Evaluation of MRI to ultrasound registration methods for brain shift correction: The CuRIOUS2018 Challenge
In brain tumor surgery, the quality and safety of the procedure can be impacted by intra-operative tissue deformation, called brain shift. Brain shift can move the surgical targets and other vital structures such as blood vessels, thus invalidating t…
Authors: Yiming Xiao, Hassan Rivaz, Matthieu Chabanas
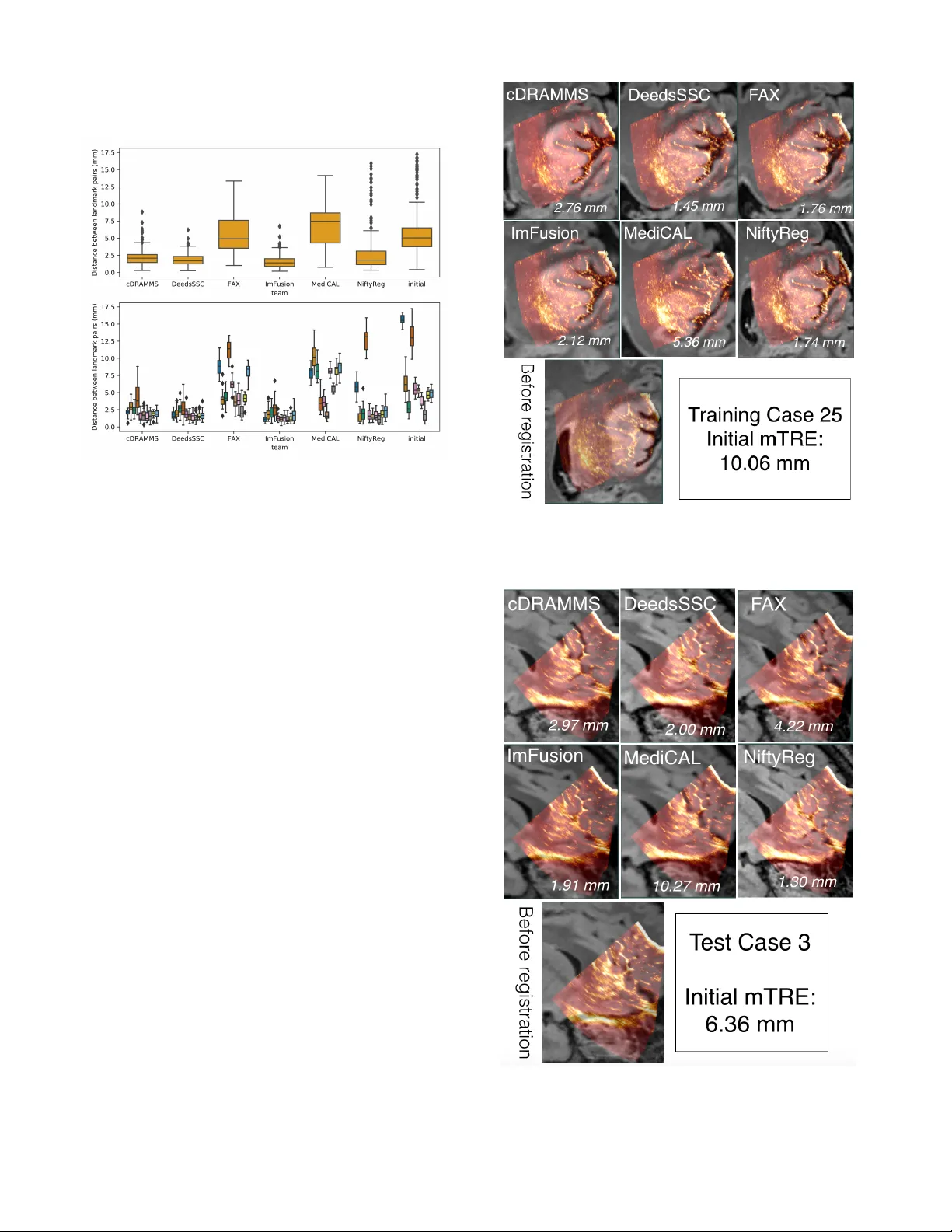
Abstract — In brain tumor surgery , th e quality an d safety of the procedure can be impacted by intra - operative tissue deformation, called brain shift. Brain shift can move the surgical t argets and other vital structures such as blood vessels, thus invalidati ng the pre - surgical plan. Intra - operative ultrasound (iUS) is a convenient and cost - effective imaging tool to track brain shift and tumor resection. Accurate image registrat ion techniques that update pre - surgical MRI based on iUS are crucial but challenging. The MICCAI Challenge 2018 for Correction of B rain shift with Intra - Operativ e UltraSound (CuRIOUS2 018) provided a public platform to benchmark MR I - iU S registration algorithms on newly released clinical datasets. In this work, we present the data, setup, eval uation, and results of CuRIOUS 2018, which received 6 fully automated algorithms from leading academic and industrial research groups. All algorithms were first trained with the public RESECT database, and then ranked based on test dataset of 10 additional cases with identical d ata cur ation and annotation protocols as the RE SECT database. The article com pares the res ults of all participating teams and discusses the insights gained from the challenge, as well as future work. Index Terms — Registration, brain, ultr asound, MRI, br ain shift, tumor *This work has been submitted to the IEEE for possi ble publication. Copyright may be transfer red without notice, afte r which t his versi on may no l onger be ac cessible. I. I NTRODUCTION LIOMAS are the most comm on brain tumors in adults, and are categorized into grade I - IV by the World Health This manuscript was submitted on xx, revised o n xx, and accepted on xx. Y. Xiao i s with the Robarts Research Institut e, Weste rn Univer sity, London, ON, Canada ( e - mail: yxiao286@uwo. ca). H. Rivaz is with the PERFORM Centre and Department of Electrical and Computer Engineering, Concordi a University , Montreal, Canada (e - mail: hr ivaz@ece.concor dia.ca ). M. Chabanas is with Universit y of Grenoble Aples, Grenobl e Inst itute of Technology, Grenoble , France (e - mail: Mat thieu.Chaba nas@univ - grenoble - alpes.fr) . M. Fortin is w ith th e Departm ent of Health, Kine siology & Applied Physiology, Concordia University, Montreal, Canada (e - mail: maryse.for tin@concordia.ca ). I. Machado is with the D epartment of Radiology, Brigham and Women’s Hospital, Harvard Medical Sch ool, Boston, MA, USA. Y. Ou is with the Department of Pediatrics and Radiology, Boston Children’s Hos pital, Harvard Medical School, Boston, MA, USA. Organi zation (WHO). Low - grade gliomas (LG G, grade I and II) are less aggressive and have slower progression tha n high - grade gliomas (H GG, grade III a nd IV) , but will eventually undergo malignant transformation into high - grad e tumors . Evidence s [1 , 2] have s hown that early tum or rese ction can effectively improve the patient’s survival rate . I mage - guidance can be a useful tool to assi st the surgeon in obtaining a maxim al safe resection of t he tumor . Image - guidance based on pre - operative MR images is in routine clinical use worldwi de. These s ystems, however, do not account for the tissue shift and deformations that occur as the resection progre sses. Due to brain shift , the surgic al targe t and other vital structures (e.g., blood vessels and ventricles) will be displac ed relat ive to the pre - surg ical plan and resulting in inaccurate im age - guidance. Mult iple fa ctors ca n contr ibute t o brain s hift, includi ng but no t limited to drug administ ration, intracranial pressure change, tissue re section. O ften such tissue shift is not directly visib le by the surgeon. Both int ra - operative ultrasound (iUS) and intra - operative magneti c reson ance imaging ( i MRI) have been employed to track tissue deformation and surgical progress . Intra - operative US has ga ined po pularity thanks to it s low cost, high portability and flexibility. However, limite d field of view and challenging image interpretation remain obstacles for widesp read use. Togethe r with iUS, automati c image registration algorithms c an b e us ed to upd ate th e su rgical p lan based on pre - operative MRI by re - aligning the pre - operative images with intra - operative imag es and offer m ore intuitive assessments of the extent of r esection. M.P. Heinrich is with the Institute of Medical Informatics, University of Luebeck, Germany. J.A. Schnabel is with the Schoo l of Biomedical Engine ering and Imaging Sciences, King’s Col lege London, UK. X. Zhang and A. Maier are with the Pattern Recogni tion Lab, Department of C omputer Science, Friedrich - Alexander - Universität Erlangen - Nürnberg, Martens str. 3, 91 058 Erlangen , Germa ny . R. Shams and S. Kadoury are with the Institute of Biomedical Engin eering, Polytechnique Montrea l & CHUM Research Centre. D. Drobny is with the Wellcome/E PSRC Cenre for Interventional and Surgical Sciences, University of College London, UK. D. Drobny and Marc Modat ar e with the School of Biomedical Engineering & Imaging Sciences, King’s College London, King’ s Health Partner s, St Thomas’ Hospital, London, SE1 7EH, UK. W. Wein is with I mFusion GmbH, Munich German y. I. Reinertsen is with SINTEF, NO - 7465 Trondheim, N orway, (e - mail: Ingerid.Reinertsen@sintef.no). Evaluation of MRI to ultrasound registration methods for brain shift correction: The CuRIOUS2018 Challenge Yiming Xiao , Member, IEEE, Hassa n Rivaz, Member, IEEE, Matthieu Chabana s, Mary se For tin, Ines Macha do, Yangmin g Ou, Mattias P . Heinrich, Julia A. Schn abel, Xia Zhong, Andreas Maier, Wolf gang We in, Roozbeh Shams, Samuel Kadoury, Davi d Drobny, Marc Modat, and Ingerid Reinertsen G Previously, a number of algorithms and strategies [3 - 8] have been developed to address iUS - MRI regi strat ion for brain s hift correction. They range from new strateg ies to map im age features to similar domains [3, 4] to n ovel cost function [6, 7] , and from different deformation models [5, 6] to improve d optimization pr ocedures [8] . How ever, partially due to the lack of relevant clinical datasets, it has been difficult to directly compare different algori thms , thu s p otentially slowing the speed of technical translation to benefit surgeons and p atients . The MICCAI Challenge 2018 f or Correction of Brain shi ft with Intra - Operati ve UltraSo und (CuRIOUS201 8) was laun ched as the fir st pu blic p latform to b enchma rk th e latest im age registration algorithms for the task, and to bring the researchers together to discuss the technical and clinical challe nges in iUS - guided brain tumor resection. For the first edition of the challenge, we focused on M RI - iUS re gistration to correct pre - resection deformation after craniotom y, as it typ ically sets the tone of brain shift for the re st of the surg ery. The challenge w as divided into two phases . In the first phase, the publicly available REtroSpe ctive Evaluation of Cerebral Tumors ( RESECT ) database [9] was used to train registr ation algorithms . The p articipating teams then submitted a descript ion of their meth od s and results on the training data. T wo week s befo re the chall enge ended, a priva te te sting database curated under the same condition as the traini ng set was suppli ed to the partic ipants who submi tted their results based on the training data. For both training and testing data, the distance between homologous anatomical landmarks across iUS and M RI were u sed to assess and rank the registration quality. The CuRIOUS2018 challenge received 8 initial submissions [10 - 17] . Seven teams validated th eir methods on the testing data, a nd six participated in the final ranking . The submissions cover a wide variety of approaches, including the latest registratio n metric s [7, 13] , op timization approaches [12] , and deep learning techniques [17] . This paper describes the organization, submitted algorithms, and results for the challenge, and further discusses the current challenges and potential future directions of tissue shift correction in US - guided brain t umor surgery. II. M ATERIALS Two datasets were included for the training and testing phases of the CuRIOUS2018 challenge. The RESECT database [9] was provided to the partici pants as the tr aining dataset for development and fine - tuning of th e alg orithms. The database contains pre - operative M R and pre - resection iUS im ages from 22 patient s who have received LGG resection surgeri es at St. Olavs Univ ersity Hospital , Trondhe im, Norway . The testi ng dataset wa s com prised of imaging data f rom 10 additional patients with LGG obtained in the same setting as t he RESECT database. The collecti on and distrib ution of both datasets were approved by the Regional C ommittee for Medical and Health Research Ethics of Central Norway, and all patients signed writt en infor med consen t. For both trai ning and testing data bases, Gd - enhanced T1w MRI and T2w fluid - attenuated inversion recovery (FLAIR) MRI scans w ere ac quired for each patient before surgery. F ive fiducial markers w ere glued to the pa tient’s h ead prior to scanning. The T1w and T2w MR Is w ere rigidly co - registered, and ali gned to t he patient’s head position on the operating table via a fi ducial - based image - to - pati ent registration. The posit ion - tracked 3D iUS scans were acqu ired with the Sonowand Invite neuronavigation system (Sonowand AS, Trondhei m, Norway), with eit her the 12FLA - L linea r transdu cer or the 12FLA flat linear array transducer for smaller superficial tumors. 3D volumes were reconstructed from the raw iUS data using the built - in proprietary reconstruction method in th e Son owand Invite system, with a reconstru ction resolutio n in the range of 0.14x0.14x0.14 mm 3 to 0.24 x0.24x0.2 4 mm 3 depending on the probe types and imaging depth. Both ultra sound transdu cers were facto ry calibrate d and equip ped with removable sterilizable re ference frames for optical tracking. A Po laris camera (NDI, Waterloo, Canada) built in the Sonowand system was used to obtain the position and pose of the ultras ound probe. Therefore, the iUS volumes reveal tissue position and deformation in t he patient’s head on the operating table. TABLE I D ETAILS OF INTRA - MODALITY LANDMARKS FOR EACH PATIENT IN THE TESTING DATASET Patient ID # of landmarks MRI vs. before US Mean in itial dist ance (range) in mm MRI vs. before US 1 17 15.66 (14.19~16.74) 3 17 6.36 (3.57~10.23) 4 17 2.98 (1.17~5.28) 5 17 13.19 (9.86~17.25) 6 18 5.52 (4.07~7.24) 7 18 5.27 (4.28~6.14) 8 18 3.73 (2.66~5.04) 9 17 1.80 (0.41~4.15) 10 17 4.66 (3.76 ~5.74) 12 17 4.89 (3.58~6.21) mean ± sd 17.3 ± 0.5 6.41 ± 4.46 The number of landmarks and mean initial Euclidean distances between landmark pairs are shown, and the range (min ~ max) of the distances is shown in parenthesis after th e mean value. Homologo us anatom ical landmarks manually labeled by two raters (authors YX and MF as Rat er 1 and 2 , resp ectively) were provided to assess registration quality, using the software ‘register’ included in the M INC toolk it ( http://bic - mni.gi thub.io ). Typical landmarks includ e the edge of the tumo r , dee p grooves of sulc i, corners of sulci, conv ex poin ts of gyri and the horns of the lateral ventricle s. After Rater 1 defined the lan dmarks in the T2w FLAIR MRIs as the refer ence s, Rater 1 and Rater 2 the n tagged th e correspo nding lan dmarks independ ently with in the co rrespond ing US volumes twice . A 1~2 - week interva l was ensured between the repetiti ons. T he final landmarks in both training and testi ng database were provided as the averaged results of two trials of landmark marked by both rater s (four 3D poin ts for each landmark). The details of the l andmarks are listed in Table I for the testing data sets . Similar det ails for the training dataset can be found in the original pu blication for th e RESEC T databa se [9] . For both sets, a wide range of brain shift s measured as mean init ial distances between correspondi ng landmarks were included to properly examine t he performance of registrat ion algorithms. We empl oyed the mean Euclidean distance between two sets of corresponding landmark points for each patient t o assess the intra - and inter - rater variability. For in tra - rater variability, we calculated the me tric b etween two trials of lan dmark pickin g for each rater; for inter - ra ter variability, th e average of tw o trials for each ra ter was first computed and used to obtain th e value between two raters. The intra - and inter - rater variability evaluations ar e presented i n Table I I for both training and testing data. TABLE II I NTER - AND I NTRA - RATER EVALUATIONS WI TH MEAN E UCLIDEAN DISTANCE BETWEEN LANDMARK SET S Type Intra - rater Rater 1 Intra - rater Rater 2 Inter - rater R1 vs. R2 Training data 0.47±0.10 mm 0.33±0.06 mm 0.33±0.08 mm Testing data 0.21 ± 0.10 mm 0.48 ± 0.22 mm 0.42 ± 0.17 mm T he results are shown as mean ± standard deviation. III. C HALLENGE SETUP The CuRIOUS2018 challenge started on April 1 st , 2018 w hen the challenge website w ent live on curious2018. grand - challenge.org. In the n ext few days, several groups who were active in the field of MRI - iUS registration were identified by literature sea rch and were in vited to participate. The challeng e was also widely adverti sed on mailing lists and on bullet in b oard of medical imaging conferences held in the first half of 2018. Another factor that lead s to a good p articipation w as the incentive of generous support of challenge sp onsors, which provided a t otal of 2,100 € for the top three win ner s. The challenge co nsisted of two phases. In ph ase I , all th e tea ms were require d to submit a short paper that elab orated the techniq ue a nd res ults on the 22 patien ts in the RESE CT datab ase. These papers were then peer - reviewed and the final camera - ready conference papers were submit ted in July 201 8. Phase II started in August 2018, wh en all the p articipants who had submitted reports and results on the training data were provided with MRI and iUS data from 10 additional patients (test data) . Th ese data sets had iden tical data curation and annotation protocols as the RES ECT database . The locatio n of landmark s in the MRI wa s provided to the teams, and the teams had to return the location s of those landma rks after MRI - iUS registration withi n 13 days of the data release . All tea ms p resented their m ethods and re sults on the training d ata at the challenge event , which took place in conjunc tion to MICCAI 2018 in Granada, Spai n. The RESECT database remains public, and ha s been downloaded 267 times since its release in April 2017. The te st datasets were only released t o the participants and the loca tion s of the ground truth lan dmarks in t hese datasets rem ains private . T he organizers will continue the challenge in 2019 by adding iUS test data collected d uring and a fter tumor re section . IV. E VALUATION The evaluation metric and ranking system are key criteri a for the success of a challenge. The metric should reflect the overall quality of the methods and the ranking system should be as fair as possible. It is worth noting that our eva lu ation meth od was published on the official websit e before the challenge took place and was not modified afterward s . A lthough such transparenc y in the ev aluation process may seem obvious, [18] reported that th is transp arency was not guaranteed in about 40% of biomedical chal lenges, which could lead t o controversy. The first component of the evaluation process is a metric to assess the quality of the registrati on methods. More than 80% of the tasks in biomedical challenges concern segmentation, with the Dice sim ilarity co efficient as the most common evaluation metric [18] . However, challenge s with image registration, e specially from different modalities, are rarer a nd we could not find any standar d metrics from the se competiti ons. We thus chose to rely sole ly on th e expert - lab eled anato mical landmark p airs, by comp uting the Euc lidian distanc es between the transfo rmed M RI lan dmarks , after reg istration, an d the ground - truth landm arks defin ed in the iU S image s. The se cond component of the evaluation process concer ns how the results for ea ch test ca se are a ggregated to rank the tea ms. The two main options are 1) aggregate the result s on all test cases, then rank; or 2) rank by test case, then aggregate the ranks. In the first scenario , we would have ranke d the teams based on the mean distance computed from all landmarks of all cases. Instead, we chose the second scenario because it i s better fitted to h andle missing cases. For each case , we also ranked fully - automatic methods over semi - automatic methods. To aggregate the cas e - by - case ranks, we simply computed the mean r ank of e ach team. The evalu ation system was as follows : 1. For each test case and for each team, compute the Euclidia n distances between landmark pairs after registration, i.e. between the transfo rmed MRI landmark s and the g round truth iUS landm arks. 2. For each test case , rank tea ms according t o their mean distance between l andmark pairs. Exceptions include : a. If one team could not provide results for a test case, or if these results could no t be processed for any reason, then that team is ranked last for the test case. b. If two mean distan ces differ by less than 0.5 mm, a team wit h a full y - automatic method is ranked higher than a team with a semi - automatic method. 3. Compute the mean rank of ever y team, which gives the final ranks of the Challen ge. V. C HALLENGE ENTRIES A. Team cDRAMMS Macha do et al. [13] extended the Deform able Reg istration via Attri bute Matchi ng and Mutual - Saliency Wei ghting (DRAM MS) algorithm [19] , a g eneral - purpose algorithm [20] , specifically fo r the US - MRI re gistr ation pro blem, whic h th ey termed as corre lation - similarity DRAMMS or cDRA MMS. They released it at https://www.nitrc.org/ projects/dramms/ (v ersion 1.5.1) . The original DRAMMS has two good properties for US - MRI regi strat ion. Firs t, repres entin g ea ch voxel with multi - scale and multi - orientation G abor attributes in DRAMMS offe rs a r icher i nformation than purely ima ge intensities. Th is helps t o establish more reliable voxel correspondences despite the differ ent image protocols and different i ntensity profiles between U S and MRI images. Second, the mutual - saliency mod ule in DRAMM S automatically assigns low confidence or weights to regions that cannot establish r eliable or cann ot find counterparts across images. This potentially reduces the negative effects of the missi ng correspo ndences between US and MRI images . Differ ent from the origina l DRAMM S, which uses the sum of square differences (SSD) between attributes for matching, the modifi ed cDRAMMS uses corr elation coeffi cient [21] and correlation ratio [22] on attributes for voxel matching. C C and CR on voxel attribute s in cDRAMMS establish voxel correspondences at a hi gher accuracy and higher reliability than SSD in DRAMMS. B. Team DeedsSSC Heinri ch et al. [14] used DeedsSSC, which comprises a linear and a non - rigid registra tion that are both based on discrete optimization and modality - invarian t image features. Specifical ly, self - similarity con text features (SSC ) are extracted for both MRI and ultrasoun d scans that are matched based on a dense displacement sampling. Fir st, the similarity maps for each considered control point are used to extract correspondences for fitting a l in ear tra nsform using least trimmed squares, simila r as done in block - matchi ng approaches. Second, new similarity maps are calculated for linearly aligned im ages and an efficien t g raphical mod el b ased discrete optimization (deeds) is used to estim ate a non line ar displacement field that avoids implausible warps and further improves the reg istration quality. All computations are performed for scans resampled to isotropi c 0.5 mm resolution and using the default parameters (see https://git hub.com/mattiaspaul/deedsBCV ) with an optimization over multiple grid - scales. Finally, the non linearly warped landmarks are again constrained to follow a rigid 6 - parameter trans form for improved robustness. The algorithm is executed within less than 10 seconds per scan pair on a multi - core CPU and ongoing work considers the huge potential for further speed - ups through parallelized GPU computations. C. Team FAX Zhong et al. [17] propose d a learning - based approach to resolve intraoperativ e brain a s an im itation game. This point - based approach predict s the deformation vectors of key points to compensate the non - rig id brain - shift. F or each key point, they extract a local 3D patch in iUS and m odel the key point distribution as the encoding of the current observation. A demonstrator is constructed providing the optimal deformation vector based on the current key point location an d the grou nd truth. An artificial neu ral n etwork is tr ained to imitate t he behavior of the demonstrator and to predict the optimal deformation vector given current obser vation. T o increase robustness, the propos ed technique use s a mu lti - tasking network wit h a rigid transformation as auxiliary output. In addition, w e use a non - rigid deformation to augmen t the 3D volume and 3D key points to facilitate t he training. D. Team ImFusion The method [16] is base d on the m ulti - modal simila rity metric LC 2 [7] and has recently been used in a first live evaluation du ring surgery [23 ] (data NOT overlapping with challeng e data) . A non - linear op timization alg orithm cha nges the valu es of a parametric transformation model to maximize it. In a pre - processing step specific to the challenge data set (cartesian 3D ultrasound volumes compounded by the SonoW and system), the volum e sides facing the ultrasou nd probe is e stimated an d the oute rmost 4m m of content are cropped accordingly. The registration algorithm is im plemented in the pro prietary ImFusion SDK w ith full OpenG L - based GPU accelerat ion. The ultrasound volume is assigned as fixed volume, resampled to 0.5mm (half the M RI voxe l size), and prope rly zero - masked. The chosen simil arity metric patch - size is 7 × 7 × 7 voxels, as optimized in prior work. Two non - linear o ptimizers successively oper ate on the parameters of a rigid pose from the initialization as pr ovided by th e navigation system. Th e first is a global DIR ECT ( DIviding R ECTangles ) su b - division met hod [24] searching on translation only, f ollowed by a local BOBYQA (Bound Opt imization BY Quadrati c Approxi mation) algori thm [25] on al l six parameters. The local optimizer then executes another search on full affine parameters in order to accommod ate non - uniform scaling and shearing of the data. E. Team MediCAL Mult imodal d eforma ble reg istra tion be tween t he MRI and intra - operative 3DUS w as achieved with a weighted version of the locally linear co rrelation metric (LC 2 ), correlating MRI intensities and gradients with ultrasound, while adapting both hyper - echoic and hypo - echoic regions within the cortex. The method [15] was initi alized with a global rotatio n of the US volume to match the orientation ob served on the MRI. This was achieved using a PCA of the extracted inferior skull region, identifying th e principal or ientation vec tors of the hea d, followed by a sc aling and translation c orrection. This fusion step uses a patch - based approach of the US vox els, co mparing intensity and gradien t magnitudes extracted from the MRI with a linear r elationship. The registration applies sequentially a rigid and non - rigid step, w ith the later integrating a w eighting t erm and controlled by a cubic 5 × 5 × 5 B - Spline inter polation grid, distribut ed uniformly in the fan - shaped U S volume . The weight ing ter m uses pre - annotated labels on the M RI, representing both the hypoec hoic (fluid c avities) and the hyperechoic (ex. choroid plexus) areas observed on ultrasound. This term is added only at the non - rigid step as it is highly specific to the internal areas in US such as the lateral ventricles, requiring a rigid pre - alignment. Registration optimization was performed using BOBYQA, w hich avoids computing the metri c’s deri vatives . F. T eam NiftyReg Drobny et al. [1 2] suggest a method which uses a block - matchi ng appro ach to a utomatic ally al ign the pre - operative MRI with the iUS ima ge. The reg istrat ion algo rithm use d is part of the Nift yReg open - source software package [26] . The block - matchi ng method of the registra tion stage itera tively estab lishes point correspondences between the reference image and the warped floati ng image and then determines the transformation parameters using least trimmed squares (LTS) regression. A two - leve l pyramidal approa ch for coarse to fine registration is used. For the block - matchi ng, both images are divided into uniform blocks of 4 voxel edge length. The 25% of blocks with the highest intensity variance in the reference image are used and the rest are discarded. Each of these im age blocks is compared to all floati ng image blocks that overlap with at leas t one voxel. The floating image matching block for ea ch reference block is determined as the one with maximum absolute normalized cross - correlation (NCC). After establishing the point - wise correspo ndences the second step is the update of trans formation parameters via LTS reg ression. At every iterati on, the c omposition of the block - mat ching correspondence and the transformation of the previous step determines the ne w transformation by LTS regression. VI. R ESULTS A. Phase I : distances on the training data The result s obtained on the training dataset were reported by e ach team in their respective contribution to the challenge proceedings [27] . Mo st authors reported their distance s obtained after registration for each case, although some re ported only averaged values. T able III summarizes the mean distance between landmark pairs after regist ration, over all landmark of all cases, compute d by e ach team. All teams but one improved from the initial distances, with three teams achieving a mean d istan ce und er 1.75 mm and tw o mo re unde r 3.35 mm. Team cDRA MMS initial report ed a m ean dista nce between landmarks of 3.35 ± 1.39 mm . With an updated ve rsion of their method, this er ror was later redu ced to 2.28 ± 0.71 mm . Sun et al. [11] provided partial results on 4 cases only , since the other 18 cases were used to train their neural network. This team eventually did not partici pate t o the second phase. TABLE III S UMMARY OF THE CHALLENGE RESULTS . Team Distances between landmark pai rs after registration ! Mean ± std, in mm Training set / Test set Mean case - by - case rank Final challenge ! rank cDRAMMS 3.35 ± 1.39 2.18 ± 1.23 3.4 3 = DeedsSSC 1.67 ± 0.54 1.87 ± 0.93 2.4 2 FAX 1.21 ± 0.55 5.70 ± 2.93 5. 3 5 = ImFusion 1.75 ± 0.62 1.57 ± 0.96 1.5 1 MedICAL 4.60 ± 3.40 6.59 ± 2.89 5. 3 5 = NiftyReg 2.90 ± 3.59 3.21 ± 3.57 3.1 3 = Hong et al . 5.60 ± 3.94 6.65 ± 4.55 - - * Sun et al. 3.91 ± 0.53 - - - Initial distances 5.37 ± 4.27 6.38 ± 4.36 - - For each team, the first columns give the mean distances betwee n landmark pairs after registration, computed over all la ndmark of all cases, for the training and test sets. The mean case - by - case rank, computed on the test set only, and the fin al c hallenge rank are then given. For c omparison, the last line contains the mean initial distances , before registra tion. Teams cDRAMMS and NiftyReg were event ually ranked tied at thi rd (=). Hong et al. sent results on the test d ata but did not attend the challenge e vent. Sun et al. sent only partial results (*) on the train ing se t, but did not participate to the second phase of the challenge. These two teams were thus not ranked. B. Phase II : distanc es on the test data This section presents the results of the 6 teams that completed phase II of the challenge, on the test dataset. Figure I first shows the results per test case, aggreg ated ac ross all teams. Test cases with th e largest in itial err or (cases 1, 5, and 3) were the most difficult to treat. Results for test ca ses w ith the smallest initial error (4 and 9) were in average improved, also several teams obtained larger distances after registration. Finally, results were consistently improved for al l other cases with an initial error in the 4 - 6 mm range. Figure I. Results per test case: box plot distribution of the distances between landmark pairs. For each test case, t he left box plot (blue) shows the in itial d istances before r egistration. while th e r ight box plot (orange) shows the distribution after registrati on, aggregated ove r all teams. Regardi ng team - by - team results , mean distan ces betwee n landmark pairs after registration are sum marize d in Table III while the di stribut ion of these distances is de tailed in Figur e II . Teams ImFusion and DeedsSSC obtained a mean distan ce between landmark pairs well below 2 mm, respectively of 1. 57 and 1.87 mm. These ex cellent results are consistent across all test c ases, with a stan dard deviation around 1 mm for both teams, which confirme d the results re ported on the trainin g dat a set. Team cD RAMMS also consistently obtained very good results, with a mean error of 2.18 mm and a single large residual error of 4.3 m m for case 5. Results of team NiftyReg are m ore contrasted. As can be seen on the lower panel of Figure II , they o btained excellent results for all cases but two, cases 1 and 5, where the dista nce was only reduced from 15.7 to 5.9 m m and 13.2 to 12.8 m m, respect ively. Without these two outliers, the mean distan ce over all cases would be reduced from 3.21 ± 3.57 mm to 1.70 ± 0.91 mm. Team FAX re ported the best res ults on the training set, with a mean distan ce between landmark pa irs of 1.21 ± 0.55 mm. However, this distance leaped to 5.70 ± 0.55 mm on the tes t dat a, which potentially shows the ir deep l earning method over fitted the data durin g the trainin g phase. F inally, team MedICA L ob tained few or no improv ement s from the initial distance s between landmark pairs. Figure II. Distribution of the distances between landmark pairs obtained by each team after registration, on t he t est set. For comparison, the last column contains the initial distances before registration. The upper panel shows the global results computed over all landmar ks of all test cases. In the lower panel, these results are split by test case. C. Phase II : com plementary criteria All met hods were fu lly automa tic . Alt h ough it was not a factor in the evaluation, sev eral team s repo rted their computation time. These valu es r ange from 1.8 sec for team F AX (architecture no t specified), to approxima tely 20 sec for bo th DeedsSSC and ImFusion on laptop s, and to 103 sec for team MedIC AL. Computation time of teams cDRAMMS and Nifty Reg were no t provid ed. D. Quali tative results To demonstrate th e d ata a nd the registration task , MR and iUS volumes of one patient was chosen from each of the training and test datasets , and are show n in Figures III and IV , respectively. The select ed cases have a relatively large initial mTRE and a substant ial variab ility betwe en the teams. Also, n ote that these two cases do not necessarily directly reflect the o verall r anking of the challenge, w hich wa s based on averaged ranking s of all cases. As no quantita tive measu res of registration quality are available in a clinical setting, visual inspection of the images is important to obtain an impression of the registratio n quality . As sho wn in F igures III and IV , the registration accuracy can be evaluated by adapted visualization and identification of homologous features such as sulci, gy ri and ventricles in the i mages. Figure III. Qualitative comparison of registration results for Training Case 25 across different teams. For each team, the ultrasound and deformed FLAIR MRI scan is overlaid together. The mTRE values for each t eam is listed at the right bottom corner of each im age ov erlay. Figure IV. Qualita tive comparison of regist ration results for Test Case 3 across different teams. For each team, the ultrasound and deformed FLAIR MRI scan is overlaid together. The mTRE values for each team is listed at the right bottom corner of each im age overlay. E. CuRIOUS2018 Challenge ranks Following the descri ption in Sectio n IV , all teams were ranked independ ently for ea ch test cas e, based o n the m ean distan ces between landmark pairs. These case - by - case ranks are summarized in Figure V , with the numbe r of times each team was ra nked at the i th place, with i from 1 to 6. The winner and runner - up are team s Im Fusion and DeedsSSC , which are perfectly consistent with their respective results reported in Figure II . No te that ImFusio n obtained the best registration for 6 of the 10 tests cas es. Despite a larger m ean registration e rror, team NiftyR eg w as ra nked third be fore team cDRAMM S as it obtained a better case - by - case rank (3.1 vs 3.4). How ever, team cDRAMMS also had very good results but consistently handled all cases, including the extreme ones. T his specific situation pointed out the fact that the challenge metric favors accuracy over precision, with a limited penalty whe n low quality results are obtained on a single case or two. To overcome this limit, as we consi der precision is a crucial fact or for the su rgeons' acceptance of a method , the chall enge’s organizers decided to declare a tie for thi rd place. Both Nifty Reg and cDRAMMS thus received the same third place prize. Finally, bot h teams FAX and MedICAL obtained a mean case - by - case rank of 5.3, and were ranked tied at the 5 th place in the chall enge. Figure V. Case - by - case r anks for each team. For example: o ver the ten test cases, team ImFusion was ranked first six times, second three times, and third one time. VII. D ISCUSSION In this challenge, the focus has been on MR - iU S registration in the context of brain tumor surg ery. As both the training and test datasets exclusively contain data from LGG surgeries, there h as been a s pecial focus on this tumor type. The resection of LGGs is particularly challeng ing as the tumor tissu e can be very similar to norm al brain tissue. In LGG surgery , there are also fewer options for additional guidance a s tools like 5 - ALA fluorescence are not available. Intraoperative ultrasoun d is therefore an attractive solution in these cases. The optimiz ation and ben chmarking of available registration algorithms on data from these tumors is therefore particularly im portant for successful future cli nical translation. Even though the em phasis has been on LGG, the resul ts f rom the challenge will generalize w ell to other tumor types such as HG Gs and metastasis as these tumors a re more distinct from normal brain tissue and d epict clearer boundaries than LGGs in ultrasound images. An import ant obsta cle for the widespr ead use of iUS is the challenging and un fami liar image inter pretation. Th e integration of iUS into the navigation system an d the visualization of corresponding slices in pre - operative MR and iUS makes this interpretation con siderably more intuitiv e. W ith accurate MR - iUS registration, the surgeon can perform the resection based on the MR ima ges even after brain shift which makes the neuro navigati on ac curate and easy to interp ret. MR - iUS registration also en ables correction of oth er ty pes of pre - operative MR data such as fMRI and DTI [28] . Image registration techniques tailored for M RI - iU S registration in this ch allenge were lan dmark - , intensity - or learning - based. The performance of landmark - based methods in non - linea r image registratio n depends on both finding enoug h landmark s that cover the en tire volume, an d correctly find ing their corresponding landmarks in the second volume. The voxel - wise attribute - based meth o d of Machado et al. [13] (team c DRAMMS) did relatively well despite the fact that iUS and MRI have drasti cally differe nt salient fea tures, and ranked third in a tie with Drobny et al. [1 2] (te am NiftyReg ). The top three algorithms in this challenge [12 - 14] were all intens ity - based te chniques, which calculate d a dense transformation map by utilizing i ntensity values at al l locations. Deep learnin g has had a large success in segmentation and classification problems in m edical image analysis , but its success in image registration ha s be en much less impressive [29] . The two submissions that used DL in this challenge were from Sun et al. [11] , w ho did not participate to the second phase, and Zhong et al. [17] (team FA X) , w ho ra nked first in the resu lts reported on the training database. How ever, their method d id not work well on the test database. A com mon culprit for such behavior is overfitt ing, where the model overfits the training data and therefore performs poorly on the unseen test data. As more training dat a becomes availa ble, this method is expected to perform better in the fu ture. Symmetric image reg istration techniques provide unbiased estimates of the transformation field and are know n to generally outperform their asymm etric counterparts [30] . Two of the top three meth o ds in this challenge [12, 1 4] com pared the performance of their techniques in symmetric and asym metric settings. They both concluded that asymmetric transformations lead to a superior performanc e in this ch allenge. This is an interesting finding and is lik ely due to the vast diff erences in physics of US and MR imaging modalities. It was noticed by some of the ch allenge participants and discussed during the event that in some cases affine transforma tions outperform ed non - linea r elastic transforma tions. T his mig ht seem surprisin g a s b rain shift is often described as a non - uniform deformation. However, before resection a large compon ent of the e xperienced m ismatch between MRI and iUS is often due to inaccurate patient - MRI registration. This is a rigid registration m ost often base d on anatomical landmarks, fiducials, surfaces or a combination of these. Co nsequen tly, an affin e transform ation m ight be sufficient to correct for m ost of the m isalignment. After resection, the situation will be different with larger and highly non - linear d ef ormations and affine transformations w ill likely not be suffic ient to register t he images. In both the training and test databases, we selected landmarks that cove r a larg e part of the iUS volume with maximal distanc e between neighboring landmarks. This strategy provides a good benchmark for comparison of image registration techniques. However, the quali ty of the alig nment closer to the tumor is more cl inicall y importan t as it be tter hel ps the neu rosurgeon to optimize the r esection size and locati on. The distance between corresponding landmarks in the two images before and after registratio n is a well - established metric for evaluation of reg istration results in the absence of a g round truth. Despite being widely used, this metric has some limitations. A s there is only a lim ited num ber of landmarks associated with each image, the registrati on error is only evaluated at a limited number of locations and will therefore not capture local displacements and d eformations in other locations. T he numb er of landm ark s and their distribution in the image vo lume are th erefore im portant. Th e landma rks in both the training and test sets have ther efore been carefu lly placed in order to capture the displ acements and deformations as good as possible. How ever, we noticed that in te st case #5 the registration resu lts were not ac curate by visual inspection even though th e mTR Es indicate d succes sful alignm ent. This emphasizes the need for both quanti tative and qualitati ve assessment of registration results. Another limitation of this metri c is the localiza tion error associat ed with manual placement of points. Also, for the landmarks to be valid for evaluation of registration results, this localization error has to be signifi cantly lower than the expected regist ration errors. W e have measured the inter - and intra - rater variability in both the training and test data a nd show n that thes e are signific antly lower than the registration errors. E ven though th e land marks do not represent the absolute ground t ruth, they a re vali d for the evaluation of the registrati on results. The use of landmarks as the only metric for the challeng e also re present a limitation as other important characteristics such as computation time are not measur ed. For implement ation in a clini cal settin g, for example, other characteristics would also be of cri tical importance. However, in the challenge setting, the use of a single well - defined metric is advantageous. A single metric enables a straightforward, comprehensible ranking sc heme and an open , fair com petition. Wit h the use of mult iple met rics , there will always be a discussion of the weighting of the different characteristics and how to aggregate the results. The rules for aggregation of the ranks in this challenge were o utlined before the challen ge and were not changed a t any p oint. Still, th e system used favors accuracy over precision. As d iscussed during the chal lenge event, for the clinical users, hi gh precision and high accuracy are equally important and preci sion can even be more import ant t han accur acy. This point should be re - designed and improved i n future editions of the challenge. VIII. F UTURE WORK In the first edition, the registration task solely focused on MRI - iUS registra tion before d ura - opening and after craniotomy. However, with the progress of tumor r esection, tissue deformation is an on - going process, and accurate tracking can ensure the complete removal of cancerous tissues, preventing any additional surgeries. Intended as a recurr ent open challenge to further improve the registratio n algorithms, we expect to introduce multiple su b - challenges in future CuRIOU S challenges to target brain shift correction at different stages of the surgery , especially d uring and a fter resection . For clinical practices, besides accuracy and robustness, processing speed is an im perative factor. In the inau gural edition of the challenge, perfor mance speed was not emphas ized in sco ring the team s because it can be a ffected by multi ple factors, inclu ding implement ation platf orms, for prototype algorithms. In future challenges, we aim to place discussions and emphasis on this topic, as well as optimization algorithms to direct the re sults of the challenge towards more realistic clinical implementations. IX. C ONCLUSION Holdin g great clinical values, MRI - iUS registration for correcting tissue shift in brain tumor resection is still a difficult task . As the first public image processing challenge to tackle this clinical problem, t he CuRIOUS2018 Challe nge provided a common platform to eva luate and discuss existing an d emerging registration algorithm s on this topic . The results of CuRIOUS2018 provided valuable insights for th e current developments and challenges from both the tech n ical and clinical perspectives. This is an important step forward to help translate researc h - grade automatic image processing into clinical practice t o benefit the patie nts and clin icians. A CKNOWLEDGMENT The work is support ed by Norwegian National Advisory Unit for Ultrasou nd and Imag e guided Therap y, NSE RC D iscovery RGPIN - 2015 - 04136. The organizers w ould like to thank Medtr onic and the Fre nch ANR within the Investissements d’Avenir program under reference s ANR - 11 - LABX - 0004 (Labex CAM I) fo r the ir ge nerous sponsorship . Th e work for the DeedsSSC algorithm was funded in part by the German Research Foundat ion (DFG) under grant number 320997906 HE 7364/2 - 1. Drobny et al. were supported by the UCL EPSRC Centre fo r Doctoral Train ing in M edical Imaging and Well come/E PSRC Cent re for Interv entio nal and Surgic al Sciences [NS/A000050/1], and the Wellcome/EPSRC Centre for M edical En gineering [WT 2031 48/Z/16/Z] and EPSR C [NS/A0000 27/1]. R EFERENCES [1] A . S. Jakola, K. S. Myrmel, R. Kloster, S. H. Torp, S. Lindal, G. Unsgard , et al. , "Com parison o f a stra tegy favorin g early surgical resection vs a strategy favoring wa tchful waiting in low - grade gliomas," JAMA, vol. 308, pp. 1881 - 8, Nov 14 2012. [2] D. A. Schomas, N. N. I. Laack, R. D. Rao, F. B. Meyer, E. G. Shaw, B. P. O'Neill , et al. , "Intra cranial low - grade gliomas in adults: 30 - year experience with long - term follow - up at Mayo Clinic," Neuro - Oncology, vol . 11, pp. 437 - 445, Aug 2009. [3] T. Arbel, X. Mor andi, R. M. Comeau, and D. L. Collins, "Automatic non - linear MRI - ultrasound registration for the correction of intra - operative brain deformations," Comput Ai ded Surg, vol. 9, pp. 123 - 36, 2004. [4] I. Reinertsen, F. Lindseth, G. Unsg aard, and D. L. Collins, "Clinical validation of vessel - based registration for correction of brai n - shift," Med Image Anal, vol. 11, pp. 673 - 84, Dec 2007. [5] P. Coupe, P. Hellier, X. Morandi, and C. Barillot, "3D Rigid Registration of Intraoperative Ultrasound and Preoperative MR Brain Images Based on Hyperechogenic Structures," Int J Biomed Imaging, vol. 2012, p. 531319, 2012. [6] D. De Nigris, D. L. Collins, and T. Arbel, "Multi - modal image registration based on gradient orientations o f minimal uncertainty," IEEE Trans Med Imagi ng, vol. 31, pp. 2343 - 54, Dec 2012. [7] W. Wein, A. Ladikos, B. Fuerst, A. S hah, K. Sharma, and N. Navab, "Global Registration of Ultrasound to MRI Using the LC2 Metric for En abling Neurosurgical Guid ance," Medical Image Computing and Computer - Assisted Int ervention (Miccai 2013), Pt I, vol. 8149, pp. 34 - 41, 2013. [8] H. Rivaz, S. J. S. Chen, and D. L. Collins, "Automatic Deformable MR - Ultrasound Registration for Image - Guided Neurosurgery," Ieee Transactions on Medic al Imaging, vol. 34, pp. 366 - 380, Feb 2015. [9] Y. Xiao, M. Fortin , G. Unsgard, H. Rivaz, and I. Reinerts en, "REtroSpective Evaluation of Cerebral Tu mors (RESECT ): A clinical database of pre - operative MRI and intra - operative ultrasound in low - grade glioma surgeries," Med Phys, vol. 44, pp. 387 5- 3882, Jul 2017. [10] J. Hong and H. Park, "Non - linear Approach for MRI to intra - operative US Registration Using Structural Skeleton: International Workshops , POCUS 2018, BIVPCS 2018, CuRIOUS 2018, and CPM 2018, Held i n Conj unction with M ICCAI 2018, Gra na da, Spain, September 16 – 20, 2018, Proceedings," ed, 2018, pp. 138 - 145. [11] L. Sun and S. Zhang, "Deformable MRI - Ultrasound Registration Using 3D Convolutional Neural Network: Internati onal Workshops, POCUS 2018, BIVPCS 2018, CuRIOU S 2018, and CPM 2018, He ld in Conjunctio n w ith M ICCAI 2018, Granada, Spain, September 16 – 20, 2018, Proceedings, " ed, 2018, pp. 152 - 158. [12] D. Drobny, T. Vercauteren, S. Ourselin, and M. Modat, "Registration of MRI and iUS Data to Compensate Brain Shift Using a Symmetric Bloc k -M atching Based Approach: International Workshops , POCUS 2018, BIVPCS 2018, CuRIOUS 2018, and CPM 2018, Held i n Conj unction with M ICCAI 2018, Gra nada, Spain, September 16 – 20, 2018, Proceedings," ed, 2018, pp. 172 - 178. [13] I. Machado, M. Toews, J. Luo, P. Un adkat, W. Essayed, E. George , et al. , "Deformable MRI - Ultrasound Registration via Attribute Matchin g and Mutual - Saliency Weighting for Image - Guided Neurosurgery: Int ernational Workshops, POCUS 2018, BIVPCS 2018, CuRIOUS 2018, and CPM 2018, Held in Conjunct ion with MICCAI 2018, Granada, Spain, September 16 – 20, 2018, Proceedings," ed, 2018, pp. 165 - 171. [14] M. P. Heinrich , "Intra - operat ive Ultrasound to M RI Fusion w ith a Public Multimodal Discrete Registration Tool: International Workshops , POCUS 2018, BIVPC S 2018, CuRIOUS 2018, and CPM 2018, Held i n Conj unction with M ICCAI 2018, Gra nada, Spain, September 16 – 20, 2018, Proceedings," ed, 2018, pp. 159 - 164. [15] R. Shams, m. - a. Boucher, and S. Kadou ry, "Intra - operative Brain Shift Correction with Weighted Locall y Linear Correlations of 3DUS and MRI: International Workshops, POCUS 2018, BIVP CS 2018, CuRIOUS 2018, and CPM 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 16 – 20, 2018, Proceedings," ed, 2018, pp. 179 - 184. [16] W. Wein, "Brain - Shif t C orrection with Imag e - Based Registrat ion and Lan dmark A ccuracy E valuation: International Workshops, POCUS 2018, BIVPCS 2018, CuRIOU S 2018, and CPM 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 16 – 20, 2018, Proceedings, " ed, 2018, pp. 146 - 151. [17] X. Zhong, S. Bayer, N. Ravikumar, N. Strobel, A. Birkhold, M. Kowarschik , et al. , "Resolve Intraoperative B rain Shift as Imita tion Game: International Workshops, POCUS 2018, BIVPCS 2018, CuRIOUS 2018, and CPM 2018, Held in Conjunction wit h MICCAI 2018, Granada, Spain, September 16 – 20, 2018, Proceedings," ed, 2018, pp. 129 - 137. [18] A. Reinke, M. Eisenmann, S. Onogur, M. Stanko vic, P. Scholz, P. M. Full , et al. , "How to Exploit Weaknes ses in Biomedical Challenge Desi gn and Organization," Ch am, 2018, pp. 388 - 395. [19] Y. Ou, A. Sotiras, N. Paragios, and C. Davatzikos, "DRAMMS: Deformable regi stration via attribut e matching and mutual - saliency weighting," Med Image Anal , vol. 15, pp. 622 - 39, Aug 2011. [20] Y. Ou, H. Akba ri, M. Bilello, X. Da, and C. Davatzikos, "Comparative ev aluation of registration algorithms in different brain databases with varying diffi culty: results and insights," IEEE Trans Med Imagin g, vol. 33, pp. 2039 - 65, Oct 2014. [21] B. B. Avants, C. L. Epstein, M. Grossman, and J. C. Gee, "Symmetric diffeomorphic imag e registration with cross - correlation: Evaluating automated labeling of elderly and neurodegenerative brain," Medical Image Analy sis, vol. 12, pp. 26 - 41, Feb 2008. [22] A. Roche, G. Mala ndain, X. Pennec , and N. Ayache, "The correlation ratio as a new similarity measure for multimodal image registration," in Medical Image Computing and Computer - Assisted Intervention — MICCAI’98 , Berlin, Heid elberg, 1998, pp. 1115 - 1124. [23 ] D. H. Iversen, W. Wein, F. Lindseth, G. Unsgard, and I. Reinertsen , "Automatic Intraoperative Correction of Brain Shift for Accurate Neuronavigat ion," World Neurosur gery, vol. 120, pp. E1071 - E1078, Dec 2018. [24] D. R. Jones, C. D. Perttunen, and B. E. S tuckman , "Lipschitzian optimization without the L ipschitz constant," Journal of Optimizati on Theory and Applications, vol. 79, pp. 157 - 181, 1993/10/01 1993. [25] M. J. D. Powell, The BOBYQA Algorithm for Bound Constrained Optimizati on without Derivat ives , 2009. [26] M. Modat, D. M. Cash, P. Daga, G. P. Winston, J. S. Duncan, and S. Ourselin, "Global image registration using a symmetric block - matching approach ," Journal of medical imaging (Bellingham, Wash.), vol. 1, pp. 024003 - 024003, 2014. [27] D. Stoyanov , Z. Tay lor, S. Aylward, J . M. R. S. Tavares, Y. Xiao, A. Simpson , et al. , "Simulatio n, Image Proces sing, and Ultraso und Systems for Assisted Diagnosis and Navigation," in POCUS 2018, BIVPCS 2018, CuRIOUS 2018, CMP 2018 , Gran ada, Spain, 2018. [28] Y. M. Xi ao, L. Eikenes, I. Reinertsen, and H. Rivaz, "Nonlinear deformation of tractography in ultrasound - guided low - grade gliomas resection," International Journal of Computer Assisted Radiology and Surger y, vol. 13, pp. 457 - 467, Mar 2018. [29] G. Litjens, T. Koo i, B. E. Bejnordi, A. A . A. Setio, F. Ciompi, M. Ghafoorian , et al. , "A survey on deep learning in medical imag e analysis," Med Image Anal, vol. 42, pp. 60 - 88, Dec 2017. [30] B. B. Avants, N. J. Tustison, G. Song, P. A. Cook, A. Klein, and J. C. Gee, "A re producible evaluation of A NTs similarity m etric performance in brain image registration," Neuroimage, vol. 54, pp. 2033 - 44, Feb 1 2011.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment