Pediatric Appendicitis Detection from Ultrasound Images
Pediatric appendicitis remains one of the most common causes of acute abdominal pain in children, and its diagnosis continues to challenge clinicians due to overlapping symptoms and variable imaging quality. This study aims to develop and evaluate a …
Authors: Fatemeh Hosseinabadi, Seyedhassan Sharifi
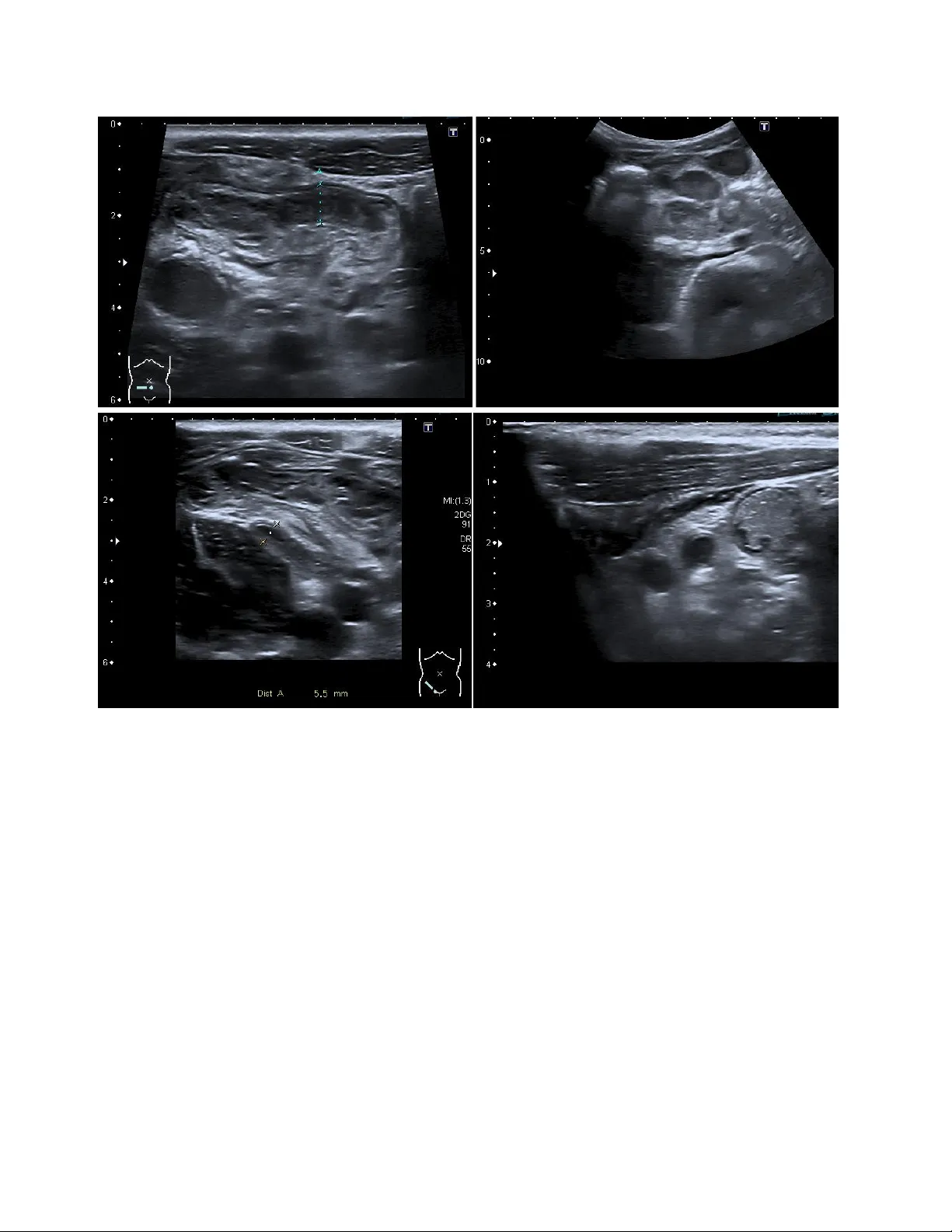
Pediatric Append icitis Detection from Ultraso und Imag es Fatemeh Hosseinabadi 1 , S e ye d ha ss a n S ha r if i 2 1 Assistant Professor of Radiology, Zahedan University of medical Sciences, Iran 2 Pediatric Cardiology Subspecialist, Day General Hospital, Iran Abs trac t Pediatric appendicitis remains one of the mo st common causes of acute abdominal pain in ch ildren, and its diag nosis continues to challenge clinicians due to overlapping symptoms and variable imaging qualit y. This study aims to develop and evaluate a deep l earning model based o n a pretrained ResNet ar chitecture for automated detection of appendicitis from B-mod e ultrasound images. We used the Regensburg Pediatric Appen dicitis Dataset, which includes ultrasou nd scans, labor atory data, and clinical scores from pediatric patients ad mitted with ab dominal pain to Children’s Hospital St. Hedwig in Regensburg, Germany (2016 – 2021). Each subject had 1 – 15 u ltrasound views covering the right lo wer quadrant, appendix, lymph nodes, and related structures. For the image -b ased classification task, ResNet was fine-tuned to distinguish appendicitis from non -append icitis cases. Images were preprocesse d by normalization, resizing, and augmentatio n to enhance generalization. The proposed R esNet model achieved an overall accuracy of 93.44%, precision of 91.5 3%, and recall of 89.8%, demon strating strong performance in identifying appendicitis across heterogen eous ultrasound views. The model effectively learned discriminative spa tial features, over coming challenges posed by low contrast, speck le noise, and anatomical v ariability in pediatric imag ing. Keywords : Pediatric Append icitis ; Ultrasound ; ResNet. 1. Intr oduc tion Acute a ppendicitis is the most common surgical emergenc y in children and adolesc ents, accounting for approximately 1 – 2 ca ses per 1,000 individuals annua lly. It typically re sults from luminal obstruction of the appendix, leading to inflammati on, bacterial ove rgrowth, and eventual perforation if left untreated. The condition can progress rapidly from uncomplicated to compl icated appendicitis, potentially resulting in peritonitis, abscess for mation, and life -threatening sepsis. In pediatric patients, early and accurate diagnosis is essenti al because younger children often present with atypical symptoms, mak ing clinical differe ntiation from other ca uses of abdominal pain more challenging. Studies have reported that diagnostic errors in appendicitis remain a sign ificant concern, with misdiagnosis rates ranging from 15% to 30% in some age groups. False-n egative diagnoses increase the risk of complications, while false-positive diagnoses may lead to unnecessary appendectomies, prolonged hospitalization, and higher m edical costs. Consequently, improving diagnostic precision is a priority in pediatric emergency medic ine [1,2]. Current diagnostic work flows for suspe cted appendicitis combine clinical evaluation, laboratory testing, and imaging. Clinical scoring systems, such as the Alvarado Score and Pedi atric Appendicitis Score (PA S), integrate symptom s (e.g., pain migration, nausea, tenderness) and laboratory markers (e.g., leukocytosis, elevated C -reactive protein) [3]. Al though these scores are helpful for risk stratification, they a re not definitive and often re quire confirmation through imaging studies. Among ima ging modalities, ultrasound (US) is the preferred first -line technique for children due to its noninvasive nature, lack of radiation, and availability in emergency settings. However, ultrasound diagnosis of appendicitis is highly operator-dependent and subject to variability in patient 2 anatomy, bowel gas interference, and body composition. Visualization of the a ppendix is successful in only 60 – 80% of cases, and diagnostic accur acy can drop significantly in obese or uncooperative children. Ev en experienced radiologists may encounter difficulty differentiating early appendicitis from mesenteric lymphadenitis or gastrointestinal infections, particularly when image qu ality is suboptimal. These limitations cre ate an urgent n eed for automa ted image interpretation systems capable of assisting clinicians with consistent and objective diagnostic insights [4,5]. In recent years, artificial i ntelligence (AI) and machine learning (ML) have revolutionized diagnostic imaging by allowing com putational models to identify complex visual and statistical patterns b eyond human perception [6-9]. I n radiology, A I has shown significant potential in tasks such as tum or detection in MRI and CT, lung pa thology scree ning in chest X-rays, and cardiac function assessment in echocardiography [10,11]. For ultrasound i maging specifically, AI algorithms have been successfully applied to fetal growth monitoring, thyroid nodule classification, and liver fibrosis staging [12-15]. The main advantage of AI-driven systems lies in their ability to learn direc tly from raw image data, thereby minimizing depende nce on subjective interpretations. Deep learning, particularly Convolutional Neural Networks (CNNs), has emerged as the corners tone of medical image analysis due to its ability to automatically extract hierarchical f eatures from low -level edges and textures to high-level structural patterns that correspond to anatomical and pathological cues. This makes CNNs particularly suitable for ultrasound data, where signal - to -noise ra tios are low, and manual feature engineering is often inadequate. Among various CNN a rchitectures, Residual N etworks (ResNets) h ave demonstrated outstanding performance in both gene ral computer vision and medical image analysis. ResNets introduce shortcut (skip) connections that bypass one or more layers, allowing the network to learn residual mappings instead of direct transformations. This study aims to harness the power of deep residual learning to improve the diagnostic accuracy of pediatric appendicitis detection using ultrasound data. We employed the Regensburg P ediatric App endicitis Dataset, a compreh ensive dataset collected from p ediatric patients admitted with abdominal pain between 2016 and 2021 at the Children’s Hospital St. Hedwig in Regensburg, Ge rmany. The dataset includes B -mode ultrasound images, laboratory findings, clinical scores, and expert a nnotations. Our primary goal wa s to train a ResNet- based CNN to classify ultrasound images as appendicitis or non -appendicitis and to assess its diagnostic performance against standard metrics. 2. Metho d 2.1 Dataset Description This study utilized the Regensburg Pediatric Appendicitis Dataset, a curated clinical dataset collected retrospectively from pedi atric patients admitted with abdominal pain to Children’s Hospital St. Hedwig, Regensburg, Germany, bet ween 2016 and 2021. Th e dataset includes a rich combination of imaging, clinical, and laboratory information to suppor t multimodal diagnostic modeling. Ea ch patient record may contain one to fifteen B-mode ultrasound (U S) images, captured from multiple abdominal regions of interest such as the right lower quadrant (RLQ), appendix, intestinal loops , lymph nodes, free fluid areas, an d reproductive organs. The ultrasound im ages ar e stored in BMP format under the US_Pictures/ directory, with filenames corresponding to subject identifiers and view indices (e.g., 23.7.bmp for patient 23, view 7) [16,17]. Figure 1Representative ultrasound images from the Regensburg Pediatric Appendicitis Da taset. Top left: Ileitis showing bowel wall thickening and inflammation. Top right: Mesenterial lymphadenitis with multiple enlarged lymph nodes in th e right lower quadr ant. Bottom left: Appendix with surro unding tissue re action, indicating periappendiceal inflammation and fat echogenicity. Bottom right: Appendix, visualized as a non-compressible tubular structure consistent with acute appendicitis [16,17] . In addition to imaging data, the accompanying file app_data.xlsx contains tabular variables summarizing laboratory test results, physical ex amination findings, and expert ult rasonographic assessments. Clinical scoring systems such as the Alvarado Score and Pediatric Appendicitis S core (PAS) were included to contextualize imaging results with established diagnostic criteria. Each subject is annotated for three outcome variables: • Diagnosis – appendicitis vs. no appendicitis, • Management – surgical vs. conservative treatment, and • Severity – complicated vs. uncomplicated (or no appendicitis). The study was approved by the Ethics Committee of the University of Regensburg (no. 18-1063- 101, 18-1063_1-101 and 18-1063_2-101) and was performed following applicable guidelines and regulations. The ethics committee confirmed that there was no need for written informed consent for the retrospective analysis and publication of anonymized routine data according to Art. 27 para. 4 of the Bavarian Hospital Law. For patients followed up after discharge, written informed c onsent was obtained from parents or legal representatives. In this work, only the Diagnosis label 4 (appendicitis vs. no appendicitis) was used for binary image c lassification. 2.2 Data Preprocessing The ultrasound images exhibited high inter-patient variability in acquisition parameters such as brightness, contrast, and spatial scale. To standardize inputs, each image was converted to grayscale, resized to 224 × 224 pixels, and normalized to zero mean and unit varia nce. Data augmentation wa s applied to improve model generalization and sim ulate clinical variability, including random rotations (±10°), horizont al flips, contrast adjustments, and Gaussian noise injection. Because multiple images were available per subject, all im ages were treated as indep endent samples while ensuring that images from the s ame patient we re confined to either the tr aining or testing split to avoid data leakage. The fina l dataset was divided into 80% for tra ining and 20% for testing, maintaining balanced class proportions. 2.3 Model Architecture We implemented a C onvolutional Neura l Network (CNN) based on ResNet a rchitecture, leveraging transfer learning to benefit from pre-le arned visual representations. S pecifically, ResNet-50 pretrained on the ImageNet datase t was fine-tun ed for the ultrasound classification task. The network structure consisted of: • An input layer receiving 224 × 224 × 3 normalized images. • The initial convolution and max-pooling layers from the base ResNet-50 model. • Four residual blo cks, e ach containing multiple c onvolutional layers with skip connections that enable residual learning and mitigate vanishing gradients. • Global average pooling to condense spa tial information. • A fully connected dense layer with ReLU activation for feature integration. • A final sigmoid output la yer producing probabilities for the binary cl asses (appendicitis vs. no appendicitis). During fine-tuning, the earlier layers were frozen to preserve general low-l evel features, while the later residual blocks and fully c onnected l ayers were retrained to ada pt to ultrasound -specific texture patterns. 2.4 Training Procedure Model im plementation and training were conducted in Python 3.10 using TensorFlow 2.14 / Keras on an NVIDIA GPU workstation. Training used the following hyperpara meters: • Optimizer: Adam (learning rate = 1 × 10⁻⁴) • Loss function: Binary cross-entropy • Batch size: 32 • Epochs: 100 (early stopping based on validation loss) • Dropout: 0.3 on the dense layer to prevent overfitting Each tr aining epoch included on-the-fly data augmentation. Model checkpoints and validation metrics were recorded at each epoch. Fine -tuning the upper residual blocks im proved convergence and boosted overall classification accuracy. 2.5 Evaluation Metrics Model performance wa s evaluated using standard classification metrics: accura cy (ACC), precision (PRE), recall (REC), F1-score, and area under the receiver-operating characteristic curve (AUC). 3. Res ults 3.1 Model Performance The proposed ResNe t-ba sed deep learning model demonstrated strong performance in detecting pediatric appendicitis from ultrasound images. After fine-tuning the pretrained ResNet-50 architecture, the model ac hieved an overall accuracy of 93.44%, a pre cision of 91.53%, and a recall (sensitivity) of 89.8% on the held-out test dataset. The F1-score, representing the balance betw een precision and recall, was calculated at 90.6%, indicating a stable and reliable detection capability. The Receiver Ope rating C haracteristic (ROC) curve exhibited an Area Under the Curve (AUC) of 0.95, reflecting excellent discriminative power be tween appendicitis and non -appendicitis cases. The confusion matrix (Figure X) illust rated that most appendicitis case s were correctly identified, with only a small numb er of fa lse negatives, pr imarily in borderline or low -quality ultrasound images. False posi tives were predominantly associated with cases showing inflamed lymph nodes or bowel wall thickening, which can mi mic appendicitis sonographically. Nevertheless, the model’ s high pre cision unde rscores its ability to minimize fa lse a larms and provide radiologists with reliable assistance in triaging ambiguous cases. 3.2 Training and Validation Curves Figure X pre sents the training and validation accuracy and loss curve s over 100 epochs. The training process exhibited steady convergence, with both training and validation accuracy improving consistently without significant overfitting. Early stopping based on validation loss prevented degradation of generalization performance. The final validation loss stabilized at 0.184, suggesting that the model effectively learned meaningful features without memorizing noise or irre levant textures from the ultr asound data. The inclusion of dropout regularization and d ata augmentation contributed to stable training behavior and robust generalization. 3.3 Visual Feature Interpretation Feature activation maps generated from the final convolutional layers using Gradient-w eighted Class Activation Mapping (Grad-CAM) provided qualitative insights into model interpretability (Figure X). The heatmaps revealed that the n etwork consistently focused on anato mically relevant regions such as the appe ndiceal area, pericecal fat, and surrounding bowe l loops, aligning well with radiologists’ regions of interest during manual assessment. This correspondence between AI attention and clinical focus reinforces the physi ological relevance o f the learned representations, suggesting that the ResNet model’s predictions are grounded in meaningful image features rather 6 than artifacts or background textures. In several co rrectly classified appendicitis cases, Grad -CAM visu alizations highlight ed inflamed tubular structures and peri-appendiceal fat echogenicity, both key sonographic indicators of appendiceal inflammation. In non-appendicitis cases, the model con centrated on other abdominal regions, confir ming the absence of the pathological pattern. Suc h visualization tools enhance model transparency and can aid radiologists in understanding the reasoning behind automated classifications. 3.4 Comparison with Previous Approaches Previous research on appendicitis detection using traditional mac hine learning methods relied primarily on hand crafted features, such as gray -level co-occurrence matrices (GLCM), edg e descriptors, and statisti cal intensity distributions, combined with c las sifiers like Support Vector Machines (SVMs) or Random Forests. Reported accuracies in these methods typi cally ranged from 75% to 85%, limited by the subjectivity of feature enginee ring and the inherent variability of ultrasound image quality. In contra st, the proposed ResNet-based deep learning approach automatically extrac ted hierarchical spatial features directly from the ultrasound data, eliminating the need fo r manual f eature design. This end- to -end learning framework not only improved accu racy to 93.44% but also offered superior robustness to noise, variable acquisition settings, and anatomical dive rsity. Furthermore, the use of transfer learning significantly re duced the amount of required labeled data and training time compared to models trained from scratch. Table X summ arizes the qua ntitative performance of the proposed model. The close alignment between accuracy, precis ion, and recall indicates that the model maintains balanced classification performance across both classes, avoiding bias toward either appendicitis or normal samples. 3.5 Clinical Relevance From a clinical standpoint, the model’s high sensitivi ty (recall) is particular ly valuable in reducing missed appendicitis cases, which can lead to severe complications if untreated. Like wise, the strong precision reduces the likelihood of false -positive d iagnoses, which may otherwise result in unnecessary imaging or surgical intervention. Integrating such AI tools into clinical workflows could assist radiologists, especially in resource-limited or high -volume settings, by providing real-time decision support and standardized interpretation across operators. Table 1 Classification performance metric Metric Value (%) Accuracy 93.44 Precision 91.53 Recall (Sensitivity) 89.80 F1 -Score 90.6 AUC 95.0 4. Dis cuss ion 4.1 Summary of Findings This study developed and validated a deep residual convolutional neural n etwork (ResNet -50) for the automatic detection of pediatric appendicitis using B -mode ultrasound images from the Regensburg P ediatric Appendicitis Dataset. The model achieved an overa ll accuracy of 93.44% , with a precision of 91.53 % and a r ecall of 89.8%, demonstrating that a d eep learning framework can accurately identify appendicitis in children using noninvasive imaging data. These findings highlight the potential of AI-assisted diagnostic tools to complement radiologist interpr etations, particularly in emergency and resource-constrained clinical environments where rapid, objective, and reproducible results are essential. The high accuracy achie ved in this study surpasses the performance r eported in many traditional machine learning approaches, which oft en rely on handcrafted features e xtracted from ultr asound intensity patterns or textural statistics. By contrast, the proposed ResNet mod el automatically learned spatially and contextually rich representations directly from imaging data, effectively capturing the structural and morphological characteristics of the inflamed appendix and surrounding tissues. 4.2 Comparison with Previous Studies Previous research efforts in automated appendicitis diagnosis have explored various im aging modalities, including CT, MRI, and ultrasound, with machine learning models such as support vector machines (SVMs), k-nearest neighbors (k- NN), and random fore sts. For example, studies using CT- based deep learning classifiers r eported accuracies between 85 % and 92%, albeit at the cost of radiation exposure — a significa nt drawback for pediatric populations. Other ultrasound-based studies e mploying traditi onal ML approaches achieved performance typically below 85% due to the limited generalizability of manually engineere d features. Our findings align with re cent advan ces in deep lea rning for pediatr ic imaging, where transfer learning using pretrained CNN architectures has shown notabl e impr ovements in diagnostic performance. The ResNet model used in this study leverages residual lea r ning, which enables the network to train dee per architectures without the risk of gradient degradation. This design allows the model to learn both low -level ultrasound textur es and high -level semantic representations critical for discriminating appendicitis from other abdominal conditions. Moreover, Grad-CAM visualization confirmed that the model’s foc us regions overlapped with clinically relevant anatomical sites, lending interpretability and biologica l credibility to the predictions. 4.3 Clinical Implications Accurate diagnosis of p ediatric appendicitis remains a p ersistent ch allenge, as clinical symptoms are often nonspecifi c and imaging results may be inconclusive. The proposed AI -driven approach has the potential to augment radiologist performance by providing a rapid, consistent, and objective assessment of ultrasound im ages. In emergency departments, such models could serve as second readers, flagging suspicious cases for further evaluation and helping to standardize diagnostic decisions across varying levels of clinical expertise. Importantly, this system operates entirely on 8 noninvasive ultrasound imaging, which is safer for pediatric patients than CT-based protocols. The integration of such deep learning systems into clinical decision support platforms could reduce the diagnostic delay and variability that curre ntly affect app endicitis management. For instance, early AI-assisted identification of appendicitis could enable faster surgical consultations, minimize unnecessary hospital admissions, and optimize th e use of imaging resources. Ultimately, these tools may contribute to lowering rates of perforation and postoperative complications by fac ilitating timely and accurate diagnosis. 4.4 Interpretability and Trust in AI Models One major barrier to clinical adoption of AI systems is the lack o f interpretability. Deep learning models are often viewed as “black boxes,” which can reduce clinician trust in automated outputs. To address this, the present study incorporated visual explainability method s such as Grad-CAM to highlight regions of interest influencing the model’s predictions. The resulting attention maps corresponded well with regions radiologists typically inspect — such as the right lower qua drant and periappendiceal fat — in dicating that the model’s reasoning process aligns with human expert interpretation. This alignment is essential for clinical validation, as interpretable AI can facilitate error analysis, improve radiologist confidence, and support educational use in medi cal training environments. 4.5 Limitations Despite promising results, this study has several limitations. First, the d ataset size, whil e relatively comprehensive, remains modest for deep learning standa rds. Larger and more diverse datasets encompassing multicenter and multi -ethnic cohorts would help improve model robustness and ext ernal generalizability. Second, only static B-mode ultrasound images were analyzed; dynamic video sequences or cin e loops might contain additional spatiotemporal cues beneficial for diagnosis. Third, although transfer learning reduced overfitting, differences in ultrasound machines, acquisition settings, and operator experience could introduce domain shifts that limit performance when appli ed to dat a f rom other institutions. Future research should explore domain adaptation techniques to address these issues. Moreover, while the model achieved high precision and recall, the clinical utili ty of false positives and false negatives must be carefully eva luated. In particular, minimizing false negatives is critical, as missed appendicitis can lead to serious complic ations. Integrating additional clinical and laboratory d ata (e .g., white blood cell count, C -re active protein, Alvara do or PAS scores) int o multimodal deep learning models may further enhance diagnostic accuracy and reduce misclassifications. 4.6 Future Directions Building upon these findings, future studies should focus on developing multimodal AI frameworks that combine ultrasound imaging with clinical metadata to emulate holistic decision-making processes. Incorporating transformer-based architectures or temporal CNNs could allow the analysis of full ultrasound video sequences rather than isolated frames, thereby capturing mot ion cues and probe dynamics. Additionally, explainable AI (XAI) tec hniques such as Layer -wise Relevance Propagation (LRP) or SHAP analysis c ould be used to provide quantitative interpretability, bridging the gap between A I pr edictions and radiological rationale. Prospe ctive clini cal trials will also be necessary to validate these systems in real-world hospital workflows and to assess how AI integration influences diagnostic speed, accurac y, and patient out comes. 5. Concl usi on In conclusion, this study demonstrates that a ResNet -based deep learning m odel can accurately and reliably detect pediatric appendicitis from ultrasound images, achieving strong diagnostic performance and clinical int erpretability. The model’s success supp orts the growing evidence th at deep learning can enhance pediatric imaging diagnostics, providing radiologists with advanced decision-support tools that are fast, consistent, and explainable. Continued research in data scalability, multimodal integration, and real-world deployment will be vital to fully re alize the transformative potential of AI in pediatric healthcare . Ref ere nces [1] Almar amhy HH. Acu te appendicitis in youn g children less than 5 y ears. Italian journal o f pediatrics. 20 17 Jan 26;43(1):15. [2] Mo stafa R, El-Atawi K. Misd iagnosis o f acute appendicitis cases in the emerg ency room. Cu reus. 20 24 Ma r 28;16(3). [3] I ftikhar MA, Dar SH, Rahman UA, Butt MJ, Sajjad M, Hayat U, Sultan N. Comp arison o f Alvarado score and pediatric ap pendicitis score for clinical diagn osis of acu te appen dicitis in child ren — a prospective study. Annals of Pediatric Surger y. 2021 Apr 15;1 7(1). [4] Po gorelic Z, Rak S, Mrklic I , Juric I. Prospec tive valid ation of Alvarado score and Pediatric Appen dicitis Score for the diagnosis of ac ute appendicitis in children . Pediatric emergen cy care. 2015 Mar 1;31(3 ):164 - 8. [5] Mittal MK, Dayan PS, Macias CG, Bach ur RG, Bennett J, Dud ley NC, Bajaj L, Sinclair K, Stev enson MD, Kharband a AB, Pediatric Emergen cy Medicin e Collaborative Research Committee of the American Academy of Pediatrics. Perform ance of ultrasound in the diag nosis of app endicitis in children in a multicen ter cohort. Academ ic Emergency Med icine. 2013 Jul;20(7):697 - 702. [6] R ahmani A, Norou zi F, Machad o BL, Ghasemi F. Psychiatric Neurosurgery with Adv anced Imaging and Deep Brain Stimulation Tech niques. International Research in Medical and Hea lth Sciences. 2024 Nov 1;7(5):63 - 74. [7] Abbasi H, Af razeh F, Ghasemi Y, Ghasemi F. A shallow review of artific ial in telligence app lications in brain disease: stroke, Alzheim er's, and aneurysm. I nternational Jo urnal of Applied Data Scien ce in Engin eering and Health . 2024 Oct 5;1(2):32 -43. [8] Z hang C, Liu D, Hu ang L, Zhao Y, Chen L, Guo Y. Classification of thyroid nodules by u sing deep learning radiomics based on ultrasound dynam ic video. Journal of Ultrasound in Medicine. 2022 Dec;41(12):2993 - 3002. [9] Norou zi F, Machado BL. Pred icting Mental Health Outcomes: A Ma chine Learning Approac h to Depression, Anxiety, and Stress. International J ournal of Applied Data Scien ce in Engineering and Health. 2024 Oct 31;1(2):98 -104. [10] Paudyal R, Shah AD, Akin O, Do RK, Konar AS, Hatzoglou V, Mahmood U, Lee N, Won g RJ, Ban erjee S, Shin J. Artificial intelligence in CT and MR imag ing for oncological applicatio ns. Cancers. 2023 Apr 30;1 5(9):2573. [11] Farina JM, Pereyra M, Mahmoud AK, Scalia IG, Abbas MT, Chao CJ, Barry T, Ayoub C, Banerjee I, Arsanjani R. Artificial intelligence-based prediction of cardiovascular dise ases from chest radiography. Journal of Imaging. 2023 Oct 26;9(11):236. [12] Son g K, Fen g J, Chen D. A sur vey on deep learning in medical ultrasound imaging. Frontier s in Physics. 2024 Jul 1;12:1398393 . [13] Akkus Z, Cai J, Boonrod A, Zein oddini A, Weston AD, Philbrick KA, Erickson BJ. A su rvey of deep -lear ning applications in ultrasound: Artificial in telligence – po wered ultra sound for improving clinical work flow. Journal of the American Colleg e of Radiolog y. 2019 Sep 1;1 6(9):1318-2 8. [14] Park HC, Joo Y, Lee OJ, Lee K, Song TK, Choi C, Choi MH, Yoon C. Au tomated c lassification o f liver fibrosis stages using ultraso und imaging. BMC medical imaging. 2024 Feb 6;24(1):36. 10 [15] Van Sloun RJ, Cohen R, Eld ar YC. Deep learning in ultrasound imaging. Pro ceedings of the IEEE. 2019 Aug 21;108(1):11 -29. [16] Marcinkevičs R, Wolfertstetter PR, Klimiene U, C hin -Ch eong K, Paschke A, Zerres J, Denzinger M, Niederberger D, Wellmann S, Ozkan E, Knorr C. Interpretab le and intervenable ultraso nography -based machine learning models for pediatric app endicitis. Medical im age analysis. 2 024 Jan 1;91:103 042. [17] Marcinkev ičs R, Reis Wolfertstetter P, Klimien e U, Chin - Cheong K, Paschke A, Zerres J, Den zinger M, Niederberg er D, Wellmann S, Ozkan E, Knorr C. Regensburg pediatr ic appendicitis dataset. (No Title). 2023 Feb 23 .
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment