Signal2Image Modules in Deep Neural Networks for EEG Classification
Deep learning has revolutionized computer vision utilizing the increased availability of big data and the power of parallel computational units such as graphical processing units. The vast majority of deep learning research is conducted using images …
Authors: Paschalis Bizopoulos, George I Lambrou, Dimitrios Koutsouris
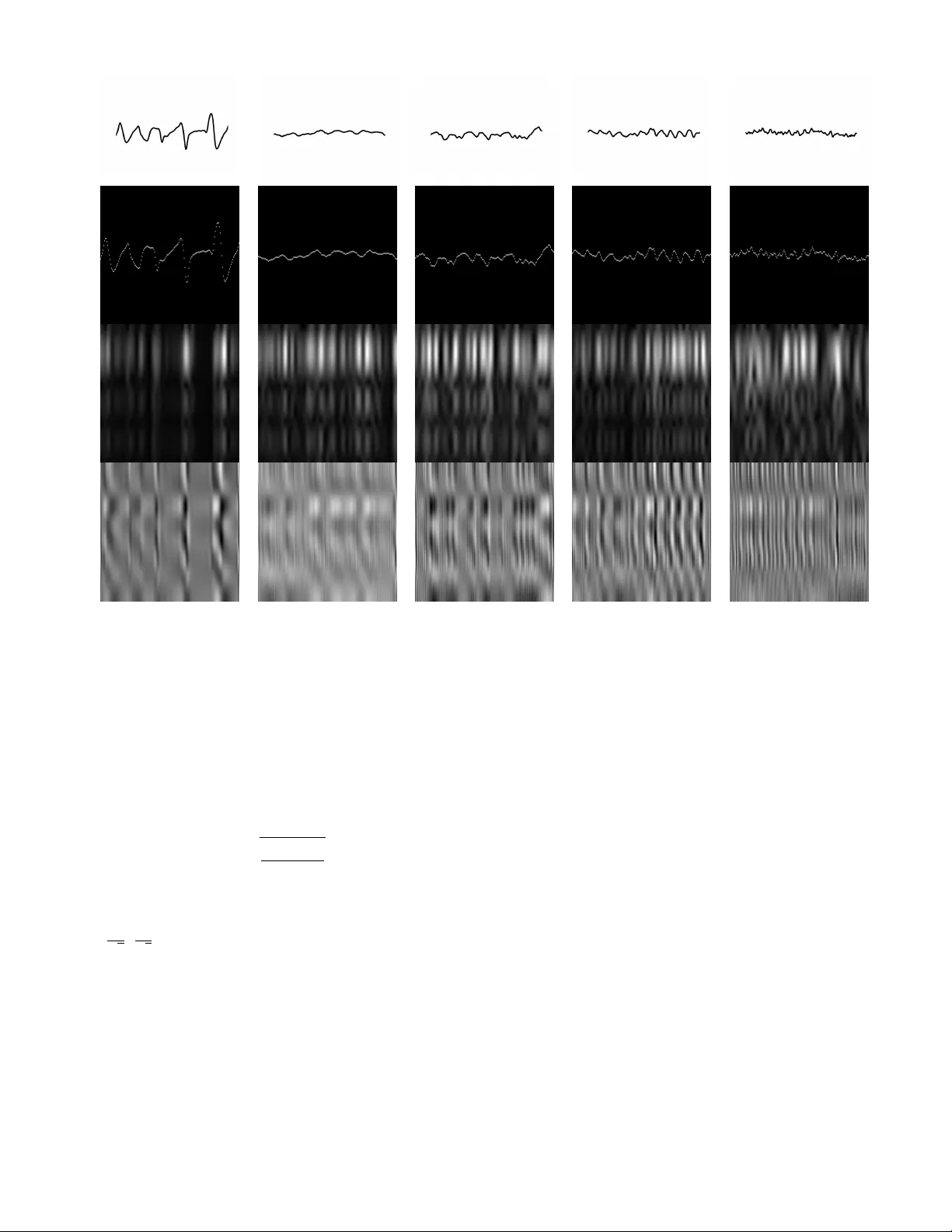
Signal2Image Modules in Deep Neural Networks for EEG Classification Paschalis Bizopoulos, George I. Lambrou and Dimitrios Koutsouris Abstract —Deep learning has r evolutionized computer vision utilizing the increased av ailability of big data and the power of parallel computational units such as graphical processing units. The vast majority of deep learning research is conducted using images as training data, howe ver the biomedical domain is rich in physiological signals that are used for diagnosis and prediction problems. It is still an open research question how to best utilize signals to train deep neural networks. In this paper we define the term Signal2Image (S2Is) as trainable or non-trainable prefix modules that conv ert signals, such as Electroencephalography (EEG), to image-like repr esenta- tions making them suitable for training image-based deep neural networks defined as ‘base models’. W e compare the accuracy and time performance of four S2Is (‘signal as image’, spectrogram, one and two layer Conv olutional Neural Networks (CNNs)) combined with a set of ‘base models’ (LeNet, AlexNet, VGGnet, ResNet, DenseNet) along with the depth-wise and 1D variations of the latter . W e also provide empirical evidence that the one layer CNN S2I perf orms better in ele ven out of fifteen tested models than non-trainable S2Is for classifying EEG signals and present visual comparisons of the outputs of some of the S2Is. I . I N T RO D U C T I O N Most methods for solving biomedical problems until re- cently in volv ed handcrafting features and trying to mimic human experts, which is increasingly prov en to be inefficient and error-prone. Deep learning is emer ging as a po werful so- lution for a wide range of problems in biomedicine achieving superior results compared to traditional machine learning. The main adv antage of methods that use deep learning is that they automatically learn hierarchical features from training data making them scalable and generalizable. This is achieved with the use of multilayer networks, that consist of million parameters [1], trained with backpropagation [2] on large amount of data. Although deep learning is mainly used in biomedical images there is also a wide range of physiological signals, such as Electroencephalography (EEG), that are used for diagnosis and prediction problems. EEG is the measure of the electrical field produced by the brain and is used for sleep pattern classification [3], brain computer interfaces [4], cogni- tiv e/af fectiv e monitoring [5] and epilepsy identification [6]. Y annick et al. [7] re viewed deep learning studies using EEG and hav e identified a general increase in accuracy when deep learning and specifically Conv olutional Neural Networks (CNNs) are used instead of traditional machine learning methods. Howe ver , they do not mention which specific char- acteristics of CNN architectures are indicated to increase The authors are with Biomedical Engineering Laboratory , School of Electrical and Computer Engineering, National T echnical Uni ver- sity of Athens, Athens 15780, Greece e-mail: pbizop@gmail.com , glamprou@med.uoa.gr , dkoutsou@biomed.ntua.gr. performance. It is still an open research question how to best use EEG for training deep learning models. One common approach that previous studies ha ve used for classifying EEG signals was feature extraction from the frequency and time-frequenc y domains utilizing the theory behind EEG band frequencies [8]: delta (0.5–4 Hz), theta (4– 8 Hz), alpha (8–13 Hz), beta (13–20 Hz) and gamma (20– 64 Hz). T ruong et al. [9] used Short-Time Fourier T ransform (STFT) on a 30 second sliding window to train a three layer CNN on stacked time-frequency representations for seizure prediction and ev aluated their method on three EEG databases. Khan et al. [10] transformed the EEGs in time-frequency domain using multi-scale wav elets and then trained a six layer CNN on these stacked multi-scale representations for predicting the focal onset seizure demonstrating promising results. Feature extraction from the time-frequency domain has also been used for other EEG related tasks, besides epileptic seizure prediction. Zhang et al. [11] trained an ensemble of CNNs containing two to ten layers using STFT features extracted from EEG band frequencies for mental w orkload classification. Giri et al. [12] extracted statistical and information measures from frequency domain to train an 1D CNN with two layers to identify ischemic stroke. For the purposes of this paper and for easier future reference we define the term Signal2Image module (S2I) as any module placed after the ra w signal input and before a ‘base model’ which is usually an established architecture for imaging prob- lems. An important property of a S2I is whether it consists of trainable parameters such as conv olutional and linear layers or it is non-trainable such as traditional time-frequency methods. Using this definition we can also deriv e that most previous methods for EEG classification use non-trainable S2Is and that no previous study has compared trainable with non-trainable S2Is. In this paper we compare non-trainable and trainable S2Is combined with well kno wn ‘base models’ neural network architectures along with the 1D and depth-wise variations of the latter . A high lev el overvie w of these combined methods is shown in Fig. 1. Although we choose the EEG epileptic seizure recognition dataset from University of California, Irvine (UCI) [13] for EEG classification, the implications of this study could be generalized in any kind of signal classification problem. Here we also refer to CNN as a neural network consisting of alternating con v olutional layers each one followed by a Rectified Linear Unit (ReLU) and a max pooling layer and a fully connected layer at the end while the term x i m b d ˆ y i Open 0.1% Closed 0.2% Healthy 0.9% T umor 34.7% Epilepsy 64.1% Fig. 1: High level overvie w of a feed-forward pass of the combined methods. x i is the input, m is the Signal2Image module, b d is the 1D or 2D architecture ‘base model’ for d = 1 , 2 respectively and ˆ y i is the predicted output. The names of the classes are depicted at the right along with the predictions for this example signal. The image between m and b d depicts the output of the one layer CNN Signal2Image module, while the ‘signal as image’ and spectrogram have intermediate images as those depicted at the second and third row of Fig. 2. Arro ws denote the flo w of the feed-forward pass. For the 1D architectures m is omitted and no intermediate image is generated. ‘layer’ denotes the number of conv olutional layers. I I . DA T A The UCI EEG epileptic seizure recognition dataset [13] consists of 500 signals each one with 4097 samples (23.5 seconds). The dataset is annotated into fi ve classes with 100 signals for each class (in parenthesis the shortened class names used in Fig. 1 and 2): 1) healthy patient while having his eyes open (Open), 2) healthy patient while having his eyes closed (Closed), 3) patient with tumor taken from healthy area (Healthy), 4) patient with tumor taken from tumor area (T umor), 5) patient while having seizure activity (Epilepsy) For the purposes of this paper we use a variation of the database 1 in which the EEG signals are split into se gments with 178 samples each, resulting in a balanced dataset that consists of 11500 EEG signals. I I I . M E T H O D S A. Definitions W e define the dataset D = { x i , y i } i =1 ...N where x i ∈ Z n and y i ∈ { 1 , 2 , 3 , 4 , 5 } denote the i th input signal with dimensionality n = 178 and the i th class with fi ve possible classes respecti vely . N = 11500 is the number of observ ations. W e also define the set of S2Is as M and the member of this set as m which include the follo wing modules: • ‘signal as image’ (non-trainable) • spectrogram (non-trainable) • one and two layers CNN (trainable) W e then define the set of ‘base models’ as B and the member of this set as b d where d = [1 , 2] denotes the 1 https://archiv e.ics.uci.edu/ml/datasets/Epileptic+Seizure+Recognition dimensionality of the con volutional, max pooling, batch nor- malization and adaptiv e average pooling layers. B includes the following b d along with their depth-wise variations and their equiv alent 1D architectures for d = 1 (for a complete list refer to first two rows of T able. I): • LeNet [14] • AlexNet [1] • VGGnet [15] • ResNet [16] • DenseNet [17] W e finally define the combinations of m and b d as the members c of the set of combined models C . Using the previous definitions, the aim of this paper is the e valuation of the set of models C , where C is the combined set of M and B i.e. C = M × B w .r .t. time performance and class accuracy trained on D . B. Signal2Image Modules In this section we describe the internals of each S2I module. For the ‘signal as image’ module, we normalized the amplitude of x i to the range [1 , 178] . The results were in verted along the y-axis, rounded to the nearest integer and then they were used as the y-indices for the pixels with amplitude 255 on a 178 × 178 image initialized with zeros. For the spectrogram module, which is used for visualizing the change of the frequency of a non-stationary signal ov er time [18], we used a T ukey windo w with a shape parameter of 0 . 25 , a segment length of 8 samples, an o verlap between segments of 4 samples and a fast Fourier transform of 64 samples to conv ert the x i into the time-frequency domain. The resulted spectrogram, which represents the magnitude of the power spectral density ( V 2 /H z ) of x i , was then upsampled to 178 × 178 using bilinear pixel interpolation. For the CNN modules with one and two layers, x i is con verted to an image using learnable parameters instead of some static procedure. The one layer module consists of one 1D con volutional layer (kernel sizes of 3 with 8 channels). The two layer module consists of two 1D conv olutional layers (kernel sizes of 3 with 8 and 16 channels) with the first layer followed by a ReLU activ ation function and a 1D max pooling operation (kernel size of 2 ). The feature maps of the last con v olutional layer for both modules are then concatenated along the y-axis and then resized to 178 × 178 using bilinear interpolation. W e constrain the output for all m to a 178 × 178 image to enable visual comparison. Three identical channels were also stacked for all m outputs to satisfy the input size requirements for b d . Architectures of all b d remained the same, except for the number of the output nodes of the last linear layer which was set to fi ve to correspond to the number of classes of D . An example of the respective outputs of some of the m (the one/two layers CNN produced similar visualizations) are depicted in the second, third and fourth row of Fig. 2. Original Signal ‘Signal as Image’ Spectrogram one layer CNN (a) Open (b) Closed (c) Healthy (d) T umor (e) Epilepsy Fig. 2: V isualizations of the original signals and the outputs of the S2Is for each class. The x, y-axis of the first row are in µ V and time samples respectively . The x, y-axis of the rest of the subfigures denote spatial information, since we do not inform the ‘base model’ the concept of time along the x-axis or the concept of frequenc y along the y-axis. Higher pixel intensity denotes higher amplitude. C. Evaluation Con v olutional layers of m were initialized using Kaiming uniform [19]. V alues are sampled from the uniform distribution U ( − c, c ) , where c is: c = s 6 (1 + a 2 ) k (1) , a in this study is set to zero and k is the size of the input of the layer . The linear layers of m were initialized using U ( − 1 √ k , 1 √ k ) . The con volutional and linear layers of all b d were initialized according to their original implementation. W e used Adam [20] as the optimizer with learning rate lr = 0 . 001 , betas b 1 = 0 . 9 , b 2 = 0 . 999 , epsilon ϵ = 10 − 8 without weight decay and cross entropy as the loss function. Batch size was 20 and no additional regularization was used besides the structural such as dropout layers that some of the ‘base models’ (AlexNet, VGGnet and DenseNet) have. Out of the 11500 signals we used 76% , 12% and 12% of the data ( 8740 , 1380 , 1380 signals) as training, v alidation and test data respectiv ely . No artifact handling or preprocessing was performed. All netw orks were trained for 100 epochs and model selection was performed using the best v alida- tion accuracy out of all the epochs. W e used PyT orch [21] for implementing the neural network architectures and train- ing/preprocessing was done using a NVIDIA Titan X Pascal GPU 12GB RAM and a 12 Core Intel i7–8700 CPU @ 3.20GHz on a Linux operating system. I V . R E S U L T S As shown in T able. I the one layer CNN DenseNet201 achiev ed the best accuracy of 85 . 3% with training time 70 seconds/epoch on av erage. In ov erall the one layer CNN S2I achiev ed best accuracies for eleven out of fifteen ‘base mod- els’. The two layer CNN S2I achiev ed worse ev en compared with the 1D variants, indicating that increase of the S2I depth is not beneficial. The ‘signal as image’ and spectrogram S2Is performed much worse than 1D v ariants and the CNN S2Is. The spectrogram S2I results are in contrary with the expecta- tion that the interpretable time-frequency representation would T ABLE I: T est accuracies (%) for combined models. The second row indicates the number of layers. Bold indicates the best accuracy for each base model. lenet alexnet vgg11 vgg13 vgg16 vgg19 resnet18 resnet34 resnet50 resnet101 resnet152 densenet121 densenet161 densenet169 densenet201 1D, 72.6 78.8 76.9 79.0 79.5 79.3 81.5 82.5 81.4 78.8 81.4 81.8 83.3 82.1 82.0 2D, signal as image 67.9 68.3 74.1 74.7 72.7 72.5 73.3 71.7 74.1 72.3 74.1 74.7 72.5 75.2 75.0 2D, spectrogram 73.2 74.0 77.9 76.3 77.5 76.0 76.2 79.0 77.2 74.6 75.3 74.1 75.2 77.0 75.4 2D, one layer CNN 75.8 82.0 84.0 77.9 80.7 78.4 85.1 84.6 83.0 85.0 83.3 84.3 80.7 85.0 85.3 2D, two layer CNN 75.0 77.9 80.7 78.8 81.1 74.9 78.3 80.0 78.3 77.1 80.9 83.2 82.3 79.0 79.1 help in finding good features for classification. W e hypothesize that the spectrogram S2I was hindered by its lack of non- trainable parameters. Another outcome of these experiments is that increasing the depth of the base models did not increase the accuracy which is inline with previous results [22]. V . C O N C L U S I O N S In this paper we hav e shown empirical e vidence that 1D ‘base model’ variations and trainable S2Is (especially the one layer CNN) perform better than non-trainable S2Is. Ho wev er more work needs to be done for full replacing non-trainable S2Is, not only from the scope of achieving higher accuracy results but also increasing the interpretability of the model. Another point of reference is that the combined models were trained from scratch based on the hypothesis that pretrained low lev el features of the ‘base models’ might not be suitable for spectrogram-like images such as those created by S2Is. Future work could include testing this hypothesis by initializ- ing a ‘base model’ using transfer learning or other initializa- tion methods. Moreover , trainable S2Is and 1D ‘base model’ variations could also be used for other physiological signals besides EEG such as Electrocardiography , Electromyography and Galvanic Skin Response. AC K N OW L E D G M E N T This work was supported by the European Union’ s Hori- zon 2020 research and innovation programme under Grant agreement 769574. W e gratefully acknowledge the support of NVIDIA with the donation of the Titan X Pascal GPU used for this research. R E F E R E N C E S [1] A. Krizhevsky , I. Sutske ver , and G. E. Hinton, “Imagenet classification with deep conv olutional neural networks, ” in Advances in neural infor- mation pr ocessing systems , 2012, pp. 1097–1105. [2] D. E. Rumelhart, G. E. Hinton, and R. J. Williams, “Learning represen- tations by back-propagating errors, ” nature , vol. 323, no. 6088, p. 533, 1986. [3] K. Aboalayon, M. Faezipour , W . Almuhammadi, and S. Moslehpour , “Sleep stage classification using eeg signal analysis: a comprehensive survey and new inv estigation, ” Entropy , vol. 18, no. 9, p. 272, 2016. [4] A. Al-Nafjan, M. Hosny , Y . Al-Ohali, and A. Al-W abil, “Revie w and classification of emotion recognition based on eeg brain-computer interface system research: a systematic revie w , ” Applied Sciences , vol. 7, no. 12, p. 1239, 2017. [5] F . Lotte, L. Bougrain, and M. Clerc, “Electroencephalography (eeg)- based brain–computer interfaces, ” W iley Encyclopedia of Electrical and Electr onics Engineering , pp. 1–20, 1999. [6] U. R. Acharya, S. V . Sree, G. Swapna, R. J. Martis, and J. S. Suri, “ Automated eeg analysis of epilepsy: a revie w , ” Knowledge-Based Systems , vol. 45, pp. 147–165, 2013. [7] R. Y annick, B. Hubert, A. Isabela, G. Alexandre, F . Jocelyn et al. , “Deep learning-based electroencephalography analysis: a systematic revie w , ” arXiv pr eprint arXiv:1901.05498 , 2019. [8] M. L ¨ angkvist, L. Karlsson, and A. Loutfi, “Sleep stage classification using unsupervised feature learning, ” Advances in Artificial Neural Systems , vol. 2012, p. 5, 2012. [9] N. D. Truong, A. D. Nguyen, L. Kuhlmann, M. R. Bonyadi, J. Y ang, S. Ippolito, and O. Kavehei, “Conv olutional neural networks for seizure prediction using intracranial and scalp electroencephalogram, ” Neural Networks , vol. 105, pp. 104–111, 2018. [10] H. Khan, L. Marcuse, M. Fields, K. Swann, and B. Y ener , “Focal onset seizure prediction using con volutional networks, ” IEEE Tr ansactions on Biomedical Engineering , vol. 65, no. 9, pp. 2109–2118, 2018. [11] J. Zhang, S. Li, and R. W ang, “Pattern recognition of momentary mental workload based on multi-channel electrophysiological data and ensemble con volutional neural networks, ” F rontier s in neuroscience , vol. 11, p. 310, 2017. [12] E. P . Giri, M. I. Fanany , A. M. Arymurthy , and S. K. Wijaya, “Ischemic stroke identification based on eeg and eog using id conv olutional neural network and batch normalization, ” in Advanced Computer Science and Information Systems (ICACSIS), 2016 International Conference on . IEEE, 2016, pp. 484–491. [13] R. G. Andrzejak, K. Lehnertz, F . Mormann, C. Riek e, P . David, and C. E. Elger , “Indications of nonlinear deterministic and finite-dimensional structures in time series of brain electrical activity: Dependence on recording region and brain state, ” Physical Revie w E , vol. 64, no. 6, p. 061907, 2001. [14] Y . LeCun, L. Bottou, Y . Bengio, and P . Haf fner , “Gradient-based learning applied to document recognition, ” Proceedings of the IEEE , vol. 86, no. 11, pp. 2278–2324, 1998. [15] K. Simonyan and A. Zisserman, “V ery deep con volutional networks for large-scale image recognition, ” arXiv preprint , 2014. [16] K. He, X. Zhang, S. Ren, and J. Sun, “Deep residual learning for image recognition, ” in Pr oceedings of the IEEE conference on computer vision and pattern r ecognition , 2016, pp. 770–778. [17] G. Huang, Z. Liu, L. V an Der Maaten, and K. Q. W einberger, “Densely connected conv olutional networks, ” in Pr oceedings of the IEEE confer- ence on computer vision and pattern recognition , 2017, pp. 4700–4708. [18] A. V . Oppenheim, Discrete-time signal pr ocessing . Pearson Education India, 1999. [19] K. He, X. Zhang, S. Ren, and J. Sun, “Delving deep into rectifiers: Surpassing human-level performance on imagenet classification, ” in Pr oceedings of the IEEE international confer ence on computer vision , 2015, pp. 1026–1034. [20] D. P . Kingma and J. Ba, “ Adam: A method for stochastic optimization, ” arXiv pr eprint arXiv:1412.6980 , 2014. [21] A. Paszke, S. Gross, S. Chintala, G. Chanan, E. Y ang, Z. DeV ito, Z. Lin, A. Desmaison, L. Antiga, and A. Lerer , “ Automatic differentiation in pytorch, ” 2017. [22] R. T . Schirrmeister , J. T . Springenberg, L. D. J. Fiederer, M. Glasstetter, K. Eggensperger , M. T angermann, F . Hutter, W . Burgard, and T . Ball, “Deep learning with con volutional neural networks for eeg decoding and visualization, ” Human brain mapping , vol. 38, no. 11, pp. 5391–5420, 2017.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment