Beat by Beat: Classifying Cardiac Arrhythmias with Recurrent Neural Networks
With tens of thousands of electrocardiogram (ECG) records processed by mobile cardiac event recorders every day, heart rhythm classification algorithms are an important tool for the continuous monitoring of patients at risk. We utilise an annotated d…
Authors: Patrick Schwab, Gaetano Scebba, Jia Zhang
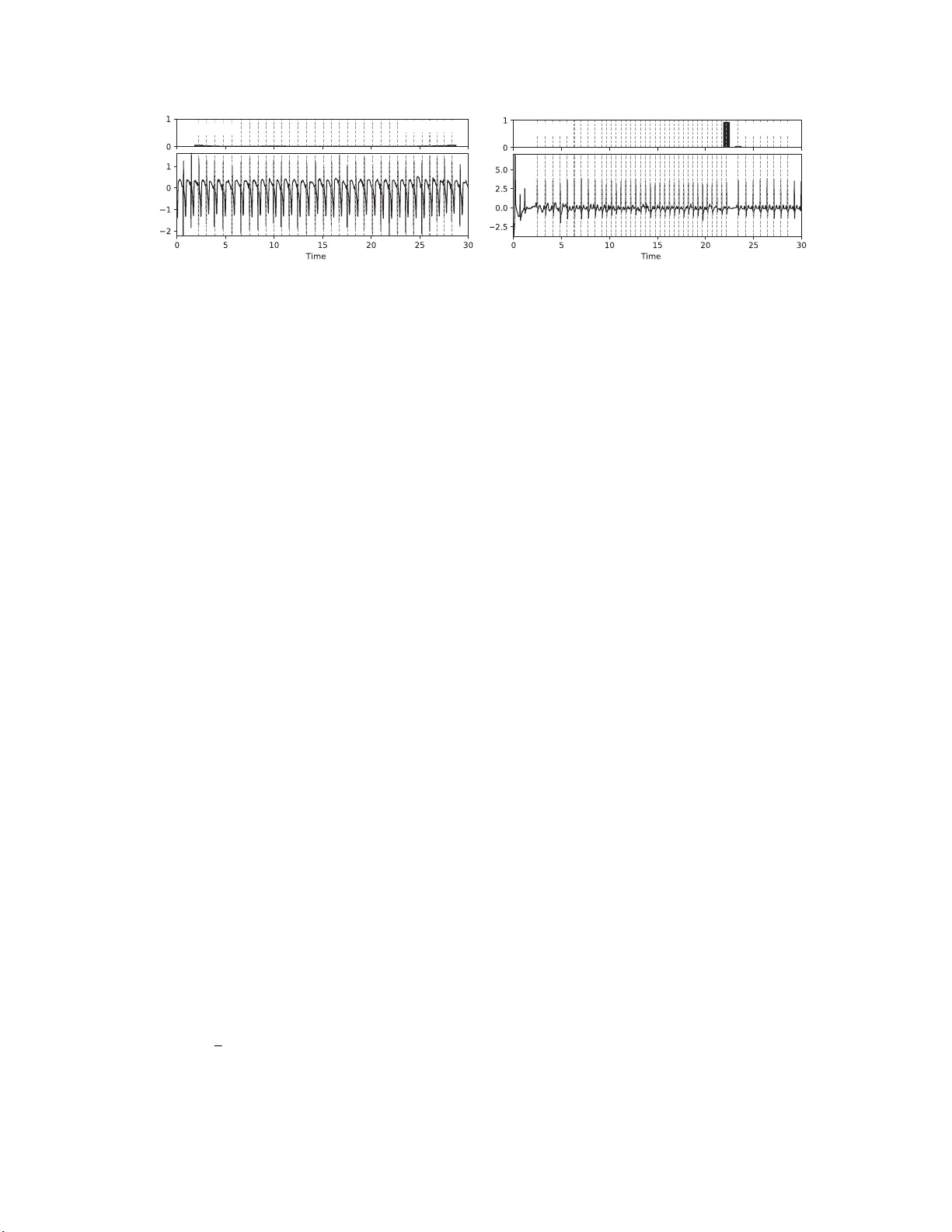
Beat by Beat: Classifying Cardiac Arrh ythmias with Recurren t Neural Networks Patrick Schwab, Gaetano C Scebba, Jia Zhang, Marco Delai, W alter Karlen Mobile Health Sys t ems Lab, Department of Health Sciences and T echnolog y ETH Zurich, Switzerland Abstract W ith tens of thousand s of electr oca r d iogram (ECG) r ec o r ds pr oce ssed b y mobile ca r d iac event r eco r d ers every day , heart rhythm classifica tio n algorithms are an impor- tant tool for the continuo us mo nitoring of patients a t risk. W e utilise an a nnotated dataset of 12,18 6 single-lead ECG r ec o r dings to build a diverse ensemble of r ecurr ent neu- ral networks ( RNNs) that is able to distinguish between normal sinu s rhythms, atrial fib rillation, other types of a r- rhythmia and signa ls that ar e too noisy to interpr et. In or der to ease learning over the temporal dimension, we in- tr odu ce a n ovel task formulation that harn e sses the natu ral se g mentation of ECG signals into h eartbeats to drastically r ed uce the number of time steps p er sequence. Ad d ition- ally , we extend our RNNs with a n attention mechanism that enables us to r eason abou t which heartbeats our RNNs fo- cus on to make their decisio n s. Thr ou gh the use of a tten- tion, our mod el main ta ins a high degr ee o f in terpr etability , while also achieving state-o f-the-art cla ssification perfor- mance with an average F1 score of 0. 79 on an unseen test set (n=3,658 ). 1. Intr oduction Cardiac arrhythmias are a heterogen ous group of con- ditions tha t is characterised by hea r t rhythms that do not follow a normal sinus pa ttern. One of the most com- mon arrh y thmias is atrial fibrillation (A F) with an age- depend ant population prev alence of 2 . 3 - 3 . 4% [1]. Due to the increased m ortality associated with arrhy thmias, re - ceiving a timely d iagnosis is of paramo u nt imp ortance for patients [1, 2]. T o diagno se card ia c arrh y thmias, medical profession a ls typically con sider a p atient’ s electr ocardio- gram (ECG) as one of the prim a ry factors [2]. In the past, clinicians recorded these ECGs main ly using m u lti-lead clinical mo n itors or Ho lter devices. Howe ver , the recent advent of m obile card ia c ev ent record ers has given patients the ability to remotely record shor t ECGs using d evices with a sin g le le a d. W e pro pose a machine-learn ing app roach based on re- current neural n etworks (RNNs) to dif ferentiate between various types of heart rhythms in this mo re challenging set- ting with just a single lead and sho rt E CG record lengths. T o ease learning of depen dencies over the tempor al dimen- sion, we introdu ce a n ovel task f ormulation that ha r nesses the natural be at-wise segmen tation of ECG signals. In ad - dition to utilising se veral he a r tbeat features that h av e be e n shown to be highly discriminative in previous works, we also u se stacked denoising au toencod e r s (SD AE) [3] to capture differences in morpholo gical structure. Further- more, we extend our RNNs with a soft attention m ech- anism [4 – 7] that en ables u s to reason ab out which ECG segments th e RNNs prioritise for their d ecision making. 2. Methodology Our cardiac rhy thm classification pipeline consists of multiple stages ( fig ure 1). The core idea of our setup is to extract a di verse set of featu res from the sequence of heartbeats in an ECG r ecord to be used a s input features to an ensemble of RNNs. W e b lend the in dividual m od- els’ p redictions into a p er-class classification scor e using a multilayer perceptro n (MLP) with a softmax o utput layer . The following paragrap h s explain the stages shown in fig- ure 1 in mor e d etail. ECG Dataset. W e use the dataset of the Phy s- ioNet Computin g in Card iology (CinC) 20 1 7 challen g e [8] which contain s 12,186 uniq ue single-lead ECG r e cords of varying length. Experts an notated each of these ECGs as b e ing either a normal sinu s rhythm, AF , an other ar- Normalise Segment Model 1 Model n ... Extract Features Blend Normal AF Other Noise ECG Preprocessing Features Level 1 Models Level 2 Blender Classification Figure 1. An overvie w of our ECG classification pipeline . rhythm ia or too noisy to classify . The ch allenge organisers keep 3,6 5 8 ( 30% ) of th ese ECG records priv ate as a test set. Add itionally , we ho ld ou t a n on-stratified r andom sub- set of 20% of the public dataset as a validation set. F or some RNN co nfiguratio ns, we further augme nt the train- ing data with labelled samples extracted from other Phy- sioNet da tab ases [9–12] in o rder to e ven ou t misbalanced class sizes in the training set. As an additional measu r e against the im balanced class distribution of the d a taset, we weight each trainin g sample’ s contr ibution to the loss func- tion to b e inv ersely proportio nal to its class’ prevalence in the overall d ataset. Normalisation. Prior to segmentation, we n ormalise the ECG reco rding to have a m ean value of zero and a standard deviation o f one. W e do no t apply any additional filters as all ECGs were b a n dpass-filtered b y the reco rding d evice. Segmentation. Follo wing normalisation, we segment the ECG into a sequence o f heartbeats. W e decide to re- formu late the given task o f classifying arrhy th mias as a se- quence classification task over he artbeats rath er than over raw ECG reading s. T he mo ti vation behind th e r eformu la- tion is that it significantly re duces the nu mber of time steps throug h which the error signal of our RNNs has to prop - agate. On the training set, the reform u lation reduces the mean number o f time steps per E CG from 9 000 to just 33 . T o perform the segmentation, we use a cu stomised QRS detector based on Pan-T ompk in’ s [1 3] that iden tifies R- peaks in the ECG r ecording . W e extend their algo rithm by adapting th e th reshold with a moving average of the ECG signal to be more r esilient against the com monly encoun - tered short bursts of noise. For the p urpose o f this w ork, we define heartb eats u sing a symmetric fixed size win dow with a total length of 0 . 66 seconds arou nd R-peaks. W e pass the extracted h eartbeat sequ ence in its original ord er to the feature extractio n stage. Feature Extract ion. W e extract a div erse set of featur es from each heartbeat in an E CG rec o rding. Specifically , we extract th e time since the last h eartbeat ( δ RR), the relative wa velet energy (R WE) over five fr equency band s, the to- tal wa velet ene rgy (TWE) over those fre quency band s, th e R amplitu d e, the Q a m plitude, QRS duration and wav elet entropy (WE). Previous works dem onstrated the efficac y of all of these featur e s in discrim inating cardiac arrh yth- mias f r om norm al heart rh ythms [14–18]. In add ition to the aforemen tioned featu res, we also train two SDAEs on the hea r tbeats in an un su pervised manne r with the goal of learning more nuanced dif ferences in morphology of in - dividual heartbeats. W e train one SD AE on the extracted heartbeats of the trainin g set and the other on their wav elet coefficients. W e then use th e encod ing side o f the SD AEs to extract low-dimension al embedd ings o f each heartbeat and each he artbeat’ s wa velet coefficients to b e used as ad- ditional input f e atures. Finally , we co ncatenate all ex- tracted f e a tures into a sing le feature vector per heartbeat and pass them to the le vel 1 mo dels in original heartbeat sequence order . Level 1 Models. W e build an ensemble of level 1 mod- els to classify th e sequence of per-beat feature vectors. T o increase th e diversity within ou r ensemble, we train RNNs in various binary classification settings and with different hyperp arameters. W e use RNNs with 1 - 5 rec urrent layers that consist of either Gated Recurrent Un its (GR U) [1 9] or Bidirectional Lon g Sho rt-T erm Memory (BLSTM ) units [20], fo llowed by an optional attention layer, 1 - 2 forward layers and a softm a x output layer . Additionally , we inf er a nonpa r ametric Hidd en Semi-Mar kov Mo del (HSMM) [21] with 64 initial states for each class in an u nsuperv ised set- ting. In total, our ensemble of le vel 1 m o dels consists of 15 RNNs and 4 H SMM s. W e concaten ate the ECG’ s nor- malised log-likelihoods under the per-class HSMMs an d the RNNs’ softmax outputs into a single prediction vector . W e pass the p r ediction vector of the lev el 1 models to the lev el 2 blend er mo d el. Level 2 Blender . W e use blen ding [22] to comb ine the prediction s of ou r le vel 1 mo d els and a set of ECG-wid e features in to a final per-class classification score. The ad- ditional featu res are the R WE and WE over the who le ECG and the absolu te average deviation (AAD) of th e WE and δ RR of all beats. W e emp loy a MLP with a softmax ou t- put layer as our le vel 2 blender model. In order to a v oid overfitting to the training set, we train the MLP on the val- idation set. Hyperparameter Selection. T o select the h y perpar am- eters of our lev el 1 RNNs, we p erforme d a g rid search o n the range o f 0 - 75 % for the dropout an d recu rrent dro pout percentag es, 60 - 5 12 for the number of u nits per h idden layer an d 1 - 5 f o r the number of recurrent layers. W e found that RNNs trained with 35% dropout, 65% recu r- rent drop out, 80 units p er hid den layer and 5 recurren t lay- ers (plus an add itio nal attention lay er) achieve consistently strong resu lts acr oss multip le binary classification settings. For ou r level 2 blen der mode l, we utilise Bayesian o ptimi- sation [23] to select the nu mber of layers, n umber of hid- den un its p er layer, dro pout an d num ber of trainin g ep ochs. W e perfo rm a 5-fold cross validation on the validation set to select th e b lender model’ s hyperp arameters. 2.1. Atte ntion over Heartbeats Attention mechanisms hav e been shown to allow for greater interp retability of neural networks in a variety of tasks in compute r vision and natural languag e proc e ss- ing [4–7]. In this w ork, we apply soft attention over the heartbeats co ntained in ECG signals in order to gain a deeper understandin g o f the dec ision -making pr ocess of our RNNs. Co n sider th e case of an RNN that is processing a sequen ce of T heartbeats. The topmo st r ecurrent layer Normal vs. all Normal: 94 % (s) a t ECG Other vs. all Other: 67 % (s) Figure 2. A visualisation of the attention values a t (top) of two different RNNs over two sample ECG recor dings (bo ttom). The graphs on top of the ECG recordin gs sh ow the attention values a t associated with each identified hear tbeat (dashe d line). Th e labels in the left and righ t corners of th e attention value g raphs show the settings the mo del was trained for and their classification co nfidence, respectively . The record ing on the left ( A02149 ) rep resents a no r mal sinus rhyth m . Due to the r egu lar hear t rhythm in the ECG, a distincti ve pattern of ap proxim a tely equ ally weighted atten tio n on e a ch heartb eat emerges from our RNN that was trained to distinguish between n ormal sinus rhy thms and all othe r types of rhythms. Th e recordin g on the right (A04661 ) is lab elled as an oth er arrh ythmia. The RNN trained to identify other arr h ythmias focu ses primarily on a sudd en, elo ngated pa use in the heart rhythm to decide that the reco r d is most likely an other arrhyth mia. outputs a hidden state h t at e very time step t ∈ [1 , T ] of the sequence. W e extend some of our RNNs w ith additive soft attention over the hidd en states h t to obtain a con text vector c that attenu ates the mo st informative hidden states h t of a heartbeat sequ ence. Based on the definition in [6], we use th e f ollowing set of equations: u t = tanh ( W beat h t + b beat ) (1) a t = sof tmax ( u T t u beat ) (2) c = X t a t h t (3) Where equation (1) is a single-layer MLP with a weight matrix W beat and bias b beat to o b tain u t as a hidd en rep- resentation of h t [6]. In equa tio n (2 ), we calculate the at- tention factors a t for each heartbeat by com puting a soft- max over the dot-prod uct similarities of every hea r tbeat’ s u t to the he a r tbeat co n text vector u beat . u beat correspo n ds to a h idden representatio n of the most in formative heart- beat [6]. W e jointly optimise W beat , b beat and u beat with the other RNN p arameters during trainin g. In figur e 2, we showcase two examples o f how qualitative analysis of the attention factors a t of equation ( 2 ) provides a deeper un- derstandin g of o ur RNNs’ decision mak ing. 3. Related W ork Our work builds on a long history of re sear ch in detect- ing cardiac ar rhythm ias from ECG re c ords by mak ing use of fe a tures that have b een shown to be h ighly discrim i- native in d istinguishing certain arr hythmias fro m normal heart r hythms [14 – 18]. Recently , Rajpurkar et al. pro- posed a 34-layer conv olutional neural network (CNN) to reach card io logist-level perfo rmance in classifying a large set of arrhythmias fr om mob ile car diac event recorder data [24]. In co ntrast, we achie ve state-of -the-art perfor m ance with significantly fewer trainable para meters by harn essing the natural heartbeat segmentation of ECGs and discrim- inativ e features from previous works. Additionally , we pay conside r ation to the fact that interp retability remains a challenge in apply ing mac h ine learning to the medical domain [25] b y extending our models with an attention mechanism that enables medical p rofessionals to reason about which heartbeats contributed most to the decision - making process of our RNNs. 4. Results and Conclusion W e present a m achine-lear ning appro a c h to distinguish- ing between multiple types o f h e a rt rhythm s. Our appr oach utilises an en semble of RNNs to join tly identify tempo r al and mor p holog ic a l p atterns in segmented ECG r ecording s of any leng th. In detail, our ap proach reaches an average F1 sco re of 0.79 on the p riv ate test set of the PhysioNet CinC Challenge 201 7 ( n = 3 , 658 ) with class-wise F1 scores of 0 . 90 , 0 . 79 and 0 . 68 f or norm al rhythms, AF and other arrhythm ias, respectively . On top of its state- of-the- art perfo rmance, o ur appro a c h main ta in s a h igh degree o f interpretab ility thro u gh the use of a soft attention mech- anism over hear tb eats. In the spirit of ope n researc h, we make a n impleme n tation of our cardiac rhythm c lassifica- tion system a vailable through the Phy sioNet 2017 Open Source Challenge. Future W ork. Based on our discussion s with a car- diologist, we hyp othesise that the accur acy of our mod- els could be fur ther improved by in corpo r ating contextual informa tio n, such as demogra p hic inf ormation , d ata fr om other clinical assessments a n d be h avioral aspects. Acknowledgeme nts This work was partially fu nded by the Swiss National Science Foundation (SNSF) p roject No . 1673 02 within the National Research Pro gram (NRP) 75 “Big Data” and SNSF pro je c t No. 1 5 0640 . W e thank Prof. Dr . med. Fi- rat Duru for providing valuable insights into the d ecision- making process of cardiologists. Refer ences [1] B all J, Carrington MJ, McMurray JJ, S tew art S. Atri al fib- rillation: Profile and burd en of an e vo lving epidemic i n the 21st century . International Journal of C ardiology 2013; 167(5):1807 –1824 . [2] C amm AJ, Kirchhof P , Lip GY , Schotten U, Sa velie va I, Ernst S, V an Gelder IC, Al-Att ar N, Hindricks G, Prender - gast B, et al. Guidelines for the management of atrial fi bril- lation. European Heart Journal 2010;31:2369 –2429 . [3] Vin cent P , Larochelle H, Lajoie I, Bengio Y , Manzag ol P A. Stacked denoising autoencoders: Learning useful representations in a deep network with a local deno ising criterion. Journal of Machine Learning Research 201 0; 11(Dec):3371– 3408. [4] B ahdanau D, Cho K, Bengio Y . Neural machine translation by jointly learning to align and translate. In International Conference on Learning Representations, 2015 . [5] Xu K, Ba J, Kiros R , Cho K, Courville A, Salakhudino v R , Zemel R, B engio Y . Show , attend and tell : Neural image caption generation with visual attention. In International Conference on Machine Learning. 2015; 2048– 2057. [6] Y ang Z, Y ang D, Dyer C, He X, Smola AJ, Hovy EH. Hi- erarchical attention networks for document classification. In Conference of the North American Chapter of the As- sociation for Computational Linguistics: Human L anguage T echnologies. 2016; 1480–1489. [7] Z hang Z, Xie Y , Xing F , McGough M, Y ang L. MD- Net: A Semantically and V isually Interpretable Medical Image Diagnosis Network. In International Conference on Computer V ision and Pattern Recognition, arXi v preprint arXiv :1707.0248 5, 2017. [8] C l ifford GD, L i u CY , Moody B, Lehman L, S ilva I, Li Q, Johnson AEW , Mark RG. AF classification fr om a short single lead ECG recording: The Physione t Computing in Cardiology Challenge 2017. In Computing in Cardiology , 2017. [9] Goldberger AL, Amaral LAN, Glass L, Hausdorf f JM , Iv anov PC, Mark RG, Mietus JE , Moody GB, Peng CK, Stanley HE. PhysioBank, PhysioT oolkit, and PhysioNet: Components of a New Research Resourc e for Complex Physiologic Signals. Circulation 2000;101(2 3):e215–e2 20. [10] Moody GB, Mark RG. The impact of the MIT -BIH arrhyth- mia database. IEEE Engineering in Medicine and Biology Magazine 2001;20(3):45 –50. [11] Moody G. A new method for detecting atrial fi bril lation us- ing RR intervals. In Computers in Cardiology . IEEE, 1983; 227–23 0. [12] Greenwald SD, Patil RS, Mark RG. Improv ed detection and classification of arrhythmias in noise-corrupted electrocar- diograms using contextual information. In Computers i n Cardiology . IEEE, 1990; 461–464. [13] Pan J, T ompkins WJ. A real-time QRS detection algo- rithm. IEEE Tra nsactions on Biomedical Engineering 1985; 3:230–23 6. [14] Sarkar S, Ritscher D, Mehra R. A detector for a chronic implantable atrial tachyarrhythmia monitor . IEEE T ransac- tions on Biomedical Engineering 2008 ;55(3):1219– 1224. [15] T ateno K, Gl ass L. Automatic detection of atri al fibrillati on using the coefficient of v ariation and density histograms of RR and δ RR i ntervals. Medical and B iological Engineering and Computing 2001;39(6):664–6 71. [16] Garc´ ıa M, R ´ odenas J, Alcaraz R, Rieta JJ. Application of the relative wav elet energy to heart rate independent detec- tion of atrial fi brillation. computer methods and programs in biomedicine 2016;131:157 –168. [17] R ´ odenas J, Garc´ ıa M, Alcaraz R , Rieta JJ. W avelet entropy automatically detects episodes of atrial fibrillation from single-lead electrocardiograms. Entropy 2015;17(9):6179– 6199. [18] Alcaraz R, V ay ´ a C, Cervig ´ on R, S ´ anchez C , Rieta J. W avelet sample entrop y: A ne w approach to predict t er- mination of atrial fibrillation. In Computers in Cardiology . IEEE, 2006; 597–600. [19] Chung J, Gulcehre C, Cho K, Bengio Y . Empirical ev alua- tion of gated recurrent neural networks on sequence model- ing. In Neural Information Processing Systems, W orkshop on Deep Learning, arXiv preprint arXiv:14 12.3555 , 2014. [20] Graves A, Jaitly N, Moham ed A r. Hybrid spee ch recog- nition wi t h deep bidirectional lstm. In Automatic Speech Recognition and Understanding , IEEE W orkshop on. IEEE , 2013; 273–278. [21] Johnson MJ, Willsky AS. Bayesian nonparametric hidden semi-marko v models. Journal of Machine L earning Re- search 2013;14(Feb):673–7 01. [22] W olpert DH. Stacked generalization. Neural network s 1992;5(2):24 1–259. [23] Bergstra J, Y amins D , Cox DD. Hyperopt: A python library for optimizing the hyperparameters of machine learning al- gorithms. In Proceedings of the 12 th Python in Science Conference. 2013; 13–20. [24] Rajpurkar P , Hannun A Y , Haghpanah i M, Bourn C, Ng A Y . Cardiologist-le vel arrhythmia detection with con volutional neural networks. arXi v preprint, arXiv:17 07.0183 6, 2017. [25] Cabitza F , Rasoini R, Gensini G. Unintended consequen ces of machine learning i n medicine. Journal of the American Medical Association 2017;318(6):51 7–518. Address for correspondenc e: Patrick Schwab, ETH Zurich Balgrist Campus, BAA D, L engghalde 5, 8092 Zurich patrick.schwab @hest.ethz.ch
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment