Multivariate analysis of Brillouin imaging data by supervised and unsupervised learning
💡 Research Summary
This paper addresses the challenge of extracting reliable mechanical parameters from Brillouin imaging (BI) datasets, which are inherently hyperspectral and often suffer from low signal‑to‑noise ratios (SNR) and spectral mixing in complex biological samples. The conventional workflow fits a single Brillouin peak in each spectrum using least‑squares methods to obtain average shift and linewidth values. While straightforward, this approach requires high SNR, is computationally intensive for large images (up to 10⁴ spectra), and can produce biased results when multiple tissue components contribute to a single voxel.
The authors propose a comprehensive suite of multivariate analysis (MVA) techniques—both unsupervised and supervised—to replace or augment traditional peak fitting. After standard preprocessing (background subtraction, standard normal variate normalization, and denoising), they explore four main algorithm families:
-
Principal Component Analysis (PCA) – An unsupervised linear dimensionality‑reduction method that rotates the data into orthogonal components capturing maximal variance. PCA effectively denoises the data by discarding high‑order components dominated by noise, but it does not directly identify pure spectral endmembers, limiting its utility for spectral unmixing.
-
Vertex Component Analysis (VCA) – A geometrical unmixing algorithm that assumes the hyperspectral data occupy a simplex in spectral space, with pure endmember spectra located at the simplex vertices. By fitting either a maximum‑volume or minimum‑volume simplex, VCA can recover endmember spectra even when no pure pixel exists in the measurement. The method is computationally faster than iterative factor‑analysis approaches and, as demonstrated, yields more accurate abundance maps than PCA, especially in mixtures with several components.
-
Linear Discriminant Analysis (LDA) – A supervised classifier that maximizes inter‑class variance while minimizing intra‑class variance. Because Brillouin spectra are often singular, the authors first project the data onto a reduced space using PCA or VCA, then apply LDA to obtain discriminant functions. With modest training data, LDA successfully labels each pixel according to predefined tissue types, enabling rapid diagnostic classification.
-
Hierarchical Cluster Analysis (HCA) – An unsupervised agglomerative clustering technique that builds a dendrogram based on pairwise distances (Ward’s method) and cuts it at a chosen number of clusters. HCA provides fully automatic segmentation without prior labeling, though it requires prior dimensionality reduction to keep the O(N²) distance computation tractable.
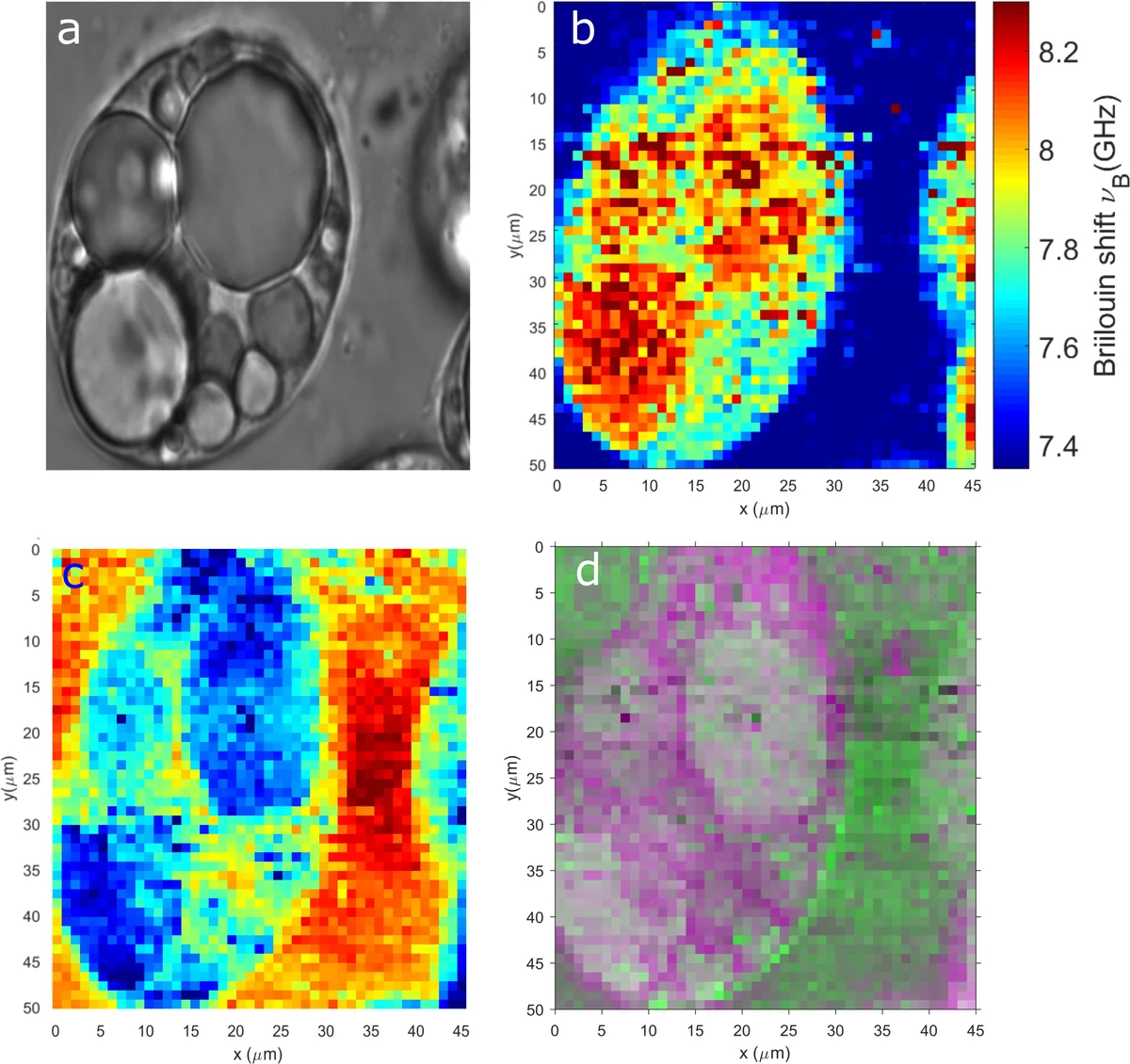
The paper validates these methods on two experimental systems: (i) a synthetic phantom composed of water and glycerol mixtures, where the known composition serves as ground truth; and (ii) live 3T3‑L1 adipocytes, a biologically relevant model with distinct subcellular compartments (cytoplasm, nucleus, extracellular matrix). In the phantom, VCA recovered the two endmember spectra with a mean‑square error ~5 % lower than PCA‑based unmixing. In the cellular data, VCA‑derived abundance maps clearly delineated subcellular regions that appeared blurred in conventional Brillouin images. LDA, trained on a small set of manually labeled spectra, achieved high classification accuracy and demonstrated the potential for disease‑state discrimination. HCA automatically segmented the cells into meaningful clusters that matched known morphological features.
A key result is the dramatic speed improvement: while traditional pixel‑wise fitting required tens of seconds to minutes per image, the combined MVA pipeline processed the entire dataset in 0.1–0.5 seconds, representing roughly a 100‑fold acceleration. This speed gain opens the possibility of near‑real‑time Brillouin imaging, essential for in‑vivo or intra‑operative applications.
In conclusion, the study shows that multivariate analysis—particularly VCA for spectral unmixing, LDA for supervised classification, and HCA for unsupervised segmentation—substantially enhances image contrast, provides quantitative abundance information, and reduces computational burden compared with standard peak fitting. The authors suggest future extensions involving nonlinear deep‑learning models to capture more complex mixing phenomena and multimodal data fusion (e.g., Raman, CARS) to further enrich the diagnostic capability of Brillouin imaging.
Comments & Academic Discussion
Loading comments...
Leave a Comment