Scalable Neural Architecture Search for 3D Medical Image Segmentation
In this paper, a neural architecture search (NAS) framework is proposed for 3D medical image segmentation, to automatically optimize a neural architecture from a large design space. Our NAS framework searches the structure of each layer including neu…
Authors: Sungwoong Kim, Ildoo Kim, Sungbin Lim
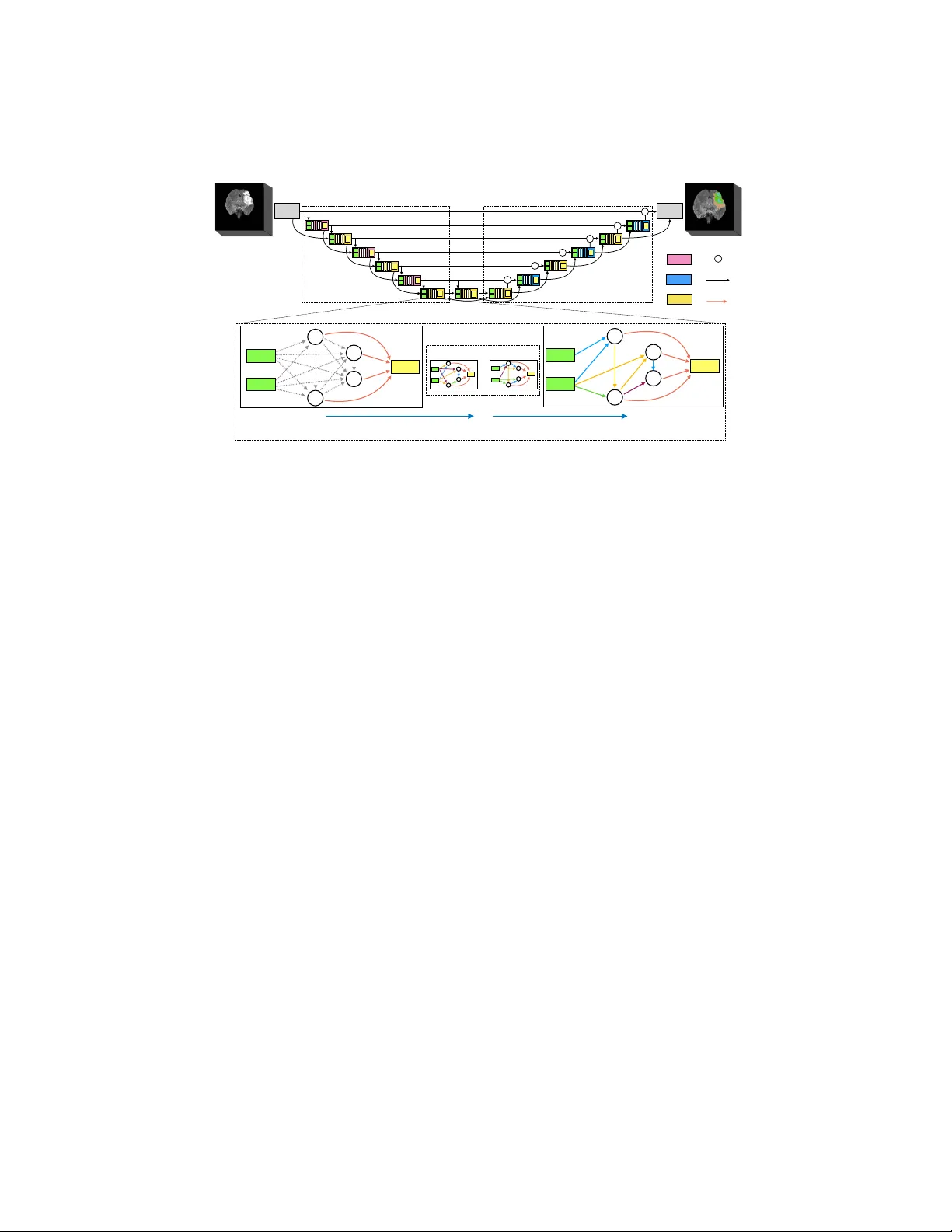
Scalable Neural Arc hitecture Searc h for 3D Medical Image Segmen tation Sungw o ong Kim 1 , Ildo o Kim 1 , Sungbin Lim 1 , W o onh yuk Baek 1 Chiheon Kim 1 , Hyung joo Cho 2 , Bo ogeon Y oon 1 , T aesup Kim 3 1 Kak ao Brain, Pangy o, Seongnam, Gyeonggi, South Korea { swkim, ildoo.kim, sungbin.lim, wbaek, chiheon.kim, eric.yoon } @kakaobrain.com 2 Departmen t of T ransdisciplinary Studies, Seoul National Univ ersity , South Korea joysquare@snu.ac.kr 3 MILA, Universit ´ e de Montr ´ eal, Canada taesup.kim@umontreal.ca Abstract. In this pap er, a neural architecture searc h (NAS) framework is prop osed for 3D medical image segmentation, to automatically opti- mize a neural arc hitecture from a large design space. Our NAS framew ork searc hes the structure of each la yer including neural connectivities and op eration t ypes in both of the encoder and deco der. Since optimizing o ver a large discrete architecture space is difficult due to high-resolution 3D medical images, a no vel sto c hastic sampling algorithm based on a contin- uous relaxation is also prop osed for scalable gradien t based optimization. On the 3D medical image segmen tation tasks with a b enchmark dataset, an automatically designed architecture by the proposed NAS framework outp erforms the h uman-designed 3D U-Net, and moreo ver this optimized arc hitecture is w ell suited to b e transferred for different tasks. Keyw ords: AutoML · Neural architecture search · Medical image seg- men tation. 1 In tro duction Recen tly , deep neural net works hav e b een extensively used for medical image segmen tation tasks. How ev er, in general, their p erformance relies on manual trial-and-error pro cesses for making decisions on the netw ork architecture, hy- p erparameters for training, and pre-/p ost-procedures. Due to b eing restricted to man ual tuning, they would ha ve limitations in performance improv emen t as well as fast transfer to related tasks. Currently , the same problem in the field of gen- eral deep learning has promoted the rapid developmen t of automated machine learning (AutoML). Y et, in contrast to the recen t studies on the use of adv anced AutoML algorithms such as neural architecture searc h (NAS) [ 5 , 9 , 12 ] and neu- ral optimizer search [ 10 ] for general computer vision tasks, only few approaches using simple hyperparameter optimization ha v e been prop osed for medical imag- ing tasks [ 6 , 7 ]. 2 S. Kim et al. Input Output Encoder Decoder Out Reduction cell Expansion cell Normal cell + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== Stem + AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== AAACGnicbVDLSgMxFE1aH7W+Wl26CRZBEMqMCLosunHZgn1AO5RM5k4bmskMSUYopV/gVnd+jTtx68a/MdPOQtseCBzOuZd7cvxEcG0c5wcXilvbO7ulvfL+weHRcaV60tFxqhi0WSxi1fOpBsEltA03AnqJAhr5Arr+5CHzu8+gNI/lk5km4EV0JHnIGTVWal0NKzWn7ixA1ombkxrK0RxWcXEQxCyNQBomqNZ910mMN6PKcCZgXh6kGhLKJnQEfUsljUB7s0XSObmwSkDCWNknDVmofzdmNNJ6Gvl2MqJmrFe9TNzoZYrSod5oBjq7thLNhHfejMskNSDZMlmYCmJikvVEAq6AGTG1hDLF7ecIG1NFmbFtlm1t7mpJ66RzXXedutu6qTXu8wJL6Aydo0vkolvUQI+oidqIIUAv6BW94Xf8gT/x13K0gPOdU/QP+PsXuvOgIg== Summation output 1 0 2 3 Forward input 1 input 0 (a) (c) output 1 0 2 3 input 0 input 1 (b) Stochastic Bi-level optimization Concat ? ? ? ? ? ? ? ? ? ⋯ Sample 1 Sample n Gumbel-Softmax Sampling Fig. 1. An ov erview of architecture search space for image segmentation tasks. Both enco der and deco der alternately stac k normal cells and resizing (reduction, expansion) cells. The directed arro ws b etw een cells indicate the forward paths. Eac h cell is rep- resen ted as a directed acyclic graph (D AG). (a) Initial candidate op erations on the edges in DA G. (b) Gum b el-softmax operation sampling on eac h edge. (c) Induced final arc hitecture from the obtained solution. In this pap er, we prop ose a NAS framework for AutoML in designing neural net works esp ecially for 3D medical image segmen tation. 3D U-Net [ 1 ] has b een p opularly used for segmen ting high-resolution 3D medical images (see [ 4 , 8 , 11 ]) since b oth semantic as well as spatial information can b e efficiently exploited through skip connections b et ween an enco der and a deco der. How ev er, a conv o- lutional blo c k for e ac h lay er in the 3D U-Net has been manually designed with v arious conv olutional filter t yp es, p ooling t yp es, skip-connections, and non-linear activ ation functions. Instead of using the artificial blo c k, we employ a NAS framew ork to obtain an automatically tuned structure of the blo c k, whic h is called a cell, for each la y er in the 3D U-Net where all cell structures and the cor- resp onding neural op eration parameters (e.g. kernel w eights) are simultaneously learned in the end-to-end manner. F or this, four types of cells, enc o der-normal , r e duction , de c o der-normal , and exp ansion cell are defined to compose the en- co der as w ell as the decoder for the learned U-Net architecture (see Figure 1 ). This is different from previous NAS approaches whic h merely use t w o types of cells (normal and reduction) for enco der-only netw orks [ 5 , 9 ]. It is imp ortant to employ a sufficien tly large search space in NAS in gener- ating an improv ed netw ork architecture on a target task. How ever, optimization o ver suc h a large space is difficult due to the extreme memory usage and the long run-time when dealing with high-resolution 3D images. Moreo ver, an ex- act bi-level formulation of NAS has to b e solved on the mixed domain of (1) discrete v ariables regarding neural connections and op eration types in eac h cell and (2) contin uous op eration parameters. This constraint restricts the use of a gradien t method for architecture searching. T o handle this problem, we prop ose Scalable Neural Architecture Searc h for 3D Medical Image Segmentation 3 a nov el contin uous appro ximation using Gum b el-softmax [ 3 ] sampling on the discrete v ariables. This mak es it p ossible to use a sto c hastic gradient descent (SGD) in bi-level optimization. This sampling pro cedure also enables to reduce the computational burden of taking the entire connectivities and operations into accoun t within an outrageously large netw ork originated from the contin uous relaxation. Namely , the prop osed differentiable NAS with sto c hastic sampling supp orts great scalabilit y in terms of solv able large search space with reduced computational cost. T o our b est kno wledge, this is the first work to exploit a complete NAS framework for automatically designing an architecture for the task of 3D medical image segmen tation. Exp erimen tal results on the b enc hmark 3D medical image segmentation dataset show that in comparison to the h uman-designed 3D U-Net [ 1 ], the net- w ork obtained by the prop osed scalable NAS leads to b etter p erformances. It is furthermore sho wn that the found architecture from a task ha ving large amoun ts of labeled data can b e transferred to build a net work for different segmentation tasks with v arious medical mo dalities including MRI and CT that ha ve small amoun ts of lab eled data and ac hieves b etter generalization p erformances. 2 Metho d In this section, we first describ e an architecture search space for 3D medical im- age segmentation. Then, w e present a SGD-based bi-level optimization to simul- taneously learn b oth of the architecture and the corresp onding neural op eration parameters. 2.1 Searc h Space for 3D Medical Image Segmen tation F ollo wing the idea of micro search space p opularly used in the state-of-the-art NAS approaches [ 5 , 9 , 12 ], U-Net-like netw orks, which is comp osed of encoder and deco der lay ers, are designed as rep eated enco der and deco der cells (see Fig- ure 1 ). Here, a cell C is one of four cell-types - enc o der-normal ( C enc ), r e duction ( C red ), de c o der-normal ( C dec ), and exp ansion ( C exp ) - and the normal cells and resizing cells are stac ked alternately with skip connections b et ween the cells in the enco der and the cells in the deco der. Note that an inter-c el l , a copy of C enc , is deplo yed b et ween the enco der and decoder. Every cell takes tw o outputs of the last previous tw o cells as inputs 4 except the first reduction cell whic h takes an output of the predefined first con volutional blo c k, called a stem cell, and then duplicates it as tw o inputs. The segmen tation output is obtained from the predefined last con volutional blo ck, referred to as an out cell. The neural structure in eac h cell C ∈ C := { C enc , C red , C dec , C exp } is repre- sen ted as a directed acyclic graph (DA G). Let G = G ( C ) = ( V , E ) b e the DA G where eac h no de i ∈ V corresp onds to an intermediate feature v ector x i in cell 4 In the deco der, b efore used as one of inputs of the curren t cell, an output of the last previous cell is summed with an output of the encoder cell at the same level. 4 S. Kim et al. C , and each directed edge ( i, j ) ∈ E stands for a connection b etw een no des i and j with a certain op eration o ( i,j ) ∈ O suc h that O denotes the set of all candidate op erations, and x j = P ( i,j ) ∈E o ( i,j ) ( x i ). The output of a cell is a c hannel-wise concatenation of all the intermediate no des. Therefore, the arc hitecture searc h problem now amounts to find the b est combination of all edge op erations in the four cell-t yp es. Basically , e v en the same type of cells can ha ve different structures according to their lay er lev els. Ho wev er, in this work, for simplicit y , all cells that ha ve a common type share a common structure regardless of lay er levels. Note that a zer o op eration is also one of the candidate op erations to optimize the neural connectivities as w ell; zer o means a disconnection b et ween tw o nodes. 2.2 Sto c hastic Bi-Level Optimization with Op eration Sampling W e first represent the selected edge op eration using the one-hot v ector z ( i,j ) = { z ( i,j ) o | o ∈ O} and op eration vector o ( i,j ) = { o ( x i ; θ ( i,j ) o ) | o ∈ O } : o ( i,j ) ( x i ) = h z ( i,j ) , o ( i,j ) i = X o ∈O z ( i,j ) o o ( x i ; θ ( i,j ) o ) , (1) where θ ( i,j ) o denotes the parameter set of the op eration o on edge ( i, j ). Then, finding the b est cell architecture corresp onds to solving the following bi-lev el optimization problem: min Z L val ( Θ ∗ ( Z ) , Z ) s.t. Θ ∗ ( Z ) = argmin Θ L train ( Θ , Z ) , (2) where ( Θ , Z ) = { ( θ ( i,j ) , z ( i,j ) ) | ( i, j ) ∈ E ( C ) , C ∈ C } , and L val and L train are the v alidation loss and training loss, resp ectively . Bi-level pragramming ( 2 ), is hard to solve since its search space is the mixed domain of contin uous v ariables Θ and discrete v ariables Z . DAR TS [ 5 ] try to circumv en t this difficulty b y relaxing Z to a contin uous logits and considering mixed op erations in the edges. This allo ws to use SGD metho d to obtain an approximate solution and derive the final architecture from the relaxed v ariables by taking the op eration with the highest strength on each edge. How ev er, this metho d is infeasible since training the mixed op erations in edge s requires the extremely large memory usage and the long run-time for desired high-resolution 3D image segmentation tasks. T o ov ercome the aforementioned problems, we prop ose a mo dified optimiza- tion, called sto chastic bi-level optimization , b y first treating Z as random discrete v ariables and then replacing ( 2 ) as min α E Z ∼ P α [ L val ( Θ ∗ ( Z ) , Z )] s.t. Θ ∗ ( Z ) = argmin Θ L train ( Θ , Z ) , (3) where P α is the discrete distribution on Z , parameterized b y contin uous v ariable α . Since it is intractable to exactly compute ∇ α E Z ∼ P α [ L val ( Θ ∗ ( Z ) , Z )], we esti- Scalable Neural Architecture Searc h for 3D Medical Image Segmentation 5 Algorithm 1 Gumbel Softmax Sampling Input: Logits α = ( α ( i,j ) o : ( i, j ) ∈ E and o ∈ O ), temp erature τ for ( i, j ) ∈ E do ¯ z ( i,j ) ← softmax (( α ( i,j ) + ( i,j ) ) /τ ), ( i,j ) o ∼ Gumb el (0 , 1) , o ∈ O foreac h p air { o 1 , o 2 } in O do q { o 1 ,o 2 } ← ( ¯ z ( i,j ) o 1 + ¯ z ( i,j ) o 2 ) / ( |O | − 1) end Sample { o 1 , o 2 } with q { o 1 ,o 2 } ˆ z ( i,j ) o 1 , ˆ z ( i,j ) o 2 ← ¯ z ( i,j ) o 1 ¯ z ( i,j ) o 1 + ¯ z ( i,j ) o 2 ¯ z ( i,j ) o 2 ¯ z ( i,j ) o 1 + ¯ z ( i,j ) o 2 , ˆ z ( i,j ) o ← 0 , o / ∈ { o 1 , o 2 } end return ˆ Z mated it through contin uous relaxation using the Gumbel-softmax reparametriza- tion tec hnique [ 3 ] as ∇ α E Z ∼ P α [ L val ( Θ ∗ ( Z ) , Z )] ≈ E ∼ Gumbel (0 , 1) [ ∇ α L val ( Θ ∗ ( ¯ Z ( α, ; τ )) , ¯ Z ( α, ; τ )] , (4) where contin uously relaxed v ariables ¯ Z ( α, ; τ ) = softmax (( α + ) /τ ), τ denotes the temp erature, and is random v ariable drawn from the Gumbel distribution. Here, the exp ectation in ( 4 ) is approximated with -sampling. It is noted that as τ → 0, the distribution of ¯ Z is identical to P α , which means that by annealing τ w e can enforce ¯ Z to be one-hot discrete v ariables Z during training; the relaxed arc hitecture is forced to b e con verged to the final architecture. When alternatively up dating Θ and α by respective gradient descen ts, we again replace ¯ Z with ˆ Z b y sampling tw o op erations on each edge from the Gum b el-softmax with rescaling of the corresponding tw o op eration w eights to b e summed to one, as shown in Algorithm 1 . Note that due to τ -annealing, the num b er of sampled op erations on each edge is naturally reduced from tw o to one during training. This sto chastic op eration sampling supp orts impro ved scalabilit y in terms of solv able large searc h space with small computational cost. 3 Exp erimen ts Dataset and Ev alua tion The prop osed scalable NAS (SCNAS) was ev alu- ated on the three 3D segmentation tasks, (1) brain tumor (MRI, 484 lab eled images, 3 classes), (2) heart (MRI, 20 labeled images, 1 class), and (3) lung (CT, 64 lab eled images, 1 class), from the Medical Segmen tation Decathlon c hallenge (MSD, http://medicaldecathlon.com ) where eac h task has differ- en t input mo dalities and sizes as w ell as differen t foreground classes, which is therefore suitable for ev aluating the generalizability and transferability of the SCNAS. Since the ground-truth labe ls for test images are not pro vided in the MSD dataset, the ev aluation was conducted by 5-fold cross-v alidation (CV) on the training images with the av erage dice similarity co efficient as the metric. Here, the authors in [ 2 ] pro vided their splitting for this 5-fold CV, and we used 6 S. Kim et al. it. F or SCNAS, the training set after the v alidation split was split again in to tw o sets with a ratio of 4:1 for resp ectiv ely optimizing the op eration parameters and the arc hitecture parameters. Implemen tation Details The p erformances of the SCNAS are compared to those obtained b y our baseline 3D U-ResNet [ 4 , 11 ], which mak es use of residual blo c ks, multiple segmentation maps [ 4 ], and attention gates [ 8 ], as well as those from the 3D nnU-Net [ 2 ] that can b e considered as the b est p erformed single mo del from the p ersp ectiv e of c hallenge results. In both of the 3D U-ResNet and the SCNAS, patch-based training and inference w ere carried out such that eac h image w as randomly cropp ed to the region of nonzero v alues with the pre- defined resolution during training, while in testing, the prediction results were obtained b y combining patch-based inference results with 50 p ercen t ov erlap. The input patch size w as basically set to 128 × 128 × 128 and mo dified for eac h task taking median shap es and memory constrain ts in to account just like that used for the 3D nnU-Net in [ 2 ]. Since ev en the same task provides 3D images with heterogeneous v oxel spacings, the input images were first resampled to ha ve an equal vo xel spacing of 0.7mm × 0.7mm × 0.7mm, and then z -normalization w as separately applied to each input channel. F ollo wing [ 2 ], w e also utilized the data augmentation tec hniques at b oth training and testing time with the the same kinds and parameters that used in [ 2 ]. How ev er, unlik e [ 2 ], net w ork- cascade, prediction-ensem bling from different architectures, and the remov al of small connected comp onen ts were not adopted in this ev aluation to solely exam- ine the effects b y the use of NAS in designing the netw ork architecture. The set of op erations O on each edge in the SCNAS consists of the following eigh t op erations: 3 × 3 × 3 conv olutions, depthwise separable dilated 3 × 3 × 3 con volutions with rate 2, 3 and 4, 3 × 3 × 3 max and av erage 3D p ooling, identit y (skip connection), and zero. Here, we used the LeakyReLU-Conv-InstanceNorm for conv olutional op erations. As shown in Figure 1 , the whole netw ork in the SCNAS is composed of 12 automatically designed cells, eac h of which has 4 no des. This num b er of stack ed cells is consistent with that of the 3D U-ResNet in terms of resp ective three times of do wnsampling and upsampling by a factor of 2. Here, all op erations in the reduction cell in the SCNAS are of stride tw o while the expansion cells p erform pre-upsampling for the inputs of the cell. Similar to the 3D U-ResNet, the reduction and expansion cells in the SCNAS resp ectiv ely double and halv e the num b er of output channels of giv en inputs. It it noted that the SCNAS first optimized all of cell architectures using 48 output channels of the stem cell in order to fit a batch size of 1 into a single GPU. Then, a larger netw ork was constructed b y increasing the num ber of stem c hannels to 68 with found cell-top ologies and was retrained from scratch. Here, 68 c hannels makes the computational complexity for inference of a found net work b y SCNAS to be similar to the baseline 3D U-ResNet, whic h has 32 output c hannels in the first conv olutional blolc k, in terms of FLOPs: 419.59 GFLOPs (3D U-ResNet) vs. 424.76 GFLOPs (SCNAS) on the brain tumor task. Scalable Neural Architecture Searc h for 3D Medical Image Segmentation 7 T able 1. Average dice similarity co efficien ts (%) on three tasks of MSD. [ 2 ] obtained their 3D nnU-Net results by mo del selection based on the v alidation loss. Brain T umor (MRI) Heart (MRI) Lung (CT) Label Edema Non-Enhancing Enhancing Av erage Left Atrium T umor 3D nnU-Net [ 2 ] 80.71 62.22 79.07 74.00 92.45 55.87 3D U-ResNet 70.74 56.69 73.23 66.89 91.48 63.28 SCNAS 80.41 59.85 78.50 72.92 91.29 64.82 SCNAS(transfer) - - - - 91.91 68.62 The SCNAS w as trained for 200 ep ochs with a batch size of 4, whic h took one da y on 4 V100 GPUs. In this SCNAS training, the ADAM optimizer w ere used where the initial learning rates / b eta parameters were as set to b e 0 . 025 / (0 . 1 , 0 . 001) for training op eration parameters Θ and 0 . 003 / (0 . 5 , 0 . 999) for training arc hitecture parameters α . If a plateau for 20 ep ochs on the training loss w as detected, the learning rate was reduced by a factor of 10. When retraining the SCNAS mo dels as well as training the 3D U-ResNet mo dels, an initial learning rate of 0 . 0003 and b eta parameters of (0 . 9 , 0 . 999) for the ADAM optimizer w ere used with a batch size of 8, where the learning rate was reduced by a factor of 5 if a training loss w as not reduced for 30 epo chs, and the iteration w as terminated either if it lasted for 500 epo c hs or if the learning rate w as smaller than 10 − 7 . The loss function for b oth 3D U-ResNet and SCNAS is the Jaccard distance [ 4 ]. Results T able 1 shows that the SCNAS pro duced b etter architectures than the (h uman-designed) 3D U-ResNet in terms of the o v erall p erformances. Especially , the p erformances of SCNAS are comparable or even better than those of the 3D nnU-Net [ 2 ]. Here, it should b e noted that the 3D nnU-Net p erformed mo del selection based on the v alidation loss during their 5-fold CV while ours did not tak e any v alidation result into account during training. On the heart and lung segmen tation tasks, which hav e only 20 and 64 lab eled images, resp ectiv ely , the 3D U-ResNet as well as the SCNAS can b e prone to ov erfitting on the train- ing set. Therefore, we transferred the found architecture by SCNAS from the first CV fold of the brain tumor task having 484 lab eled images to these tasks. F or this, w e modified the stem cell arc hitecture to match the num b er of input c hannels according to each task, and the op eration parameters in the trans- ferred architecture were retrained from scratch on eac h task. As a result, the transferred arc hitecture from the brain tumor task ac hieved b etter ge neraliza- tion p erformances in comparison to their own NAS results. Figure 2 shows the optimized cell arc hitectures by SCNAS on the brain tumor task. W e conjecture that the selected dilated con volutions are helpful to reflect a more global con text for improving segmentation results. Example input images and the corresp ond- ing segmentation outputs from the brain tumor task are prese n ted in Figure 3 , whic h sho ws b etter segmentation results b y SCNAS compared to 3D U-ResNet. 8 S. Kim et al. Encoder Normal Expansion Reduction Decoder Normal k 1 0 2 3 k-1 k-2 skip connection conv(3x3x3) avg_pooling(3x3x3) dilated_conv(3x3x3, rate=2) concatenation k 1 0 2 3 k-1 k-2 k 1 0 2 3 k-1 k-2 k 1 0 2 3 k-1 k-2 dilated_conv(3x3x3, rate=3) Fig. 2. The found cell arc hitectures b y SCNAS on the brain tumor task (first CV fold). Flair T1w T1gd T2w Ground T ruth 3D U-ResNet SCNAS Edema Non-Enhancing Enhancing Fig. 3. Segmentation results on the brain tumor task from the 3D U-ResNet and the SCNAS. The first four images from left show the MRI sequences used as input channels. The last image shows the ground truth. 4 Conclusion In this w ork, a complete NAS framework for automatically designing an archi- tecture is prop osed and demonstrated on the b enc hmark dataset of 3D medical image segmentation tasks. In the prop osed framework, NAS is formulated as finding the optimal structure of four types of cells comp osing an enco der as w ell as a deco der. W e introduce a nov el sto c hastic sampling algorithm which re- sults in significant improv ement in terms of the scalability suitable for handling high-resolution 3D medical images. Empirical ev aluation demonstrates that the automatically optimized netw ork via the prop osed NAS outp erforms the man u- ally designed 3D U-Net, and the learned arc hitecture is successfully transferred to differen t segmentation tasks. Bibliograph y [1] C ¸ i¸ cek, ¨ O., Abdulk adir, A., Lienk amp, S.S., Bro x, T., Ronneberger, O.: 3d u- net: Learning dense volumetric segmen tation from sparse annotation. MIC- CAI, Springer, LNCS 9901 , 424–432 (2016) [2] Isensee, F., Petersen, J., Klein, A., Zimmerer, D., Jaeger, P .F., Kohl, S., W asserthal, J., Ko ehler, G., Nora jitra, T., Wirk ert, S., Maier-Hein, K.H.: nn u-net: Self-adapting framew ork for u-net-based medical image segmenta- tion (2018) [3] Jang, E., Gu, S., P o ole, B.: Categorical reparameterization with gumbel- softmax. ICLR (2017) [4] Ka yaliba y , B., Jensen, G., v an der Smagt, P .: Cnn-based segmen tation of medical imaging data. CoRR abs/1701.03056 (2017), http://arxiv. org/abs/1701.03056 [5] Liu, H., Simony an, K., Y ang, Y.: Darts: Differentiable arc hitecture search. arXiv preprin t arXiv:1806.09055 (2018) [6] Mortazi, A., Bagci, U.: Automatically designing cnn architectures for med- ical image segmen tation. MLMI (2018) [7] Naceur, M.B., Saouli, R., Akil, M., Kac houri, R.: F ully automatic brain tumor segmentation using end-to-end incremental deep neural netw orks in mri images. Computer Metho ds and Programs in Biomedicine, Elsevier 166 , 39–49 (2018) [8] Okta y , O., Schlemper, J., F olgo c, L.L., Lee, M.C.H., Heinrich, M.P ., Misaw a, K., Mori, K., McDonagh, S.G., Hammerla, N.Y., Kainz, B., Glo ck er, B., Ruec kert, D.: Atten tion u-net: Learning where to lo ok for the pancreas. MIDL (2018) [9] Pham, H., Guan, M.Y., Zoph, B., Le, Q.V., Dean, J.: Efficien t neural ar- c hitecture search via parameter sharing. ICML (2018) [10] Wic hrowsk a, O., Mahesw aranathan, N., Hoffman, M.W., Colmenarejo, S.G., Denil, M., de F reitas, N., Sohl-Dickstein, J.: Learned optimizers that scale and generalize. ICML (2017) [11] Y u, L., Y ang, X., Chen, H., Qin, J., Heng, P .A.: V olumetric convnets with mixed residual connections for automated prostate segmentation from 3d mr images. AAAI (2017) [12] Zoph, B., V asudev an, V., Shlens, J., Le, Q.V.: Learning transferable archi- tectures for scalable image recognition. CVPR (2018)
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment