A Multimodal Deep Network for the Reconstruction of T2W MR Images
Multiple sclerosis is one of the most common chronic neurological diseases affecting the central nervous system. Lesions produced by the MS can be observed through two modalities of magnetic resonance (MR), known as T2W and FLAIR sequences, both prov…
Authors: Antonio Falvo, Danilo Comminiello, Simone Scardapane
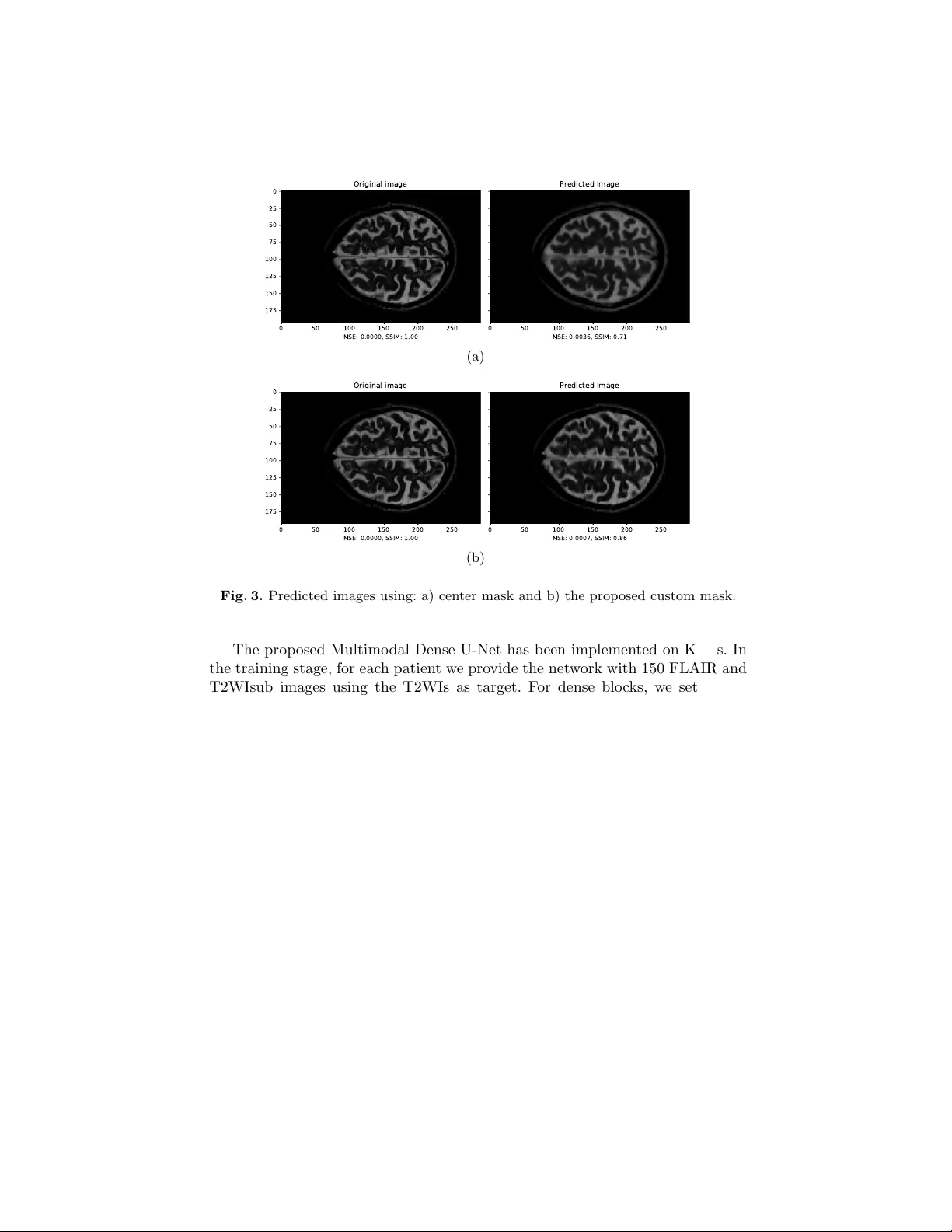
A Multimo dal Deep Net w ork for the Reconstruction of T2W MR Images An tonio F alv o, Danilo Comminiello, Simone Scardapane, Mic hele Scarpiniti, and Aurelio Uncini Dept. Information Eng., Electronics and T elecommunications (DIET) Sapienza Universit y of Rome, Via Eudossiana 18, 00184 Rome, Italy danilo.comminiello@uniroma1.it Abstract. Multiple sclerosis is one of the most common c hronic neuro- logical diseases affecting the cen tral nerv ous system. Lesions produced b y the MS can b e observ ed through tw o modalities of magnetic resonance (MR), kno wn as T2W and FLAIR sequences, b oth providing useful infor- mation for form ulating a diagnosis. How ever, long acquisition time makes the acquired MR image vulnerable to motion artifacts. This leads to the need of accelerating the execution of the MR analysis. In this pap er, we presen t a deep learning metho d that is able to reconstruct subsampled MR images obtained by reducing the k -space data, while maintaining a high image quality that can b e used to observe brain lesions. The pro- p osed metho d exploits the multimodal approach of neural netw orks and it also fo cuses on the data acquisition and processing stages to reduce execution time of the MR analysis. Results pro ve the effectiv eness of the prop osed metho d in reconstructing subsampled MR images while sa ving execution time. Keyw ords: magnetic resonance imaging, fast MRI, m ultiple sclerosis, deep neural netw ork 1 In tro duction Nuclear magnetic resonance (NMR) is a transmission analysis tec hnique that allo ws to obtain information on the state of matter, exploiting the in teraction b et w een magnetic fields and atoms n uclei. In the biomedical field, information deriving from the NMR is represen ted in the form of tomographic images. No wa- da ys, the NMR plays an imp ortan t role in the health field, and it allows to carry out a whole typology of diagnostic exams, from traditional to functional neuro- radiology , from in ternal diagnostic to obstetrics and p ediatric diagnostics [1]. During the acquisition stage of an MR signal, it is necessary to sample the en tire k -space to obtain images that are as m uch detailed as possible [4, 10]. Data in the k -space enco de information on spatial frequencies and are generally captured line by line. Therefore, the acquisition time for a given sequence de- p ends on the n umber of lines sampled in the k -space, th us leading to a rather slo w acquisition pro cess. Moreov er, significan t artifacts ma y o ccur in the MR 2 F alvo et al. images caused by to slow mov ements of the patien t, due to physiological factors or to fatigue, e.g., to o muc h time in the same p osition [4, 10]. Moreov er, the long scan time also increases the healthcare cost for the patient, b esides limiting the a v ailability of MR scanners. Ov er the years, several metho ds, suc h as compressed magnetic resonance and parallel magnetic resonance [3, 7, 11, 12], ha ve b een proposed to accelerate MRI scans b y skipping the k -space phase co ding lines and a void the aliasing phenomenon introduced by subsampling. The problem of accelerating magnetic resonance can also be tackled through deep learning tec hniques. In particular, the reconstruction of tomographic images has been often efficiently addressed b y using conv olutional neural netw orks (CNNs) [8, 13, 14, 16, 17]. Most of the state-of-the-art metho ds fo cus on the reconstruction of MR im- ages using a unimodal neural architecture where a subsampled image to b e reconstructed is pro vided as input. In this pap er, w e prop ose a new deep learn- ing metho d for reconstructing MR images by exploiting additional information pro vided b y FLAIR images. Such images are widely used in the MR diagnosis as they allow to enhance the brain lesions due to the disease. FLAIR images are highly correlated with the T2 w eighted images (T2WIs), thus the joint use of such images increases the efficiency of the reconstruction and also presents m uch more information in the lesion region. T o exploit b oth the images, we pro- p ose a m ultimo dal deep neural netw ork, inspired by the w ell-kno wn U-Net, a con volutional-based mo del that w as dev elop ed for biomedical image segmenta- tion [15]. In the l iterature, sev eral studies hav e b een prop osed using a multimodal approac h for image reconstruction. In [19], T2WIs were attempt to b e estimated from T1WI, while other w orks fo cus on impro ving the quality of the subsam- pled images with the help of high resolution images with different contrast [6, 9]. Ho wev er, to the b est of our knowledge, no attempt has ever b een made to re- construct T2WIs from subsampled T2WIs (T2WIsub) and FLAIR images while main taining high image qualit y in the area of lesions. Exp erimen tal results prov e that the proposed method is able to accelerate the MR analysis four times, while preserving the image qualit y , with a high detail of any lesion and negligible aliasing artifacts. The rest of the pap er is organized as follows. In Section 2, we introduce the prop osed approac h, including a new subsampling mask, while the prop osed Multimo dal Dense U-Net is presented in Section 3. Results are shown in Section 4 and, finally , our conclusion is drawn in Section 5. 2 Prop osed Approac h: Main Definitions W e first fo cus on the images to b e pro vided as input in order to reconstruct the T2WI. 2.1 Problem F orm ulation W e denote with X T 2 the k -space for the T2WI that represen ts the target. Mul- tiplying the k -space X T 2 for a suitably designed mask M , it is p ossible to obtain Multimo dal Deep Netw ork 3 a subsampled v ersion of the k -space, i.e., X T 2 sub = M · X T 2 (1) The bidimensional inv erse F ourier transform allows to achiev e data in to the space domain. Therefore, we define the fully-sampled target image Y T 2 and the subsampled T2 image Y T 2 sub to be used for reconstruction through the proposed deep netw ork. Finally , we denote the FLAIR image to b e provided as input with Y F . W e w ant to reconstruct the fully-sampled T2 image Y T 2 , giv en the only a v ailability of subsampled Y T 2 sub and Y F . The reconstructed T2 image is denoted as ˆ Y T 2 . T o this end, w e build and train a deep netw ork to minimize the following loss function: arg min { MSE + DSSIM } (2) in whic h the MSE denotes the mean-square error and DSSIM the structural dissimilarit y index. The former is defined as: MSE = 1 N N X i =1 Y T 2 ,i − ˆ Y T 2 ,i 2 . (3) On the other hand, the DSSIM is complementary to the structural similarity index (SSIM), whic h is often adopted to assess the p erceiv ed qualit y of television and film images as well as other types of digital images and videos. It was designed to improv e traditional metho ds such as the signal-to-p eak noise ratio (PSNR) and the mean square error (MSE), and it is defined as: DSSIM Y T 2 , ˆ Y T 2 = 1 2 − 2 µ Y µ ˆ Y + c 1 2 σ Y ˆ Y + c 2 2 µ 2 Y + µ 2 ˆ Y + c 1 σ 2 Y + σ 2 ˆ Y + c 2 (4) where µ Y , µ ˆ Y represen t the mean v alues, σ 2 Y and σ 2 ˆ Y the v ariances, and σ Y ˆ Y the co v ariance. 2.2 Customization of a New Subsampling Mask Most of the existing literature dealing with MRI acceleration mainly focuses on the reconstruction of images. How ever, the quality of the reconstruction dep ends significan tly on ho w k -space is sampled. This problem can b e faced essentially by adopting one of the following approaches: 1) a dynamic appr o ach based on deep learning in which cells of fixed width are allo wed to mov e in the k -space and c hange p osition based on the reconstruction performance; 2) a static appr o ach in whic h fixed sampling masks are used that go to select only certain areas of the k -space. In this work, we choose the static approach since the dynamic one do es not guaran tee that the p ow er sp ectrum of a reference image is similar to that of the test image. The adopted sampling method consists of a mask that acts along 4 F alvo et al. Kx 50 100 150 200 250 Ky 20 40 60 80 100 120 140 160 180 (a) Kx 50 100 150 200 250 Ky 20 40 60 80 100 120 140 160 180 (b) Fig. 1. Subsampling masks: a) center mask and b) the prop osed custom mask. the direction of the phase co ding of the k -space, in whic h it is p ossible, once the subsampling factor is set, to c ho ose the p ercen tage of samples that will o ccup y the central part of k -space, thus leaving the rest of the samples equidistant from eac h other. Figure 1 shows tw o different types of mask b oth obtained by setting a subsam- pling factor k = 4. In the center mask of Fig. 1(a), samples are taken exclusively in the central area of the k -space, where most of the low-frequency comp onents can b e found pro viding useful information on the contrast of the image [19]. Ho wev er, in this w ork, we prop ose a new mask, depicted in Fig. 1(b), which selects the 80% of the total samples from the center and the remainder in an equidistan t manner so as to ha ve information even in the high frequencies. 3 Multimo dal Dense U-Net The prop osed neural net work is multimodal architecture: on one branc h we pro- vide the T2WIsub as input while on the other branc h we provide the FLAIR image to be used to impro ve the reconstruction quality . W e exp ect all the spatial information in the FLAIR image to help estimate the anatomical structures in T2WI. Both inputs initially undergo separate con traction transformations to then merge later and follow the classic coding-deco ding approach of U-Net models. The prop osed Multimo dal Dense U-Net is depicted in Fig. 2. The netw ork consists essentially of 4 comp onen ts, namely conv olutive la yers, p ooling lay ers, decon volutiv e la y ers and dense blo c ks. The size of the charac- teristic map decreases along the contraction path through the p ooling blo c ks as it increases along the expansion path by decon v olution. Pooling partitions the input image into a set of squares, and for eac h of the resulting regions re- turns the maxim um v alue as output. Its purp ose is to progressively reduce the size of the representations, so as to reduce the num b er of parameters and the computational complexit y of the netw ork, at the same time counteracting any o verfitting. Multimo dal Deep Netw ork 5 C O N V O L U T I O N 3 X 3 P O O L I N G 2 X 2 D E C O N V O L U T I O N 2 X 2 C O N C A T E N A T E M E R G E P O O L I N G F E A T U R E S C O N V . F E A T U R E S D E C O N V . F E A T U R E S D E N S E B L O C K 6 4 T 2 W I _ s u b F L A I R T 2 W I C O N V 1 6 4 6 4 6 4 6 4 6 4 6 4 6 4 6 4 6 4 6 4 6 4 Fig. 2. Scheme of the prop osed Multimo dal Dense U-Net architecture. Decon volutiv e la y ers act inv ersely with respect to p o oling and aim to increase the spatial dimensions of the inputs. This allows to obtain images of a size com- parable to those of the input images from the net w ork. In the simplest case these lev els can b e implemented as static ov ersampling with bilinear interpolation. The dense block, prop osed in [5], allo ws to effectiv ely increase the depth of the entire netw ork while main taining a low complexity . Moreo ver, it requires less parameters to b e trained. The dense blo c k consists of three consecutiv e op erations: batc h normalization (BN) , ELUs activ ation functions [2] and 3 × 3 con volution filters. The hyper-parameters for the dense blo ck are the gro wth rate (GR) and the n umber of con volutional la yers (NC). The netw ork ends with a reconstruction level consisting of a dense blo c k follow ed by a 1 × 1 conv olutional la yer that yields the reconstructed T2WI. 4 Exp erimen tal Results 4.1 Dataset and Netw ork Setting W e test the prop osed net work on a dataset con taining MRIs of m ultiple sclerosis patien ts [18]. In particular, the dataset is related to 30 patients and it con tains axial 2D-T1W, 2D-T2W and 3D-FLAIR images. The final v oxel size of suc h images is 0 . 46 × 0 . 46 × 0 . 8 mm 3 . In our work, a further prepro cessing has b een p erformed in MA TLAB to make the vo xel size isotropic to 0 . 8 × 0 . 8 × 0 . 8 mm 3 , to extract slices of size 192 × 292, and to shrink in tensity to the range [0 , 1]. T2WIsub images were created by considering tw o types of masks, the center mask and the prop osed custom mask, with a subsampling factor k = 4. 6 F alvo et al. 0 50 100 150 200 250 MSE: 0.0000, SSIM: 1.00 0 25 50 75 100 125 150 175 Original image 0 50 100 150 200 250 MSE: 0.0036, SSIM: 0.71 Predicted Image (a) 0 50 100 150 200 250 MSE: 0.0000, SSIM: 1.00 0 25 50 75 100 125 150 175 Original image 0 50 100 150 200 250 MSE: 0.0007, SSIM: 0.86 Predicted Image (b) Fig. 3. Predicted images using: a) cen ter mask and b) the prop osed custom mask. The prop osed Multimo dal Dense U-Net has b een implemented on Keras. In the training stage, for each patient w e provide the netw ork with 150 FLAIR and T2WIsub images using the T2WIs as target. F or dense blo c ks, we set a zero gro wth rate and a num b er of levels equal to 5 with feature maps size of 64 and ELU activ ation lev els. W e use Adam as an optimizer for training. A total of 80 ep ochs are p erformed, with early stopping. The duration of each ep och is ab out 15 minutes, having set a batch size of 4 and using a desktop PC with an In tel Core i5 6600-K 3.50 GHz CPU, 16 GB of RAM and NVIDIA GeF orce GTX 970 GPU. T o quan titatively ev aluate reconstruction p erformance, w e use the MSE and DSSIM metrics. 4.2 Ev aluation of the Proposed Mask W e wan t to ev aluate first the effectiveness of the prop osed custom subsampling mask compared to the center mask on the quality of reconstruction in terms of the SSIM using a Dense U-Net net work. Results are shown in Fig. 3, where it is clear that the prop osed custom mask allo ws us to obtain a reconstruction of the image with outstanding p erformance. In particular, using the cen ter mask we get a 71% reconstruction p ercen tage com- pared to the target (Fig. 3(a)), while using the proposed custom mask (Fig. 3(b)) the similarity index rises up to 86%. Multimo dal Deep Netw ork 7 Fig. 4. Predicted T2WI reconstructed by the prop osed Multimo dal Dense U-Net. 0 20000 40000 60000 80000 100000 Iteration 10 − 1 10 0 Loss Fig. 5. Loss function b eha vior. 4.3 Ev aluation of the Proposed Deep Arc hitecture Conceptually , the prop osed arc hitecture and the standard Dense U-Net might app ear similar, but the former one manages the tw o inputs differently . Moreo ver, the h yp er-parameters of the dense blo c ks chosen for our netw ork considerably c hange the concept of dense blo ck as the whole of gro wth was set to zero thus a voiding internal expansion in dense blo c ks. W e compare the reconstruction quality of the tw o netw orks in terms of SSIM ha ving used the mask that provided the best p erformance for the subsampling, i.e., the prop osed custom mask. Results are shown in Fig. 4, where it is clear that the quality of reconstruction has been considerably impro ved compared to Dense U-Net. In particular, the degree of similarity with resp ect to the target is 94% rather than 86% of the Dense U-Net. By using the prop osed arc hitecture, high image qualit y is achiev ed, thus enabling the recognition of brain injuries caused by the disease. W e also show the loss function b eha vior for the prop osed metho d in Fig. 5. 8 F alvo et al. 5 Conclusion In this w ork, we prop ose a deep learning model exploiting the capabilities of b oth m ultimo dal netw orks and dense blocks. In particular, the prop osed approach allo ws to reconstruct T2WIs, subsampled by a factor of 4, thus leveraging the correlation that exists with FLAIR images. At the same time, the prop osed metho d is able to maintain a high quality of image reconstruction, in particular in the area of the brain lesions due to multiple sclerosis. The comparison with a state-of-the-art Dense U-Net architecture has shown that the prop osed netw ork outp erforms both in terms of p erceptiv e quality and in terms of execution times. F uture w orks will fo cus on increasing the sp eed of the MRI scan, with the goal of achieving an acceleration of at least 10 times, and on further impro ving the reconstruction quality . References 1. Beall, P .T., Amtey , S.R., Kasturi, S.R.: NMR Data Handbo ok for Biomedical Ap- plications. Pergamon Bo oks Inc., Elmsford, NY (1984) 2. Clev ert, D.A., Unterthiner, T., Ho c hreiter, S.: F ast and accurate deep netw ork learning by exp onen tial linear units (ELUs). In: In ternational Conference on Learn- ing Representations (ICLR). pp. 1–14. San Juan, Puerto Rico (Ma y 2016) 3. Gamp er, U., Boesiger, P ., Kozerke, S.: Compressed sensing in dynamic MRI. Mag- netic Resonance in Medicine 59(2), 365–373 (F eb 2008) 4. Haac ke, E.M., Bro wn, R.W., Thompson, M.R., V enk atesan, R.: Magnetic Reso- nance Imaging: Physical Principles and Sequence Design, v ol. 82. Wiley-Liss, New Y ork, NY (1999) 5. Huang, G., Liu, Z., v an der Maaten, L., W ein b erger, K.Q.: Densely connected con volutional net works. In: IEEE Conference on Computer Vision and P attern Recognition (CVPR). pp. 2261–2269. Honolulu, HI (Jul 2017) 6. Huang, J., Chen, C., Axel, L.: F ast multi-con trast MRI reconstruction. Magnetic Resonance Imaging 32(10), 1344–1352 (Dec 2014) 7. Jaspan, O.N., Fleysher, R., Lipton, M.L.: Compressed sensing MRI: A review of the clinical literature. The British Journal of Radiology 88(1056) (Dec 2015) 8. Jin, K.H., McCann, M.T., F roustey , E., Unser, M.: Deep conv olutional neural net- w ork for inv erse problems in imaging. IEEE T ransactions on Image Pro cessing 26(9), 4509–4522 (Sep 2017) 9. Kim, K.H., Do, W.J., Park, S.H.: Impro ving resolution of MR images with an adv ersarial netw ork incorp orating images with different contrast. Medical Physics 45(7), 3120–3131 (Jul 2018) 10. Liang, Z.P ., Lauterbur, P .C.: Principles of Magnetic Resonance Imaging. A Signal Pro cessing Perspective. The Institute of electrical and Electronics Engineers, New Y ork, NY (2000) 11. Lustig, M., Donoho, D., Pauly , J.M.: Sparse MRI: The application of compressed sensing for rapid MR imaging. Magnetic Resonance in Medicine 58, 1182–1195 (Oct 2007) 12. Lustig, M., Donoho, D.L., Santos, J.M., P auly , J.M.: Compressed sensing MRI. IEEE Signal Pro cessing Magazine 25(2), 72–82 (Mar 2008) Multimo dal Deep Netw ork 9 13. McCann, M.T., Jin, K.H., Unser, M.: Conv olutional neural netw orks for in verse problems in imaging: A review. IEEE Signal Pro cessing Magazine 34(6), 85–95 (No v 2017) 14. Qin, C., Schlemper, J., Caballero, J., Price, A.N., Ha jnal, J.V., Rueck ert, D.: Con- v olutional recurrent neural netw orks for dynamic MR image reconstruction. IEEE T ransactions on Medical Imaging 38(1), 280–290 (Jan 2019) 15. Ronnen b erger, O., Fischer, P ., Bro x, T.: U-Net: Conv olutional netw orks for biomedical image segmentation. In: International Conference on Medical Image Computing and Computer-Assisted In terven tion (MICCAI). Lecture Notes in Computer Science, vol. 9351, pp. 234–241. Springer, Cham (2015) 16. Ro y , S., Butman, J.A., Reic h, D.S., Calabresi, P .A., Pham, D.L.: Multiple sclerosis lesion segmentation from brain MRI via fully con volutional neural net works. arXiv preprin t arXiv:1803.09172v1 (Mar 2018) 17. Sc hlemp er, J., Caballero, J., Ha jnal, J.V., Price, A.N., Rueck ert, D.: A deep cas- cade of conv olutional neural netw orks for dynamic MR image reconstruction. IEEE T ransactions on Medical Imaging 37(2), 491–503 (F eb 2018) 18. ˇ Ziga, L., Galimzianov a, A., Koren, A., Lukin, M., Pern u ˇ s, F., Lik ar, B., ˇ Spiclin, v.: A nov el public MR image dataset of multiple sclerosis patients with lesion segmen tations based on m ulti-rater consensus. Neuroinformatics 16(1), 51–63 (Jan 2018) 19. Xiang, L., Chen, Y., Chang, W., Zhan, Y., Lin, W., W ang, Q., Shen, D.: Deep leaning based multi-modal fusion for fast MR reconstruction. IEEE T ransactions on Biomedical Engineering (Early Access) (2018)
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment