MIScnn: A Framework for Medical Image Segmentation with Convolutional Neural Networks and Deep Learning
The increased availability and usage of modern medical imaging induced a strong need for automatic medical image segmentation. Still, current image segmentation platforms do not provide the required functionalities for plain setup of medical image se…
Authors: Dominik M"uller, Frank Kramer
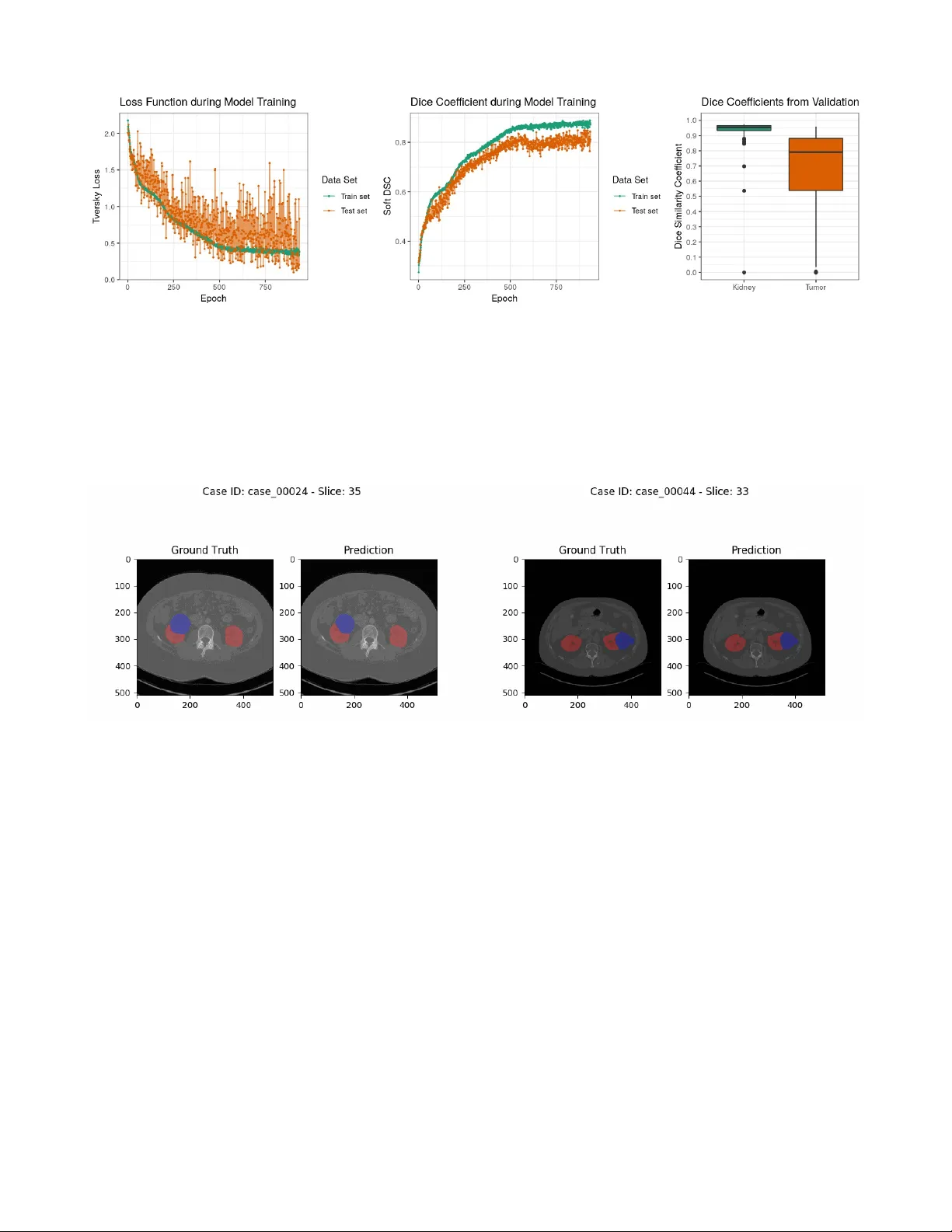
MIScnn: A Framework for Medical Image Segmentation with C onvolutional Neural Networks and Deep Learning Dominik Müller 1 and Frank Kramer 1 1 IT -Infrastructure for T ranslational Medical Research, Faculty of A pplied Computer Science, Faculty of Medicine, University of Augsburg, Germany Abstract. The increased availability and usage of modern medical imaging induced a strong need for automatic medical image segmentation. Still, current image segmentation platforms do not provide the required functionalities for plain setup of medical image segmentation pipelines. Already implemented pipelines are commonly standalone software, optimized on a specific public data set. Therefore, this paper introduces the open-source Python library MIScnn. The aim of MIScnn is to provide an intuitive API allowing fast building of medical image segmentation pipelines including data I/O, preprocessing, data augmentation, patch-wise analysis, metrics, a library with state-of-the-art deep learning models and model utilization like training, prediction, as well as fully automatic evaluation (e.g. cross-validation). Similarly , high configurability and multiple open interfaces allow full pipeline customization. Running a cross- validation with MIScnn on the Kidney Tumor Segmentation Challenge 2019 data set (multi-class semantic segmentation with 300 CT scans) resulted into a powerful predictor based on the standard 3D U-Net model. With this experiment, we could show that the MIScnn framework enables researchers to rapidly set up a complete medical image segmentation pipeline by using just a few lines of code. The source code for MIScnn is available in the Git repository: https://github.com/frankkramer-lab/MIScnn . Keywords: Medical image analysis; segmentation; computer aided diagnosis; biomedical image segmentation; u-net, deep learning; convolutional neural network; open-source; framework. 1 Intr oduction Medical imaging became a standard in diagnosis and medical intervention fo r the visual rep resentation of the functionalit y of or gans and tissues. Through the increased availability and usa ge of modern medical imaging like Magnetic Resonance Imaging (MRI), the need for automated processing of scanned imaging data is quite strong [1]. Currently , the evaluation of medical images is a manual process performed by physicians. Larger numbers of slices require the inspection of even more image material by doctors, especially regarding the i ncreased usage of high-resolution medical imaging. In order to shorten the time-consuming inspection and evaluation process, an autom atic pre-segmentation of abnormal features i n medical images would be required. Image segmentation i s a popular sub- field of image processing within computer science [2]. The aim of semantic segmentation is to identify common features in an input image by learning and then labeling each pi xel in an image with a class (e.g. background, kidney or tumor). Ther e is a wide range of algorithms to solve segmentation problems. However , state-of-the-art accuracy w as accomplished by convolutional neural networks and deep learning models [2–6], which are used extensively today . Furthermore, t he newest convolutional neural networks are able to exploit local and global features in im ages [7–9] and they can be trained to use 3D image information as well [10,1 1]. In recent yea rs, medical image segmentation models with a convolutional neural network architecture have become quite powerful and achieved similar results performance-wise as radiologists [5,12]. Nevertheless, t hese models have been standalone applications with optimized architectures, preprocessing procedures, data augm entations and metrics specific for their data set and corresponding segmentation problem [9]. Also, the pe rformance of such optimized pipelines varies drastically between different medical conditions. However , even for the same medical condition, evaluation and comparisons of these models are a persistent challenge due to the variety of t he size, shape, localization and distinctness of different data sets. In order to objectively compare two segmentation model architectures from the sea of one-use standalone pipelines, e ach specific for a single public data set, it would be required to implement a complete custom pipeline with preprocessing, data augment ation and batch creation. Frameworks for general image segmentation pipeline building can not be fully utilized. Th e reason for this are their missing medical image I/O interfaces, their preprocessing methods, as well as their lack of handling highly unbalanced class distributions, which is standard in medical imaging. Recently developed medical image segmentation platforms, like NiftyNet [13], are powerful t ools and an excellent first step for standardized medical image segmentation pipelines. However, they are designed more like configurable software instead of frameworks. They lack modular pipeline blocks to offer researchers the opportunity for easy customization and to help dev eloping their own software for their specific segmentation problems. In this work, we push towards constructing an intuitive and easy-to-use fra mework for fast setup of state-of-the-art convolutional neural network and deep learning models for medical image segmentation. The aim of our framework MIScnn ( M edical I mage S egmentation with C onvolutional N eural N etworks) is to provide a complete pipeline for preprocessing, data augmentation, patch slicing and batch creation steps in order to start straightforward with training and predicting on diverse medical imaging data. Instead of being fixated on one model architecture, MIScnn allows not only fast switching between multiple modern convolutional neural network models, but it also provides the possibility to easily add custom model architectures. A dditionally , it facilitates a simple deployment and fast usage of new deep learning models for medical image segmentation. S till, MIScnn i s highly con figurable to adjust hyperparamet ers, general training par ameters, preprocessing procedures, as well as include or exclude data augmentations and evaluation techniques. 2 Methods The open-source Python library MIScnn is a framework to setup medical image segmentation pipelines with convolutional neural networks and deep learning models. MIScnn is providing several core features: • 2D/3D medical image segmentation for binary and multi-class problems • Data I/O, preprocessing and data augmentation for biomedical images • Patch-wise and full image analysis • State-of-the-art deep learning model and metric library • Intuitive and fast model utilization (training, prediction) • Multiple automatic evaluation techniques (e.g. cross-validation) • Custom model, data I/O, pre-/postprocessing and metric support 2.1 Data Input NIfTI Data I/O Interface MIScnn provides a data I/O interface for the N euroimaging Informatics T echnology Initi ative (NifTI) [14] file format for loading Magnetic Resonance Imaging and Computed T omography data into the framework. This format was initially created to speed up the development and enhance the utility of informatics tools related to neuroimaging. Still, it is now commonly used for sharing public and anonymous MRI and CT data sets, not only for brain imaging, but also for all kinds of human 3D imaging. A NIfTI file contains the 3D image matrix and diverse metadata, like the thickness of the MRI slices. Custom Data I/O Interface Next to the i mplemented NIfTI I/O interface, MIScnn allows the usage of custom data I/O interfaces for other imaging data formats. This open interface enables MIScnn to handle specific biomedical imaging features (e.g. MRI slice thickness), and therefore it avoids losing these feature information by a fo rmat conversion requirement. A custom I/O i nterface must be committed t o the preprocessing function and i t has to return the medical image as a 2D or 3D matrix for i ntegration i n the workflow . It is advised to add format specific prep rocessing procedures (e.g. MRI slice thickness normalization) in the format specific I/O interface, before returning the image matrix into the pipeline. Figure 1: Flowchart diagram of the MIScnn pipeline starting with the data I/O and ending with a deep learning model. 2.2 Pre processing Pixel Intensity Normalization Inconsistent signal intensity ranges of images can drastically influence the performance of segmentation methods [15,16]. The signal ranges of biomedical imaging data are highly varying betw een data sets due to different image formats, diverse hardware/instruments (e.g. diff erent scanners), technical discrepancies, and simply biological variation [5]. Additionally , the machine learning algorithms behind image segmentation usually perform better on features which follow a normal distribution. In order to achieve dynamic signal intensity rang e consistency , it is advisable to scale and standardize imaging data. Th e signal intensity scaling projects the original value range to a predefined range usually between [0,1] or [-1,1], whereas standardization centers the values close to a normal distribution by computing a Z-Score normalization. MIScnn can be configured to include or exclude pixel intensity scaling or standardization on the medical imaging data in the pipeline. Clipping Similar to pixel intensity normalization, it i s also common to clip pixel intensities t o a certain range. Intensity values outside of this range will be cli pped to t he minimum or maximum range value. Especially in computer tomography images, pixel intensity values are expected to be i dentical for the same organs or tissue types even in di f ferent scanners [17]. This can be exploited through orga n-specific pixel intensity clipping. Resampling The resampling technique is used to modify t he width and/or height of images. This results into a new image with a modified number of pixels. Magnetic resonance or computer tomography sc ans can have diff erent slice thickness. However , training neural network models r equires the images to have the same slice thickness or voxel spacing. This can be accomplished through resampling. Additionally , downs ampling images reduces the required GPU memory for training and prediction. One Hot Encoding MIScnn is able to handle binary (background/cancer) as well multi-class (background/kidney/liver/lungs) segmentation problems. The representation of a binary segmentation is being m ade quite simple by using a variable with two states, zero and one. But for the processing of multiple categorical segmentation labels in machine learning algorithms, like deep learning models, it is required to convert the classes into a more mathematical representation. This can be achieved with the One Hot encoding method by cre ating a single binary variable for each segmentation class. MIScnn automatically One Hot encodes segmentation labels with more than two classes. 2.2 Patch-wise and Full Image Analysis Depending on the resolution of medical images, the available GPU hardware plays a large role in 3D segmentation analysis. Currently , it is not possible to fully fit high-resolution MRIs with an example size of 400x512x512 into state-of-the-art convolutional neural network models due to the enormous GPU memory requirements. Therefore, the 3D medical imaging data can be either sliced into sm aller cuboid patches or analyzed slice-by-slice, similar to a set of 2D images [5,6,18]. In order to fully use the information of all three axis, MIScnn slices 3D medical images into patches with a configurable size (e.g. 128x128x128) by default. Depending on the model architecture, these patches can fit into GPUs with RAM si zes of 4GB to 24GB, which are commonly used in research. Nevertheless, the slice-by-slice 2D analysis, as well as the 3D patch analysis is supported and can be used in MIScnn. It is also possible to configure the usage of full 3D images in case of analyzing uncommonly small medical images or having a large GPU cluster . B y default, 2D medical images are fitted completely into the convolutional neural netwo rk and deep learning models. Still, a 2D patch-wise app roach for large resolution images can be also applied. 2.3 Data Augmentation for T r aining In the machin e learning field, dat a augmentation covers t he artificially increase of training data. Especi ally in medical imaging, commonly only a small num ber of samples or images of a studied medical condition i s available for training [5,19–22]. Thus, an image c an be modified with multiple techniques, like shifting, to exp and the number of plausible examples for training. The aim is to create re asonable variations of the desired pattern i n order to avoid overfitting in small data sets [21]. For state-of-the-art data augmentation, MIScnn integrated the batchgenerators package from the Division of Medical Image Computing at the German Canc er Research Center (DKFZ) [23]. It offers various data augmentation techniques and was used by the winners of the l atest medical image processing challeng es [9,17,24]. It supports spatial translations, rotations, scaling, elastic deformations, brightness, contrast, gamma and noise augmentations like Gaussian noise. 2.4 Sampling and Batch generation Skipping Blank Patches The known problem i n medical images of the large unbalance between the relevant segments and the background results into an extensive amount of parts purely labeled as background and without any learning information [5,19]. Especially after data augmentation, there is no ben efit to multiply these blank parts or patches [25]. Therefore, in the patch-wise model training, all patches, which are completely labeled as background, can be excluded in order to avoid wasting time on unnecessary fitting. Batch Management After the data preprocessing and the optional data augmentation for training, sets of full i mages or patches are bundled into batches. One batch contains a number of prep ared images which are processed in a single step by the model and GPU. Sequential for each batch or proc essing step, the neural network updates i ts internal weights accordingly with the predefined learning rate. The possible number of images inside a single batch highly depends on the available GPU memory and has to be configured properly in MIScnn. Every batch is saved to disk in order to allow fast repeated access during the training process. This approach drastically reduces the computing time due to the avoidan ce of unnecessary repeated preprocessing of the batches. Nevertheless, this approach is not ideal for extr emely large data sets or for researchers without the required disk spa ce. In order to bypass this problem, MIScnn also supports “on-the-fly” generation of the next batch in memory during runtime. Batch Shuffling During model training, the order of batches, which are going to be fit ted and processed, is shuffled at the end of each epoch. This method r educes the vari ance of t he neural netwo rk during fitting over an epoch and lowers the risk of overfitting. Still, it must be noted, that only the processing sequence of the batches is shuffled and the data itself is not sorted into a new batch order . Multi-CPU and -GPU Support MIScnn also supports the usage of multiple GPUs and parallel CPU batch loading next to the GPU computi ng. Particularly , the storage of already prepared batches on disk enables a fast and parallelizable processing with CPU as well as GPU clusters by eliminating the risk of batch preprocessing bottlenecks. 2.5 Deep Learning Model Cr eation Model Ar chi tectur e The selection of a deep learning or convolutional neural network model is the most important step in a medical image segmentation pipeline. There is a variety of model architectures and each has different strengths and weaknesses [7,8,10,1 1,23–29]. MIScnn features an open model interfac e to l oad and switch between provided state-of- the-art convolutional neural network models like the popul ar U-Net model [7]. Models are represented with the open-source neural network library Keras [33] which provides an user-friendly API for commonly used neural-network building bl ocks on top of T ensorFlow [34] . The already implemented m odels are highly configurable by defin able number of neurons, custom input sizes, optional dropout and batch normalization l ayers or enhanced architecture versions like the Opti mized High Resolution Dens e-U-Net model [10] . Additionally , MIScnn of fers architectures for 3D, as well as 2D medic al ima ge segmentation. Besides the flexibility i n switching between already implemented models, the open m odel interface enables the ability for custom deep learning model implementations and simple integrating t hese custom models i nto the MIScnn pipeline. Metrics MIScnn offers a large quantity of various metrics which can be used as loss function for training or for evaluation in figures and manual performance analysis. The Dice coefficient, also known as the Dice similarity index, is one of the most popular metrics for medical image segmentation. It scores the similarity between the predicted segmentation and the ground truth. However , it also penaliz es false positives comparable to the precision metric. Depending on the segmentation classes (binary or multi-class), there is a simple and class-wise Dice coef ficient implementation. Whereas the simple implementation only accumulates the overall number of correct and false predictions, the class-wise implementation accounts the prediction performance for each segmentation class which is strongly recommended for commonly class- unbalanced medical images. Another popular supported metric is the Jaccard Index. Even though it is sim ilar t o the Dice coefficient, it does not only emphasize on precise segmentation. How ever , it also penalizes under- and over-segme ntation. Still, MIScnn uses the Tversky loss [35] for training. Comparable t o t he Dice co efficient, the Tversky loss function addresses data imbalance. Even so, it achieves a much better t rade-off between precision and r ecall. Thus, the Tversky loss function ensures good performance on binary , as well as multi-class segmentation. A dditionally , all standard metrics which are included in Keras, like accuracy or cross-entropy , can be used in MIScnn. Next to the already implemented metrics or loss fun ctions, MI Scnn of fe rs t he int egration of custom m etrics for training and evaluation. A custom metric can be implemented as defined in Keras, and simply be passed to the deep learning model. 2.6 Model Utilization W ith the initialized deep learning model and the fully preprocessed data, the model c an now be used for training on the data to fit model weights or for pr ediction by using an already fitted model. Alternatively , the model can perform an ev aluation, as well, by running a cross-validation for example, with multiple training and prediction calls. The model API allows saving and loading models in order to subsequently reuse already fitted models for prediction or for sharing pre-trained models. T raining In the pro cess of training a convoluti onal neural network or deep learning model, divers e sett ings have to be configured. At this point in the pipeline, the data augmentation options of the data set, which have a large influence on the training in medical image segmentation, must be already defined. Sequentially , the batch management configuration covered the settings for the batch size, and also the batch shuffling at the end of each epoch. Therefore, only the lea rning rate and the number of epochs are required to be adjusted before running the training process. The learning rate of a neural network model is defined as the extend in which the old weights of the neural network model are updated in each iteration or epoch. In contrast, the number of epo chs de fines how many times the complete data set will be fitted i nto the model. Sequentially , the resulting fitted model can be saved to disk. During t he training, the unde rlying Keras fr amework gives insights int o the current model performanc e with the predefined metrics, as well as the remaining fitting time. Additionally , MIScnn offers the usage of a fitting-evaluation callback functionality in which the fitting scores and metrics are stored into a tab separated file or directly plotted as a figure. Pr ediction For the segmentation prediction, an already fitted neural network model can be directly used after training or it can be loaded from file. The model predicts for ev ery pixel a Sigmoid value for each class. The Sigmoid value represents a probability estimation of t his pixel for the associated label. Sequentially , the a r gmax of the One Hot encoded class are identified for multi-class segmentati on problems and then converted back to a single result variable containing the class with the highest Sigmoid value. When using the overlapping patch-wise analysis approach during the training, MIScnn supports t wo methods for patches in the prediction. Either the prediction process plainly creates distinct patches and treats the overlapping patches during the training as purely data augmentation, or overlapping pat ches are created for prediction. Due to the lack of prediction power at pat ch edges, computing a second prediction for edg e p ixels in patches, by using an overl ap, is a commonly used approach. In the following merge of patches back to the original medical image shape, a merging strategy for the pixels is required, in the ove rlapping part of two patches and with multiple predictions. By default, MIScnn cal culates the mean between the predicted Sigmoid values for each class in every overlapping pixel. Figure 2: Flowchart visualization of the deep learning mod el creation and architecture selection process. The resulting image matrix with the segment ation prediction, which has the identical shape as the original medical image, is saved into a file structure according to the provided data I/O interface. By default, usi ng the NI fTI data I/O interface, the predicted segmentation matrix is saved in NIfTI format without any additional metadata. Evaluation MIScnn supports multiple automatic evaluation techniques to investigate medical image segmentation performance: k-fold cross-validation, leave-one-out cross-validation, per centage-split validation (data set split into test and train set with a given percentage) and detailed validation in which it can be spe cified which images should be used for training and t esting. Except for the detailed validation, all other evalu ation techniques use random sampling to c reate training and testing data sets. During the evaluation, the predefined metrics and loss function for the model a re automatically plotted in figures and saved in t ab separated files for possible further analysis. Next to the performance metrics, the pixel value range and segmentation cl ass frequency of medical images can be analyz ed in the MIScnn evaluation. Also, the resulting predi ction can be compared directly next to the ground truth by creation image visualizations with segmentation overlays. For 3D images, like MRIs, the sl ices with the segmentation overlays are automatically vi sualized in the Graphics Inte rchange Format (GIF). 3 Experiments and Results Here, we analyze and evaluate data from the Kidney T umor Segmentation Challenge 2019 using MIScnn. All results were obtained using the scripts shown in the appendix. 3.1 Kidney T umor Segmentation Challenge 2019 (KiTS19) W ith more than 400 000 kidney cancer diagnoses worldwide in 2018, kidney cancer is under the t op 10 most common cancer types in men and under t he top 15 i n woman [36]. The development of advanced t umor visualization t echniques is highly important for ef fi cient surgical planning. Through the variety in kidney and kidney tumor morphology , the automati c image segmentation is challenging but of great interest [24] . Figure 3: Computed T omography image from the Kidney T umor Segmentation Challenge 2019 data set showing the ground truth segmentation of both kidneys (red) and tumor (blue). The goal of the KiTS19 challenge is the development of reliable and unbiased kidney and kidney tumor semantic segmentation methods [24]. Therefore, the challenge built a data set for arteri al phase abdominal CT scan of 300 kidney cancer patients [24]. The original scans have an image resolution of 512x512 and on average 216 slices (highest slice number is 1059). For all CT scans, a ground truth semantic segmentation was created by exp erts. This semantic segmentation labeled each pi xel with one of three classes: Background, kidney or tumor . 210 of these CT scans with the ground truth segmentation were published during t he training phase of the challenge, whereas 90 CT scans without published ground truth were released afterwards i n the submission phas e. The submitted user predictions for these 90 CT scans will be objectively evaluated and the user models ranked according to their performance. The CT scans were provided in NIfTI format in original resolution and also in interpolated resolution with slice thickness normalization. 3.2 V alidation on the KiTS19 data set with MIScnn For the evaluation of the MIScnn framework usability and data augment ation quality , a subset of 120 CT scans wit h slice thickness normalization were retrieved from the KiTS19 data set. An automatic 3-fold cross-validation was run on this KiTS19 subset with MIScnn. MIScnn Configurations The MIScnn pipeline was configured to perform a multi-class, patch-wise analysis with 80x160x160 patches and a batch size of 2. The pixel value normalization by Z-Score, clipping to the range -79 and 304, as well as resampling to the voxel spacing 3.22x1.62x1.62. For data augmentation, all implemented techniques were used. This includes creating patches through random cropping, scaling, rotations, elastic de formations, mirroring, brightness, contrast, gamma and Gaussian noise augmentations. For prediction, overlapping patches were created with an overlap size of 40x80x80 in x,y ,z directions. The standard 3D U-Net with batch normalization layers were used as deep l earning and convolutional neural network model. The training was performed using the Tversky loss for 1000 e pochs with a starting learning rate of 1E-4 and batch shuffling after each epoch. The cross-validation was run on two Nvidia Quadro P6000 (24GB memory each), using 48GB memory and taking 58 hours. Results W ith the MIScnn pipeline, it was possible to successfully set up a complete, working medical image m ulti-class segmentation pipeline. The 3-fold cross-validation of 120 CT scans for kidney and tumor segmentation were evaluated through several metrics: Tversky loss, soft Dice coeff icient, class-wise Dice coefficient, as well as the sum of categorical cross-entropy and soft Dice coefficient. These scores were computed during the fitting itself, as well as for the prediction with the fitted model. For each cross-validation fold, the training and predictions scores are visualized in figure 4 and sum up in table 1. T able 1: Performance results of the 3-fold cross-validation for tumor and kidney segmentation with a standard 3D U-Net model on 120 CT scans from the KiTS19 challenge. Each metric is computed between the provided ground truth and our model prediction and then averaged between the three folds. Metric T raining V alidation Tversky loss 0.3672 0.4609 Soft Dice similarity coef ficient 0.8776 0.8235 Categorical cross-entropy - 0.8584 - 0.7899 Dice similarity coef ficient: Background - 0.9994 Dice similarity coef ficient: Kidney - 0.9319 Dice similarity coef ficient: T umor - 0.6750 The fitted model achieved a very strong performance for kidney segmentation. The ki dney Dice coeff icient had a m edian around 0.9544. The tumor segmentation predi ction showed a considerably good but weaker performance than the kidney with a median a round 0.7912. S till, the predictive power is ve ry im pressive in the context of using only the standard U-Net architecture with mostly default hyperparameters. In the medical perspective, through t he variety in kidney tumor morphology , which is one of the reasons for the KiTS19 challenge, the weaker t umor results are quite reasonable [24]. Also, the models were trained with only 38% of the original KiTS19 data set due to 80 images for t raining and 40 for testing were randomly selected. The r emaining 90 CT s were excluded in order to reduce run time in the cross-validation. Nevertheless, it was possible to build a powerful pipeline for kidney t umor segmentation with MIScnn resulting into a model with high performance, which is directly comparable with modern, optimized, standalone pipeline [7,8,1 1,26 ]. Figure 4: Performance evaluation of the standard 3D U-Net model for kidney and tumor prediction with a 3-fold cross-validation on the 120 CT data s et from the KiTS19 challenge. LEFT: Tversky loss against epochs illustrating loss development during training for the corresponding test and train data sets. Each point represents the average Tversky loss between the cross-validation folds. CENTER: Class-wise Dice coefficient against epochs illustrating soft Dice similarity coefficient development during training for the corresponding test and train data sets. Each point represents the average soft Dice similarity c oefficient between the cross-valid ation folds. RIGHT : Dice similarity coefficient distribution for the kidney and tumor for all samples of th e cross-validation. Besides the computed metrics, MIScnn c reated segmentation visualizations for manual comparison between ground truth and prediction. As illustrated in figure 5, the predicted semantic segmentation of kidney and tumors is highly accurate. 4 Discussion 4.1 MIScnn framework W ith excellent performing convolutional neu ral network and de ep l earning models like the U-Net, the urge to m ove automatic medical image segment ation from the research l abs i nto practical application in cli nics is uprising. Still, the landscape of standalone pipelines of top performing models, designed only for a single specific public data set, handicaps this progress. The goal of MIScnn is t o provide a high-level API to setup a medical image segmentation pipeli ne with preprocessing, data augmentation, model architectur e selection and model utilization. MIScnn offers a highly configurable and open-source pipeline with several int erfaces for custom deep learning models, image formats or fitting metrics. The modular structure of MIScnn allows a medical image segmentation novice to setup a fun ctional pipeline for a custom data set in just a f ew l ines of code. Addit i onally , switchable models and an automatic ev aluation functionality allow robust and unbiased comparisons between deep learning models. An universal framework for medical image segmentation, following the Python phi losophy of si mple and intuitive modules, is an important step in contributing to practical application development. Figure 5: Computed T omography scans of kidney tumors from the Kidney Tumor Segmentation Challenge 2019 data set showing the kidney (red) and tumor (blue) segmentation as overlays. The images show the segmentation differences between the ground truth provided by the KiTS19 challenge and the prediction from the standard 3D U-Net models of our 3-fold cross-validation . 4.2 Use Case: Kidney T u mor Segmentation Challenge In order to show the reliability of MIScnn, a pipeline was setup for kidney tumors segmentation on a CT image data set. The popular and state-of-the-art standa rd U-Net were used as deep learning model with up-to-date data augmentation. Only a subset of the original KiTS19 data set were used to reduce run time of the cross-validation. The resulting performance and predictive power of the MIScnn showed impr essive predictions and a very good segmentation performance. W e proved that with just a few lines of codes using the MIScnn framework, it was possible to successfully build a powerful pipeline for medical image segmentation. Y et, this performance was achieved using only the standard U-Net model. Fast switching the model to a more precise architecture for hi gh resolution images, like the Dense U-Net model, would probably result into an even better performance [10]. However , this gain would go hand in hand with an increased fitting time and higher GPU memory requirement, which was not possible with our current s haring schedule for GPU hardware. Nevertheless, the possibility of swift switching between models to compare their performance on a data set is a promising step forward in the field of medical image segmentation. 4.3 Road Map and Futur e Dir ection The active MIScnn development is currently focused on multiple key features: Adding further data I/O i nterfaces for the most common medical image formats li ke DICOM, extend pr eprocessing and data augmentation m ethods, implement more ef ficient patch skipping techniques instead of excluding e very blank pa tch (e.g. de noising patch skipping) and implementation of an open interface for custom preprocessing techniques for specific image types like MRIs. Next to the planned feature implementations, the MIScnn road map includes the model library extension with more state-of-the-art deep learning models for medical image segmentation. Additi onally , an objective comparison of the U-Net m odel version variety is outl ined t o get m ore insights on different model performances with the same pipeline. Community contributions in terms of implementations or critique are w elcomed and can be included a fter evaluation. Currently , MIScnn already offers a robust pipeline for medical image segmentation, nonetheless, it will still be regularly updated and extended in the future. 4.4 MIScnn A vailability The MIScnn framework can be di rectly installed as a Python library using pip i nstall miscnn. Additional ly , the source code is av ailable in the Git repository: https://github.com/frankkramer-lab/MIScnn . MIScnn is licensed under the open -source GNU General Public License V ersion 3. The cod e of the cross-validation experiment for the Kidney T um or Segmentation Challenge is available as a Jupyter Notebook in the official Git repository . 5 Conclusion In this paper , we have introduced the open-source Python library MIScnn: A framework for medical image segmentation with convolutional neural networks and deep learning. The intuitive API allows fast building medical image segmentation pipelines including dat a I/O, preprocessing, data augmentation, patch-wise analysis, metrics, a library with state-of-the-art deep learning models and model utilization like training, prediction, as w ell as fully automatic evaluation (e.g. cross- validation). High configurability and multiple open interfaces allow users to fully customize the pi peline. Thi s framework enables researchers to rapidly set up a complete medical image segmentation pipeline by using j ust a fe w l ines of cod e. W e proved the MIScnn functionality by running an automatic cross-validation on the Kidney Tumor S egmentation Challenge 2019 CT data set resulting i nto a powerful pr edictor . W e hope that it will help migrating medical image segmentation from the research labs into practical applications. Acknowledgments W e want to thank B ernhard Bauer and Fabian R abe for sharing their GPU hardware (2x Nvidia Quadro P6000) with us which was used for this work. W e also want to thank Dennis Klonnek, Florian Auer, Barbara Forastefano, Edmund Müller and Iñaki Soto Rey for their useful comments. Conflicts of Inter est The authors declare no conflicts of interest. References [1] P . Agg arwal, R. V ig, S. Bhadoria, A. C.G.Dethe, Role of Segmentation in Medical Imaging: A Com parative Study , Int. J. Comput. Appl. 29 (201 1) 54–61. doi:10.5120/3525- 4803. [2] Y . Guo, Y . Liu, T . Georgiou, M.S. Lew , A review of semantic segmentation using deep neural networks, Int. J. Multimed. Inf. Retr . 7 (2018) 87– 93. doi:10.1007/s13735-017-0141-z. [3] S. Muhammad, A. Muhammad, M. Adnan, Q. Muhammad, A. M ajdi, M.K. Khan, Medical Image Analysis using Convolutional Neural Networks A Review , J. Med. Syst. (2018) 1–13. [4] G. W ang, A perspective on deep imaging, IEEE Ac cess. 4 (2016) 8914–8924. doi:10.1 109/AC CESS.2016.2624938. [5] G. Litjens, T . Kooi, B.E. Bejnordi, A.A.A. Setio, F . Ciompi, M. Ghafoorian, J.A.W .M. van der Laak, B. van Ginneken, C.I. Sánchez, A survey on deep learning in medical image analysis, Med. Image Anal. 42 (2017) 60–88. doi:10.1016/j.media.2017.07.005. [6] D. Shen, G. W u, H.-I. Suk, Deep Learning in Medical Image Analysis, Annu. Rev . Bio med. Eng. 19 (2017) 221–248. doi:10.1 146/annurev- bioeng-071516-044442. [7] O. Ronneberger , Philipp Fischer , T . Brox, U-Net: Convolutional Networks for Biomedical Image Segmentation, Lect. Notes Comput. Sci. (Including Subser . Lect. Notes Ar tif. Intell. Lect. Notes Bioinformatics). 9351 (2015) 234–241. doi:10.1007/978-3-319-24574-4. [8] Z. Zhou, M.M.R. Siddiquee, N. T ajbakhsh, J. Liang, UNet++: A Nested U-Net Architecture for Medical Image Segmentation, (2018). http://arxiv .org/abs/1807.10165 (accessed July 19, 2019). [9] F . Isensee, J. Petersen, A. K lein, D. Zimmerer , P .F . Jaeger , S. Kohl, J. W asserthal, G. Koehler , T . Norajitra, S. W i rkert, K.H. Maier-Hein, nnU- Net: Self-adapting Framework for U-Net-Based Medical Image Segmentation, (2018). http://arxiv .org/abs/1809.10486 (accessed July 19, 2019). [10] M. Kolařík, R. Burget, V . Uher , K. Říha , M. Dutta, Optimized High Resolution 3D Dense-U-Net Network for Brain and Spine Segmentation, Appl. Sci. 9 (2019) 404. doi:10.3390/app9030404. [1 1] Ö. Çiçek, A. Abdulkadir, S.S. Lienkamp, T . Brox, O. Ronneberger , 3D U-net: Lear ning dense volumetric segmentation from sparse annotation, Lect. Notes Comput. Sci. (Including Subser . Lect. Notes Artif. Inte ll. Lect. Notes Bioinformatics). 9901 LNCS (2016) 424–432. doi:10.1007/978-3-319-46723-8_49. [12] K. Lee, J. Zung, P . Li, V . Jain, H.S. Seung, Superhuman Accuracy on the SNEMI3D Connectomics Challenge, (2017) 1–11. http://arxiv .org/abs/1706.00120. [13] E. Gibson, W . Li, C. Sudre, L. Fidon, D.I. Shakir , G. W ang, Z. Eaton-Rosen, R. Gray , T . Doel, Y . Hu, T . Whyntie, P . Nachev , M. Modat, D.C. Barratt, S. Ourselin, M.J. Cardoso, T . V ercauteren, Nif tyNet: a deep-learning platform for medical imaging, Comput. Methods Programs Biomed. 158 (2018) 11 3–122. doi:10.1016/ j.cmpb.2018.01.025. [14] Neuroimaging Informatics T echnology Initiative, (n.d.). https://nifti.nimh.nih.gov /background (accessed July 19, 2019). [15] S. Roy , A. Carass, J.L. Prince, Patch bas ed intensity normalization of brain MR images, in: Proc. - Int. Symp. Biome d. Imaging, 2013. doi:10.1 109/IS BI.2013.6556482. [16] L.G. Nyú, J.K. Udupa, On standardizing the MR image intensity scale, Magn. Reson. Med. (1999). doi:10.1002/(SICI)1522- 2594(199912)42:6<1072::AID-MRM1 1>3.0.CO;2- M. [17] F . Isensee, K.H. Maier-Hein, An attempt at beating the 3D U-Net, (2019) 1–7. http://arxiv .org/abs/1908.02182. [18] G. Lin, C. Shen, A. V an Den Hengel, I. Reid, Efficient Piecewise T raining of Deep Structured Models for Semantic Segmentation, Proc. IEEE Comput. Soc. Conf. Comput. V is. Pattern Recognit. 2016- Decem (2016) 3194–3203. doi:10.1 109/CVPR.2016.348. [19] Z. Hussain, F . Gimenez, D. Y i, D. Rubin, Diff erential Data Augmentation T e chniques for Medical Imaging Classification T asks., AMIA ... Annu. Symp. Proceedings. AMIA Symp. 2017 (2017) 979–984. https://www .ncbi.nl m.nih.gov/pubmed/29854165. [20] Z. Eaton-rosen, F . Bragman, Improving Data Augmentation for Medical Image Segmentation, Midl. (2018) 1–3. [21] L. Perez, J. W ang, The Ef fective ness of Data Augmentation in Image Classification using Deep Learning, (2017). http://arxiv .org/abs/1712.04621 (accessed July 23, 2019). [22] L. T aylor , G. Nitschke, Improving Deep Learning using Generic Data Augmentation, (2017). http://arxiv .org/abs/1708.06020 (accessed July 23, 2019). [23] G.C.R.C. (DKFZ) Division of Medical Image Computing, batchgenerators: A framework for data augmentation for 2D and 3D image classification and segmentation, (n.d.). https://github.com/MIC-DKFZ/batchgenerators (accessed October 9, 2019). [24] N. Heller , N. Sathianathen, A. K alapara, E. W alczak, K. Moore, H. Kaluzniak, J. Rosenberg, P . Blake, Z. Rengel, M. Oestreich, J. Dean, M. T radewell, A. Shah, R. T ejpaul, Z. Edgerton, M. Peterson, S. Raza, S. Regmi, N. Papanikolopoulos, C. W eight, The KiTS19 Challenge Data: 300 Kidney T umor Cases with Clinical Context, CT Semantic Segmentations, and Surgical Outcomes, (2019). http://arxiv .org/abs/1904.00445 (accessed July 19, 2019). [25] P . Coupé, J. V . Ma njón, V . Fonov , J. Pruessner , M. Robles, D.L. Collins, Patch-based segmentation using expert priors: Application to hippocampus and ventricle segmentation, Neuroimage. 54 (2011 ) 940–954. doi:10.1016 /j.neuroimage.2010.09.018. [26] Z. Zhang, Q. Liu, Y . W ang, Road Extraction by Deep Residual U-Net, IEEE Geosci. Remote Sens. Lett. (2018). doi:10.1 109/LGRS.2018.2802944. [27] V . Iglovikov , A. Shvets, T ernausNet: U-Net with VGG1 1 Encoder Pre-Trained on ImageNet for Image Segmentation, (2018). http://arxiv .org/abs/ 1801.05746 (accessed July 19, 2019). [28] N. Ibtehaz, M.S. Rahman, MultiResUNet : Rethinking the U-Net A rchitecture for Multimodal Biomedical Image Segmentation, (2019). http://arxiv .org/abs/1902.04049 (accessed July 19, 2019). [29] K. Kamnitsas, W . Bai, E. Ferrante, S. McDonagh, M. Sinclair , N. Pawlowski, M. Rajchl, M. Lee, B. Kainz, D. Rueckert, B. Glocker, Ensembles of multiple models and architectures for robust brain tumour segmentation, Lect. Notes Comput. Sci. (Including Subser . Lect. Notes Artif. Intell. Lect. Notes Bioinformatics). 10670 LNCS (2018) 450–462. doi:10.1007/978-3-319-75238-9_38. [30] S. V alverde, M. Salem, M. Cabezas, D. Pareto, J.C. V ilanova, L. Ramió-T orrentà, À. Rovira, J. Salvi, A. Oliver, X. Ll adó, One-shot domain adaptation in multiple sclerosis lesion segmentation using convolutional neural networks, NeuroImage Clin. 21 (2019) 101638. doi:10.1016/j.nicl.2018.101638. [31] T . Brosch, L.Y .W . T ang, Y . Y oo, D.K.B. Li, A. T raboulsee, R. T am, Deep 3D Convolutional Encoder Networks W ith Shortcuts for Multiscale Feature Integration Applied to Multiple Sclerosis Lesion Segmentation, IEEE Trans. Med. Im aging. 35 (2016) 1229–1239. doi:10.1 109/TMI.2016.2528821. [32] G. W ang, W . Li, S. Ourselin, T . V ercauteren, Automatic brain tumor segmentation using cascaded anisotropic convolutional neural networks, Lect. Notes Comput. Sci. (Including Subser . Lect. Notes Artif. Inte ll. Lect. Notes Bioinformatics). 10670 LNCS (2018) 178–190. doi:10.1007/978-3-319-75238-9_16. [33] Chollet, François, others, Keras, (2015). https://keras.io. [34] Martin~Abadi, Ashish~Agarwal, Paul~Barham, Eugene~Brevdo, Zhifeng~Chen, Craig~Citro, Greg~S.~Corrado, Andy~Davis, Jeffrey~Dean, Matthieu~Devin, Sanjay~Ghemawat, Ian~Goodfellow , Andrew~H arp, Geoffrey~Irving, Michael~Isard, Y . Jia, Rafal~Jozefowicz, Lukasz~Kaiser , Manjunath~Kudlur, Josh~Levenber g, Dandelion~Mane , Rajat~Monga, Sherry~Moore, Derek~Murray , Chris~Olah, Mike~Schuster , Jonathon~Shlens, Benoit~Ste iner , Ilya~Sutskever , Kunal~T a lwar , Paul~Tucker , V incent~V anhoucke, V ijay~V asudevan, Fernanda~V iegas, Oriol~V inyals, Pete~W arde n, Martin~W attenberg, Martin~W ic ke, Y uan~Y u, Xiaoqiang~Zheng, T ensorFlow: Large -Scale Machine Learning on Heterogeneous Systems, (2015). https://www . tensorflow .org/. [35] S.S.M. Seyed, D. Erdogmus, A. Gholipour , Tversky loss function for image segmentation using 3D fully convolutional deep networks, (2017). https://arxiv .org/abs/1706.05721. [36] J. Ferlay , M. Colombet, I. Soerjomataram, C. Mathers, D.M. Parkin, M. Piñeros, A. Znaor, F . Bray , Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods, Int. J. Cancer . (2018) ijc.31937. doi:10.1002/ijc.31937.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment