Exact Bayesian inference for discretely observed Markov Jump Processes using finite rate matrices
We present new methodologies for Bayesian inference on the rate parameters of a discretely observed continuous-time Markov jump processes with a countably infinite state space. The usual method of choice for inference, particle Markov chain Monte Car…
Authors: Chris Sherlock, Andrew Golightly
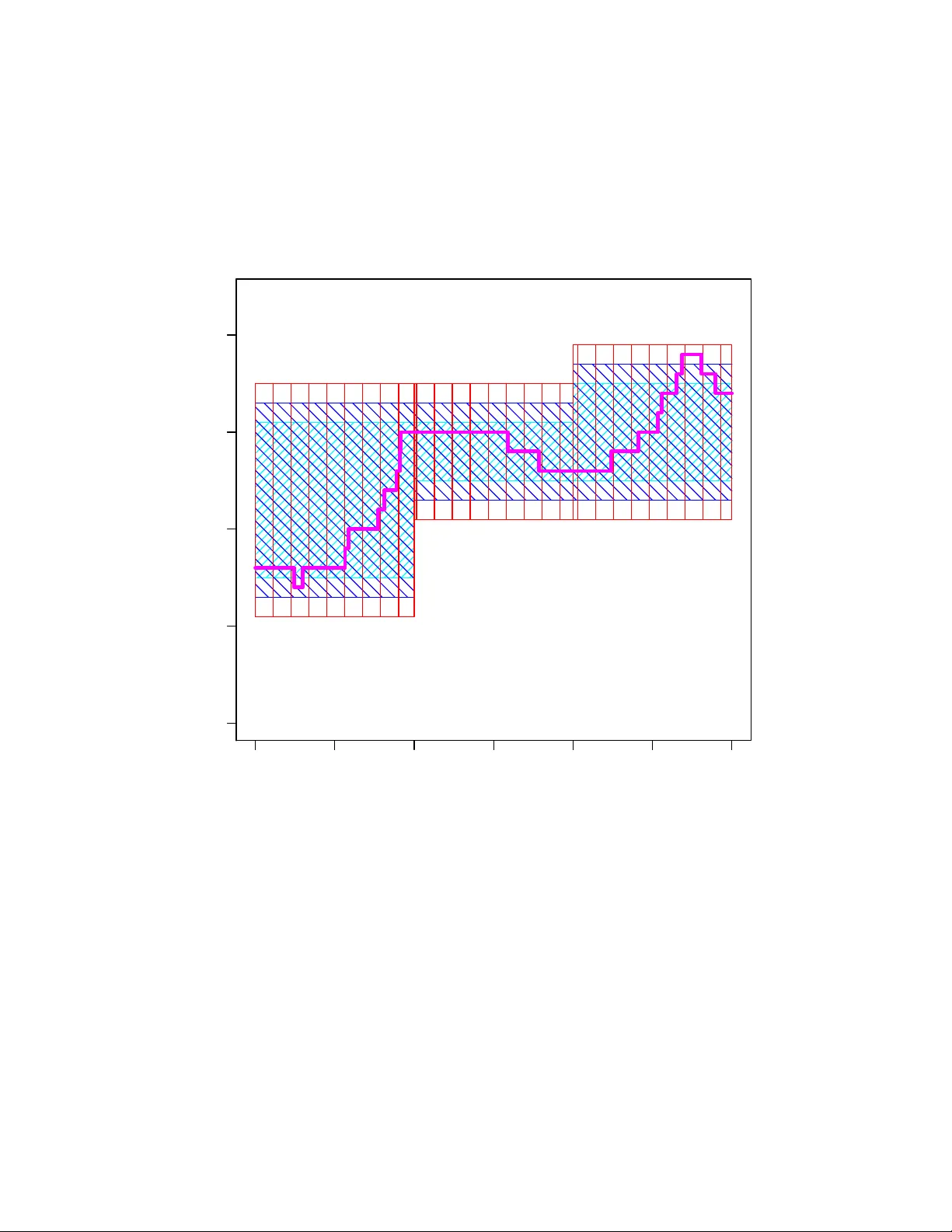
Exact Ba y esian inference for discretely observ ed Mark o v Jump Pro cesses using finite rate matrices Chris Sherlo c k 1 ∗ and Andrew Goligh tly 2 1 Departmen t of Mathematics and Statistics , Lancaster Univ ersity , UK 2 Departmen t of Mathematical Sciences, Durham Univ ersity , UK Abstract W e presen t new metho dologies for Ba yesian inference on the rate parameters of a discretely observ ed contin uous-time Marko v jump pro cess with a countably infinite statespace. The usual method of choice for inference, particle Mark ov chain Mon te Carlo (particle MCMC), struggles when the observ ation noise is small. W e consider the most c hallenging regime of exact observ ations and provide t wo new metho dologies for inference in this case: the minimal extended statespace algorithm (MESA) and the nearly minimal extended statespace algorithm (nMESA). By extending the Mark ov c hain Monte Carlo statespace, b oth MESA and nMESA use the exponentiation of finite rate matrices to p erform exact Bay esian inference on the Marko v jump pro cess ev en though its statespace is countably infinite. Numerical experiments sho w improv ements o ver particle MCMC of betw een a factor of three and several orders of magnitude. Keywor ds: MCMC, contin uous-time Mark ov c hain, coffin state, correlated pseudo-marginal. ∗ c.sherlo c k@lancaster.ac.uk 1 1 In tro duction W e consider exact inference for Marko v jump pro cesses (MJPs), contin uous-time Marko v c hains which arise from reaction netw orks: sto c hastic mo dels for the joint ev olution of one or more p opulations of sp e cies . These sp ecies may be biological sp ecies (e.g. Wilkinson, 2018), animal species (e.g. Dro v andi and McCutchan, 2016), in teracting groups of individuals at v arious stages of a disease (e.g. Andersson and Britton, 2000), or coun ts of sub-p opulations of alleles (Moran, 1958), for example. The state of the system is encapsulated by the n umber of eac h sp ecies that is presen t, and in man y cases this n umber is unbounded and, consequen tly , the statespace is coun tably infinite. The system evolv es via a set of r e actions whose rates dep end on the current state. Section 1.1 describ es three examples of reaction net works. The n umber of possible ‘next’ states giv en the current state is bounded b y the num b er of reactions, whic h is typically small; thus the infinitesimal generator of the process, whic h can b e viewed as a coun tably infinite ‘matrix’, is sparse. The metho ds which w e dev elop in this article can b e applied to any MJP , but they are particularly effective when the generator of the MJP is sparse. The usual metho d of c hoice for inference on discretely observed MJPs with a countably infinite statespace is particle Mark o v c hain Mon te Carlo (particle MCMC, Andrieu et al., 2010) using a b o otstrap particle filter (e.g. Andrieu et al., 2009; Golightly and Wilkinson, 2011; McKinley et al., 2014; Ow en et al., 2015; Koblen ts and Miguez, 2015; Wilkinson, 2018). P aths from the prior distribution of the pro cess are simulated and then resampled according to weigh ts that dep end on the lik eliho o d of the next observ ation given the simulated path. T ypically , as the precision of an observ ation increases, its compatibilit y with most of the paths plummets, leading to lo w w eights; consequen tly , the efficiency of bo otstrap particle MCMC decreases substan tially . W e consider the situation that is most c hallenging of all for a particle filter: when the observ ations are exact. Recen tly , paths prop osed from alternativ e sto c hastic pro cesses whic h tak e the next observ ation in to account ha v e successfully mitigated against this issue within particle MCMC (Goligh tly and Wilkinson, 2015; Golightly and Sherlo c k, 2019), alb eit at an increased computational cost. The first of these pro vides the primary particle-filter comparator for the very differen t inference metho dology , inspired by Georgoulas et al. (2017), that we will in tro duce. The likelihoo d for a discretely observed con tinuous-time Mark o v chain with a large but finite statespace and a rate matrix (or infinit esimal generator) Q is the pro duct of a set of transition probabilities, each of whic h requires the ev aluation of v > e Qt for an inter-observ ation time t and a non-negativ e v ector v representing the state at an observ ation time. F ast algorithms for exactly this calculation, some sp ecifically designed for sparse Q , are av ailable (e.g. Sidje, 1998; Sidje and Stew art, 1999; Moler and V an Loan, 2003; Al-Moh y and Higham, 2011) and some are applicable ev en when the num b er of p ossible states, d , is in the tens of thousands; how ev er, man y pro cesses of in terest hav e a coun tably infinite n um b er of states, and an exact, matrix- exp onen tial approac h might appear imp ossible for suc h systems. Refuting this conjecture, Georgoulas et al. (2017) describes an ingenious pseudo-marginal MCMC algorithm (Andrieu and Rob erts, 2009) that uses random truncations (e.g. McLeish, 2011; Glynn and Rhee, 2014) and coffin states to explore the parameter p osteriors for MJPs with infinite statespaces 2 using exp onentials of finite rate matrices. Unlik e other pseudo-marginal algorithms whic h use random truncation (e.g. Lyne et al., 2015), the algorithm of Georgoulas et al. (2017) is guaranteed to produce un biased estimates of the lik eliho o d which are non-negative. The algorithm, ho w ever, suffers from several issues: the most imp ortant arises from the need to use a prop osal distribution for the truncation lev el (see Section 2.2). As a result, in some examples the algorithm is less efficient than the most appropriate particle MCMC algorithm (see Section 4). W e describ e the minimal extended statespace algorithm (MESA) and the nearly minimal extended statespace algorithm (nMESA), inspired b y the key nov el idea in Georgoulas et al. (2017). These are fast and efficient algorithms for exact inference on Marko v jump pro- cesses with infinite statespaces through exp onentiation of finite-dimensional rate matrices. Essen tially , a sequence of nested regions is defined, ∅ = R 0 ⊂ R 1 ⊆ R 2 ⊆ . . . , with lim r →∞ R r = X , the statespace of the MJP . The statespace of the MCMC Marko v c hain is then extended to include ˜ r , the index of the smallest region that con tains the MJP , and the corresp onding extended p osterior can b e calculated using only finite rate matrices. In the examples w e inv estigate w e find that MESA and nMESA are anything from a factor of three to several orders of magnitude more efficien t than the most efficien t particle MCMC algorithm. W e conclude this section with three motiv ating examples of reaction net works; these three examples will b e used to b enchmark our algorithms, the algorithm of Georgoulas et al. (2017) and particle MCMC in Section 4. In Section 2 we describ e the algorithm of Georgoulas et al. (2017), separating out the k ey idea of nested regions whic h is shared by our algorithms. Section 3 describ es MESA and nMESA themselv es, and the article concludes in Section 5 with a discussion. 1.1 Reaction net w ork examples In a reaction netw ork, the state vector, X , consists of the (non-negativ e) counts of one or more physical, c hemical or biological sp ecies. The state vector is piecewise constan t o ver time, and up dates only when a reaction o ccurs. F or a giv en system, let there b e R p ossible reactions. The state up date is deterministic given the curren t state and the particular reaction. Reaction r ( r = 1 , . . . , R ) o ccurs according to a P oisson pro cess with a rate of h r ( X ; θ r ) for some function h r and unkno wn parameter θ r ; i.e. , for some small time in terv al δ t , P (reaction r o ccurs in ( t, t + δ t ] | X = x ) = h r ( x ; θ r ) δ t + o ( δ t ). F or example, the Lotk a- V olterra mo del, b elow, has R = 3 reactions, the first of whic h changes the state vector x = ( x 1 , x 2 ) to ( x 1 − 1 , x 2 ) and o ccurs with a rate of h 1 ( x ; θ 1 ) = θ 1 x 1 . T o motiv ate the imp ortance of inference on reaction netw orks w e no w present: the Lotk a- V olterra predator-prey mo del, the Schl¨ ogel mo del, whic h is one of the simplest bistable net- w orks, and a simple mo del of gene auto-regulation in prok ary otes. W e will p erform inference on these reaction netw orks in our sim ulation study in Section 4. Examples 1. The Lotk a-V olterra predator-prey mo del (e.g. Boys et al., 2008). Two sp ecies, predators, p red , and prey , p rey , with coun ts of X 1 and X 2 resp ectiv ely ev olve and 3 in teract through the follo wing three reactions (with asso ciated rates): Pred θ 1 X 1 − → ∅ Prey θ 2 X 2 − → 2 Prey Pred + Prey θ 3 X 1 X 2 − → 2 Pred . Examples 2. The Sc hl¨ ogel mo del (e.g. V ellela and Qian, 2009) has t w o stable meta states, and the frequency of transitions b etw een the meta states is muc h low er than the frequency of transitions b etw een states within a single meta state. The in teractions betw een the single sp ecies, whose frequency is X , and t wo c hemical ‘p o ols’, A and B are: A + 2X θ 1 X ( X − 1) / 2 − → 3 X , B θ 3 − → X 3X θ 2 X ( X − 1)( X − 2) / 6 − → 2 X + A , X θ 4 X − → B . Examples 3. The autoregulatory gene net w ork of Goligh tly and Wilkinson (2005) mo d- els the production of RNA from DNA and of a protein P from RNA , as well as the extinction of both RNA and P , the rev ersible dimerisation of P and the rev ersible binding of the dimer, P 2 to DNA , where the binding inhibits pro duction of RNA . The total num b er of copies of DNA , G , is fixed, and the reactions are: DNA + P 2 θ 1 ( G − X 4 ) X 3 − → DNA · P 2 , DNA θ 3 ( G − X 3 ) − → DNA + RNA , DNA · P 2 θ 2 X 4 − → DNA + P 2 , RNA θ 4 X 1 − → RNA + P , 2 P θ 5 X 2 ( X 2 − 1) / 2 − → P 2 , RNA θ 7 X 1 − → ∅ , P 2 θ 6 X 3 − → 2 P , P θ 8 X 2 − → ∅ , where X 1 , . . . , X 4 denote the coun ts of RNA , P , P 2 , and DNA · P 2 resp ectiv ely . The p oten tially coun tably infinite set of p ossible states of a reaction net work can be placed in one-to-one correspondence with the non-negativ e integers. The i, j th en try of the corre- sp onding rate ‘matrix’ Q ( i 6 = j ) is the rate for moving from state i to state j . 1.2 Notation Throughout this article, a scalar operation applied to a vector means that the operation is applied to eac h elemen t of the vector in turn, leading to a new vector, e.g. , for the d -vector θ , log θ ≡ (log θ 1 , . . . , log θ d ) > . W e denote the v ector of 1s b y 1. The symbol 0 denotes the scalar 0 or the vector or matrix of 0s as dictated b y the context. There is a one-to-one correspondence b et ween any vector state x , suc h as num b ers of preda- tors and prey in Example 1, and the asso ciated non-negative integer state, whic h we denote b y k ( x ). Throughout this article, for simplicity of presen tation, w e abuse notation b y abbre- viating the ( k ( x ) , k ( x 0 )) elemen t of a matrix M , strictly [ M ] k ( x ) ,k ( x 0 ) , to [ M ] x,x 0 . 4 2 Inference for MJPs with infinite statespaces using the rate matrix F or simplicity of presen tation we assume a known initial condition, x 0 , though the metho dol- ogy is trivially generalisable to an initial distribution. As in Georgoulas et al. (2017), we then consider the observ ation regime where particle filters t ypically struggle most: exact coun ts of all sp ecies are observed at discrete points in time, t 1 , t 2 , . . . , t n ; for simplicit y w e present the case where t i = it for some inter-observ ation interv al t . Throughout this article, θ denotes the vector of p ositiv e reaction-rate parameters, and Ba yesian inference is p erformed on ψ := log θ , to which a prior densit y π 0 ( ψ ) is assigned. F or a finite-statespace Marko v c hain, whilst the rate matrix, Q , is the natural descriptor of the pro cess, the lik eliho o d for t ypical observ ation regimes inv olv es the transition matrix, e Qt , the ( i, j )th element of whic h is exactly P ( X t = j | X 0 = i ). By the Marko v prop erty , the lik eliho o d for the exact observ ations x 1 , . . . , x n is then L ( ψ ; x 1: n ) = n Y i =1 [ e Q ( ψ ) t ] x i − 1 ,x i . (1) The ab ov e likelihoo d is used within the algorithm of Georgoulas et al. (2017), and within MESA and nMESA. All three algorithms share the same construction of ne sted regions which enables the use of (1) and which w e now describe. 2.1 Set up for coun tably infinite statespaces Let the MJP , { X s } s ≥ 0 , ha ve a statespace of X , start from X 0 = x and b e observed pre- cisely at time t : X t = x 0 . Consider an infinite sequence of regions, {R r } ∞ r =0 with R 0 = ∅ ⊂ R 1 ⊆ R 2 ⊆ R 3 . . . and lim r →∞ R r = X ; w e permit equalit y so that the description also applies to MJPs with finite statespaces. F urthermore, R 1 should be chosen such that P ( { X s } 0 ≤ s ≤ t ∈ R 1 | X 0 = x, X t = x 0 ) > 0. Let d r = |R r | b e the num b er of states in R r . Let Q ( ψ ) be the infinitesimal generator for the MJP on X . F or finite A, B ⊆ X , let Q ( A, B ) b e the submatrix of Q that in volv es transitions from A to B , and let Q r b e the b e the rate matrix for transitions inside R r except that it has an additional coffin state, C , whic h receiv es all transitions that, under Q , would exit R r . Sp ecifically Q r := Q ( R r , R r ) c 0 0 , where, here and henceforth, c denotes the scalar or column v ector (as appropriate) such that P d r +1 j =1 Q i,j = 0 for eac h i . W e will, in fact, hav e a differen t sequence of regions defined for eac h in ter-observ ation interv al. W e will denote the r th region for the i th in ter-observ ation in terv al b y R ( i ) r and the asso ciated transition matrix b y Q ( i ) r . F or clarit y of exposition, we will often suppress the sup erscript ( i ). 5 F or eac h region R r ⊆ X , there is a one-to-one map k r : R r → { 1 , . . . , d r } and we add that k r : C → d r + 1. Using the shorthand of X for { X s } t s =0 , for 0 ≤ t 1 < t 2 ≤ t w e define: B ( X ; t 1 , t 2 , R ) := 1 { X s ∈ R ∀ s ∈ [ t 1 , t 2 ] } ( r ≥ 0) . W e set ˜ r ( X ; t 1 , t 2 ) to b e the unique index where X s ∈ R ˜ r ( X ; t 1 ,t 2 ) ∀ s ∈ [ t 1 , t 2 ] and ∃ s ∈ [ t 1 , t 2 ] suc h that X s / ∈ R ˜ r ( X ; t 1 ,t 2 ) − 1 ; i.e. , the smallest index r such that B ( x ; t 1 , t 2 , R r ) = 1. 2.2 The metho d of Georgoulas et al. (2017) In Georgoulas et al. (2017), henceforth abbreviated to GHS17, the random-truncation metho d of McLeish (2011) and Glynn and Rhee (2014) leads to an un biased estimator of the likelihoo d of a set of observ ations, whic h feeds in to a pseudo-marginal MCMC algorithm (Andrieu and Rob erts, 2009) targetting the p osterior π ( ψ ) ∝ π 0 ( ψ ) L ( ψ ). Unlik e other uses of random truncation within MCMC (e.g. Lyne et al. (2015); see also Jacob and Thiery (2015)), ho w ever, the un biased estimator of GHS17 can never be negative. W e first briefly describ e the random truncation metho d, b efore detailing the algorithm of GHS17. Let z 0 , z 1 , . . . b e a sequence, with z := lim i →∞ z i < ∞ . Let R ∈ { 1 , 2 , . . . , } be sampled from some mass function. Then ˆ Z := z 0 + R X j =1 z j − z j − 1 P ( R ≥ j ) is an un biased estimator of z . This is b ecause, sub ject to the condition of F ubini’s Theorem, the order of sum and exp ectation can b e exc hanged, so E h ˆ Z i = z 0 + ∞ X j =1 ( z j − z j − 1 ) P ( R ≥ j ) E [ I ( R ≥ j )] , and the result follows from the telescoping sum. A t eac h iteration of the algorithm of GHS17, a v alue for r is sampled at random from some discrete probabilit y mass function { p ( r ) } ∞ r =1 . In GHS17, P ( R > r | R ≥ r ) := q r = aq r − 1 for some a < 1 and with q 0 = 1. Consequently , P ( R > r ) = a r ( r +1) / 2 . (2) Since [ e Q r ( ψ ) t ] x,x 0 = P ( X t = x 0 , B ( X ; 0 , t, R r ) = 1 | X 0 = x ), lim r →∞ [ e Q r ( ψ ) t ] x , x 0 = L ( ψ ; x, x 0 ). Th us, if X 0 = x and X t = x 0 are consecutiv e observ ations, ˆ L ( ψ ; x, x 0 , r ) = r X j =1 1 P ( R ≥ j ) [ e Q j ( ψ ) t ] x , x 0 − [ e Q j − 1 ( ψ ) t ] x,x 0 , (3) is a realisation from an unbiased estimator for the likelihoo d contribution L ( ψ ; x, x 0 ) = [ e Qt ] x , x 0 (here [ e Q 0 t ] x,x 0 = 0). 6 One estimator of the form (3), with its own indep endently sampled r , is created for eac h in ter-observ ation in terv al, and ˆ L ( ψ ; r 1: n ) := Q n i =1 ˆ L ( ψ ; x i − 1 , x i , r i ) then pro vides a realisation from an un biased estimator for the full likelihoo d. Giv en a current p osition ψ = log θ and a set of region indices r 1: n w e ha ve a realisation of an un biased lik eliho o d estimator, ˆ L ( ψ ; r 1: n ). One iteration of the pseudo-marginal algorithm of GHS17 pro ceeds as follo ws: first, prop ose a new position from some densit y q ( ψ 0 | ψ ) then sam- ple r 0 1 , . . . , r 0 n indep enden tly from the mass function p ( r ) to obtain a realisation, ˆ L ( ψ 0 ; r 0 1: n ) of an un biased estimator for L ( ψ 0 ). The pseudo-marginal Metropolis-Hastings acceptance prob- abilit y for the prop osal ( ψ 0 , r 0 1: n ) is then α ( ψ , ψ 0 ) = 1 ∧ [ π 0 ( ψ 0 ) ˆ L ( ψ 0 ; r 0 1: n ) q ( ψ | ψ 0 )] / [ π 0 ( ψ ) ˆ L ( ψ ; r 1: n ) q ( ψ 0 | ψ )]. The pseudo-marginal algorithm can b e viewed as a Metrop olis-Hastings Mark ov c hain on ψ and r 1: n with a target distribution prop ortional to π 0 ( ψ ) ˆ L ( ψ ; r 1: n ) n Y i =1 p ( r i ) , and a prop osal of q ( ψ 0 | ψ ) Q n i =1 p ( r 0 i ). Because ˆ L ( ψ ; R 1: n ) is unbiased, in tegrating out all of the auxiliary v ariables from the target lea ves π ( ψ ) ∝ π 0 ( ψ ) L ( ψ ), as desired. T ypically , b ecause they arise from a sequence of differences, lik eliho o d estimates obtained via random-truncation might b e negativ e and hence un usable ( e.g. Lyne et al., 2015; Jacob and Thiery, 2015). How ev er, for observ ations X 0 = x and X t = x 0 , [ e Q r ( ψ ) t ] x,x 0 − [ e Q r − 1 ( ψ ) t ] x,x 0 = P ( X 1 = x 0 , B ( X ; 0 , t, R r ) = 1 | X 0 = x, ψ ) − P ( X 1 = x 0 , B ( X ; 0 , t, R r − 1 ) = 1 | X 0 = x, ψ ) (4) = P ( X 1 = x 0 , ˜ r ( X ; 0 , t ) = r | X 0 = x, ψ ) , (5) whic h is non-negativ e; so, by construction, a negativ e likelihoo d estimate is imp ossible. Although it can nev er b e negative, the random truncation algorithm in (3) suffers from sev eral related problems. The prop osal p ( r i ) should reflect the patterns in the terms in the sum in (3): if [ e Q j ( ψ ) t ] x , x 0 − [ e Q j − 1 ( ψ ) t ] x,x 0 / P ( R ≥ j ) → ∞ as j → ∞ then the unbiased estimate will b e unstable and hav e a high, or even infinite, v ariance; if, on the other hand the ratio goes to zero then unnecessarily large regions will frequen tly b e used, resulting in the exp onen tiation of unnecessarily large rate matrices with unnecessarily high rates, increasing the computational exp ense. GHS17 states that the distributional form of p ( r ), was chosen partly since it describ es the steady state of man y simple queuing systems. Ho wev er, the M/M/1 queue, for example, has a geometric stationary distribution (e.g. Grimmett and Stirzak er, 2001, Ch.11). F rom (2) the tails of p ( r ) are v ery ligh t compared to geometric tails. Even if a hea vier-tailed prop osal w ere used, ho wev er, there is no obvious choice for its shap e, or reason to b eliev e the shape w ould b e consistent across in ter-observ ation in terv als. F urther, some sp ecies migh t hav e a differen t spread than others, requiring differently shap ed regions. Finally , this shap e would t ypically dep end on θ , as exemplified in the follo wing remark. Remark 1. Consider a Lotk a-V olterra system with mo derate v ariabilit y b et w een X 0 = x and X 1 = x 0 , but no w divide the true reaction rates b y 1000, which is equiv alen t to slowing 7 do wn time by a factor of 1000. The most likely paths would b e those with close to a minimal n umber of even ts to get from x to x 0 , so P ( ˜ r ( X ; 0 , 1) = 1) ≈ 1; on the other hand, a large increase in all of the rates would see an appro ximately quasi-stationary distribution for most of the time interv al so larger regions would be more lik ely . W e will reform ulate the lik eliho o d, creating an explicit extended statespace, and giving a differen t distribution for r and r 0 ; as a result, there is no division by P ( R ≥ r ) and, indeed, no requirement for a generic prop osal p ( r ). The p oten tial for different amoun ts of v ariation b et w een sp ecies and across interv als is allo wed for b y letting the shap e of the cub oidal regions v ary and for the nature of the cub oids themselv es to v ary b et ween observ ations, all go verned b y tw o tuning parameters. F urther, instead of requiring a random num b er, R of matrix exp onen tials for eac h in ter-observ ation interv al, our algorithm requires just one. 3 New Algorithms W e employ the same idea of a sequence of nested regions as in GHS17, with one sequence for eac h in ter-observ ation interv al. The ne arly minimal extende d statesp ac e algorithm (nMESA) has one auxiliary v ariable p er in terv al as in GHS17, whereas the minimal extende d states- p ac e algorithm (MESA) has a single auxiliary v ariable. In eac h case w e explicitly extend the statespace from Ψ to include the index of the outermost region visited by X ov er the observ ation windo w (MESA) or b etw een each pair of observ ations (nMESA), and p erform MCMC directly on this extended statespace. This partitioning of the space was defined in GHS17 but then unbiasedly in tegrated out via random truncation using auxiliary v ariables to set the truncation levels. Figure 1 depicts a realisation of an MJP together with the rele- v an t regions for MESA and nMESA. It pro vides a graphical representation of three regions (MESA, see Section 3.2) or three regions p er in ter-observ ation in terv al (nMESA, see Section 3.3) together with a realisation of the MJP . 3.1 New regions F or simplicity , eac h region, {R ( i ) r : i = 1 , . . . , n, r = 1 , . . . } , is cub oid. Let the upp er and lo wer b ounds for sp ecies s in region r for in ter-observ ation interv al i b e u ( i ) r,s and l ( i ) r,s ; w e refer u ( i ) r,s − l ( i ) r,s + 1 as the width of region R ( i ) r for sp ecies s . In GHS17, for an in terv al b et ween observ ations of x and x 0 , R 1 is the smallest cuboid that con tains both x and x 0 . How ever this cub oid do es not necessarily allow a path b etw een x and x 0 . F or example, since no reaction increases predator num b ers by 1, a Lotk a-V olterra system with x = ( x 1 , x 2 ) and x 0 = ( x 1 + 1 , x 2 ) m ust ha v e left the rectangle with corners at x and x 0 . W e therefore define a scalar parameter w min , which sp ecifies, for Region R 1 , the smallest width for every sp ecies; if, for an y species, the smallest cuboid that con tains the t wo observ ations is narrow er than w min then the recursions b elow are p erformed for that species until this is no longer the case, leading to region R 1 . Subsequent regions are obtained recursiv ely from the previous region 8 ● ● ● ● 0.0 0.5 1.0 1.5 2.0 2.5 3.0 0 5 10 15 20 time state ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● Figure 1: Realisation (thic k line) of X , a one-dimensional MJP , o ver the time in terv al [0 , 3], with x 0 = 8, x 1 = 15, x 2 = 12 and x 3 = 16 (solid circles). F or MESA, region 1 is the union of the three rectangles filled with diagonal increasing lines, and is contained within region 2 whic h is the union of the three rectangles filled with diagonal decreasing lines, and is contained within and region three which is the union of the three rectangles filled with v ertical lines. F or nMESA, the same line types apply to the regions for eac h in ter-observ ation in terv al. 9 with the increase in region width for a given sp ecies prop ortional to the curren t width. F or region k + 1, u ( i ) r +1 ,s = u s ∧ u ( i ) r,s + [1 ∨ γ ( u ( i ) r,s − l ( i ) r,s + 1)] , l ( i ) r +1 ,s = l s ∨ l ( i ) r,s − [1 ∨ γ ( u ( i ) r,s − l ( i ) r,s + 1)] where, γ > 0 is a tuning parameter and l s and u s are hard lo w er and upp er b ounds. T ypically l s = 0, whilst u s is only required when there is an upp er b ound on the num b ers for sp ecies s . The regions in GHS17 use the ab ov e formulation, with γ = w min = 0. 3.2 The minimal extended statespace and target F or the observ ation regime giv en at the start of Section 2, define ˜ B r ( X ) := n Y i =1 B ( X ; t i − 1 , t i , R ( i ) r ) ( r ≥ 0) . Th us ˜ B r ( X ) = 1 if for ev ery in ter-observ ation interv al, X is entirely con tained within that in terv al’s region r . W e denote the smallest index r for whic h ˜ B r ( X ) = 1 b y ˜ r ( X ). In every in ter-observ ation interv al the pro cess is confined to that interv al’s region ˜ r ( X ) but in at least one in ter-observ ation interv al it is not confined to that interv al’s region ˜ r ( X ) − 1 ( ˜ r = 3 in Figure 1). W e target the extended posterior ˜ π ( ψ , r ) ∝ π 0 ( ψ ) P ( x 1: n , ˜ r ( X ) = r | ψ , x 0 ) , where P ( x 1: n , ˜ r ( X ) = r | ψ , x 0 ) = n Y i =1 e Q r ( ψ ) t x i − 1 ,x i − n Y i =1 e Q r − 1 ( ψ ) t x i − 1 ,x i . The marginal for ψ is π 0 ( ψ ) P ∞ r =1 P ( x 1: n , ˜ r ( X ) = r | ψ , x 0 ) = π 0 ( ψ ) P ( x 1: n | ψ , x 0 ), as re- quired. 3.3 Nearly minimal extended statespace and target Let ˜ r i ( X ) ≡ ˜ r ( X ; t i − 1 , t i ) b e, for the i th in ter-observ ation interv al, the index of the smallest region to completely con tain the MJP o ver that in terv al (in Figure 1, ( ˜ r 1 ( X ) , ˜ r 2 ( X ) , ˜ r 3 ( X )) = (2 , 1 , 3)). W e target the extended p osterior ˜ π ( ψ , r 1: n ) ∝ π 0 ( ψ ) n Y i =1 P ( x i , ˜ r i ( X ) = r i | ψ , x i − 1 ) where P ( x i , ˜ r i ( X ) = r i | ψ , x i − 1 ) = [ e Q r i ( ψ ) t ] x i − 1 ,x i − [ e Q r i − 1 ( ψ ) t ] x i − 1 ,x i . As for the MESA, the marginal for ψ is the desired posterior, in this case since X r 1: n n Y i =1 P ( x i , ˜ r i ( X ) = r i | ψ , x i − 1 ) = n Y i =1 ∞ X r i =1 P ( x i , ˜ r i ( X ) = r i | ψ , x i − 1 ) = n Y i =1 P ( x i | ψ , x i − 1 ) . 10 3.4 The MCMC algorithms Both MESA and nMESA use Metrop olis-within-Gibbs algorithms. First, either r | ψ , x 0: n (MESA) or r 1: n | ψ , x 0: n (nMESA) is up dated, then, respectively , ψ | r , x 0: n or ψ | r 1: n , x 0: n . F or eac h algorithm, the ψ up date can, in principle, use any MCMC mo ve that w orks on a con tinuous statespace; for simplicit y and robustness w e employ the random w alk Metrop olis. W e propose ψ 0 from a multiv ariate normal distribution centred on ψ and with a v ariance matrix prop ortional to the v ariance of ψ obtained from an initial tuning run. F or MESA w e up date r using a discrete random w alk, proposing r 0 = r − 1 or r 0 = r + 1 eac h with a probabilit y of 0 . 5 (when r = 1, the down ward prop osal is immediately rejected). F or nMESA, conditional on ψ , eac h comp onen t, r i , is up dated indep enden tly via this symmetric discrete random-w alk mo v e. The random walk mo ves b y a single region so as to sav e on computational cost. The new likelihoo d for a region r 0 in volv es quantities of the form v > [ e Q r 0 ] x s ,x e and v > [ e Q r 0 − 1 ] x s ,x e ; when r 0 is either r + 1 or r − 1, one of these quan tities is already a v ailable from the lik eliho o d calculations for the current region. Both stages of b oth algorithms require the computation of the exp onen tial of at least n rate matrices. As with the algorithm of GHS17, therefore, our algorithm is, w ell suited to parallelisation if the rate matrices and/or the largest required matrix p o wer are large; w e do not pursue this here. 3.5 Algorithm tuning and relativ e efficiency W e now consider the behavioural differences b et ween MESA and nMESA and the effects of the tuning parameters. MESA uses a single additional auxiliary v ariable, which sp ecifies the region n um b er for all in ter-observ ation interv als, whereas nMESA has one v ariable p er inter-observ ation interv al. It is kno wn (e.g. Rob erts et al., 1997) that man y MCMC algorithms, including the random walk Metrop olis mix more slo wly on larger statespaces. This suggests that the MESA algorithm should b e more efficient in terms of effectiv e samples p er iteration. On the other hand, conditional on any given parameter v ector ψ , consider the p osterior distribution of the MJP X o v er the set of of inter-observ ation in terv als, and the v aribilit y in the definition of the regions from one interv al to the next. Setting w min to the minimum allow able for the system (e.g. 1 for the Lotk a-V olterra mo del and 0 for the Schl¨ ogel mo del) leads to the smallest R 1 sizes and the resulting matrix exp onen tials are v ery c heap to calculate. Ho w ever, the larger the b et ween-observ ation sto chasticit y the larger the region that is likely to b e needed to contain the pro cess b etw een the observ ation times, y et some consecutiv e observ ations in the sequence will just happ en to be similar, so the minimal R 1 s will b e to o small. This is not an issue for nMESA whic h allows eac h in ter- observ ation interv al to find its own region level ˜ r i ; but MESA forces all in ter-observ ation in terv als to use the same region n um b er, ˜ r . The disparity b et w een some of these R 1 s very probably con taining the pro cess, and others v ery probably not containing it leads to p o or 11 mixing for MESA when w min is at its minim um, and suggests increasing w min to a sensible minim um v alue for an y in terobserv ation in terv al. Increasing w min to o far, for example beyond the range where the sto chastic pro cess is likely to lie, w ould lead to unnecessary computational exp ense, suggesting there may be an optimal w min v alue. Ev en with this larger w min , for many in ter-observ ation in terv als, the less flexible MESA will t ypically exponentiate larger matrices than nMESA. The numerical exp erimen ts in Section 4 sho w that in the scenarios considered, nMESA’s flexibility is more adv an tageous than MESA’s smaller statespace, but typically b y a factor of less than 2. When the tuning parameter γ = 0, for eac h sp ecies R r +1 is 2 units wider than the R r . Ho wev er, for regions of size >> 10, sa y , this is a v ery small relativ e increase in width, and as such, we might exp ect an asso ciated very small probabilit y of the pro cess sta ying within R r +1 \R r . In other w ords, the range of region indices ov er which the process is likely to need to mo v e is large. Since in MESA and nMESA the MCMC mov e to change region prop oses either an increase or decrease of the region index by 1 (see Section 3.4), region n umber mixes slo wly . Since larger region indices are asso ciated with larger reaction rates, for example, this also affects the mixing of ψ , all of which suggests c ho osing γ > 0. On the other hand, consider a γ so large that the pro cess is almost certain to be in R 1 ; the dimension of R 2 will b e appro ximately (1 + 2 γ ) n s times the size of that of R 1 , and may contain unnecessarily large rates, making matrix exp onen tials exp ensiv e to calculate, yet a mov e to R 2 is prop osed ev ery other iteration. These heuristics suggest that there should b e an optimal γ ∈ (0 , ∞ ), 4 Numerical comparisons F or eac h of the reaction netw orks given in Section 1.1 and a known initial condition, x 0 , w e simulated a realisation from the sto chastic pro cess from time 0 to an appropriate t end and then recorded the states at regular interv als so that there were 50 observ ations eac h from a realisation of the Sc hl¨ ogel pro cess and a realisation of the autoregulatory pro cess, and 20 observ ations of a Lotk a-V olterra pro cess; we name these data sets Sch50, AR50 and L V20, resp ectively . T o inv estigate the effect of altering the in ter-observ ation interv al, for the Lotk a-V olterra pro cess, t wo further data sets, L V40 and L V10, w ere generated with 40 and 10 observ ations, resp ectively . T o suppress the effect of in ter-realisation v ariability , L V40 and L V10 w ere generated from the same realisation as L V20 by , resp ectiv ely , halving and doubling the in ter-observ ation time interv al; thus LV 10 ⊂ LV 20 ⊂ LV 40. Figure 2 sho ws the realisations of the sto chastic pro cesses from whic h L V20, L V40, L V10 and Sch50 arose, together with the observ ations in L V20 and Sc h50. The realisation from the autoregulatory pro cess and the observ ations AR50 are pro vided in Figure 4 in App endix B.1, whic h also pro vides v alues for x 0 , t end and the true parameters for all three pro cesses (see T able 5) as w ell as the prior distributions assigned to the parameters. F or eac h data set the output from a tuning run of 10 4 iterations of nMESA w as used to create an estimate, b Σ, of the v ariance matrix of the parameter v ector, ψ . F or comparabilit y , for all algorithms, prop osals for the random w alk on ψ w ere of the form: ψ 0 = ψ + λ b Σ 1 / 2 z , where z is 12 0 5 10 15 20 20 30 40 50 60 70 80 90 Lotka−V olterra predators time species count 0 5 10 15 20 10 20 30 40 50 Lotka−V olterra prey time species count 0 50 100 150 200 0 5 10 15 20 25 30 35 Schlogel time species count Figure 2: Left (predators) and cen tral (prey) panels: the realisation of the sto c hastic pro cess from which datasets L V10, L V20 and L V40 arose, together with the L V20 data. Right panel: the Sc h50 data with the Sc hl¨ ogel pro cess from whic h it arose. a realisation of a v ector of standard Gaussians, b Σ 1 / 2 is defined so that b Σ 1 / 2 b Σ 1 / 2 > = I , and λ is a tuning parameter. The scaling, λ , of the random w alk proposal w as c hosen using standard acceptance-rate heuristics (e.g. Rob erts et al., 1997; Sherlock et al., 2010, 2015). The n um b er of particles w as chosen so that the v ariance of log ˆ π at p oin ts in the main p osterior mass was not m uch abov e 1 (Sherlo ck et al., 2015; Doucet et al., 2015). No tuning advice is given in GHS17 so w e pro ceeded by first tuning γ and a for a fixed, sensible ψ , and then tuning λ ; see App endix B.2 for further details. Unless otherwise stated, eac h algorithm w as run for 10 5 iterations. In all cases, the first 100 iterations w ere remo v ed as burn in, as trace plots sho wed that this w as all that was necessary . Results for MESA are presen ted in terms of the CPU time in seconds, T , the acceptance rate for the random w alk up date on the parameters, α ψ , the acceptance rate for the in teger random walk up date on the region, α r , and the num b er of effectiv e samples p er minute (rounded to the nearest in teger) for the parameters, the region index and log π . The n um b er of effective samples w as calculated using effectiveSize command in the coda pac k age in R (Plummer et al., 2006). Quantities are the same for nMESA except that α r is the mean of the acceptance rates for the random w alks on eac h of the region indices, and the effectiv e samples p er minute of the a verage region index is recorded. Neither particle MCMC nor GHS17 has a ‘region’ auxiliary v ariable, so the t wo fields for this are left blank. GHS17 strictly should ha ve γ = 0, but we also report the results for larger γ where this did not reduce the efficiency to o muc h. 4.1 Numerical and computational issues Calculations of the form v > e Qt w ere p erformed using the more efficien t of tw o p ossible algo- rithms, chosen automatically at runtime on a case-by-case basis. The uniformisation metho d (e.g. Sidje and Stew art, 1999) and a v ariation on the scaling and squaring method (e.g. Moler 13 and V an Loan, 2003). In our examples, Q is a sparse d × d matrix with O ( d ) non-zero en tries; define ρ := max i =1 ,...d | Q ii | . With this set up, the uniformisation method takes O ( ρtd ) op- erations and has a memory fo otprin t of O ( d ), whereas the scaling and squaring metho d has tak es O ( d 3 log ρ ) op erations with a footprint of O ( d 2 ). The latter was typically only used for some of the calculations for the Sc hlo¨ gel mo del where for some of the observ ation interv als, with the MESA and GHS17 algorithms, t ypically , ρ ' 10 8 but d / 10 3 . See App endix A for more details on the metho ds. The maxim um size of an unsigned integer in C++ is ≈ 4 × 10 9 . Both metho ds of exp o- nen tiation require the ev aluation of v > M j for some matrix, M with no negativ e entries, for some in teger p o w er j ∼ ρ + O ( √ ρ ). Straigh tforw ard, exact (to a prescribed small tolerance) ev aluation requires storing j as an in teger. F or the Schl¨ ogel system using GHS17 or using MESA with small w min , for some in ter-observ ation interv als on some iterations ρ > 4 × 10 9 , sometimes considerably so. In such cases ρ w as truncated to 4 × 10 9 so that inference w as no longer exact, but could at least con tin ue; in the remainder of this section w e refer to this as the inte ger overflow problem. With an increase in co de and algorithm complexit y this issue could be ov ercome, but on the o ccasions when it o ccurred the algorithms for which it o ccurred, ev en with the inexact inference, w ere muc h less efficien t than MESA with a larger w min or nMESA, so we did not pursue this further. Unless stated otherwise runs were performed on a desktop mac hine using a single thread of a single i7-3770 core. Code for MESA, nMESA and GHS17 is a v ailable from https: //chrisgsherlock.github.io/Research/publications.html . 4.2 Lotk a-V olterra mo del T ables 1, 2 and 3 show the simulation results for a selection of the runs p erformed, resp ec- tiv ely , for the L V20, L V40 and L V10 data sets. W e fo cus on the L V20 data set, and p oin t out an y differences eviden t in the other t wo data sets. Firstly , particle MCMC using the bridge of Goligh tly and Wilkinson (2015), henceforth re- ferred to as GW15, w as appro ximately a factor of 4 times as efficien t as the algorithm of GHS17 (factors of approximately 3 and 7 for L V40 and L V10 respectively). F urther, runs of GHS17 with γ ≈ 0 . 1 (b est performance w as for L V10, which is shown) w ere m uch less efficien t than with γ = 0 . 0. The most efficien t MESA tuning w as a factor of 3 more efficient than particle MCMC (factors of approximately 7 and 2 . 5 for L V40 and L V10), whilst the most efficient nMESA w as a factor of nearly 5 times more efficient than particle MCMC (factors of appro ximately 12 and 4 . 5 for L V40 and L V10). F or MESA and nMESA, the v ariations in efficiency with γ and many of the v ariations with γ visible in T ables 1, 2 and 3 are explained b y the heuristics in Section 3.5. F urther, an y trend (or drift) in the pro cess b et ween a pair of observ ations w ould be approximated b y the observ ations differences, so the choice of w min should b e driven b y the exp ected sto c hasticity , and so should increase as the inter-observ ation time interv al increases, as also observ ed in the three tables. 14 T able 1: T uning parameter settings and results for GHS17, MESA and nMESA for the L V20 dataset. 1 GW15 used 100 particles. 2 GHS17 used a = 0 . 98. ESS/min ute Algorithm λ γ w min T α ψ α r ψ 1 ψ 2 ψ 3 r or ¯ r log π GW15 1 1 . 6 8157 15 . 5 24 25 26 10 GHS17 2 1 . 6 0 . 0 1 17874 6 . 3 6 6 6 4 MESA 1 . 5 0 . 0 1 8270 28 . 2 73 . 5 64 65 62 42 58 MESA 1 . 5 0 . 1 1 12634 28 . 4 72 . 2 42 42 42 36 42 MESA 1 . 5 0 . 1 7 8023 28 . 2 68 . 9 65 65 64 85 66 MESA 1 . 5 0 . 0 10 7579 28 . 2 70 . 8 68 69 68 54 66 MESA 1 . 5 0 . 1 10 8035 28 . 2 67 . 1 67 68 69 95 72 MESA 1 . 5 0 . 1 20 7557 28 . 2 51 . 4 72 73 71 147 74 MESA 1 . 5 0 . 1 30 9396 28 . 9 1 . 3 62 64 62 194 61 nMESA 1 . 4 0 . 0 1 2887 27 . 8 69 . 4 106 111 105 78 140 nMESA 1 . 4 0 . 1 1 2991 27 . 7 69 . 2 106 105 104 78 157 nMESA 1 . 4 0 . 2 1 4164 27 . 6 59 . 5 81 85 84 90 264 nMESA 1 . 4 0 . 3 1 5553 27 . 9 50 . 2 69 70 71 91 252 nMESA 1 . 4 0 . 0 10 3054 28 . 8 53 . 6 115 117 115 75 93 nMESA 1 . 4 0 . 1 10 3126 28 . 7 53 . 6 116 117 115 89 115 nMESA 1 . 4 0 . 2 10 3745 28 . 9 40 . 9 114 114 108 276 180 nMESA 1 . 4 0 . 1 20 4458 30 . 6 4 . 2 120 119 118 192 171 nMESA 1 . 4 0 . 1 30 9480 31 . 6 0 . 07 63 62 60 177 65 T able 2: T uning parameter settings and results for GHS17, MESA and nMESA for the L V40 dataset. 1 GW15 used 170 particles. 2 GHS17 used a = 0 . 98. ESS/min Algorithm λ γ w min T α ψ α r ψ 1 ψ 2 ψ 3 r or ¯ r log π GW15 1 1 . 5 14039 16 . 9 15 14 15 6 GHS17 2 1 . 4 0 . 0 19320 6 . 7 5 6 5 5 MESA 1 . 5 0 . 0 1 5252 27 . 5 59 . 2 97 101 101 145 105 MESA 1 . 5 0 . 1 1 5372 27 . 7 59 . 3 103 102 99 131 103 MESA 1 . 5 0 . 0 7 4964 27 . 5 57 . 3 105 106 106 151 111 MESA 1 . 5 0 . 1 7 5073 27 . 6 57 . 3 98 108 112 148 112 nMESA 1 . 4 0 . 0 1 1672 25 . 6 59 . 1 164 162 161 155 312 nMESA 1 . 4 0 . 1 1 1680 25 . 7 59 . 2 169 170 169 162 317 nMESA 1 . 4 0 . 0 7 1805 27 . 3 37 . 8 186 196 202 196 220 nMESA 1 . 4 0 . 1 7 1807 27 . 3 37 . 9 180 185 186 179 203 15 T able 3: T uning parameter settings and results for GW15, GHS17, MESA and nMESA for the L V10 data set. 1 GW15 used 140 particles. 2 GHS17 used a = 0 . 98; with γ = 0 . 1, GHS17 w as run for 10 4 iterations. ESS/min Algorithm λ γ w min T α ψ α r ψ 1 ψ 2 ψ 3 r or ¯ r log π GW15 1 1 . 6 10324 14 . 6 14 16 17 6 GHS17 2 1 . 4 0 . 0 19800 3 . 6 2 3 2 2 GHS17 2 1 . 4 0 . 1 10773 4 . 1 0 . 6 0 . 5 0 . 7 0 . 5 MESA 1 . 5 0 . 0 1 18615 27 . 9 85 . 9 24 25 29 7 20 MESA 1 . 5 0 . 1 1 56925 27 . 9 72 . 4 9 9 9 7 9 MESA 1 . 5 0 . 1 7 33932 27 . 9 72 . 3 16 15 16 14 16 MESA 1 . 5 0 . 1 14 18172 27 . 7 68 . 1 27 28 27 34 31 MESA 1 . 5 0 . 0 20 14274 28 . 1 82 . 6 32 34 34 12 25 MESA 1 . 5 0 . 1 20 14911 27 . 9 65 . 2 36 36 35 53 37 MESA 1 . 5 0 . 1 30 13144 27 . 8 51 . 7 41 40 41 82 44 MESA 1 . 5 0 . 1 40 14659 28 . 1 6 . 4 36 37 37 90 38 nMESA 1 . 4 0 . 0 1 6355 28 . 7 79 . 7 49 53 58 19 37 nMESA 1 . 4 0 . 1 1 12244 28 . 7 75 . 1 29 30 31 15 45 nMESA 1 . 4 0 . 2 1 21082 28 . 9 62 . 1 20 20 21 24 44 nMESA 1 . 4 0 . 1 14 7570 29 . 3 60 . 0 48 52 57 37 78 nMESA 1 . 4 0 . 0 20 6132 29 . 4 43 . 2 54 56 58 23 36 nMESA 1 . 4 0 . 1 20 6469 29 . 4 37 . 6 65 71 67 81 94 nMESA 1 . 5 0 . 2 20 7857 29 . 7 29 . 4 62 63 66 148 126 nMESA 1 . 5 0 . 1 30 8177 27 . 6 7 . 1 60 63 63 96 86 nMESA 1 . 5 0 . 1 40 13652 28 . 1 0 . 7 40 40 40 83 40 16 4.3 Sc hl¨ ogel mo del F or the Schl¨ ogel mo del, nMESA is slightly more efficien t than MESA which is appro ximately fort y times as efficient as GHS17. The particle MCMC sc heme of Goligh tly and Wilkinson (2015) failed to conv erge, but a b o otstrap particle filter scheme with a large num b er of particles did con verge, although it was o v er t wo and a half orders of magnitude less efficien t than MESA and nMESA. There are several reasons for these striking results. Firstly , for typical rate parameters, θ , and large v alues of X the rates of the t w o reactions in volving the reserv oir A are extremely high. An y particle filter must sim ulate all the reactions that o ccur, a n umber of the same order as the sum of the four reaction rates. F or Sc h50, GHS17, MESA and nMESA all used the scaling and squaring algorithm (see Section 4.1), with a cost prop ortional to the logarithm of the of total rate; for reference, runs using used the uniformisation, whic h is linear in the rate, led to sp eed reductions by factors of 270, 105, and 54 for GHS17, MESA and nMESA resp ectively . The bridge of Goligh tly and Wilkinson (2015) tries to driv e the path for the sto c hastic process along an appro ximate straigh t line from the curren t p osition to the next observ ation. F or transitions b oth b et ween meta states and within the higher meta state this is an extremely p o or approximation to the b ehaviour (see Figure 2). Ho wev er, particle MCMC using a b o otstrap particle filter did mix. As well as needing to simulate every single one of the reactions, the large computational cost arose from the filter needing an order of magnitude more particles than was used on the Lotk a-V olterra examples. Figure 2 shows that the largest changes from one observ ation to the next o ccur during tran- sitions from one meta state to the other, but that it is in the higher meta state that the largest expansion in region co v erage from the smallest cuboid con taining adjacen t observ a- tions is required. T o fit the latter a relativ ely high region num b er (MESA with w min = 0) or a proposal distribution that leads to a relatively high region num b er (GHS17) is required, but this leads to a large statespace size as well as larger ρ arising from the upp er end of this large statespace. F or MESA, this problem is o vercome using a larger w min . The biggest problem with GHS17 was the requirement for the same prop osal distribution for the truncation index whatev er the meta-state of the pro cess, whereas in realit y the pro cess is likely to need a high index when in the high meta state and a low index when in the lo w meta state. F or lo wer a v alues, suc h as 0 . 95 (not shown), small region num b ers w ere t ypically prop osed, the calculations were relativ ely c heap but the c hain mixed exceedingly p o orly b ecause of the enormous (p ossibly infinite) v ariance of the logarithm of the un biased estimator of the lik eliho o d through the quotien t term in (3); the c hains failed to con verge. T o bring the v ariance under con trol, the probabilit y of proposing larger region num b ers had to b e increased substan tially , leading to exp ensive calculations b ecause, for each in ter-observ ation in terv al, the final region is larger, and b ecause there are typically more terms in the random truncation. Increasing γ similarly reduces the v ariance of the truncation estimator of the lik eliho o d and increases the computational effort. F urthermore, for γ ≥ 0 . 2 with a large enough for visible mixing, integer ov erflo w (see Section 4.1) b ecame more and more frequen t since the states in the larger prop osed regions led to larger rates. Sp ecifically , for the b est 17 T able 4: T unings and results for GHS17, MESA and nMESA for the Sch50 dataset. 1 The b o otstrap particle filter used 1400 particles. 2 F or GHS17, a was resp ectively 0 . 99, 0 . 998 (both with γ = 0) and 0 . 98; the run with a = 0 . 998 used only 2 × 10 4 iterations; the “ − ” indicates mixing so p o or that ESS could not b e estimated even approximately (estimated ESS < 20). 3 MESA with w min = 0 w as run for 10 4 iterations. 4 MESA with w min = 60 nev er exited region 1. ESS/min Algorithm λ γ w min T α ψ α r ψ 1 ψ 2 ψ 3 ψ 4 r or ¯ r log π BS 1 1 . 2 840782 18 . 0 0 . 30 0 . 30 0 . 25 0 . 27 0 . 21 GHS17 2 1 . 0 0 . 0 3271 0 . 019 - - - - - GHS17 2 1 . 0 0 . 0 4158 4 . 69 2 . 58 2 . 55 2 . 43 2 . 17 2 . 76 GHS17 2 1 . 0 0 . 2 110333 0 . 342 0 . 01 0 . 01 0 . 01 0 . 01 0 . 01 MESA 3 1 . 2 0 . 4 0 31182 27 . 7 28 . 9 1 . 21 1 . 28 1 . 07 1 . 04 0 . 27 1 . 01 MESA 1 . 2 0 . 4 10 4879 27 . 3 17 . 7 85 86 80 74 158 85 MESA 1 . 2 0 . 4 20 4596 27 . 4 8 . 8 91 91 81 83 158 78 MESA 1 . 2 0 . 4 40 3648 27 . 9 5 . 8 116 114 114 110 196 104 MESA 4 1 . 2 0 . 4 60 7912 29 . 1 12 . 4 58 58 53 53 ∗ 43 nMESA 0 . 8 0 . 8 0 1322 26 . 2 29 . 9 120 140 55 81 47 102 nMESA 0 . 8 0 . 2 20 1276 29 . 1 9 . 4 145 157 116 96 69 103 nMESA 0 . 9 0 . 4 20 1420 28 . 8 5 . 5 170 179 150 137 132 362 nMESA 0 . 9 0 . 4 40 3325 39 . 8 0 . 22 118 117 105 107 78 98 run, which used γ = 0 . 2, integer ov erflo w occurred on 120 of the iterations with a maxim um true ρ ≈ 1 . 2 × 10 11 and d v alues in excess of 3000. F or MESA with w min = 0, integer o verflo w o ccurred on 10 of the 10 4 iterations, with a maxim um true ρ ≈ 5 . 7 × 10 9 , and d v alues up to ≈ 1400; no o v erflo w occurred when w min ≥ 10. F or nMESA the maximum v alue of ρd w as < 1 . 1 × 10 8 . 4.4 Autoregulatory system Finally , we applied nMESA to the AR50 dataset. A run of 2 × 10 5 iterations to ok approx- imately 40 hours and ga ve a minim um (o v er all parameters) effective sample size of 1491. T uning runs for GW15 suggested that the same num b er of iterations would tak e around 48 da ys, so the algorithm w as not run. How ever, alternativ e, app oximate inference is a v ailable via particle MCMC using the c hemical Langevin equation (CLE), a sto chastic differential equation (SDE) appro ximation to the ev olution of the spatially discrete Mark ov jump pro- cess (e.g. Wilkinson, 2018). The mo dified diffusion bridge of Durham and Gallant (2002) w as used to propose paths b etw een the observ ation within a particle MCMC sc heme. When sim ulating from an approximation to the conditioned SDE using a bridge, a discretisation time step must be c hosen; the larger the time step, the smaller the computational cost of eac h iteration. In addition to the Mon te Carlo error inheren t in an y MCMC sc heme, the CLE 18 −4.0 −3.0 −2.0 −1.0 0.0 0.2 0.4 0.6 0.8 1.0 1.2 Comparison for θ 1 log θ 1 density −2 −1 0 1 0.0 0.5 1.0 1.5 Comparison for θ 2 log θ 2 density −1.5 −0.5 0.5 0.0 0.5 1.0 1.5 2.0 Comparison for θ 3 log θ 3 density −2.5 −1.5 −0.5 0.0 0.5 1.0 1.5 Comparison for θ 4 log θ 4 density −3.0 −2.0 −1.0 0.0 0.5 1.0 1.5 Comparison for θ 5 log θ 5 density −0.5 0.5 1.5 0.0 0.5 1.0 1.5 Comparison for θ 6 log θ 6 density −2.5 −1.5 −0.5 0.0 0.5 1.0 1.5 Comparison for θ 7 log θ 7 density −3.0 −2.0 0.0 0.5 1.0 1.5 Comparison for θ 8 log θ 8 density Figure 3: Kernel densit y estimates of the parameter p osteriors using nMESA (solid), and the CLE with ∆ t = 0 . 05 (dashed), with the true parameter v alue (dotted). approac h introduces error due to the approximation of the MJP b y a spatially con tinuous pro cess and then due to the approximation of the temp orally contin uous SDE by discretising time. F earnhead et al. (2014) observed that a coarser discretisation can lead to a premature decrease in the righ t tail of the p osterior for some parameters, essen tially b ecause doubling a rate parameter is equiv alen t to doubling the in ter-observ ation in terv al and k eeping the parameter the same, thus effectiv ely doubling the discretisation interv al. Using ∆ t = 0 . 2 w e observed a severe truncation in the righ t tails of the four parameters in volv ed in reversible reactions ( ψ 1 , ψ 2 , ψ 5 , ψ 6 ) so we decreased the time step to ∆ t = 0 . 05, a run whic h took appro ximately 90 hours for 2 × 10 4 iterations and ga v e a min ESS of 1404. P osteriors resulting from the final discretisation are compared with the true p osteriors in Fig- ure 3. Even with this discretisation a clear premature decay is visible in the four parameters in volv ed in reversible reactions. The issue is likely to b e comp ounded for ψ 1 , ψ 2 and ψ 3 b y the error in appro ximating an MJP with an SDE since the first three reaction rates dep end on the n umber of DNA molecules, which, with the set up detailed in App endix B.1, can only 19 tak e v alues of 0, 1 or 2. Decreasing the discretisation in terv al of the CLE still further w ould reduce (but not en tirely remo ve) the error resulting from the CLE appro ximation; how ever, this would reduce the computational efficiency still further, and particle MCMC using the current discretisation is already only half as efficient as nMESA. 5 Discussion W e ha v e described the minimal extended statespace algorithm (MESA) and the nearly min- imal extended statespace algorithm (nMESA) for inference on discretely and precisely ob- serv ed Mark ov jump processes, a setting in whic h standard inference b y particle MCMC is sev erely c hallenged. Our algorithms use the same key idea of nested regions that was used in the random-truncation algorithm of Georgoulas et al. (2017) but, in practice, are one or more orders of magnitude more efficient than that algorithm. On the three Lotk a-V olterra data sets MESA was 2 . 5 - 7 times as efficien t as the b est particle MCMC algorithm, and nMESA 4 . 5 - 12 times as efficien t as particle MCMC. On the Sc hl¨ ogel mo del, where the scaling and squaring matrix-exp onentiation algorithm w as used, the im- pro vemen t o v er the b est particle MCMC algorithm w as o v er t wo orders of magnitude. F or the autoregulatory gene mo del, nMESA w as able to p erform exact inference in a reasonable time, where no other metho d could. The sim ulations suggest that the greater flexibilit y of nMESA is more important for efficiency than the reduced statespace of MESA, though the adv an tage is only decisive for the autoregulatory mo del. Both MESA and nMESA can b e framed within the technique in tro duced in W alk er (2007) of rendering an infinite n umber of p ossibilities finite, and hence computable, by in tro ducing one or more auxiliary slic e v ariables. Lik e nMESA, W alker (2007) uses one auxiliary v ariable p er observ ation, but a single auxiliary is also possible. In our case, the region n umber b ounds the dimension of the statespace of the MJP . The EA3 of Beskos et al. (2008) uses a similar set of nested regions and the idea of a minimal region con taining a sto c hastic process, but the region is used to b ound a Radon-Nik o dym deriv ative and hence allo w the exact sim ulation of a skeleton of a diffusion with unit v olatilit y . In Rao and T eh (2012), which, lik e MESA and nMESA, applies to con tin uous-time Mark ov c hains, new states are sampled conditional on a finite set of possible even t times (which are resampled in another step); if the initial condition is kno wn and if the num b er of p ossible transitions from an y state is finite then the set of p ossible states after finitely many jumps is also finite. P article MCMC and the random truncation algorithm of Georgoulas et al. (2017) are ex- amples of pseudo-marginal MCMC (Andrieu and Rob erts, 2009). Such algorithms can b e view ed as introducing an extended statespace and targetting a p osterior on this extended space such that the marginal, in tegrating out the auxiliary v ariables, is the target of interest. New auxiliary v ariables are prop osed at eac h iteration: in the case of Georgoulas et al. (2017) these are a set of truncation v ariables, whereas in particle MCMC the auxiliary v ariables are all of the v ariables used b y the particle filter to estimate the lik eliho o d. Both MESA and 20 nMESA, ho w ever, are examples of correlated pseudo-marginal algorithms (Murra y and Gra- ham, 2016; Deligiannidis et al., 2018; Dahlin et al., 2015) since fresh auxiliary v ariables are not prop osed at eac h iteration, but instead a random w alk Metrop olis mov e from the existing v ariables is applied. Other matrix exp onentiation algorithms are a v ailable and, in particular, Krylo v subspace- based techniques (e.g. Sidje and Stewart, 1999) are often used for calculating e A b for a general sparse square matrix A and a v ector b . In b oth the Lotk a-V olterra example and the autoregulatory gene example w e found this tec hnique to b e betw een a factor of three and an order of magnitude slow er than the uniformisation tec hnique. Our algorithms are designed for the challenging exact-observ ation regime, but it would be straighforw ard to extend them to deal with noisy observ ations: nMESA via additional latent v ariables for the states at observ ation times, MESA b y considering all paths that sta y en tirely within R r but not R r − 1 , and including a lik eliho o d term at eac h observ ation time. How ever, as the observ ation noise increased, the efficiency of either algorithm w ould decrease gradually , whereas the efficiency of particle MCMC w ould increase, so that, for large-enough noise, PMCMC would b e more efficient. Of more interest, is the p otential for including the nested- region construction within particle MCMC, and this is the sub ject of current inv estigations. References Al-Moh y , A. H. and N. J. Higham (2011). Computing the action of a matrix exp onential with an application to exp onential in tegrators. SIAM J. Sci. Comput. 33 (2), 488–511. Andersson, H. and T. Britton (2000). Sto chastic epidemic mo dels and their statistic al anal- ysis , V olume 151 of L e ctur e Notes in Statistics . New Y ork: Springer-V erlag. Andrieu, C., A. Doucet, and R. Holenstein (2009). P article Mark ov c hain Mon te Carlo for efficien t n umerical sim ulation. In P . L’Ecuy er and A. B. Owen (Eds.), Monte Carlo and Quasi-Monte Carlo Metho ds 2008 , pp. 45–60. Spinger-V erlag Berlin Heidelb erg. Andrieu, C., A. Doucet, and R. Holenstein (2010). Particle Marko v chain Monte Carlo metho ds. J. R. Statist. So c. B 72 (3), 269–342. Andrieu, C. and G. O. Rob erts (2009). The pseudo-marginal approac h for efficien t Mon te Carlo computations. Ann. Statist. 37 (2), 697–725. Besk os, A., O. P apaspiliop oulos, and G. O. Roberts (2008, Mar). A factorisation of diffusion measure and finite sample path constructions. Metho dolo gy and Computing in Applie d Pr ob ability 10 (1), 85–104. Bo ys, R. J., D. J. Wilkinson, and T. B. L. Kirkwoo d (2008, Jun). Ba yesian inference for a discretely observ ed stochastic kinetic mo del. Statistics and Computing 18 (2), 125–135. Dahlin, J., F. Lindsten, J. Kronander, and T. B. Sc h¨ on (2015). Accelerating pseudo-marginal Metrop olis-Hastings by correlating auxiliary v ariables. 21 Deligiannidis, G., A. Doucet, and M. K. Pitt (2018). The correlated pseudomarginal metho d. Journal of the R oyal Statistic al So ciety: Series B (Statistic al Metho dolo gy) 80 (5), 839–870. Doucet, A., M. K. Pitt, G. Deligiannidis, and R. Kohn (2015, 03). Efficien t imple- men tation of Mark o v c hain Monte Carlo when using an unbiased lik eliho o d estimator. Biometrika 102 (2), 295–313. Dro v andi, C. C. and R. McCutc han (2016). Aliv e SMC 2 : Ba yesian mo del selction for lo w- coun t time series mo dels with in tractable lik eliho o ds. Biometrics 72 , 344–353. Durham, G. B. and R. A. Gallant (2002). Numerical tec hniques for maximum likelihoo d estimation of con tinuous time diffusion pro cesses. Journal of Business and Ec onomic Statistics 20 , 279–316. F earnhead, P ., V. Giagos, and C. Sherlo ck (2014). Inference for reaction net works using the linear noise appro ximation. Biometrics 70 (2), 457–466. Georgoulas, A., J. Hillston, and G. Sanguinetti (2017, Jul). Unbiased Bay esian inference for p opulation mark ov jump pro cesses via random truncations. Statistics and Comput- ing 27 (4), 991–1002. Glynn, P . W. and C.-H. Rhee (2014, 12). Exact estimation for Marko v c hain equilibrium exp ectations. J. Appl. Pr ob ab. 51A , 377–389. Goligh tly , A. and C. Sherlock (2019, Sep). Efficien t sampling of conditioned Marko v jump pro cesses. Statistics and Computing 29 (5), 1149–1163. Goligh tly , A. and D. J. Wilkinson (2005). Bay esian inference for sto chastic kinetic models using a diffusion approximati on. Biometrics 61 (3), 781–788. Goligh tly , A. and D. J. Wilkinson (2011). Ba yesian parameter inference for sto c hastic bio- c hemical netw ork mo dels using particle Mark ov c hain Mon te Carlo. Interfac e F o cus 1 (6), 807–820. Goligh tly , A. and D. J. Wilkinson (2015). Bay esian inference for Mark o v jump pro cesses with informativ e observ ations. SAGMB 14 (2), 169–188. Grimmett, G. and D. Stirzaker (2001). Pr ob ability and r andom pr o c esses , V olume 80. Oxford univ ersity press. Jacob, P . E. and A. H. Thiery (2015, 04). On nonnegative unbiased estimators. A nn. Statist. 43 (2), 769–784. Koblen ts, E. and J. Miguez (2015). A p opulation Monte Carlo scheme with transformed w eights and its application to sto c hastic kinetic mo dels. Statistics and Computing 25 (2), 407–425. Lyne, A.-M., M. Girolami, Y. Atc had ´ e, H. Strathmann, and D. Simpson (2015, 11). On rus- sian roulette estimates for Ba yesian inference with doubly-in tractable lik eliho o ds. Statist. Sci. 30 (4), 443–467. 22 McKinley , T. J., J. V. Ross, R. Deardon, and A. R. Co ok (2014). Sim ulation-based Bay esian inference for epidemic mo dels. Computational Statistics and Data Analysis 71 , 434–447. McLeish, D. (2011). A general method for debiasing a Mon te Carlo estimator. Monte Carlo Metho ds and Applic ations 17 , 301–315. Moler, C. and C. V an Loan (2003). Nineteen dubious wa ys to compute the exp onential of a matrix, t wen ty-fiv e y ears later. SIAM R eview 45 (1), 3–49. Moran, P . (1958). Random pro cesses in genetics. Mathematic al Pr o c e e dings of the Cambridge Philosophic al So ciety 54 (1), 60–71. cited By 468. Murra y , I. and M. Graham (2016, 09–11 May). Pseudo-marginal slice sampling. In A. Gretton and C. C. Rob ert (Eds.), Pr o c e e dings of the 19th International Confer enc e on A rtificial Intel ligenc e and Statistics , V olume 51 of JMLR: W&CP , Cadiz, Spain, pp. 911–919. Ow en, J., D. J. Wilkinson, and C. S. Gillespie (2015). Likelihoo d free inference for Marko v pro cesses: a comparison. Statistic al Applic ations in Genetics and Mole cular Biolo gy 14 (2), 189–209. Plummer, M., N. Best, K. Cowles, and K. Vines (2006). Co da: Conv ergence diagnosis and output analysis for MCMC. R News 6 (1), 7–11. Pulungan, R. and H. Hermanns (2018). T ransien t analysis of CTMCs: Uniformization or matrix exp onential? IAENG International Journal of Computer Scienc e 45 , 267–274. Rao, V. and Y. T eh (2012). MCMC for con tinuous-time discrete-state systems. In F. Pereira, C. J. C. Burges, L. Bottou, and K. Q. W einberger (Eds.), A dvanc es in Neur al Information Pr o c essing Systems , V olume 25. Curran Asso ciates, Inc. Reibman, A. and K. T riv edi (1988). Numerical transient analysis of marko v mo dels. Com- puters and Op er ations R ese ar ch 15 (1), 19 – 36. Rob erts, G. O., A. Gelman, and W. R. Gilks (1997, 02). W eak con v ergence and optimal scaling of random walk Metropolis algorithms. Ann. Appl. Pr ob ab. 7 (1), 110–120. Sherlo c k, C. (2021). Direct statistical inference for finite Marko v jump pro cesses via the matrix exp onential. Computational Statistics 36 , 2863–2887. Sherlo c k, C., P . F earnhead, and G. O. Rob erts (2010, 05). The random walk Metrop olis: Linking theory and practice through a case study . Statist. Sci. 25 (2), 172–190. Sherlo c k, C., A. H. Thiery , G. O. Rob erts, and J. S. Rosen thal (2015, 02). On the efficiency of pseudo-marginal random w alk Metrop olis algorithms. Ann. Statist. 43 (1), 238–275. Sidje, R. B. (1998). EXPOKIT. Softw are pack age for computing matrix exponentials. A CM- T r ans. Mathematic al Softwar e 24 (1), 130 – 156. Sidje, R. B. and W. J. Stewart (1999). A numerical study of large sparse matrix exp onen tials arising in Mark o v chains. Computational Statistics and Data Analysis 29 (3), 345 – 368. 23 V ellela, M. and H. Qian (2009). Sto chastic dynamics and non-equilibrium thermodynamics of a bistable c hemical system: the Schl¨ ogel mo del revisited. Journal of The R oyal So ciety Interfac e 6 (39), 925–940. W alk er, S. G. (2007). Sampling the diric hlet mixture mo del with slices. Communic ations in Statistics - Simulation and Computation 36 (1), 45–54. Wilkinson, D. J. (2018). Sto chastic Mo del ling for Systems Biolo gy (3rd ed.). Bo ca Raton, Florida: Chapman & Hall/CRC Press. A Matrix Exp onen tiation Consider ν > exp Q = ν > P ∞ i =0 Q i /i !, for some non-negative d -v ector ν and d × d rate matrix Q . In our case for region R r , from the j − 1 to the j th observ ation, ν r is the d r + 1 v ector whic h is 1 at k r ( x j − 1 ) and 0 ev erywhere else. F or a general matrix the calculation is especially difficult to ev aluate efficien tly yet to a pre-defined tolerance, , because of the p ossibility of very large negativ e and p ositiv e n um b ers cancelling during the ev aluation of the series; ho w ever, when Q is a rate matrix this issue can b e circum ven ted as follows. Let ρ = max i =1 ,...,d | Q i,i | then P := I d + Q/ρ has non-negativ e en tries and is, in fact, a Mark o v transition matrix. F urthermore: ν > e Q = ν > e ρ ( P − I d ) = e − ρ ν > e ρP . The uniformisation metho d (e.g. Sidje and Stewart, 1999) ev aluates ν > e P = P ∞ i =0 ν > P i /i ! ≈ P m i =0 ν > P i /i !, where, given ν > P i , and the fact that P is sparse, the calculation of ν > P i +1 = ( ν > P i ) P is an O ( d ) op eration. The truncation p oint m is c hosen so that ≤ 1 − F ( m + 1; ρ ), where F is the cumulativ e distribution function of a Poisson( ρ ) random v ariable, since then ν > e Q 1 − [ ν > e Q 1 = e − ρ ν > ∞ X i = m +1 ρ i i ! P i 1 = P (P oisson( ρ ) ≥ m + 1) ≤ ; see Sherlo ck (2021) for further details. The sc aling and squaring metho d (e.g. Moler and V an Loan, 2003) uses: e M ≡ e M / 2 s 2 s , for an y square matrix M . Th us, e M can be obtained can be obtained from e M / 2 s b y squaring s times. W e calculate e ρP / 2 s using the uniformisation metho d (but without the ν > term, as in Reibman and T rivedi, 1988; Pulungan and Hermanns, 2018), and revert to v ector-matrix m ultiplications b efore the final squaring; see Sherlo ck (2021) for further details. 24 T able 5: Observ ation and parameter information for our five data sets of exact observ ations, together with the parameter v alues used and the abbreviated name for the data set. Pro cess θ x 0 t end ∆ t n obs Name Lotk a-V olterra 0 . 3 , 0 . 4 , 0 . 01 (30 , 40) 20 . 0 1 . 0 20 L V20 0 . 5 40 L V40 2 . 0 10 L V10 Sc hl¨ ogel 3 . 0 , 0 . 5 , 0 . 5 , 3 . 0 (0) 200 . 0 4 . 0 50 Sc h50 Autoregulatory 0 . 1 , 0 . 7 , 0 . 7 , 0 . 2 (5 , 5 , 5 , 1 , 1) 25 . 0 0 . 5 50 AR50 0 . 1 , 0 . 9 , 0 . 3 , 0 . 1 B Details of n umerical exp erimen ts B.1 P arameter v alues, pro cess settings, and the AR50 data set T able 5 sho ws the observ ation and parameter information for the fiv e sim ulated data sets used in the simulation study in Section 4. The realisation of the autoregulatory system and the asso ciated AR50 dataset are provided in Figure 4. F or the Lotk a-V olterra mo del w e assigned indep endent a priori distributions of: ψ 1 ∼ N (log(0 . 2) , 1), ψ 2 ∼ N (log(0 . 2) , 1) and ψ 3 ∼ N (log(0 . 02) , 1). The Sc hl¨ ogel mo del parame- ters were a priori indep enden t with ψ i ∼ N (log (1) , 1), i = 1 , . . . , 4. F or the autoregularory mo del, parameters w ere a priori indep enden t with ψ i ∼ N (log 0 . 2 , 1), i = 1 . . . , 4 , 6 , . . . 8, and ψ 5 ∼ N (log 0 . 2 , 0 . 1). Both θ 5 and θ 6 describ e the rates for the rev ersible dimerisation of P and are very p o orly iden tified b y the data, although their quotien t is w ell identified (e.g. Goligh tly and Wilkinson, 2005); the tigh ter prior for ψ 5 ensures that the behaviour of the MCMC algorithm is not almost entirely dominated b y this one reaction. B.2 T uning GHS17 The algorithm of GHS17 w as tuned b y first fixing ψ at some sensible v alue (here the kno wn true v alue, but in practice it w ould be set to the p osterior mean from a training run) and recording the log-posterior, log π , at each iteration. F or an y giv en choice of γ , the parameter a was adjusted to achiev e the maxim um ESS/sec for log π . Still with ψ fixed, the ESS/sec of log π w as then in vestigated for differen t γ at the optimal a for eac h. In all cases it was found that γ = 0 gav e optimal p erformance. Then with the optimal γ and a parameters, λ w as adjusted to giv e an approximately optimal ESS. 25 0 10 20 30 40 50 0 1 2 3 4 5 6 7 Autoregulatory − RNA time species count 0 10 20 30 40 50 0 5 10 15 Autoregulatory − P time species count 0 10 20 30 40 50 0 2 4 6 8 Autoregulatory − P2 time species count 0 10 20 30 40 50 0.0 0.5 1.0 1.5 2.0 Autoregulatory − DNAP2 time species count Figure 4: The sto chastic pro cess that led to the AR50 dataset together with the AR50 data v alues: RNA (top left), P (top righ t), P 2 (b ottom left) and DNA · P 2 (b ottom right). Quan tities of DNA ma y b e obtained deterministically as D N A = 2 − D N A · P 2 26
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment