dri ven agricultural forecasting. Open EO programmes such as the Copernicus Sentinel missions provide optical, radar , and atmospheric measurements at high spatial and temporal resolution, enabling detailed mon- itoring of crop conditions [4, 5]. Long-term vegetation indices from MODIS facilitate phenological analysis across large regions [6]. Climate datasets such as ERA5-Land and NASA PO WER o ff er globally consistent information on temperature, precipitation, radiation, humidity , and wind, key dri vers of agricultural producti vity [7]. Complementary en vironmental datasets from SoilGrids and SR TM further describe soil texture, carbon content, pH, elev ation, slope, and microclimatic influences [8]. Despite these adv ances, the practical dev elopment of yield prediction models remains hindered by a persistent data-engineering bottleneck. Most studies construct bespoke pipelines tailored to a specific crop, region, or time frame, requiring substantial manual e ff ort to integrate heterogeneous datasets and harmonise spatial and temporal resolutions [1]. Even state-of-the-art multimodal deep learning approaches, such as recently proposed ensemble and fusion systems for rice yield prediction [9], rely on comple x, labour-intensi ve preprocessing workflows. As noted in recent revie ws, the central challenge is increasingly one of scalable and reproducible data preparation rather than modelling innov ation [3]. T o address this challenge, we introduce UniCrop , a uni versal, configuration-driven data pipeline that automates the acquisition, harmonisation, and transformation of multi-source en vironmental data for crop yield prediction. Uni- Crop separates the specification of required v ariables from implementation, enabling users to adapt the pipeline to new crops or regions by modifying a simple configuration file. The system integrates optical and radar EO data, ve getation indices, climate reanalysis, agro-climatological variables, soil composition, and topographic layers into a unified, analysis-ready dataset. W e validate UniCrop using a real-world rice yield dataset comprising 557 field observations. The case study demonstrates that high-quality and consistent multi-source features generated by UniCrop support accurate predictions using standard machine-learning models, while interpretability analysis confirms that the pipeline captures agronom- ically meaningful relationships. T ogether , these contributions establish UniCrop as a robust and scalable foundation for multi-crop, multi-region yield prediction. UniCrop is not intended to replace crop-specific modelling expertise or advanced learning architectures. Instead, it aims to provide a robust, reusable, and transparent data foundation upon which a wide range of statistical and 2 machine-learning models can be applied consistently across crops and regions. 2. Background Crop yield prediction has evolv ed substantially over the last two decades, driv en by advances in remote sensing, agro-meteorological modelling, and machine learning (ML). Early approaches predominantly relied on statistical regressions and empirical models calibrated for individual crops and regions. Recent studies, howe ver , increasingly integrate multi-source en vironmental data with ML architectures to capture non-linear crop–en vironment interactions. Despite strong progress, persistent challenges remain in scalability , data heterogeneity , and operational generalisation. This section revie ws the ke y de velopments in three interconnected domains: (i) satellite-based remote sensing, (ii) en vironmental and agro-climatic data integration, and (iii) machine-learning approaches for yield prediction. W e conclude by identifying the unresolved limitations in current practice that moti v ate the UniCrop framew ork. 2.1. Satellite-Based Optical and Radar Remote Sensing Remote sensing has become central to agricultural monitoring, supporting ve getation assessment, phenology tracking, flood mapping, and yield forecasting. Optical sensors, particularly the Multispectral Instrument aboard Sentinel-2, deliv er high-resolution reflectance data used to deriv e ve getation indices such as the Normalised Dif- ference V e getation Index (NDVI) and Enhanced V e getation Index (EVI). These indices provide proxies for canopy greenness, biomass accumulation, and stress responses [5, 4]. Howe ver , optical data su ff er from cloud contamination, especially in monsoon-a ff ected or tropical regions where agricultural monitoring is most needed. Synthetic Aperture Radar (SAR) addresses this limitation by providing cloud-penetrating micro wav e observ ations. Sentinel-1 SAR data capture structural and moisture-related properties of crop canopies, enabling robust monitoring under all-weather conditions [10]. Studies increasingly fuse optical and SAR observations to exploit their comple- mentary characteristics. Recent transformer -based fusion models demonstrate performance gains for rice mapping and phenology recognition [4], while multimodal satellite–weather systems hav e been sho wn to improv e rice yield prediction accuracy through ensemble deep learning architectures [9]. These trends highlight the power of EO fusion but also underscore the data-engineering w orkload inherent in multi-sensor integration. 2.2. Coarse-Resolution T ime Series and V e getation Dynamics Beyond high-resolution imagery , coarse-resolution products such as MODIS remain essential due to their high temporal frequency and global co verage. MODIS vegetation indices (MOD13), leaf area index (MOD15), e vapotran- spiration (MOD16), and gross primary productivity (MOD17) have been widely used to monitor seasonal dynamics of crop dev elopment [6]. Their consistent, cloud-screened composites enable long-term trend analysis and facili- tate large-scale or multi-region yield modelling. T ime-series modelling using NDVI curves, phenological metrics, or harmonised Landsat–MODIS series has been applied to wheat, maize, rice, soybean, and potato systems worldwide 3 [11, 12]. Howe ver , harmonising MODIS with higher-resolution datasets requires careful temporal alignment and spatial reduction, which is often implemented manually . 2.3. Inte grating Climatic, Soil, and T opographic Driver s Crop development is strongly influenced by weather variability and en vironmental conditions. Climate reanaly- sis products such as ERA5-Land provide consistent global records of temperature, radiation, precipitation, and soil moisture, critical for modelling crop–climate interactions [2]. Similarly , N ASA PO WER o ff ers agro-climatology- ready variables including vapour pressure deficit (VPD), dewpoint temperature, wind speed, and diurnal temperature range (DTR). These variables capture k ey stressors associated with heatwaves, humidity fluctuations, and atmospheric dryness. Static en vironmental properties further influence crop growth. SoilGrids provides global soil texture, carbon, pH, and bulk density maps at 250 m resolution, enabling soil–water and nutrient av ailability modelling [8]. T errain data from the Shuttle Radar T opography Mission (SR TM) inform hydrological behaviour , slope-driv en erosion, and microclimate [13]. Integrating these di verse datasets requires careful standardisation, reprojection, temporal matching, and prov enance tracking, steps that are rarely automated in existing agricultural analytics workflo ws. 2.4. Machine Learning Appr oaches for Y ield Prediction Machine learning has become a dominant tool in data-driv en yield prediction, o ff ering the ability to model com- plex, non-linear relationships across di verse input features. Ensemble tree-based models such as Random Forests, Gradient Boosting Machines, and LightGBM frequently outperform classical regressions due to their robustness and ability to handle noisy inputs [1, 3]. Recent studies have also explored neural architectures, including con volutional neural networks (CNNs) for spatial imagery and recurrent or LSTM networks for time-series climate and phenology modelling [14]. Deep ensembles combining EO imagery , meteorology , and crop-gro wth indicators ha ve demonstrated strong performance for rice, maize, and wheat yield prediction [9, 15]. Despite these modelling inno vations, sev eral challenges persist. Deep learning architectures require lar ge datasets and are sensitiv e to missing or misaligned inputs. More importantly , many state-of-the-art models perform only marginally better than simpler baselines when trained on datasets that are inconsistently engineered or incomplete [3]. This underscores a critical insight: data quality and harmonisation often determine predictive performance more than model complexity . 2.5. Limitations of Curr ent Pipelines and Motivation for UniCr op Across the literature, most studies de velop bespoke workflo ws tailored to a single crop, region, or satellite product. These workflows are often complex, manually curated, and di ffi cult to reproduce. The lack of standardised pipelines leads to siloed methodologies, limited transferability , and substantial entry barriers for agricultural researchers. As noted in sev eral re vie ws [1, 7], the primary obstacle to scalable yield prediction is not model design but the engineering required to gather , harmonise, and prepare multi-source en vironmental data. 4 UniCrop directly addresses these limitations by providing a universal, configuration-driven, multi-source data pipeline that automates acquisition, cleaning, temporal alignment, spatial aggreg ation, feature engineering, and selec- tion across EO, climate, soil, and topographic datasets. By standardising the data-engineering foundation, UniCrop enables reproducible and transferable yield modelling regardless of crop type or geographic conte xt. 3. Materials and Methods UniCrop is designed as a uni versal, reusable, and configuration-dri ven data pipeline that automates the process of acquiring, cleaning, harmonising, and transforming multi-source en vironmental data into an analysis-ready dataset for crop yield prediction. The system decouples the choice of data sources from the underlying implementation, enabling rapid portability across crops, regions, and temporal windo ws. This section details the full UniCrop methodology , from pipeline design and data ingestion to feature engineering, selection, and model training. 3.1. High-Level Pipeline Ar c hitectur e Sentinel-1 (SAR) Sentinel-2 (Optical) MODIS (ET , ND VI, fP AR) ERA5-Land NASA PO WER SoilGrids SR TM (T opography) Multi-source Data Acquisition Data Harmonisation & T emporal / Spatial Alignment Feature Engineering mRMR Feature Selection (Compact 15 Feature Set) Baseline ML Modelling (LightGBM, RF , SVR, ElasticNet) Ensemble & Interpretability (SHAP , Metrics) Figure 1: Hybrid schematic of the UniCrop pipeline. Multiple environmental data sources are ingested in parallel, harmonised into a unified structure, enriched with agronomic features, reduced via mRMR into a compact predictor subset, and finally used for baseline modelling and ensemble ev aluation. The UniCrop architecture (Figure 1) is structured into fiv e modular stages: 1. Configuration and Planning : Users specify required features through a structured feature mapping file and provide field-le vel observ ations (latitude, longitude, dates). These inputs drive a dynamically constructed fetch plan . 5 2. Data Acquisition : Multi-source en vironmental data are collected from satellite, climate, soil, and topographic repositories through Google Earth Engine (GEE) and public APIs. 3. Data Harmonisation : All source outputs are temporally and spatially aligned, cleaned, standardised, and merged into a unified table with complete pro venance records. 4. Featur e Engineering and Selection : Statistical screening, agronomic feature engineering, and minimum re- dundancy maximum rele vance (mRMR) selection are applied to deriv e a compact subset of informati ve predic- tors. 5. Baseline Model T raining and Evaluation : Selected features are used to train multiple baseline models under rigorous cross-validation, follo wed by a constrained ensemble. This modular design ensures that extending UniCrop to new crops, regions, or temporal resolutions requires changes only to the configuration layer , without modification of the underlying pipeline logic. 3.2. F eatur e Mapping and F etch Plan Generation The feature mapping file serves as the central configuration layer that defines all en vironmental variables required by UniCrop. Each entry specifies the key variable name, the API parameter used for retriev al, the source dataset, the platform (e.g. GEE, NASA PO WER, SoilGrids), and any associated deriv ation or calculation rules. This structured mapping enables UniCrop to separate data specification from data implementation, ensuring that users can adapt the pipeline to ne w crops or regions simply by modifying a human-readable configuration file rather than altering code. An illustrativ e excerpt is shown in Figure 2, which highlights how variables such as EVI, ev apotranspiration, elev ation, and total precipitation are declaratively defined. During execution, UniCrop expands this mapping into a comprehensive fetch plan that enumerates all field–variable combinations, forming the basis for automated multi- source data acquisition. 3.2.1. F eatur e Mapping Specification All features required by the user are defined in a single CSV specification, unicrop_feature_mapping.csv . Each row corresponds to a data v ariable and includes: • Key V ariable : Human-readable identifier (e.g., NDVI , T2M , soil_carbon ). • API Parameter : Data-source-specific tok en controlling retrie val (e.g., Sentinel-2 band names or NASA POWER API codes). • Source Dataset : The product identifier (e.g., ERA5-Land, MODIS MOD13Q1, Copernicus Sentinel-1). • Platform : Retriev al route (e.g., GEE collection ID, API endpoint). • Notes / Derivation : Optional deriv ation instructions, including formulas for indices. 6 Key V ariable API Param Source Dataset Platform Notes / Derivation Aspect aspect SR TM (DEM) USGS / SR TMGL1_003 Deriv ed from DEM using terrain() func- tion in GEE. ET (Evapotranspiration) ET MOD16A2 (MODIS) MODIS / 006 / MOD16A2 Daily ev apotranspiration (mm / day). EVI Deriv ed Sentinel-2 COPERNICUS / S2 EVI = 2 . 5 N I R − RE D N I R + 6 RE D − 7 . 5 BLU E + 1 . Elev ation elev ation SR TM (DEM) USGS / SR TMGL1_003 Elev ation (m) from SR TM DEM. Irrigation Derived ERA5-Land ECMWF / ERA5_LAND / D AIL Y Irrigation = max(0, PEV – TP). T otal Precipitation total_precipitation ERA5-Land ECMWF / ERA5_LAND / DAIL Y Accumulated precipitation over time step. Excerpt fr om unicrop_feature_mapping.csv Fetch Plan (Fields × V ariables) Automated Data Acquisition Figure 2: Schematic representation of UniCrop’ s feature mapping system, illustrated with real e xamples from the mapping file. The mapping table defines variables, API parameters, datasets, and deri vations used to b uild the automated fetch plan. By separating data specification from implementation, the feature mapping mechanism also improves repro- ducibility , as the exact data requirements for a study can be shared, audited, and reused independently of code. 3.2.2. F etch Plan Construction The system automatically generates a fetch plan that enumerates all required retriev al operations. This is con- structed by: 1. Cleaning and standardising the input field table (coordinates, dates, identifiers); 2. Normalising the feature mapping (column names, dataset identifiers, API parameters); 3. Creating a Cartesian product between input records and feature requests. The resulting fetch_plan.csv defines, for each field-date record, which v ariable to retriev e, from which source, and under what parameters. Duplicates, in valid coordinates, and malformed ro ws are automatically remo ved. 3.3. Data Sour ces and Acquisition Methods UniCrop integrates six major data categories: optical satellite imagery , radar backscatter , coarse-resolution vege- tation products, meteorological reanalysis, agro-climatological data, and environmental context (soil and topography). T able 1 summarises all sources. 3.4. Data Harmonisation and Master T able Construction All source-specific outputs are merged into a master time-series table keyed by latitude, longitude, and date. UniCrop applies: 7 T able 1: Summary of data sources inte grated in UniCrop. Source T ype Data Pr ovided Sentinel-2 MSI Optical RS Reflectance bands; ND VI, EVI, SA VI, NDRE, CI red-edge ; canopy biophysical indices Sentinel-1 SAR Radar RS VV , VH backscatter; VV / VH ratio; Radar V egetation Index (R VI); texture features MODIS (MOD13, MOD15, MOD16, MOD17) V egetation ND VI / EVI (16-day), fP AR, LAI, ET , GPP ERA5-Land Climate T emperature, precipitation, soil moisture, radiation, ev aporation (hourly) N ASA PO WER Agro-climate T2M, T2M_MIN / MAX, RH2M, VPD, DTR, wind speed, solar radiation SoilGrids Soil Soil texture, SOC, b ulk density , nitrogen, pH (0–30 cm) SR TM T opography Ele v ation, slope, aspect, terrain deriv ati ves • ISO-standard date formatting, • numeric casting and float precision enforcement, • duplicate remov al with prioritisation of data completeness, • su ffi x restoration for consistent naming (e.g., elev ation, slope), • generation of unicrop_columns_manifest.csv for prov enance. Missingness is preserved until imputation within cross-v alidation to pre vent leakage. 3.5. F eatur e Engineering UniCrop generates additional features capturing agronomic processes: • Growing Degr ee Days (GDD) : GDD base10 = max 0 , T max + T min 2 − 10 • Chill nights : count of days where T min < 15 ◦ C. • V egetation dynamics : seasonal amplitude of NDVI / EVI. • SAR texture metrics : variability in VV / VH backscatter . 8 • Soil–climate interactions : e.g., clay × radiation, elev ation × temperature. All engineered features undergo type v alidation and schema alignment, and the y are designed to be crop-agnostic and deriv ed solely from en vironmental signals, ensuring applicability across di ff erent cropping systems without re- quiring crop-specific calibration. 3.6. Statistical Scr eening and mRMR F eatur e Selection Feature screening proceeds in sev eral stages designed to remov e redundant or uninformativ e predictors before applying a more principled selection method. First, features with near-zero variance are discarded. Second, highly collinear v ariables ( r ≥ 0 . 98) are pruned by retaining only the feature with the highest bi variate association with yield as measured by mutual information. Third, a family-preserv ation heuristic ensures that at least one feature from each major en vironmental data family (meteorology , vegetation, SAR, soil, topography) is retained in the candidate pool, prev enting ov er-selection from an y dominant modality . The final selection step emplo ys the minimum redundanc y maximum relev ance (mRMR) algorithm, a widely used filter-based technique for identifying compact feature subsets. Let S denote the selected feature set and f a candidate feature. The r elevance of f to the target v ariable y is quantified using mutual information, I ( f ; y ) = Z Z p ( f , y ) log p ( f , y ) p ( f ) p ( y ) d f d y , which captures general (non-linear) dependencies beyond simple correlation. In practice, an ensemble rele vance score combining mutual information, Pearson correlation, and Spearman rank correlation is used to enhance robustness across varying feature types. The r edundancy between a candidate feature f and an existing selected feature s ∈ S is computed as R ( f ; S ) = 1 | S | X s ∈ S I ( f ; s ) , penalising features that provide overlapping information. The mRMR objectiv e seeks to maximise the di ff erence between relev ance and redundanc y: max f < S I ( f ; y ) − R ( f ; S ) , or equiv alently , maximise the relev ance-to-redundanc y ratio, max f < S " I ( f ; y ) R ( f ; S ) + ϵ # , where ϵ is a small constant prev enting division by zero. Features are added iterativ ely until the desired subset size is reached. In this study , a compact set of 15 pre- dictors (UniCrop default value is 15, and can be set by users to any value) is selected independently within each cr oss-validation fold to ensure that no information from the test partitions influences feature selection, thereby pre- venting leakage. The resulting subset preserves div ersity across en vironmental data families while maintaining a strong aggregate rele v ance to rice yield. 9 The resulting compact feature set prioritises interpretability and generalisability ov er mar ginal performance gains, aligning to produce robust baseline models rather than maximally tuned predictors. 3.7. Model T r aining and Evaluation 3.7.1. Cr oss-V alidation Strate gy Model e valuation is conducted using a 5-fold shu ffl ed cross-validation scheme at the field level. T o ensure an unbiased assessment, all preprocessing steps, including imputation, scaling, statistical screening, feature engineering, and mRMR feature selection, are performed independently within each training fold . V alidation folds remain com- pletely unseen during preprocessing, pre venting information leakage and yielding a realistic estimate of out-of-sample performance. 3.7.2. Imputation and Scaling Missing values and scale heterogeneity are addressed using a family-aw are preprocessing strategy designed to preserve physical meaning while preventing information leakage. All imputation and scaling steps are performed within each cr oss-validation fold . • Meteorological variables : multiv ariate iterativ e imputation is applied to e xploit cross-variable dependencies among temperature, humidity , radiation, and precipitation, improving robustness under partial missingness. • V egetation indices : K-nearest neighbours (KNN) imputation is used with district–season contextual grouping, ensuring that gap-filling respects local agro-climatic conditions and phenological patterns. • Soil and topographic variables : median imputation is emplo yed, reflecting their quasi-static nature and reduc- ing sensitivity to outliers. • Outlier handling : extreme values are winsorised at the 1% lev el to limit the influence of measurement noise and retriev al artefacts. • Featur e scaling : robust scaling based on median and interquartile range is applied to all numeric features, ensuring comparability across heterogeneous data sources while maintaining resistance to heavy-tailed distri- butions. 3.7.3. Baseline Models T o v alidate the quality and representati veness of the datasets generated by UniCrop, a set of widely used and well- understood machine-learning models is emplo yed. These models are not introduced as methodological contributions, but rather as r eference baselines commonly reported in the crop yield prediction literature. The following models are trained: • LightGBM : gradient-boosted decision trees suitable for heterogeneous, non-linear feature spaces. 10 • Random For est : an ensemble of bagged decision trees providing rob ust performance with minimal tuning. • Support V ector Regression (RBF ker nel) : a kernel-based method capturing non-linear relationships. • ElasticNet : ℓ 1 – ℓ 2 regularised linear re gression serving as a transparent linear baseline. Hyperparameter tuning is intentionally limited to reasonable def ault ranges to maintain methodological simplicity and reproducibility . The objectiv e of this step is not to maximise predictiv e performance, but to demonstrate that the features constructed by UniCrop are su ffi ciently informative to support competitiv e baseline modelling. Users are encouraged to apply more advanced models or extensiv e tuning strategies according to their specific research objectiv es, as improv ed performance is likely achie vable be yond the scope of this study . 3.7.4. Ensemble Modelling T o assess whether complementary strengths of individual baseline models can be exploited, a simple ensemble is constructed using out-of-fold predictions. For each training fold, predictions from all base learners are retained and combined only at the validation stage, ensuring that ensemble optimisation does not introduce information leakage. The ensemble weights are estimated by solving a constrained least-squares optimisation problem: min w y − 4 X i = 1 w i ˆ y i 2 s.t. w i ≥ 0 , X i w i = 1 , where ˆ y i denotes the out-of-fold predictions from the i -th base model. Non-negati vity and sum-to-one constraints ensure interpretability and pre vent ov er-reliance on any single model. Optimisation is performed using the Sequential Least Squares Programming (SLSQP) algorithm. The ensemble is not intended to maximise performance, but to provide a stable and interpretable aggregation of baseline predictors, further illustrating the consistency and usefulness of the UniCrop-generated feature set. 3.7.5. Evaluation Metrics Model performance is assessed using multiple complementary metrics to capture both absolute and relative pre- diction errors: RMSE , MAE , R 2 , MAPE . Root Mean Squared Error (RMSE) emphasises large de viations and is sensitiv e to extreme errors, while Mean Absolute Error (MAE) provides a more robust measure of average prediction accuracy . The coe ffi cient of determi- nation ( R 2 ) quantifies the proportion of yield variance explained by the model, and Mean Absolute Percentage Error (MAPE) o ff ers an intuitiv e relati ve error measure suitable for agronomic interpretation. 11 In addition to predicti ve accuracy , model interpretability is ev aluated using SHAP (Shapley Additive Explana- tions). SHAP values are computed for the strongest-performing baseline model to analyse both global feature im- portance and local, instance-lev el contrib utions, enabling assessment of whether predictions rely on agronomically meaningful signals across data sources. 4. Case Study: V alidation of the UniCrop Pipeline Using Rice Y ield Data T o ev aluate the e ff ectiveness, robustness, and generalisability of UniCrop, a validation study w as conducted using a rice yield dataset comprising 557 field-lev el observ ations. This case study demonstrates ho w harmonised, multi-source en vironmental data generated by UniCrop can be utilised to support accurate and interpretable yield predictions using standard machine-learning models. Detailed information on the dataset is av ailable through the original challenge documentation (link) and the accompanying academic publication by Y ewle et al. [9]. The objectiv e of this analysis is not to maximise predictive accuracy , b ut rather to demonstrate that UniCrop reliably produces high-quality , analysis-ready datasets that enable robust baseline modelling without manual data engineering. 4.1. Study Ar ea and Spatial Context The dataset spans multiple agricultural districts representing distinct climatic and soil conditions. T o provide spatial conte xt, Figure 3 sho ws the distrib ution of the 557 field parcels, colour-coded by season. Spatial heterogeneity in field locations underscores the importance of harmonising multi-source en vironmental predictors. 4.2. Dataset Description The reference dataset includes geolocated field parcels with recorded yield measurements (kg / ha), administrativ e attributes (district, season), and harv est dates. These serve as the seed inputs for UniCrop; all en vironmental variables are obtained e xclusiv ely through automated pipeline execution. After running the full UniCrop process, the result- ing master dataset contained approximately 160 candidate features spanning meteorology , v egetation indices, radar backscatter , soil composition, and topography . 4.3. Exploratory Data Assessment The distribution of rice yield values exhibited a unimodal shape centred around approximately 6,600 kg / ha, with mild right skewness. Extended analysis shows that the di ff erences across growing seasons (Summer–Autumn vs. W inter–Spring) reflect kno wn climatic and spatial heterogeneity in the re gion. V egetation-related features exhibited higher missingness during the Summer–Autumn season due to monsoon- related cloud co ver , while meteorological and soil properties were lar gely complete. This informed the f amily-specific imputation strategy applied during cross-v alidation. 12 Figure 3: Spatial distribution of field parcels included in the case study , colour-coded by growing season. 4.4. F eatur e Reduction and mRMR Selection Follo wing schema alignment, UniCrop applies a multi-stage feature reduction strategy designed to remove re- dundancy , enhance interpretability , and ensure that all major en vironmental data families remain represented. The initial filtering step eliminates near-zero variance features and prunes highly collinear variables ( r ≥ 0 . 98), retaining only those with the strongest biv ariate association with yield based on mutual information. Agronomic feature engi- neering further enriches the dataset by incorporating thermal time, v egetation dynamics, SAR v ariability metrics, and soil–climate interaction terms. T o obtain a compact yet informative set of predictors, we employ the minimum redundancy maximum relev ance (mRMR) algorithm. mRMR ranks features by balancing two criteria: ( i ) high relev ance to the response variable, quantified using a combination of mutual information and rank-based correlations; and ( ii ) low redundancy with respect to features already selected. This ensures that the final subset retains div erse information content rather than concentrating on a single data family or feature type. The algorithm is run independently within each training fold to 13 Figure 4: Distribution of rice yield (kg / ha). Figure 5: Seasonal missingness patterns for vegetation indices. av oid information leakage. Figure 6 visualises the top 15 selected predictors. Bars are colour-coded by feature f amily to illustrate the breadth of en vironmental signals captured by the subset, including v egetation indices, SAR backscatter statistics, meteorolog- ical indicators, soil properties, and topographic descriptors. This f amily-lev el di versity reflects the multi-source nature 14 of UniCrop’ s feature space and demonstrates that the selection process draws from all major data domains rather than ov erfitting to a single modality . Figure 6: T op 15 predictors selected by the mRMR process. Bars are colour-coded by variable family (meteorology , ve getation, SAR, soil, topography). The resulting 15-feature set achieves a balance between interpretability and predictive strength. By reducing the original ∼ 160 candidate variables to a well-structured and div erse subset, mRMR provides a principled foundation for do wnstream modelling, while maintaining representation across all major en vironmental driv ers of rice yield. The presence of v egetation, meteorological, SAR, soil, and topographic v ariables among the selected predictors highlights UniCrop’ s ability to preserve complementary information from heterogeneous data sources. 4.5. Model P erformance Four baseline models, LightGBM, Random F orest, Support V ector Re gression, and ElasticNet, were trained under a strict 5-fold cross-validation framew ork, where all preprocessing and feature selection were carried out indepen- dently within each fold. This ensures honest ev aluation without information leakage. T able 2 summarises the out-of-fold performance metrics. LightGBM achiev ed the lowest RMSE and highest R 2 among single models, while a constrained linear ensemble of all base learners provided a small but consistent improv ement. Figure 7 visualises the relationship between predictions and observations. Predictions from LightGBM and Elas- ticNet aligned most closely with the identity line, indicating lower error v ariance. T o further examine model behaviour , Figure 8 presents a residual distribution plot and residuals versus fitted values. Residuals were approximately centred around zero, with no major heteroskedastic patterns, suggesting that the engineered and selected features provide stable model beha viour . 15 T able 2: Cross-validated performance metrics for baseline and ensemble models. Model RMSE (kg / ha) MAE (kg / ha) R 2 MAPE (%) LightGBM 465.1 378.6 0.6576 5.72 Random Forest 480.0 392.7 0.6353 5.92 SVR (RBF) 526.0 425.9 0.5621 6.43 ElasticNet 467.6 380.2 0.6539 5.73 Ensemble 463.2 375.0 0.6604 5.66 Figure 7: Out-of-fold predictions vs. observed yields. 4.6. Interpr etability via SHAP Analysis Global SHAP (Shapley Additi ve Explanations) values were computed for the LightGBM model to understand feature influence. Sev eral patterns were consistent with agronomic knowledge: 16 Figure 8: Residual distributions and residuals vs. fitted v alues for the LightGBM model. • high maximum relativ e humidity and stable night-time temperatures were associated with higher yield, • high temperature variability and e xtreme heat had negati ve impacts, • SAR texture metrics highlighted structural di ff erences in the canopy link ed to lodging or unev en growth, • solar radiation interacted non-linearly with diurnal temperature range. Figure 9: Global SHAP Importance plot for the T op 5 features. 17 These interpretability results confirm that the model relies on agronomically meaningful signals across meteorol- ogy , vegetation, SAR, soil, and topography , reflecting the multi-modal nature of the UniCrop dataset. These patterns are consistent with established agronomic understanding of rice growth, indicating that UniCrop-derived predictors capture meaningful en vironmental dri vers rather than spurious correlations. 4.7. Summary of V alidation F indings The case study demonstrates four key aspects of UniCrop’ s e ff ecti veness: 1. High-quality , harmonised predictors : The pipeline reliably integrates div erse en vironmental datasets into a compact, informativ e predictor set. 2. Strong baseline predicti ve perf ormance : Ev en without model-specific tuning, the processed dataset supports competitiv e performance among classic ML models. 3. Stable and interpretable residual structure : Residual diagnostics sho w well-behav ed model error distribu- tions. 4. Robust generalisation : Performance remains stable under seasonal and spatial shifts. T aken together , these findings validate UniCrop as a robust, univ ersal data pipeline suitable for scalable yield modelling across di ff erent regions and crops. 5. Discussion and Limitations The UniCrop pipeline provides a unified, scalable, an d data-centric foundation for crop yield prediction. By automating multi-source data acquisition and harmonisation, the framew ork addresses a persistent bottleneck in agri- cultural analytics: the need for bespoke, time-consuming pipelines tailored to specific studies. The case study on rice yield prediction demonstrates that UniCrop can produce high-quality , analysis-ready datasets capable of sup- porting competitiv e predictive performance using only standard machine-learning models. This section synthesises the broader implications of UniCrop, its practical value for agricultural modelling, and the remaining limitations that motiv ate future work. 5.1. Contributions and Pr actical Implications A key contribution of UniCrop is the decoupling of data specification from implementation. Through a flexible configuration file, users can modify crop type, re gion, or temporal co verage without altering the underlying codebase. This design enables UniCrop to function as a reusable tool rather than a single-use pipeline, facilitating reproducibil- ity and lowering technical barriers for researchers and practitioners. As agricultural modelling becomes increasingly reliant on multi-modal environmental data, such standardisation o ff ers a pragmatic route to ward operational deploy- ment. 18 The pipeline’ s multi-source inte gration, spanning Sentinel-1 / 2 imagery , MODIS vegetation dynamics, reanaly- sis climate records, agro-climatological variables, soil properties, and terrain information, ensures that the resulting dataset captures di verse en vironmental dri vers. The mRMR-based reduction to a compact set of 15 predictors demon- strates that much of the predictiv e power can be distilled into a small, interpretable subset while still lev eraging mul- tiple data families. The robustness of the case study results, including the consistency of SHAP interpretations with agronomic kno wledge, emphasises the importance of well-structured data engineering in supporting model reliability . From an applied perspecti ve, UniCrop can support w orkflows such as re gional monitoring systems, early warning tools, or large-scale benchmarking studies. The ability to generate consistent feature sets across di ff erent crops and geographies provides a basis for comparati ve analysis and transferable modelling frame works. 5.2. Limitations and Operational Challenges Despite its strengths, UniCrop exhibits sev eral limitations that require consideration. First, the claim of crop- independence has been validated using only a single crop (rice) in a specific region. Although the pipeline is techni- cally generalisable, empirical validation across div erse agro-climatic contexts, including temperate cereals, horticul- tural crops, and drought-prone systems, is necessary to confirm univ ersality . Second, the accuracy of UniCrop outputs is inherently constrained by the quality and availability of public data sources. Optical imagery remains susceptible to persistent cloud cover , which can reduce the reliability of vegetation indices in tropical or monsoon-dominated re gions. While the inclusion of SAR data mitig ates this issue, SAR-deri ved features capture di ff erent aspects of crop structure and do not fully substitute for optical metrics. Third, UniCrop does not currently include management v ariables such as irrigation schedules, fertiliser use, plant- ing density , or cultiv ar information. These factors can substantially influence yield variation, particularly at the field scale. W ithout such management data, model performance may plateau, especially for fine-grained predictions. Fourth, although UniCrop performs extensi ve preprocessing, the pipeline presently produces a single “snapshot” of features per field-date pair , rather than incorporating full temporal trajectories. As interest gro ws in spatio-temporal deep learning, future versions of UniCrop may require extensions to extract sequential, multi-date time series from satellite and climate sources. Finally , the computational cost of lar ge-scale GEE queries, especially when co vering multiple seasons or high- resolution Sentinel products, may pose practical constraints for countries or institutions with limited cloud resources. 5.3. Futur e Dir ections Sev eral av enues for future de velopment arise from these limitations: • Multi-crop and multi-r egion validation : Applying UniCrop to wheat, maize, soybean, and other major crops across div erse climates will empirically test uni versality . • Integration of management data : Incorporating farmer-reported or farm-management system inputs could greatly enhance fine-scale modelling. 19 • Spatio-temporal extensions : Adding modules for extracting time series (e.g., multi-date ND VI / SAR / ERA5) would support deep sequence models such as LSTMs, TCNs, and transformer -based architectures. • SAR–optical fusion : Cloud-robust composites that combine Sentinel-1 and Sentinel-2 could improve ve geta- tion monitoring in cloud-prone en vironments. • Local high-resolution refinement : UniCrop could be adapted into a hybrid framework combining global sources with user-supplied U A V imagery or on-farm sensor data. • Scalable deployment : Containerised or cloud-nativ e implementations (e.g., using Kubernetes or serverless functions) may improv e performance for national-scale applications. Overall, UniCrop represents a significant step toward reproducible and transferable agricultural analytics. By ad- dressing the data-engineering bottleneck at the core of yield prediction w orkflows, the frame work lays the foundation for robust, scalable, and interpretable modelling across a wide range of agricultural systems. 6. Conclusion This study introduced UniCrop , a univ ersal and configuration-dri ven data pipeline designed to automate the prepa- ration of multi-source en vironmental data for crop yield prediction. By integrating satellite imagery , climate reanal- ysis, agro-climatological variables, soil composition, and topographic information into a unified and harmonised structure, UniCrop directly addresses one of the most persistent challenges in agricultural analytics: the construction of reproducible, scalable, and generalisable data-engineering workflo ws. The pipeline’ s design separates the specifi- cation of required features from implementation, o ff ering a modular and extensible architecture that can be rapidly adapted to new crops, re gions, and temporal windo ws. The validation case study using 557 rice field observations demonstrates that UniCrop generates analysis-ready datasets capable of supporting strong predictiv e performance from standard machine-learning models. The compact set of 15 predictors selected through mRMR balances interpretability and predictive power while maintaining repre- sentation across all major data families. The resulting models, including a constrained ensemble, achiev ed competiti ve accuracy without the need for crop-specific tuning, underscoring the importance of a high-quality data pipeline over model complexity . SHAP-based interpretability further confirmed that the models rely on agronomically meaningful relationships, reinforcing the credibility and scientific soundness of the UniCrop-deriv ed feature set. While the pipeline exhibits strong generality and robustness, several limitations remain. The absence of manage- ment data, reliance on public EO sources, and lack of temporal-sequence extraction constrain its applicability in some contexts. Empirical validation across additional crops and agro-climatic regions will also be important to fully estab- lish univ ersality . Nonetheless, these limitations highlight natural directions for future work, including the integration of high-resolution local data, SAR–optical fusion, and support for spatio-temporal deep learning architectures. 20 Overall, UniCrop represents a significant contribution to data-centric agricultural machine learning. By automat- ing the most labour-intensiv e stages of data preparation, it enables researchers and practitioners to focus on model innov ation, scenario analysis, and decision support. As global agriculture faces intensifying climatic and economic pressures, tools like UniCrop provide a practical and scalable foundation for transparent, transferable, and impactful yield prediction systems. References [1] M. A. Jabed, M. A. A. Murad, Crop yield prediction in agriculture: A comprehensi ve re vie w of machine learning and deep learning approaches, with insights for future research and sustainability , Heliyon 10 (7) (2024) e40836. doi:10.1016/j.heliyon.2024.e40836 . [2] H. Li, Z. Gan, H. Lu, The impact of climate risk on agriculture, Sustainability 17 (16) (2025) 7566. doi: 10.3390/su17167566 . [3] A. Morales, F . V illalobos, Using machine learning for crop yield prediction in the past or the future, Frontiers in Plant Science 14 (2023). doi:10.3389/fpls.2023.1128388 . [4] C. He, J. Song, H. Xu, Optical and sar data fusion based on transformer for rice identification: A comparativ e analysis from early to late integration, Agriculture 15 (7) (2025) 706. doi:10.3390/agriculture15070706 . [5] A. Nit , u, et al., Ndvi and beyond: V egetation indices for crop recognition (2025). doi:10.3390/s250203817 . [6] H. Md-T ahir , et al., Localized crop classification by ndvi time series, AgriEngineering 6 (3) (2024) 2429–2444. doi:10.3390/agriengineering6030142 . [7] J. W ang, et al., Integration of remote sensing and ml for precision agriculture, Agronomy 14 (9) (2024) 1975. doi:10.3390/agronomy14091975 . [8] X. Geng, J. He, V . Grima, Y . Jiang, M. T etreau, S. Crittenden, S. Kiley , A. J. V andenBygaart, J. V anrobaeys, 100 m soil landscape grids of canada, Scientific Data 12 (2025) 1178. doi:10.1038/s41597- 025- 05460- 4 . [9] A. Y ewle, L. Mirzaye v a, O. Karaku ¸ s, Multi-modal data fusion and deep ensemble learning for accurate crop yield prediction, arXiv preprint arXi v:2502.06062 (2025). [10] N. Keerthana, S. Shaik, B. Dodamani, Identifying rice crop flooding patterns using sentinel-1 sar , Journal of the Indian Society of Remote Sensing 50 (6) (2022) 1569–1584. doi:10.1007/s12524- 022- 01553- 4 . [11] A. Alsaber, A. Satpathi, M. Alsabah, P . Setiya, Optimizing potato yield predictions in uttar pradesh, india: a comparati ve analysis of machine learning models, Scientific Reports 15 (2025) 26897. doi:10.1038/ s41598- 025- 12719- 8 . 21 [12] H. Liang, et al., Ev aluating soybean drought tolerance using uav multimodal data, Remote Sensing 16 (11) (2024) 2043. doi:10.3390/rs16112043 . [13] N. JPL, Nasa shuttle radar topography mission global 1 arc second v003, NASA EOSDIS Land Processes D AAC (2013). doi:10.5067/MEaSUREs/SRTM/SRTMGL1.003 . [14] A. Joshi, et al., Explainable bi-lstm model for winter wheat yield prediction, Frontiers in Plant Science 15 (2025) 1491493. doi:10.3389/fpls.2024.1491493 . [15] S. Das, et al., Machine learning model ensemble for predicting sugarcane yield, Remote Sensing Applications: Society and En vironment 30 (2023) 100962. doi:10.1016/j.rsase.2023.100962 . Appendix A. F eatur e Mapping Resources and Documentation The UniCrop pipeline uses a configuration-driven feature mapping file that specifies the en vironmental v ariables to be retrie ved from each data source. This mapping defines the dataset identifiers, API parameters, temporal resolutions, and any required deriv ations (e.g., ve getation indices, radiation interactions). The mapping file enables users to e xtend UniCrop to new crops or re gions by modifying configuration entries rather than pipeline code. T o ensure correctness and reproducibility , the mapping was constructed using the o ffi cial technical documentation for all data sources integrated into UniCrop. These resources define band names, variable identifiers, quality flags, spatial resolutions, and recommended preprocessing methods. The most relev ant documentation sources are listed below . Appendix A.1. Earth Observation and Remote Sensing Documentation • Sentinel-1 (SAR): Copernicus Data Space Ecosystem documentation for GRD products, polarisation modes, and radiometric calibration procedures. • Sentinel-2 (Optical): ESA MSI User Guides, band specifications, Scene Classification Layer (SCL), and Lev el- 2A product processing parameters. • MODIS Products (MOD13, MOD15, MOD16, MOD17): NASA LP DAA C Algorithm Theoretical Basis Documents (A TBDs) describing ND VI / EVI, LAI / fP AR, ev apotranspiration, and producti vity algorithms. Appendix A.2. Climate and Agr o-Climatology Documentation • ERA5-Land: Copernicus Climate Data Store documentation for hourly land-surface variables, aggregation methods, and variable definitions. • NASA PO WER: Agroclimatology Data Dictionary , API reference, and variable descriptions for temperature, humidity , radiation, and wind. 22 Appendix A.3. Soil and T opographic Data Documentation • SoilGrids: ISRIC documentation on soil depth layers, soil property definitions, and machine-learning based spatial predictions. • SR TM: NASA / USGS documentation for ele vation data, v oid-filling, and terrain deriv ati ves (slope, aspect, hillshade). Appendix B. Supplementary Material and GitHub Release T o support transparency , reproducibility , and reuse, the UniCrop softw are frame work is released as an open-source repository alongside this study . The GitHub version of UniCrop demonstrates the full pipeline using a publicly av ailable maize yield dataset from Spain , rather than the proprietary rice dataset used in the main case study . All files required to reproduce the GitHub e xample are included in the repository . These comprise the sample yield dataset, location-to-coordinate mapping files, configuration scripts, feature-mapping tables, and the full data acquisition and modelling code. The example dataset is deri ved from the W ageningen Uni versity & Research (WUR) AI sample data repository and is temporally subsampled to include harvest years from 2010 onwards. The GitHub release includes: • A complete example dataset with latitude–longitude coordinates and annual maize yield v alues • The feature-mapping file ( unicrop_feature_mapping.csv ) specifying data sources, v ariables, API parame- ters, and deriv ation rules • Configuration files ( config.py , paths.py ) required to initialise data downloads and modelling • Automated pipelines for data acquisition from NASA PO WER, Sentinel-2, MODIS, ERA5, SoilGrids and SR TM • A modular modelling and benchmarking workflo w that can be rerun without repeating data downloads This open example is intended as a methodological demonstration of UniCrop rather than a claim of optimal yield prediction performance for maize in Spain. Researchers are encouraged to adapt the configuration files, feature mappings, and data sources to other c rops, regions, and temporal settings. The separation between data downloading and modelling allows UniCrop to be readily extended while maintaining reproducibility and clarity of experimental design. The full UniCrop repository , documentation, and sample data are av ailable at: [https://github.com/CoDIS- Lab/UniCrop] 23 Please note that the rice yield dataset used in the primary case study of this paper cannot be publicly shared due to data usage restrictions. The dataset is subject to proprietary rights held by Ernst & Y oung (E&Y) and was made av ailable to the authors under specific research and confidentiality agreements. As a result, redistribution of the raw rice yield data or associated parcel-lev el information is not permitted. Researchers interested in accessing the rice dataset used in this study should contact Ernst & Y oung directly to enquire about data av ailability and applicable licensing conditions. T o ensure transparency and reproducibility of the UniCrop methodology despite these restrictions, we provide a fully open and self-contained GitHub example using a public maize yield dataset, along with all configuration files, feature mappings, and pipeline code required to reproduce the software workflo w . 24
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment