Hear the Heartbeat in Phases: Physiologically Grounded Phase-Aware ECG Biometrics
Electrocardiography (ECG) is adopted for identity authentication in wearable devices due to its individualspecific characteristics and inherent liveness. However, existing methods often treat heartbeats as homogeneous signals, overlooking the phase-s…
Authors: Jintao Huang, Lu Leng, Yi Zhang
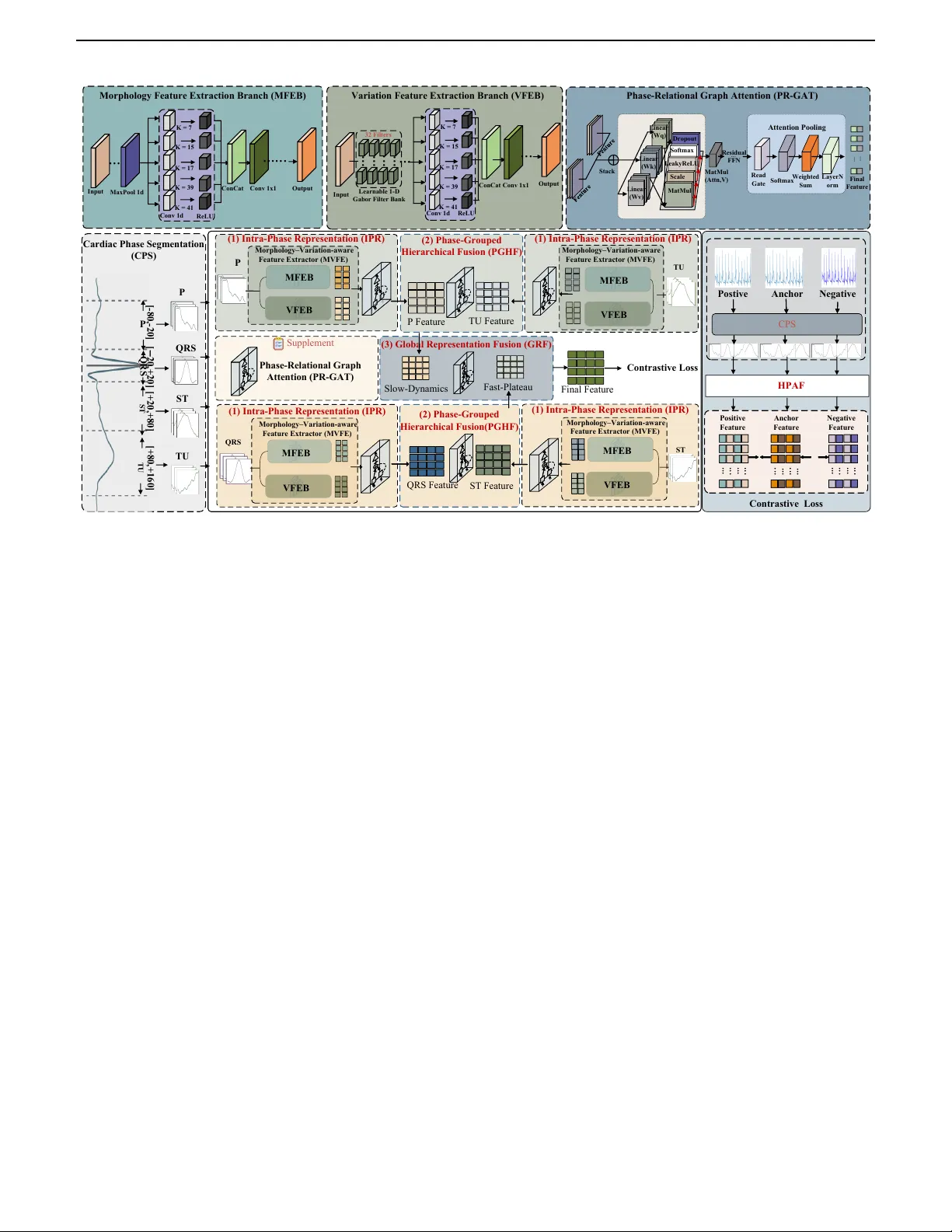
IEEE TRANSACTIONS AND JOURNALS TEMPLA TE 1 Hear the Heartb eat in Phases: Ph ysiologically Grounded Phase-A w are ECG Biometrics Jin tao Huang, Lu Leng, Member, IEEE, Yi Zhang, Senior Member, IEEE, and Ziyuan Y ang, Member, IEEE Abstract— Electro cardiograph y (ECG) is adopted for iden- tit y authen tication in wearable devices due to its individual- sp ecic characteristics and inheren t liveness. How ev er, existing metho ds often treat heartb eats as homogeneous signals, o ver- lo oking the phase-sp ecic characteristics within the cardiac cycle. T o address this, we propose a Hierarchical Phase- A ware F usion (HP AF) framework that explicitly av oids cross- feature en tanglement through a three-stage design. In the rst stage, In tra-Phase Represen tation (IPR) independently extracts representations for each cardiac phase, ensuring that phase-sp ecic morphological and v ariation cues are preserved without interference from other phases. In the second stage, Phase-Group ed Hierarchical F usion (PGHF) aggregates phys- iologically related phases in a structured manner, enabling reliable in tegration of complemen tary phase information. In the nal stage, Global Represen tation F usion (GRF) further com bines the group ed representations and adaptiv ely balances their contributions to pro duce a unied and discriminative iden tity represen tation. Moreov er, considering ECG signals are con tinuously acquired, m ultiple heartb eats can b e collected for eac h individual. W e prop ose a Heartbeat-A w are Multi- protot yp e (HAM) enrollment strategy , which constructs a m ulti-prototype gallery template set to reduce the impact of heartb eat-sp ecic noise and v ariabilit y . Extensive exp eriments on three public datasets demonstrate that HP AF achiev es state-of-the-art results in the comparison with other metho ds under b oth closed and op en-set settings. Index T erms— ECG Biometrics, Multi-View Represen tation, Graph Neural Netw orks, Identit y V erication. I . I N T R O D U C T I O N Electro cardiograph y (ECG) has long pla y ed a funda- men tal role in clinical diagnostics and, with the rise of w earable devices, is increasingly recognized as a physio- logically grounded biometric signal. Compared to other This work has b een submitted to the IEEE for p ossible publication. Copyrigh t ma y b e transferred without notice, after which this version may no longer be accessible. This work was supported in part by the National Natural Science F oundation of China under Grants 62466038; in part by the Jiangxi Provincial Key Lab oratory of Image Pro cessing and Pattern Recog- nition under Grants 2024SSY03111 and ET202404437; and in part by the High Performance Computing Service of Information Center at Nanchang Hangkong Universit y . (Corresponding author: Lu Leng and Ziyuan Y ang) Jintao Huang is with the Jiangxi Provincial Key Lab oratory of Image Processing and Pattern Recognition, Nanchang Hangkong Universit y , 330063, China (e-mail: 2520083500018@stu.nc hu.edu.cn). Lu Leng is with the Jiangxi Provincial Key Lab oratory of Image Processing and Pattern Recognition, Nanchang Hangkong Univer- sity , 330063, China (email: leng@nch u.edu.cn). Yi Zhang and Ziyuan Y ang are with the Sc ho ol of Cyb er Sci- ence and Engineering, Sich uan Universit y , Chengdu 610065, China (yzhang@scu.edu.cn and cziyuany ang@gmail.com). mo dalities such as face or ngerprint, ECG acquisition originates directly from cardiac activity and thus inher- en tly provides liveness detection [1]–[3]. These charac- teristics make ECG a promising mo dality for priv acy- preserving and con tin uous authen tication [4]–[7]. Recen tly , a gro wing b o dy of research has in vesti- gated the use of ECG for identit y recognition. These metho ds hav e progressed from handcrafted feature-based approac hes to deep learning-based ones [8], [9]. Most existing approac hes use a single mo del to enco de dierent cardiac phases. F or instance, Ibtehaz et al. [10] employ ed a m ulti-resolution Siamese net work on whole b eats for ECG authen tication. Chu et al. [11] utilized a parallel m ulti-scale ResNet with cen ter and margin losses to extract discriminative features. W ang et al. [12] prop osed a self-sup ervised conv olutional neural netw ork (CNN) framew ork that compares dierent ECG segmen tation strategies. Allam et al. [13] adopted a parallel CNN-LSTM arc hitecture to learn representations from R-p eak-aligned b eats. Although promising in ideal settings, these meth- o ds uniformly treat heartb eats con taining the P w av e, QRS complex, and ST/T(U) segments as a homogeneous sequence pro cessed b y a shared enco der. These metho ds rely on the basic assumption that dieren t phases follow the homogeneous signal patterns, ignoring their inherent structural and physiological heterogeneity . This leads to po or discriminativ e feature learning and sub optimal p erformance in the iden tication task. Sp ecically , the ECG exhibits distinct wa veform phases that originate from dierent electrophysiological activities of the heart, including the P wa ve, QRS complex, and T wa ve. Due to their distinct electrophysiological mecha- nisms, these phases exhibit markedly dierent morpholog- ical and spectral characteristics, as illustrated in Fig. 1. The P wa ve reects atrial dep olarization with relatively smo oth and lo w-amplitude morphology , while the QRS complex corresp onds to rapid ven tricular dep olarization and presents the highest energy and steepest transitions due to the large my o cardial mass and fast conduction v elo city . The T w av e represents v entricular repolarization, c haracterized b y slow er electrical recov ery and smoother w a v eform patterns. Hence, treating these physiologically distinct phases as homogeneous signal patterns would sev erely limit the mo del’s ability to capture stable and discriminativ e iden tit y features. Inspired by the intrinsic physiological structure of car- diac cycles, w e prop ose a multi-gran ularity cardiac-phase 2 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE Gr aph A tt e n ti on (c ) Our E C G - b a se d b io m e trics Fra m e wo rk s He ar t - r ate v ar iabi li ty ( HR V ) L e ad / de v ice s hi ft M oti on nois e / dr if t : He ar t - r ate v ar iabi li ty ( H R V ) : R hy thm - r e lat e d : C ondit ion - r e lat e d : I de nti ty - r e lat e d (a ) Da ta Ac q u isit io n (b ) Ba sli n e M e th o d E nc onde r Gr anph A t te nt ion I ntr al F e atur e P / Q R S/ T /U P QR S TU ST T ime banc h F r e que nc y br anc h P QR S TU ST T ime banc h F r e que nc y br anc h P QR S TU ST T ime ban c h F r e que nc y br anc h T ime ban c h F r e que nc y br anc h D is e nt an glem e nt D is e nt an glem e nt F i nal F e atur e P QR S (c ) Cro ss Fu sio n Fra m e wo rk I ntr al F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on I ntr al F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on S low F u s ion (d ) I n fe r e n c e Gr aph A tt e n ti on I ntr al F e atur e Gr aph A tt e n ti on I ntr al F e atur e Gr aph A tt e n ti on F as t F u s ion Gr aph A tt e n ti on Gr aph A tt e n ti on Gr aph A tt e n ti on (a ) Da ta Ac q u isit io n (b ) Va n il la E C G - b a se d b io m e trics Fra m e wo rk s E nc onde r P QR S ST T ime banc h F r e que nc y br anc h P / Q R S/ T /U He ar t - r ate v ar iabi li ty ( HR V ) L e ad / de v ice s hi ft M oti on / nois e / dr if t : He ar t - r ate v ar iabi li ty ( H R V ) : R hy thm - r e lat e d : C ondit ion - r e lat e d : I de nti ty - r e lat e d (a ) Da ta Ac q u isit io n TU TU TU T ime banc h T ime banc h T ime banc h T ime banc h T ime banc h T ime banc h TU T ime banc h T ime banc h Gr aph A tt e n ti on QR S F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on ST F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on S low F u s ion P F e atur e Gr aph A tt e n ti on T U F e atur e Gr aph A tt e n ti on F as t F u s ion Gr aph A tt e n ti on TU QR S P T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h ST T ime banc h F r e que nc y br anc h Slow F e atur e F as t F e atur e Gr aph A tt e n ti on F i nal F e at ur e F in al F u s ion C r o s s F us i o n F r a m e w o r k Tra i n a b le M o d e l Gr aph A tt e n ti on QR S F e atur e Gr aph A tt e n ti on ST F e atur e Gr aph A tt e n ti on S low F u s ion P F e atur e Gr aph A tt e n ti on T U F e atur e Gr aph A tt e n ti on F as t F u s ion Gr aph A tt e n ti on TU QR S P T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h ST T ime banc h F r e que nc y br anc h Slow F e atur e F as t F e atur e Gr aph A tt e n ti on F i nal F e at ur e F in al F u s ion C r o s s F us i o n F r a m e w o r k Tra i n a b le M o d e l ST S i n u s N o d e At ri a l Mu sc l e A - V N o d e C o m m o n Bra n c h e s Bu n d l e Bu n d l e Ve n t ri c u l a r Mu sc l e Q R S P ST TU Pu rki n j e Fi b e rs Q R S P ST TU C a rdia c c onduc ti on pa t hw a y to EC G Gr aph A tt e n ti on (c ) Our E C G - b a se d b io m e trics Fra m e wo rk s He ar t - r ate v ar iabi li ty ( HR V ) L e ad / de v ice s hi ft M oti on nois e / dr if t : He ar t - r ate v ar iabi li ty ( H R V ) : R hy thm - r e lat e d : C ondit ion - r e lat e d : I de nti ty - r e lat e d (a ) Da ta Ac q u isit io n (b ) Ba sli n e M e th o d E nc onde r Gr anph A t te nt ion I ntr al F e atur e P / Q R S/ T /U P QR S TU ST T ime banc h F r e que nc y br anc h P QR S TU ST T ime banc h F r e que nc y br anc h P QR S TU ST T ime ban c h F r e que nc y br anc h T ime ban c h F r e que nc y br anc h D is e nt an glem e nt D is e nt an glem e nt F i nal F e atur e P QR S (c ) Cro ss Fu sio n Fra m e wo rk I ntr al F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on I ntr al F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on S low F u s ion (d ) I n fe r e n c e Gr aph A tt e n ti on I ntr al F e atur e Gr aph A tt e n ti on I ntr al F e atur e Gr aph A tt e n ti on F as t F u s ion Gr aph A tt e n ti on Gr aph A tt e n ti on Gr aph A tt e n ti on (a ) Da ta Ac q u isit io n (b ) Va n il la E C G - b a se d b io m e trics Fra m e wo rk s E nc onde r P QR S ST T ime banc h F r e que nc y br anc h P / Q R S/ T /U He ar t - r ate v ar iabi li ty ( HR V ) L e ad / de v ice s hi ft M oti on / nois e / dr if t : He ar t - r ate v ar iabi li ty ( H R V ) : R hy thm - r e lat e d : C ondit ion - r e lat e d : I de nti ty - r e lat e d (a ) Da ta Ac q u isit io n TU TU TU T ime banc h T ime banc h T ime banc h T ime banc h T ime banc h T ime banc h TU T ime banc h T ime banc h Gr aph A tt e n ti on QR S F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on ST F e atur e Gr aph A tt e n ti on Gr aph A tt e n ti on S low F u s ion P F e atur e Gr aph A tt e n ti on T U F e atur e Gr aph A tt e n ti on F as t F u s ion Gr aph A tt e n ti on TU QR S P T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h ST T ime banc h F r e que nc y br anc h Slow F e atur e F as t F e atur e Gr aph A tt e n ti on F i nal F e at ur e F in al F u s ion C r o s s F us i o n F r a m e w o r k Tra i n a b le M o d e l Gr aph A tt e n ti on QR S F e atur e Gr aph A tt e n ti on ST F e atur e Gr aph A tt e n ti on S low F u s ion P F e atur e Gr aph A tt e n ti on T U F e atur e Gr aph A tt e n ti on F as t F u s ion Gr aph A tt e n ti on TU QR S P T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h T ime banc h F r e que nc y br anc h ST T ime banc h F r e que nc y br anc h Slow F e atur e F as t F e atur e Gr aph A tt e n ti on F i nal F e at ur e F in al F u s ion C r o s s F us i o n F r a m e w o r k Tra i n a b le M o d e l ST S i n u s N o d e At ri a l Mu sc l e A - V N o d e C o m m o n Bra n c h e s Bu n d l e Bu n d l e Ve n t ri c u l a r Mu sc l e Q R S P ST TU Pu rki n j e Fi b e rs Q R S P ST TU C a rdia c c onduc ti on pa t hw a y to EC G Fig. 1: An illustration of the relationship b et w een the dieren t phases of the cardiac cycle and the corresponding ECG signal. represen tation framew ork that phase-aw are mo dels the heterogeneous phase patterns. Sp ecically , at rst, we prop ose a Cardiac Phase Segmen tation (CPS) metho d to segmen t each heartb eat in to four distinct phases, including P , QRS, ST, and T/U, whic h is anc hored at the R-p eak. In this w ay , our metho d obtains physiologically meaningful temp oral dynamics phases and preserves phase-sp ecic morphological c haracteristics, providing a reliable basis for subsequen t phase-sp ecic mo deling. As previously discussed, employing a single shared enco der ignores the physiological heterogeneity across phases, hindering stable and discriminative representation extraction. T o address this issue, we prop ose a Hierarc hical Phase-A w are F usion strategy (HP AF), including the Intra- Phase Represen tation (IPR), Phase-Group ed Hierarchical F usion (PGHF), and Global Represen tation F usion (GRF) mo dules. In IPR, to av oid inter-phase feature entan- glemen t, we design phase-sp ecic Morphology–V ariation- a w are F eature Extractors (MVFEs), where each phase is pro cessed by an indep endent MVFE to extract discrimi- nativ e features. MVFE consists of a standard conv olution branc h for morphology extraction and a learnable Gab or branc h for v ariation mo deling. The standard conv olution branc h can eectively capture the w av eform morphology and temp oral structures, while the learnable Gab or lters are capable of mo deling ne-grained lo cal v ariations such as subtle oscillations and abrupt transitions. Then, in PGHF, w e divide the phases into t w o phys- iologically coheren t groups and p erform phase-group ed hierarc hical fusion separately: one PGHF mo dule for the P and T/U features, and another for the QRS and ST features. Sp ecically , the P and T/U features are group ed as they are b oth related to rep olarization and exhibit smo oth, low-amplitude wa veforms. In con trast, the QRS and ST features are group ed due to their asso ciation with ven tricular dep olarization and early rep olarization, c haracterized by higher amplitude, sharp er transitions, and more complex morphologies. Then, we design a cross-phase gated fuser that selectively couples phases through gated calibration rather than directly fusing them, enabling stable calibration for dieren t phase groups. Finally , the tw o features fused by the PGHF mo dules are fed into the GRF mo dule, where a cross-phase gated fuser is applied again to adaptively fuse them into the nal representation. Concretely , GRF rst projects eac h fused feature into a shared scoring space and computes a scalar attention-lik e gate that measures the relative imp ortance of the QRS–ST group versus the ST–T/U group. This gate then controls a weigh ted mixture of the tw o fused features, follow ed by a linear pro jection to obtain the nal global feature. In this wa y , the mo del can dynamically decide whether fast transient cues (QRS–ST) or slow rep olarization cues (ST–T/U) should dominate under dieren t noise conditions and rh ythm patterns. T o optimize our netw ork and extract discriminative fea- tures, we introduce a contrastiv e loss. This loss constrains global fused em b eddings in the feature space to encourage balancing the phase-sp ecic identit y cues and cross-phase consistency . In the implementation stage, since ECG signals are contin uously acquired, multiple heartb eats can b e collected for each individual. W e prop ose a Heartb eat- A w are Multi-prototype (HAM) enrollment strategy . Un- lik e previous w orks that represen t each individual with a single feature vector, HAM represents eac h sub ject using a set of prototypes. In this wa y , our metho d can obtain a gallery set that improv es the matching reliability and the p erformance through reducing the impact of heartb eat- sp ecic noise and v ariability . Our main contributions can b e summarized as follo ws: • W e prop ose HP AF, a physiologically grounded phase- a w are ECG iden tication framew ork that a v oids cross-phase feature entanglemen t through a three- stage design. • W e design a dual-branch MVFE to capture the morphology and v ariation features, and in tro duce PR- GA T to mo del their interactions for reliabilit y-a w are renemen t explicitly . • W e prop ose a nov el enrollmen t strategy HAM, which lev erages multiple heartb eats within ECG signals to construct a robust m ulti-protot yp e gallery template for eac h individual. • Extensiv e experiments on the public datasets v alidate the eectiveness through comprehensive exp eriments under closed-set and op en-set proto cols. I I . R E L A T E D W O R K S A. Representation Learning for ECG Iden tication 1) Fiducial-Based F eature Learning : Fiducial-based ECG iden tication metho ds rely on the precise detection of salien t landmarks (P wa ve, QRS complex, T w a ve) and construct time-domain features. F or example, Biel et al. [2] prop osed to extract amplitudes, durations and inter-w av e in terv als from 12-lead resting ECG and use a SIMCA classier for subject v erication. Israel et al. [3] derive JINT AO HUANG et al.: HEAR THE HEAR TBEA T IN PHASES: PHYSIOLOGICALL Y GROUNDED PHASE-A W ARE ECG BIOMETRICS 3 normalized temp oral features from carefully annotated P/R/T p oints on high-rate ECG and com bine LDA with ma jority voting to achiev e robust recognition under v arying electro de lo cations and anxiety states. Accurate delineation is therefore a key prerequisite for ducial pip elines; for instance, Martínez et al. [14] developed a w a v elet-transform-based ECG delineator to lo calize QRS complexes and delineate P/QRS/T p eaks and b oundaries on standard databases. These landmark-based sc hemes are ph ysically interpretable but highly sensitive to b oundary detection errors, motion and EMG noise, as summarized b y F ratini et al. [15], whereas W ang et al. [16]. w av elet– based time–frequency represen tation with stack ed sparse auto enco ders disp enses with explicit ducial detection, underscoring the limitations of traditional ducial ap- proac hes in noisy and cross-condition scenarios. 2) Non-Fiducial Represen tation Learning : Non-ducial feature extraction bypasses explicit detection of P/QRS/T landmarks and instead derives descriptors from the whole ECG or its transformed/reconstructed domains. Repre- sen tativ e metho ds include Jung et al. [17], who seg- men t preprocessed single-lead ECG into xed windows and compute auto correlation- and DCT-based features with a windo w-remov al strategy and classical classiers; F ang et al. [18], who reconstruct ECG phase space and quantify similarity/dissimilarit y b etw een p ortraits; Sriv astv a et al. [19], who com bine auto correlation with DCT/DFT/WHT transforms and PCA/LDA; and Gosh- v arp our et al. [20], who use Matching Pursuit–based statistical and nonlinear features with PNN/kNN and feature selection. Gutta et al. [21], who reformulate multi- class ECG identication as one-vs-all multi-task learning and use a feature-scaled probabilistic kernel classier with sparse–lo w-rank weigh t decomp osition for joint feature selection and classication.These approac hes av oid errors caused by inaccurate ducial detection, but typically pro- duce high-dimensional representations that require feature selection or dimensionalit y reduction. 3) Deep learning–based feature extraction : In recen t y ears, deep learning has b ecome the mainstream solution for ECG-based identit y recognition, enabling end-to-end learning of discriminative features. Most approaches rely on time-domain CNNs op erating on R-p eak–anchored b eats or rhythm segments, while frequency information is added via xed time–frequency transforms (STFT, CWT) or handcrafted Gab or descriptors fused a p osteriori, whic h can introduce time–frequency crosstalk and mak e p erformance sensitive to window length and sampling rate. Representativ e examples include single-b eat time– frequency mo deling around the R p eak [22], 2D CNNs on STFT sp ectrograms for rhythm classication [23], multi- scale residual CNNs and Siamese arc hitectures for iden tit y v erication [10], [11], [24], and LSTM-based mo dels that capture long-range temp oral dep endencies across multiple b eats. B. Atten tion mec hanisms for ECG biometrics Recen t ECG biometric systems increasingly exploit atten tion to emphasize informative temp oral patterns and heartb eats. Chee et al. [25] employ a T ransformer-style self-atten tion enco der on ECG sequence pairs to learn exible identit y embeddings that supp ort both identi- cation and verication across multiple databases. Jyotishi et al. [26] design a hierarchical LSTM with an in ternal atten tion mo dule that assigns larger weigh ts to ECG complexes carrying richer biometric information, outper- forming traditional ducial and non-ducial baselines on on-b o dy and o-b o dy datasets. At the con volutional lev el, Hammad et al. [27] integrate an attention blo ck into a residual CNN (ResNet-A ttention) so that discriminativ e ECG segments are highlighted in feature maps, while Pan et al. [28] introduce a m ulti-branc h domain-adaptive fusion net w ork where a weigh ted adaptive attention mechanism reinforces session-robust heartbeat features on ECGID, PTB, CYBHi, and Heartprin t. Bey ond purely temp oral attention, several works apply atten tion to channel, feature, or graph spaces. Sun et al. [29] prop ose RandSaD, a dual-path residual mo del with split attention that selectively aggregates m ulti-scale c hannel features from wa v elet-denoised ECG windows and reac hes ≈ 99 . 6% identication accuracy . W ang et al. [30] fuse 1D and 2D ECG represen tations within a collab orativ e em b edding framework and use dimensional atten tion to up-weigh t discriminative laten t features. Ma et al. [31] present OR GNNFL, whic h couples a multi- feature atten tion module with graph neural net work fusion to adaptively aggregate top ology-aw are ECG features under distribution shifts. Al Al et al. [32]construct a 12-lead ECG graph whose edges are w eigh ted by mutual- information-based “atten tion-lik e” scores. I I I . P RO P O S E D M E T H O D A. Overview In this pap er, we prop ose a nov el multi-gran ularity cardiac-phase representation framework for ECG iden ti- cation, and the ov erview of our prop osed metho d can b e found in Figure 2. Employing a single shared enco der to extract features across dierent cardiac phases w ould neglect the physiological heterogeneity , which ma y hinder the extraction of stable and discriminative features. T o mitigate this issue, we prop ose the Cardiac Phase Seg- men tation (CPS) metho d to segment the ECG in to four cardiac phases to av oid phase distortion in the subsequent feature extraction. Then, w e propose a Hierarchical Phase- A w are F usion (HP AF) strategy . HP AF consists of Intra- Phase Represen tation (IPR), Phase-Group ed Hierarchical F usion (PGHF), and Global Represen tation F usion (GRF) mo dules. Sp ecically , in IPR, each phase is pro cessed by a phase- sp ecic Morphology–V ariation-aw are F eature Extractor (MVFE) to av oid inter-phase feature entanglemen t. Each MVFE contains a conv olution branch for wa veform mor- phology and a learnable Gab or branc h for ne-grained 4 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE Mutil - Cscale Front - end Key path ... Ti m e - do m a i n i nfo rm a ti o n F reque ncy - do m a i n i nfo rm a ti o n 1D - Gabor 1D - CNN Features Features ... ... ... Slow - Phase Cross - Phase Fusion Final Feature ... ... ... ... ... ... ... ... ... ... ... ... Fast - Phase Cross - Ph ase Fusion Intra - Phase Dual - Encoder Fusion En cond e r A nchor Postive N egat i ve ... ... ... ... ... ... DP DN LOS S = (D P ,D N ) Beat - level, phase - aligned tri p let construct ion End - to - E nd co ntra s ti v e tr a i ni n g o n fus ed em beddi ng s ... ... ... ... ... ... F i na l Featur e F i na l Featur e F i na l Featur e Graph - based Late Fusion I nput ... ... M a x P o o l 1d K = 7 K = 15 K = 17 K = 39 K = 41 R eLU C o n C a t C o n v 1 d Parallel Mutil - scale Con volu ti on s Out put N o r m a l i z a ti o n & N o n l i nea ri ty Li nea r (Wq ) Li nea r (Wk ) Li nea r (Wv ) M a tM ul Sca l e So ftm a x D ro po u t Linear R ea d Ga te Stack Lea ky R eLU M a tM ul (A ttn,V ) So ftm a x We i g hted Sum Final Feature Attention Pooling Multi - Head Gr ap h Attention ... R es i dua l F F N Gr aph Fusion E ncon d e r Within - batch H ar d - Example Mining ★ P osit ive N e ga ti ve ★ ha r de st ne ga ti ve ... P QR S ... Ti m e - do m a i n i nfo rm a ti o n F reque ncy - do m a i n i nfo rm a ti o n 1D - CNN Features Features ... ... ... ... Intra - Phase Dual - Encoder Fusion ... ... 1D - Gabor ... ... ... ... ... R - cent ered Tempo ral Phase Windowing P QR S ST TU ... TU ST La y erN o rm P Fea t ure TU Feature TU QRS Feat ure ST Fea ture Slow - Dynamics Final Fea ture P (1) Intra - Phase Repre s e n tati on (IPR) P QRS Cardiac Phas e Segme n t ation (CPS) HPAF A nchor Postive N egat i ve DP DN Contrast i ve L oss A ncho r F ea ture P o s i ti v e F ea ture N eg a ti v e F ea ture End - to - E nd co ntra s ti v e tr a i ni n g o n fus ed em beddi ng s Morphology Fe atu r e Extrac ti on Branch ( MFEB ) In p u t ... M axPool 1d K = 7 K = 15 K = 17 K = 39 K = 41 Re L U ConCat Conv 1d Ou t p u t Nor m ali z ation & Nonl i n e ar ity L i n e ar (Wq) L i n e ar (Wk) L i n e ar (Wv) M at M u l S c ale S oft m ax Dr op ou t S tac k L e ak yRe L U M at M u l (A tt n ,V) Fin al Fe at u r e ... Re si d u al FFN ... QR S ST ST TU (1) Intra - Phase Re p r ese ntation (IPR) (3) Global Repre s e n tati on Fus i on (GRF) Tri - Stage DeCoCo Graph Fusion for ECG Biometrics ST P QRS TU [+ 80, + 160] [ - 80, - 20] [ − 20, + 20] [+ 20, + 80] S l o w – S l o w (P T/ U ) : low – m i d (1 – 7 H z ), l o w s l o p e; g r a p h co - co u p l ed cue s . Fast → Plat e au: Q RS → ST — Pair f as t QR S ( 10 – 40 Hz ) w i th t he ST ( ~0 slope ) plat e au Introduction Re ad Gate S oft m ax We igh te d S u m A ttent i o n Poo l i ng L aye r N or m ... In p u t ... ... ... ... L e ar n ab l e 1 - D Gab or Fil t e r B an k 32 F il te r s K = 7 K = 15 K = 17 K = 39 K = 41 Re L U Conv 1d Conv 1x1 ... ... Conv 1x1 ... ConCat Ou t p u t ... Variat i on Feature Extrac t ion Branch (VFEB) ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ... CP S Phase - Relat i on al G raph A tte n ti on (PR - GAT) Phase - Relat i on al G raph Attention (PR - GAT) Supplement Fast - Plateau ha r de st posi t ive ★ An ch o r N eg ativ e ★ H a r de st N e ga ti ve H a r de st P osit ive Contrast i ve Los s P o sit iv e (2) Ph as e - Grouped Hierarchic al F u sion ( PG H F) (2) Ph as e - Grouped Hierarchic al F u sion (PGHF) (1) Intra - Phase Re p r ese ntation (IPR) (1) Intra - Phase Re p r ese ntation (IPR) (1) Intra - Phase Re p r ese ntation (IPR) (1) Intra - Phase Re p r ese ntation (IPR) REB VMB M o rpho l o g y – V a r i a ti o n - a w a re F ea ture Ex tra cto r (M V F E) M o rpho l o g y – V a r i a ti o n - a w a re F ea ture Ex tra cto r (M V F E) M o rpho l o g y – V a r i a ti o n - a w a re F ea ture Ex tra cto r (M V F E) M o rpho l o g y – V a r i a ti o n - a w a re F ea ture Ex tra cto r (M V F E) MFEB VFEB MFEB VFEB VFEB MFEB MFEB VFEB VFEB Fig. 2: W e presen t the prop osed HP AF framew ork for phase-a w are ECG iden tication. CPS detects R-p eaks and partitions each heartb eat into four phase-aligned segments, namely P , QRS, ST, and T/U. Eac h phase is enco ded by MVFE with MFEB and VFEB, and fused by PR-GA T to form phase representations, which PGHF and GRF then aggregate into a single b eat-level identit y em bedding. The encoder is trained with the contrastiv e loss to increase in ter-sub ject separability . lo cal v ariations. PGHF then groups the phases in to P–T/U and QRS–ST, and p erforms hierarc hical fusion within eac h group via a cross-phase gated fuser. GRF then aggregates the tw o group-level representations into a unied b eat- lev el presen tation as the nal feature. Moreov er, a con- trastiv e loss is further introduced to enhance cross-phase consistency while preven ting ov er-reliance on any single phase. These comp onen ts are detailed in the follo wing. B. Hierarchical Phase-A w are F usion (HP AF) 1) Cardiac Phase Segmentation (CPS) : T o alleviate the heterogeneit y across dierent ECG phases, we propose the CPS metho d for explicit phase segmentation, which eectiv ely prev en ts feature entanglemen t among dierent phases during subsequent feature extraction. Sp ecically , CPS treats the R peak as the anchor to explicitly partition eac h heartb eat into dieren t physiological phases, instead of the conv entional av erage segmen ting of the entire ECG signal. Each heartb eat is divided in to four stages, including atrial dep olarization, ven tricular dep olarization and early plateau, and v en tricular rep olarization, corresponding to the P w a v e, QRS complex, ST segmen t, and T/U w av e. Sp ecically , CPS rst detects the R peak in the prepro cessed ECG signal and then delineates four time windo ws around each R p eak, whic h appro ximately cor- resp ond to the P , QRS, ST, and T/U phases based on predened relative osets and durations. The four xed-length segments are extracted by selecting samples with predened oset windows, which are measured in samples relative to the R p eak: [–80, –20] for P , [–20, +20] for QRS, [+20, +80] for ST, and [+80, +160] for TU. In this wa y , CPS pro duces four-phase segmen ts with clear physiological meaning and consistent temporal scale, whic h can eectively preven t feature entanglemen t during subsequen t feature extraction. each segmented phase is indep enden tly encoded using a dedicated phase-sp ecic MVFE. 2) Intra-Phase Represen tation (IPR) : F ollo wing CPS, eac h segmented phase is indep endently enco ded by a phase-sp ecic MVFE. While commonly used con v olutional lters eectiv ely capture coarse morphological informa- tion, they often fail to preserv e subtle v ariations in ECG signals. Gab or lters are eective at capturing subtle v ariations in ECG signals [?], as the y emphasize lo cal- ized w av eform v ariations within short temporal regions. Motiv ated by these complemen tary prop erties, MVFE is designed as a dual-branch architecture, including a CNN- based Morphology F eature Extraction Branc h (MFEB) and a learnable Gab or lter-based V ariation F eature Ex- traction Branc h (VFEB). Moreo ver, to model the relation- ships among morphology and v ariation represen tations, w e in tro duce a Phase-Relational Graph A tten tion (PR-GA T) mo dule. By treating the morphology and v ariation repre- sen tations as graph nodes, PR-GA T explicitly captures JINT AO HUANG et al.: HEAR THE HEAR TBEA T IN PHASES: PHYSIOLOGICALL Y GROUNDED PHASE-A W ARE ECG BIOMETRICS 5 their relational dep endencies to obtain the comprehensive and discriminativ e feature. The VFEB is tailored to extract ne-grained v ariations. In contrast to conv entional Gab or lters with manually sp ecied h yp erparameters, w e in tro duce a Learnable Ga- b or Conv olution (LGC) lay er, where the lter parameters are directly from data without requiring prior domain kno wledge. The k -th channel of the 1D Gab or k ernel g at the t -th relative position within the kernel is dened as: g k ( t ) = exp − t 2 2 σ 2 k · cos(2 π f k t + ψ k ) , (1) where σ k , f k , and ψ k are learnable, denoting the scale, cen ter frequency , and phase shift of the Gab or kernel, resp ectiv ely . T o preven t the mo del from b eing inuenced b y slow-v arying signal osets and to emphasize the lo cal v ariations, w e apply a zero-mean constrain t as follo ws: ˆ g k ( t ) = g k ( t ) − 1 | T | T τ =1 g k ( τ ) , (2) where T is the length of the lter, and ˆ g k denotes the zero-mean v ersion of g k . Then, to simultaneously extract short-term and long- term features, w e further design a Multi-Scale F eature Blo c k (MSFB), whic h consists of parallel con v olutional la y ers with k ernel sizes k = { 7 , 15 , 17 , 39 , 41 } . Then, the extracted multi-scale features are concatenated along the c hannel dimension, follow ed by feature fusion using a 1 × 1 con v olution and further downsampling. Finally , subsequen t feature lay ers map the represen tation into a compact em b edding z v . Besides, MFEB fo cuses on extracting morphological features from ECG signals. MFEB shares a similar ov erall arc hitecture with VFEB, with the key dierence lying in the rst feature extraction stage: MFEB employs con v en tional conv olutional lay ers, whereas VFEB adopts the LGC lay ers. Then, through MFEB, the extracted morphological features are represen ted as z m . Unlik e static concatenation that treats z v and z m equally , PR-GA T explicitly mo dels interactions b etw een morphology and v ariation representations via graph at- ten tion. Sp ecically , we rst stack z v and z m in to H ∈ R 2 × d , where d is the feature dimension. Then, we pro ject H into query , key , and v alue embeddings with learnable matrices W Q , W K , W V as follo ws: Q = HW Q , K = HW K , V = HW V , (3) where Q , K , and V denote query , k ey , and v alue repre- sen tations, resp ectively . Then, the nal attention feature A is obtained through atten tion-based aggregation as follo ws: A = softmax LeakyReLU QK ⊤ √ d k , (4) where LeakyReLU( · ) denotes the Leaky ReLu activ ation function. d k is the scaling factor. Then, A is fed into a standard attention blo ck with residual connections and Lay er Norm la y ers, which yields the rened features H ′ = [ h ′ v ; h ′ m ] . The t wo c hannels of H ′ represen t the rened v ariation and morphology features, resp ectiv ely . T o fuse h ′ v and h ′ m , we design one attention p o oling mo dule. F or k ∈ { v , m } , a light weigh t MLP is used to predict a scalar score s k , which is normalized into a weigh t. Then, the pro cess can b e formulated as: α k = exp( s k ) j ∈{ v ,m } exp( s j ) , k ∈ { v , m } , (5) where α denotes the normalized feature. Then, the nal em b edding can b e obtained as follows: h phase = LN( α v h ′ v + α m h ′ m ) , (6) where LN( · ) denotes the Lay er Normalization op eration. h phase ∈ R D is the fused phase representation. In practice, this mechanism dynamically assigns dierent w eigh ts to dieren t features, thereb y enhancing robustness. 3) Phase-Group ed Hierarchical F usion (PGHF) : Up on ob- taining the disen tangled phase-sp ecic representations from IPR mo dules, PGHF is designed to fuse these fea- tures based on the physiologically-driv en fusion strategy . W e rst partition the embeddings of four phases in to tw o coheren t groups based on their morphological c haracter- istics. Sp ecically , the P and T/U phases are assigned to the slow-w av e group, since they exhibit relatively smo oth and lo w-amplitude w a v eform proles asso ciated with dep olarization-rep olarization transitions. Conv ersely , the QRS and ST phases constitute the fast-w av e group, distinguished by high-amplitude, sharp transient dynamics driv en by ven tricular activity . These groups are then pro- cessed by the PR-GA T mo dule, which facilitates selective cross-phase coupling via gated calibration, ensuring stable feature adaptation across distinct ph ysiological mo des. 4) Global Representation F usion (GRF) : T o bridge the slo w-w a v e and fast-wa ve groups into a discriminative em- b edding, GRF serves as the nal stage of our hierarchical arc hitecture. Similar to the previous step, we form ulate the global fusion as a high-level graph interaction task, rather than naiv e feature concatenation. Our proposed PR-GA T mo dule treats the embeddings of the tw o groups as graph no des and facilitates pairwise interactions to assess the relative imp ortance. This enables our metho d to dynamically mo del the relationship b etw een slo w and fast dynamics, adaptiv ely syn thesizing a holistic beat-level em b edding that is robust to inter-beat v ariability . C. Contrastiv e Loss V erication diers fundamentally from classication in that it inv olves a matching pro cess b et ween a query co de and the gallery template, rather than directly assigning a class lab el. Hence, a conv entional classication loss is insucien t for v erication tasks, as it mainly encourages features to cluster around class centers. Moreov er, such a loss tends to cause ov ertting, resulting in p o or general- ization to unseen identities, whic h is a common c hallenge 6 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE in verication scenarios. This loss fails to adequately constrain the feature space, making it ill-suited for ECG biometrics, where sub jects are represented by only a few enrolled b eats and must b e distinguished from many highly similar negativ e individuals. T o alleviate this issue, we optimize the enco der with a margin-based contrastiv e loss rather than the classication loss. F or i -th sample in the mini-batc h, w e use its paired p ositiv e P (same individual) and mine the negativ e sample N from the candidate p o ol consisting of all anchors and p ositiv es in the mini-batch. N is selected based on the highest cosine similarity (excluding the anchor and its paired positive). Let s ( · , · ) denote the cosine similarity form ulation. Then, our loss can b e form ulated as follows: L CL = 1 B B i =1 m + s u i , u N i − s u i , u P i + , (7) where m is the margin , [ · ] + = max( · , 0) . and u i is denoted the b eat-lev el feature pro duced by GRF. D. Implementation Since ECG signals are contin uously acquired, multiple heartb eats can b e collected for each individual. Then, w e adopt a Heartb eat-A ware Multi-prototype (HAM) Enrollmen t strategy to ensure the verication robustness. Unlik e the previous works, they represent each individual via the single feature vector, HAM represen ts each subject with a set of protot yp es. The em b edding set of the s -th individual can b e form ulated as E s = { u s, 1 , ..., u s,N s } , where N s denotes the n um b er of the heartb eats. T o construct a multi-protot yp e represen tation, we cluster E s in to K groups and represent eac h group b y a protot yp e. Sp ecically , the k -th prototype p s,k is computed as the mean of the em b eddings assigned to the k -th cluster: p s,k = N s n =1 I ( c s,n = k ) u s,n N s n =1 I ( c s,n = k ) , k = 1 , . . . , K, (8) where c s,n ∈ { 1 , . . . , K } denotes the cluster assignment of the n -th em b edding, and I ( · ) is the indicator function. During v erication, each query embedding is matc hed against all prototypes stored in the gallery . The iden tit y asso ciated with the prototype that yields the minimum distance to the query is assigned as the nal matching result. This design helps reduce the impact of noise and bias arising from individual heartbeats, thereby impro ving the robustness of the v erication pro cess. I V . E X P E R I M E N T S A. Datasets and Prepro cessing 1) Datasets : W e ev aluate our proposed metho d un- der dieren t exp erimental settings in three public ECG datasets. ECGID Dataset: Ninet y sub jects with 310 sessions are pro vided in the ECGID dataset. Each session contains a 20s single-lead I record with tw o columns (ra w/ltered as column 0/1) sampled at 500Hz, 12-bit. Sessions span 1 da y–6 months under unconstrained conditions with coarse R/T annotations [34]. MIT-BIH Dataset: In the MIT-BIH dataset, forty-eigh t half-hour Holter excerpts from 47 sub jects, typically MLI I and V1/V2 leads, at 360 Hz (11-bit, 10 m V range) with man ually v eried b eat lab els ( ∼ 110k b eats) [35]. PTB ECG Dataset: In the PTB dataset, ve h undred fort y-nine records from 290 subjects (1–5 p er subject) with 15 synchronous leads (12 standard + F rank XYZ) at 1000 Hz/16-bit plus clinical metadata [36]. 2) Pre-Pro cessing Details : T o accoun t for v ariations in lead conguration, sampling rate, and signal quality across ECG datasets, we adopt a unied prepro cessing pip eline to standardize mo del inputs. The ra w signals are denoised using a band-pass lter b et ween 0.5–40Hz to remov e baseline drift and high- frequency noise, and are then uniformly resampled to 200Hz. The CPS is applied to detect R-p eaks and seg- men t each b eat into four phases, including P , QRS, ST, and TU phases. A t the unied sampling rate of 200Hz, eac h sample represen ts 0.005s. A ccordingly , the segmen t lengths assigned to the P , QRS, ST, and T/U phases are 60, 40, 60 and 80 samples, corresp onding to temp oral durations of 0.30s, 0.20s, 0.30s, and 0.40s, resp ectiv ely . B. Exp erimen tal Settings and Comparison Metho ds 1) T raining and Ev aluation Settings : W e ev aluate our metho d under b oth closed-set and op en-set settings. In b oth cases, the data are split at the sub ject level instead of randomly splitting all samples. All metho ds were implemen ted in PyT orch and trained for 40 ep o chs using SGD with momen tum 0.9, a batc h size of 32, and a cosine annealing learning rate sc hedule initialized at 1 × 10 − 4 . Exp eriments were conducted with an AMD R yzen 7 5800X CPU and a single NVIDIA R TX 3060Ti GPU. Similar to the op en-set protocol, each query ECG is matc hed against all enrollment ECGs, and the predicted iden tit y is determined based on the minimum distance criterion. 2) Ev aluation Metrics and Comparison Methods : Sp eci- cally , we ev aluate our metho d under t wo tasks, including the verication and iden tication tasks. F or the v erica- tion task, we use Equal Error Rate (EER), dened as the v alue where the false acceptance and false rejection rates are equal. Besides, Receiver Op erating Character- istic (ROC) is also used to ev aluate the verication p erformance. Besides, for the iden tication task, T op-1 A ccuracy (Acc), Area Under the ROC Curve (AUC), and Cum ulativ e Match Characteristic (CMC) curv es are used to ev aluate the p erformance. Among them. CMC is used to measure the probability that the correct identit y ranks within the top- k predictions. T o ensure fair and comprehensive comparisons, we compare sev en baselines, including BAED [13], RDS- CNN [24], EDITH [10], PMS-ResNet [11], ADAFFN [28], JINT AO HUANG et al.: HEAR THE HEAR TBEA T IN PHASES: PHYSIOLOGICALL Y GROUNDED PHASE-A W ARE ECG BIOMETRICS 7 T ABLE I: Closed-set segment-lev el iden tication across three datasets. Our method consisten tly outp erforms state- of-the-art baselines, demonstrating sup erior represen tation learning under in tra-sub ject conditions. Method ECGID MIT-BIH PTB Acc (%) ↑ A UC (%) ↑ EER (%) ↓ A cc (%) ↑ AUC (%) ↑ EER (%) ↓ Acc (%) ↑ AUC (%) ↑ EER (%) ↓ PMS_RESNET 96.8700% 93.1958% 12.4092% 96.9625% 99.8303% 1.2697% 94.3336% 93.1958% 12.4092% ESN 82.9200% 85.0162% 21.2550% 95.4948% 98.5032% 4.9168% 92.1900% 92.1200% 14.3600% BAED 89.2708% 90.1200% 13.8700% 98.7302% 99.9106% 0.7120% 98.0616% 93.6600% 11.9300% ADAFN 83.6000% 90.6300% 13.4800% 96.8900% 99.7700% 0.9200% 92.9600% 96.5300% 9.7000% EDITH 91.8700% 90.1500% 14.5600% 95.6700% 94.0900% 10.8000% 98.8500% 96.1100% 8.3800% SSL_CNN 88.4347% 94.2343% 12.1726% 98.7169% 12.1726% 0.4241% 98.7069% 92.9626% 13.5638% RDS_CNN 88.3745% 90.5300% 14.9500% 98.6600% 99.1208% 4.0864% 92.5735% 90.8400% 15.8600% Ours 97.4829% 95.4247% 9.9058% 99.7553% 99.9692% 0.1114% 99.1678% 96.8652% 8.6857% T ABLE I I: Op en-set segment-lev el identication across ECGID, MIT-BIH, and PTB. Our mo del shows robust generalization to unseen sub jects across datasets with div erse pathologies and acquisition setups. Method ECGID MIT-BIH PTB Acc (%) ↑ AUC (%) ↑ EER (%) ↓ Acc (%) ↑ AUC (%) ↑ EER (%) ↓ Acc (%) ↑ AUC (%) ↑ EER (%) ↓ PMS_RESNET 82.4300% 84.6029% 22.7084% 89.2250% 85.0718% 23.7836% 80.7700% 82.3452% 24.5576% ESN 71.7800% 71.2400% 34.3700% 69.4326% 66.0120% 39.5100% 70.4300% 69.5938% 35.2144% BAED 71.2361% 69.8344% 34.1535% 89.3586% 87.3643% 19.0133% 85.4745% 94.7807% 9.6553% ADAFN 64.6900% 65.3800% 34.9100% 57.7700% 66.6600% 37.9600% 60.5900% 63.6300% 38.0400% EDITH 87.8700% 92.2100% 12.4300% 96.6600% 97.4600% 6.0400% 98.6500% 95.1100% 10.2800% SSL_CNN 57.1344% 66.9551% 38.8534% 90.0811% 90.0811% 17.8188% 87.8999% 87.0397% 20.0727% RDS_CNN 89.3680% 83.1503% 22.1800% 91.4315% 91.0700% 15.9090% 94.5760% 91.6559% 15.5920% Ours 94.4709% 92.5548% 15.1465% 98.2179% 97.6833% 8.0360% 98.9339% 96.2483% 9.6234% 2.5 5.0 7.5 10.0 12.5 15.0 17.5 20.0 R ank 0.92 0.93 0.94 0.95 0.96 0.97 0.98 0.99 1.00 A ccuracy CMC Curves on Close ECGID Dataset AD AFN BAED EDITH ESN Ours PMS-R esNet RDS-CNN SSL -CNN (a) ECGID 2.5 5.0 7.5 10.0 12.5 15.0 17.5 20.0 R ank 0.92 0.93 0.94 0.95 0.96 0.97 0.98 0.99 1.00 A ccuracy CMC Curves on MIT Dataset AD AFN BAED EDITH ESN Ours PMS-R esNet RDS-CNN SSL -CNN (b) MIT-BIH 2.5 5.0 7.5 10.0 12.5 15.0 17.5 20.0 R ank 0.92 0.93 0.94 0.95 0.96 0.97 0.98 0.99 1.00 A ccuracy AD AFN BAED EDITH ESN Ours PMS-R esNet RDS-CNN SSL -CNN (c) PTB 2.5 5.0 7.5 10.0 12.5 15.0 17.5 20.0 R ank 0.92 0.93 0.94 0.95 0.96 0.97 0.98 0.99 1.00 A ccuracy BAED AD AFN EDITH ESN Ours PMS-R esNet RDS-CNN SSL -CNN (d) ECGID 2.5 5.0 7.5 10.0 12.5 15.0 17.5 20.0 R ank 0.92 0.93 0.94 0.95 0.96 0.97 0.98 0.99 1.00 A ccuracy BAED AD AFN EDITH ESN Ours PMS-R esNet RDS-CNN SSL -CNN (e) MIT-BIH 2.5 5.0 7.5 10.0 12.5 15.0 17.5 20.0 R ank 0.92 0.93 0.94 0.95 0.96 0.97 0.98 0.99 1.00 A ccuracy BAED AD AFN EDITH ESN Ours PMS-R esNet RDS-CNN SSL -CNN (f) PTB Fig. 3: CMC comparisons of dierent methods across three datasets. Subplots (a)–(c) denote the p erformance of dieren t metho ds on the MIT-BIH, ECGID, and PTB datasets under the closed-set setting, resp ectively , while (d)–(f) sho w the corresp onding results under the op en-set setting. SSL-CNN [12], and ESN [37]. All baselines share the same prepro cessing pip eline, input conguration, and training sc hedule as our metho d. C. Closed-Set Exp erimen t A t rst, w e ev aluate our prop osed metho d under the closed-set setting, where the identities in the training and test sets are iden tical. F or eac h sub ject, half of the samples are used for training and the other half for testing. 8 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE T ABLE I I I: Ablation on basic segmentation with tw o branches across datasets. Protocol Method ECGID MIT-BIH PTB Acc(%) ↑ A UC(%) ↑ EER(%) ↓ Acc(%) ↑ AUC(%) ↑ EER(%) ↓ A cc(%) ↑ AUC(%) ↑ EER(%) ↓ Closed-set Only-CNN 88.4347 94.2343 12.1726 90.9963 99.8905 2.0421 98.7069 92.9626 13.5638 Only-Gabor 87.9113 94.7270 11.5600 99.6345 99.6345 1.9013 98.0266 94.6215 11.7848 Ours 97.4829 95.4247 9.9058 99.7553 99.9692 0.1114 99.1678 96.8952 8.6857 Open-set Only-CNN 57.1344 66.9551 38.8534 90.0811 90.0811 17.8188 87.8999 87.0397 20.0727 Only-Gabor 67.5981 79.7807 27.7885 85.3700 88.8950 19.3803 88.7787 89.0093 19.3619 Ours 94.4709 92.5548 15.1465 98.2179 97.6833 8.0360 98.9339 96.2483 9.6234 (a) ECGID (b) MIT-BIH (c) PTB (d) ECGID (e) MIT-BIH (f) PTB Fig. 4: R OC comparisons of dieren t v ariants. Subplots (a)– (c) sho w the ROC curv es on ECGID, MIT-BIH, and PTB under the closed-set setting, resp ectively , whereas (d)– (f) depict the corresp onding results under the open-set setting (same dataset order). T ABLE IV: Ablation on staged mo dule stacking (IPR is alw a ys enabled). Dataset CPS PGHF GRF A UC(%) ↑ EER(%) ↓ ECGID 3 7 7 92.4538 15.4915 3 3 3 92.5548 15.1465 MIT-BIH 3 7 7 97.0113 9.7294 3 3 3 97.6833 8.0360 PTB 3 7 7 95.7800 10.3812 3 3 3 96.2483 9.6234 The quan titativ e results are listed in T able I, with CMC curv es plotted in Fig. 3 (a)–(c). Across all three public datasets, our metho d yields the highest identication ac- curacy and low est EERs. On ECGID, our method ac hieves 97.48% accuracy , surpassing both PMS-ResNet (96.87%) and BAED (89.27%). The improv ement is most eviden t on the MIT-BIH dataset, where our mo del records a T op- 1 accuracy of 99.75% and an EER of 0.11%. Notably , this EER is signicantly low er than that of the closest comp etitors, BAED (0.71%) and PMS-ResNet (1.27%). 1 3 5 7 9 原型个数 80.0% 82.5% 85.0% 87.5% 90.0% 92.5% 95.0% ACC ECGID 开集 ACC Fig. 5: Ablation study ab out the num b er of enrollment protot yp es in HAM. D. Op en-Set Exp eriment Besides, w e ev aluate our proposed metho d under a more c hallenging setting, the op en-set setting, whic h assumes that the test sub jects are unseen during training. All sub jects w ere randomly split into disjoint training and JINT AO HUANG et al.: HEAR THE HEAR TBEA T IN PHASES: PHYSIOLOGICALL Y GROUNDED PHASE-A W ARE ECG BIOMETRICS 9 test sets, with mo del parameters trained exclusively on the training sub jects. F or each sub ject in the test set, half of the samples are used for enrollment, and the remaining half are used as query samples. The feature of each query ECG is matched against all enrollment ECGs’ features, and the identit y corresp onding to the minimum distance is assigned as the nal prediction. T able I I and Fig. 3 (d)–(f) summarize the results, the general p erformance decline is observed across baseline metho ds due to the distribution shift in unseen sub jects. SSL-CNN and ADAFN are particularly aected, yielding only 57.13% and 64.69% accuracy on ECGID, respectively . It can b e seen that our metho d is pro v en robust to this issue. It ac hieves the b est results on all public datasets: 94.47% accuracy on ECGID (surpassing the second best metho d by 5.11%), 98.21% on MIT-BIH, and 98.93% on PTB. Besides, we can observ e that our method can achiev e the low est EER in almost all settings. The CMC curv es reect this adv antage, showing higher identication rates at Rank-1 compared to comp eting metho ds. These results v alidate the eectiveness and robustness of our strategy , the previous metho ds achiev e satisfactory p erformance in the closed-set setting but they generally cannot keep it to the unseen individuals. In contrast, our metho d preserves robust p erformance by eectively mo deling phase-sp ecic information. E. Ablation Exp erimen ts As shown in T able I I I and Fig 4, we ev aluated the eectiv eness of each branch in MVFE. All v ariants adopt the same NPD segmentation proto col under the same training proto col. It can b e noticed that single-branch- based v ariations exhibit clear limitations in the op en- set proto col, while fusing the morphology and v ariation features improv es discrimination across datasets most no- tably on ECGID. Sp ecically , AUC increases from 66.96% (b est single-branch) to 92.55%. These results suggest that the tw o branches capture complementary cues that are dicult to obtain from a single view alone. Moreo v er, to v erify the h yp erparameter setting ab out the n umber of the prototypes in HAM, w e v alidate the p erformance in the ECGID dataset under the open-set setting, and the results can b e found in Fig. 5. W e observ e that the accuracy consisten tly improv es as the n um b er of prototypes increases from 1 to 3, indicating that the proposed multi-protot yp e design eectively reduces noise introduced by individual heartb eats and leads to more robust p erformance. How ever, when the num b er of protot yp es is further increased, the p erformance tends to saturate and shows no signicant v ariation. Therefore, w e empirically recommend using 3 as the n um b er of enrollmen t prototypes to balance p erformance and the storage cost. Finally , the results of the ablation study ab out the hierarc hical fusion strategy can b e found in T able IV. It can b e noticed that adding PGHF and GRF to the CPS and IPR yields consisten t gains, indicating that eac h mo dule con tributes to rening the global embedding and tightening the feature space structure. Imp ortan tly , the combined mo dules achiev e the strongest ov erall p er- formance among the compared v ariants under the same proto col, supporting the eectiv eness of in tegrating phase- guided hierarc hical fusion and global renemen t for robust op en-set iden tication. V . C O N C L U S I O N In this work, we prop ose a multi-gran ularity cardiac- phase represen tation framework for ECG identication b y segmenting each ECG signal in to four physiologically meaningful phases. W e then prop ose a hierarchical phase- a w are feature extraction framew ork, HP AF, which consists of the IPR, PGHF, and GRF mo dules. Sp ecically , phase- sp ecic MVFEs are assigned to each individual phase to extract phase-sp ecic representations in IPR, after which similar phases are group ed in PGHF and global features are progressively extracted in GRF. Our metho d achiev es comp etitiv e performance on public datasets. Our results suggest that representing an ECG signal as a comp osi- tion of phase-sp ecic cues, rather than a homogeneous whole-b eat pattern, can improv e the robustness of ECG- based identit y . Though our metho d achiev es impressive p erformance, a current limitation is that our metho d relies on the accurate R-p eak lo calization, which may degrade under severe noise or ectopic b eats. In our future w ork, we will explore adaptive phase lo calization and R- p eak-free segmen tation metho ds to ac hieve more robust p erformance. V I . R E F E R E N C E [1] M. Komeili, N. Armanfard, and D. Hatzinakos, “Liveness de- tection and automatic template up dating using fusion of ECG and ngerprint,” IEEE T rans. Inf. F orensics Security , vol. 13, no. 7, pp. 1810–1822, Jul. 2018. [2] L. Biel, O. Pettersson, L. Philipson, and P . Wide, “ECG analysis: A new approach in human iden tication,” IEEE T rans. Instrum. Meas., vol. 50, no. 3, pp. 808–812, Jun. 2001. [3] S. A. Israel et al., “ECG to identify individuals,” P attern Recognit., vol. 38, no. 3, pp. 133–142, 2005. [4] K. W eimann and T. O. F. Conrad, “F ederated learning with deep neural networks: A priv acy-preserving approach to en- hanced ECG classication,” IEEE J. Biomed. Health Inform., vol. 28, no. 11, pp. 6931–6943, Nov. 2024. [5] J. Ma, T. Zhang, and M. Dong, “A nov el ECG data com- pression method using adaptive F ourier decomp osition with security guarantee in e-health applications,” IEEE J. Biomed. Health Inform., vol. 19, no. 3, pp. 986–994, May 2015, doi: 10.1109/JBHI.2014.2357841. [6] S.-C. W u et al., “ECG biometric recognition: Unlinkability , irreversibilit y , and securit y ,” IEEE Internet Things J., vol. 8, no. 6, pp. 487–500, 2020. [7] M. A. Serhani et al., “ECG monitoring systems: Review, architecture, pro cesses, and key c hallenges,” Sensors, v ol. 20, no. 6, Art. no. 1796, 2020. [8] D. Jy otishi and S. Dandapat, “An LSTM-based model for person iden tication using ECG signal,” IEEE Sensors Lett., vol. 4, no. 8, pp. 1–4, Aug. 2020. [9] G. Petmezas et al., “State-of-the-art deep learning metho ds on electro cardiogram data: Systematic review,” JMIR Med. Informat., vol. 10, no. 8, Art. no. e38454, 2022. 10 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE [10] N. Ibtehaz et al., “EDITH: ECG biometrics aided by deep learning for reliable individual authentication,” IEEE T rans. Emerg. T opics Comput. In tell., v ol. 6, no. 3, pp. 928–940, 2021. [11] Y. Chu et al., “ECG authentication metho d based on parallel multi-scale one-dimensional residual net work with cen ter and margin loss,” IEEE Access, vol. 7, pp. 51598–51607, 2019. [12] G. W ang, S. Shank er, A. Nag, Y. Lian, and D. John, “ECG biometric authentication using self-sup ervised learning for IoT edge sensors,” IEEE J. Biomed. Health Inform., vol. 28, no. 11, pp. 6606–6618, 2024, doi: 10.1109/JBHI.2024.3455803. [13] A. J. Prakash et al., “BAED: A secured biometric authen- tication system using ECG signal based on deep learning techniques,” Bio cyb ern. Biomed. Eng., vol. 42, no. 4, pp. 1081– 1093, 2022. [14] J. P . Martinez et al., “A wav elet-based ECG delineator: Ev alu- ation on standard databases,” IEEE T rans. Biomed. Eng., vol. 51, no. 4, pp. 570–581, Apr. 2004. [15] A. F ratini et al., “Individual identication via electro cardiogram analysis,” Biomed. Eng. Online, vol. 14, no. 1, p. 78, 2015. [16] D. W ang et al., “A no vel electrocardiogram biometric identi- cation method based on temporal-frequency auto enco ding,” Electronics, vol. 8, no. 6, p. 667, 2019. [17] W. H. Jung and S. G. Lee, “ECG identication based on non-ducial feature extraction using window remo v al metho d,” Appl. Sci., vol. 7, no. 11, p. 1205, 2017. [18] S. C. F ang and H. L. Chan, “Human identication by quan- tifying similarity and dissimilarity in electrocardiogram phase space,” Pattern Recognit., vol. 42, no. 9, pp. 1824–1831, 2009. [19] R. Sriv astv a and Y. N. Singh, “ECG analysis for human recognition using non-ducial methods,” IET Biometrics, v ol. 8, no. 5, pp. 295–305, Sep. 2019. [20] A. Goshv arp our and A. Goshv arp our, “Human identication using a new matc hing pursuit-based feature set of ECG,” Comput. Methods Programs Biomed., vol. 172, pp. 87–94, Apr. 2019. [21] S. Gutta and Q. Cheng, “Joint feature extraction and classier design for ECG-based biometric recognition,” IEEE J. Biomed. Health Inform., vol. 20, no. 2, pp. 460–468, Mar. 2016. [22] D. A. AlDuw aile and M. S. Islam, “Using conv olutional neural netw ork and a single heartbeat for ECG biometric recognition,” Entrop y , vol. 23, no. 6, Art. no. 733, 2021. [23] J. Huang et al., “ECG arrhythmia classication using STFT- based spectrogram and con volutional neural netw ork,” IEEE Access, vol. 7, pp. 92871–92880, 2019. [24] E. Ihsanto et al., “F ast and accurate algo rithm for ECG authentication using residual depthwise separable conv olutional neural netw orks,” Appl. Sci., vol. 10, Art. no. 3304, 2020. [25] K. J. Chee and D. A. Ramli, “Electrocardiogram biometrics using transformer’s self-attention mechanism for sequence pair feature extractor and exible enrollment scope identication,” Sensors, vol. 22, no. 9, Art. no. 3446, 2022. [26] D. Jyotishi and S. Dandapat, “An ECG biometric system using hierarchical LSTM with attention mechanism,” IEEE Sens. J., vol. 22, no. 6, pp. 6052–6061, 2022. [27] M. Hammad et al., “ResNet-atten tion mo del for human authen- tication using ECG signals,” Exp ert Syst., vol. 38, no. 6, Art. no. e12547, 2021. [28] P . Yi et al., “ECG biometrics based on attention enhanced domain adaptiv e feature fusion net work,” IEEE A ccess, v ol. 12, pp. 1291–1307, 2023. [29] L. Sun et al., “Randomized attention and dual-path system for electro cardiogram iden tity recognition,” Eng. Appl. Artif. Intell., vol. 132, Art. no. 107883, 2024. [30] K. W ang and N. W ang, “ECG biometrics via dual-lev el features with collab orative embedding and dimensional attention weigh t learning,” Sensors, vol. 25, no. 17, Art. no. 5343, 2025. [31] T. Ma et al., “Out-of-distribution representation and graph neural netw ork fusion learning for ECG biometrics,” IEEE T rans. Biom. Behav. Identit y Sci., vol. 7, no. 2, pp. 225–233, 2025. [32] M. Al Al et al., “Enhancing biometric identication using 12- lead ECG signals and graph con volutional net works,” F ront. Digit. Health, vol. 7, Art. no. 1547208, 2025. [33] T. S. Kumar and V. Kanhangad, “Gabor lter-based one- dimensional local phase descriptors for obstructive sleep apnea detection using single-lead ECG,” IEEE Sensors Lett., vol. 2, no. 1, pp. 1–4, 2018. [34] T. Lugov ay a, “Biometric human identication based on electro- cardiogram,” M.S. thesis, Saint P etersburg State P olytechnical Univ., Saint Petersburg, Russia, 2005. [35] G. B. Mo o dy and R. G. Mark, “The impact of the MIT-BIH arrhythmia database,” IEEE Eng. Med. Biol. Mag., vol. 20, no. 3, pp. 45–50, 2001. [36] R. Bousseljot et al., “Nutzung der EK G-Signaldatenbank CAR- DIODA T der PTB üb er das Internet,” Biomed. T ech., v ol. 40, no. 1, pp. 317–318, 1995. [37] M. Hazratifard et al., “Ensemble siamese netw ork (ESN) using ECG signals for human authentication in smart healthcare system,” Sensors, vol. 23, no. 10, Art. no. 4727, 2023. First A.JinT ao Huang receiv ed his Bac helor of Engineering degree from the School of Soft- ware, Nanchang Hangkong Universit y in 2025. He is currently studying for a master’s degree at Nanchang Hangkong Universit y , China. His research interests include computer vi- sion, biometric recognition, and applications of generative articial intelligence. Second B.Lu Leng received his Ph.D de- gree from Southw est Jiaotong Universit y , Chengdu, P . R.China, in 2012. He p erformed his p ostdo ctoral research at Y onsei Univ ersity , Seoul, South Korea, and Nanjing University of Aeronautics and Astronautics, Nanjing, P . R. China. He w as a visiting sc holar at W est Vir- ginia Universit y , USA, and Y onsei Universit y , South Korea. Currently , he is a full professor and the dean of Institute of Computer Vision at Nanchang Hangkong Universit y . Prof. Leng has published more than 100 international journal and conference papers. He has been gran ted several scholarships and funding pro jects, including six pro jects supp orted by National Natural Science F oundation of China (NSFC). He serves as a reviewer of more than 100 international journals and 50 conferences. His researc h interests include computer vision, biometric template protection, biometric recognition, medical image pro cessing, data hiding, etc. JINT AO HUANG et al.: HEAR THE HEAR TBEA T IN PHASES: PHYSIOLOGICALL Y GROUNDED PHASE-A W ARE ECG BIOMETRICS 11 Third C.Yi Zhang received the B.S., M.S., and Ph.D. degrees in computer science and tech- nology from the College of Computer Science, Sich uan Universit y , Chengdu, China, in 2005, 2008, and 2012, resp ectively . F rom 2014 to 2015, he w as with the Depart- ment of Biomedical Engineering, Rensselaer Polytec hnic Institute, T roy , NY, USA, as a Postdoctoral Researcher. He is currently a F ull Professor with the School of Cyber Science and Engineering, Sich uan Universit y , and is also the Director with the Deep Imaging Group (DIG). He authored more than 140 papers in the eld of image pro cessing. These papers were authored or co-authored in several leading journals and conferences, including IEEE T ransactions on Medical Imaging, IEEE T ransactions on Information F orensics and Security , MedIA, IJCV, and CVPR; and rep orted by the Institute of Ph ysics (IOP) and during Lindau Nob el Laureate Meeting. His researc h interests include medical imaging, compressive sensing, and deep learning. He was a recipient of the major funding from the National Key Research and Developmen t Program of China, the National Natural Science F oundation of China, and the Science and T echnology Supp ort Project of Sic huan Province, China. He is also a Guest Editor of International Journal of Biomedical Imaging and Sensing and Imaging and an Asso ciate Editor of IEEE T ransactions on Medical Imaging and IEEE T ransactions on Radiation and Plasma Medical Sciences. F ourth D.Ziyuan Y ang is currently a postdo c- toral fellow in the Departmen t of Electronic Engineering at the Chinese Universit y of Hong Kong. He receiv ed the M.S. degree in com- puter science from the School of Information Engineering, Nanc hang Universit y , Nanchang, China, in 2021, and the Ph.D. degree from the College of Computer Science, Sic huan Univ er- sity , China. He was a Research In tern at the Centre for F rontier AI Research, Agency for Science, T echnology and Research (A*ST AR), Singapore. In the last few years, he has published over 50 pap ers in leading machine learning conferences and journals, including CVPR, AAAI, IJCV, IEEE T- IFS, IEEE T-NNLS, IEEE T-SMCS, and IEEE T-AI. He was a reviewer for leading journals or conferences, e.g. IEEE T-P AMI, IEEE T-TIP , IEEE T-IFS, IEEE T-MI, CVPR, and ICCV.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment