A Survey on Active Learning and Human-in-the-Loop Deep Learning for Medical Image Analysis
Fully automatic deep learning has become the state-of-the-art technique for many tasks including image acquisition, analysis and interpretation, and for the extraction of clinically useful information for computer-aided detection, diagnosis, treatmen…
Authors: Samuel Budd, Emma C Robinson, Bernhard Kainz
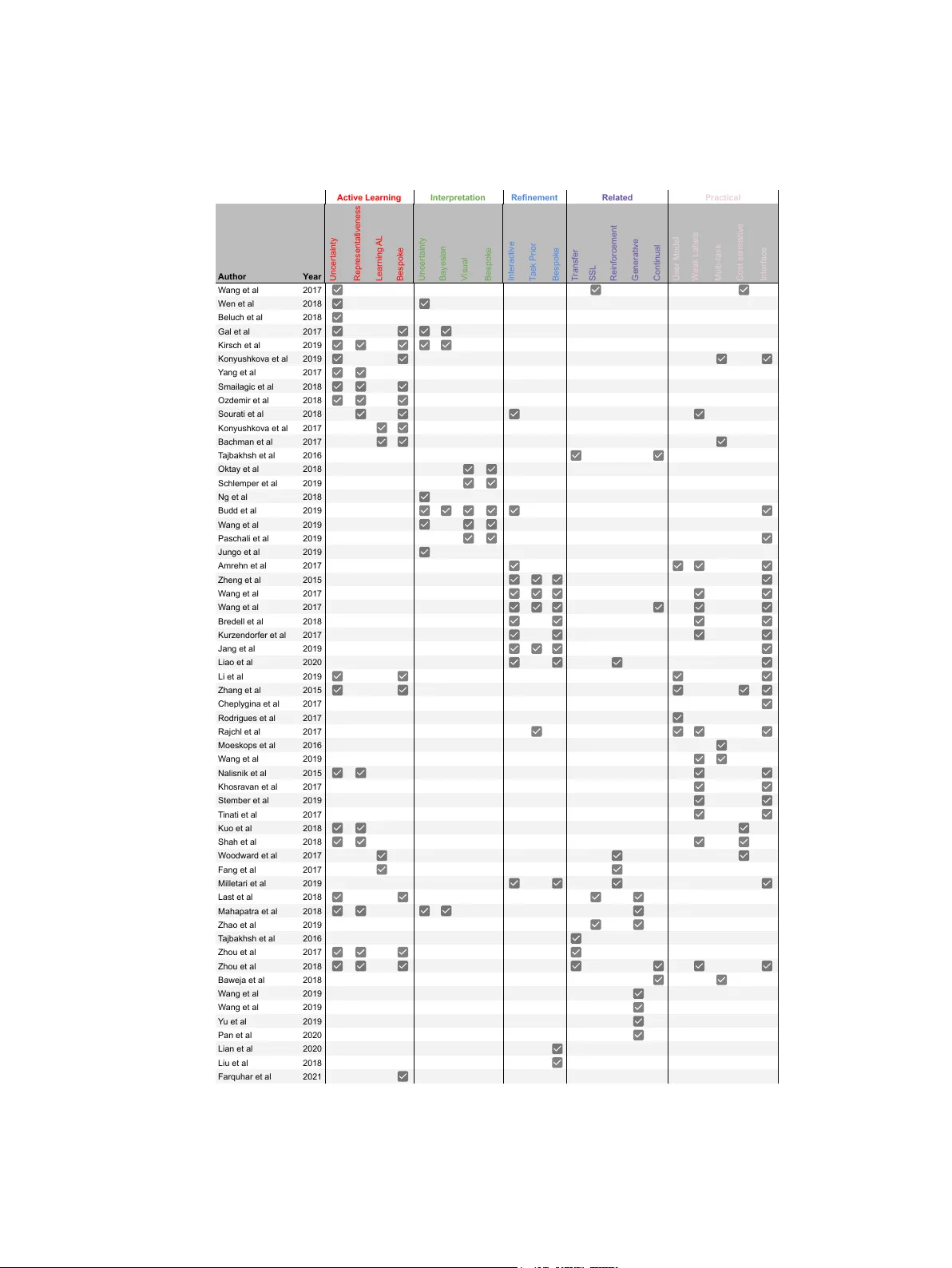
A Surve y on Acti ve Learning and Human-in-the-Loop Deep Learning for Medical Image Analysis Samuel Budd a, , Emma C Robinson b,1 , Bernhard Kainz a,1 a Department of Computing, Imperial Colle ge London, UK b Department of Imaging Sciences, King’ s Colle ge London, UK Abstract Fully automatic deep learning has become the state-of-the-art technique for many tasks including image acquisition, analysis and interpretation, and for the extraction of clinically useful information for computer-aided detection, diagnosis, treatment planning, intervention and therapy . Howe ver , the unique challenges posed by medical image analysis suggest that retaining a human end- user in any deep learning enabled system will be beneficial. In this revie w we in vestigate the role that humans might play in the dev elopment and deplo yment of deep learning enabled diagnostic applications and focus on techniques that will retain a significant input from a human end user . Human-in-the-Loop computing is an area that we see as increasingly important in future research due to the safety-critical nature of working in the medical domain. W e ev aluate four key areas that we consider vital for deep learning in the clinical practice: (1) Active Learning to choose the best data to annotate for optimal model performance; (2) Interaction with model outputs - using iterativ e feedback to steer models to optima for a giv en prediction and o ff ering meaningful ways to interpret and respond to predictions; (3) Pr actical considerations - developing full scale applications and the ke y considerations that need to be made before deployment; (4) Futur e Pr ospective and Unanswered Questions - knowledge gaps and related research fields that will benefit human-in-the-loop computing as they ev olve. W e o ff er our opinions on the most promising directions of research and how v arious aspects of each area might be unified towards common goals. 1. Introduction Medical imaging is a major pillar of clinical decision making and is an inte gral part of many patient journe ys. Information ex- tracted from medical images is clinically useful in many areas such as computer-aided detection, diagnosis, treatment plan- ning, interv ention and therap y . While medical imaging remains a vital component of a myriad of clinical tasks, an increasing shortage of qualified radiologists to interpret complex medical images suggests a clear need for reliable automated methods to alleviate the growing burden on health-care practitioners ( of Ra- diologists , 2017 ). In parallel, medical imaging sciences are benefiting from the dev elopment of novel computational techniques for the analysis of structured data like images. De velopment of algorithms for image acquisition, analysis and interpretation are dri ving inno- vation, particularly in the areas of registration, reconstruction, tracking, segmentation and modelling. Medical images are inherently di ffi cult to interpret, requiring prior expertise to understand. Bio-medical images can be noisy and contain many modality-specific artefacts, acquired under a wide variety of acquisition conditions with di ff erent protocols. Thus, once trained models do not transfer seamlessly from one clinical task or site to another because of an often yawning do- main gap ( Kamnitsas et al. , 2017 ; Ben-David et al. , 2010 ). Su- pervised learning methods require extensiv e relabelling to re- gain initial performance in di ff erent workflo ws. The e xperience and prior knowledge required to work with such data means that there is often lar ge inter- and intra- observer variability in annotating medical data. This not only raises questions about what constitutes a gold-standard ground truth annotation, b ut also results in disagreement of what that ground truth truly is. These issues result in a large cost associ- ated with annotating and re-labelling of medical image datasets, as we require numerous expert annotators (oracles) to perform each annotation and to reach a consensus. In recent years, Deep Learning (DL) has emerged as the state-of-the-art technique for performing man y medical im- age analysis tasks ( T ajbakhsh et al. , 2020 ; T izhoosh and Pan- tanowitz , 2018 ; Shen et al. , 2017 ; Litjens et al. , 2017 ; Suzuki , 2017 ). Dev elopments in the field of computer vision ha ve shown great promise in transferring to medical image analy- sis, and sev eral techniques have been shown to perform as ac- curately as human observ ers ( Haenssle et al. , 2018 ; Mar and Soyer , 2018 ). Ho wever , uptake of DL methods within the clin- ical practice has been limited thus far , largely due to the unique challenges of working with complex medical data, regulatory compliance issues and trust in trained models. W e identify three k ey challenges when de veloping DL en- abled applications for medical image analysis in a clinical set- ting: 1. Lack of Training Data: Supervised DL techniques tra- ditionally rely on a lar ge and ev en distribution of accu- rately annotated data points, and while more medical im- age datasets are becoming av ailable, the time, cost and ef- fort required to annotate such datasets remains significant. 2. The Final Percent: DL techniques have achiev ed state-of- the-art performance for medical image analysis tasks, but in safety-critical domains e ven the smallest of errors can cause catastrophic results do wnstream. Achie ving clini- cally credible output may require interacti ve interpretation of predictions (from an oracle) to be useful in practice, i.e users must hav e the capability to correct and o verride auto- mated predictions for them to meet any acceptance criteria required. 3. Transparency and Interpretability: At present, most DL applications are considered to be a ’black-box’ where the user has limited meaningful ways of interpreting, under- standing or correcting how a model has made its predic- tion. Credence is a detrimental feature for medical appli- cations as information from a wide variety of sources must be ev aluated in order to make clinical decisions. Further indication of how a model has reached a predicted con- clusion is needed in order to foster trust for DL enabled systems and allo w users to weigh automated predictions appropriately . There is concerted e ff ort in the medical image analysis re- search community to apply DL methods to various medical im- age analysis tasks, and these are sho wing great promise. W e refer the reader to a number of revie ws of DL in medical imag- ing ( Hesamian et al. , 2019 ; Lundervold and Lundervold , 2019 ; Y amashita et al. , 2018 ). These w orks primarily focus on the de- velopment of predictive models for a specific task and demon- strate state-of-the-art performance for that task. This revie w aims to give an overvie w of where humans will remain in volved in the dev elopment, deployment and practical use of DL systems for medical image analysis. W e focus on medical image segmentation techniques to explore the role of human end users in DL enabled systems. Automating image interpretation tasks like image se gmen- tation su ff ers from all of the dra wbacks incurred by medical image data described abov e. There are many emerging tech- niques that seek to alleviate the added complexity of working with medical image data to perform automated segmentation of images. Segmentation seeks to divide an image into seman- tically meaningful regions (sets of pixels) in order to perform a number of downstream tasks, e.g. biometric measurements. Manually assigning a label to each pixel of an image is a la- borious task and as such automated segmentation methods are important in practice. Advances in DL techniques such as Ac- tiv e Learning (AL) and Human-in-the-Loop computing applied to segmentation problems have shown progress in overcoming the key challenges outlined abov e and these are the studies this revie w focuses on. W e categorise each study based on the na- ture of human interaction proposed and broadly divide them between which of the three key challenges the y address. Section 2 introduces Acti ve Learning, a branch of Machine Learning (ML) and Human-in-the-Loop Computing that seeks to find the most informative samples from an unlabelled distri- bution to be annotated next. By training on the most informa- tiv e subset of samples, related work can achiev e state-of-the-art performance while reducing the costly annotation burden asso- ciated with annotating medical image data. Section 3 ev aluates techniques used to refine model predic- tions in response to user feedback, guiding models towards more accurate per-image predictions. W e ev aluate techniques that seek to improve interpretability of automated predictions and how models pro vide feedback on their own outputs to guide users tow ards better decision making. Section 4 e valuates the k ey practical considerations of de vel- oping and deploying Human-in-the-Loop DL enabled systems in practice and outlines the work being done in these areas that addresses the three key challenges identified above. These ar- eas a re human focused and assess how human end users might interact with these systems. In Section 5 we discuss related areas of ML and DL research that are having an impact on AL and Human-in-the-Loop Com- puting and are beginning to influence the three key challenges outlined. W e o ff er our opinions on the future directions of Human-in-the-Loop DL research and how many of the tech- niques ev aluated might be combined to work to wards common goals. 2. Active Learning In this section we assume a scenario in which a lar ge pool of un-annotated data U is av ailable to us, and that we ha ve an ora- cle (or group of oracles) from which we can request annotations for ev ery un-annotated data point x U to add to an annotated set L . W e wish to train some model f ( x | L ∗ ) where L ∗ ⊆ L and consider methods that rely on annotated data to do so. A brute- force solution to this problem would be to ask the oracle(s) to annotate e very x U such that L ∗ = L , b ut this is rarely a practical or cost-e ff ecti ve solution due to the unique challenges associ- ated with annotating biomedical image data. It is theorised that there is some L ∗ that achiev es equiv alent performance to L , i.e. f ( x | L ∗ ) ≈ f ( x | L ). A model trained on some optimal subset L ∗ of a dataset might achie ve equiv alent performance to a model trained on the entire, annotated dataset. Active Learning (AL) is the branch of machine learning that seeks to find this opti- mal subset L ∗ giv en a current model f 0 ( x | L 0 ), where L 0 is an intermediate annotated dataset, and an un-annotated dataset U . AL methods aim to iteratively seek the most informativ e data- points x ∗ i for training a model, under the assumption that both the model and the un-annotated dataset will evolv e over time, rather than selecting a fix ed subset once to be used for training. In a wider context and before the advent of DL, Settles ( 2009 ) revie wed this field as a state-of-the-art ML methodology . A typical AL frame work, as outlined in Figure 1 , consists of a method to ev aluate the informativeness of each un-annotated data point x U giv en f 0 ( x U | L 0 ), tied hea vily to the choice of query type , after which all chosen data-points are required to be an- notated. Once new annotations have been acquired, the AL framew ork must use the new data to improv e the model. This is normally done by either r etraining the entire model using 2 Annotate for Training Annotators Predict Present Performance Threshold Active Learning Sample Selection Automatic Predictions Unlabelled Data Intermediate Model T rained Model Fig. 1. Over view of Active Learning frameworks. all available annotated data L 0 , or by fine-tuning the netw ork using the most recently annotated data-points x ∗ i . Using this approach, state-of-the-art performance can be achie ved using fewer annotations for sev eral bio-medical image analysis tasks, as shown in the methods discussed in this section, thus widen- ing the annotation bottleneck and reducing the costs associated with dev eloping DL enabled systems from un-annotated data. 2.1. Query T ypes In every AL frame work the first choice to be made is what type of query we wish to mak e using a model and un-annotated dataset. There are currently three main choices av ailable and each lends itself to a particular scenario dependant on what type of un-annotated data we hav e access to, and what question we wish to ask the oracle(s). Str eam-based Selective Sampling assumes a continuous stream of incoming un-annotated data-points x U ( Atlas et al. ( 1990 ); Cohn et al. ( 1994 )). The current model and an informa- tiveness measure I ( x U ) are used to decide, for each incoming data-point, whether or not to ask the oracle(s) for an annotation ( Dagan and Engelson ( 1995 )). This query type is usually com- putationally inexpensi ve but o ff ers limited performance benefits due to the isolated nature of each decision: the wider context of the underlying distribution is not considered, thus balancing exploration and e xploitation of the distribution is less well cap- tured than in other query types. Another disadv antage of this query type is calibrating the threshold to use for the chosen in- formativ eness measure such that we do not request annotations for ev ery incoming data-point, and that we do not reject annota- tions for too man y data-points resulting in valuable information being lost. Membership Query Synthesis assumes that rather than draw- ing from a real-world distribution of data-points, we instead generate a data-point x ∗ G that needs to be annotated ( Angluin ( 1988 )). The generated data-point is what the current model ’believ es’ will be most informativ e to itself. This data-point is then annotated by the oracle(s) ( Angluin ( 2001 )), this can be very e ffi cient in finite domains. This approach may su ff er from the same drawbacks as Str eam-based methods as a model may ha ve no kno wledge of unseen areas of the distribution, and thus be unable to request annotations of those areas. Issues can arise where queries can request annotations for data-points that mak e no sense to a human oracle ( Lang and Baum ( 1992 )), and are not representativ e of the actual distribution that is be- ing modelled, stream based and pool based sampling methods were proposed to ov ercome these issues ( Settles ( 2009 )). Nev- ertheless, recent advances of Generative Adversarial Networks (GANs) have shown great promise in generating data-points that mimic real-world distributions for many di ff erent types of data, including biomedical images, that may go some way to ad- dressing the ke y issue with using query synthesis for complex distributions, which we discuss in Section 2.2.3 . This query type can be adv antageous in scenarios where the distrib ution to generate is fully understood, or domains in which annotations are acquired autonomously instead of from humans ( King et al. ( 2004 , 2009 )). P ool-based Sampling assumes a large un-annotated real- world dataset U to draw samples from and seeks to select a batch of N samples x ∗ 0 , ..., x ∗ N from the distrib ution to request labels for ( Lewis and Catlett ( 1994 ). P ool-based methods usu- ally use the current model to make a prediction on each un- annotated data point to obtain a ranked measure of informative- ness I ( x U | f 0 ( x U | L 0 )) for ev ery data-point in the un-annotated set, and select the top N samples using this metric to be an- notated by the oracle(s). Pool based sampling has been applied to several real world tasks, prior to the advent of deep learning ( Lewis and Catlett ( 1994 ); McCallum and Nigam ( 1998 ); Set- tles and Cra ven ( 2008 ); Zhang and Chen ( 2002 ); Hauptmann et al. ( 2006 ). These methods can be computationally expensi ve as ev ery iteration requires a metric ev aluation for every data- point in the distribution. Howe ver , these methods have shown to be the most promising when combined with DL methods, which inherently rely on a batch-based training scheme. Pool based sampling is used in the majority of methods discussed in the rest of this section unless stated otherwise. While pool- based methods hold adv antages ov er other methods in terms of finding the most informativ e annotations to acquire, scenarios in which stream based or synthesis based queries are advanta- geous are also common, such as when memory or processing power is limited for e xample in mobile or embedded de vices ( Settles ( 2009 )). 2.2. Evaluating Informativeness In dev eloping an AL frame work, once a query type has been selected, the next question to ask is how to measure the infor- mativ eness I ( x U ) of each of the data-points? Many varying ap- proaches hav e been taken to quantifying the informati veness of a sample gi ven a model and an underlying distribution. Here we sort these metrics by the le vel of human interpretability they o ff er . T raditionally , AL methods employ hand-designed heuristics to quantify what we as humans belie ve makes something infor - mativ e. A v ariety of model specific metrics seek to quantify what the e ff ect of using a sample for training w ould ha ve on the 3 model, e.g., the biggest change in model parameters. Howe ver , these methods are less pre valent than human designed heuris- tics due to the computational challenge of applying these to DL models with a large number of parameters. Finally some meth- ods are emerging that are completely agnostic to human inter- pretability of informativ eness and instead seek to learn the best selection policy from av ailable data and previous iterations, as discussed in detail in Section 2.2.4 . 2.2.1. Uncertainty The main family of informativ eness measures falls into cal- culating uncertainty . It is argued that the more uncertain a pre- diction is, the more information we can gain by including the ground truth for that sample in the training set. There are se veral ways of calculating uncertainty from dif- ferent ML / DL models. When considering DL for segmentation the most simple measure is the sum of lowest class probability for each pixel in a giv en image segmentation. It is argued that more certain predictions will hav e high pixel-wise class proba- bilities, so the lower the sum of the minimum class probability ov er each pixel in an image, the more certain a prediction is considered to be: x ∗ LC = argmax x 1 − P θ ( ˆ y | x ) where ˆ y = argmax y P θ ( y | x ). This is a f airly intuiti ve way of thinking about uncertainty and o ff ers a means to rank uncer- tainty of samples within a distribution. W e refer to the method abov e as least confident sampling where the samples with the highest uncertainty are selected for labelling ( Settles , 2009 ). A drawback of least confident sampling is that it only consid- ers information about the most probable label, and discards the information about the remaining label distrib ution. T wo alter- nativ e methods have been proposed that alleviate this concern. The first, called mar gin sampling ( Settles , 2009 ), can be used in a multi-class setting and considers the first and second most probable labels under the model and calculates the di ff erence between them: x ∗ M = argmin x P θ ( ˆ y 1 | x ) − P θ ( ˆ y 2 | x ) where ˆ y 1 and ˆ y 2 are the first and second most probable labels under the current model, respectiv ely . The intuition here is that the larger the margin is between the two most probable labels, the more confident the model is in assigning that label. The second, more popular approach is to use entr opy ( Shannon ) as an uncertainty measure: x ∗ E = argmax x − X i P ( y i | x ) log P ( y i | x ) where y i ranges across all possible annotations. Entropy is used to measure the amount of information required to encode a dis- tribution and as such, is often thought of as a measure of uncer- tainty in machine learning. For binary classification, all three methods reduce to querying for the data-point with a class pos- terior closest to 0.5. The ability of entr opy to generalise easily to probabilistic multi-class annotations, as well as models for more complex structured data-points has made it the most pop- ular choice for uncertainty based query strategies Settles and Crav en ( 2008 ). Using one of the above measures, un-annotated samples are ranked and the most ’uncertain’ cases are chosen for the next round of annotation. There hav e been many recent uses of un- certainty based sampling in AL methods in the DL field and these are discussed next. W ang et al. ( 2017 ) propose the Cost-E ff ective Acti ve Learn- ing (CEAL) method for deep image classification. The CEAL methods is initialised with a set of unlabelled sample U , ini- tially labelled samples L , a choice of pool size K , a high con- fidence sample selection threshold ω , a threshold decay rate d r , a maximum iteration number T and a fine-tuning interval t . After initialisation, CNN weights W are initialised with L and the model is used to make predictions on each data-point in U . CEAL explores using each of the three uncertainty methods described abov e to obtain K uncertain data-points to be man- ually annotated and added to D L . So far the CEAL method follows very closely the approach outlined in traditional activ e learning methods as described abov e, but they introduce an ad- ditional training step where the most confident samples (whose entr opy is less than ω ) from U are added to D H . D L and D H are then used to fine-tune W for t iterations. CEAL then updates ω before the pseudo-labels from D H are discarded and each data-point is added back to U , while D L is added to L . This pro- cess repeats for T iterations. The authors describe this approach of simultaneously learning from manual labels of the most un- certain annotations and predicted labels of the least uncertain annotations as complementary sampling . The CEAL method showed that state-of-the-art performance can be achiev ed using less than 60% of a vailable data for tw o non-medical datasets (CA CD and Caltech-256) for face recognition and object cate- gorisation. W en et al. ( 2018 ) propose an active learning method that uses uncertainty sampling to support quality control of nucleus seg- mentation in pathology images. Their work compares the per- formance improv ements achiev ed through acti ve learning for three di ff erent f amilies of algorithms: Support V ector Machines (SVM), Random Forest (RF) and Con volutional Neural Net- works (CNN). They sho w that CNNs achiev e the greatest accu- racy , requiring significantly fewer iterations to achiev e equiv a- lent accuracy to the SVMs and RFs. Another common method of estimating informativ eness is to measure the agreement between multiple models perform- ing the same task. It is argued that more disagreement found between predictions on the same data point implies a higher lev el of uncertainty . These methods are referred to as Query by consensus and are generally applied when Ensembling is used to improve performance - i.e, training multiple models to per- form the same task under slightly di ff erent parameters / settings Settles ( 2009 ). Ensembling methods ha ve shown to measure in- formativ eness well, but at the cost of computational resources - multiple models need to be trained and maintained, and each of these needs to be updated in the presence of newly selected training samples. Nev ertheless, Beluch Bcai et al. ( 2018 ) demonstrate the 4 power of ensembles for activ e learning and compare to al- ternativ es to ensembling. They specifically compare the per- formance of acquisition functions and uncertainty estimation methods for acti ve learning with CNNs for image classification tasks and sho w that ensemble based uncertainties outperform other methods of uncertainty estimation such as ’MC Dropout’. They find that the di ff erence in activ e learning performance can be explained by a combination of decreased model capacity and lower di versity of MC dropout ensembles. A good performance is demonstrated on a diabetic retinopathy diagnosis task. K onyushko va et al. ( 2019 ) propose an activ e learning ap- proach that exploits geometric smoothness priors in the image space to aid the segmentation process. They use traditional un- certainty measures to estimate which pixels should be annotated next, and introduce nov el criteria for uncertainty in multi-class settings. They exploit geometric uncertainty by estimating the entropy of the probability of supervox els belonging to a class giv en the predictions of its neighbours and combine these to en- courage selection of uncertain regions in areas of non-smooth transition between classes. They demonstrate state-of-the-art performance on mitochondria se gmentation from EM images and on an MRI tumour segmentation task for both binary and multi-class segmentations. They suggest that exploiting geo- metric properties of images is useful to answer the questions of where to annotate next and by reducing 3D annotations to 2D annotations provide a possible answer to how to annotate the data, and that addressing both jointly can bring additional ben- efits to the annotation method, howe ver they ackno wledge that it would impossible to design bespoke selection strategies this way for e very ne w task at hand. Gal et al. ( 2017 ) introduce the use of Bayesian CNNs for Activ e Learning with ’Bayesian Active Learning by Disagree- ment’ or B ALD, and show that the use of Bayesian CNNs out- perform deterministic CNNs in the context of Active Learning, and exploit this through the use of a new acquisition function that chooses data-points expected to maximise the information gained about the model parameters i.e maximise the mutual in- formation between predictions and model posterior . This ap- proach uses a Bayesian CNN (induced using Dropout during inference Gal and Ghahramani ( 2016 )), to produce a single pre- diction using all parameters of the network for each unlabelled data-point, and a set of stochastic predictions for each unla- belled data-point, generated with dropout enabled. The BALD acquisition function is then calculated as the di ff erence between the entrop y of the av erage prediction and av erage entropy of stochastic predictions. Intuitiv ely this function selects data- points for which the model is uncertain on av erage, b ut there exist model parameters that produce disagreeing predicted an- notations with high certainty . They demonstrate their approach for skin cancer diagnosis from skin lesion images to sho w sig- nificant performance improvements ov er uniform sampling us- ing the B ALD method for sample selection. While this method has been sho wn to be particularly e ff ectiv e for AL, when query- ing batches of data-points, it often results in many very similar , redundant data-points being acquired when used in a greedy fashion, as such BatchB ALD was introduced to alle viate this problem Kirsch et al. ( 2019 ). The BatchB ALD approach in- stead no longer calculates the mutual information between a single sample predictions and model posterior , but instead cal- culates the mutual information between a batch of samples and the model posterior to jointly score the batch of samples, en- abling BatchB ALD to more accurately ev aluate the joint mutual information and select batches of samples for annotation that re- sult in less redundant data-points being selected together in an acquired batch. This extension is an example of the motiv ation behind Section 2.2.2 in which we discuss methods that move beyond pure uncertainty based methods and begin to measure div ersity among selected samples to reduce redundant annota- tion. 2.2.2. Representativeness Many AL framew orks extend selection strategies to include some measure of repr esentativeness in addition to an uncer- tainty measure. The intuition behind including a representati ve- ness measure is that methods only concerned with uncertainty hav e the potential to focus only on small regions of the distri- bution, and that training on samples from the same area of the distribution will introduce redundanc y to the selection strategy , or may ske w the model towards a particular area of the distri- bution. The addition of a representativeness measure seeks to encourage selection strategies to sample from di ff erent areas of the distribution, and to increase the diversity of samples, thus improving AL performance. A sample with a high represen- tativ eness cov ers the information for many images in the same area of the distrib ution, so there is less need to include many samples cov ered by a representativ e image. T o this end, Y ang et al. ( 2017 ) present Suggestive Anno- tation, a deep active learning framework for medical image segmentation, which uses an alternative formulation of uncer - tainty sampling combined with a form of representativeness density weighting. Their method consists of training multi- ple models that each exclude a portion of the training data, which are used to calculate an ensemble based uncertainty mea- sure. They formulate choosing the most representative exam- ple as a generalised version of the maximum set-cov er problem (NP Hard) and o ff er a greedy approach to selecting the most representativ e images using feature vectors from their models. They demonstrate state-of-the-art performance using 50% of the av ailable data on the MICCAI Gland segmentation chal- lenge and a lymph node segmentation task. Smailagic et al. ( 2018 ) propose MedAL , an activ e learning framew ork for medical image segmentation. They propose a sampling method that combines uncertainty , and distance be- tween feature descriptors, to extract the most informativ e sam- ples from an unlabelled data-set. Once an initial model has been trained, the MedAL method selects data-points to be labelled by first filtering out unlabelled data-points with a predictiv e en- tropy below a threshold. From this set the CNN being trained is used to generate feature descriptors for each data-point by taking the output of intermediate layers of the CNN, these fea- ture descriptors are then compared amongst each other using a v ariety of distance functions (e.g ’Euclidian’, ’Russellrao’, ’Cosine’) in order to find the feature descriptors which are most distant from each other . The data-point with the highest average 5 distance to all other unlabelled data-points (above the entropy threshold) is selected for annotation. In this way , the MedAL acquisition function finds the set of data-points that are both in- formativ e to the model, and incur the least redundancy between them by sampling from areas of the input distrib ution most dis- tant from each other . MedAL method initialises the model in a novel way by leveraging existing computer vision image de- scriptors to find the images that are most dissimilar to each other and thus cover a larger area of the image distribution to use as the initial training set after annotation. They sho w good results on three di ff erent medical image analysis tasks, achieving the baseline accuracy with less training data than random or pure uncertainty based methods. Ozdemir et al. ( 2018 ) propose a Borda-count based combi- nation of an uncertainty and a representativ eness measure to se- lect the next batch of samples. Uncertainty is measured as the vox el-wise v ariance of N predictions using MC dropout in their model. They introduce new representativeness measures such as ’Content Distance’, defined as the mean squared error be- tween layer activ ation responses of a pre-trained classification network. They extend this contribution by encoding represen- tativ eness by maximum entropy to optimise network weights using an nov el entropy loss function. Sourati et al. ( 2018 ) propose a novel method for ensuring di- versity among queried samples by calculating the Fisher Infor- mation (FI), for the first time in CNNs. Here, e ffi cient compu- tation is enabled by the gradient computations of propagation to allo w FI to be calculated on the large parameter space of CNNs. They demonstrate the performance of their approach on two di ff erent flavours of task: a) semi-automatic segmentation of a particular subject (from a di ff erent group / di ff erent pathol- ogy not present in the original training data) where iterati vely labelling small numbers of voxels queried by AL achie ves ac- curate segmentation for that subject; and b) using AL to build a model generalisable to all images in a gi ven data-set. They show that in both these scenarios the FI-based AL improv es performance after labelling a small percentage of voxels, out- performed random sampling and achie ved higher accurac y than entropy based querying. 2.2.3. Generative Adversarial Networks for Informativeness Generativ e Adv ersarial Network (GAN) based methods hav e been applied to se veral areas of medical imaging such as de- noising, modality transfer, abnormality detection, and for im- age synthesis, directly applicable to AL scenarios. This o ff ers an alternative (or addition) to the man y data augmentation tech- niques used to expand limited data-sets Y i et al. ( 2019 ) and a DL approach to Membership Query Synthesis . Last et al. ( 2018 ) propose a conditional GAN (cGAN) based method for activ e learning where they use the discriminator D output as a measure of uncertainty of the proposed se gmenta- tions, and use this metric to rank samples from the unlabelled data-set. From this ranking the most uncertain samples are pre- sented to an oracle for segmentation and the least uncertain im- ages are included in the labelled data-set as pseudo gr ound truth labels. They show their method approaches increasing accurac y as the percentage of interactively annotated samples increases - reaching the performance of fully supervised benchmark meth- ods using only 80% of the labels. This work motiv ates the use of GAN discriminator scores as a measure of prediction uncer- tainty . Mahapatra et al. ( 2018 ) also use a cGAN to generate chest X- Ray images conditioned on a real image, and using a Bayesian neural network to assess the informativ eness of each generated sample, decide whether each generated sample should be used as training data. If so, is used to fine-tune the network. They demonstrate that the approach can achiev e comparable perfor- mance to training on the fully annotated data, using a dataset where only 33% of the pixels in the training set are annotated, o ff ering a huge saving of time, e ff ort and costs for annotators. Zhao et al. ( 2019 ) present an alternative method of data syn- thesis to GANs through the use of learned transformations. From a single manually segmented image, they lev erage other un-annotated images in a SSL like approach to learn a trans- formation model from the images, and use the model along with the labelled data to synthesise additional annotated sam- ples. T ransformations consist of spatial deformations and in- tensity changes to enable to synthesis of complex e ff ects such as anatomical and image acquisition variations. They train a model in a supervised way for the segmentation of MRI brain images and show state-of-the-art improvements over other one- shot bio-medical image segmentation methods. The utility of GAN based approaches in AL scenarios goes beyond single-modality image synthesis. Many works hav e demonstrated the capabilities of GANs to perform cross- modality image synthesis, which directly addresses not only problems of limited training data, but also issues of miss- ing modalities which occur in multi-modal analysis scenarios. Methods by which missing modalities can be generated to fill missing data-points enabling the full suite of AL methods to be applied to multi-modal analysis problems. W ang et al. ( 2019 ) introduce a GAN based method for super- resolution across di ff erent microscopy modalities. This work uses GANs to transform di ff raction limited input images into super-resolv ed ones, impro ving the resolution of wide-field im- ages acquired using low-numerical-aperture objectives to match the resolution acquired using high-numerical-aperture objec- tiv es. This work extends this approach to demonstrate cross- modality super-resolution to transform confocal microscop y images to the resolution acquired with a stimulated emission depletion microscope. This approach enables man y types of images acquired at lower resolutions to be super-resolved to match those of higher resolutions, enable greater performance of multi-modal image analysis methods in both AL and be yond. W ang et al. ( 2018 ) introduce a GAN based method for the generation of high-quality PET images which usually require a full dose radioacti ve tracer to obtain. This w ork enables a low dose tracer to be used to obtain a low-quality PET im- ages, from which a high quality PET image can be generated using a 3D conditional GAN, conditioned on the low-dose im- age. Additional to this, a 3D c-GANs based progressi ve re- finement scheme is introduced to further improve the quality of estimated images. Through this work the dose of radioactiv e tracer required to acquire high-quality PET images is greatly re- 6 duced, reducing the hazards to patients and enabling low-dose PET images to be used alongside high-dose images in do wn- stream analysis. Y u et al. ( 2019 ) extend existing GAN based methods for improv ed cross-modality synthesis of MR images acquired under di ff erent scanning parameters. Their work introduces edge-aware generativ e adversarial networks (Ea-GANs), which specifically integrate edge information reflecting the textural structure of image content to depict the boundaries of di ff er- ent objects in images, which goes beyond methods which focus only on minimising pixel.vox el-wise intensity di ff erences. Us- ing tw o learning strate gies they introduce edge information to a generator-induced Ea-GAN (gEa-GAN) and to a discriminator- induced Ea-GAN (dEa-GAN), incorporating edge information via the generator and both generator and discriminator respec- tiv ely , so that the edge similarity is also adversarially learned. Their method demonstrates state-of-the-art performance for cross-modal MR synthesis as well as excellent generality to generic image synthesis tasks on facades, maps and cityscapes. Pan et al. ( 2020 ) explore the use of GANs to impute missing PET images from corresponding MR images for brain disease identification using a GAN based approach, to av oid discarding data-missing subjects, thus increasing the number of training samples av ailable. A hybrid GAN is used to generate the miss- ing PET images, after which a spatially-constrained Fisher rep- resentation network is used to extract statistical descriptors of neuroimages for disease diagnosis. Results on three databases show this method can synthesise reasonable neuroimages and achiev e promising results in brain disease identification in com- parison to other state-of-the-art methods. The abo ve works demonstrate the po wer of using synthetic data conditioned on a very small amount of annotated data to generate ne w training samples that can be used to train a model to a high accuracy , this is of great value to AL methods where we usually require a initial training set to train a model on be- fore we can employ a data selection policy . These methods also demonstrate the e ffi cient use of labelled data and allow us to generate multiple training samples from a indi vidually an- notated image, this may allow the annotated data obtained in AL / Human-in-the-Loop methods to be used more e ff ectiv ely through generating multiple training samples for a single re- quested annotation, further reducing the annotation e ff ort re- quired to train state-of-the-art models. 2.2.4. Learning Active Learning The majority of methods discussed so far employ hand de- signed heuristics of informati veness, but some w orks hav e emerged that attempt to learn what the most informative sam- ples are through experience of pre vious sample selection out- comes. This o ff ers a potential way to select samples more e ffi ciently but at the cost of interpretability of the heuristics employed. Many factors influence the performance and op- timality of using hand-crafted heuristics for data selection. K onyushko va et al. ( 2017 ) propose ’Learning Acti ve Learning’, where a regression model learns data selection strategies based on e xperience from previous AL outcomes. Arguing there is no way to foresee the influence of all factors such as class imbal- ance, label noise, outliers and distribution shape. Instead, their regression model ’adapts’ its selection to the problem without explicitly stating specific rules. Bachman et al. ( 2017 ) take this idea a step further and propose a model that leverages labelled instances from di ff erent b ut related tasks to learn a selection strategy , while simultaneously adapting its representation of the data and its prediction function. Reinforcement learning (RL) is a branch of ML that enables an ’agent’ to learn in an interactiv e en vironment, by trial and er- ror , using feedback from its own actions and experiences, work- ing to wards achieving the defined goal of the system. Active Learning has recently been suggested as a potential use-case of RL and sev eral works ha ve begun to e xplore this area. W oodward et al. ( 2017 ) propose a one-shot learning method that combines with RL to allow the model to decide, during inference, which examples are worth labelling. A stream of im- ages is presented and a decision is made either to predict the la- bel, or pay to receive the the correct label. Through the choice of RL re ward function they are able to achie ve higher predic- tion accuracy than a purely supervised task, or trade prediction accuracy for fe wer label requests. Fang et al. ( 2017 ) re-frame the data selection process as a RL problem, and e xplicitly learn a data selection polic y . This is agnostic to the data selection heuristics common in AL frame- works, providing a more general approach, demonstrating im- prov ements in entity recognition, howe ver this is yet to be ap- plied to medical image data. RL methods o ff er a di ff erent approach to AL and Human- in-the-Loop problems that is well aligned with aiding real-time feedback between a DL enabled application and its end users, howe ver it requires task specific goals that may not be general- isable across di ff erent medical image analysis tasks. 2.3. Fine-tuning vs Retr aining The final step of each AL framework is to use ne wly acquired annotations to impro ve a model. T wo main approaches are used to train a model on new annotations. These are retraining the model using all a vailable data including the ne wly acquired an- notations or to fine-tune the model using only new annotations or the new annotations plus a subset from the existing annota- tions. T ajbakhsh et al. ( 2016 ) in vestigate using transfer learning and fine-tuning in se veral medical image analysis tasks and demon- strate that the use of a pre-trained CNN with fine-tuning outper- formed a CNN trained from scratch and that these fine-tuned CNNs were more robust to the size of the training sets. The y also sho wed that neither shallow nor deep tuning was the opti- mal choice for a particular application and present a layer-wise training scheme that could o ff er a practical way to reach opti- mal performance for the chosen task based on the amount of data a vailable. The methods emplo yed in this work perform one-time fine-tuning where a pre-trained model is fine-tuned just once with av ailable training samples, ho wev er this does not accommodate an activ e selection process or continuous fine- tuning. Zhou et al. ( 2017 ) propose a continuous fine-tuning method that fine-tunes a pre-trained CNN with successi vely larger 7 datasets and demonstrate that this approach con verges faster than repeatedly fine-tuning the pre-trained CNN. They also find that continuously fine-tuning with only ne wly acquired anno- tations requires careful meta-parameter adjustments making it less practical across many di ff erent tasks. An alternati ve approach to retraining from new data that is inspired by the two main approaches described above is to re- train a model using all available data, but using the previous parameters as initialisation, howe ver this approach has not been applied to AL in any works the authors are a ware of. Retraining is computationally more expensiv e than fine- tuning b ut it pro vides a consistent means to e valuate AL frame- work performance. Fine-tuning is used across a number of dif- ferent ML areas such as one or few shot learning, and transfer learning and the best approach to this is still an open question and as such is less prev alent in AL frameworks, as fine tuning improv es we may see a shift to wards its use in AL frame works. It is important to establish baseline fine-tuning and retraining schemes to e ff ectiv ely compare the DL / AL methods in which they are applied in order to isolate the e ff ects of these schemes from the improv ements made in other areas. 3. The Final Per cent: Interactive refinement of model out- puts So far we have considered the role of humans in annotating data to be used to train a model, b ut once a model is trained, we still require a human-in-the-loop to interpret model predictions and potentially to refine them to acquire the most accurate re- sults for unseen data, as outlined in Figure 2 . In Human-in-the- loop scenarios, a model makes predictions on unseen input, and subject to acceptance criteria, automated predictions may need manual adjustment to meet those acceptance criteria. Commu- nication of information about the prediction is important to al- low acceptance criteria to be met with confidence, and form an understanding of the limitations of automated predictions. This communication is two fold i.e. a user must be able to commu- nicate with the model being used to guide predictions to more accurate results or to correct erroneous predictions, and a model must be able to communicate with the user to provide meaning- ful interpretation of model predictions, enabling users to take the best course of action when interacting with model outputs and to mitigate human uncertainty . This creates the feedback loop as shown in Figure 2 . 3.1. Interactive Refinement If we can dev elop accurate, robust and interpretable mod- els for medical image applications we still cannot guarantee automated predictions meet acceptance criteria for ev ery un- seen data-point presented to a model. The ability to generalise to unseen input is a cornerstone of deep learning applications, but in real world distributions, generalisation is rarely perfect. As such, methods to rectify these discrepancies must be built into applications used for medical image analysis. This itera- tiv e refinement must save the end user time and mental e ff ort ov er performing manual annotation or purely manual correc- tion. Many interactiv e image segmentation systems have been proposed, and more recently these hav e b uilt on the advances in deep learning to allow users to refine model outputs and feed- back the more accurate results to the model for improv ement. Amrehn et al. ( 2017 ) introduced UI-Net, that builds on the popular U-Net architecture for medical image segmentation Ronneberger et al. ( 2015 ). The UI-Net is trained with an ac- tive user model , and allows for users to interact with proposed segmentations by providing scribbles o ver the image to indicate areas that should be included or not, the network is trained us- ing simulated user interactions and as such responds to iterati ve user scribbles to refine a se gmentation towards a more accurate result. Conditional Random fields hav e been used in various tasks to encourage segmentation homogeneity . Zheng et al. ( 2015 ) propose CRF-CNN, a recurrent neural network which has the desirable properties of both CNNs and CRFs. W ang et al. ( 2019 ) propose DeepIGeoS, an interactiv e geodesic framew ork for medical image segmentation. This framework uses two CNNs, the first performs an initial automatic segmentation, and the second takes the initial segmentation as well as user inter- actions with the initial segmentation to provide a refined result. They combine user interactions with CNNs through geodesic distance transforms Criminisi et al. ( 2008 ), and these user in- teractions are integrated as hard constraints into a Conditional Random Field, inspired by Zheng et al. ( 2015 ). The y call their two networks P-Net (initial segmentation) and R-Net (for re- finement). They demonstrate superior results for segmentation of the placenta from 2D fetal MRI and brain tumors from 3D FLAIR images when compared to fully automatic CNNs. These segmentation results were also obtained in roughly a third of the time taken to perform the same segmentation with traditional interactiv e methods such as GeoS or ITK-SN AP . Graph Cuts ha ve also been used in se gmentation to incor- porate user interaction - a user provides seed points to the al- gorithm (e.g. mark some pixel as foreground, and another as background) and from this the segmentation is calculated. W ang et al. ( 2018 ) propose BIFSe g, an interactiv e segmenta- tion frame work inspired by graph cuts. Their work introduces a deep learning framew ork for interactive segmentation by com- bining CNNs with a bounding box and scribble based segmen- tation pipeline. The user provides a bounding box around the area which they are interested in se gmenting, this is then fed into their CNN to produce an initial segmentation prediction, the user can then provide scribbles to mark areas of the image as mis-classified - these user inputs are then weighted heavily in the calculation of the refined segmentation using their graph cut based algorithm. Bredell et al. ( 2018 ) propose an alternativ e to BIFSeg in which two networks are trained, one to perform an initial seg- mentation (they use a CNN but this initial segmentation could be performed with any existing algorithm) and a second net- work they call interCNN that takes as input the image, some user scribbles and the initial segmentation prediction and out- puts a refined segmentation, they show that with several itera- tions over multiple user inputs the quality of the segmentations improv e over the initial segmentation and achieve state-of-the- art performance in comparison to other interactiv e methods. 8 Refiners Predict Feedback to Model Refine Present Intermediate Prediction Accurate Prediction Unseen Data T rained Model Fig. 2. Over view of Refinement frameworks. The methods discussed above ha ve so far been concerned with producing segmentations for individual images or slices, howe ver many segmentation tasks seek to extract the 3D shape / surface of a particular region of interest (ROI). Kurzen- dorfer et al. ( 2017 ) propose a dual method for producing seg- mentations in 3D based on a Smart-brush 2D se gmentation that the user guides to wards a good 2D segmentation, and after a few slices are segmented this is transformed to a 3D surface shape using Hermite radial basis functions, achieving high ac- curacy . While this method does not use deep learning it is a strong example of the ways in which interactive segmentation can be used to generate high quality training data for use in deep learning applications - their approach is general and can produce segmentations for a large number of tasks. There is potential to incorporate deep learning into their pipeline to im- prov e results and accelerate the interactiv e annotation process. Jang and Kim ( 2019 ) propose an interactiv e segmentation scheme that generalises to any previously trained segmenta- tion model, which accepts user annotations about a target ob- ject and the background. User annotations are con verted into interaction maps by measuring the distance of each pixel to the annotated landmarks, after which the forward pass out- puts an initial segmentation. The user annotated points can be mis-segmented in the initial segmentation so they propose BRS (back-propogating refinement scheme) that corrects the mis-labelled pixels. They demonstrate that their algorithm out- performs conv entional approaches on sev eral datasets and that BRS can generalise to medical image segmentation tasks by transforming existing CNNs into user -interactiv e versions. Liao et al. ( 2020 ) propose modelling the dynamics of it- erativ e interactive refinement as a Markov Decision Process (MDP) and solve this with multi-agent RL. T reating each v oxel as an agent with a shared vox el-lev el behaviour strategy they make vox el-wise prediction tractable in this w ay . The multi- agent method successfully captures the dependencies among vox els for se gmentation tasks, and by passing prediction un- certainty of pre vious segmentations through the state space can deriv e more precise and finer segmentations. Using this method they significantly outperform existing state-of-the-art methods with fewer interactions and a f aster con vergence. In this section we focus on applications concerned with it- erativ ely refining a segmentation towards a desired quality of output. In the scenarios above this is performed on an un-seen image provided by the end user , but there is no reason the same approach could not be taken to generate iterativ ely more accu- rate annotations to be used in training, e.g., using active learning to select which samples to annotate next, and iterativ ely refin- ing the prediction made by the current model until a su ffi ciently accurate annotation is curated. This has the potential to accel- erate annotation for training without any additional implemen- tation o verhead. Much w ork done in AL ignores the role of the oracle and merely assumes we can acquire an accurate label when we need it, but in practice this presents a more significant challenge. W e foresee AL and HITL computing become more tightly coupled as AL research improves it’ s consideration for the oracle providing the annotations. It is fairly intuitiv e how a user might refine segmentations of medical images, but this is not the case for other medical im- age analysis tasks. Refinements of predictions on clinical tasks in volving classification and regression have seen less dev elop- ment than those in segmentation and remains an open area of research. The following works ha ve taken steps to wards ad- dressing interacti ve refinement strategies for classification and regression tasks. Lian et al. ( 2020 ) explore the use of CNN methods for auto- mated diagnosis of Alzheimer’ s disease and identify that many state-of-the-art methods rely on the pre-determination of infor- mativ e locations in structural MRI (sMRI). This stage of dis- criminativ e localisation is isolated from the latter stages of fea- ture e xtraction and classifier construction. Their work proposes a hierarchical fully con volutional CNN (H-FCN) to automati- cally identify discriminati ve local patches and regions in whole brain sMRI, from which multi-scale feature representations can be jointly learned and fused to construct classification models. This work enables interacti ve refinement of patch choice and classifier construction which, if interv ened on by human end users could guide the network to wards more discriminativ e re- gions of interest and thus more e ff ectiv e classifiers. Similarly , Liu et al. ( 2018 ) introduce a landmark-based deep multi-instance learning (LDMIL) framew ork for brain disease diagnosis. Firstly , by adopting a data-driv en approach to dis- cov er disease related anatomical landmarks in brain MR im- ages, along with nearby image patches. Secondly the frame- work learns an end-to-end MR image classifier for capturing local structural information in the selected landmark patches, and global structure information deri ved from all detected land- marks. By splitting the steps of landmark detection and classi- fier construction, a human-in-the-loop can be introduced to in- tervene on selected landmarks and to guide the network tow ards maximally informativ e image regions. Thus, the resulting clas- sifier can be refined via updating which regions of the image are used as input. 9 3.2. Interactive Interpr etation In the previous section we discussed methods by which the user of a human-in-the-loop system might communicate with a predictiv e model, in this section we consider methods by which a model might communicate with the user , thus completing the feedback loop in Figure 2 . ’Interpretation’ can mean many dif- ferent things depending on the conte xt, so here we focus on interpretation of model outputs with the goal of appropriately weighting automated predictions in downstream analysis (e.g uncertainty of predictions) and to enable users to mak e the most informed corrections or manual adjustments to model predic- tions (e.g ’Attention Gating’ Oktay et al. ( 2018 )). While DL methods hav e become a standard state-of-the-art approach for many medical image analysis tasks, the y largely remain black-box methods where the end user has limited meaningful ways of interpreting model predictions. This fea- ture of DL methods is a significant hurdle in the deployment of DL enabled applications to safety-critical domains such as medical image analysis. W e w ant models to be highly accu- rate and robust, but also explainable and interpretable. This interpretability is vital to mitigate human uncertainty and foster trust in using automated predictions in downstream tasks with real-world consequences. Recent EU law 1 has led to the ’ right for explanation’, whereby any subject has the right to have automated decisions that have been made about them explained. This even fur- ther highlights the need for transparent algorithms which we can reason about [ Goodman and Flaxman ( 2017 ), Edwards and V eale ( 2017a ), Edwards and V eale ( 2017b )]. It is important for users to understand ho w a certain decision has been made by the model, as even the most accurate and ro- bust models aren’t infallible, and false or uncertain predictions must be identified so that trust in the model can be fostered and predictions are appropriately weighted in the clinical decision making process. It is vital the end user , regulators and auditors all ha ve the ability to contextualise automated decisions pro- duced by DL models. Here we outline some di ff erent methods for providing interpretable ways of reasoning about DL models and their predictions. T ypically DL methods can provide statistical metrics on the uncertainty of a model output, many of the uncertainty mea- sures discussed in Section 2 are also used to aid in intepretabil- ity . While uncertainty measures are important, these are not suf- ficient to foster complete trust in DL model, the model should provide human-understandable justifications for its output that allow insights to be drawn elucidating the inner workings of a model. Chakraborty et al. ( 2017 ) discuss many of the core con- cerns surrounding model intepretability and highlight various works that hav e demonstrated sophisticated methods of making a DL model interpretable across the DL field. Here we ev alu- ate some of the works that hav e been applied to medical image segmentation and refer the reader to [ Stoyanov et al. ( 2018 ), 1 Regulation (EU) 2016 / 679 on the protection of natural persons with reg ard to the processing of personal data and on the free movement of such data, and repealing Directi ve 95 / 46 / EC (General Data Protection Regulation) [2016] OJ L119 / 1 Holzinger et al. ( 2017 )] for further reading on interpretability in the rest of the medical imaging domain. Oktay et al. ( 2018 ) and Schlemper et al. ( 2019 ) introduce ’Attention Gating’ to guide networks towards giving more ’at- tention’ to certain image areas, in a visually interpretable way - potentially aiding in the subsequent refinement of annotations. Attention Gates are introduced into the popular U-Net architec- ture ( Ronneberger et al. ( 2015 )), where information extracted from coarse scale layers is used in gating to disambiguate ir- relev ant and noisy responses in skip connections, prior to con- catenation, to merge only relev ant layer activ ations. This ap- proach eliminates the need for applying external object locali- sation models in image segmentation and regression tasks. Co- e ffi cients of Attention Gate layers indicate where in an image feature acti vations will be allo wed to propagate through to final predictions, providing users with a visual representation of the areas of an image that a model has weighted highly in making predictions. In Budd et al. ( 2019 ) we propose a visual method for in- terpreting automated head circumference measurements from ultrasound images, using MC Dropout at test-time to acquire N head segmentations to calculate an upper and lower bound on the head circumference measurement in real-time. These bounds were displayed over the image to guide the sonogra- pher towards views in which the model predicts with the most confidence. This upper lower bound is presented as a measure of model compliance of the unseen image rather than uncer- tainty . Finally , variance heuristics are proposed to quantify the confidence of a prediction in order to either accept or reject head circumference measurements, and it is sho wn these can improv e overall performance measures once ’ rejected’ images are remov ed. Milletari et al. ( 2019 ) propose the application of RL to ul- trasound care, guiding a potentially inexperienced user to the correct sonic window and enabling them to obtain clinically relev ant images of the anatomy of interest. This human-in-the- loop application is an example of the novel applications pos- sible when combining DL / RL with real-time systems enabling users to respond to model feedback to acquire the most accurate information av ailable. W ang et al. ( 2019b ) propose using test-time augmentation to acquire a measure of aleatoric (image-based) uncertainty and compare their method with epistemic (model) uncertainty mea- sures and show that their method provides a better uncertainty estimation than a test-time dropout based model uncertainty alone and reduces ov erconfident incorrect predictions. Jungo and Reyes ( 2019 ) ev aluate sev eral di ff erent voxel- wise uncertainty estimation methods applied to medical im- age segmentation with respect to their reliability and limitations and show that current uncertainty estimation methods perform similarly . Their results show that while uncertainty estimates may be well calibrated at the dataset le vel (capturing epistemic uncertainty), they tend to be mis-calibrated at a subject-lev el (aleatoric uncertainty). This compromises the reliability of these uncertainty estimates and highlights the need to de velop subject-wise uncertainty estimates. They show auxiliary net- works to be a valid alternati ve to common uncertainty methods 10 as they can be applied to any previously trained segmentation model. Dev eloping transparent systems will enable faster uptake in clinical practice and including humans within the deep learn- ing clinical pipelines will ease the period of transition between current best practices and the breadth of possible enhancements that deep learning has to o ff er . W e suggest that ongoing work in improving interpretability of DL models will also ha ve a positi ve impact on AL, as the majority of methods to improv e intepretability are centred on providing uncertainty measures for a models prediction, these same uncertainty measures can be used for AL selection strate- gies in place of existing uncertainty measures that are currently employed. As intepretability and uncertainty measures improv e we expect to see a similar improvement of AL frame works as they incorporate the most promising uncertainty measures. The methods discussed in Section 3 remain open areas of re- search interest with great implications for the progress of AL dev elopment and greater uptake of DL and HITL methods in clinical practice. The study of interaction between users and models is of growing importance and is having a significant impact on the e ffi cacy of Deep Acti ve Learning systems and their deployment to real-world applications, especially in clin- ical scenarios ( Beede et al. ( 2020 ); Amrehn et al. ( 2019 )). The wider study of interpretability in ML and the study of Human Computer Interaction may seem distinct and di verging, ho w- ev er we expect to see these two research fields con ver ge through Activ e Learning as the feedback loop between human users and machine models becomes of increasing importance. 4. Practical Considerations W e have so far discussed the core body of work behind AL, model interpretation and prediction refinement, and while the works discussed above go a long way in covering the major- ity of research being done, there are several practical consid- erations for dev eloping and deploying DL enabled applications that we must consider . In this section we outline the main prac- tical research areas that are impacting DL enabled application dev elopment pipelines and suggest where we might look next. 4.1. Noisy Oracles Gold-standard annotations for medical image data are ac- quired by aggre gating annotations from multiple expert oracles, but as previously discussed, this is rarely feasible to obtain for large comple x datasets due to the expertise required to perform such annotations. Here we ask what e ff ect on performance we might incur if we acquire labels from oracles without domain expertise, and what techniques can we use to mitigate the sus- pected degradation of annotation quality when using non-expert oracles, to av oid any potential loss in accurac y . Zhang and Chaudhuri ( 2015 ) propose acti ve learning method that assume data will be annotated by a crowd of non-e xpert or ’weak’ annotators, and o ff er approaches to mitigate the in- troduction of bad labels into the data set. They simultaneously learn about the quality of individual annotators so that the most informativ e examples can be labelled by the strongest annota- tors. Li et al. ( 2019 ) propose methods for crowd-sourced learning in two scenarios. Firstly , they aim at inferring instances ground truth gi ven the crowd’ s annotations by modelling the crowd’ s expertise and label correlations from two di ff erent perspecti ves: firstly they model expertise based on indi vidual labels, based on the idea that labeller’ s annotations for similar instances should be similar , and secondly through modelling the crowd’ s exper- tise to distinguish the relev ance between label pairs. They ex- tend their approach to the activ e paradigm and o ff er criteria for instance, label and labeller selected in tandem to minimise an- notation cost. Cheplygina et al. ( 2016 ) explore using Amazon’ s MT urk to gather annotations of airways in CT images. Results showed that the novice oracles were able to interpret the images, but that instructions provided were too complex, leading to many unusable annotations. Once the bad annotations were remo ved, the annotations did show medium to high correlation with ex- pert annotations, especially if annotations were aggregated. Rodrigues and Pereira ( 2018 ) describe an approach to assess the reliability of annotators in a cro wd, and a crowd layer used to train deep models from noisy labels from multiple annota- tors, internally capturing the reliability and biases of di ff erent annotators to achie ve state-of-the-art results for several crowd- sourced data-set tasks. W e can see that by using a learned model of oracle annota- tion quality we can mitigate the e ff ects of low quality annota- tions and present the most challenging cases to most capable oracles. By providing clear instructions we can lower the bar- riers for non-expert oracles to perform accurate annotation, but this is not generalisable and would be required for every new annotation task we wish to perform. 4.2. W eakly Supervised Learning Most segmentation tasks require pixel-wise annotations, but these are not the only type of annotation we can give an image. Segmentation can be performed with ’weak’ annotations, which include image le vel labels e.g. modality , organs present etc. and annotations such as bounding boxes, ellipses or scribbles. It is argued that using ’weak er’ annotation formulations can make the task easier for the human oracle, leading to more accurate annotations. ’W eak’ annotations have been shown to perform well in sev eral segmentation tasks, Rajchl et al. ( 2016a ) demon- strate obtaining pixel-wise segmentations gi ven a data-set of images with ’weak’ bounding box annotations. The y propose DeepCut, an architecture that combines a CNN with an iterative dense CRF formulation to achiev e good accuracy while greatly reducing annotation e ff ort required. In a later study , Rajchl et al. ( 2017 ) examine the impact of expertise required for di ff erent ’weak’ annotation types on the accuracy of liver segmentations. The results showed a decrease in accuracy with less expertise, as expected, across all annotation types. Despite this, segmen- tation accurac y was comparable to state-of-the-art performance when using a weakly labelled atlas for outlier correction. The robust performance of their approach suggests ’weak’ annota- tions from non-expert crowds could be used to obtain accurate 11 User Qulaity Scores Weak Labels? Strong Labels? Single T ask/ Multi T ask? Cost Assessment Aggregated Label(s) Unlabelled Data Noisy Oracles Interface Fig. 3. Over view of practical considerations segmentations on many di ff erent tasks, howe ver their use of an atlas makes this approach less generalisable than is desired. In Rajchl et al. ( 2016b ) they examine using super pixels to accelerate the annotation process. This approach uses a pre-processing step to acquire a super-pix el segmentation of each image, non-experts are then used to perform the an- notation by selecting which super-pixels are part of the tar- get region. Results showed that the approach largely reduces the annotation load on users. Non-expert annotation of 5000 slices was completed in under an hour by 12 annotators, com- pared to an expert taking three working days to establish the same with an advanced interface. The non-expert interface is web-based demonstrating the potential of distributed annotation collection / crowd-sourcing. An encouraging aspect of this paper is that the results sho wed high performance on the segmentation task in question compared with e xpert annotation performance, but may not be suitable for all medical image analysis tasks. It has been shown that we can de velop high performing mod- els using weakly annotated data, and as weak annotations re- quires less expertise to perform, the y can be acquired faster and from a non-expert crowd with a smaller loss in accuracy than gold-standard annotations. This is v ery promising for future research as datasets of weakly annotated data might be much easier and more cost-e ff ectiv e to curate. 4.3. Multi-task learning Many works aim to train models or acquire training data for sev eral tasks at once, it is argued that this can sav e on cost as complementary information may result in higher performance ov er multiple di ff erent tasks ( Moeskops et al. , 2016 ). W ang et al. ( 2019a ) propose a dual network for joint se gmentation and detection task for lung nodule segmentation and cochlea segmentation from CT images, where only a part of the data is densely annotated and the rest is weakly labelled by bounding boxes, using this the y show that their architecture out-performs sev eral baselines. At present this w ork only handles the case for two di ff erent label types but they propose extending the frame- work for a true multi-task scenario. This is a promising area but, as of yet, it has not been in- corporated into an active learning setting. As such, it may be elucidating to analyse the di ff erences in samples chosen by dif- ferent AL methods when the model is being training for mul- tiple tasks simultaneously . Ho wever , Lowell et al. ( 2019 ) raise concerns over the transferability of acti vely acquired datasets to future models due to the inherent coupling between activ e learning selection strategies and the model being trained, and show that training a successor model on the actively acquired dataset can often result in worse performance than from ran- dom sampling. The y suggest that, as datasets begin to outliv e the models trained on them, there is a concern for the e ffi cacy of acti ve learning, since the acquired dataset may be disadvan- tageous for training subsequent models. An exploration of how activ ely acquired datasets perform on multiple models may be required to e xplain the e ff ects of an acti vely acquired dataset coupled with one model on the performance of related models. 4.4. Annotation Interface So far the majority of Human-in-the-loop methods assume a significant lev el of interaction from an oracle to annotate data and model predictions, but fe w consider the nature of the in- terface with which an oracle might interact with these images. The nature of medical images require special attention when proposing distributed online platforms to perform such annota- tions. While the majority of techniques discussed so far hav e used pre-existing data labels in place of newly acquired labels to demonstrate their performance, it is important to consider the e ff ects of accuracy of annotation that the actual interface might incur . Nalisnik et al. ( 2015 ) propose a framework for the online classification of Whole-slide images (WSIs) of tissues. Their interface enables users to rapidly build classifiers using an activ e learning process that minimises labelling e ff orts and demonstrates the e ff ectiveness of their solution for the quan- tification of glioma brain tumours. Khosrav an et al. ( 2017 ) propose a no vel interface for the se g- mentation of images that tracks the users gaze to initiate seed points for the segmentation of the object of interest as the only means of interaction with the image, achieving high segmen- tation performance. Stember et al. ( 2019 ) extend this idea and compare using eye tracking generated training samples to tradi- tional hand annotated training samples for training a DL model. They show that almost equiv alent performance was achie ved using annotation generated through e ye tracking, and suggest that this approach might be applicable to rapidly generate train- ing data. They acknowledge that there is still improvements to be made integrate eye tracking into typical clinical radiology work flo w in a faster , more natural and less distracting way . T inati et al. ( 2017 ) e valuate the player motiv ations behind EyeW ire, an online game that asks a crowd of players to help segment neurons in a mouse brain. The gamification of this 12 task has seen ov er 500,000 players sign up and the segmenta- tions acquired ha ve gone onto be used in sev eral research w orks [ Kim et al. ( 2014 )]. One of the most exciting things about gam- ification is that when surveyed, users were moti vated most by making a scientific contribution rather than any potential mon- etary re ward. Howe ver this is very specialised towards this par- ticular task and would be di ffi cult to apply across other types of medical image analysis tasks. There are many di ff erent approaches to dev eloping annota- tion interfaces and the ones we consider above are just a fe w that hav e been applied to medical image analysis. As dev elop- ment increases we expect to see more online tools being used for medical image analysis and the chosen format of the inter- face will play a lar ge part in the usability and o verall success of these applications. 4.5. V ariable Learning Costs When acquiring training data from various types of oracle it is worth considering the relativ e cost associated with querying a particular oracle type for that annotation. W e may wish to acquire more accurate labels from an expert oracle, b ut this is likely more expensi ve to obtain than from a non-expert oracle. The trade o ff , of course, being accuracy of the obtained label - less expertise of the oracle will likely result in a lower quality of annotation. Several methods ha ve been proposed to model this and allow dev elopers to trade o ff between cost and overall accuracy of acquired annotations. Kuo et al. ( 2018 ) propose a cost-sensiti ve acti ve learning ap- proach for intracranial haemorrhage detection. Since annota- tion time may vary significantly across examples, the y model the annotation time and optimize the return on inv estment. They show their approach selects a div erse and meaningful set of samples to be annotated, relati ve to a uniform cost model, which mostly selects samples with massive bleeds which are time con- suming to annotate. Shah et al. ( 2018 ) propose a budget based cost minimisation framew ork in a mixed-supervision setting (strong and weak an- notations) via dense segmentation, bounding boxes, and land- marks. Their frame work uses an uncertainty and a represen- tativ eness ranking strategy to select samples to be annotated next. They demonstrate state-of-the-art performance at a signif- icantly reduced training b udget, highlighting the important role of choice of annotation type on the costs of acquiring training data. The above works each show an improved consideration for the economic burden that is incurred when curating training data. A valuable research direction would be to assess the ef- fects of oracle expertise lev el, annotation type and image an- notation cost in a unified frame work as these three factors are very much linked and may hav e a profound influence over each other . 5. Future Prospectiv e and Unanswered Questions In Sections 2 & 3 we discuss methods through which a user might gather training data to build a model, use their model to predict on new data and receiv e feedback to iterati vely refine the model output towards a more accurate result. Each of these techniques assume some human end user will be present to in- teract with the system at the point of initial annotation, inter- pretation and refinement. Each of these areas seeks to achieve a shared goal of achieving the highest performing model from as little annotated data as possible - with a means to weigh con- clusions of models predictions appropriately . AL is not the only area of research that aim to learn from limited data. Semi-supervised learning, and Transfer Learning both mak e significant contributions to e xtracting the most v alue from limited labelled data. In the presence of large data-sets, but the absence of labels, unsupervised and semi-supervised approaches o ff er a means by which information can be extracted without requiring labels for all the data-points. This could potentially ha ve a massiv e impact on the medical image analysis field where this is often the case. In a semi-supervised learning (SSL) scenario we may have some labelled data, but this is often very limited. W e do how- ev er have a large set of un-annotated instances (much like in activ e learning) to draw information from, the goal being to im- prov e a model (trained only on the labelled instances) using the un-labelled instances. From this we deriv e two distinct goals: a) predicting labels for future data (inductive SSL) and b) pre- dicting labels for the av ailable un-annotated data (transductive SSL) ( Cheplygina et al. ( 2019 ); Xia et al. ( 2020 )). SSL methods provide a powerful way of extracting useful information from un-annotated image data and we believe that progress in this area will be beneficial to AL systems that desire a more accu- rate model for initialisation to guide data selection strategies. T ransfer Learning (TL) is a branch of DL that aim to use pre- trained netw orks as a starting point for new applications. Giv en a pre-trained network trained for a particular task, it has been shown that this network can be ’fine-tuned’ tow ards a target task from limited training data. W e refer the reader to Morid et al. ( 2021 ); Raghu et al. ( 2019 ); Cheplygina et al. ( 2019 ) for a more general overvie w of transfer learning in medical imag- ing, and focus on the use of TL in AL scenarios in the follow- ing. T ajbakhsh et al. ( 2016 ) demonstrated the applicability of TL for a v ariety of medical image analysis tasks, and show , de- spite the large di ff erences between natural images and medical images, CNNs pre-trained on natural images and fine-tuned on medical images can perform better than medical CNNs trained from scratch. This performance boost was greater where fewer target task training examples were a v ailable. Many of the meth- ods discussed so far start with a network pre-trained on natural image data. Zhou et al. ( 2018a ) propose AFT*, a platform that com- bines AL and TL to reduce annotation e ff orts, which aims at solving sev eral problems within AL. AFT* starts with a com- pletely empty labelled data-set, requiring no seed samples. A pre-trained CNN is used to seek ’w orthy’ samples for anno- tation and to gradually enhance the CNN via continuous fine- tuning. A number of steps are taken to minimise the risk of catastrophic forgetting. Their previous work Zhou et al. ( 2017 ) applies a similar but less featureful approach to se veral medi- cal image analysis tasks to demonstrate equiv alent performance can be reached with a heavily reduced training data-set. The y 13 then use these tasks to ev aluate sev eral patterns of prediction that the network exhibits and how these relate to the choice of AL selection criteria. Zhou et al. ( 2018b ) hav e gone onto to use their AFT frame- work for annotation of CIMT videos, a clinical technique for characterisation of Cardio vascular disease. Their extension into the video domain presents its own unique challenges and thus they propose a new concept of an Annotation Unit - reducing annotating a CIMT video to just 6 user mouse clicks, and by combining this with their AFT frame work reduce annotation cost by 80% relati ve to training from scratch and by 50% rel- ativ e to random selection of new samples to be annotated (and used for fine-tuning). Kushibar et al. ( 2019 ) use TL for supervised domain adapta- tion for sub-cortical brain structure segmentation with minimal user interaction. They significantly reduce the number of train- ing images from di ff erent MRI imaging domains by le veraging a pre-trained network and improve training speed by reducing the number of trainable parameters in the CNN. They show their method achieves similar results to their baseline while using a remarkably small amount of images from the target domain and show that using even one image from the tar get domain was enough to outperform their baseline. The above methods and more discussed in this revie w demonstrate the applicability of TL to reducing the number of annotated sample required to train a model on a new task from limited training data. By using pre-trained networks trained on annotated natural image data (there is an abundance) we can boost model performance and further reduce the annotation ef- fort required to achiev e state-of-the-art performance. A related sub-field of TL worth exploring is domain adapta- tion (DA). Many DL techniques used in medical image analysis su ff er from the domain shift problem caused by di ff erent distri- butions between source data and target data, often due to med- ical images being acquired on a v ariety of di ff erent scanners, scanning parameters and subject cohorts etc. D A has been pro- posed as a special type of transfer learning in which the domain feature space and tasks remain the same while marginal distri- butions of the source and target domains are di ff erent. W e refer the reader to Guan and Liu ( 2021 ); Choudhary et al. ( 2020 ) for an overvie w of D A methods used for medical image analysis, and hope to see greater application of DA methods in AL sce- narios in the future. In many of scenarios described in this re view , models con- tinuously receiv e new annotations to be used for training, and in theory we could continue to retrain or fine-tune a model in- definitely , but is this practical and cost e ff ectiv e? It is important to quantify the long term e ff ects of training a model with new data to assess how the model changes over time and whether or not performance has improv ed, or w orse, declined. Learning from continuous streams of data has proven more di ffi cult than anticipated, often resulting in ’catastrophic forgetting’ or ’in- terference’ Parisi et al. ( 2019 ). W e face the stability-plasticity- dilemma . A voiding catastrophic forgetting in neural networks when learning from continuous streams of data can be broadly divided among three conceptual strategies: a) Retraining the the whole network while regularising (to prev ent forgetting of previously learned tasks). b) selecti vely train the network and expand it if needed to represent new tasks, and c) retaining pre- vious experience to use memory replay to learn in the absence of new input. W e refer the reader to Parisi et al. ( 2019 ) for a more detailed ov erview of these approaches. Baweja et al. ( 2018 ) inv estigate continual learning of two MRI segmentation tasks with neural networks for counter- ing catastrophic forgetting of the first task when a new one is learned. They inv estigate elastic weight consolidation, a method based on Fisher information to sequentially learn seg- mentation of normal brain structures and then segmentation of white matter lesions and demonstrate this method reduces catastrophic forgetting, but acknowledge there is a large room for improvement for the challenging setting of continual learn- ing. It is important to quantify the performance and robustness of a model at ev ery stage of its lifespan. One way to consider stopping could evaluate when the cost of continued training out- weighs the cost of errors made by the current model. An exist- ing measure that attempts to quantify the economical value of medical intervention is the Quality-adjusted Life year (QAL Y), where one QAL Y equates to one year of healthy life NICE ( 2013 ). Could this metric be incorporated into models? At present we cannot quantify the cost of errors made by DL med- ical imaging applications but doing so could lead to a deeper understanding of how accurate a DL model really ought to be. As models are trained on more of the end user’ s o wn data, will this cause the network to perform better on data from that user’ s system despite performing worse on data the model was initially trained on? Catastrophic forgetting suggests this will be the case, but is this a bad thing? It may be beneficial for models to gradually bias themselves tow ards high performance for the end user’ s own data, even if this results in the model becoming less transferable to other data. Farquhar et al. ( 2021 ) explore the role of bias in AL methods. Bias is introduced be- cause the training data no longer follows the population distri- bution in AL. The authors pro viding a general method by which unbiased AL estimators may be constructed using novel correc- tiv e weights to remove bias. Further to this, an explanation of the empirical successes of existing AL methods which ignore this bias is provided. It is shown that bias introduced by AL methods can be actively helpful when training ov erparameter- ized models like neural networks with relatively little data. This further moti vates future work to better understand when the bias introduced by AL could ha ve a positi ve influence on the perfor- mance of AL methods, to the detriment of generalisability to other data sources. Activ e learning assumes the presence of a user interface to perform annotations b ut is only concerned with which data to annotate. Refinement assumes we can generate an annotation through iterati ve interaction with the current model prediction. Hence, it would be desirable to combine these two in future work. If we can train a model with a tiny amount of training data, and then ask annotators to refine model predictions to- wards a more accurate label, we can expedite the annotation process by reducing the initial annotation workload and reduce additional interface work for use with unseen data. This would 14 be the same interface used to create the training annotations. By combining the e ff orts of active learning and iterati ve refinement into a unified frame work we can rapidly produce annotations to train our model, as well as acquiring high quality results from our models from the beginning. This should also hav e the added side e ff ect of training the model on data from the same distribu- tion that it will be predicting on, reducing domain shift e ff ects in unseen distributions. By incorporating our end user at each stage of the model life cycle we could also use human feedback on model performance to add a more ’human interpretable’ metric of model confidence as each user could rank the performance of the model for each input as it sees it, potentially giving a metric of confidence based on human interpretation of the model output. This of course requires e xperts to be using the system.One might argue that the models initial predictions may impart some influence ov er the human user b ut by crowd-sourcing the initial annota- tions to a less expert multi-label cro wd we could reduce this bias. Dev elopments in uncertainty quantification will benefit both AL selection heuristics and interpretation of model outputs, b ut there is no guarantee that the best performing uncertainty met- rics for selecting new samples to be annotated will be the same metrics that are the most interpretable to a human user . Figure 4 outlines the core methods being used in human-in- the-loop computing for each of the papers discussed in this re- view . This figure shows that there is significant overlap of re- search goals for many areas of human-in-the-loop computing but there are large gaps that need to be filled in order to un- derstand the relationships between di ff erent methods and ho w these might a ff ect their performance. As the many areas of DL research conv erge towards shared goals of working with limited training data to achiev e state- of-the-art results, we expect to see more systems emerge that exploit the advances made in the range of sub-fields of ML de- scribed here. W e hav e already seen the combination of se v- eral methods into indi vidual frame works b ut as of yet no w orks combine all of the approaches discussed into a single frame- work. As di ff erent combinations of approaches be gin to ap- pear it is important to consider the measure by which we as- sess their performance, as isolating indi vidual dev elopments be- comes more di ffi cult. Developing baseline human-in-the-loop methods to compare to will be vital to assess the contributions of individual works in each area and to better understand the influences of competing improv ements in these areas. 6. Conclusions In this revie w we hav e explored the large body of emerging medical image analysis work in which a human end user is at the centre. Deep Learning has all the ingredients to induce a paradigm shift in our approach to a plethora of clinical tasks. The direct in volvement of humans is set to play a core role in this shift. The works presented in this revie w each o ff er their own approaches to including humans in the loop and we sug- gest that there is su ffi cient o verlap in many methods for them to be considered under the same title of Human-in-the-Loop com- puting. W e hope to see new methodologies emerge that com- bine the strengths of AL and HITL computing into end-to-end systems for the development of deep learning applications that can be used in clinical practice. While there are some practi- cal limitations as discussed, there are many proposed solutions to such issues and as research in these directions continues it is only a matter of time before deep learning applications blos- som into fully-fledged, accurate and robust systems to be used for daily routine tasks. W e are in an exciting era for medical image analysis, with endless opportunity to innov ate and im- prov e the current state-of-the-art and to leverage the powers of deep learning to make a real impact in health care across the board. With diligent research and development we should see more and more applications boosted by deep learning capabili- ties finding their way onto the mark et, allo wing users to achie ve better results, faster , and with less expertise than before, freeing up expert time to be used on the most challenging cases. The field of Human-in-the-loop computing will play a crucial role to achiev e this. Acknowledgments SB is supported by the EPSRC Centre for Doctoral T rain- ing in Smart Medical Imaging EP / S022104 / 1. This work was in part supported by EP / S013687 / 1, Intel and Nvidia. W e thank Innov ate UK: London Medical Imaging & Artificial In- telligence Centre for V alue-Based Healthcare [104691] for co- funding this research. References Amrehn, M., Gaube, S., Unberath, M., Schebesch, F ., Horz, T ., Strumia, M., Steidl, S., Ko warschik, M., Maier , A., 2017. UI-Net: Interactiv e Artificial Neural Networks for Iterative Image Segmentation Based on a User Model, in: Bruckner, S., Hennemuth, A., Kainz, B., Hotz, I., Merhof, D., Rieder, C. (Eds.), Eurographics W orkshop on V isual Computing for Biology and Medicine, The Eurographics Association. doi: 10 . 2312/vcbm . 20171248 . Amrehn, M., Steidl, S., Kortekaas, R., Strumia, M., W eingarten, M., K owarschik, M., Maier, A., 2019. A semi-automated usability eval- uation framework for interactive image segmentation systems. In- ternational Journal of Biomedical Imaging 2019. URL: https:// pubmed . ncbi . nlm . nih . gov/31582963/ , doi: 10 . 1155/2019/1464592 . Angluin, D., 1988. Queries and Concept Learning. Machine Learning 2, 319–342. URL: https://link . springer . com/article/10 . 1023/A: 1022821128753 , doi: 10 . 1023/A:1022821128753 . Angluin, D., 2001. Queries revisited, in: Lecture Notes in Com- puter Science (including subseries Lecture Notes in Artificial Intelli- gence and Lecture Notes in Bioinformatics), Springer V erlag. pp. 12– 31. URL: https://link . springer . com/chapter/10 . 1007/3- 540- 45583- 3 3 , doi: 10 . 1007/3- 540- 45583- 3{ \ }3 . Atlas, L.E., Cohn, D.A., Ladner , R.E., 1990. T raining connectionist networks with queries and selective sampling, in: T ouretzky , D.S. (Ed.), Advances in Neural Information Processing Systems 2. Mor gan- Kaufmann, pp. 566–573. URL: http://papers . nips . cc/paper/ 261- training- connectionist- networks- with- queries- and- selective- sampling . pdf . Bachman, P ., Sordoni, A., T rischler , A., 2017. Learning Algorithms for Active Learning. T echnical Report. URL: http://proceedings . mlr . press/ v70/bachman17a/bachman17a . pdf . Baweja, C., Glocker , B., Kamnitsas, K., 2018. T ow ards continual learning in medical imaging. T echnical Report. URL: https://www . doc . ic . ac . uk/ ~ bglocker/public/mednips2018/med- nips 2018 paper 82 . pdf . 15 Beede, E., Baylor , E., Hersch, F ., Iurchenko, A., W ilcox, L., Ruamviboon- suk, P ., V ardoulakis, L.M., 2020. A Human-Centered Evaluation of a Deep Learning System Deployed in Clinics for the Detection of Diabetic Retinopathy , in: Proceedings of the 2020 CHI Conference on Human Factors in Computing Systems, Association for Computing Machinery (A CM), New Y ork, NY , USA. pp. 1–12. URL: https://dl . acm . org/doi/10 . 1145/ 3313831 . 3376718 , doi: 10 . 1145/3313831 . 3376718 . Beluch Bcai, W .H., N ¨ urnberger , A., Bcai, J.M.K., 2018. The power of ensembles for active learning in image classification. T echnical Re- port. URL: http://openaccess . thecvf . com/content cvpr 2018/ CameraReady/1487 . pdf . Ben-David, S., Blitzer , J., Crammer, K., Kulesza, A., Pereira, F ., V aughan, J.W ., 2010. A theory of learning from di ff erent domains. Machine learning 79, 151–175. Bredell, G., T anner, C., Konukoglu, E., 2018. Iterative interaction training for segmentation editing networks, in: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), Springer V erlag. pp. 363–370. URL: https:// doi . org/10 . 1007/978- 3- 030- 00919- 9 42 , doi: 10 . 1007/978- 3- 030- 00919- 9{ \ }42 . Budd, S., Sinclair , M., Khanal, B., Matthew , J., Lloyd, D., Gomez, A., T ous- saint, N., Robinson, E.C., Kainz, B., 2019. Confident head circumference measurement from ultrasound with real-time feedback for sonographers, in: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), Springer . pp. 683–691. URL: https://link . springer . com/chapter/10 . 1007/978- 3- 030- 32251- 9 75 , doi: 10 . 1007/978- 3- 030- 32251- 9{ \ }75 . Chakraborty , S., T omsett, R., Raghav endra, R., Harborne, D., Alzan- tot, M., Cerutti, F ., Srivasta va, M., Preece, A., Julier, S., Rao, R.M., Kelle y , T .D., Braines, D., Sensoy , M., Willis, C.J., Gurram, P ., 2017. Interpretability of deep learning models: A survey of re- sults, in: 2017 IEEE SmartW orld, Ubiquitous Intelligence & Comput- ing, Adv anced & T rusted Computed, Scalable Computing & Communi- cations, Cloud & Big Data Computing, Internet of People and Smart City Innov ation (SmartW orld / SCALCOM / UIC / A TC / CBDCom / IOP / SCI), IEEE. pp. 1–6. URL: https://ieeexplore . ieee . org/document/8397411/ , doi: 10 . 1109/UIC- ATC . 2017 . 8397411 . Cheplygina, V ., de Bruijne, M., Pluim, J.P ., 2019. Not-so-supervised: A survey of semi-supervised, multi-instance, and transfer learning in medi- cal image analysis. Medical Image Analysis 54, 280–296. doi: 10 . 1016/ j . media . 2019 . 03 . 009 . Cheplygina, V ., Perez-Rovira, A., Kuo, W ., Tiddens, H.A., de Bruijne, M., 2016. Early experiences with crowdsourcing airway annotations in chest CT, in: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), Springer V erlag. pp. 209–218. URL: https://link . springer . com/chapter/ 10 . 1007/978- 3- 319- 46976- 8 22 , doi: 10 . 1007/978- 3- 319- 46976- 8{ \ }22 . Choudhary , A., T ong, L., Zhu, Y ., W ang, M.D., 2020. Adv anc- ing Medical Imaging Informatics by Deep Learning-Based Do- main Adaptation. Y earbook of medical informatics 29, 129– 138. URL: /pmc/articles/PMC7442502//pmc/articles/ PMC7442502/?report=abstracthttps://www . ncbi . nlm . nih . gov/ pmc/articles/PMC7442502/ , doi: 10 . 1055/s- 0040- 1702009 . Cohn, D., Atlas, L., Ladner , R., 1994. Improving generalization with activ e learning. Machine Learning 15, 201–221. URL: https: //link . springer . com/article/10 . 1007/BF00993277 , doi: 10 . 1007/ bf00993277 . Criminisi, A., Sharp, T ., Blake, A., 2008. GeoS: Geodesic Image Seg- mentation, Springer , Berlin, Heidelberg, pp. 99–112. URL: http: //link . springer . com/10 . 1007/978- 3- 540- 88682- 2 9 , doi: 10 . 1007/ 978- 3- 540- 88682- 2{ \ }9 . Dagan, I., Engelson, S.P ., 1995. Committee-Based Sampling For Training Prob- abilistic Classifiers, in: Machine Learning Proceedings 1995. Elsevier , pp. 150–157. doi: 10 . 1016/b978- 1- 55860- 377- 6 . 50027- x . Edwards, L., V eale, M., 2017a. Enslaving the Algorithm: From a Right to an Explanationn to a Right to Better Decisionss? SSRN Electronic Jour- nal URL: https://www . ssrn . com/abstract=3052831 , doi: 10 . 2139/ ssrn . 3052831 . Edwards, L., V eale, M., 2017b . Slave to the Algorithm? Why a Right to Explanationn is Probably Not the Remedy Y ou are Looking for. SSRN Electronic Journal URL: https://www . ssrn . com/abstract=2972855 , doi: 10 . 2139/ssrn . 2972855 . Fang, M., Li, Y ., Cohn, T ., 2017. Learning how to acti ve learn: A deep reinforcement learning approach, in: Proceedings of the 2017 Confer- ence on Empirical Methods in Natural Language Processing, Associa- tion for Computational Linguistics, Copenhagen, Denmark. pp. 595–605. URL: https://www . aclweb . org/anthology/D17- 1063 , doi: 10 . 18653/ v1/D17- 1063 . Farquhar , S., Gal, Y ., Rainforth, T ., 2021. On statistical bias in active learning: How and when to fix it, in: International Conference on Learning Represen- tations. URL: https://openreview . net/forum?id=JiYq3eqTKY . Gal, Y ., Ghahramani, Z., 2016. Dropout as a Bayesian approximation: Rep- resenting model uncertainty in deep learning, in: Proceedings of the 33rd International Conference on Machine Learning (ICML-16). Gal, Y ., Islam, R., Ghahramani, Z., 2017. Deep bayesian active learning with image data, in: Proceedings of the 34th International Conference on Ma- chine Learning - V olume 70, JMLR.org. p. 1183–1192. Goodman, B., Flaxman, S., 2017. European union regulations on algo- rithmic decision-making and a “right to explanation”. AI Magazine 38, 50–57. URL: https://www . aaai . org/ojs/index . php/aimagazine/ article/view/2741 , doi: 10 . 1609/aimag . v38i3 . 2741 . Guan, H., Liu, M., 2021. Domain Adaptation for Medical Image Analysis: A Survey URL: http://arxiv . org/abs/2102 . 09508 . Haenssle, H.A., Fink, C., Schneiderbauer , R., T oberer , F ., Buhl, T ., Blum, A., Kalloo, A., Hassen, A.B.H., Thomas, L., Enk, A., Uhlmann, L., Alt, C., Arenbergero va, M., Bakos, R., Baltzer, A., Bertlich, I., Blum, A., Bokor- Billmann, T ., Bo wling, J., Braghiroli, N., Braun, R., Buder-Bakhaya, K., Buhl, T ., Cabo, H., Cabrijan, L., Cevic, N., Classen, A., Deltgen, D., Fink, C., Georgiev a, I., Hakim-Meibodi, L.E., Hanner , S., Hartmann, F ., Hart- mann, J., Haus, G., Hoxha, E., Karls, R., Koga, H., Kreusch, J., Lallas, A., Majenka, P ., Mar ghoob, A., Massone, C., Mekokishvili, L., Mestel, D., Meyer , V ., Neuberger , A., Nielsen, K., Oliviero, M., Pampena, R., Paoli, J., Pawlik, E., Rao, B., Rendon, A., Russo, T ., Sadek, A., Samhaber, K., Schnei- derbauer , R., Schweizer , A., T oberer, F ., T rennheuser , L., Vlahov a, L., W ald, A., W inkler , J., W ¨ olbing, P ., Zalaudek, I., 2018. Man against machine: di- agnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Annals of Oncology 29, 1836–1842. URL: https://academic . oup . com/ annonc/article/29/8/1836/5004443 , doi: 10 . 1093/annonc/mdy166 . Hauptmann, A., Lin, W .H., Y an, R., Y ang, J., Chen, M.y ., 2006. Extreme video retriev al: Joint maximization of human and computer performance, pp. 385– 394. doi: 10 . 1145/1180639 . 1180721 . Hesamian, M.H., Jia, W ., He, X., K ennedy , P ., 2019. Deep Learning T echniques for Medical Image Segmentation: Achievements and Challenges. Jour- nal of Digital Imaging 32, 582–596. URL: http://link . springer . com/ 10 . 1007/s10278- 019- 00227- x , doi: 10 . 1007/s10278- 019- 00227- x . Holzinger , A., Malle, B., Kieseber g, P ., Roth, P .M., M ¨ uller , H., Reihs, R., Zatloukal, K., 2017. T owards the Augmented Pathologist: Challenges of Explainable-AI in Digital Pathology URL: http://arxiv . org/abs/ 1712 . 06657 . Jang, W .D., Kim, C.S., 2019. Interacti ve image segmentation via backprop- agating refinement scheme, in: Proceedings of The IEEE Conference on Computer V ision and Pattern Recognition. Jungo, A., Reyes, M., 2019. Assessing Reliability and Challenges of Uncertainty Estimations for Medical Image Segmentation. T echni- cal Report. URL: https://github . com/alainjungo/reliability- challenges- uncertainty . Kamnitsas, K., Baumgartner, C., Ledig, C., Newcombe, V ., Simpson, J., Kane, A., Menon, D., Nori, A., Criminisi, A., Rueckert, D., et al., 2017. Unsu- pervised domain adaptation in brain lesion segmentation with adversarial networks, in: International conference on information processing in medical imaging, Springer . pp. 597–609. Khosrav an, N., Celik, H., T urkbey , B., Cheng, R., McCreedy , E., McAuli ff e, M., Bednarov a, S., Jones, E., Chen, X., Choyke, P ., W ood, B., Bagci, U., 2017. Gaze2Segment: A Pilot Study for Integrating Eye-T racking T echnology into Medical Image Segmentation, Springer, Cham, pp. 94– 104. URL: http://link . springer . com/10 . 1007/978- 3- 319- 61188- 4 9 , doi: 10 . 1007/978- 3- 319- 61188- 4{ \ }9 . Kim, J.S., Greene, M.J., Zlateski, A., Lee, K., Richardson, M., T uraga, S.C., Purcaro, M., Balkam, M., Robinson, A., Behabadi, B.F ., Campos, M., Denk, W ., Seung, H.S., EyeW irers, t., 2014. Space–time wiring specificity supports direction selectivity in the retina. Nature 509, 331–336. URL: http:// www . nature . com/articles/nature13240 , doi: 10 . 1038/nature13240 . 16 King, R.D., Ro wland, J., Oliver , S.G., Y oung, M., Aubrey , W ., Byrne, E., Liakata, M., Markham, M., Pir, P ., Soldatova, L.N., Sparkes, A., Whelan, K.E., Clare, A., 2009. The automation of science. Science 324, 85–89. URL: https://pubmed . ncbi . nlm . nih . gov/19342587/ , doi: 10 . 1126/science . 1165620 . King, R.D., Whelan, K.E., Jones, F .M., Reiser, P .G., Bryant, C.H., Mug- gleton, S.H., Kell, D.B., Oliver , S.G., 2004. Functional genomic hy- pothesis generation and experimentation by a robot scientist. Nature 427, 247–252. URL: https://pubmed . ncbi . nlm . nih . gov/14724639/ , doi: 10 . 1038/nature02236 . Kirsch, A., van Amersfoort, J., Gal, Y ., 2019. Batchbald: E ffi cient and div erse batch acquisition for deep bayesian active learning, in: Ad- vances in Neural Information Processing Systems 32. Curran Associates, Inc., pp. 7026–7037. URL: http://papers . nips . cc/paper/8925- batchbald- efficient- and- diverse- batch- acquisition- for- deep- bayesian- active- learning . pdf . K onyushkov a, K., Sznitman, R., Fua, P ., 2017. Learning Active Learning from Data. URL: https://papers . nips . cc/paper/7010- learning- active- learning- from- data . K onyushkov a, K., Sznitman, R., Fua, P ., 2019. Geometry in ac- tiv e learning for binary and multi-class image segmentation. Computer V ision and Image Understanding 182, 1–16. URL: https://www . sciencedirect . com/science/article/pii/ S107731421930013X , doi: 10 . 1016/J . CVIU . 2019 . 01 . 007 . Kuo, W ., H ¨ ane, C., Y uh, E., Mukherjee, P ., Malik, J., 2018. Cost-Sensitive Activ e Learning for Intracranial Hemorrhage Detection, Springer, Cham, pp. 715–723. URL: http://link . springer . com/10 . 1007/978- 3- 030- 00931- 1 82 , doi: 10 . 1007/978- 3- 030- 00931- 1{ \ }82 . Kurzendorfer , T ., Fischer , P ., Mirshahzadeh, N., Pohl, T ., Brost, A., Steidl, S., Maier , A., 2017. Rapid Interactive and Intuitive Segmentation of 3D Medical Images Using Radial Basis Function Interpolation †. Annual Conference on Medical Image Understanding and Analysis , 11–13URL: www . mdpi . com/ journal/jimaging , doi: 10 . 3390/jimaging3040056 . Kushibar , K., V alverde, S., Gonz ´ alez-V ill ` a, S., Bernal, J., Cabezas, M., Oliv er, A., Llad ´ o, X., 2019. Supervised Domain Adaptation for Automatic Sub-cortical Brain Structure Segmentation with Minimal User Interaction. Scientific Reports 9, 6742. URL: http://www . nature . com/articles/ s41598- 019- 43299- z , doi: 10 . 1038/s41598- 019- 43299- z . Lang, K., Baum, E., 1992. Query learning can work poorly when a human oracle is used, in: Proceedings of the IEEE International Joint Conference on Neural Networks, IEEE. pp. 335—-340. Last, F ., Klein, T ., Ra vanbakhsh, M., Nabi, M., Batmanghelich, K., Tresp, V ., 2018. Human-Machine Collaboration for Medical Image Segmentation. T echnical Report. URL: https://pdfs . semanticscholar . org/4e0c/ 535386e3a3d307cee45e97b9417eff4da92e . pdf . Lewis, D.D., Catlett, J., 1994. Heterogeneous uncertainty sampling for super - vised learning, in: In Proceedings of the Ele venth International Conference on Machine Learning, Morgan Kaufmann. pp. 148–156. Li, S., Jiang, Y ., Chawla, N.V ., Zhou, Z., 2019. Multi-label learning from crowds. IEEE Transactions on Knowledge and Data Engineering 31, 1369– 1382. doi: 10 . 1109/TKDE . 2018 . 2857766 . Lian, C., Liu, M., Zhang, J., Shen, D., 2020. Hierarchical fully conv olutional network for joint atrophy localization and Alzheimer’ s disease diagnosis us- ing structural MRI. IEEE Transactions on Pattern Analysis and Machine Intelligence 42, 880–893. doi: 10 . 1109/TPAMI . 2018 . 2889096 . Liao, X., Li, W .H., Xu, Q., W ang, X., Jin, B., Zhang, X., Zhang, Y ., W ang, Y ., 2020. Iteratively-refined interactiv e 3d medical image segmentation with multi-agent reinforcement learning. 2020 IEEE / CVF Conference on Com- puter V ision and Pattern Recognition (CVPR) , 9391–9399. Litjens, G., Kooi, T ., Bejnordi, B.E., Setio, A.A.A., Ciompi, F ., Ghafoorian, M., van der Laak, J.A., van Ginneken, B., S ´ anchez, C.I., 2017. A survey on deep learning in medical image analysis. Medical Image Analysis 42, 60– 88. URL: https://www . sciencedirect . com/science/article/pii/ S1361841517301135 , doi: 10 . 1016/J . MEDIA . 2017 . 07 . 005 . Liu, M., Zhang, J., Adeli, E., Shen, D., 2018. Landmark-based deep multi- instance learning for brain disease diagnosis. Medical Image Analysis 43, 157–168. doi: 10 . 1016/j . media . 2017 . 10 . 005 . Lowell, D., Lipton, Z.C., W allace, B.C., 2019. Practical obstacles to deploying activ e learning, in: Proceedings of the 2019 Conference on Empirical Meth- ods in Natural Language Processing and the 9th International Joint Confer- ence on Natural Language Processing (EMNLP-IJCNLP), Association for Computational Linguistics, Hong K ong, China. pp. 21–30. URL: https:// www . aclweb . org/anthology/D19- 1003 , doi: 10 . 18653/v1/D19- 1003 . Lundervold, A.S., Lunderv old, A., 2019. An overvie w of deep learning in med- ical imaging focusing on MRI. Zeitschrift f ¨ ur Medizinische Physik 29, 102– 127. URL: https://www . sciencedirect . com/science/article/ pii/S0939388918301181 , doi: 10 . 1016/J . ZEMEDI . 2018 . 11 . 002 . Mahapatra, D., Bozorgtabar , B., Thiran, J.P ., Reyes, M., 2018. E ffi cient Active Learning for Image Classification and Segmentation Using a Sample Se- lection and Conditional Generative Adversarial Network, Springer , Cham, pp. 580–588. URL: http://link . springer . com/10 . 1007/978- 3- 030- 00934- 2 65 , doi: 10 . 1007/978- 3- 030- 00934- 2{ \ }65 . Mar , V .J., Soyer , H.P ., 2018. Artificial intelligence for melanoma diagno- sis: how can we deliver on the promise? Annals of Oncology 29, 1625–1628. URL: https://academic . oup . com/annonc/article/29/ 8/1625/5004449 , doi: 10 . 1093/annonc/mdy193 . McCallum, A., Nigam, K., 1998. Emplo ying em and pool-based activ e learning for text classification, in: Proceedings of the Fifteenth International Confer- ence on Machine Learning, Morgan Kaufmann Publishers Inc., San Fran- cisco, CA, USA. p. 350–358. doi: 10 . 5555/645527 . 757765 . Milletari, F ., Birodkar , V ., Sofka, M., 2019. Straight to the Point: Reinforce- ment learning for user guidance in ultrasound, in: Lecture Notes in Com- puter Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), Springer. pp. 3–10. URL: https:// doi . org/10 . 1007/978- 3- 030- 32875- 7 1 , doi: 10 . 1007/978- 3- 030- 32875- 7{ \ }1 . Moeskops, P ., W olterink, J.M., van der V elden, B.H.M., Gilhuijs, K.G.A., Leiner , T ., V ierge ver , M.A., I ˇ sgum, I., 2016. Deep Learning for Multi- task Medical Image Segmentation in Multiple Modalities, Springer , Cham, pp. 478–486. URL: http://link . springer . com/10 . 1007/978- 3- 319- 46723- 8 55 , doi: 10 . 1007/978- 3- 319- 46723- 8{ \ }55 . Morid, M.A., Borjali, A., Del Fiol, G., 2021. A scoping review of transfer learning research on medical image analysis using ImageNet. doi: 10 . 1016/ j . compbiomed . 2020 . 104115 . Nalisnik, M., Gutman, D.A., Kong, J., Cooper, L.A., 2015. An Inter- activ e Learning Framew ork for Scalable Classification of P athology Images. Proceedings : ... IEEE International Conference on Big Data. IEEE International Conference on Big Data 2015, 928–935. URL: http://www . ncbi . nlm . nih . gov/pubmed/27796014http: //www . pubmedcentral . nih . gov/articlerender . fcgi?artid= PMC5082843 , doi: 10 . 1109/BigData . 2015 . 7363841 . NICE, 2013. Judging whether public health interv entions o ff er value for money — Guidance and guidelines — NICE URL: https://www . nice . org . uk/ advice/lgb10 . Oktay , O., Schlemper, J., Folgoc, L., Lee, M., Heinrich, M., Misawa, K., Mori, K., McDonagh, S., Hammerla, N., Kainz, B., Glocker, B., Rueckert, D., 2018. Attention u-net: Learning where to look for the pancreas URL: https://arxiv . org/pdf/1804 . 03999 . pdf . Ozdemir , F ., Peng, Z., T anner , C., Fuernstahl, P ., Goksel, O., 2018. Activ e Learning for Segmentation by Optimizing Content Information for Maximal Entropy , Springer, Cham, pp. 183–191. URL: http:// link . springer . com/10 . 1007/978- 3- 030- 00889- 5 21 , doi: 10 . 1007/ 978- 3- 030- 00889- 5{ \ }21 . Pan, Y ., Liu, M., Lian, C., Xia, Y ., Shen, D., 2020. Spatially-Constrained Fisher Representation for Brain Disease Identification with Incomplete Multi-Modal Neuroimages. IEEE T ransactions on Medical Imaging 39, 2965–2975. URL: https://pubmed . ncbi . nlm . nih . gov/32217472/ , doi: 10 . 1109/TMI . 2020 . 2983085 . Parisi, G.I., Kemk er, R., Part, J.L., Kanan, C., W ermter , S., 2019. Con- tinual lifelong learning with neural networks: A revie w . doi: 10 . 1016/ j . neunet . 2019 . 01 . 012 . of Radiologists, T .R.C., 2017. Clinical radiology UK workforce cen- sus 2017 report. T echnical Report. URL: https://www . rcr . ac . uk/ system/files/publication/field publication files/ bfcr185 cr census 2017 . pdf . Raghu, M., Zhang, C., Kleinberg, J., Bengio, S., 2019. T ransfusion: Under- standing transfer learning for medical imaging, in: NeurIPS. Rajchl, M., Koch, L.M., Ledig, C., Passerat-Palmbach, J., Misawa, K., Mori, K., Rueckert, D., 2017. Employing W eak Annotations for Medical Image Analysis Problems URL: https://arxiv . org/pdf/1708 . 06297v1 . pdf . Rajchl, M., Lee, M.C.H., Oktay , O., Kamnitsas, K., Passerat-Palmbach, J., Bai, W ., Damodaram, M., Rutherford, M.A., Hajnal, J.V ., Kainz, B., Rueckert, D., 2016a. Deepcut: Object segmentation from bounding box annotations using conv olutional neural networks. IEEE Transactions on Medical Imag- 17 ing 36, 674–683. Rajchl, M., Lee, M.C.H., Schrans, F ., Davidson, A., Passerat-Palmbach, J., T arroni, G., Alansary , A., Oktay , O., Kainz, B., Rueck ert, D., 2016b . Learn- ing under Distributed W eak Supervision URL: https://arxiv . org/pdf/ 1606 . 01100v1 . pdf . Rodrigues, F ., Pereira, F .C., 2018. Deep learning from crowds, in: The Thirty- Second AAAI Conference on Artificial Intelligence (AAAI), 2018, AAAI Press. pp. 1611–1618. Ronneberger , O., Fischer , P ., Brox, T ., 2015. U-Net: Conv olutional Net- works for Biomedical Image Segmentation, Springer , Cham, pp. 234–241. URL: http://link . springer . com/10 . 1007/978- 3- 319- 24574- 4 28 , doi: 10 . 1007/978- 3- 319- 24574- 4{ \ }28 . Schlemper , J., Oktay , O., Schaap, M., Heinrich, M., Kainz, B., Glocker, B., Rueckert, D., 2019. Attention gated networks: Learning to leverage salient regions in medical images. Medical Image Analysis 53, 197–207. doi: 10 . 1016/j . media . 2019 . 01 . 012 . Settles, B., 2009. Acti ve learning literature surve y . T echnical Report. University of W isconsin-Madison Department of Computer Sciences. Settles, B., Crav en, M., 2008. An analysis of active learning strategies for sequence labeling tasks, in: Proceedings of the Conference on Empirical Methods in Natural Language Processing, Association for Computational Linguistics, USA. p. 1070–1079. doi: 10 . 5555/1613715 . 1613855 . Shah, M.P ., Bhalgat, Y .S., A wate, S.P ., 2018. Annotation-cost Mini- mization for Medical Image Segmentation using Suggestiv e Mixed Supervision Fully Conv olutional Networks. T echnical Report. URL: https://www . doc . ic . ac . uk/ ~ bglocker/public/mednips2018/med- nips 2018 paper 30 . pdf . Shannon, C.E., . A Mathematical Theory of Communication. T echnical Report. Shen, D., Wu, G., Suk, H.I., 2017. Deep Learning in Medical Image Analysis. Annual re view of biomedical engineering 19, 221–248. URL: http://www . ncbi . nlm . nih . gov/pubmed/28301734http: //www . pubmedcentral . nih . gov/articlerender . fcgi?artid= PMC5479722 , doi: 10 . 1146/annurev- bioeng- 071516- 044442 . Smailagic, A., Noh, H.Y ., Costa, P ., W alawalkar , D., Khandelwal, K., Mirshekari, M., Fagert, J., Galdran, A., Xu, S., 2018. MedAL: Deep Activ e Learning Sampling Method for Medical Image Analysis. unde- fined URL: https://www . semanticscholar . org/paper/MedAL%3A- Deep- Active- Learning- Sampling- Method- for- Smailagic- Noh/ fa23dc7a8b3927953d83f5ce46e0b622b7cac456 . Sourati, J., Gholipour , A., Dy , J.G., K urugol, S., W arfield, S.K., 2018. Activ e Deep Learning with Fisher Information for Patch-W ise Semantic Segmentation, in: Deep learning in medical image analysis and multi- modal learning for clinical decision support : 4th International W orkshop, DLMIA 2018, and 8th International W orkshop, ML-CDS 2018, held in conjunction with MICCAI 2018, Granada, Spain, S.... volume 11045, pp. 83–91. URL: http://www . ncbi . nlm . nih . gov/pubmed/30450490http: //www . pubmedcentral . nih . gov/articlerender . fcgi?artid= PMC6235453http://link . springer . com/10 . 1007/978- 3- 030- 00889- 5 10 , doi: 10 . 1007/978- 3- 030- 00889- 5{ \ }10 . Stember , J.N., Celik, H., Krupinski, E., Chang, P .D., Mutasa, S., W ood, B.J., Lignelli, A., Moonis, G., Schwartz, L.H., Jambawalikar , S., Bagci, U., 2019. Eye Tracking for Deep Learning Segmentation Using Con volutional Neural Networks. Journal of Digital Imaging 32, 597–604. URL: http: //link . springer . com/10 . 1007/s10278- 019- 00220- 4 , doi: 10 . 1007/ s10278- 019- 00220- 4 . Stoyanov , D., T aylor, Z., Kia, S.M., Oguz, I., Reyes, M., Martel, A., Maier- Hein, L., Marquand, A.F ., Duchesnay , E., L ¨ ofstedt, T ., Landman, B., Car- doso, M.J., Silv a, C.A., Pereira, S., Meier , R. (Eds.), 2018. Understanding and Interpreting Machine Learning in Medical Image Computing Applica- tions. volume 11038 of Lecture Notes in Computer Science . Springer Inter- national Publishing, Cham. URL: http://link . springer . com/10 . 1007/ 978- 3- 030- 02628- 8 , doi: 10 . 1007/978- 3- 030- 02628- 8 . Suzuki, K., 2017. Overvie w of deep learning in medical imag- ing. Radiological Physics and T echnology 10, 257–273. URL: http://www . ncbi . nlm . nih . gov/pubmed/28689314http: //link . springer . com/10 . 1007/s12194- 017- 0406- 5 , doi: 10 . 1007/ s12194- 017- 0406- 5 . T ajbakhsh, N., Jeyaseelan, L., Li, Q., Chiang, J.N., W u, Z., Ding, X., 2020. Em- bracing imperfect datasets: A revie w of deep learning solutions for medical image segmentation. Medical Image Analysis 63, 101693. doi: 10 . 1016/ j . media . 2020 . 101693 . T ajbakhsh, N., Shin, J.Y ., Gurudu, S.R., Hurst, R.T ., Kendall, C.B., Got- way , M.B., Liang, J., 2016. Con volutional Neural Networks for Medical Image Analysis: Full Training or Fine Tuning? IEEE Transactions on Medical Imaging 35, 1299–1312. URL: http://ieeexplore . ieee . org/ document/7426826/ , doi: 10 . 1109/TMI . 2016 . 2535302 . T inati, R., Luczak-Roesch, M., Simperl, E., Hall, W ., 2017. An in vestigation of player motivations in Eyewire, a gamified citizen science project. Comput- ers in Human Behavior 73, 527–540. URL: https://ac . els- cdn . com/ S0747563216309037/1- s2 . 0- S0747563216309037- main . pdf? tid= abaecc23- 8276- 4fab- ad47- 300db95a3b56&acdnat= 1523979569 9e91cf49a50416e7c8772bfb73614a6b , doi: 10 . 1016/ j . chb . 2016 . 12 . 074 . T izhoosh, H.R., Pantano witz, L., 2018. Artificial Intelligence and Digital Pathology: Challenges and Opportunities. Journal of pathology infor - matics 9, 38. URL: http://www . ncbi . nlm . nih . gov/pubmed/30607305 , doi: 10 . 4103/jpi . jpi{ \ }53{ \ }18 . W ang, D., Li, M., Ben-Shlomo, N., Corrales, C.E., Cheng, Y ., Zhang, T ., Jayen- der , J., 2019a. Mixed-Supervised Dual-Network for Medical Image Seg- mentation, in: Lecture Notes in Computer Science (including subseries Lec- ture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), Springer . pp. 192–200. URL: https://doi . org/10 . 1007/978- 3- 030- 32245- 8 22 , doi: 10 . 1007/978- 3- 030- 32245- 8{ \ }22 . W ang, G., Li, W ., Aertsen, M., Deprest, J., Ourselin, S., V ercauteren, T ., 2019b . Aleatoric uncertainty estimation with test-time augmentation for medical image segmentation with con volutional neural networks. Neurocomputing 338, 34–45. doi: 10 . 1016/J . NEUCOM . 2019 . 01 . 103 . W ang, G., Li, W ., Zuluaga, M.A., Pratt, R., Patel, P .A., Aertsen, M., Doel, T ., David, A.L., Deprest, J., Ourselin, S., V ercauteren, T ., 2018. Interactiv e medical image segmentation using deep learning with image-specific fine tuning. IEEE Transactions on Medical Imaging 37, 1562–1573. W ang, G., Zuluaga, M.A., Li, W ., Pratt, R., Patel, P .A., Aertsen, M., Doel, T ., David, A.L., Deprest, J., Ourselin, S., V ercauteren, T ., 2019. Deepigeos: A deep interacti ve geodesic frame work for medical image segmentation. IEEE T ransactions on Pattern Analysis and Machine Intelligence 41, 1559–1572. W ang, H., Riv enson, Y ., Jin, Y ., W ei, Z., Gao, R., G ¨ unaydın, H., Bentolila, L.A., Kural, C., Ozcan, A., 2019. Deep learning enables cross-modality super-resolution in fluorescence microscopy . Nature Methods 16, 103– 110. URL: https://www . nature . com/articles/s41592- 018- 0239- 0 , doi: 10 . 1038/s41592- 018- 0239- 0 . W ang, K., Zhang, D., Li, Y ., Zhang, R., Lin, L., 2017. Cost-E ff ective Ac- tiv e Learning for Deep Image Classification. IEEE Transactions on Cir- cuits and Systems for V ideo T echnology 27, 2591–2600. doi: 10 . 1109/ TCSVT . 2016 . 2589879 . W ang, Y ., Y u, B., W ang, L., Zu, C., Lalush, D.S., Lin, W ., W u, X., Zhou, J., Shen, D., Zhou, L., 2018. 3D conditional generativ e adversarial net- works for high-quality PET image estimation at low dose. NeuroImage 174, 550–562. URL: https://pubmed . ncbi . nlm . nih . gov/29571715/ , doi: 10 . 1016/j . neuroimage . 2018 . 03 . 045 . W en, S., Kurc, T .M., Hou, L., Saltz, J.H., Gupta, R.R., Batiste, R., Zhao, T ., Nguyen, V ., Samaras, D., Zhu, W ., 2018. Comparison of Di ff erent Classi- fiers with Active Learning to Support Quality Control in Nucleus Segmen- tation in Pathology Images. AMIA Joint Summits on T ranslational Science proceedings. AMIA Joint Summits on Translational Science 2017, 227–236. URL: http://www . ncbi . nlm . nih . gov/pubmed/29888078 . W oodward, M., Finn, C., Research, B.A., 2017. Active One-shot Learning. T echnical Report. URL: https://arxiv . org/pdf/1702 . 06559 . pdf . Xia, Y ., Liu, F ., Y ang, D., Cai, J., Y u, L., Zhu, Z., Xu, D., Y uille, A., Roth, H., 2020. 3d semi-supervised learning with uncertainty-aw are multi-view co-training, in: 2020 IEEE W inter Conference on Applications of Computer V ision (W A CV), pp. 3635–3644. Y amashita, R., Nishio, M., Do, R.K.G., T ogashi, K., 2018. Convolutional neu- ral networks: an overvie w and application in radiology . Insights into Imag- ing 9, 611–629. URL: https://insightsimaging . springeropen . com/ articles/10 . 1007/s13244- 018- 0639- 9 , doi: 10 . 1007/s13244- 018- 0639- 9 . Y ang, L., Zhang, Y ., Chen, J., Zhang, S., Chen, D.Z., 2017. Sugges- tiv e Annotation: A Deep Active Learning Framework for Biomedical Image Segmentation, Springer, Cham, pp. 399–407. URL: http:// link . springer . com/10 . 1007/978- 3- 319- 66179- 7 46 , doi: 10 . 1007/ 978- 3- 319- 66179- 7{ \ }46 . Y i, X., W alia, E., Babyn, P ., 2019. Generativ e adversarial network in medical imaging: A revie w. Medical Image Analysis 58, 101552. doi: 10 . 1016/ j . media . 2019 . 101552 . 18 Y u, B., Zhou, L., W ang, L., Shi, Y ., Fripp, J., Bourgeat, P ., 2019. Ea-GANs: Edge-A ware Generative Adversarial Networks for Cross-Modality MR Im- age Synthesis. IEEE Transactions on Medical Imaging 38, 1750–1762. doi: 10 . 1109/TMI . 2019 . 2895894 . Zhang, C., Chaudhuri, K., 2015. Active learning from weak and strong label- ers, in: Proceedings of the 28th International Conference on Neural Infor- mation Processing Systems - V olume 1, MIT Press, Cambridge, MA, USA. p. 703–711. doi: 10 . 5555/2969239 . 2969318 . Zhang, C., Chen, T ., 2002. An acti ve learning framework for content-based information retriev al. IEEE Trans. Multimedia 4, 260–268. Zhao, A., Balakrishnan, G., Durand, F ., Guttag, J.V ., Dalca, A.V ., 2019. Data augmentation using learned transformations for one-shot medical image seg- mentation, in: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 8543–8553. Zheng, S., Jayasumana, S., Romera-Paredes, B., V ineet, V ., Su, Z., Du, D., Huang, C., T orr, P .H.S., 2015. Conditional Random Fields as Recurrent Neural Networks. T echnical Report. Zhou, Z., Shin, J., Feng, R., Hurst, R.T ., Kendall, C.B., Liang, J., 2018a. Integrating Active Learning and T ransfer Learning for Carotid Intima-Media Thickness V ideo Interpretation. Journal of Digital Imag- ing URL: http://www . ncbi . nlm . nih . gov/pubmed/30402668http: //link . springer . com/10 . 1007/s10278- 018- 0143- 2 , doi: 10 . 1007/ s10278- 018- 0143- 2 . Zhou, Z., Shin, J., Feng, R., Hurst, R.T ., Kendall, C.B., Liang, J., 2018b . Integrating Acti ve Learning and T ransfer Learning for Carotid Intima-Media Thickness V ideo Interpretation. Journal of Digital Imag- ing URL: http://www . ncbi . nlm . nih . gov/pubmed/30402668http: //link . springer . com/10 . 1007/s10278- 018- 0143- 2 , doi: 10 . 1007/ s10278- 018- 0143- 2 . Zhou, Z., Shin, J., Zhang, L., Gurudu, S., Gotway , M., Liang, J., 2017. Fine-tuning Conv olutional Neural Networks for Biomedical Image Analysis: Actively and Incrementally . Proceedings. IEEE Computer Society Conference on Computer V ision and Pattern Recognition 2017, 4761. URL: http://www . ncbi . nlm . nih . gov/pubmed/30337799http: //www . pubmedcentral . nih . gov/articlerender . fcgi?artid= PMC6191179 , doi: 10 . 1109/CVPR . 2017 . 506 . 19 Supplementary Material 20 Active Learning Interpretation Refinement Related Practical Author Year Uncertainty Representativeness Learning AL Bespoke Uncertainty Bayesian V isual Bespoke Interactive T ask Prior Bespoke T ransfer SSL Reinforcement Generative Continual User Model W eak Labels Multi-task Cost sensistive Interface W ang et al 2017 W en et al 2018 Beluch et al 2018 Gal et al 2017 Kirsch et al 2019 Konyushkova et al 2019 Y ang et al 2017 Smailagic et al 2018 Ozdemir et al 2018 Sourati et al 2018 Konyushkova et al 2017 Bachman et al 2017 T ajbakhsh et al 2016 Oktay et al 2018 Schlemper et al 2019 Ng et al 2018 Budd et al 2019 W ang et al 2019 Paschali et al 2019 Jungo et al 2019 Amrehn et al 2017 Zheng et al 2015 W ang et al 2017 W ang et al 2017 Bredell et al 2018 Kurzendorfer et al 2017 Jang et al 2019 Liao et al 2020 Li et al 2019 Zhang et al 2015 Cheplygina et al 2017 Rodrigues et al 2017 Rajchl et al 2017 Moeskops et al 2016 W ang et al 2019 Nalisnik et al 2015 Khosravan et al 2017 Stember et al 2019 T inati et al 2017 Kuo et al 2018 Shah et al 2018 W oodward et al 2017 Fang et al 2017 Milletari et al 2019 Last et al 2018 Mahapatra et al 2018 Zhao et al 2019 T ajbakhsh et al 2016 Zhou et al 2017 Zhou et al 2018 Baweja et al 2018 W ang et al 2019 W ang et al 2019 Y u et al 2019 Pan et al 2020 Lian et al 2020 Liu et al 2018 Farquhar et al 2021 Fig. 4. T able of features demonstrated by work discussed in this re view 21
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment