Medical Image Registration Using Deep Neural Networks: A Comprehensive Review
Image-guided interventions are saving the lives of a large number of patients where the image registration problem should indeed be considered as the most complex and complicated issue to be tackled. On the other hand, the recently huge progress in t…
Authors: Hamid Reza Boveiri, Raouf Khayami, Reza Javidan
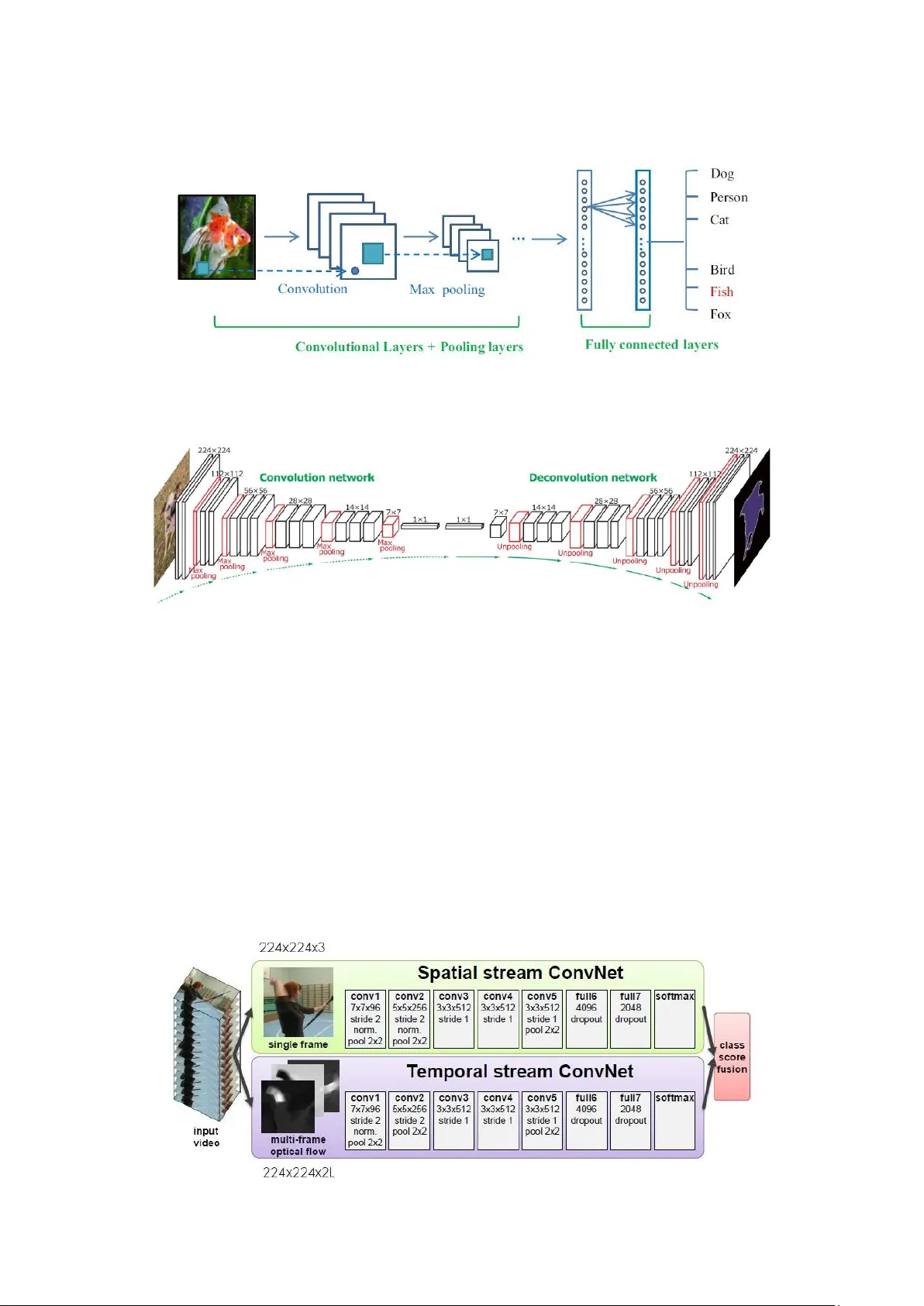
- 1 - Medical I m a ge Registratio n Using Deep Neural Net works: A Comprehensive Revie w Hamid Reza Boveiri a ,* , R aouf Kh ayami a , R e za Ja vi d a n a , Ali Re za MehdiZadeh b a Depar tment of Com puter Eng ineering and I T, Shiraz Univ ersity of Techno logy , Shi raz , Iran b Depa rtment of Med ical Phy sics and Eng i neer ing, Shiraz U niver sity of Medi cal Scienc e, Sh iraz, I ran * Correspond ing A uthor Ema il: hr. bov ei ri @sutech. ac.ir Abstract — I mage-guide d inte r ve n tio n s are sav ing the live s of a large n u mber o f pati e n ts w h e r e the image r egis tr ation prob lem should indee d be considered as the most complex and co m plicated issue to b e tackled. On the o ther hand, the recent ly h uge prog ress in t he field of m ac hine le arning m ade by th e pos sibility of impleme n ting dee p neural netwo r ks o n the co nt empo rary man y - co r e GPUs o p ened up a promis ing w in dow t o challenge w ith many m edic al applic ations, w her e the r egis tration is not an exce pt ion. In thi s pape r, a compre h ensiv e revie w on the state- of -the-art li te rature know n as medical image regist ratio n using dee p n eu ral netwo r ks is pre sented. The r ev iew i s sy stematic and enco mpasses all the related w orks p r ev iously published in the f ield . Key conce pts, statis tical a nalys is from dif fe ren t po ints of view , confiding challe n ges, n ov elties and main co n trib utio n s, key - en abli n g te chniques, future directio n s and prospe ctive tre n ds all are discuss ed and survey ed in deta ils in this co mpre h ensiv e revie w . T h is r ev iew allow s a dee p understa nding and ins ight fo r the reade r s active in the field who are inves tigating t h e st at e- of-the-art a nd see kin g to co ntri bute th e futu re literatu r e. Keywords — Co n vo lutional Neu ral Netw ork (CNN) ; Deep Le arning; De ep Re in fo r cement Lea rning; Def ormable Re gistr atio n; Ge n e r ative Adve rsari al Netw ork (GAN) ; Im age -guided Interve n tio n; Medica l Im age Registratio n; One- shot Re gistration; Staked Auto-Enco ders (SAEs). ——————— —————— —————— — ♦ —— ————— ————————— ———— 1. Intr oduction In most medical interventio n s , ther e are a n umb er of cases i n which so me images need t o be captured fo r diag n o sis, prog n os is, treatment an d f ollow-up purpos es. These images can be var y in terms of temporal, spat ial, dime nsional or modula r. Ima ge fus ion causing in fo r matio n sy n ergy can h ave a sig nifica n t cont r ibutio n to guide and suppo r t phy sician s in the proc ess of decisio n m aki n g, mostly in online and r e al -time fasion . Lack of a lignment is unavo i dable fo r th es e images taken in d ifferent t imes and co n ditio n s; h e n ce , ca n c ha lle n ge the q uality and acc u r acy of th e subs equent analy ses. Image r egistratio n is th e p r oc ess of a ligning two (or mo r e) give n im ages based on an identica l geo m etrical co ordinatio n sy stem. Th e aim i s at f indin g an o ptimum sp atial tra n sf ormation th at registe r s the structures- of -interest in th e bes t way . This prob l em i s important in numerous way s in the fie ld o f machine vision e.g. for r e m o te sensing, objec t traci n g , sa tellite imaging and so on (Gos htasby 2017) . Image r egist ration is also fu n dame ntal to the image -guided interve ntion w her e e.g. telesu r ger y , i mage-guided radiot h e r apy (IGR T), and precision medicine ca nn ot b e o pera tio nal without using image r eg istratio n techniques (Peters & Cleary 2008) . To exemplify , in IGR T, a pre-inte r ventio nal im age (typic all y hi gh -qu ality 3D image), on w hich the treatme nt pla nning is co n duc ted, needs to be r egiste r ed on an ope r atio nal image (ty pically l ow-quality and noisy 2D) s o that the lin ea r acce lerator (li n ac) machine can be calibrated, and th e radiatio n fragment can be c o n ducted wit h ma xim al precisio n and m inim al ri sk o f radiatio n to the adjace nt healt hy organs refe r red to as mi n im all y in vasiv e proc edure . In this proce ss, the chal lenges li ke diff er ent modalities of inputted images, low -qual ity and n oise of i nt e rventio nal images , defo rm atio n o f abdo m inal cavity ’s o r gans (b ecause of t he spo n taneous c ontractio n/in f latio n ), mo ve ment of th o ra x cav it y ’s o r gans (b ec ause of th e re spi r atio n a nd heartb eats), c hanging t h e size o f organs and reg ions- of -in terest (R oIs) due to th e w eight loss /gain duri ng the treatme n t proce ss can co mpromise s th e quality o f solving the p r ob l em. In practice , spec ial co n side rations s h ould be taken int o acco unt, and o ther image proce ssing tec hn iques n ee d to collab orate, w h ich makes t h e issue very challenging and co m plic ated (H ajnal 20 01). Bas icall y , co n ve n tional image r egis tration is an i terativ e optimizatio n p r oc ess th at r equi r es e xtra ct ing prope r f eatures, sele ctin g a simil arity measure (to ev al uate the r egist ration quality ), choo sing the mode l of tr ansfo r mat ion , and finally , a mechanism to inve stigate the sea rch sp ace (Olive ira & Tav ares 2014). As illustrated in Fig. 1, a co uple o f im ages are inputted to the sy stem, o f w h ic h one is co n sidered as fixe d and t h e o ther as mov in g image . The optim al alig nment ca n be - 2 - achiev ed via iterative l y sliding the mov in g image over the fixe d i mage. A t fi rst, the c onsidered simi larity m easu r e identifies th e amou n t of corresponde nce be tween the inputt ed images. A n optimiza tion algo rithm, using an u pdating mechanism , calculates th e new tran sf ormatio n’s paramete r s . Appling these paramete r s o n the mo ving image leads a new suppo sedly be tter -aligned im age. If t h e te rminatio n criteria a r e sa tisfied, t he algo ri thm i s te rminate d, else a new iteratio n should b e s tarted. In eac h iteratio n , t h e moving image ge t a b etter c orrespo n de n ce with t h e fixe d im age, and t h e it e r atio n s co nt inue unti l no f urther regist ration can b e ac h iev ed, or some predef in ed c ri te r ia are satisf i ed. T h e sy stem output can b e either the transf ormation para meters o r fi nal interpo l ated fuse d i mage . Figure 1: The wo rkflow of c onv ent ional imag e reg istra ti on techn i ques based o n optim izatio n proc e dures There a r e tw o main drawb acks to this st r ategy as follows : This iterative ma nn e r is ve r y slow w her e r untime s i n the te n s of min u tes a r e n o rm f or c ommon def ormable image registratio n technique s eve n with a n e f ficie n t impleme ntation on the conte mporary GPUs (like a NVIDIA TitanX) ; w hile th e p ractical use in cli n ical o perations is real-ti me, and such a prolo n ged wasting time is n ot appreciated . Mo st simi la rity m easures have a lots of loc al optima around t h e g lob al one, spec ially where deali n g wit h images from dif fe r ent mo dalities (ref err ed to as multi modal im age regist ration); they lose th ei r ef ficiency c ausin g prematu r e co n ve r gence o r stag n ation w hich are tw o prevalent co n f inin g dilemmas in the optim ization field . Acco rdin g ly , t o circumvent these two confining p r ob l ems, lea rning-based regis tratio n approac h es have gained increasing po pularity i n the r ec ent y ear s; me anwhile, deep n eu ral netw orks (DNNs ) , as o n e o f the m o st pow er f ul technique s ev er see n by th e co mmunity of machi ne int el ligence, have be en a pplie d t o the various image proc ess in g applicat ions. Of c ourse, medical image registratio n is not an exc eption, and a numbe r of deep learning base d approac h es were propo sed i n th e lite r atu r e; h ow e v er, the number of wor ks an d the used t ec hniques is ve r y lim ited, an d th e re is a promisi n g potential fo r inve stigation (Litjens e t al. 2017). In th is pape r , a co mprehensive sy stematic review o n th e medical image r egis tr atio n using dee p n eu r al n etw orks is presented. We ha v e gath e r ed all the r elev ant st ate- of -th e-art s, from the first one in 2013 up the last one in 2019. Th e wo r ks ar e analy ses based on the dif ferent statist ical perspec tive s an d me asures of in te r est e.g. y ear s, pub lication titles , public at io n ty pes, pub lishe r s, aut h o r s (t o tal n u mber of publicatio ns and cit ati o n s), key wo r ds, t ec hnique s, co m parison metrics, datasets, orga n s of int e rest, and mo da lities. The app roaches are ca tego r ized to the t hree majo r gene r atio n s base d on t h e breakthroug hs af fec tin g t he cont ri butio n s. Also , the se m inal w orks in eac h catego ry an d ge n e r atio n are int roduce d and analy z ed in dept h, and key co ntr ibutio n s and philo sophie s b ehind th e m are p r ese n ted in det ails. Finally , there is a discuss ion introduc ing confi ning c hallenges, open p r ob l ems a nd prospe ctive direc ti o n s. This rev iew all o ws a deep No Fixed Image Moving Image Similarity M easure Cal culation Parameter Up dating Tra nsformation Applyi ng Terminati on? Fused Image Tra nsform Parameter s - 3 - understa n di ng and insig h t fo r the r e aders activ e i n the fiel d who ar e inve stigating the st ate - of -th e-a rt and seeki n g t o co nt ribute th e f uture lite rature. Re st of th e paper is orga n ized as fo l lows . In the n ext Se ction, r ef erence gath e rin g methodo l ogy a nd the challenge d fac ed ar e de scribe d. Se ction 3 an d 4 are devo t ed to t he t axonomy of me dical image r egist ration, a nd t h e prob lem formula ti o n, r espe ctive l y . Th e arc h itec tures of dee p n eural networks use d in the lite r ature are inve stigated in Sec tion 5 . Sec tion 6 i s an in -dept h l ite rature r ev iew on semi n al wo r ks. Statistica l analy sis on th e st at e- of -the-art is prese nted in Sec tion 7 . Sec tion 8 is devo ted to the disc ussio n on the co n fining c hallenges a nd open p r ob l ems i n the field. A nd fi nally, the co n clusio n and futu r e t rends a r e prese nted in the last Sect ion. 2. Refe re n ce Gatherin g Met h odology To inves tigate the lite r atu r e, the sy stematic sea r c h w as co n d ucted on t w o ma jo r scientific datab ases i. e. “ SCOP US ” and “ Pub Med ” (since the topic is multidisciplina ry be t w een enginee r ing and medicine) . “Me dical Image Regist ra tion” w as the main key wo r d sea r c hed acco mpanied wit h one of the fo llowings as th e unde rly in g technique i .e. “Dee p Learni n g,” “Dee p Neural Ne two rk, ” o r “Convo lutional.” Mo r eo ver, anothe r sea rch w as co n ducted usi ng the af o r ementio n ed keyw ords in majo r scient ific datasets i.e. “Go ogle Sc h olar” and “C r ossRef ” t o validate th e co mpr e h e n siveness of t he results. T h e se ar c h es w er e restricted to the Title, Ab stract and Keywo r ds whenev er it w as feasib le. A total n u mber of 25 refe r ences we r e detec ted, am o n g them, 6 cases was irr elev ant (e.g . just the r es ult s we r e compa r ed to the deep l ea rning approac h es) a nd 21 case s was c ompletely mat c h ed w ith our cri te ria. At this po int, w e detecte d that some r ef eren ce s w ere ov er looked, and that these keyw ords are insuf fic ient to co nduct a co mpre h ensive sea r ch; fo r example, there w as a large numb er of confe r ence papers wit h n o “Dee p Le arn ing,” “Dee p Neural N etw ork,” o r “ Convo l utional” i n t h e Title, Ab str act or Ke ywo r ds, but e.g. st art w i th CNN , SA Es or GAN w ithout def in ing t h ese abb reviations . Additio nally , we found t h at there a re a numb er of r ef erences that use e.g. “Im age Re gistr atio n,” “I mage Correspo n dence ,” “P ose E sti mat ion” etc. i n stead of o ur main keyw ord “Me dical Image Re gistr atio n, ” but the paper is complete ly w ith in th e sco pe of t he r ev iew. Such case was t rue fo r our backup sear c h in “Goo gl e S cholar” and “Cros sR ef.” On this basis, with a c ompr e h en s ive li st of keywo rds extr acted from the already detec ted r ef erences, we conducted a co m prehe n sive sy stematic searc h in the “SCOPU S” an d “Pub Me d” as w ell as an exhaustiv e ad- hoc sea r c h in “Go o gle Schola r ” an d “Cross Ref .” We also r ev iewe d all the citatio n s and refe r ences of all the selecte d pape r s, an d sea r ched wit h in the jo urn als/co n fere nces in which the sele cted papers we re publis h ed. Totally , we r ev iewe d mo r e th an 500 pote n tial pape r s, and a mo n g t hem w e co uld ext ract a c omple te list o f 80 papers that co nstitute the sta te- of -th e-a rt lite r atu r e. These r ef erences along with their u nderly in g t ec hniques, datasets, ev aluatio n metric s and other rel ated info r matio n are listed in the Tab le 1. In this reg ards, w e fac ed fo ur determinat ive ch alle nges t o constitu te this sy stemat ic rev iew as fo llows: Routi n ely , we we r e h ighly c o n ce rn ed not to overloo k an y work in o rd er to draw justif iable co n clusions, w h ere this topic is in its infa ncy , and the numb er of r elated w o r ks is limited. Seco ndl y , ther e w as not an y a pplicat ion/sof tware to analy ze such dat a to suppo rt the aut h o r s of syste ma tic review s, an d w e were fo r ce d t o develop a simple deskto p applicatio n , and use embe dded-SQL t o dra w our statistical analy sis. Thirdly , w e ar e not ab l e to co n duct any expe r ime n tal b enchmark rev iew o n this to pic bec ause, as we hav e already co n tacted mo st of th e aut h o r s, this t o pic is rel ated to the medica l scie n ce where mos t of the utili zed datase ts ar e p r ivate, and the imple mentatio ns have co p y ri gh t, and cannot be a cqui r ed by us. Finally and mos t impo rtantly , we we r e highly c o n ce rn ed abo ut the soundness and usef ulness of o ur asserts and co nclusions spec ially fro m th e m edica l/cli n ical point of view . Fortunately , our co-Autho r Dr. MehdiZade h, the asso ciate Prof esso r and Dean of the Departme nt of Medical Ph y sics an d Engineering, SU MS, has be en with us in all the p hases, a nd meticulous ly read, revise d, a n d c e r tified all t h e d iscuss ions and co n clus i ons t o be so un d and valuab le from t h e medica l/ cl inical pe r spec tive. - 4 - Table 1 : Founded ref erences along with thei r used techniqu e s, da t asets and rel ated ev aluatio n metric s Au th or & Year Reference T ec hnique Dataset (O rga n) Metrics Description Wu et al. (2013) G. Wu, M. Kim, Q. Wan g, Y. Gao, S. Liao , and D. Shen, “Unsupervised Deep Fe ature Learning for De formable Registration of MR Brain I mages,” Lecture Notes in Computer Science, pp. 649 – 656, 2 013. CNN IXI - T1 and T2- w ei ghted MRI (Adult Brain) Dice Using hybrid ISA and CNN to automatically extract featu res and feed them to HAMME R for final deformable reg i stration ANDI - MRI (Adult Br ain) Zhao and Jia (2015) L. Zhao and K. Jia, “Deep Adaptive Log -De mons: Diffeo morphic Image Registration with Ve ry Large Deform ations,” Computational and M athematical Methods i n Medicine, vol. 2015, pp. 1 – 16, 2015. CNN BrainWeb – MRI (Brain) Dice Using CNN to a uto matically extract features and fee d them to Demons for final reg i stration Empire 10 – 3D CT (Lu ng) Cheng et al. (2016) X. Cheng, L. Zhan g, and Y . Zheng, “Deep si milarity learnin g for mu lt imodal medi cal i mages,” C ompu te r Methods in Biomechanics and Bio med i cal Engineeri ng : Imaging & Visualization, vol. 6, no. 3, pp. 248 – 252, Apr. 2018. (Ac cepted & Onlin e from 201 6) SA Es CT and MRI (Head) - Multimodal CSPE Using SAEs as a multi modal similarity for rigid regis tration. Yang e t al. (2016) X. Yang, X. Han, E. P ark, S. Aylward, R. Kwitt, and M. Niethammer, “Re g ist ration of Patholo g ical Im ages,” Lecture Notes in Computer Scien ce, pp. 97 – 107, 2016. SA Es OASIS – MRI (B rain ) TRE and Deformation Error Directly re gr essin g the unimodal deform ab le reg i stration p arameters via a convolutional S AE. BRATS – MRI (Brain) Yang e t al. (2016) X. Yang, R. Kwitt, an d M. Nie thammer, “Fast Predictive Image Registration,” Lecture Notes in Computer S cienc e , pp. 48 – 57, 2016. CNN OASIS – MRI (B rain ) Deformation Error Directly re gr essin g the deformable re g ist ration parameters via h ybrid C NN and LDDMM Wu et al. (2016) G. Wu, M. Kim, Q. Wan g, B. C. Munsell, and D. She n, “Scalable High-Perform anc e I mage Registration Frame w or k by Unsupervised Deep Fe ature Representations Learning, ” IEEE Transactions on Bio med i cal Engineering, vol. 6 3, no. 7, pp. 1505 – 1516, Jul. 2016. SA Es LONI - MRI (Adult B rain) Dice Using SAEs to extra c t features and feed them to a traditi onal approaches for final deformable re g ist ration ANDI - MRI (Adult Br ain) Simonovsky et a l . (2016) M. Simonovsky, B. Gutiérrez-Becker, D. Ma teus, N. N avab, and N. Komodakis, “A Dee p Metri c for M ultimodal Re g istratio n,” Medical Image Computi ng and Compute r-Assisted Interve ntion -- MICCAI 2016, pp. 10 – 18, 2016. CNN IXI - T1 and T2- w ei ghted MRI (Adult s’ Br ain) Dice and Jaccard Using CNN as a multimod al similarity measure to guide conventional iter a ti ve approaches ALBERTs - T1 and T 2- weighted MRI (Neonatal s’ Brain) Miao et al. (2016) S. Miao, Z. J. W a ng, Y. Zheng, and R. Liao, “Real -time 2D/3D reg i stration vi a CNN regression,” 201 6 IEEE 13th International Symposium on Biomedical Im aging (ISBI), Ap r. 2016. CNN XEF (X-ra y Echo Fu sion) mTREproj Directly re gr essin g the rigid reg i stration p arameters using CNN based on the implante d par ti cles VIPS (Visual Impl an t Planning System) TKA (Total Knee Arthroplasty) Miao et al. (2016) S. Miao, Z. J. W a ng, an d R. Liao, “A CNN Regression Approach for Real- Time 2D/3D Re gistration,” IEEE Tr ansactions on Medical Imaging, vol. 3 5, no. 5, pp. 1352 – 1 363, M ay 2 016. CNN XEF (X-ra y Echo Fu sion) mTREproj Directly re gr essin g the rigid reg i stration p arameters using CNN based on the implante d par ti cles VIPS (Visual Impl an t Planning System) TKA (Total Knee Arthroplasty) Liao et al. (2017) Rui Liao, Shun Mi ao, P ie rre de Tournemire , Sasa Grbic, Ali Kamen, Tommaso M ansi, and Dorin Com an ici u. “An Artificial Agent for Robust Image Regist ration,” in AAAI , pp. 4168-4 175, 2017. CNN E1 – CT and CBCT (Spine) TRE and MME Using reinforcement le arning to train CNN to be used as ag e nts for approxim ating Affine registration’s parameters in an iter ative manner E2 – CT and CBCT (Cardiac) Krebs et al. (2017) J. Krebs, T. Mansi, H. Delingette, L. Zhang, F. C. Ghesu, S. Miao, A. K. Maier, N. Ayache, R . Liao, and A. Kamen, “Robust Non - rig i d Registr ation T hrough Agent - Based Action Le arning,” Lecture Notes in Comp uter Science, pp. 344 – 3 52, 201 7. CNN PROMISE12 – MRI – Prostate Dice and Hausdorff Using reinforcement le arning to train CNN to be used as ag e nts for approxim ating Affine registration’s parameters in an iter ative manner Prostate-3T – MRI – (Prostate) Yang e t al. (2017) X. Yang, R. Kwitt, M. St yner, and M. Nietham mer, “Fast pr e dictive multim odal image registr ati on,” 2017 IEEE 14th International Symposiu m on Biomedical Imagin g (ISBI 2017), Apr. 2017. CNN IBIS 3D Autism – TI and T2 MRI (multimodal) (Brain) SSD Directly re gr essin g the deformable re g ist ration parameters via h ybrid C NN and LDDMM Wang et al. (2017) S. Wang, M. Kim, G . Wu, and D. She n, “ S calable High Performance Im ag e Re g i stration Frame work by Unsupervised Deep Feature Represent a ti ons Learnin g,” Deep Learning for Medical Image An alysis, pp. 245 – 269, 2017. SA Es LONI - MRI (Adult B rain) Dice Using SAEs to extra c t features and feed them to co nventional approaches for final deformable re g ist ration ANDI - MRI (Adult Br ain) Miao et al. (2017) S. Miao, J. Z. W a ng, an d R. Liao, “Convolutional Neural Networks for Robust and Real-Time 2 -D/3- D Registr ation, ” Deep Learning for Medical Im age Analysis, pp. 271 – 296, 20 17 . CNN XEF (X-ra y Echo Fu sion) mTREproj Directly re gr essin g the rigid reg i stration p arameters using CNN based on the i mp l anted par ti cles VIPS (Visual Impl an t Planning System) TKA (Total Knee Arthroplasty) Sokooti et al. (2017) H. Sokooti, B. de Vos, F. Berendsen, B. P. F. Lelievel dt, I. Išgum, and M . Starin g , “Nonrigid Im ag e Re g i stration Using M ulti-scale 3D Convolution al Ne ural Networks,” Lectu re Notes in Computer Science, pp. 232 – 239, 2 017. CNN SPREAD - 3D CT (Che st) MAE and TRE Directly re gr essin g the defprmable re g ist ration parameters using CNN de Vos et al. (2017) B. D. de Vos, F. F. Be rendsen, M. A. Viergever, M. St aring, and I. Išgum, “End - to - End Unsupe rvised Deformable Ima ge CNN Sunnybrook Cardi ac D ata – 3D MRI (Cardia c) Dice and MAD Directly re gr essin g the deformable re g ist ration parameters using CNN - 5 - Registration with a Convol utional Neur al Net work,” Lecture Notes in Computer Scien ce, pp. 204 – 2 12, 2017. Bhatia et al. (2017) P. S. Bhatia, F. Reda, M. Hard e r, Y. Zh an , and X. S. Zh ou , “Real time coarse orientation dete ction in MR s can s u sing multi -planar deep convolution al neural networks,” Medical Im ag i ng 2017: Image Processing, Feb. 2 017. CNN Private – MRI (Elbo w) Accuracy Directly re gr essin g the unimodal rigid re g i stration parameters using CNN Zh e ng et al. (2017) J. Zh e ng, S. Mi ao, and R. Liao, “Learning CNNs w it h Pairwise Domain Adaption for Re al-Time 6D oF Ult rasound Transd uc e r Detection and Tr ack i ng from X- R ay Imag e s,” Medi cal Image Computing and Compute r- Assisted Interve ntion − MICCAI 2 017, pp. 646 – 654, 2017. CNN Private – X-Ray and TEE (Transducer) Projected TRE Directly re gr essin g the mu lt imodal ri g i d registration parameters (pose) of TEE Transducer using CNN Pei et al. (2017) Y. Pei, Y. Zhang, H. Qin, G. Ma, Y. Guo, T. X u, and H. Zha, “Non -rigid Craniofacial 2D -3D Registr ation Using CNN -Based Regression,” Lecture Notes in Computer Science, pp. 117– 125, 2017. CNN NewTom – CBCT (Craniofacial) MCD and MID Directly re gr essin g the unimodal deform ab le CT and X-Ray registration p arameters using CNN Eppenhof and Pluim (2017) K. A. J. Eppenho f and J. P. W. Plui m, “Supervised lo cal e rror estimation for nonlinear ima ge registration usin g convolutional neural networks,” Medic al I maging 2017: Im ag e Processin g , Feb . 2017. CNN Private – 2D DSA (Br ain) RMSD, NRMSD and PCC Proposing a supervise d method for the estim a ti on of the unimodal registr ation e rror map for deformable i mag e reg i stration usin g CNN DIRLAB – 3D CT ( Lung ) Ghosal and Ray (2017) S. Ghosal and N. Ray, “Dee p deformable re g ist ration: Enhancing accuracy by fully con vol utional neural net,” Patte rn Recognition Letters, vol. 94, pp. 81 – 8 6, Jul. 2017. CNN (VGG-net) (Simonyan and Zisserman 2015) IXI - T1 and T2- w ei ghted MRI (Adults’ Brain) SSIM, PSNR, and SSD Learn a CNN to work as S SD as a n e w uni modal similarity metrics to work with any conventio nal deformable registr ation method. ANDI - MRI (Adult Br ain) Ma et al. (2017) K. Ma, J. Wang, V. Sin gh, B. Tamerso y , Y. -J. Ch ang, A. Wimmer, and T. Che n, “ M ultimod al Ima ge Registration with Deep Context Reinfo rc e ment Le arning,” Lecture Notes in Computer Scien ce, pp. 240 – 248, 2017. DRL (DuelingNet) (Wang et al. 2016) ABD – depth and CT (Chest and Abdomen) Hausdorff Directly re gr essin g the rigid mu lt imodal registr a ti on parameters using Deep Reinforcement Learning (DRL) Salehi et al. (2017) M. Salehi, R. Prevost, J.- L. Moctezu ma, N. Navab, and W. Wein, “Precise Ultrasound Bone Registration with Le arn i ng -Based Segm e ntation and Speed of Sound Calibr ation,” Medical Image Computing and Compute r- Assisted Interve ntion − MICCAI 2017, pp. 682 – 690, 2017. CNN Private – CT and US (Bone) Precision, Reca ll and Dice Directly re gr essin g the deformable multimodal (CT - US) registration pa ram ete rs using a weakly-s up e rvised train e d CNN Rohe et al. (2017) M. - M. Rohé, M. Datar, T. Heimann, M. Se rmesant, and X. Pennec, “SVF -Net: Le arning Deform ab le Image Registration Using Shape Matchin g,” Lecture Notes in Compute r Science, pp. 266 – 274, 2017. CNN Private - 3D MRI (Car diac) Dice, Hausdorff, LCC and RVLJ Directly re gr essin g the 3D unimodal deform ab le reg i stration p arameters using CNN Yoo et al. (2017) I. Yoo, D. G. C. Hildebran d, W. F. Tobin, W. -C. A. Lee, and W.- K. Jeong, “ssEMnet: Se rial -Section Ele ctron Microscopy Image Registration Using a S patial Transformer Netwo rk with Le arn ed Featur es,” Lecture Notes i n Computer Scien ce, pp. 249– 257, 2017. SA Es CREMI TEM – EM (Brain) Dice Using SAEs to extra c t unimodal structu ral features and feed to a STN f or final deformable re g ist ration Cao et al (2017) X. Cao, J. Yan g, J . Zhang, D. Nie, M. Kim, Q. Wan g, and D. Shen, “Deformable Im ag e Registration B ased on Simil ar ity - Steered CNN Regressi on ,” Lecture Notes in Co mputer Science, pp. 300 – 308, 2017. CNN LONI LPBA40 (Br ain MRI) Dice and AS S D Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN (Transfer Learning) ANDI (Brain MRI) Uzunova et al. (2017) H. Uzunova, M. Wilms, H. Handels, and J. Eh rhard t , “Training CNNs for Image Registr ation from Fe w S amples with Model - based Data Au gm ent ation,” Lecture Notes i n Computer S cience, pp. 223 – 231, 2017. CNN (FlowNet) (Dosovitskiy et a l . 2015) LONI LBPA40 (Br ain MRI) Jaccard and ASCD Directly re gr essin g the 2D unimodal deform ab le reg i stration p arameters using a weakly-supervised CN N Private - cine Cardiac MRI Yang e t al. (2017) X. Yang, R. Kwitt, M. St yn e r, and M. Niethammer, “ Q ui cksilver: Fast predictive im ag e re gistration – A deep learning approach,” NeuroImage, vol. 158, p p. 378 – 396, Se p. 2017. CNN OASIS – MRI (B rain ) SSD and Deformation Error Directly re gr essin g the deformable re g ist ration parameters via h ybrid C NN and LDDMM IBIS 3D Autism – TI and T2 MRI (multimodal) (Brain) Zh e ng et al. (2018) J. Zh e ng, S. Mi ao, Z. Jane Wang, and R. Liao, “Pairwise dom ain adaptation module fo r C NN -b ased 2-D/3- D re g i stration,” Journ al of Medical Imaging, vol. 5, no. 02, p. 1, Jan. 2018. CNN TEE – X-ray (Spine) TRE Directly re gr essin g the rigid unimodal registration parameters using CNN b as e d on the implanted p articles Spin e – X-ra y and CT (Spine) Hu et al. (2018) Y. Hu, E. Gibson, N. Ghavami, E. Bonmati, C. M. M oore, M. Emberton, T. Ver cau te ren, J. A. Noble, an d D. C. Barratt, “Adversarial Deform ation Regularization fo r Tra i ning Image Registration Neural Net works,” Lecture Notes in Co mputer Science, pp. 774 – 782, 2 018. GAN SmartTarget – MRI -T2 and TRUS (Prostate) - Multimodal TRE and Dice Directly re gr essin g the mu lt imodal def ormable reg i stration vi a a weakly- supervised anatomic al-label- dr i ven GAN Hu et al. (2018) Y. Hu, M. Modat, E. Gib son, N. Gh a va mi, E . Bonmati, C. M. Moore, M. Emberton, J. A. N oble, D. C. B arra tt , and T. Vercauteren, “Label - dr i ven we akly-supervised le arn i ng for mu lt imodal def ormarle image registration,” 20 18 IEE E 15th International Symposiu m on Biomedical Imagin g (ISBI 2018), Apr. 2018. CNN SmartTarget – MRI -T2 and TRUS (Prostate) - Multimodal TRE and Dice Directly re gr essin g the deformable multimodal reg i stration p arameters using a weakly-supervisedl y trained CNN Hu et al. (2018) Y. Hu, M. Modat, E. Gib son, W. Li, N. Ghavam i , E. Bonm ati , G. Wang, S. Bandula, C. M. M oore, M. Embert on , S. O urselin, J. A. Noble, D. C. Barratt, and T . Ver cau te ren, “Weakly -supervised convolutional neural netwo rks for multimod al i mage registration,” Medical Image An alysis , vol. 49, pp. 1 – 13, Oct. 2018. CNN SmartTarget – MRI -T2 and TRUS (Prostate) - Multimodal TRE and Dice Directly re gr essin g the deformable multimodal reg i stration p arameters using a weakly-supervised t rain e d CNN Dalca et al. (2018) A. V. Dalca, G. Bala kr i shnan, J. Gutt ag, and M. R. Sabun cu , “Unsupervised Learning for Fast Probabilisti c Di ffeomorphic CNN ADNI OASIS Dice Directly re gr essin g the unimodal deform ab le ABIDE ADHD200 - 6 - Registration,” Lecture Notes in Computer S cienc e , pp. 729– 738, 2018. MCIC PPMI diffeomorphic re g ist ration parameters using CNN with unsupervised learning HABS Harvard GS P MRI (Brain) Ba l akrishnan et a l . (2018) G. Balakrishnan, A. Zhao, M. R. Sabun cu , A. V. Dalca, and J . Guttag, “An Unsupe rv i sed Learning M odel for Deformable Medical Image Registratio n,” 2018 IEEE/CVF Co nf e rence on Computer Vision a nd Pattern Re cogn itio n, Jun. 2018. CNN ADNI OASIS Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN with unsupervised learning ABIDE ADHD200 MCIC PPMI HABS Harvard GS P MRI (Brain) Shu et al. (2018) C. Shu, X. Chen, Q . X ie, and H. Han, “An unsupervised network for fast microscopic image reg i stration,” Medi cal I ma g i n g 2018: Digital Pathology, M ar. 2018. CNN Private – EM (Br ain ) Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN Awan and Ra j poot (2018) R. Awan and N. Rajpoot, “Deep Autoencoder Fe atures for Registration of Histology Im ages,” Medical I mage Underst anding and Analysis, pp. 37 1 – 378, 2018. SA Es Bioimaging Challen g e 2015 – EM (Breast) RMSE Using a convolutio nal SAEs as a multimodal similarity for rig i d registration. Stergios et al (2018) C. Stergios, S. Mihir, V. M aria, C. Guillaume, R. Marie -Pierre, M. Stavroula, and P. Nikos, “Linear and Deformable Image Registration with 3D Co nvolutional Neural Net works,” Lecture Notes in Computer Scien ce, pp. 13 – 22, 2018. CNN Private – MRI ( Lung ) Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN in unsupervised manner Onieva et al. (2018) J. Onieva Oniev a, B. Marti-Fuste r, M. Pedrero de l a Puente, and R. San José Estépar, “Diff e omorphic L ung R e gistration Usin g Deep CNNs and Rei nforc e d Learning,” Le cture N otes in Computer Scien ce, pp. 284 – 294, 2018. CNN (Re g Net) (Sokooti et al. 2017) COPDGene – CT ( Lung ) Deformation Error Directly re gr essin g the unimodal diffeomorphic deformable registr ation parameters using CNN Mahapatra et al. (2018) D. Mahapatra, Z. Ge, S. Sed ai, and R. C hakravorty, “Joint Registration And Se gmen t ation Of Xr ay Images Using Generative Adversarial Networks,” Le ctur e Notes i n Computer Scien ce, pp. 73 – 80, 2 018. GAN NIH ChestXray14 – X -Ray (Chest) TRE, Dice and Hausdorff Directly re gr essin g the unimodal deform ab le reg i stration p arameters using GAN Mahapatra et al. (2018) D. Mahapatra, B. Anton y, S. Sedai, and R. Garnavi, “Deformable medical image registr ation using generative adversarial net works,” 2018 IEEE 15th Intern ational Symposium on Biomedi cal I maging (ISBI 2018), Apr. 2018. GAN Private – CFI and F A (Retina) Dice, Hausdorff, MAD, MSE, Deformation Error Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using GAN Sunnybrook – 3D MRI (Cardiac) Sentk e r et al. (2018) T. Sentker, F. Madest a, and R. Werner, “GDL -FIRE 4D: Deep Learning- Based F as t 4D CT Image Re g is tration,” Lecture Note s in Computer Scien ce, pp. 765 – 773, 2 018. CNN DIRLAB – 4D CT ( Lung and Liver) TRE Directly re gr essin g the unimodal deform ab le 4D CT reg i stration p arameters using CNN CREATIS – 4D CT (Lu ng and Liver) Ito and Ino (2018) M. Ito and F. Ino, “An Au t omated Method fo r Gen e ratin g Training Sets for Deep Learning based Image Re g i stration,” Proc ee din gs of the 11th Inte rna ti onal Joint C onference on Biomedical Engineerin g Systems and Technol og ie s, 2018. CNN (GoogLeNet) (Szeg e dy et al. 2015) ANDI - MRI (Brain) Precision and Recall Automated method fo r generating training set for imag e registration, aiming at realizing non- rigid reg i stration with deep learning. Yan et al. (2018) P. Yan, S. Xu, A. R. R astinehad, and B. J. Wood, “Adversarial Image Registration with App l ication for MR and T RUS I mag e Fusion,” Lecture Notes in Computer Science, pp. 1 97– 20 4, 2018. GAN NIH MSH – T2 M RI and TRUS (Prostate) TRE and D- Scor e Directly re gr essin g the mu lt imodal ri g i d registr ati on parameters using GAN Abanovie et al. (2018) E. Abanovie, G. Stankeviei us, and D. M atu ze vieius, “Deep Ne ura l Network-based Fe ature De scriptor fo r Retinal Im ag e Registration,” 2018 IEE E 6th Workshop on Advances in Information, Ele ctronic and Ele ctrical En g i neering (AIEEE) , Nov. 2018. CNN Chas e DB Diaret DB Matching Performance Using CNN as a uni modal similarity measure to guide conventional rigid iter ative approaches DTSET1 DTSET2 HRF-base Messidor1 RODREP FIRE CFI and OCT (Retin a) Miao et al. (2018) S. Miao, S. Piat, P. Fis cher, A. Tuysuzoglu, P. Me wes, T. Mansi, R. Liao, “Dilated FCN for multi -ag e nt 2D/3D medical image reg i stration,” in Thi rty -Second AA A I Conferen ce on Artificial Intelligence, Apr. 2018. CNN Private – CBCT and X- Ray (Spine) TRE Directly re gr essin g the mu lt imodal ri g i d registr ati on parameters using CNN Toth et al. (2018) D. Toth, S. Miao, T. Kurzendorfer, C. A. Ri naldi, R. Liao, T. Mansi, K. Rhode, and P . Mountney, “3D/ 2D model - to -image reg i stration b y im i tation learning for cardiac pro cedur e s,” International Journal o f C omputer Assiste d Radiology and Surgery, vol. 13, no. 8, p p. 1141 – 1149, May 2018. CNN LIDC-IDRI – CT ( Cardiac) TRE Directly re gr essin g the mu lt imodal ri g i d model to imag e registration p arameters using CNN Private - MRI (Cardi ac) Blendowski and H e inrich (2018) M. Blendowski and M. P . Heinrich, “3D -CNNs for Deep Binary Descriptor Learning i n Medical Volume Data,” In formatik aktuell, pp. 23 – 28, 2018. CNN DIRLAB – 3D CT ( Lung ) Retrieval Ra te Using CNN as a uni modal similarity measure to guide conventional iter a ti ve approaches Liu et al. (2018) X. Liu, D. Jiang, M. Wang, and Z. Song, “Image synthesis - based mu lt i-modal image registration framework b y using deep fully convolutional netwo rks,” Me dical & Biolo g i cal Engineerin g & Computing, vol. 57, n o. 5, pp. 1037 – 1048, Dec. 2018. CNN BrainWeb – T1, T2 and P D MRI (Brain) Mean TRE Using CNN as a multi modal to unimodal similarit y m e asure convertor to guide conventional deform able iterative approaches o r dee p learning. IXI - T1 and T2- w ei ghted MRI (Adult Brain) Eppenhof et al. (2018) K. A. J. Eppenho f, M. W. Lafarge, P. Moesk ops, M. Veta, and J. P. W. Pluim, “Deform ab le image re gistration using convolutional CNN DIRLAB – 3D CT ( Lung ) TRE Directly re gr essin g the unimodal deform ab le CREATIS – 3D CT (Lu ng) - 7 - neural networks,” Medic al I maging 2018: Im ag e Processin g , M ar. 2018. reg i stration p arameters using a combination of TPS and CNN Krebs et al. (2018) J. Krebs, T. Mansi, B. M ailhé, N. Ayache, and H . Delingette, “Unsupervised P robab il istic Deformation Mo deling for Robust Diffeomorphic Re gistration,” Lecture Notes in Computer Science, pp. 101 – 109, 2018. SA E ACDC – MRI (C ardiac) Dice, RMSE, MDM, MDG and Hausdorff Directly re gr essin g the unimodal deform ab le diffeomorphic re g ist ration parameters using SAEs Kearney et al. (2018) V. Kearney, S. Haaf, A. Sudhyadhom, G. V ald e s, and T. D. Solb e rg, “An unsu pervised convolution al neural network - based algorithm for defo rmable i mage registration,” P hysics in Medicine & Biology, vol. 63, no. 18, p. 185017, Sep. 2018. CNN Private – CBCT and CT (Head and Neck) NMI, FSIM, and RMSEc Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN Sheikhjafari et a l . (2018) A. Sh ei khj afar i, M. N oga, K. Punith akumar, N. Ray, “Unsupervised deform ab le image re gistration with fully connec te d generative neural net work,” in proc. the 1st Confe rence on Medical Imaging with Deep Learning (MIDL 201 8), The Netherlands, 2018. SA E ACDC – MRI (C ardiac) Dice Using SAEs to produ ce a l ow - dim e nsional vecto r from imag e and feed the m to a optimizer and full y-connected network for final re gistration Sun et al. (2018) Y. Sun, A. Moelker, W. J. Niessen, and T. v an Walsum, “To ward s Robust CT-Ultr a s ound Registration Usi ng Deep Learning Methods,” Lecture N otes i n Computer Scien ce, pp. 43– 5 1, 2018. CNN Privat e – CT and US (Liver) MAE Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using CNN Sun and Zhang (2018) L. Sun and S. Zh ang, “Deformable MRI -Ultr as ou nd Registration Using 3D Convolutio nal Neural Network,” Le ctur e Notes in Computer Scien ce, pp. 152 – 158, 2018. CNN RESECT – T1 an d T2 M RI and U S (Br ain ) TRE Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using CNN Cao et al. (2018) X. Cao, J. Yan g, L. Wang, Z. Xue, Q. Wa ng, and D. Shen, “Deep Learning Based Inter -modality Image Registr ation Supervised by Intra- modality Simil arity,” Lecture Notes in C omputer Science, pp. 55 – 63, 2018. CNN (U -Net) (Ronneberger et a l . 2015) Private – CT and MRI (Prostate) Dice and AS D Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using CNN Cao et al. (2018) X. Cao, J. Yan g, J . Zhang, Q. Wan g , P. -T. Yap, and D. Shen, “Deformable Ima g e Re gistration Usin g a Cue -Aw are Deep Regression Network,” IEEE Tr ansactions on Biomedi cal Engineering, vol. 65, no. 9, p p. 1900 – 1911, Sep. 2018. CNN ANDI - MRI (Brain) Dice and AS D Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN LONI - MRI (Brain) IXI - MRI (Brain) Ferrante et al. (2018) E. Ferrante, O. Oktay, B. Glocker, and D. H. Milone, “On the Adaptability of Unsupervise d CNN -Bas e d Deform ab le Image Registration to Unseen Im ag e Domains,” Lectu re Notes in Computer Scien ce, pp. 294 – 302, 2018. CNN Sunnybrook Cardi ac D ata – 3D MRI (Cardia c) Dice, MAD, and M CD Using transfer-le arning for zero-shot multimodal deformable image registr a ti on JSRT – X-Ray (Chest) Li and Fan (2018) H. Li and Y. Fan, “Non -rigid image re g i stration usin g s el f- supervised full y conv olutional net w or ks without trainin g da t a, ” 2018 IEEE 15th Intern ational S ympo s ium on Biomedi cal I maging (ISBI 2018), Apr. 2018. CNN LONI LPBA40 - MR I (Brain) Dice Deformable unimod al i mage reg i stration usin g a s el f - supervised CNN ( without training data) ANDI – MRI (Brain) Fan et al. (2018) J. Fan, X. Cao, Z. Xue, P .-T. Y ap, and D . Shen, “Adv e rsarial Similarity Network f or Evaluating Im ag e Ali gnm e nt in Deep Learning Based Re g i stration,” Lecture Note s in Computer Science, pp. 739 – 746, 2 018. GAN LPBA40 IBSR18 Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using GAN CUMC12 MGH10 MRI (Brain) Zhu et al. (2018) X. Zhu, M. Ding, T. Huang, X . Jin, and X. Zhang, “PCANet - Based Structural Rep resen t ation for Nonrigid Multim odal Medical Image Registration,” Senso rs, vol. 18, no. 5, p. 147 7, May 2 018. CNN (PCANet) (Chan et al. 2015) BrainWeb – T1, T2 and P D MRI (Brain) TRE Using PCANet to automatically extract structu ral features and feed them to L- BFGS for final deformable reg i stration AANLib – MRI (Br ain) RIRE – CT and MRI (Brain) Sloan et al. (2018) J. M. Sloan, K. A. Goa t man, and J. P. Siebe rt, “Learning Ri g id Image Registration - Utilizing Convolutio nal Neural Networks f or Medical Image Registratio n,” Proceedings of the 1 1th International Joint Con fer e nce on Biomedi cal Engineeri ng Systems and Te chno lo gies, 2018. CNN OASIS – MRI (B rain ) MSE Directly re gr essin g the mu lt imodal ri g i d registration parameters using CNN IXI - T1 and T2- w ei ghted MRI (Adult Brain) ISLES2015 – MRI (B rain) Schaffert et al. (2019) R. Schaffert, J. W ang, P. Fischer, A. Borsdorf, and A. Maier, “Metric -Driven Learning of Correspondence Wei gh ti ng for 2 -D/3- D Image Registr ati on,” Lecture Notes in Co mputer Science, pp. 140 – 152, 2019. CNN (PointNet) (Qi et al. 2017) C-arm – CT (Spine) Mean TRE, Mean RPD, Success Ra te and Capture Range Directly re gr essin g the unimodal rigid re g i stration parameters using CNN de Vos et al. (2019) B. D. de Vos, F. F. Be rendsen, M. A. Viergever, H. Sokooti, M. Staring, and I. Iš gum, “A deep learning frame work for unsuperv ised af fin e a nd deformable im ag e registration,” Medical Image Analysis, vol. 52, p p. 128 – 1 43 , Feb. 2019. CNN NLST - 3D CT (Chest) Dice, Hausdorff, and ASSD Directly re gr essin g the Affine and deformable registr ation parameters using CNN Sunnybrook Cardi ac Data – 3D MRI (Cardia c) Zhu et al (2019) N. Zhu, M. Najafi, B. Ha n, S. Hancock, and D. Hri stov, “Feasibility of Image Re gistration for Ultrasound - Gu i ded Prostate Radiotherap y B as e d on Simil ar ity Me asurement b y a Convolutional Neural Net work,” Tec hnology in Cancer Researc h & Trea t ment, v ol. 18, pp. 153303381882196, J an. 2019. CNN Private – 3D US (Prost ate) Registration Error Using CNN as a uni modal similarity measure to guide conventional ri g id patch- based approaches Blendowski and H e inrich (2019) M. Blendowski and M. P . Heinrich, “Combinin g MRF - based deformable registr ation and deep binary 3D -CNN des crip t ors for larg e lung motion esti mation in COPD p ati e nts,” Internatio nal Journal of Computer Assisted Radiology and S urger y, vol. 14, no. 1, pp. 43 – 52, 2019. CNN DIRLAB – 3D CT ( Lung ) TRE Using CNN as a uni modal similarity measure to guide conventional deform able iterative approaches Haskins et al. (2019) G. Haskins, J. Kruecke r, U . Kruger, S. Xu, P . A. Pinto, B. J. Wood, and P. Yan, “Le arn i ng deep similarity metric for 3D MR– TRUS image registr ation ,” Internation al J ournal of Comp uter Assisted R a diolog y and Surgery, vol. 14, no. 3, pp. 417 – 4 25, 2019. CNN NIH – MRI and TRUS (Prostate) TRE Using CNN as a multimod al similarity measure to guide conventional iter a ti ve approaches - 8 - Sun et al. (2019) S. Sun, J. Hu, M. Yao, J. Hu, X. Yang, Q. S ong, and X. Wu, “Robust Multimodal Ima ge R e gistratio n Us i ng Deep Rec urrent Reinforcement Learning,” Le ctur e Notes i n Computer Scien ce, pp. 511 – 526, 2019. RNN and CNN Private – MRI -C T (Nasopharynx) TRE Directly re gr essin g the rigid mu lt imodal registr a ti on parameters using the combination of CNN an d RNN (LSTM) Salehi et al. (2019) S. S. Mohseni Salehi, S. Khan, D. Erdogmus, and A. Gho l ipour, “R e al -Time Deep P ose Estimation With Geo desic Loss for Image - to - Te mplate Rigi d R e gistration,” IEE E Transactions on Me dical Imaging, vol. 38, no. 2, pp. 470 – 481, Feb . 2019. CNN Private – T1 and T2 MRI (Newborn Brain) Registration Error (Degree) Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters ( pose) using CNN Private – T1 and T2 MRI (Fetus Brain) Fan et al. (2019) J. Fan, X. Cao, P. - T. Yap, and D. Shen, “ BIRNet: Brain image reg i stration usin g dual-supervised full y convolutional net works,” Medical Image An alysis , vol. 54, pp. 193 – 206, May 2 019. CNN LONI LPBA40 - MR I (Brain) Dice Directly re gr essin g the unimodal deform ab le diffeomorphic re g ist ration parameters using a dual- supervised CNN IBSR18- MRI (Brain) CUMC12- MRI (Brain) MGH10- MRI (Brain) IXI - MRI (Brain) Krebs et al. (2019) J. Krebs, H. e Delin g ette, B. Mailhe, N. Ayache, and T. Mansi, “Learning a Probabilistic Mo del for Diffeomorphic Re gistration,” IEEE Tra ns action s on Medi cal Imaging, E ar ly Ac cess, pp. 1 – 12, 2019. SA Es ACDC – MRI (C ardiac) Dice, RMSE, MDM, MDG, Hausdorff and Grad Det-Jac Directly re gr essin g the unimodal deform ab le diffeomorphic re g ist ration parameters using SAEs Ba l akrishnan et a l . (2019) G. Balakrishnan, A. Zhao, M. R. Sabun cu , J. Gutt ag , and A. V. Dalca, “VoxelMorph: A Learning Framework for Def ormable Medical Image Registratio n,” IEEE Transactions on Me dical Imaging, Early Access, p p. 1 – 13, 2019. CNN Buckner40 OASIS Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN with unsupervised learning ABIDE ADHD200 MCIC PPMI HABS Harvard GS P MRI (Brain) Elmahdy et al. (2019) M. S. Elmahdy, T. J agt, R. T. Zinkstok, Y. Qi ao, R. Shahzad, H. Sokooti, S. Yousefi, L. I ncrocci, C. A. M. M ar ijne n, M. Hoogeman, and M. St aring, “Robust contour prop agation using deep le arning and ima ge registration for online adaptive proton therapy of prost a te canc e r,” Medical Ph ysics, May 2019. CNN and GAN LUMC – CT (Prost ate) Dice, MSD, and Hausdorff Directly re gr essin g the unimodal deform ab le diffeomorphic re g ist ration parameters using a combination of CNN an d GAN EMC – CT (Prostate) HMC – CT (P ro s tate) Yu et al. (2019) H. Yu, X. Zhou, H. Jiang, H. Kan g, Z. Wang, T. Ha ra, and H. Fujita, “Learning 3D non -rigid deformation b ased on an unsupervised deep le arning for PET/CT ima g e re gistration,” in Medical Imaging 2019: Bio medical Applications in M olecular, Structural, and Functio nal Imaging, Mar. 2019. CNN Private – PET and CT (Body) NCC and MI Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using CNN Che et al. (2019) T. Ch e , Y. Zhe ng , X. Sui, Y. Ji ang, J. Cong, W. Jiao, and B. Z hao, “DGR -Net: Deep Groupwise Registration of Multis pectral Images,” Informat ion Processing in Medical Im aging, pp. 706 – 717, 2019. CNN (U -Net) (Ronneberger et a l . 2015) Annidis RHA – MSI (Retina) Dice, Ratio of Registration, and T RE Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using CNN with unsupervised learning Van Kranen et a l . (2019) S. R. Van Kranen, T. Kanehira, R. Rozendaal, and J. Sonke, “Unsupervised deep le arn i ng for fast an d accurate CBCT to CT deformable image registr a ti on,” Radiother apy and Oncology, vol. 133, pp. S267 – S26 8, Apr. 2019. CNN Private – CBCT and CT (Head and Neck) Accuracy Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN Che et al. (2019) T. Ch e , Y. Zhe ng , J. Cong, Y. Jian g, Y . Niu, W. Ji ao , B. Zhao, and Y . Din g, “Deep Group -Wise Registration for Multi -Spectral Images From Fundus I mages,” IEEE Access, vol. 7, pp. 27650– 27661, 2019. CNN (U -Net) (Ronneberger et a l . 2015) Annidis RHA – MSI (Retina) Dice, Ratio of Registration, and CPD Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using CNN Hering and Heldmann (2019) A. Hering and S. Heldman n, “Unsupervised le arning for large mo ti on thora cic CT follow - up reg i stration,” Medi cal I maging 2019: Image Processin g, Mar. 2019. CNN Private – CT (Lung) Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN Hering et al. (2019) A. Hering, S. Ku ck e rtz, S. Heldm ann, and M. P. Heinrich, “Enhancing Label -D riv e n Deep De formable Image Registr ation with Local Distance Met rics for State - of - the- Art Cardiac Moti on Tracking,” Bildverarbeitun g für die Medizin 2 019, pp. 309– 314, 2019 CNN (U -Net) (Ronneberger et a l . 2015) ACDC – MRI (C ardiac) Dice Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN Liu et al. (2019) C. Liu, L. Ma, Z. Lu, X. Jin, and J. Xu, “M ultimodal medical imag e registration via common represent ations learning and differentiable geometri c constraints,” Ele c t ronics Letters, vol. 55 , no. 6, pp. 316 – 318, M ar. 2019. CNN (Xception) (Chollet 2017) APCH - DRR and DR (Body) Success Ra te Directly re gr essin g the mu lt imodal def ormable reg i stration p arameters using differentiable geometri c constraints and CNN (incorporating b ackground knowledge to CNN) Foo te et al. (2019) M. D. Foote, B. E. Zim merman, A. Sawant, and S. C. J oshi, “R e al -Time 2D-3D Deformable Re gistration with Deep Lear ning and Application to Lung Radiotherapy Targeting,” I nformation Proc e ssin g in Medical I maging, pp. 265 – 276, 2 019. CNN (DenseNet) (Gao et al. 2017) RCC T – 4D CT (L ung) Distance Error Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN CNN ANDI LPBA40 - 9 - Duan e t al. (2019) L. D uan, G. Yuan, L. Gong, T. Fu, X. Y ang, X. Chen, and J. Zh e ng, “Adversari al le arning for de formable registration of b rain MR image using a mu lt i-s cale fu ll y convolutional netwo rk,” Biomedical Signal Pro c es sing and Contr ol, vol. 53, p. 101562, Aug. 2019. (U -Net) IBSR18 CUMC12 Dic e and Distance Error Directly re gr essin g the unimodal deform ab le reg i stration p arameters using CNN MGH10 MRI (Brain) 3. Tax o nomy of Image Re gistration Ty pi call y , a go od i mage r egistra tion n eeds selec tin g prope r fe atur es, a similarity m etric (t o a sse ss the quality ), a transfo rmati o n mode l, and a se arch strategy in the state spa ce. So far, a large number o f co n ventio n al medica l image registratio n methods ha v e been propo sed in the literatu r e which can be classifie d base d o n diff erent metrics; A popular ye t stil l-alive taxo n omy is r epresente d in Fig. 2, whe r e th e classificatio n is base d on th e image dimensio n (2D, 3D , 4D, etc.), modality , so ur ce of th e f eatur es (int r insic v s. ext r ins ic), t ran sf ormatio n do mai n, t r ansfo r matio n mode l (rigid , af fine, and def ormable), k ind of th e fus ion (interpolatio n v s. appro ximation), user in te raction (m anual, semiau tomatic, and automatic) , a n d paramete r in v estigation met h o d (iterativ e v s. direct) (Maints & Vie r geve r, 1998) . Figure 2: A t axo nomy on imag e reg istratio n metho ds (Ma ints & Vi ergev er, 1998) At the be ginning in machine visio n fie ld, i mage r egis tration was refe rr ed to align 2D images; how ever, med ical imaging as illust rat ed in Fig. 3 is 3D in natu r e so th at mos t of the medical images captu r ed by current imagi n g dev ices like MR I and CT ar e 3-dime n tional . Co n v ent io n al 2D im aging modal ities l ike mammo graphy of X-ray can also be represe n ted as 3D w ith out l o sin g the generality . Be sides, there ar e plenty o f pr oc edures in w h ich a num be r of discrete (Fluo r os copy ) or co ntinuous (So n o graphy ) images are take n by th e phy sician s so that the transfo r matio n needs t o b e co nsider ed as 4D, addi ng t ime as th e fo ur th dime n sio n . Th ese interventio n al 4D images are o ften in low -qua lity w ith much amount of n oise due t o th e restrictiv e n atu r e o f th e ope r atio n al dev ices , w hi c h also challe nges th e r egist ration proce ss. Figure 3: A n example o f reg istering a 2D plane in a 3D vo l ume ( Fe rr ant e & Pa ragious, 2017) The pre-inte r ve n tio n al i mage on which th e treatme n t pla n is deve l ope d is r outi n ely a 3D high -qu ality MRI or C T image si n ce i t is taken by mo dern diag n os is devices with n o o per atio n al limitatio n , w h il e int ra-ope r ational i mages taken for tr ea tment and th e r apy pur pos es can be captur ed wit h the same (u n imo dal) o r dif ferent m oda lities (multimodal ). A co mmon case is to r egiste r a pre-i nter v entional 3D T1 -w eighted MRI with operatio na l Tra n s r ec tal U ltrasound (TRU S) images of t h e prostate gland . Often, same modalities w ith diff erent imaging paramete r s e.g . T1 and T2 we ighted MRI, as - 10 - illust r ated in Fig . 4., are conside r ed as mu ltimodal . Of c our se , co n sidering di ff er ent modalities, multi modal image registratio n is far muc h co m plicated si nce each modality i s sensitiv e to so me spec i fic pa r amete r s var y in g in the diff eren t body ’s tissues to generate co n trast w h ile ma y n ot be detec tib le in ot h e r modalities. Moreo ve r, so me similarity metrics e.g. S um o f S quared Diff er ences (SSD) c annot b e applie d to multi modal r egist ration regarding diff eren t colo r s o r inte n sit ies’ r ange of point s fo r diff erent modalitie s. Figure 4: Sagi ttal imag e s of human brain: T1 -w eighted , T2-weig hted, an d Pro ton-Densi ty MRI from left to right, resp e ctiv ely (I mages from BrainWe b projec t) Medic al image r egist ration ca n be int r ins ic o r ext r in s ic bas ed on the nature of extracted featu r es from th e image s . Althoug h intri n sic r egistratio n that is base d o n the anatomic a l s tructures in the bo d y i s by far much commo n , ext r ins ic registratio n base d o n the exte rnal o bj ects implanted in th e bo d y is still alive w ith num e rous fan s specia lly for skin registratio n . Th ese o bj ects, like what can be see n in the Fig. 5, can be im pl anted in the b ody for diff erent propo ses, y et are go od i n dicato rs to v erify the quality of registratio n . B esides , they may be r ef lective tiny ob jects or senso r s t o fac ilitate and accu r ate the r egist ration p r oc ess . E xtrin s ic r eg istratio n met h ods si mply us e the loc ations of these ob j ec ts a nd ca n identify the transf ormation p aramete r s ve ry sim ple, acc urate and fast . Figure 5: A n example o f impla nted ex terna l obj ects used f or extrin sic reg istratio n (Miao et al. 2016) Anot h er pa r amete r t o classify r egistration met h o ds is the part of image evo l ve d in the regist ration p r o ces s. A metho d is considered as glob al where all the image ’s points are use d i n the proce ss, a nd can be subjec ted to th e prospe ctive manipulat ion, while ot h e r app r oaches use only a part/so me parts of the image (at each iteratio n ) ar e refe rr ed t o as l oc al approac h es. Inte nsit y -bas ed methods ar e usually glo bal, w hile featu r e-base d o n es ar e typic all y loc al. Also, n ume r ous hy brid approac h es reporte d in t h e lite r ature h ave adopted multistage st rategy ; that is, a glo bal pre-regis tration is made at the be ginning fo l lowed by m uch loc al ly r egistratio n s at e ach s ta ge. Capability to c o n sider def ormation is anothe r decisiv e facto r to c ategorize image r egistrat ion methods spe cially in the fie ld of medicine. From this point of view , the unde r ly in g t r ansfo r matio n of image regist ration methods ar e divided into the t hr ee ki n ds o f r igid , Aff i ne and defo r mab le . The rigid t ransfo r matio n mode l only considers t ranslatio n and r ot ation o f image alo ng and arou n d t h e coo rdinate axes, respe ctive l y , s o th at the tra n sf ormatio n can be m o deled using 6 pa r amete r s - 11 - (also c alled deg r ee s-of-freedo m ) . Aff i ne tra n sf orma tio n also c o n siders scaling and s h ee r ing so th at t h e n umb er of parameters is doub led to the 12 o nes. Aff i ne tra n sf ormatio n i s no n li n ea r tra nsfo r matio n that preserve s points , straight lines and planes i.e. sets of para lle l lines r emai n pa r allel after the Aff in e transf ormation; h ow ever , angles be twee n lines or d istances be twee n po ints m ay not b e p r ese r ve d. Alt houg h Affine t ransfo r matio n is n o n li n ear, li ke ri gid t ransfo rm atio n, it is n o t able to capture th e deformatio n complexi ty i nh e r it in th e flexib le m emb ers and o r ga n s in side th e h um an bo d y . Bas icall y , rigi d and Aff in e tran sf ormatio ns have no more than the tw o fo llowing applicatio ns: These a pp r oac h es ca n be used for t o ugh structures of the bo dy l ike bones and skull registratio n, whe r e the r e are a lots o f fan s a mong the expe r ts and phy sician s b ec ause of the simplicity an d spee d. They can b e explo ited as a glo b al p r e-regist ration fo r muc h co mplex multistage defo r ma b le app r o aches in order to avo i d stalking i n l oc al mi n ima, and increase the co n vergenc e spee d. Practical ly, i ncreasi n g in t h e numb er of transf ormation p aramete r s i s in ev itable to mo d el the defo r mable orga ns i nside the b ody so t hat a la rge number of effo r ts have b een made to i ntroduc e de formable models matc h ed w ith d i ff er ent o r ga n s. Defo r ma b le models are almost alway s local, and ta r get pa r t of the image in eac h step. U suall y , at the be ginning, a mesh of c ontr ol points is co n side red fo r mi n g i n this w a y a defo r matio n netwo r k , a s illust r ate d in Fig . 6 . The n u mber of control points (a n d th e spac e amo ng them) i de ntifies the amount o f deformation th e m ode l can captu r e. In eac h ite r ation, some of th ese control points are subj ecte d to displacement, w h ic h cause a cha n ge in the place and inte n sity of oth er r elate d points in the image (usually using an inte rpolation mecha nism). In a commo n def o r mable approach named Thi n -Plate Spline (TPS) (Duchon 1977), a cha n ge in a co n trol point b r oadc asts to al l othe r poi n ts in th e image; this is a global approac h causing high ov erload on th e sy stem. In contrast , B-Splin e base d approac hes (Ruec kert et al. 1999) have bee n introduc ed in w hich displ acing a co ntr o l poi n t o nly a ff ect th e adjace nt po int s, fo r n ot only i ncreasi n g the loc ality of transfo rmati o n in o r de r to b etter cap ture t h e u n de rl y i ng def ormation, but also de creasing the co mputatio nal ov er load. Figure 6: A mesh of co ntr o l poin ts as a def ormation netwo r k used in defo r mab le im age reg istrat ion Generally , thr ee lev els of interactio n can b e co n side r ed f or image r egist ration tec hn ique s. I n manual approac h es, use r can co mpl etely in te r act w ith the sy stem, and may identify some paramete rs or prov ide a n approxima tio n of the transfo rmati o n for th e met h od . In co n trast, there is n o inte r ac tion w ith the mac hin e fo r ful ly -automated approaches, and w h o le the p r oc ess is do n e by th e a pproac h itself. S emi-auto mated app r oaches try to explo it the k n o wle dge of e xpert use r, though minimal ly . This minimal explo itation can be actually of high value , causi n g a tremendous decrease in the search space to decrease the risk of failed registratio n, and inc r ease the overall spee d; howev er, huma n interve ntion ca n c h alle nge w h o le th e p r o ces s sin ce th e diff er ent lev els of in te r actio ns c ann ot be validat ed, cont r o lled o r m easu r ed qu antitative ly . - 12 - 4. Prob le m Formulation The pu r pos e of image r egis tration is to fi nd a geo m etrical transf ormatio n T : I F → I M t o align a mov i ng image I M to the f ixed image I F in the be st way . This alig n me n t can be achiev ed conve n tio n ally by optimizing a similarity m easure L so tha t ) , ; ( m i n ˆ M F I I T L (1) w h ere T µ is the transfo r matio n model reco gni zed by the para meters µ . Th e re are a numb er of simila rity measures d ivided into t he tw o catego r ies of intensity - based and featu r e-base d ones. Ge n erally , in te n si ty-base d measures, e.g . Mea n Squa re Diffe ren ce (MSD), co n side r a c o mpl ete mechanical co rr espo ndence be tween the giv en image s, w h ic h may not be required from the perspective of h uman expe r ts. O n th e ot her hand, feature-based measu res, e.g. Mutual Info rmation w h ic h i s bas ed o n t h e info r matio n theo r y , lo ok fo r a s atisfy i ng struc tural co rr espo n de n ce be t wee n t h e O r g ans- of -In terest (Oo Is) i n the input ted images; practically , the approac h es b ased on the feature -b ased measures tr y to detect peers of str uctura l fe atur es like lines, co rn e r s, landmarks, co n tours etc. i n b oth t he fixed and m ov in g images, and to align t h em. T h e detectio n and se lecting these st r uctu ral f eatures can be ma nually (r ef erre d to as handc r afted fe atures), o r co mpletely automated like w h at is see n using de ep Co n vo luti onal Neu ra l Ne two rks (CNNs). Whatev er the simila rit y m easu r e is, the Eq. (1) n ee ds to be opti mized. The optimiza tion proce ss can b e co nducted in tw o w a ys ; co n ventio n ally , t h e af o r ementio n ed E q . is optimiz ed usi ng iterativ e approaches like h ill -climbi n g an d gradient dec ent; w hile in mac h ine learni n g a pp r oac h es, a transfo rmati o n m ode l is c r eated base d on some previo usly learn ed sa mpl es, and t h e p aramete r s are regress ed directly i n o n e shot using (2). ) , ( M F I I f (2) w h ere is the set o f t he m ac h ine-lea rning mode l’s parame ters ( the n etw ork’s weights) i dentifie d base d on th e trai n o n the learning data. I n oth e r wo r ds, the L a cts as a loss-f unction fo r th e l ea rning m ode l to be o ptimized usi ng some co nventional trai ning algo rithms, like back-p r opaga tio n , to l ea rn so that th e si milarity between the co uple o f th e inputted im ages ca n be maximized. 5. Deep Neu ral Ne twork s The theo r y of deep neural n etw orks backs to the late-70s (Fukushim a 1980), and was first applied t o medical image proce ssing by Lo in 1992 (Lo et al. 1992). Nev ertheless, since the proper infrast r uctu r e f or suc h a huge co mputation was not avai lable thos e day s, the f ir st o peratio n al impleme n tatio n dates to 1 998 w h er e in (Lec un et al. 1998) a Convo lutional Neural Netw ork (CNN) was utilized to reco gn ize hand-writte n n ume r ical characte r s fo r po st o ffice appli ca tio n . Numeric al character reco gn ition was such a simp le machi n e visio n task fo r w h ic h the co mputatio n al pow er p r ov ided by th e har dw are of that time allow s to effective l y use dee p learn ing. Unfo rt unately , the propo sed appr o ach l o st it s way to be appli ed to other much complex and co m plicated prob l ems ti ll 2012 whe r e Krizh ev sk y et al. (2012) w as pros per ed to train a deep CNN on g raphic card equip ped wit h a ma n y -core G PU, a n d won the g ran d w orl d image proc ess in g c hampio n s h ip n amed ImageNet (K r iz h ev sk y et al. 2012) . Since the n, other c h am pions w ere in the same f am ily of dee p learni ng t ec hn iques, each of which w ith a contrib uting n ov elty , ti ll it was announced the r ec ognition po we r of the deep learni n g propo sed approac h es o utpe r fo r m the hum an expe r t, a n d the champ ionship w as practica lly closed (Russakov sky et al. 2015) . Acco rdin g ly , deep n eural n etw orks of vario us kinds penet rated to all areas of ma c h ine visio n, and turned to be the domi n ant t ec hnique use d by a large n umber of experts activ e i n the field. Some activ e domains in the fie ld of medical image analy sis c an b e e n ume rat ed as orga n dete ction, la n d mar k lo cali zatio n, lesio n dete ction and c lassif ication, treatment planning an d fo llow-up (Litje n s et al. 2017). Of co ur se, image registratio n a s o ne of the mo st importa nt and challe n g ing dilemma in image-guide d in te r v ention was not an exceptio n, and a numbe r of approac h es as listed i n Table 1 h ave be en introduc ed so far. De ep n eu ral n etw orks can have di ff eren t arc hitectures and to polo gies mad e each of which suit able fo r so me specific a pplicatio n s . As s h ow n in Fig. 7 , f ive kinds of dee p n eu r al netw orks have a lready b een applie d t o m edic al image regist ration namely CNN ( 66 times, 80.5%), Stak ed Auto-E n co ders (SAEs) (8 time s, 9.8%), Ge n erativ e Adv ersarial Netw o rk (GA N) (6 times . 7.3 %) , Re current Neural Netw ork (R NN) (1 time , 1.2%) and Dee p Re info r ce ment Le arn ing (DRL ) (1 time, 1.2%). - 13 - Figure 7: Deep learning techn i ques used in the lit erature with the i r f requenc ies An Auto -Enco ders (AEs), as illus trated in Fig. 8-(a ) , i s a very simple netwo rk whic h trie s to r ec onstruct th e input pattern a s output us ing a si ngl e h idde n lay er. Of co ur se, the hidden laye r should be smalle r than the inputted pat tern so that it ma ps to a co m pacte r space of the h idden lay er w ith the most disc riminati n g capab ility . De n oisi ng AEs (DAEs) i s a resemb le netw ork t r y i ng to rec onstruct the i n pu tted patte rn s w ith so me noise applied. Apply in g so me noise to the input elev a tes th e gene r ali zatio n capab i lity of th e mode l. Dee p architecture of AEs called Staked Auto-E n co ders (SAEs), as presented in Fig. 8-(b), h as more h idde n lay ers staked on the top of one other. Ge n erally , th e computatio nal bu r de n of training suc h a n etw ork is not a f fordable; h e n ce , to make it practical, usual ly each laye r is trained separately , and a fi nal low - cost i n teg rated training fi n e t u n es w h ole the n e two rk. In t h e literature o f medical image r egis tration, this n etw ork has only bee n used to prov ide the m o st signific ant and disc riminati n g featu r es from th e im ages to fee d to an alternativ e registratio n method, i n s tead o f using ha n dcraf ted fe atures. Figure 8: (a) L eft: A uto-Encoders (A Es) ne twork (b) R ight: Staked A ut o- Encoders (SA Es) netwo rk CNNs should be c o n side red a s o n e of th e mo st succ essful and powe r ful dee p learni n g tec hniques in whic h whole th e give n image (or so me extracted p atches) is fe eded dir ec tly to th e n etw ork. And, this i s v ersus the tradit ional n eural n etw ork bas ed image proc essing a pp r oac h es in w h ic h so me handc rafted feature s we r e extracte d at fi rst, an d prov ided t o the netwo r k. As r ep r ese nted in Fig. 9, a ty pi cal CNN has some interleav ing kernel and poo lin g lay ers to be ended wit h a ty pi cal fully - connected tw o o r three-lay er n etwo r k . Ke rn els ar e trained to extrac t t h e most sig nificant f eatures v i a co nvolving wit h the input , while poo ling lay er s decrease th e curse of dimen s ionality , an d make the results invaria n t to the diff er ent geo metrical transfo r matio n s . The output of each laye r so-c all ed a fe atu re -map is in p utted to the n ex t lay er , and w her e the numbe r laye rs i s high, a hie r archical fe ature-se t can be achiev ed, and th e netw ork can b e r egarde d as dee p CNN. The featu r e-m aps of the last lay er i s co n catenated an d vec torized to fee d a f ully-co n n ec ted two o r thr ee-lay er netwo r k fo r the fi nal classificat ion. In a large n umbe r of case s, e.g. in defo r ma b le image r egist rati o n , w e are wit n ess ed CNN , 66 , 80.5% SAEs , 8 , 9.8% GAN , 6 , 7.3% RNN , 1 , 1.2% DRL , 1 , 1.2% CNN SAEs GAN RNN DRL - 14 - the so -c all ed U-Ne t (Ro n nebe r ger et a l. 2015) whe r e th e fina l fully-co n nected lay er can be droppe d out, li ke Fig. 10 , so that a di r ec t end- to -e nd regist ration field ca n be a chiev ed. Figure 9 : Co nvo luti o nal Neura l Netwo rk (CNN ) arc hitect ure Figure 10: Ful ly Convo luti o nal Neura l Netwo rk (fCNN o r U-N et ) arc hitectu re CNN h as a lso t h e capab ility of ge tti ng h eteroge n eo us patterns wit h dif fe r ent r ep r ese ntati o n s as input. Ea c h represe n tat ion is r eg ar ded as a c h annel, a nd t h e netwo r k h aving thi s capab ility is c alled mu lti-channel . To exe mplify , let ’s co nsider s mall-sized patc h es extracted from images and in p utted to the n etwo r k fo r the classificat ion issue. S h ould th e co nt ext be info r mative , one ca n co n side r so me large r pa tches a r ou n d the sele cted patch, co mpact them, and fe ed th e m to the n etw ork as a separate channe l . Of co ur se, these compac ted larger-sized p at c h es ca nn o t be pr oc esse d by th e netwo r k via th e same c h anne l with the o ri g inal-sized patc h es. Anot her examp le is w h e r e we are facing co lor images so that ar e able t o use thr ee chann e l s o f RGB i nstead of one ch anne l of intensity for each i mage’s point. The n etw ork ca n fuse the channel in ea rly layers, or po stpone it t o the last lay ers r egarded as multi-stea m n etwo r k. Mo st of th e CNNs applied to medical i mage regist ratio n are of thi s type where a couple of patc hes ext racted from th e give n fix ed and mov in g images (almo st with diff er ent moda lities a nd rep r ese ntati o n s) a r e fe eded sepa rately as t w o channels , p roce ssed separately in tw o pathway s (pipelines) where t h e info r mation f usion is usually postpo n ed to the la te laye rs (Fig. 11) . Figure 11 : Multi-stream CNN archi tecture - 15 - Propo sed by Goo dfellow et al. i n 2014 (Goo d f ellow et al. 2014), a Generative Adversarial Netw ork (GAN) is co mposed of two compe ting subnetw orks, th e generator and the discrimina tor, as de monstrated in the F ig. 1 2. The generato r i s trai n ed on g r ound-trut h datase t to sy n thesize fake samples, while th e discrim inator s h ould discrim inate be t w een fake (sy n thesized) data and the r ea l o n e, a s a bi n ary output. Base d o n th e su r viv a l co m petition be twee n th e generato r and the disc rimi n ato r, just like the game theo r y , th e n etw ork can be trained on a small set of da ta so that th e generated samples canno t b e discrim inated, and th e n etw ork g oes fo r equilib r ium. As the ge nerato r is trained adve r saria lly bas ed on th e discrimi nat o r ’s fe edbac k, the n etw ork takes its name. While the origi nal GAN w as applied for the i mage noise remov al, i t has gained increasi n g popula r ity in r ec ent y ears, and applied to a lmo st a ll the prob lems in medical imaging (Yi et al. 2018) . I n the context of image r egistratio n, the generato r t akes th e inputted fixe d and mov in g images , and try to p r oduc e suc h transfo r matio n pa rameters so that t h e tra nsfo r med mo ving image ca lled w ar ped image ca nn o t b e discrimina ted from t he grou n d-t r uth by the disc r imi nat o r , the sit uat ion expe cted f r om an expe r t r egist ration a ge n t . Figure 12 : Gener ative A dversarial Netwo rk (GA N) arc hitectu re Rec urr ent N eu r al N etw ork (R NN) is just like t he trad itional Mult i-Lay er Pe r ce ptron (MLP ) o r AEs wit h an ext ra data loo p (fee dback) f r o m the hidde n -lay er n o des to themselv es, as illus trated i n Fig. 13. W h ile the other n etw orks like CNNs or AEs are su itable fo r spatial analy sis, these feedb ack lo ops make the RN Ns the mos t pow erful for temporal analy sis. Routi n ely , th e p revio us states a re h eld ac r os s the hidden lay ers, and a cc ordingly , the next s tate ca n b e e stima ted usi n g the curre n t s tate i nputted to the netwo rk, and t he p r ev ious sto r ed ones. Where t h e numb er of h idde n lay ers go es more to sto r e farer s tates, the n e t w ork is conside r ed as deep. Again, just like SAEs, the comput ational burde n of traini n g suc h h eavily co nn ected n etw o r k is u naffo r dable i n traditio nal manner; h e nce , rese ar c h w a s co nstan tly fo llowe d by th e co mmunity , and fortunately simpli fie d m emo r y m o dels such as Lo n g S h o r t Te r m Memory (LS TM) (Ho chr eite r and Schmid h ub er 1997) and Gated Re curre n t U nit s (GRD) (Cho et al. 2014) we r e introduce d an d popula rly applied in th e r ece n t y ears. Wor se mentioni n g that in th e co ntext of i mage r eg istratio n, the R NNs are mostly used fo r opt ical -f low , and whe r e one o f the modalities are asso ciated wit h tempo r al dime n sio nalit y e.g. TRU S or X -ray fluorosco p y . Figure 13 : Recu rrent Neu r al N etwork (RNN ) ar chi tecture - 16 - Last but n ot least is the Dee p Re in fo rcement Lea rning (DRL ). It is b ased on th e th eo r y of stochastic proce sses an d Markov cha in. The r e is an a ge n t with so me inte rnal st ates, tran sitio n p r ob a b il ities , and a r ew ar d/pe n alty rate. It learns iterative l y to interact w ith the enviro nm e nt. A t e ach iteration, the DRL machine c hoo s e an ac tion f r om its actio n - list base d on the enviro n ment’s fee dback, its cu rr ent inte rn al s tates and tra n sitio n prob abilities vi a a prob a bilistic decision making proce ss. The sele cted action is a pplie d to th e enviro n ment, and based on the desirability of its feedback, the m ac hine gai n s reward o r pe n alty . I n this ma nn e r, t he DRL m achine le arns to sele ct the be st action in eac h si tuation, whe re the b est actio n is the one wit h th e mos t prob ability to get r ew ar d from the e nvironme n t. In the co ntext of ima ge r egist ration, suc h DRL agents we r e a pp lied to spe cific al l y rigid or aff in e transf ormatio n s where the n umb er of states are r es tri cte d, and affo r dable for the age nt to be conve r ged. Fo r example , th e age nt ca n selec t the actio ns of 1 deg r ee cloc kw is e/co un ter clo ckw i se rotation o r 1 m m (mi llimete r ) tra n slat ion in all the directio ns. These sele cted actio n s are applied t o the mov i ng image, and based o n th e desirability of actions e.g. a simila rity metri c , the age nt gets r ew ar d/penalty . It update s its inte rn al transitio n prob a b il ities b a sed on a learni ng algo r ithms suc h as Q- Le arn ing to m aximize its pe r fo r man ce . Fi g. 14 is a clea r illust r ation o f this co n ce pt. Figure 14 : Deep Reinf orc ement Lea rning (DRL ) architec ture a pplied to medic al image reg istration (Sun et al. 2019 ) 6. Lite rature Re v iew Conv ent io n al i mage registratio n is conducted usi ng i terati ve optimizatio n alg orit hms. In eac h ite ratio n , a b etter alig n me n t is suppo sed t o be ach iev ed base d on a p r ede fined simila rity m easure. The operatio n s co n tinue till n o be tter registratio n ca n b e achiev ed, o r so me p r ede fined crite r ia are satisfied. P r olo nged running time, and defec tive nature of the introduc ed simil arity measures spec ially for mult im o dal r eg istrat ion, which causes getti n g trapped in local mi n ima, are of m os t challengi n g issue s to be tackled for exploiting this paradigm. To r eso l ve th e afo rementioned prob lems, deep learning base d app r oaches have gained inc r easing popularity in r ece nt y ear s. The underly in g philo so ph y behind th ese approac h es a re divide d into the two follow in g catego ries: Dee p n eu ral n etwo r k acts as an app roximator o f the simil arity betwee n in putted images as a co mplete and no- faulty similarity m etric in orde r to help ot h e r r eg istratio n methods . Deep n eural netwo r k acts as a reg resso r to di r ec tly estimate the t r ansf ormatio n paramete r s in o ne- shot in orde r t o maximize t h e runtime speed. Bas ed on the literat ure breakthroug h s , a taxonomy w ith fiv e gen erat ions ca n be concluded n amed Deep Simila rit y Metrics (DSM) , Supe rvised End- to -End Re gistratio n (SE2ER ), Deep Re in fo r ceme n t Le arning (or Agent -Base d Re gistr atio n ) (DR L), U n supe r vise d End- to -End R egistratio n (U E 2ER), We akly /Semi -Supe r vise d En d- to -En d Re gistr atio n (WSE2ER ) (Fig. 15) . Insp ired by e.g. (No w ak an d Jurie 2007) and (Huang et al. 2012), the first generatio n of wo r ks w ere base d o n the utiliz ation of diff erent kinds o f DNNs to lea rn visual similarity metrics from a large set of paired annotate d grou nd -trut h s . We call ed th em dee p similarity m easures/met ri cs . As illustrated in Fig. 16, the learned mode l after th e train i s suppo sed to be able to precise l y and meaningfully model the struc tur al dif fe r ences betwee n th e - 17 - inputted p air s of images/patc h es. (Wu e t al. 2013) and (Che ng et al. 201 6) are t he mos t impo rtant r eprese ntatives f or this primary generatio n . De ep similarity measur es are b a sica lly prov ided to th e co n ve n tio nal ite r ative defo rmable r egist ration algorithms in order t o produc e f inal tra n sf ormatio n para meters. As a large n umb er o f sim ila r a pp r oac h es ha v e b een co nducted so far, n ow ada y s, we can def in itely argue that this paradigm, in its basic fo r m , can be a po ten t rival to th e co nventional multimo dal simila rity measur es e .g. Mutu al Info rm atio n (MI) if and o n ly if th e ade quate n umb er of clear ly annotated grou n d-t r ut hs are available , which is a sev ere confining fac tor to deve lop su ch approac h es. Mo r eov er , it has bee n rev ealed that, fo r u nimodal regis tration, if the simila r ity m easu r e can be p r o perly se lected base d on th e co n text and modality , th e uti lization o f dee p sim ilarity m easu r es has no s trong just ificatio n. A l ist of wo r ks be long to thi s ge neratio n are prese n ted in Tab le 2 . Figure 15: The taxo nomy on deep lea rning ap proac hes fo r medic al imag e reg istratio n Figure 16 : A de ep sim ilarity metri c b ased o n the CNN (Zago r uy ko and Kom odaki s 2015 ) Table 2 : T he Firs t Gene ration of Deep Learn ing A pproac hes for M e dical Im age Registra tion (De ep Sim ilarity Metrics) Reference Title T ec hnique Modality Transformation Wu et al. ( 2013) Unsupervised deep feature le arning for de formable registration of MR b rain images CNN Multimodal Deformable Zhao et al. (2015) Deep adaptive lo g-d e mons: diffe omorphic image registration with v e ry lar g e deformations CNN Unimodal Deformable Simonovsky et al. (2016) A deep metric for multim odal registration CNN Multimodal Deformable Cheng et al. (2016) Deep similarity learnin g for multimodal me dical images SA Es Multimodal Rigid Wu et al. ( 2016) Scalable high-per formance image re g istratio n framework b y unsupervised deep feature representations le arn i ng SA Es Unimodal Deformable Yang e t al. (2016) Registration of patholo gical images SA Es Unimodal Deformable Wang et al. (2017) Scalable high performa nce image registration fr amework by unsupervised deep feature representations le arn i ng SA Es Unimodal Deformable Ghosal et al. (2017) Deep deformable re g i stration: Enhan cing accuracy by fully c onvoluti onal neural net CNN Unimodal Deformable Awan et al. (2018) Deep Autoencoder Fe atur e s for Re g i stra ti on of His t ology Images CNN Unimodal Rigid Blendowski et a l. (2018) 3D -CNNs for deep binary descriptor learning in medi cal volume dat a CNN Unimodal Deformable Abanovie et al. (2 018) Deep Neural Net wor k -based Fe atur e Descriptor for Retinal Im age Registratio n CNN Unimodal Rigid Zhu et al. (2018) PCANet-Based Str uctural Representation fo r Nonrigid Multimodal Medi cal Image Registration CNN Multimodal Deformable Liu et al. (2019) Image synthesis-based m ulti-mod al i mage registration fr amework by using deep fu l ly convolutional netwo rks CNN Unimodal Deformable Deep Learning App roaches for Medical Image Registration Deep Simili arity Metrics (DSM) Supe rvised End- to -End Registration (SE2ER) Deep Reinforceme nt Learning ( Agent-Based Approaches) (D RL) Unsupe rvised End- to -End Registration (U E2ER) Weakly/Se mi-Supervise d End- to -End Regi stration (WSE2ER) - 18 - Zhu et al. (2019) Feasibility of Image Re g i stration for Ultrasound -Guided Prostate Radi otherapy Based on Similarity Me asur e ment b y a C onvolutional Neur a l Ne twork CNN Unimodal Rigid Haskins et al. (2019) Learning deep similarit y metr i c for 3D MR -TRUS i mage registration CNN Multimodal Deformable Blendowski et a l. (2019) Combining MRF -based deformable registr a ti on and deep bin ary 3D -CNN descriptors for l arge lung motio n estimation in COPD patients CNN Unimodal Deformable The seco n d generatio n be longs to the e n d- to -e n d supe r v ised r e gi stratio n , w h e r e the dif f eren t kin ds of DN Ns are trained on grou nd -trut h to co n struct r egress ion mode ls to produc e the tra n sf ormation p arameters in one -shot. Fo r a ff i ne and defo rm able transfo r matio n models, CNN and U -Net (i.e. fully CNN) ar e predo minant t ec hniques , r espec tively (Miao et al. 2016) and (So koo ti et al. 2017 ). T he mai n def orm able fra mewo r k is illust rated i n Fig. 17 . First of a ll, a grid o f co n trol points is co nsidered as a Dense Displ acement Field (DDR ). Eac h co n trol poi n t ca n be freely tr anslated in horizo n tal and ve rt ical d irections in order t o capture the underly in g def ormation. Th e n umbe r of co n tr ol poi nts and th e space amo n g them gov ern th e accuracy of mode l to capture the defo r mat ion. In the so-c all ed Thin-Plate Spli n e (TPS) eac h mo ve ment in a co ntrol point is b roadcasted to the all o n es (glob al t ransfo r mation), while so-c all ed B-Spli n e approac h es only co nsider ed a djac ent co ntrol poi nt s (lo cal tra n sf ormatio n ) to tract th e co mputatio n al o ve rh ead. Def ormations ar e c ontrolled by th e regul arizer that pe n alizes i n acc eptable t ran sf ormatio n s. Jus t like c onve n tio nal ap proaches the r e is a heavy disputat ion a n d dis agreeme n t o n t h e r egul arization te rm to b e dictat ed to th e DNN, w hich t urned to b e th e so ur ce o f many innov a tions in t h e fie ld (Soti ras et al. 201 3). Figure 17 : The m ai n f ramewo r k fo r supe rvised end- to -end medic al imag e reg istratio n Anot h er sou r ce of i nn ov a tions in this gene r atio n is belo ng ed t o the introductio n o f Spatial Transfo r mer Netw ork (STN) by Jaderbe r ag et al. i n 2015 (Jade r be rag et al. 201 5). It is an explicit module that can b e injec ted in the dif fe r ent kinds o f DNNs to m ake the f low of data ac r oss the h idde n laye rs b eing tra n sf ormatio n invariant; w h en si tuated n ext to and co llabora ted wit h po oling la y ers that ar e implic itly translatio n and scali n g in v ariance, th ey can be compleme n ta r y to introduc e a complete set of spatia l invariance whos e sy n ergic im pac t can drastic ally enh ance the pe r fo r mance of CNNs applied to m any di ffe rent i mage proc essing applicatio n s , whe r e the medical image r egistrat ion cannot be an exceptio n . The STN, as illust r ate d in F ig. 18, is compo sed of thr ee sequential co mponents. Fi r st, a loc alization n etw ork, w ith very fle xible structure e.g. a regula r MLP, w h ic h lea rns to r eg ress the transfo rmation paramete r s fo r the inputted f eature -m ap bas ed on a predef in ed similarity measure as the l os s functio n . Seco n d, a grid ge n erato r w h o se aim i s at apply in g the estimate d tr ansf ormatio n parame ters by the loc al izatio n n etw ork to the inputte d fe ature -map. Fi n ally , a sample r that wo rks as an interpolato r to co nstruct the final outp utted warped image. Since the STN is fully di f feren tiab le, i t can be inse r ted anyw h ere in the netwo r k, and its locatio n i s co n text -spec ific, and source of disagr eeme nt in the co mmunity . STN is n ot flaw -free; large tra n sf ormatio ns can cause sev ere disto r tio n in t h e o utput, as it is n ot to le ra b le by t he sampler, and boundary int e rpolating is also ve r y hard fo r the sample r si nce some of th e output should be bring from outside of th e input, w h ich is not exist. Rec ently , Li n an d L ucey in 2017 (L in and Luc ey 2017) introduc ed I n ve r se Compos it ional STN (IC -STN) , and ar gue d that we can postpo n e the r ec onstructio n by th e sample r , and se nding transfo r mat ion paramete r s acco m panied w ith t h e o utput w h ere the CNN itself decides how to tr eat wit h the t ransfo r matio n . I n dee d, t h e issue is in its infancy , and the p r o blem is still o pen. A list of works be long to this gene ratio n are p r ese nted in Tab le 3. Loss Train Line L o s s Fixed Image Moving Image Deep Neura l Net work Output Warped Image Transform Parameters Regulari ze r Ground Tru th - 19 - Figure 18 : Spat ial Tr ansform er Netwo r k (STN ) (Jaderbe r g et al. 20 15) Table 3 : T he Sec ond Gen eratio n of Deep L earning A pproac hes fo r Med ical I mage Registrat ion (Superv ised End- to -End Reg istration) Reference Title T ec hnique Modality Transformation Miao et al. (2016) A CNN regression appr oach for real-time 2D/3D re g ist ration CNN Unimodal Rigid Yang e t al. (2016) Fast predictive im ag e re gistration CNN Unimodal Deformable Miao et al. (2016) Rea l -time 2D/ 3D registration vi a CNN regression CNN Unimodal Rigid Miao et al. (2017) Convolutional Neural Net works for Robust and Real-Ti me 2-D/3-D Re g i stration CNN Unimodal Rigid Yang e t al. (2017) Fast predictive multim odal image registr ation CNN Multimodal Deformable Yang e t al. (2017) Quicksilver: F as t p redictive image registratio n - A deep learnin g ap proach CNN Multimodal Deformable Yoo et al. (2017) ssEMnet: Serial-sectio n electron micros copy image re gistration using a spatial transformer network with le arned features SA Es Unimodal Deformable Sokooti et al. (2017) Nonrigid image re g ist ration using multi -scale 3D co nvolutional neural net works CNN Unimodal Deformable Cao et al. (2017) Deformable image reg i stration b as e d on similarit y-steered CNN regressi on CNN Unimodal Deformable Rohe et al. (2017) SVF-Net: learnin g d e formable image re g i stration usin g shap e matching CNN Unimodal Deformable Bhatia et al. (2017) Rea l time coarse orie ntation dete ction in MR scans usin g multi-planar deep convolutional neural netwo rks CNN Unimodal Rigid Zh e ng et al. (2 017) Learning CNNS wit h p airwise dom ain adaption for real -time 6dof ultrasound transducer detection a nd t racking from x -ray im ag e s CNN Multimodal Rigid Pei et al. (2017) Non-rigid craniof acial 2D-3D registr ation using CNN-b ased regression CNN Unimodal Deformable Eppenhof et al. (2 017) Sup e rvised l ocal error estimation for nonlinea r i mage re g i stra ti on usin g co nvolutional neural networks CNN Unimodal Deformable Eppenhof et al. (2 018) Deformable image re g ist ration using convolutional neural networks CNN Unimodal Deformable Zh e ng et al. (2 018) Pairwis e do main adaptation module for CNN -bas e d 2 -D/3-D registration CNN Multimodal Rigid Sloan et al. (2018) Learning Rigid Image Re gis t ration -Utilizing Convolutio nal Neural Networks f or Medical Image Registratio n CNN Multimodal Rigid Sun et al. (2018) Deformable mri -u l trasound re gistration using 3d convolutio nal n e ural net work CNN Multimodal Deformable Yan et al. (2018) Adversarial image reg i stration with applicatio n for mr and trus image fusion GAN Multimodal Rigid Cao et al. (2018) Deep learning based inte r-modalit y im age registration su pervis e d by intra -mod ality similarity CNN Multimodal Deformable Cao et al. (2018) Deformable image re g ist ration using a c ue -a ware deep regression network CNN Unimodal Deformable Onieva et al. (201 8) Diffeomorphic Lun g Re gistration Using Dee p CNNs and Reinfo rced Learning CNN Unimodal Deformable Mahapatra et al. (2018) Joint registration and se gmentation of xray im ag e s using generative adv e rsarial networks GAN Unimodal Deformable Mahapatra et al. (2018) Deformable medi cal i mag e registration u sing generative a dv e rsarial networks GAN Multimodal Deformable Sentk e r et al. ( 2018) GDL-FIRE 4D: Dee p Learning-Based Fast 4D CT Image Re gistration CNN Unimodal Deformable Sun et al. (2018) Towards Robust CT-Ul trasound Re g i stration Using Deep Learning Methods CNN Multimodal Deformable Salehi et al. (2019) Rea l -time deep p ose e stimation with geodesic loss for image - to -template rigid reg i stration CNN Multimodal Deformable Elmahdy et al. (201 9) Robust contour prop aga ti on using deep le arning and image registration fo r online adaptive proton ther apy of prostate can cer CNN GAN Unimodal Deformable Liu et al. (2019) Multimodal medical image reg i str a tion via common representatio ns learning and differentiable geometri c constraints CNN Multimodal Deformable Foo te et al. (2 019) Rea l -Time 2D-3D De formable Re gistration with Deep Lea rning and A ppli cation to Lung Radiotherapy T arg eti ng CNN Unimodal Deformable The third generatio n belo n gs to the Dee p Re in fo r ce ment Lea rning (DRL ), w h ere just like Fig . 1 4, the deep age n t (o r multiple agents) l ea rn s to produce the final transf ormatio n step-by -st ep so that the positiv e fee dbac k from the envi ronment - 20 - (here f r om the si milarity measure) c an b e maxi mized. I n stead of th e f ir st deep simila r ity measure paradigm, t h e s imilarity measu r es are r outi n e ly pr ov i ded in a co n venti o n al w a y e.g. N orm alize d M I (NMI) o r Lo cal Cross Co rr elatio n (L CC). Th e mos t co n fini n g fac tor to extinct the gene r atio n o f t hi s parad igm is t he inab ility of the a ge nt s to inte ra ct w ith t he h uge sta te space introduc ed by deformable regist r atio n field. Wit h out the ability to captu r e def ormation ess ential fo r prospe r ous registratio n of elastic o r gans as w ell as r elativ el y prolonge d registrat ion time, the paradig m is doo med to co n structio n. A list of wo r ks be long t o this ge n e r atio n ar e prese n ted in Table 4 . Table 4 : T he Th ird Gener atio n of Deep Lear ning A pproac hes fo r Medic al I mag e Registratio n (Deep Reinfo r cem ent L earning) Reference Title T ec hnique Modality Transformation Ma et al. (2017) Multimodal ima g e re gistration with deep con te xt reinfo rcem e nt le arning DRL Multimodal Rigid Krebs et al. (2017) Robust non-rigid registr ation through ag e nt-based actio n learning CNN Unimodal Rigid Liao et al. (2017) An Artificial Agent fo r Robust Image Registratio n CNN Unimodal Rigid Toth et al. (2018) 3D/2D model- to - image re gistration by imit ati on le arning for cardi ac procedures CNN Multimodal Rigid Miao et al. (2018) Dilated FCN for multi -agent 2D/3D medical image registration CNN Multimodal Rigid Sun et al. (2019) Robust Multimod al Im age Re g istratio n Usin g Deep Recurrent Reinfor c e ment Learning CNN and RNN Multimodal Rigid As th e previo us generatio ns were based on th e g r ou n d -truth to co n st r uct the mode l, and gene rall y the ann o tated datase ts i n medici n e, and spec ifically for image r egistratio n , a r e small-sized , n ot sui ta b le fo r t h e exhaustiv e deep l earning, the fo urt h gen e ra tio n belo n gs to the u n supe r vise d end- to -e n d r eg istratio n , w h ere diff eren t kinds o f DNNs ar e trained w ith out a n y gr o und-truth to co n struct r egress ion mo dels to pro duce the t r ansf or matio n pa r amete r s i n o n e-shot . Instead of using enormo us grand-truth set, they use data augmenta tion techniques on a fe w num be r of in putte d samples as see ds w h ere a traditional similarity measur e (or a comb in atio n of th em) is use d as los s function to guide the lea rning p r oc ess , as illustrated i n Fig . 1 9 . Mo st of th e approac h es in thi s gene rati o n ha v e bee n succ essf ul on unimo dal r egist ration, w hile multimo dal r egistrat ion is fa r more co mplicated as multimo dal simila rit y measu r es are still ineff icient, and a n etw ork trained o n them inh e rits this inef ficie n cy , acco rdin gly . Wu et al. (2016) ca n be a goo d r eprese ntative w h e r e a SAEs is trained t o extract the fe atures, and a CNN makes the final tra n sf ormation esti mation. U tilizatio n of SAEs instead of co nventional mu ltimodal simi larity measu r es l ike M I is an e vidence to ou r af o r ementio n ed argume n t. W hi le sy n th esi z ing fake samples by data a ug mentatio n tec hn iques, the rule of regularizatio n term is c r itica l to co n trol the applied defo rm atio n s in orde r to be r ea listic. Yet, practitio n e rs and e xperts are dub ious on this regard, and we expec t a w in di n g w a y ah ead wit h a signif icant researc h focus in th e near f uture. A list o f works be long to this generatio n ar e presente d in Table 5 . Figure 19 : The m ai n f ramewo r k fo r unsu pervised end- to -end me dical im age reg istratio n Train Line Loss Fixed Image Moving Image Deep Neural Net wor k Conventio nal Similari ty Metrics e.g. NMI or LCC Output Warped Image Transform Parameters Regulari ze r Sampler - 21 - Table 5 : T he Fo urth Gene r atio n of Dee p Lea rning Appro aches fo r Medic al I mag e Registratio n (Unsu perv ised End- to -End Reg istration) Reference Title T ec hnique Modality Transformation de Vos et al. (2017) End- to -end uns up e rvised defor mab le image re gistration with a convolution al neural network CNN Unimodal Deformable Li et al. (2018) Non-rigid image reg i stration usin g sel f-supervised fully convolution al networks without training data CNN Unimodal Deformable Dalca et al. (2018) Unsupervised le arn i ng for fast prob ab ilisti c diffeomorphic re gistration CNN Unimodal Deformable Ba l akrishnan et al. (2018) An unsupervised learnin g model for deform ab le medical ima g e r e gistration CNN Unimodal Deformable Shu et al. (2018) An unsupervised net work for fast microscopic im age registration CNN Unimodal Deformable Stergios et al. (2018) Lin e ar and Deformable I mage Registr ati on with 3D Convoluti onal Neural Netwo rks CNN Unimodal Deformable Krebs et al. (2018) Unsupervised pr obabilistic deformation modelin g for robust dif feomorph i c registration SA Es Unimodal Deformable Kearney et al. (2018) An unsupervised convoluti onal neur al net work-b ased algorithm for deform able image reg i stration CNN Unimodal Deformable Sheikhjafari et al. (2018) Unsupervised deform ab le image registr ation with fully connected gene rative neural network SA Es Unimodal Deformable Ferrante et al. (2018) On the adaptability of u nsupervised CNN -based deform ab le image registr ation t o unsee n imag e domains CN N Multimodal Deformable Ito et al. (2018) An Automated Metho d for Generating Trainin g Sets for Deep Learning b ased Image Registration. CNN Unimodal Deformable Ba l akrishnan et al. (2019) VoxelMorph: a lear ning framework for deform ab le medical im age registration CNN Unimodal Deformable Yu et al. (2019) Learning 3D non- rig i d deformation based on an unsupervised deep learnin g for PET/C T imag e registration CNN Multimodal Deformable Che et al. (2019) DGR -Net: Deep Groupw i se Registration of M ultispectr al Images CNN Multimodal Deformable Van Kranen et al. (2019) Unsupervised deep le arn i ng for fast and accurate CBCT to CT de formable image reg i stration CNN Unimodal Deformable Che et al. (2019) Deep Group-Wise Re gistration for Multi -Spectral Images Fr om Fundus Images CNN Multimodal Deformable Hering et al. (2019) Unsupervised le arn i ng for large motion th orac i c CT follow -up re gistration CNN Unimodal Deformable Duan e t al. (2019) Adversarial learning for de formable registration o f brain MR image using a multi-scale fu ll y convolution al network CNN Unimodal Deformable de Vos et al. (2019) A Dee p Lea rn i ng Framework for Unsu pervised Affine and De formable Image Registration CNN Unimodal Deformable Krebs et al. (2019) Learning a probabilisti c model for di ff e omorphic regist ration SA Es Unimodal Deformable Finally , sin ce bo t h supervised and unsupe r vised end - to -en d image r egist rati o n have th ei r o wn drawb acks, th e fif th generatio n b elongs t o the w eakl y /semi -supervise d app r oache s. There are two di ffe ren t key para digms in this catego r y . So me approaches a re based o n t h e ful ly ann ot ated g r and-trut h data w ith as man y la n dma r ks as po ssible . Ro ut in ely , these landmarks are co n tours , l egio n s, co rner s , lin es, turni ng points and so on eac h of which get s a d istinct class label. The netwo r k trained on these fully labe led data ; h ow e v er, to be a few. B esides it s mai n duty i. e. image r egis tration, it learns to detect la n dma r ks in an y pair of in putted i mages . Dete ctin g suc h kin d s of l andm arks ar e key to co n st r uct ef ficie n t mode ls, an d to e nhance the accurac y of s yste m. In additio n, Target Re gistration Error (TRE) that i s the most precio us structu r al si milarity measure ca n be used a s th e los s function t o train the netwo r k , w h ich is n on-t r ivia l . Hu et al. (2018 ) can be co n sidered a s the most co mplete r ep r ese ntative for this paradigm w h e re the framew ork is illust rated in Fig. 20 . Anot h er paradigm is base d on th e utili zation of Gene r ative Adv ersari al Netwo r k (GAN) (Goo d fellow et al. 2014) (Fig. 12) w h e r e the gene r ato r takes th e input ted fixed a n d mov ing images , and try to produce such transfo r matio n pa ramete r s so th at the tran sf ormed mov in g image ca nn ot be discriminate d from the ground-truth by th e discrim inator, the situ a tio n expe cted from an expert r egist ration a ge nt. Base d on th e survi val compe ti tio n be twee n the ge n erato r and the disc r iminato r, just like the game theo r y , the n etw ork can be trained o n a small se t of data so that the gene rated samp les ca nn ot be discrimina ted, and t h e netw ork goes fo r equi li b rium. A list of wo r ks be long t o t his gene r atio n are prese nt ed in Tab le 6. - 22 - Figure 20 : The m ai n f ramew ork fo r weak l y- supervised label-dr iven med ical im age reg istrat ion ( Hu et a l. 2018) Table 6 : The Fourth G eneration o f Dee p Lea rning A ppr oac hes fo r Medic al I mag e Reg ist ratio n (Weak ly /Semi-Supe rvised En d- to - End R egistratio n) Reference Title T ec hnique Modality Transformation Uzunova et al. (2017) Training CNNs for im ag e r e gistratio n from few samples with model -based data augme ntation CNN Unimodal Deformable Salehi et al. (2017) Precise ultrasound bo ne registration with learnin g-based segment ation and spee d of sound calibration CNN Multimodal Deformable Hu et al. (2018) Weakly-supervised co nvolutional neural net works for multimodal image registration CNN Multimodal Deformable Fan et al. (2018) Adversarial simil ar it y network for evalu ating i mage alignment in dee p learning based reg i stration GAN Unimodal Deformable Hu et al. (2018) Lab el -driven weakly-supervise d learning for multimodal def ormarle image re gistration CNN Multimodal Deformable Hu et al. (2018) Adversarial defor mation regularization for trainin g image registr a tion neural networks GAN Multimodal Deformable Hering et al. (2019) Enhancing Label-D riven Deep Deform ab le Image Registration with Local Distance Me trics for State- of - the -Art Cardi ac Mo t ion Trackin g CNN Unimodal Deformable Fan et al. (2019) BIRNet: Brain image registr ation using dual -supervised f ully convolutional net works CNN Unimodal Deformable In th e fo llowing, some breakthroug h s , ge n eratio n s’ turning points and represe n tative s ar e r ev iewe d in more detail, and thei r netwo r k st r uctu r es, advantage s, disadv antages, novel ideas and key c ontributio n s are p r ese nted. (Wu et a l. 2013) should be pr ope rly conside red as the fir s t tr y to apply dee p n eu ra l netwo r ks to medical image registratio n. The autho r s used a comb ination of CNN an d Inde pendent Subs et Analy sis (ISA) t o extr act prope r featu r es from in pu tted i mage s. As illustrated in Fig. 21, the unde r ly in g ar c h itectu r e was a 2-lay er n etw ork w h ic h takes input via both the lay ers. A lso, a h iera r chical trai n ing m ec h anism w a s used where sm all-sized patc h es in the size of 13 ×13×13 voxe ls were fee ded to the fi r st la y er, an d th e first laye r w as trained ac co r dingly . Af terwards, bigge r -sized patc h es in the size o f 21×21 ×21 vo xels using sliding w indow w i th o ve r lap were f ee ded to the sec ond laye r to train t h e se co nd lay er . Th e output of the netwo r k f or e ach inputted patc h w as a 1 50 -fe ature ve cto r that w as prov ided for two co n v enti onal regist ration algorithms namely H AMMER (Vercauteren, et al. 20 09) and Dif feo morphic Demo ns (S h e n 2007), a n d t he f i nal registratio n was ac hi ev ed . T h e expe rimental study was co n duc ted on two differ ent datasets of IXI and ANDI co n taining MR i mages of human brain co n side r ing Dice co efficie n t as th e compa r iso n metric . R esults from both the registratio n algorithms on bo t h the afo rementioned datasets s h o w ed tan gible imp r ov ement consideri n g si gnifica n ce test with 95% co nfidence inte r val. - 23 - Figure 21: The architec ture u sed in (Wu et al . 2 013) Cheng et al. (2016) trained a 5-laye r SDA Es to approxim ate t h e amou n t of simila r ity betwee n the couple of inputted CT and M R images of b r ain. Actually , the co n ve n tio n al s imilarity measure w as sub stituted by th e p r opo sed netwo r k. The underly in g r easo n w a s t h at multimodal similarity measures l ike No r malize d Mu tual I n fo r matio n (NMI) and L oc al Cros s- Correlatio n (LCC) are far fro m the co mpleteness, and have a lot o f l o cal minima in so me applica tio n s. Th e input of th e netwo r k, as illustra ted in Fig. 22 , is a co uple of corr espo n di ng patches from inputted f ix ed and m ov i ng images, and the binary output indicates the co rr es ponde n ce /non -co rrespondence b etw een these p atches. In addi tio n , t h e o utput befo r e the Sigmoid activat ion fu n ctio n was extracted an d used to compute the simi larity measure. To trai n the bi n ary n etw o r k, the inputted CT a n d MR images we r e al igned rigidly by the au tho r s. Then, t h e co uple o f in putted images we re normalize d to the zero mean and unit variance. To co n fi ne training data, patc h es w ere ext racted from the central parts and arou n d the scull co n sideri ng the fact that edges an d co rn e rs have the most info r mative and disc r imi native data. 2000 co rr espo n ding patc h es an d 2000 no n -co rr espo n ding ones in the size of 17×17 vo xels were extracted to train the sys tem. The propo sed netwo r k w as evaluated ve rsus LCC a n d NM I o n 300 patc h es as test data using cumu lative sum o f predictio n e rr o r a s t h e perfo rm ance metric, and it show ed significa ntly better pe rformance. Figure 22: The architec ture u sed in (Ch eng e t al. 2016) Simonov sk y et al . (201 6) also p r o pose d a deep learni n g m o de l to use a s a m ult imodal si milarity measure, but despite the (C h eng et al. 2016) , a CNN was e xploited. Agai n , the r easo n was the same i. e . def iciency o f th e conventional multimo dal simi larity measu r es for all th e moda lities a n d all the o r gans o f interest . At first, a rigid a li g nment was applied to the inputted images manually . The n, a la r ge numbe r of co r respo n di ng and n on-correspo n di n g patc hes in the size of 17×17×17 vo xels we re extracted from input ted image s, and the n etw ork was t r ai n ed acco r dingl y . The n etw ork was a 5- lay er 2 -channel CNN w i th a b out two mill io n w eight s . It was dete cted using stride in t h e po oling laye rs a s w ell a s H in ge los s- functio n (i nstead o f cro ss-entr opy ) can co n tribute to t he traini n g co nvergence and t h e ove r all perfo r ma n ce . Since the sy stem w as feeded by th e co uple of pat ches f r om fix ed and mo ving images as an i n teg r atio n , it could not exploit t h e fac t that the p at c h f rom fix ed i mage doe s n o t n eed an y ma n ipul ation. This fact was exp loited in the n ext wo r ks by o th er a utho r s - 24 - using late fusio n in th e n etw ork. Using late fus ion, the pro ces sed da ta over the c hann els / pipeli ne do n ot fused until the late lay ers of netwo rk i.e. w e postp o n e sharing t he cha nn els’ weights u n til t he last la y ers . ALB ERTs data se t w as used fo r the ev a luation study containing aligned and labe led T1 and T2 -we igh ted MR images fro m the b rain of 20 infants where there w ere 50 se gmented anato mical regio n s l abele d in e ach image. To w orsen t h e s ituatio n, t h e sy stem w as trained b ased on the IXI datase t th at co n tai n s 600 alig n ed T1 and T2 - w eighted MR i mages of adults’ b rain. The images of i nfants and adults h ave some differences in anato my helping us b ette r estimating the generalizatio n potenti ality of th e p r opo s ed sy stem. The propo sed approac h w as co mpared versus Mutu al I n fo r matio n (MI ) , and th e r esults showe d its su periority w ith 99 % confide n ce inte rval. In addition, the running t ime used to r es ponse the netw ork was a bo ut 2 time s slow er than MI w h ic h sti ll indicates ac ceptable t ime fo r cl inical use . In all t h e afo r ementio n ed w orks, dee p learni ng m o dels were used to w ork as a s imila rit y measure b eside s a co nventional ite r ativ e approac h exp loit ing this measure. For the first time, in a seminal wo r k, Mia o et al. (2016) used a CNN to di rectly r eg ress the r est ricted Af fine transfo r matio n p arameters to r egis t er 2D X -ray ima ges o n 3D CT o n es; how ever, the ir app roach w as b ased on the pa r ticle s impla nted i n t h e p a tie nts’ b od ies, as illust rated i n F i g. 5. A t fi r st, t hey co nsider ed an at te ntion map f r o m the 3D CT image , and ex tract ed a Digi tally Reconstructed R adiog r aph (DDR ) from it. The duty of th e pr o pose d approach was to r egis t er th e extracte d DDR on the operatio n al X-ray ima ges. To enhance the sy stem, th ey used the follo w in g three st rat egie s: Fi r st o f all, a n ov el similarity m easu r e named Loc al Im age Re sidual (LIR ) was used, whic h could ef fec ti v el y keep its correl ati o n w ith the d ifference cause d by changes in tra n sf ormatio n parameters . I n o ther wo r ds, it was a be tter es timatio n o f diffe r ent alig nment s of the inputted images . S eco n dly, to dec r ease the co mplexity of regist ration, th e images w ere seg mented and deco mpos ed to a predef in ed n umb er of r egio n s w h ere each regio n had i ts ow n CNN; h ow ever, thi s strategy significantly inc r ease s the comput ational burde n. Thirdly , parameters regress ion was m ade h iera r chical ly to in crease the precisio n. That i s , the 6 transfo rm atio n paramete r s w ere divide d t o 3 diff er ent sets of eas y , mode r ate and h ard, and would be calculated fro m easy to h ard. As illustrated in Fig. 23, a t fi r st, the propo sed a pproac h ext ract ed N patc h es ar ou nd the implanted partic le. The r e was a CNN fo r each p atch, co mposed of 2 convo lutiona l la y er s, 2 poo l in g laye r s, and a fully -conn ecte d laye r w ith a 100 -bi t output. The outputs of all t h e CNNs w ere gat h e r ed, concate nated a n d f eede d to a final 2-lay er fully -c onnected n etw ork that results i n the fi nal 6 transfo rmati o n p arameters, 2 fo r t ran s lation, 1 fo r scali ng, an d 3 fo r r otatio n around the co ordination axe s. T hr ee datase ts named Total K n ee Arthroplasty (TKA), V isual Impla n t Planning Sy stem (VIPS), and X -ray Ech o Fusio n (XEF) we r e used fo r co mpariso n study . Also , a nov el ev aluat io n metric named mTREproj denoting mean target r egis tration e rr or fo r image projec ti o n from 8 corne r s of im planted particle was considered fo r e valuating the approac h es. The propo sed approac h was compa r ed ve r sus dif fe ren t variants of th e co n ventio n al Pow ell m et h o d, and it show ed sensible supe ri ority ove r all t h e t hree datasets. The r unni n g time w a s ab out 100ms on a ty pical syste m show i n g hig h potentially for r eal-time clinical use . Figure 23: The architec ture u sed in (Mi ao et a l. 2016) While the p r ev ious wo r k co n sidered only Affine tr ansf ormatio n , in anot her seminal wo r k, Soko oti et al. (2017) w ere succe ssful to trai n a CNN to reg r es s a Displac ement V ecto r Fie ld (DVF) cap able of defo r mablely register ing t h e input ted images in o n e shot. An ov erview o n the mecha ni s m of DVF is illus trated in Fig. 24 . They extra cted a large number of co rr espo n ding patc h es (abo ut 2100 patches eac h image) based on the semi -auto matically dete ct ed landma r ks f r o m the co uple of in putted ima ges, and fee ded th em to th e netwo r k (Fig. 25). The patches we r e in the size of 29×29×29 voxe ls, w h ile so me other case s w ere extracted in the size of 54×54×54 vo xels to k eep th e r ece ptive field. These bigger patc h es were co mpr esse d to the half (to decrease the c omput ational inte n sity ), and t h en w er e fe eded to the n etw ork v ia a diffe ren t - 25 - pipeline. Fo r the f ir st time, late fus ion w as used so tha t the diffe r ent -size d patc h es used diff erent pipeli n es in t he netw ork until the la s t lay ers, whic h could have a go od co ntr ibutio n to the applicabil ity an d pe r fo r mance. The first and sec ond pipelines (fo r 29 ×29×29 a n d 54×54 ×54 v oxels patches, r espe ctive l y ) we r e co mposed o f 3 s equential co n v olutional lay ers w ith 3×3×3 ke rn els fo l low ed by 2×2×2 po oling lay er s. Afte rwards, the fi rst a n d se conds pipeli n es w o uld be subje cted to 6 and 2 1×1×1 co n v olutional lay ers, r espec tive l y , to b eco m e the same-s ized. Concate natio n was do ne at this poi n t, and after 4 o ther convo l utional lay ers, the re was a 2-lay er fully -conn ecte d n etw ork to p r oduce t he f inal 3 tran s latio na l parameters fo r each patc h es. Th e SPR E AD da tase t containi n g 19 co upl es of 3D CT images from chest w as used to ev aluat e the propo se d appr oac h whe r e 10, 2, and 7 of them were used fo r training, validat ion and testing, respec tively . The Mean Ab so lute Err o r (MAE) metric w as used fo r sy stem traini n g w h ile Target Re gistration Erro r (TR E) was the ev aluat o r o f th e approac h es in the compa r ison study . Th e pr o pose d appr o ach was ev al uated versus Affine, r e gula r B- Spline, and 3-reso luti o n B-Spli n e , and it outpe r fo rm ed the Affine and r egu lar B-Spline m et h ods , an d w as co mpetit ive w ith the 3- resulation B -Spline, one o f the stro n gest met hods in the lite r ature. Figure 24: A n overv iew on the mechan i sm of DV F F igure 25 : The archit ecture use d in ( Soko oti et al. 2017) - 26 - H u et a l. (2018) trained a CNN f or defo r ma ble ly registeri ng p r e-interve ntional multi-paramet ric MR images to ope ra tio n al Transrectal Ultraso un d (TR US ) of th e pros tate gland in order to decrease the ri sk o f i mage-guided inte r ve n tio n. Si n ce the captu red TRU S i mages are in l ow -qua lity with a high leve l of n oise , fusi n g the m w ith th e h igh- quality 3D MR im ages can cause s y n ergic effec t to des ir ably gui de and suppo rt the inte r ve n tio n p r oc ess . The prob lem w as that MR I and T R US do n ot have represe n tatio n al a bo nn eme nt exc ept in the l imite d cases, whic h challe nges th e registratio n proc ess . A cc o r dingly , utiliz ation of conve n tio n al a pproac h es w as n ot possib le, and it justified t h e exploitatio n of machi n e lea rning approac h es. Since the re was not a n y compre h ensive l abe led dataset in this regard, a n d utilizatio n of inte n sity -base d similarity measures ca using mec h anical r egist rat io n s has ambigui ty aspects fo r the phy sicians and expe r ts, a datase t that w a s partial ly man ually labe led by th e experts w as used to train t he netwo r k. Final ly , co r r espo n ding labeled structu r es w ere exploited by th e n etw ork to produc e an exact vo xel - vise r egistration. Th e promisi ng point is that the sy stem im p licitly learn ed to detec t the structures- in -interest afte r trai ning; h ence , the need to manual operation in th e utilizatio n phase is o bviated. The similarity measu re used b y th e propo sed approac h w as multiscale and base d o n th e registratio n distance of the correspo n ding lab eled l andm arks an d structu r es in th e inputted images . As illust rated in F ig. 26, th e propo sed 3D CNN was compo sed of 4 dow n sampling bloc ks fo l lowed by 4 upsamp ling ones jus t like t y pical U- Nets, but with fa r more co nn ec ti o n s to keep the impact of back-pr opag atio n over a la r ge numbe r of lay ers . Th e n etw ork output was a Dense Displ acement Field (DD F) fo r r eg istering and f using whole the MR image o n the co ntinues TRU S ones . Th e Sma rtTarget datase t conta ining 1 08 couple of T2 -we ighted MR I and TRU S w as used in whic h eve r y pat ie n t has three ki n ds of images base d o n h is/he r treatment pla n. R ese ar cher fe llowships and students co n su med 200 h ours in total to detect an d labe l 83 4 couple o f a n ato mical landm arks. 12 -fo ld c r o ss-validatio n a s we ll as Dice a n d T R E me asures were used to t est the sy stem, whe r e the r esults , su pported by stat istical Wilco xon r ank -sum test with 95% co nfide n ce , ce r tified the significant supe r io rity of th e propo sed approach. F igure 26 : T he arc hitectur e used in ( Hu et a l. 2018) While the prev ious wo rk is h ighly appreciated a n d v aluab le since the aut h o r s e n te r ed the time as the 4 th dime nsion of TRU S in o r de r to made th e prob l em of registration far m o re c omplica ted, the w ork of de Vos et a l. (2019) s h o uld be co nsider ed as the mos t co mpr e h e nsive deep learning frame work base d o n the CNN to directly r egress the defo rmable transfo rmati o n pa r amete r s in o n e shot. In additio n to r egres s th e transf ormatio n paramete r s , their n etw ork w as able to learn a predef in ed simila r ity m easure so that the neces sit y t o util izatio n of s y nt hesized and labe led da taset is obv i ated, w h ich is a big progress in apply i ng DNNs to the field of medic al image an aly sis w h ere we a re faci ng sm all -sized annotated datase ts as r egu l ar. As illustrate d in Fig. 27, the propo sed m et h o d w as a multi -reso l utio n m ulti-stage 2-channel CNN- bas ed approach. In the fi r st stage, a 2 -channel CNN with 5 laye rs of 3×3× 3 kernels fo ll ow ed by 2×2× 2 ave r age poo l ing lay ers too k th e inputted images. The we igh ts w er e shared be twee n lay ers to dec r ease the numbe r o f the netw orks’ w eigh ts. At the e nd, a co n cate n ating lay er followe d by a 2 -lay er fully-c onn ec ted n etw orks pro duced th e 12 pa rameters of A ffine transfo rmati o n . I n the seco n d stage, anot h e r CNN w as use d to made the final def ormable r egist ration base d on th e B - - 27 - Spline. At first, a n umb er of pat c hes based on the specif ied landma r ks were extracted and fe eded t o the n etw ork. The segme n tat ion w as b ased o n t he (Lo ng et al. 2015) p r ov i ding i mage analy sis w ith arbit rary size. T h e netwo r k w as co mposed of in te rl eav ing 3×3×3 c onvo lutional laye r s fo ll ow ed by 2×2×2 poo l ing ones. The n umb er of lay ers wa s variable bas ed o n the n umb er of contr o l points in the def orm able mesh (i.e. based on the space am o n g control po ints) . Af ter the final poo l in g la y er, there w ere 2 la y ers of co n vo luti on to in c rease the rece ptive field, whic h is n ov el contrib ution to set th e CNN arc h itec ture. Finally , a 2 -lay er f ully-co n nected n et work reg r ess ed th e B-Spli n e co n trol poi nt loc ati o n s that was used a s a refe r ence t o produce t he correspo n de n ce DVF. Tw o different datasets w ere used to evaluate the propo sed approac h. The first o n e was Sunny brook Cardiac D ata (SCD) co n taini ng 900 MR images of 45 patie nts wit h 4 d ifferent states of heart attack. Fo r eac h patie nt, 20 images c ove r ed a full cy cle of heartbeat . The sec ond dataset was co mpose d of 2060 CT image s randomly selecte d from th e Natio n al Lu n g Sc r ee n ing Trial (NL ST) w ith diff erent size a n d qua lity . T he propo sed a pproac h was ev al uated versus the we ll -kn ow n Ela stix (Marst al et al. 2016) using D ice coeff icient, H ausdorff distance and Av er age Su r face Distance (ASD) as the c ompari so n metri cs, w here t he propo sed app r oac h o utperfo r med the Elastix in ma n y cases while 350 time faste r . A f ull def ormab le r egist r atio n on th e 4 -co r e Xeo n E5-1620 proc essors and NVIDIA Titan-X GPU was reported to take abo ut 39 mil li se co n d (ms), w h ich is highly appreciated fo r r eal-time cli n ical use. Figure 27: The architec ture u sed in (de Vo s et al. 2019 ) 6. 1. The Co mparative Analysis , Adv antages, Disad vantages, Main Co ntributio ns and No veltie s As stated in Sec tion 6, we have faced five diffe r ent ge n e r at ions o r ca tego r ies fo r medical image r egist rati o n using dee p n eu ral netwo r ks base d o n e the achiev ements and b r eakt hroughs face d by the commu nity. Th ese catego ri es can be enumerated as D eep Simila rity Metrics (DSM) , Supervised End - to -End Re gistratio n (SE2 ER), D eep Re in fo r cement Le arn ing (or Age nt -Bas ed Regist r atio n ) (DRL ), Unsupe rvised End- to -E nd Re gistratio n (UE2ER), We akly/Semi- Supervis ed End- to -End R egistration (WSE2ER ), eac h of which w ith diff eren t paradigm t o enco unt e r the r eg istratio n prob lem. The numb er o f works in each ca tegory a nd a timeli n e from start to the end are dep icted in the Fig. 28 and Fig . 29, respe ctiv el y . - 28 - Figure 28: D ifferent g enerat ions of deep le arning a ppro aches fo r medic al imag e registrat ion w it h their fre quenci es Figure 29: D ifferen t gen e rat ions in timelin e At first, the topic starte d w ith Dee p Similarity Metr ic (DS M) a pp r oac h es in 2013 where diff erent kinds of DNNs trained to learn visual similarity metr ics f r om a l arge set of pa ired annotate d grou n d -truths . Th e learned mode l a fte r the train was ab le to prec isel y a n d mea ningfully m o del th e struc tural diff er ence be t w een the input ted pair of i mage s/patches spec iall y fo r de fo r ma b le tra n sf ormation with diff erent modali ties, w h e r e co n ve n tio nal similarity metrics (supposedly with the exc eption of MI) had a lot o f diff i culties. Two main draw bac ks to this paradig m are that It i s depe n dent o n a large set of paired annotated g r ound-trut h s to trai n th e netwo r k, w h ic h rarely is the case fo r medical a pplica tio n s. It is still depe n de n t to co nventional i te r ativ e optimization-b a sed app roaches that are ve r y slow and im prac tical for clinical use. Supervis ed End- to -End Re gistration (SE2ER), sta rted in 201 6, w as a signif icant milesto n e that the commu n ity face d since it obv iat es t h e co mputatio nal burde n and ti me inconv eniency of co n ve n tional i terative r egis tratio n approac h es, and by conducting the registratio n p roce ss in o n e-sho t, p racticall y makes the rea l-time c linical use poss i ble. It started in 2016 by Mia o et al (2016) fo r rigid registratio n, exte n ded in 2017 by Sokooti et al. (2017) fo r deformable registratio n, and is the mains tream catego ry th at is rem ain ed active so far. Agai n, the main p r ob lem to this paradigm is that It i s depe n dent on a large set o f pair ed a nnotated g r ound-trut h s to train th e n etw o r k, w h ic h is a sev ere hindra n ce to dev el op any appr oac h in this catego r y . Actually , a lar ge n umbe r o f authors abandoned this catego r y , a n d w ent to try th eir c h ance o n o ther diff erent paradigms w h ere the data annotatio n is out of th e case. Deep Re info r cement Le arnin g (DRL ) (or Agent-B ased Re gistra tio n) is one of such paradigms , where a deep agent (or multiple age nts) learns to produce the final tra n sf ormatio n step -by - step so t hat the po siti v e fee d bac k from the e n viro n me nt (here from a simil arity measure) ca n be ma xim ized. I n stead of the fi r st deep si milarity measure approac hes, the si milarity measur es are rout inely pr ov i ded i n a conve n tio nal w ay like NMI o r LCC. The most co n f inin g facto r to dev elop the generatio n of thi s p ara d igm is DSM , 16 , 20 .0% SE2ER , 30 , 37.5% DRL , 6 , 7.5% UE2ER , 20 , 25.0% WSE2ER , 8 , 10.0% Deep Si milarity Me trics (DSM) Super vised End-t o-End Registra tion (SE2ER) Deep Rein forcement Learning (Agent- Based) ( DRL) Unsupe rvised End-to-End Registration (UE2ER) Weakly/Semi-S upervised E nd-to- End Re gistration (WSE2E R) DSM SE2ER DRL UE2ER WSE2ER 2013 2014 2015 2016 2017 2018 2019 Deep Si miliarity Met rics (DSM) Super vised End-t o-End Reg istration (SE2ER) Deep Rein forcement Learning (Agent-Ba sed) (DRL) Unsupe rvised End-to-End Registration (UE2ER) Weakly-S emi-Supe rvised End-to -End Re gistration (WSE2ER) - 29 - Th e in ability o f th e age n ts to i nteract w ith t h e huge st at e-spac e introduc ed by def ormable r egistrat ion f i eld w here, as it can b e see n in Tab le 4, al l the app r oac h es w er e p r opo sed fo r ri gid registrat ion. While th ey were sti ll depe n dent to gr ou n d-truth to tra in the agents. To circumv en t th is prob l em, Unsupervise d End- to -End Regist r atio n (UE2ER) paradi gm w as in troduc ed, where dif ferent kinds o f DNNs ar e trained wit h o ut any gr ound - truth to const r uct r eg ressio n mode ls t o produce th e transfo r mation pa ram ete r s in o ne -shot. I n ste ad of using enormo us grand-t r uth se t, they use data augmenta tion tec hni que s on a few n umb er of in put ted samples as seeds where a traditional similarity measure (o r a co mbination o f t h e m ) i s used a s los s fu n ction to g uide the l ea rning proc ess. Most o f the approac h es in this ge n eratio n have bee n succ essf ul on unimo da l r egist ration, w h ile multimo dal r egist ration is fa r mo re co mplicated, and ca n be regarde d a s the main chal lenge o f this catego r y , be cause Multimo dal simi larity measu r es that are us ed as lo ss f unction to co n duct t h e n etw ork ’s le arning are s ti ll ineff icient, and a n etw ork trained on them , acco rdin gly , inh e ri ts t h is i n ef fic ienc y . On this basis, this catego ry needs t o be loo kin g fo r w ard to the in troduc tio n of muc h ef ficie n t and pow erful n ov el similarity measures in the n ear future. Where bo th supe r v ised and unsupe r v ised ma nner have their o w n drawb a cks , We akl y /Semi-Supervise d End- to -E n d R egistratio n (WS E2ER ) found it s way from 2017. It o bviates th e s h ortc omi n gs asso ciated to th e two afo r ementio n ed paradig ms while inheri ts the stre ngths of bo th. So me approac h es are label -driven, i.e. b a sed on a f ew f ull y -ann otated g r ound-t r ut h sa mples, they can impl icitly lea rn to de tect m any paired lan dm arks in the inputted image, and conduct the r egist r atio n proce ss, a cco rdin gly . So me oth er a p proaches are dual-supe r vise d base d on the simil arity measures (jus t like unsupe rvised a pp r oac h es) an d a few grou n d-t r uth samples to fin e-tune the n etw ork. I n addition, it has be en demons trated that h avi n g a few gr ound-t r uth sample s, tran sf er l ea rning from ot her bo d y organs o r modalities , is fully pra ctical fo r medica l image registratio n ( Cao et al. 2017) and (Ferrante et al. 2018). Finally , anot h e r approac h es use GANs as u nderly i ng technique , whe r e the compe titive interactio n be tween the ge n erato r and discri minator needs a few g r o un d-t r ut h samples to cons truct and mature the mode l. Ac tually , w eakl y / semi supe r visio n ca n be co n side red as th e be st practica l paradigm so fa r, w h ere w e expect the signif icant researc h fo cus in the n ear futu r e. The prog ress of this paradi gm is h eav ily dependent of th e t h eo retical progress and b reakthroug h s i n the b r oade r fields of ma chi ne vision, image proc essing, mac hine learning and pa tte rn r eco gn itio n , from w hich m os t of the p r ogress es in the fiel d of m edic al image r egistra tion have bee n inspired. Table 7 is a big pictu r e on these fiv e di ff eren t catego ries as th e t axonomy c onducted on the lite r atu r e. Table 7 : D eep L earning Appro aches for Medic al I mag e Registrat ion: Par adig ms, Fram ewo rks, Ref erenc es, Adv a ntages and Disadv a ntages Paradigm Framework List o f References Advantages Disadvantage Deep Similarity Metrics (DSM) Fig. 16 Tab le 2 The constructed model afte r t he train on ground -truth, whenever sufficient, were able to outperform the trad i tional similarity metrics specially for multi modal reg i stration. 1. The approaches are depe nd e nt on a l arg e set of paired annotated ground -truths to train t he network, which rarely is the case for medi cal applications. 2. The p aradigm is still dependent to conventio nal iterative optimization-b as e d approaches, wh ic h were very slow and impra c ti cal for clinical use. Sup e rvised End - to -End Registration (SE2ER) Fig. 17 Tab le 3 It obviates the comput ationa l burden of co nv e ntional iterative registration a pp roaches, and b y conducting t he reg i stration pro c es s in one -shot, practically m ak e s the real-time clinical use possible . The paradigm is still de pendent on a l arg e set of paired annotated ground -truths to train t he network, which is a se ver e hindrance to develop the approaches i n this category. Deep Reinforcement Learning (Agent-B ased Registration) (DR L) Fig. 19. Tab le 4 Instead of DSM approaches, which are d e pendent to conventional iter a ti ve optimization -based approaches to construct the final tr ansformation, this step - by - step reg i stration was still ve ry faster for clinic a l use. The inability of the ag e nts to intera ct w it h the huge state spa c e in troduced by defo rmable reg i stration fiel d wh e re, as it can be seen in Tab le 4, all the appr oaches were proposed for rig i d registration. Unsupervised End- to - End Registration (UE2ER) Fig. 20 Tab le 5 Instead of using enorm ous grand-truth set, t hey use d ata augmentation techniques on a f e w nu mb e r of inputted samples as seeds whe re a trad i tional similarity measure (or a combination of them) is used as loss function to guide the learning pro cess. Multimodal similarity measures that are used as loss function to conduct the network learning are still ineffi cient, and a network train e d on them, accordingly, inherits this inefficiency. Weakly-Semi- Sup e rvised End - to -End Registration (WSE2ER) Fig. 21 Tab le 6 It obviates the short comings associated to the SE 2ER and UE 2ER p aradigms while inherits t he s t rengths of both. T he approaches are l abel -driven, dual-s up e rvised, or bas e d on adverbial learning (GAN). The best practical par ad igm s o far, where we expect the signific a nt re search focus in nea r fu t ure. - 30 - On the ot h er h and, from technica l poi nt o f view , we can have a diffe r ent analy sis on the introduce d deep learning approac h es fo r medical i mage r egis tration. As stated, so me autho r s us e dee p l ea rning techniques to elicit the most influentia l and disc rimi n ative fe atu res to be fe eded to the co n ve n tio n al o ptimizatio n -base d medical r egistratio n approac h es in o r der to maximize the pe r fo r man ce, while the other s use deep neural n etw orks as th e r eg resso r to direc tly estimate t he transf ormatio n p aramete r s in one-shot in orde r to maxi mize th e r untime s peed. Since the techniques are ab out same i n nature , it c an b e s ta ted t hat all t he afo r eme n tio n ed wo r ks on medic al i mage r eg istratio n exploit the adv antages of utilizatio n of deep learning; the m ain contrib uting adv ant age s are as fo l lows : Dee p learni n g obv i ates t he burde n o f choosing, r educ ing, sele cting, a nd n o rm alizi ng handc r afted fe atures t hat ar e the most importa n t f actor to ac h ieve h ig h pe r fo r mance i n registration. Dee p L earning techniques do not s talk in p r em atu r e co n ve rgen ce o r s tagnatio n w hich ar e two prev al ent co n fining dilemmas in t h e c onve ntional o ptimiz ati o n -based approac h es spe cially where deal ing w ith im ages f rom diff er ent modalities (multimo dal image r eg istratio n ). Dee p learning r espo n se time is eff ective l y low (below a sec ond in mos t t h e c ases) instead of co n v ent ional iterativ e manner w h e r e r untime s in the tens of minutes are n orm fo r co m mon def ormable i mage r eg istratio n technique s. This is an actually decisive factor where pract ical use in clinica l ope ratio n s i s r eal-time and suc h a prolo n ged w astin g time is not a pp r eciated. On t he o ther hand, b ased on t he n o-f r ee -lun c h theo rem (W olpe r t and Mac r eady 1997), we a r e lo si n g some paramete r s w h ile getting so m e o th ers, and de ep l ea rning c annot b e a n exc eption. A cco r ding ly , ut ilizatio n o f dee p neural netw orks impose s some di sadv antages frequently r eported by th e auth ors as f ollow s: Dee p learning n eeds large-sc ale t r ai n ing data to e lev at e its perfo rmance and av oid ove r -f it ting p h e n o mena, w h ile medical image datasets are inhe r ently sma ll-sized. Curre ntl y , e n deav ors from t h e bo t h pe r spec tive s of expanding datase ts, and devising n ov el ideas to circumv ent the prob lem ar e in age n da; n ev erthele ss, th e prob l em is still reported as irritati ng by m o st the aut h o r s (L it je n s et al. 2017) . Computa ti o nal bu rden of tr ai n ing dee p n eural n etw orks is r eally high, a nd c annot be a ffo r dable b ut w ith the multiple co nt em porary GPUs (like a NVIDIA Ti tanX); ac tual ly , the aut h o r s n ee d to rest r ict their set of expe r iments whic h is an influe ntial ob stacle to in ve stigate novel i deas. Curre ntly , c o mmer cial cloud computing enviro n me n ts have a go od potenti al to contrib ute the issue; h ow ever, i t is n ot available or a ffo r dable by th e all, and the p r ob l em is sti ll ope n (A grawal et al. 2015). Indeed, a nothe r info r mativ e know ledge fo r the r eade r s is the m ain cont r ibutio n a nd nov elt y pr opo sed by each afo rementioned se minal w ork; Tab le 8 i s a collec tion of this in f ormatio n in an ov er view , w h ic h ca n be used as refe r ence to compa r e the contrib ution of e ach wo r k. Table 8 : The main co ntribut ion and no velty propo s ed by each afo rement ioned sem i nal wo rk in an o verv iew Author & Year Deep Lear ning Technique Main Co ntribution a nd Nove lty Wu et al. (201 3) CNN The first utilizat ion of DNN s o n m edical image r egistratio n. T he CNN w i th I SA (I nd ependent Su bset Analy sis) exploited to extract feature s fro m multimodal i mages to fee d t o th e conventional H AMMER an d D e mons approache s. T hey u sed patches in the size s of 13×1 3× 13 as well as 21× 21× 21 voxels to broa d en th e network’s receptive fi eld. Cheng et al. (201 6) SAEs Utilizatio n of S AEs to extra ct feature s a n d to work a s a similarity metri c for multim odal i mage r egistration, where t he p rop osed a pproa ch outp erformed c onventio nal simila rity m etrics like N MI (N or malized Mutua l Informati on) and LC C (Lo cal Cr oss -Correlat ion). T he patch es were selected arbi trary from the c enter a nd corners of skull in th e size of 17×1 7 voxels. Simonovsk y et al . (201 6) CNN Utilizatio n o f CNN to extra ct feature s a nd to work as a similarit y m etric for mul timodal image r egi stration, where the proposed a pproac h compare d with MI (Mutual Infor mation) and out performed it with 99 % confidence. Th e patches were sele cted a rbitrar y from t he center and c orners of skull in the siz e of 17× 17 ×17 voxels. It w a s d etected u sing stride in t he pooling layers a s well a s Hi nge l oss -function i nstead of cross -entr opy can contribut e to the train ing conv ergence a nd the ov erall perfor mance. Miao et al. (201 6) CNN For t he fir st tim e, a CNN w a s use d to dir ectly r egr ess th e re stricte d Affin e tra n sfo r mation para meter s to r egister 2D X- ray i mages on 3 D CT o nes based on the parti cles impla nted in the pa tients’ bodi es. T he pat ches wer e selected arbitrary from t he center and corners of i mpla nt i n the size of 5 2× 52 voxels. T he runti me w as reporte d as 100ms whi ch is appr eciable fo r clini cal use. Sokooti et al. (201 7) CNN While t he p reviou s work co nsidered o nly Affine transfor mation, t his work was succ essful to tra in a CNN to regress a D VF (D i splacement Vector Field) ca pable of deforma blely r egiste ring i nputted ima ges i n one shot; however, the r egi stration was u ni modal. The patc hes were i n t he size of 2 9×2 9×29 voxels, while som e ot her cases w ere extracte d in the si ze of 54× 54× 54 voxels to keep the receptiv e field. - 31 - Hu et al. (201 8) CNN A CNN was trai ned f or deforma blely registering and f u sing pre -inter ventiona l multi -param etric M R ima ge s t o the operational T ransrectal Ultra sound (TR US) o f t he pro st at e gland in ord er to decreas e t he ri sk of i mage - guided i ntervention. The work is i mportant considerin g it is m ult imodal, deformabl e, and one sh ot. In a ddition, for t he fir st ti me, T R US was used where th e ti me as a n e xtra d imen sion exacerbat e th e situatio n, a nd extra consideratio ns should be taken into account. de Vos et al . (201 9) CNN This work sh ould i ndeed be con sidered a s the most co mp r ehensive de ep l earning framework ba sed on the CNN to directly regr ess t he defor mable tra nsformation para meter s i n o ne shot. I n a ddition to regre ss t he transfor mation para meters, their multistag e multiresolut ion a pproach was ab le to learn a pre defined sim ilarity measure so that the necessity to utiliza tion of synthe sized a nd la beled da taset i s obviated, whic h i s a big progress i n ap plyin g DNN s t o th e f i eld of medi cal ima ge a naly sis wh ere we a re facing small - si ze d an notated dataset s a s regular. 7. Lite rature Re v iew A nal ys is Fig. 30 presents the di st ribution of publications ov er the y ear s from 2013 w hen the fi r st related pape r was published in the confe r e nce of Medical Image Co mputing and Compute r -Assisted Inte r ve ntion (MICCA I-2013) up to June 2019 (the sub mission t ime of th e 2 nd r ev ise of this review ). The r e are a 1-y ear gap in 2 014 w ith n o publicatio n , and a pick of public at io n s in 2018. Altho ugh this topic is in its infa n cy , and any co n clusio n may be premature at this time , base d on a simple regressio n o n the numbe r o f wo rks publis h ed i n eac h y ear , w e e xpec t this topic to ultima tely find s i ts w a y , and w e co nt inue w itnessi n g more pub l icatio n s a nd wo r ks in this c h alle n gi n g area. Figure 30: The numbe r of publ icatio ns based o n the y ear The numbe r of publications base d on the pub l icatio n ty pe is pr ese nted in Fig. 31 . Mo r e than 60% of publications are from co n fe r ence proc eedin gs where journal articles and book chapte r s ar e in the n ext ran ks . Acco r dingly , Table 9 lists top jo urn als, co nferences, an d bo oks in th e fie ld based on the n u mber o f r elated public atio ns. As the first pape r of th e literatu r e w as pub lished i n, the c onfe r ence of Medical Im age Computing a nd Co mputer -Assisted Inte r ve n tio n (MICCAI ) has b een a valu able f orum fo r t h e r el at ed researc h e r s and the ir c ontributio ns. IEEE Internat ional S y mposium o n Bio m edical Imaging, IEEE Tr ans actio n s on Medical Imagi n g, I n te rnati o n al Wo r ks h op o n Deep Learning in Medica l Image A nalys is, and Medical Imagi n g : Image P r oc ess in g are in the sec ond an d third ran ks . Ac co r dingly , Fig. 32 s h ow s top pub licati o n titles b ased th e n umb er of t he publis h ed w orks where Springer Nat ure, IEE E, Elsev ier B V, and SPIE a r e co nsider ed as t o p pub li cat ions pub lished ov er 85% o f th e w or ks in this a r ea . 0 5 10 15 20 25 30 35 2013 2015 2016 2017 2018 2019 The No. of publications Year - 32 - Figure 31: The numbe r of publ icatio ns based o n the pub li c ation ty p e Table 9 : To p journ als, c onf erences, and boo ks in the field b ased o n the number of r elat ed publica t ions Internati on al Confer ence on M edical I mage Computi ng and Computer -Assi sted Interventio n 14 Biomedi cal Signal Pr ocessing a nd Control 1 IEEE Internati onal Sym posium on Biomedi cal Imagi ng (ISBI) 5 Computational a nd Mathematical Method s in Medici ne 1 IEEE T rans a ction s on Medical I maging 4 Computer M ethods i n Biome ch a nics and Bi omedical Engine ering: Imaging & Visualizati on 1 Medical I maging: I mage Pro cessing 4 Conferen ce on Medi cal Imagi ng with De ep Learning 1 Internati on al Worksh op on Ma chine L earning in M edical Imaging 4 Simulation, I mage Pro cessing, an d Ultra so u nd Sy stems for Assisted Dia gnosi s an d Navigati on 1 Deep Learnin g in Me dical I mage An al ysis an d Multimodal Learning fo r Clinical Deci sion Support 3 Sensor s 1 Internati on al Jour nal of Co mputer Assisted Radi ology and Surgery 3 Electronic s Letters 1 Medical I mage Anal ysis 3 Radiothera py and On cology 1 Bildverar beitung fur di e M edizin 2 Internati on al Worksh op on Simulati on a nd Synthe sis in Medical Imaging 1 Deep Learnin g for M edical Image Analysis 2 Pattern R ecognition Lette rs 1 IEEE Tra ns a ction s on Biom edic al Engin eering 2 IEEE Works hop on Ad vances in In formatio n, Electroni c a nd Electrical En gineering 1 Image An al ysis fo r Movi ng Organ, Br east, an d Thoracic I mages 2 IEEE/C VF confer ence on comput er vision an d pattern r ecognition 1 AAAI C onference on Artificial Int elligence 2 NeuroIma ge 1 Internati on al Joint C onferen ce on Bio medical Engineeri ng Systems an d Technol ogies 2 Medical ph ysics 1 Internati on al Worksh op on De ep Learni ng in Medical Image Analysi s 2 Medical I maging: Digita l Pathol ogy 1 Internati on al Confer ence on In formation Pr oce ssing in Medi cal Imaging 2 Understan ding and Int erpreting Ma chine L earning in M edical Image Computi ng A pplications 1 Physics in M edicine & Biology 1 Medical I maging: Biom edical Ap plicatio ns in Molecular, Structural, a nd Functional Imagi ng 1 Annual Con ference o n Medical I mage Un de r standing an d Analysi s 1 Medical & Bi olo gical Engineer ing & Com puting 1 Asian Co nference on C omputer V ision 1 Journal of M edical Ima ging 1 Technolo gy in canc er research & tr eatm en t 1 IEEE Access 1 Conferen ce Proceed ings , 51 , 64% Journal Articl e , 27 , 34% Book Section , 2 , 2% - 33 - Figure 32 : T op pub licatio n titles based the numbe r of the pub l ishe d wo r ks Table 10 s h ow s a list of top 100 a u thors active in th e field based on th e numbe r of th ei r publicatio n s as we ll as total numbe r o f citati o n s to thei r w orks . To tally 264 au thors w ere detecte d in the f i eld w h e r e o t her 16 4 aut ho r s acc ommo dated in the Ap pendix 2 be cause of th e space inco n ve n ie nce . The n u mber of citatio n s have been gathe r ed from “ Sc h o lar Goo gle. ” Rui Li ao and S h u n Miao f r om Me dical Imag ing Te chnologie s, Sieme n s H ealthcare, U SA can b e co n sidere d as the to p autho r s b oth f r om the numbe r o f pub l ications and the total n umb er of citations perspectiv es. Also , D ingga n g S h en at t h e Depa rtme nt o f Radio logy , U n ive r sity of North Carolin a at C hapel Hill is w orkin g in t h e nex t rank. Table 10 : To p 100 autho r s ac ti v e in this fie ld Au th or Pub.s Cite.s Au th or Pub.s Cite.s Au th or Pub.s Cite.s Au th or Pub.s Cite.s Liao, Rui 9 269 Ghavami, Nooshin 3 34 Sedai, Suman 2 12 Comaniciu, Dorin 1 48 Miao, Shun 9 269 Gibson, Eli 3 34 Mahapatra, D. 2 12 Grb i c, Sasa 1 48 Shen, Dinggang 8 205 Hu, Yipeng 3 34 Ray, Nilanjan 2 12 de Tournemire, P. 1 48 Mansi, T ommaso 6 113 Yap, Pew-Thi an 3 32 Zh e ng, Jiann an 2 6 Pennec, Xavier 1 47 Wang, Qian 5 178 Heinrich, Mattias P 3 7 Wood, Brad J 2 5 Sermesant, Maxime 1 47 Cao, Xiaohuan 5 56 Wu, Guorong 3 0 Xu, Sheng 2 5 Heimann, Tobias 1 47 Wang, Z Jane 4 170 Styn e r, M ar tin 2 97 Yan, Pingkun 2 5 Datar, Manasi 1 47 Kim, Mi njeong 4 168 Vierg e ver, Max A 2 85 Heldmann, Stefan 2 4 Rohe, Marc-Michel 1 47 Niethammer, Marc 4 158 Kamen, Ali 2 83 Hering, Alessa 2 4 Cheng, Xi 1 45 Kwitt, Roland 4 158 Navab, Nassir 2 82 Blendowski, Max 2 4 Maier, Andreas K 1 35 Yang, Xiao 4 158 Zhang, Li 2 80 Zhao, Bojun 2 1 Ghesu, Florin C 1 35 Staring, Marius 3 143 Zh e ng, Yefeng 2 75 Jiao, Wanzhen 2 1 Nie, Dong 1 19 Išgum, I vana 3 143 Zhao, Amy 2 59 Cong, Jinyu 2 1 Fan, Y on g 1 15 Berendsen, Floris 3 143 Delingette, Herve 2 45 Jiang, Yanyun 2 1 Li, Hongmin g 1 15 de Vos, Bob 3 143 Zhang, Jun 2 30 Zh e ng, Yu an jie 2 1 Bandula, Steven 1 13 Sabuncu, Mert R 3 87 Yang, Jianhua 2 30 Che, Tongtong 2 1 Wang, Guotai 1 13 Guttag, John 3 87 Modat, Marc 2 30 Yang, Xiaodong 2 0 Li, Wenqi 1 13 Ba l akrishnan, Gu ha 3 87 Fan, Jingfan 2 21 Muns el l, Brent C 1 81 Ehrhardt, Jan 1 12 Dalca, Adrian V 3 87 Barratt, Dean C 2 21 Komodakis, Nikos 1 72 Handels, Heinz 1 12 Sokooti, Hessam 3 69 Noble, J Alison 2 21 Mateus, Diana 1 72 Wilms, Matthias 1 12 Ayache, Nicholas 3 51 Vercauteren, Tom 2 21 Gutierrez-Becker, B. 1 72 Uzunova, Hristina 1 12 Krebs, Julian 3 51 Pluim, Josien PW 2 20 Simonovsky, Martin 1 72 Veta, Mitko 1 12 Emberton, Mark 3 34 Eppenhof, Koen 2 20 Liao, Shu 1 62 Moeskops, Pim 1 12 Moore, Caroline M 3 34 Xue, Zhong 2 17 Gao, Yaozong 1 62 Lafarge, Maxime 1 12 Bonmati, Ester 3 34 Mailhe, Boris 2 16 Leliev e ldt, Boude wijn 1 58 Jeong, Won- Ki 1 11 Springer , 38 , 48% IEEE , 14 , 18% Elsevier , 9 , 11% SPIE , 7 , 9% SCITEPRESS , 2 , 3% AAAI , 2 , 3% Wiley Onli ne Librar y , 1 , 1% Taylor & Francis , 1 , 1% SAGE Publications , 1 , 1% MDPI , 1 , 1% IOP Publi shing , 1 , 1% IET , 1 , 1% Hindawi , 1 , 1% Unkno wn , 1 , 1% - 34 - Fig. 33 is a TagC loud visuali zation base d o n the freque ncies of keywo r ds used by th e aut h o r s. The most freque n t keyw ords a re Dee p L earning (22 time s), Convo lutional Ne ural Netwo r k (16 times), I mage R egistration (12 times), Defo r ma b le Image Re gistration (8 times), a n d Def ormable Re gistration ( 6 times). Figure 33: Tag Cloud key wo rd frequency diagram Top m et rics used to ev aluate the approac h es are illustrated in Fig. 34. They really n eed special conside ratio n s w h e re Dice Coeffic ient (DSC) and T arget R egistration E rr o r (TRE) are the most f r eque n t ly used metrics. Dice c oeff icient is an accredited no n -pa r amet ri c measu re to quantify the amo unt o f over lappi n g r egions in th e inputted fix ed and moving images . I t can b e calculate d using (3). (3) w h ere A and B are in the inte n sity histog r am of the inputted image s. It s value is in range of [0, 1] whe r e 0 indicates n o ove rl ap, w hil e the v alue o f 1 indicates perfe ct o n e. T h e most i nteresting po int ab out DSC i s i ts n on-pa rametric n atu r e an d no n ee d t o any manual o peration. On the ot h e r hand, t h e re i s Ta r get Registratio n Erro r (TR E) f or which to be computed, a n umb er o f c orrespo n ding landmarks should b e specified in bo t h th e inputted images . The sum of distan ce s between these points at the e nd o f th e registrat ion is co nsidered as TR E using ( 4) r egul arly utte red in mm. TRE is also of th e most accredited perfo r mance metrics w h ose only dra w bac k is its n eed to manual o perations; h ow ever, the co rr es pondi n g landmarks can be specif ied automatically to remo ve the b ur de n . (4) w h ere A and B are th e inputted i mages , an d are the i -the c orr espo n ding la n dma rks in A and B , and n i s the tot al numbe r of lan d marks i n bo t h image s. - 35 - Figure 34: Top me tri c s used to evalua t e th e appro aches Top da tase ts used for th e imple mentatio n and ev al uatio n stu dies are i llust rated in Fig. 35. Th e mos t frequent ly used datase ts ar e the set of {Pri v ate, ANDI, LON I, IXI , OASIS , and DIR LAB }, each of w h ich has bee n utilized in more than 5 papers. P rivate st ands fo r th e priva te datasets that w ere not pub l icly av ailable at the time th e pape r had bee n publis h ed. Also , in the sec ond ra n k is the set of { Sunn yb r oo k, ACD C, MCIC, M GH 10, XEF, Harvard GSP, HA BS, PPMI, CU MC12, IBSR 18, BrainWeb , SmartTarge t, ADH D200,AB IDE, TKA , and VIPS }. Figure 35 : Top d atasets used to im plemen t the app roac hes based on the number of publicatio ns The top organs o f in te r es t based on the n u mber of publicatio n s are prese nted in the Fig. 36 where the brai n is by far the mo st i nteresti n g o r gan to w h ich 77 pape r s are b elong ed . It is the mos t likely bec ause the b rain is e nclose d by th e skull that is so lid and ma kes t he proc ess of ali g nment faste r a nd simple r . I n addi tion, the structures of interest ar e mostly v isi b le for brain i n diff erent modalities, which also ma kes the val idation proc ess m o r e accurate. 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 32 34 36 38 Dice TRE Hausdorff MAD/MAE Deformation Err or ASD/ASSD ASCD/MCD SSD RMSE mTREproj MSE/M SD MDM Accuracy Jaccard MDG Distance Erro r Ratio of Registrati on Recall Registrati on Error Precision FSIM Grad Det-Jac LCC D-Score CSPE Matching Performance CPD Projected TRE Sucess Rate SSIM RVLJ RMSEc RMSD MID PSNR Mean TRE PCC NRMSD NMI NCC MME MI Retrival Rate The No. of Publicat ions Used This Metric 0 2 4 6 8 10 12 14 16 18 20 22 24 Private ANDI LONI IXI OASIS DIRLAB Sunnybrook ACDC MCIC MGH10 XEF Harvard GSP HABS PPMI CUMC12 IBSR18 BrainWeb SmartTarget ADHD200 ABIDE TKA VIPS Annidis RHA IBIS 3D Autism CREATIS APCH HRF-base CREMI TEM Prostate-3T COPD-Gene Chase DB Buckner40 BRATS Diaret DB Bioimaging Challenge NLST RIRE RODREP ALBERTs Spine SPREAD TEE ABD RCCT NIH MSH LIDC-IDRI JSRT Messidor1 ISLES2015 NewTom NIH AANLib PROMISE12 NIH ChestXray14 LUMC FIRE Empire 10 EMC E2 E1 DTSET2 DTSET1 HMC - 36 - Figure 36: Top o rgans of inte rest Fi g. 37 show s th e pie diagram of top modalities use d in the literatu r e. MR imagin g was used fo r abo ut 50%, CT and X-ray which ar e in the n ext ranks co nsti tute abo ut other 27%, a n d t h e re a r e CFI and OC T f o r the retina imaging accumulate ly a b out 10%. B ased on th e Fig. 38, mult imodal registratio n i s the case fo r 29 publicatio ns (36%) and the remaini n g 51 w orks ( 64 %) we r e on unimoda l registratio n . Wo r se m e n ti o n ing that r egist ration of co n sub stan tial modalit ies like CT and X-ray is co nsidered as unimodal while the utilizatio n of same modal ities wit h diff eren t parame ters like T1 and T2-we ighted MRI is co n side red a s multimoda l. Figure 37: Top m oda lities us ed in the lite rature 0 5 10 15 20 25 30 35 40 45 50 55 60 65 70 75 80 Brain Cardiac Lung Retina Prostate Chest Spine Head and Neck Knee Wrist Body Bone Breast Chest an d Abdomen Elbow Liver Nasopharynx Transduce r Craniofacial The No. of Publicat ions Organ MRI , 96 , 52% CT , 35 , 19% X-ray , 15 , 8% CFI , 9 , 5% OCT , 8 , 4% CBCT , 5 , 3% TRUS , 4 , 2% EM , 3 , 2% US , 3 , 2% MSI , 2 , 1% FA , 1 , 1% DSA , 1 , 1% TEE , 1 , 1% PET , 1 , 1% DR , 1 , 1% DRR , 1 , 1% - 37 - Figure 38: Mu ltimo dal versu s unimo dal regi stration Finally , Fig. 39 shows the pie diagram of transfo r mat ion mo dels used in th e lite r ature. Acco r dingly , in 78 % of cases (62 times), aut h ors tried fo r def ormable regist ration, while only 22% (18 case s) be longs to th e ri g id registratio n; w h ere we se e that mostly th e o r gan- of -in te r est is th e brain t hat has relati v ely a rigid fo r m, it c an be tter rev eal us the importa n ce of de formable regist ration f or almost al l the orga n s. Figure 39: Tran sfo rmation mo dels used in the litera ture 8. Discu ss ion on Confi di ng Ch allenges, Open Pr oblems and Pr o m is ing Dire ctions The stu dy on the publicatio ns in th e field o f a pply in g deep l e arn ing approac h es on medical imagi n g rev eals us some freque n t challe n ges. O n t h e o ther hand, the se t o f nove l so lut io n s p r opo sed by the r el ated authors, experts a nd researche r s in th ei r publicatio n s ca n be a rich r efe ren ce and guideli n e t o th e p r os pectiv e prob lems in the area o f medical image registratio n s using dee p neural netw orks. So me of th ese challenges and so l utio n s are enume r at ed as fo llows: Challeng e: Med ical datase ts are of t en small-si zed. So l u tion: U til izatio n of augme n tatio n techniques to artifici al ly inc rease the n umbe r o f sample s. U til iza tion of transfe r learni n g to tr ai n the netwo rk from oth e r datase ts and the n fi n e-tu ne it w ith th e datase t in conside ratio n . U til izatio n of weakl y-s upervise d learning to train the n etwork from semi-annota ted data. F inally , utiliz ation of dropo ut whic h is a technique to s tochastical ly drop s ome inputs out o f e ach lay er to dec rease th e ov erfitting ef fect . Challeng e: Medical datase ts’ annotated lab els ar e noisy t o a large exte n t since th e phy sician s and expe rts do not have a co n se nt in a lots of case s. So l u tion: Mo delin g t h e no ise dist ribution and fe edin g it to the n etw ork, o r use e.g. f uzzy logic t o t ack le the issue . Unimod al , 51 , 64% Multimodal , 29 , 36% Rigi d , 18 , 22% Defo rmable , 62 , 78% - 38 - Challeng e: I n co ntr ast to Dec ision Support Sy stems (DSSs), dee p learning a pp r oac h es do n ot present the r ule - chain fo r their inferences w h ere it is may be unacce ptable for th e phy sician s ev en thoug h the sy st em h as had a high p r ecis ion and accu r acy . So l u tion: S ome pr o mising stu dies ha ve already bee n co n ducte d to r eprese nt t he w ay of netw ork’s i n fe r ence bas ed on the visual izatio n of n etw ork’s in te rnal (hidde n ) lay ers, but the ov erco me s a re limite d, and the p r ob l em is still ope n. Challeng e: Backgrou nd know l edge an d co n text can be h ighly in f ormative a s th e phy sician s ask the patients a numbe r of r el ated ques ti o n s , and see their diff erent r ec ords a n d expe rime nt s’ results . So l u tion: Likew ise, p a tie nts’ clinical rec ords, thei r genomics , bio psies and o th e r expe r ime n ts’ results can al so be gathe r ed an d fee ded to t he n etw o r k v ia diff er ent channels to enhance the pe rformance; w h il e to in ve stigate the impact, t h e re are too few i nt eg rated datase ts whic h wo rsen s the situatio n . Challeng e: Medica l i magi ng is inhe r e ntly 3D, but is proce ssed 2D or 2 .5D by most of th e current dee p n eu ral netwo r ks. The reaso n is that 3D proc ess in g with 3D DNN s is co mputationally un affo r dable in many case s. So l u tion: No th ing t o do! We should seat and see whet h e r the progress in the inf r ast ructure w ill fi n ally en a ble us to at tra ct the comput ational intensity requisite to do that. The r evie w o n the literatu r e o f medical image r egist ratio n b ased o n the deep n eu ral n etw orks r ev eals t hat t he propo sed approac h es try to technically enhance the two follow in g paramete r s: 1. Registratio n Ru ntime: Th e a r c hi tectu res of th e propo sed appr oac h es are so t hat to dec r ease the registratio n r un- time while keeping th e perfo r m ance ; in othe r w ords, th e aut h ors did n o t w ant to over-engineer a n e two rk by increasing th e n umb er of l ay er s, co nn ec tio n s and pa r amete r s to drastic ally improv e th e perfo r ma n ce . In (de Vos et al. 2019), i t h as b ee n asserted that t he propo sed appr o ach is able registe r a ty pical co uple of in putted image s in less t h an 50 m s , w h ich is hig hl y suitable a nd appreciate d f or real-time cl inical use. 2. Netwo rk Rec eptive Field: T h e ext r acted patc h es fro m the c ouple of in putted ima ges are ty pi cally selecte d usin g a small sliding windo w with so me o verlap. The patches are usually small in r ange of 13×13×13 up to 30×30× 30 voxe ls fo r r educ ing t h e co mputational i n te n sity to be tractabl e, and thi s c o n fide s the n etw ork rece ptive field while the bac kgr ou n d context may be fully in fo r mative. To issue the prob lem, in some stud ies, so me bigger p atches are extracted apa r t f r om t h e regu lar patc h es; th ese patc hes a r e co m press ed and shrunke n (t o reduce t h e computa ti o nal burden), a n d f ee d ed to th e ne two r k via a diff erent channel . 9. Conc lusions and Future Trends In this pape r, a taxo n omy w as developed o n deep lea rning b ase d a pp r oac h es f or medical im age registrat ion w ith fiv e catego ri es n amed Deep Simila r ity Metrics (DSM), Supe r vis ed End - to -End Regist r atio n (SE2ER ), Deep Re in fo rcement Le arn ing (or Age n t -B ased Regist r atio n ) (DR L), U n supe rvised End - to -End Re gistratio n (U E2ER), Weakly /Semi- Supervis ed End- to -End Re gistra tio n (WSE2 ER). T h e appro aches in e ach catego r ies sha res some ide n tic al spec ifications , underly in g philos ophies, pa ra d igms, advantages and di sadv antages. Ge n e r ally , deep r ei n fo r ceme n t l ea rn ing prov ed n ot be optimistic as t h e huge state-space assoc iat ed to defo rm abl e regist r atio n i s out- of -tole r ate fo r the lea rning age nts avo id them to b e prope r ly converged. Amo ng others, u n supe r vise d and w eakly-supervise d approac h es h ave less depe n dency t o the grou n d-truth that is very costy - to -obtain f or medic al imag ing applica tio n s; hence, b eside t h e hind r ance and co n f ining challe n ges t h ey ar e f aced, w e expe ct in c r easi ng attentio n and r ese ar c h foc us on th e m. U n less, we ca n be en co un tered t he public at io n of h uge publicly -availa b le annot ated d a tase ts fo r dif fe r ent o r ga ns - of -in terest w ith diff erent m o dal ities . Transfe r learni n g is an o ther choice fo r supe r vise d approac h es, whic h w a s suc ces sfull y applied fo r so me o r g ans and mod ali ti es , but its ge n eral izati o n needs muc h mo re ev iden ces. Amo n g we akly-supervise d end- to -e nd regist r atio n , adve r b ia l learni n g e.g . ut ilizatio n of GA Ns has the maj or co n tributio n; also , dual-supe r v ision t hat is base d o n the learning from a similarity measur e l ike MI (just like unsupe r vise d approac hes) and a few gr ound -t ruth samples to fine-t u n e th e netwo r k is very promising, a n d needs a fu rth e r co n side r atio n ; howe ver, as we go farer f r om the real-wo r ld ground-t r ut h, the r es ults get questionab le fo r cli nician s who wa n t to know abo ut h ow realistic , fe asible an d practica l ar e the m. We are see in g a gap here, and h ope fo r furthe r r ese arc h on the r egistratio n validat ion to o bviate t his doub t . The fo llowing stateme n ts can also be concluded f rom this r ev i ew: Multistage policy , w h ere w e de fine a rigid r egist ration be fore go es fo r de fo rm able o n e, h as a po sit ive imp act as repo r ted by m any authors. - 39 - Multireso lutio n policy , w h ere w e gr a du ally c onduct t h e registratio n p r oc ess f r om low-r eso luti o n to the highest o n e, also h as r epo r t ed as a strong pa r adig m to inc rease the regist ration p r ec ision. As r epo rted i n the l iteratu r e, havi n g a few grou nd -t r uth s ample s, tra nsfer lea rning from o th e r bo dy organs o r modalities , is fully pr actical fo r medical im age regist ration. Incorpo r ating the th eo r y of geometry to the app r o ach es i s a nov el idea, and nee d future co n sideratio n . Spatial T ransfo r mer Netw ork (STN) is one the key contributo r s, w h e n co upled to a e.g. CNN , it ca n have a sig n ifica n t imp rov ement o n the pe rfo r mance. Ov er-engin ee ring, i.e. inc r easi n g the numbe r o f la y ers, co n nections an d p aramete r s to dras ti cal ly improve the perfo rmance, is o f the case her e since th e applicatio n is re al -time. Since t he b ackgrou n d co n text ca n be f ully in f ormative, increasi n g t h e netw ork’s r ec ep tive field is a pos it ive co nt ributi n g f actor fo llowe d by m an y authors. We b elie ve t hat most o f the f ut u r e trends and co n trib utions cannot b e i n tri nsic in the fie ld o f medica l image registratio n, b ut they will co me f r om other medical imag ing prob lems, or ev en a bit farer i.e. co mputer visio n and mac h ine learning fields. From the applicat ion p r os pectiv e, deep learni n g techniques a pplied to the medical image r egis tration are restricted t o th e CN Ns, SAEs, GAN s, DRL , and dee p R NN w h il e othe r mode ls have a hig h po tential f or c ontributio n. To exe mpli fy , Gated Rec ur rent Units (GRD), w hich is a r ecu rren t dee p learni n g model, has a high pote nt ial to be appli ed w h ere the t ime i s the case as the 4t h dimensio n e.g. in co n ti nues U S images or disc rete fluo r os co p y. On the othe r hand, from the tec hn ique po int of v iew, th e fie lds of machine learni n g and co mputer visio n are in a co nt inuous prog ress w her e promisi ng tec hn ique s are introd ucing co n stantly . Fo r exa mple, Spi king Neural Ne two r ks (SNNs ), as th e 3rd gene ration of n eu r al netwo r ks, aim at bridgi n g the gap be twee n neuros cience and machine lea rn ing via utilizat ion of b i ologic all y -r ealistic mode ls o f n eu r ons for c omput ation. A SNN is b asically diffe ren t from t h e co nventional n eu ral n etw orks know in g by th e co mmunity of machine l ea rning. SNN s o perate using spikes, w hic h ar e discrete ev en ts oc curring at time in st ances, rat h e r than to be co n tinuous values . Taking pl ace of a spike is de termi n ed by diffe r ential equ atio n s t h at rep resent various b iolo gical pro cesse s where the mos t impo r ta n t is t h e membrane po tential. In general, o n ce a n eu ron r eaches a ce r tain potenti al, it spike s, and the potential of t hat neuron is r eset. The most commo n mode l fo r this is t h e Le ak y Integrate-a n d-F ire (LIF ) m ode l . Mo r eo ver, SNNs are of ten spa r sely conn ec ted and take adva n tage o f sparse netw ork to polo gies. Refe re nce s A . A . Goshtasby , Theory and Ap plicat ions of I ma ge Re gistrat ion . Jo hn Wiley & Sons, 2017 . J. Ha jnal, D . Haw ke s, and D . H ill, Medical Image Registrat ion . Bio medical Eng i nee ring, J un. 2001 . T. Peter s, and K . Cleary , Image -guided interve ntion s: technol ogy a nd applic ations . Spri nger Science & Busin ess Me dia, 20 08. F. P. O liveira , and J. M. R. Tavares , “ Med ic al imag e reg istration : a review ” Com pute r me thods i n biome chanics and biomedic al engine ering , vo l . 17, no . 2, pp . 73-93, 2 014. G. Li tjens, T. Ko oi, B. E . Be jnordi, A. A . A. Set io, F. Ciom pi, M. Ghaf oo rian, J. A. W. M. van der Laak, B. van Ginn eken, and C. I . Sánchez , “A survey o n de ep learning in m edical imag e ana ly sis ,” Me dical Image Analy sis , vo l. 42, pp. 60 – 88, Dec. 2017 . G. Wu , M. Kim, Q . Wang , Y. Gao , S. Li ao, and D. Shen, “Unsu perv ised De ep Feature Learn ing fo r Defo rmab le Reg istratio n of MR Brain I mages,” Lectu re No tes in Com pute r Sci ence , pp. 649 – 656 , 2013. X. Cheng , L. Zhang , and Y . Zh eng, “D eep si m ilarity learn ing fo r multimo dal medic al images, ” Comp ut er Meth ods i n Biomech anics and Biome dica l Engine ering: Imag ing & Visualiz ation , vo l . 6, no . 3, pp. 248 – 2 52, Apr. 2018. (A ccepted & On line fro m 2016) X. Yang , R. Kw itt, and M. Nietham mer, “Fast P redictive I mag e Registratio n,” Le cture No tes in Compu ter Sc ience , pp. 48 – 57, 2016 . G. W u, M. Kim, Q. Wang, B. C. Mu nsell, a nd D . Shen, “Scalabl e Hig h -Perfo r manc e Im age Reg istrat ion Fram ework by Unsupervise d Deep Fea ture R epresent atio ns Lear ning ,” IEEE T ransact ions on Biome dical Eng ineering , vo l. 6 3, no. 7, pp. 1505 – 1516, Jul. 2016. M. Simo nov sk y , B. Gut iérrez- B ecker, D. M ate us, N. Nav ab, and N. Ko mo dakis, “A Deep Me tr ic fo r Multimo dal R egistrat ion,” Medic al Image Compu ting and Comp uter-Assi sted In tervent ion (MICC AI 2016), pp. 10 – 18, 2016 . S. Miao , Z. J. Wang , Y . Zheng , and R. Liao , “Real - t ime 2D/3D registrat ion via CNN reg ression, ” 20 16 I EEE 13t h Inte rnationa l Sympos ium on Biome dical Im aging (I SBI ), A pr. 2016 . S. Miao , Z. J. Wang , and R . Liao, “A CNN Reg ression A ppr oac h fo r Real - T ime 2D/ 3D Regi stration, ” IEEE Transac t ions on Me dical Imag ing , vol. 35, no . 5, pp. 1352 – 136 3, May 2016. Rui L iao, S hun M iao, P ierre de T ourn emire, Sasa G rbic, A li K amen , To mm aso Man si, and Do ri n Co manic iu. “A n A r tificial A gent f o r Ro bust I mag e Registra tion,” i n AAA I , pp. 41 68-4175 , 2017. J. Krebs, T. M ans i , H. D eli ngette , L. Zh ang, F. C. Ghesu , S. Miao, A. K. Maier , N . Ay ache, R. Liao , a n d A. Kame n, “Robus t Non - rigid Reg i st ration T hroug h Ag ent - B ased A ction L earning,” Le cture Note s in Compu t er Science , pp. 344 – 35 2, 2017 . - 40 - X. Yang , R. Kwi tt, M. Sty ner, and M. N iethamme r , “ F ast p redict iv e multimo dal i mag e regist ration,” 2017 IE EE 14 th Interna tiona l Sympos ium on Biome dical Im aging (I SBI 2017), A pr. 2017. S. Wang , M. Kim, G. Wu, and D . Shen , “Sc alable High Perfo r manc e I mage Reg istration F ramewo rk by Unsupervise d Deep Fea tu re Represen tat ions Lear ning ,” in De ep Le arning fo r Medica l Image Analy sis , pp. 24 5 – 269, 20 17. S. Miao, J. Z. Wang , and R . L i ao , “Co nvo luti o nal Ne ural Netwo r ks f or Robust and Real -Tim e 2- D/3- D Reg istratio n,” in De ep Learn ing for Me dical Ima ge Ana l ysi s , pp. 271 – 296 , 2017. H. Soko oti, B . de Vo s, F . Berendse n, B. P. F. Leliev el dt, I . Išg um, and M. Staring , “Nonr igid I mag e Registra ti o n Usi ng Multi -sc ale 3D Co nvo lutional Neu ral Netwo r k s,” Lec ture Note s in Com puter S cience , pp . 232 – 239 , 2017. B. D. de Vos, F . F. B erendsen , M. A . V ierg ever, M. Stari ng, and I . I šgum , “End - to -End Unsupe rvised D efo rmable I mage Reg istratio n with a Conv olutional Neural Netwo r k,” Lecture Notes in Comp uter Scie nce , pp. 204 – 212 , 2017. X. Yang, R. Kwi tt, M. Sty ner, and M. Nie tham mer, “Quicks ilver: Fast pred ictiv e imag e registrat ion – A deep learning appro ach,” Ne uroImage , v ol. 158 , p p. 378 – 39 6, Sep. 2017 . J. Zheng , S. M iao, Z. Jane W ang, and R. L iao, “P airwise domain adaptat ion mo dule fo r CNN -based 2-D/3- D regis tration,” J ournal o f Medic al Imaging , vo l. 5, no. 02 , p. 1 , Jan. 201 8. Y. Hu , M. Modat , E. G ibson, W. Li, N. Gh avam i, E. Bo nmati, G . Wang , S. Bandu la, C . M. Moo r e, M. Emberto n, S. Ourselin, J. A. No ble, D . C . B arratt, and T. Vercau t eren , “Weakly -supervised co nvolutional ne ural networks f or multimodal image re g istra ti on, ” Medic al Image Analy sis , vo l. 49, pp. 1 – 13, Oct . 2018 . X. Zh u, M. D i ng, T. Huang , X. Jin , and X . Zhang , “PCA Net -Based Structur al Rep r esen tation fo r No nrigid Multimo dal Me dical I mag e Reg istration,” Se ns ors , vo l. 18, no . 5, p. 1477, May 2018. J. M. S loan, K. A. Goatman , and J. P. Siebe rt, “Learn ing Rigid Im age Registra ti o n - U tilizing Co nvo l utional N eural Ne tworks for Medic al I mage Reg istration,” in Pro ceeding s o f the 11th In t erna tion al Jo int C onfere nce on B iome dical Eng ineering Systems and Technolo gies , 2018 . B. D. d e Vo s, F . F . Berend sen, M. A. Vierg ever, H. Soko oti, M. S taring , and I. I šgum , “A deep learning fram ework fo r unsup erv ised aff ine and defo rmab le image regist ration, ” Medi cal Image Anal ysis , v ol. 52, pp . 128 – 143 , Feb. 2019. J. B. A . Maintz and M. A. V iergev er, “A sur v ey of medical imag e reg istration, ” M e dical Im age An alysis , vo l . 2, no . 1, pp. 1 – 36, Ma r. 1998. E. Ferrant e a nd N. P arag ios, “Slice - to - vo lume med ical imag e r egis tratio n: A sur v ey ,” Medi cal Image A nalysis , vo l . 39, pp. 101 – 123, Jul. 201 7. J. Duc hon, “Sp lines m inimizi ng rota tion -invar iant sem i- norms in So bolev spaces, ” Lec ture Note s in Mathem atics , pp . 85 – 100 , 1 977. D. R ueckert , L . I . S ono da, C. Hay es, D. L. G . Hill, M. O. Leach, and D. J . Haw kes, “Nonr igid regist ration using free -f orm defo rmations: applicat ion to breast MR im ages,” IEEE Transa ctions on Med i cal Imagin g , v ol. 18, no . 8, pp. 712 – 721, 1999 . K. Fukus him a, “Neoc og nitron: A se lf -o rganiz ing neura l netwo rk mo de l for a mechan ism o f pattern recog ni tion unaffec ted by shift in positio n,” Biologi cal C yberne tics , vo l. 36, no . 4 , pp. 193 – 202, A pr . 1980 . S. -C. B. Lo , M. T . Freedma n, J. -S. Lin, B . K rasne r, and S. K. Mu n , “Com puter -a ssisted diagno si s fo r lung nodule detection using a neural network techniq ue,” in Me dical Im aging VI : Image Process ing , Jun. 1992 . Y. Lec un, L . Bo ttou, Y . B engio , and P . Haff ner, “G radient - ba sed learning applied to do cum ent reco gnition,” P roc eedings of the IEEE , v ol. 86 , no. 11, pp . 2278 – 232 4, 19 98. Krizhev sky , I. Sutskev er, and G. E . Hinto n, “I mag eNet class ificatio n with de ep c onvo luti ona l neu ral netwo rks,” Comm unicatio ns of the AC M , vo l. 60, no. 6, pp. 8 4 – 90, M ay 2017. O. Ru ssakov sk y , J. Deng , H . Su , J. Kr ause, S. Satheesh , S. Ma, Z. Huang, A . Karpa t hy , A . Khosla, M. Bernste in, A . C. Berg , and L. Fei- Fe i, “Im ageN et Large Sc ale Visua l Reco gnition Challeng e,” I nte rnational Jour nal o f Compu ter Vision , vo l. 115, no. 3, pp. 211 – 252, Apr. 20 15. T. Ve rcauteren, X. P ennec, A . Perchan t, and N . Ay a che, “D iffeo morphic de mo ns : Effic ient non - parame tric i mag e re g istratio n,” Ne uroImage , v ol. 45, no . 1, pp. S61 – S7 2, Mar. 2009. D. Shen , “I mag e registration by loc al histo gram ma t ching ,” Patte rn Re cognition , vo l . 4 0, no. 4, pp. 11 61 – 1172 , A pr. 2007 . J. L ong , E. Shelham er, and T. D arrell, “F ully co nvolutional netwo r ks fo r sem antic segm entatio n,” 2015 IEEE Confe rence on Compu ter Vision and Pat tern Reco gnition (CVPR ), Jun . 2015. D. H. Wo l per t and W. G. M acrea dy , “ No free lunch t heor ems for optimizatio n,” I EEE Transact ions on Evolu tionary Comp utation , vo l . 1, no. 1, p p. 67 – 82 , A pr. 1997 . H. Ag rawal, C. S. Mathialag an, Y . Goy al, N. Chav ali, P. Ban i k, A. Mohap atra, A. Osm an, a nd D . B atra, “Clo udCV: Larg e -S cale Distr i bu ted Com puter Vision as a Cloud Serv i ce,” in M obi le Cloud Visual Media Com puting , p p. 265 – 2 90, 2015 . Z. W ang, T. Schau l, M . He ssel, H . V an H asse lt, M . L anctot, and N. De Fr eitas, “Duel ing netwo rk arc hitectu res f or deep r einfo r cement learn ing,” i n proc . the 33r d In ternatio nal C onf ere nce on In t erna tion al Con f ere nce on Machine Le arning , v ol. 48, pp. 19 95-2003 , 2016. A . Do sov itski y , P . F ischer , E. I l g , P. Ha usser , C. H azirb as, V. Golko v, P . v an der Smag t, D . Cr emers, and T. Bro x, “ Flow Net: Lear ning Optica l Flo w with Co nvo l utional Netwo r ks,” in proc . 2015 IEEE Interna tional Confe rence on Compu t er Vis ion ( I CCV), Dec. 2015. H. Uz unov a, M. Wilms, H . Han dels, and J. Eh rhard t, “Training CN Ns fo r I mag e Reg istratio n fro m F ew Samp les w ith Mode l -based Data A ugmentatio n,” Lecture Notes in Compu ter Sc i ence , pp. 223 – 231, 2017 . K. Ma , J. Wa ng, V. Si ngh, B. T amersoy , Y. - J. Cha ng, A . Wimm er, and T . C hen, “Mu ltimo dal I mag e Registratio n with Deep Contex t Reinfo rcement Learn ing,” Lec ture Note s in Com puter Scie nce , pp. 240 – 2 48, 2 017. - 41 - M. - M . Rohé, M . D atar, T. Heim ann, M. Serm es ant , and X . Pennec, “SVF -Net: L earning Def ormab l e I mag e Registratio n Using Sh ape Matc hing,” Lect ure No tes in Compute r Scie nce , pp. 26 6 – 274, 20 17. X. C ao, J. Ya ng, J. Zh ang, D. Nie, M. K i m, Q . Wa ng, and D. Sh en, “Defo rmable I mage Registrat ion B ased o n Simil arity -Steered CN N Reg ression,” Le cture No tes in Compute r Scie nce , pp. 30 0 – 308, 2017. A . V. Dalc a, G. Balakri shnan, J. Gut tag, and M. R. S abunc u, “Unsupe rvised Lear ning f or Fast Probabi listic Diff eom orphic Reg istration,” Lec t ure N otes i n Compute r Scie nce , pp. 729 – 738, 20 18. J. Fan , X . Cao, Z. Xue, P. -T. Yap, and D. Shen, “Adv ersarial Simi larity Network fo r Evalu ating Im age Alignm ent i n D eep Learning Based Reg istration, ” Lec ture N otes in Compute r Scie nce , pp. 739 – 746, 20 18. G. Ba lakrish nan, A . Zhao , M. R. Sabunc u, A. V . Dalc a, and J. Gu ttag, “An Unsupe rvised Lea rning Model fo r Defo rmable M edic al Im age Reg istration,” in 2018 IEEE /CVF Con fere nce on Com pute r Vision and Pat tern Reco gnition , Jun. 201 8. T. S entker, F. Madesta , and R. Werne r, “GDL - FIR E 4D: Deep L earning- B ased Fast 4D CT I mag e Reg istration,” Le cture No tes in Compute r Scie nce , pp . 7 65 – 773 , 2018 . Y. Hu, E . Gibso n, N. Ghav ami, E. Bonm ati, C . M. Mo ore, M. Em berton , T. V ercauter en, J. A . Noble , and D. C. Barrat t, “Adv er sar ial Defo r mation Regu l ariz ation for Tra ining I mag e Registratio n Ne ural N etworks ,” Lec ture N otes in Compute r Science , pp. 774 – 782, 2018 . M. I to and F . I no, “A n Auto mated Method fo r Gene rating Tra ining Sets fo r De ep L earning based I mag e Regist ration, ” in Procee di ngs of the 11th In ternatio nal Join t Con ference on Biom edical Engine ering S yste ms and Te c hno logies , 2018. S. Ghosal and N. Ray , “Deep deformab l e reg istratio n: Enhancing acc uracy by fully convolutiona l neura l net,” Pat tern Recogn i tion Lette rs , v ol. 94 , pp. 81 – 86 , Jul. 2017 . K. Simo ny an a nd A. Zis serman , “ Very deep co nvo lut ional netwo rks fo r large -sc ale imag e recog ni tion, ” I CL R, 2015 . C. Szeg edy , Wei L iu, Yangqi ng J ia, P. Se rmanet , S . Reed, D. A nguelo v, D. Erhan , V. Vanho ucke, and A . Rabino vic h, “Go ing deepe r with c onvo luti ons , ” in 2015 IEEE Confere nce o n Com puter V ision and Pat tern Re cognit ion (CV PR), Jun . 2015. H. Li and Y . Fan, “No n-rig id i mag e registr ation using self- superv ised ful ly convo l utional netwo rks wi thout train ing data,” in 2018 IEEE 15 t h In ternatio nal Sympo si um on B iomedical Imag ing (I SBI 2018), A pr. 2018. E. Fe r rant e, O . Ok t ay , B . Glo cker , and D. H . M ilone, “O n th e A dapt ability of U nsuperv ised C NN-Bas ed D efo rmable I mag e Reg istration to Unseen I mag e Domains,” Lecture Note s in Comp ut e r Science , pp. 294 – 302, 2018. J. K r ebs, T. M ansi , B. M ailhé, N . A y a che, an d H . De linge tte, “Unsup ervised Probab ilist ic D efo rmation Modeling f o r R obust Diff eomo rphic Regis tration, ” Lec ture No tes in Com pute r Scie nce , pp. 101 – 109 , 2018. I . Yoo , D. G. C . Hilde brand , W. F . Tobi n, W . -C. A . Lee, and W.- K . Jeong , “ssEMne t : Ser ial -Sectio n Elec tro n Mic rosco p y I mage Reg istration Using a Spatia l Tran sform er Netwo rk w ith Lear ned Feat ures,” Lec ture Notes i n C ompu ter Scie nce , pp. 249 – 25 7, 2017. M. Salehi , R. Pr evo s t, J.- L. Moc tezuma, N . N avab , and W. We in, “Preci se Ultraso und Bo ne Regis tratio n w ith Learn ing -Based Segm entation and Speed of Sound Ca librat ion,” i n Medical Image Comput ing and Com puter- As sisted Interve ntion − MICCAI 2017 , pp. 682 – 6 90, 2017 . S. Sun, J . Hu, M. Y ao, J. Hu, X. Ya ng, Q. So ng, and X. Wu , “Robust Multimo dal I mag e Reg i stratio n U sing Deep R ecurren t Reinfo rcement Learn ing,” Lec ture Note s in Com puter Scie nce , pp. 511 – 5 26, 2 019. J. K rebs, H . e Deling ette, B . M ailhe, N . A yache, and T . Mansi , “L earni ng a Pro babilis tic Mode l fo r Diffeo morphic Reg istratio n ,” I EEE Transac tions on Medica l Ima ging , Ear ly A ccess, pp. 1 – 12 , 2019. K. A . J. Eppenho f, M. W. Laf arge, P. Moesko ps, M. Ve ta, and J. P. W . P luim, “Defo rmab le imag e registra tion using co nvolutiona l neural networks ,” Medic al Im aging 2018: Ima ge P rocessing , Ma r. 2018 . K. A . J. Epp enhof and J. P. W. Pl u im, “S uperv ised local erro r e sti mation fo r non linea r imag e r egis tratio n using co nvolutiona l neura l network s,” Me dical Im aging 2017: Ima ge P rocess ing , Feb . 2017 . X. C ao, J. Y ang, J. Zhang, Q. Wang , P. - T. Yap, and D. Shen , “Def ormable I mage Registra tion Using a Cue -A ware De ep Regress ion Netwo rk,” IEEE Trans actions on Biom edica l Engine ering , v ol. 65, no. 9, pp. 1900 – 1911 , Sep. 201 8. Y. Hu, M. M o dat, E. G ibso n, N. Ghav ami, E. Bonm ati, C. M. Moo re, M. Emb erto n, J. A . No ble, D. C. B arratt , and T. Vercau teren, “L abel -dr iven w e ak ly- supervis ed learning fo r multimo dal defo rmarl e i mag e re g istratio n,” in 2018 IEE E 15th Internat ional Sympos ium on Biome dical Im aging (I SBI 2018), A pr. 2018. A . She ikhjafari , M. No ga, K . Pun i thakum ar , N . R ay , “Unsupe rvised defo r mab le imag e regi stration w ith fu l ly co nnected gene rativ e neural network ,” in p roc . the 1 st Confe rence on Me dical Imag ing w ith Dee p Learn ing (MI DL 2018), T he Net herlands , 2018. H. Yu, X. Zhou, H. Jia ng, H. K ang, Z . Wang , T. Ha ra, and H. Fuji ta, “L earning 3D non -rigid d efo rmation ba sed o n an unsu pervised deep lea r ning fo r PET/C T imag e registrat ion,” in Medica l Imag ing 2019 : Biome dical Appl icat ions in M olecular , Struc tural, and Func tiona l Imagin g , Mar. 2019. Y. Pei , Y. Zha ng, H . Q in, G. Ma, Y. Guo , T. Xu, and H. Zh a, “No n -rigid Cran iof acial 2D-3D R e g istration Using CNN-Based Reg ression,” Le cture No tes in Compu ter Sc ience , pp. 11 7 – 125, 2017. S. R. Van Kranen , T. Kaneh ira, R . Rozendaa l, and J. Sonk e, “ Un superv ised deep lear ning fo r fast and acc urate CB CT t o CT defo rmable im age reg i strat ion,” Radio therapy and Onco l ogy , vo l . 1 33, pp. S267 – S268, Apr. 2019. A . Her ing and S . Heldm ann, “Unsupe rvised learning for la rge mo tio n t horac ic CT fo ll ow - up regist ration,” in Medica l Imagin g 2019 : Image P r ocess ing , Mar . 2019. M. Blendo wsk i and M. P. He inrich , “Com bining M RF -ba sed def orm able regis tratio n and dee p binary 3D -CN N descrip tors fo r large lung mo t ion es timat ion in COPD patien ts,” Inte rnationa l Jou rnal of Com pute r A ssiste d Radio logy and Surge ry , vo l. 14, no . 1, pp. 43 – 52 , 2019 . - 42 - G. Hask ins, J . K ruecke r, U. K ruger , S. Xu, P . A. Pinto , B. J. Wo od, and P. Yan , “ L earning deep similar ity m etric fo r 3D MR – TRUS imag e registr ation,” Interna tiona l Journa l of Comp ut er Assiste d Ra diolog y and Sur gery , vo l. 14 , no . 3, pp . 417 – 425 , 2019 . X. Liu , D . Jiang , M. W ang, and Z. Song , “I mage sy nthesis -based mult i-mo dal image regist ration fr amewo rk by using d eep ful ly co n volutiona l netwo rks,” Med ical & Biologica l Engi nee ring & Computing , vo l. 57, no . 5, pp . 1037 – 10 48, Dec. 2018 . J. Fan, X. Cao , P. - T . Yap , and D. Shen, “BI RNet: Brain imag e regis tratio n using d ual - superv ised fu lly co n vo l utional netw orks,” Medic al Image Analy sis , vo l. 54, pp. 19 3 – 206, M ay 2019. S. S. Mo hsen i S alehi, S. Khan , D. Erdo gm us, and A . Ghol ipour, “R eal -Tim e D eep Pose Estima tion with Geodes ic L oss fo r Im age - to - Templa te Rig id Reg istrat ion,” I EEE Tran s acti ons on Medical Imag ing , v ol. 38 , no. 2, pp. 470 – 481 , Feb . 2019. M. S. E lmahdy , T. Jagt, R . T. Z inkstok, Y. Qiao , R . Shahz ad, H. So kooti, S. Yo us efi, L . Inc rocci , C. A. M. Mari jnen, M. Hoo geman, and M . Staring , “Ro bust co ntour pro pagat ion using deep learning and im age reg istrat ion fo r online a d aptive proton therapy of prostate canc er,” Me dical P hysics , May 2019. T. -H. Chan , K. Jia, S. Gao , J . L u, Z. Z eng, and Y. Ma, “PCA Net: A Si mp le De ep L earning Ba se line fo r I mage Class ificat ion , ” IEEE Transac tions on Image Proce ssing , vo l. 24, no . 12, pp. 5017 – 5032 , Dec . 2015 . D. To th, S. Miao , T . Ku r zendo r fer , C . A. R inaldi, R . Liao , T. M ansi, K. Rho de, an d P . Mountney , “3D /2D m ode l - to -imag e regis tration by imitation l earning fo r cardiac pro cedure s,” Intern ational Jou rnal of Com put er As sisted Radio logy and Sur gery , v ol. 13 , no . 8 , pp. 1141 – 1 149, May 2018. S. Miao , S. Piat, P. Fischer , A . Tuy suzog lu, P . Mewes, T. Mansi, R. Liao , “D ilated FCN for mu lti -ag ent 2D/3D medic al image registr ation,” in T hirty-Seco nd AAAI Con ference on Ar tificia l Intell ige nce , A pr. 2018. J. Zheng , S. Miao , and R. Liao, “L earning CNNs with P airwi se Do main A daptio n fo r Real -T i me 6DoF Ultraso und Transduc er Detect ion and Tracking from X- Ray I mag es, ” i n Medical Im age Computing and Compute r - Ass isted Interve ntion − MICCA I 2017 , pp. 646 – 6 54, 2017 . O. Ronne be rge r, P. F ischer, a nd T. Brox , “U - N et: Co nvolutional ne tworks for biom edical image segmen tation, ” in Proc . Medi cal Image Com put ing and C omputer-Assi sted I nte rvent ion (MI CCAI 2 015), 201 5, pp. 234-24 1. X. C ao, J. Yang, L . Wang , Z. Xue , Q. Wang , and D. Shen , “Deep L earni ng Based I nter -mo dality Im a ge Regis tration Supe r vise d by I ntra- mo dali ty Simila rity ,” Lecture Note s in Com puter S cience , pp . 55 – 63 , 2018. P. Y an, S. Xu , A . R. Rast inehad , an d B . J. W o od, “A dversar ial I mage Reg istra ti o n w ith A pplication fo r MR and TRUS I mag e Fusion ,” Lecture Notes in Compu ter Sc i ence , pp. 197 – 20 4, 2018 . T. Che, Y. Z heng, J. Co ng, Y. J iang, Y. Ni u, W . Jiao, B. Zhao , and Y . D ing, “Deep Gro up -Wise Reg istra ti o n fo r Mul ti-Spectral I mages From Fundus Im ages,” IE EE Acce ss , v ol. 7, pp . 27650 – 276 61, 201 9. T. C he, Y . Zhe ng, X . Sui , Y . J iang , J. Co ng, W . Jiao , a nd B. Zh ao, “ DGR - Net: Deep G roupwis e R egistra tion of Mult ispectr al I mages,” Informa tion Pro cessi ng in Medical I ma ging , pp. 706 – 717, 2019. G. Ba lakr ishnan, A. Z hao , M. R . S abu ncu, J. Gut tag, and A . V. D alc a, “Vo xelMorph: A L e arning Framewo r k fo r Def ormable M e dical Im age Reg istration,” IEEE Tr ansaction s on M ed i cal Imaging , Ea rly Acc ess, pp. 1 – 13 , 2019. A . Her ing, S . Kuc kertz, S. H eldm ann, an d M . P. Hein ri ch, “En hanci ng L abel -Driv en Dee p Def orm able I mage Reg istration w it h Lo cal Distanc e Met rics fo r State- of -the- A rt Cardi ac Mo tion T racking, ” Bi ldvera rbeitung für die Medizi n 2019 , pp . 309 – 314 , 2019 C. Liu, L . Ma , Z . L u, X . Jin , a nd J. Xu, “Multimo dal medic al i mag e reg istration via c o mmo n representa tions learning and diff erentiab le geo metri c c onstrain ts,” Electr onics Le tters , v ol. 55, no . 6, pp. 316 – 318, Mar. 2019. F. Chollet , “Xcep ti o n: Deep Learn ing with Depthwise S eparable C onvo luti o ns,” in 2017 IEEE Co nference on C ompute r Vis ion and P att ern Re cogni tion (CVPR 2017) , Jul. 2 017. E. A ba no vie, G . S tankevi eius, and D. Matuzev ieius, “ Deep Neu ral Net wo rk - based Feature D escripto r f or R etinal I mag e Reg istration ,” in IEEE 6th W orkshop on A dvance s in Info rma ti o n, Ele ctronic and Electric al Engi ne ering (A I EEE 2018), No v. 2018. R. Sc haffer t, J. Wa ng, P. F ischer, A . Borsdo rf, and A. Maier, “M etric -Driv en L earning o f Correspondenc e Weig hting fo r 2 -D/3- D Im age Reg istration,” Lec ture N otes in Compute r Scie nce , pp. 1 40 – 1 52, 2019. C. R. Qi, H. Su , M. K aic hun, and L. J. Guibas, “Poin tNet: Deep L earning on Po int Sets fo r 3D Cla ssific ation and Segm entatio n,” in 2017 IEEE Co nference on Comp ut er Vision and P attern Re cogni tio n ( CVPR ), Jul. 201 7. L . Sun an d S . Zha ng, “D e fo r mable MRI - Ultrasound Reg i strat ion Us i ng 3D Co nvo lut ional Neu r al N et wo rk,” Le cture No t es in Compute r Scie nce , pp . 1 52 – 158 , 2018 . X. Y ang, X. Han, E. Park , S. A y lward, R. Kw itt, and M. Nieth am mer, “ R egis tration of Patho log ical I mage s,” Le cture Note s in Compute r Scie nce , pp . 9 7 – 107, 20 16. P. S. Bhati a, F . Reda , M. Hard er, Y . Zhan , and X. S. Zho u, “Real ti me co ar se orient ation detect ion in MR scans using mu lti -pl anar deep c onvo luti o nal neu ral netwo rks,” in Medical Imagin g 2017 : Image Proce ssing , Feb . 2017 . C. Shu, X. Chen , Q. Xie , and H. Han , “A n unsuperv ised netwo rk for fas t microsc opic image regist ration, ” in Medi cal Imagin g 2018: Digita l P athology , Mar . 2018. C. Ste rgio s, S. Mihi r, V. M ari a, C. Guil laume, R . Marie-Pie rre, M. Stav r o u l a, and P . Niko s, “Linea r and Defo r mab le Im age Registratio n with 3D Conv olutional Neu ral Netwo r k s,” Lec ture Note s in Com pu ter Scie nce , pp. 13 – 22, 2018 . J. On ieva Oniev a, B . Mar ti - Fu ster, M. Pedr ero de la P uente , and R. San José Estépa r, “Diff eomo r phic Lung Regis tratio n Using Deep CNN s an d Reinf orc ed Learn ing,” Lecture Note s in Com puter S cien ce , pp. 284 – 294 , 2018. L . Duan, G . Yuan, L. Gong , T. Fu, X. Yang , X . Chen , and J. Zheng , “Adv ersarial learning fo r deformabl e registrat ion of brain MR imag e using a mu lti - sc ale fully co nvolutional netwo rk,” Biome dical Signa l Proce ssing and Control , v ol. 53, p. 10156 2, Aug . 2019. - 43 - V. Kea rney , S. Ha af, A . Sudhy a dho m, G. V a ldes, and T. D. Solb erg, “A n unsu pervised co nvo l utional n eural netwo rk -based alg orithm fo r deform able imag e reg istratio n,” Physi cs in Me dicine & B iology , v ol. 63, no . 18, p. 185017 , Sep. 2018. M. D. Foo te, B . E. Zimm erman , A . Sawant , and S. C. Joshi, “Re al -Time 2D-3D Def orm able Reg istra tion with Dee p L earn ing and A pplication t o L ung Radio therapy Target ing,” Informa tion Process ing in Medical I m aging , pp. 265 – 2 76, 2019 . H. G ao, Z. L iu, L. van d er Maaten, and K. Q. Weinbe r ger, “Densel y C onnec ted Conv olutional Netwo rks,” in 2017 IEEE Con f ere nce on Com puter Vision and Patter n Recog nition (CVPR ), Jul. 2017 . R. Awan and N . Rajpo ot, “ De ep A utoenc oder Feat ures fo r Regist ration of Histolo gy I mag es, ” Medica l Image Unde rstandin g and Analys is , pp. 37 1 – 378, 20 18. Y. Sun, A . M o elker, W . J. Niess en, and T. van Wa l sum , “To wards Ro bust C T - Ul trasound Reg i st ration Using Deep L e arning Methods, ” Lecture Notes in Compu ter Sc i ence , pp. 43 – 51 , 2018. M. Blendow ski and M . P. He inrich, “3D - CNNs fo r Dee p Bin ary Descrip tor Learn ing i n Medica l Vo lume D ata,” Informa tik aktuel l , pp. 23 – 28 , 2018 . L . Zhao and K. J i a, “D eep A dapti ve Lo g - Demons: Diff eomo rphic I mag e Reg istration wit h Ve ry Larg e Def ormations , ” Com puta tiona l and Ma themat ical Methods in Med icine , v ol. 20 15, pp. 1 – 16, 2015 . N. Zhu , M. N ajafi, B. Han , S. Hanco ck, a nd D. Hris tov , “F easi bility o f I ma g e Reg i strat ion fo r U l trasound -Gu ided Pro state Radioth erapy B ased on S i mi larity Measu reme nt by a Conv olutional Neur al Netwo rk,” Te chnol ogy i n C ancer Re search & Treatme nt , vo l. 18, pp. 15 3303381 882196, Jan . 2019 . I . Goodf ellow, J. Po uget-A badie, M. Mirz a, B. Xu, D. Warde-Far ley , S. Ozair , A . Co urvill e, and Y. Bengio , “Gene r ativ e adv ersarial nets,” i n Advance s in neur al inform ation proce ssing sy st e ms , 2014, pp. 26 72-2680 . X. Yi , E. W ali a, and P . Baby n, “Gener ative adv ersarial netwo rk in medic al imag i ng: A review, ” arX iv pre print arXiv :1809.0729 4, v2, pp. 1-19 , 2018. S. Hoc hreiter and J. Sc hmid huber, “L ong Sho rt - Term Memo ry ,” Neural Com putation , vo l. 9, no . 8, pp . 1735 – 178 0, No v. 1997. K. Cho, B . van Mer rienboer , C. Gulc ehre , D. Bah danau , F. Bo ugar es, H. Schw enk, and Y. B engio , “L earn ing Phras e Represen tat ion s using RNN Enc oder –Dec oder fo r Statis ti ca l Machine Tran slatio n,” in pr oc . 2014 Co nference on Empi rical Meth ods in Natural Langua ge Proce ssing (E MNL P), 2014 . S. Zago ruy k o and N. Kom odakis, “Learning t o co mpare i mag e patches v ia co nvo lut ional neu ral netwo rks,” i n pro c. 20 15 IEEE Confere nce on Compute r Vi sion and Pattern Recog nition (CVPR) , Jun . 2015 . E. Nowak and F. Jur ie, “Learn i ng Visual Sim ilarity M easure s fo r Comparing Nev er Seen Objec ts,” in 2007 I EEE C onfe rence on Compute r Visi on and Pattern Recog nition , Jun . 2007. G. Huang , M. M a tt ar , H. Lee , and E.G. Lea r ned- Mil ler, “Learning to align from scratch,” in 2012 Advance s in Neura l I nforma tion P rocessing Sys t em s , 201 2, pp. 764 -77 2. A . So ti ras , C . Davatz ikos, and N. Parag ios, “D efo rmable Medical Im age Reg is tratio n: A Sur vey ,” IEE E T ransactio ns on Medical Imag ing , vol. 32, no . 7, pp. 1153 – 119 0, Jul. 2013 . M. Jaderberg , K. S i mo ny an, A . Z isserm an, and K . Kav ukcuog l u, “S patia l tra nsfo rmer ne tworks ,” i n 2015 Adv ances in ne ural informa tion pro cess ing system s , 2015, pp. 2017-20 25 . C. - H. Lin and S. Luc e y , “I nverse Co mpo sitional Spatia l Transf orm er Netwo rks,” in 2017 IEEE Con ference on Comp uter Vision and P att ern Re cogni tion (CVPR ) , Jul . 2017. - 44 - Appendi x 1: A cronym s CNN Convolutio nal Neural N etwork s PCC Pearson 's Correlatio n Coefficie nt DNN Deep N eural Networks RMSD Root M ean Square d E rr or SNN Spiking N eural Net work NRMSD Normaliz ed RMSD AEs Auto-En coders MDM Mean De formatio n M a gnitude DAEs Denoising A Es MDG Mean De formatio n Gradient SAEs Staked AEs MID Mean Inte nsity Di fference GAN Generative Adversarial N etwork CSPE Cumulati ve Sum of Pr ediction Err or DRL Deep Rei nfo r cem ent Learning TRE Ta rge t Re gistrati on Error LIR Local Ima ge Residua l D-Scores Discriminator Scores LSTM Long Sh ort-Term M emory SSIM Structural S imilarity In dex ISA Independent Subset Anal ysis PSNR Peak Signal To Nois e Ratio LDDMM Large De formation Di ffeomorp hic Metric Map ping DSC Dice Coe ff i cient SVF Stationary V elocity Fi eld ASD Average Sur face Di stance TPS Thin-Plat e Spline ASSD Average Sym metric Sur face Dista nce LIF Leaky Integrat e-a nd-Fire ASCD - or - MCD Average Sym metric C ontour D istance - or - Mean Contour Di stance DR Digital Ra diograph RVLJ Relative Var iance Lo g-Jaco bian DDR Digitally R econstru cted Radiogra ph Grad Det -Jac Mean ma gnitude o f the Gradi ents of the Determinant of the Ja cobian DVF Displac ement Vector Fi eld LLC Local Correlat ion Coe fficient IGRT Image-Guid ed Radio - T he ra py MI Mutual Infor mation RoI Region of Int erest NMI Normaliz ed Mutual In formation OoI Organ of Int erest FSIM Feature Si milarity Inde x Metric Linac Linear Acc elerator Mac hine RMSE c Root M ean Square d E rr or of t he 3D Cann y Edg e DSS Decision Su pport Sy stem LCC Local Cros s-Correlat ion TPS Thin-Plat e Spline NCC Normaliz ed Cros s-Correlation CT Computed To mograp hy RPD Re -Projectio n Distanc e MRI Magnetic Re sonanc e Imaging CPD Closest P oint Distan ce (CPD) PET Positron E mission To mography DSM Deep Similarit y Metric s TEE Tra nsesophageal Ech ocardiograph y SE2ER Supervi sed End - to -End Regi stration EM Electron Mi croscopy DRL Deep Rei nfo r cem ent Learning FA Fluoresc ein Angiograph y UE2ER Unsupervi sed End- to -End Re gistrati on US Ultrasound WSE2ER Weakly/Se mi-Super vised En d- to - End R egistratio n MSI Multi-Spe ctral Imagin g TRUS Tra ns- R ectal Ultra -Sou nd DSA Digital Su btraction Angiograph y SSD Sum of Squa red Differe nce s CFI Color Fun dus Image s MSE - or - MSD Mean Squar ed Error - or - Dista nce OCT Optical Coh erence Tom ography MAE - or - MAD Mean Ab solute Error - or - Dista nce DoF Degree of Fr eedom - 45 - Appendi x 2: Oth er 164 autho rs act ive in th is field Au th or Pub.s Cite.s Au th or Pub.s Cite.s Au th or Pub.s Cite.s Au th or Pub.s Cite.s Lee, Wei-Chun g A. 1 11 Zha, Hongbin 1 5 Ito, Masato 1 1 Rozendaal, R 1 0 Tobin, Willie F 1 11 Xu, Tianmin 1 5 Werner, Rene 1 1 Kanehira, T 1 0 Hildebrand, David 1 11 Guo, Yuke 1 5 Mad e sta, Frederi c 1 1 Van Kranen, SR 1 0 Yoo, Inwan 1 11 Ma, G e ngyu 1 5 Sentk e r, Thilo 1 1 Sui, Xiaodan 1 0 Garnavi, Rahil 1 10 Qin, Haifang 1 5 Nikos, Paragios 1 1 Fujita, Hiroshi 1 0 Antony, Bhavna 1 10 Zhang, Yungeng 1 5 Stavroula, M. 1 1 Hara, Takeshi 1 0 Wein, Wolfgang 1 10 Pei, Yuru 1 5 Marie-Pierre, Revel 1 1 Wang, Zhiguo 1 0 Moctezuma, J-L 1 10 Rastinehad, Ardeshir R 1 4 Guillaume, C. 1 1 Kang, Hongjian 1 0 Prevost, Raphael 1 10 Kuckertz, Sven 1 3 Maria, Vakalopoulou 1 1 Jiang, Huiyan 1 0 Salehi, Mehrdad 1 10 Gholipour, Ali 1 3 Mihir, Sahasrabudhe 1 1 Zhou, Xiangrong 1 0 Solb e rg, Timoth y D 1 9 Erdogmus, Deniz 1 3 Stergios, C. 1 1 Yu, Hengjian 1 0 Valdes, Gilmer 1 9 Khan, Shadab 1 3 Han, Hua 1 1 Hoogeman, Mischa 1 0 Sudhyadhom, At char 1 9 Salehi, Seyed Sade gh 1 3 Xie, Qiwei 1 1 Marijnen, CAM 1 0 Haaf, Samuel 1 9 Pun i thakuma r, K. 1 3 Chen, Xi 1 1 Incrocci, Luca 1 0 Kearney, Vasant 1 9 Noga, Michelle 1 3 Shu, Chang 1 1 Yousefi, Sahar 1 0 Mewes, Philip 1 9 Sheikhjafari, Ameneh 1 3 Zhang, Xuming 1 1 Shahzad, Rahil 1 0 Tuysuzoglu, Ahmet 1 9 Milone, Diego H 1 2 Jin, Xiaomen g 1 1 Qiao, Yuchuan 1 0 Fischer, Peter 1 9 Glocker, Ben 1 2 Huang, Tao 1 1 Zinkstok, R Th 1 0 Pia t , Seb as tie n 1 9 Oktay, Ozan 1 2 Ding, Mingyue 1 1 Jagt, Thyrza 1 0 Ghosal, Sayan 1 9 Ferrante, Enzo 1 2 Zhu, Xingxing 1 1 Elmahdy, Mohamed S 1 0 Jia, Kebin 1 8 Zhang, Songtao 1 2 Hristov, Dimitre 1 0 Wu, Xi 1 0 Zhao, Liya 1 8 Sun, Li 1 2 Hancock, Steven 1 0 Song, Qi 1 0 Aylward, Stephen 1 7 Chakravorty, Rajib 1 2 Han, Bin 1 0 Hu, Jinrong 1 0 Park, Eunbyung 1 7 Ge, Zongyuan 1 2 Najafi, M ohamm ad 1 0 Yao, Mingqin g 1 0 Han, Xu 1 7 Siebert, J Paul 1 2 Zhu, Ning 1 0 Hu, Jing 1 0 e Delingette, Herve 1 6 Goatman, Keith A 1 2 Zh e ng, Jian 1 0 Sun, Shanhui 1 0 Chen, Terrence 1 6 Sloan, James M 1 2 Chen, Xinjian 1 0 Song, Zhijian 1 0 Wimmer, Andreas 1 6 Ding, Yanhui 1 1 Fu, Ti anxiao 1 0 Wang, Manning 1 0 Chang, Yao-Jen 1 6 Niu, Yi 1 1 Gong, Lun 1 0 Jiang, Dongsheng 1 0 Tam e rsoy, Birgi 1 6 van Walsum, Theo 1 1 Yuan, Gang 1 0 Liu, Xueli 1 0 Singh, Vivek 1 6 Niessen, Wiro J 1 1 Duan, Luwen 1 0 Estepar, Raul S an J. 1 0 Wang, Jiangping 1 6 Moelker, Adriaan 1 1 Joshi, Sarang C 1 0 de la Puente, Mari a P . 1 0 Ma, Kai 1 6 Sun, Yuanyuan 1 1 Sawant, Amit 1 0 Marti-Fuster, Berta 1 0 Wang, Shaoyu 1 6 Pinto, Peter A 1 1 Zimmerman, Blake E 1 0 Onieva, Jorge Onie va 1 0 Wang, Li 1 5 Kruger, Uwe 1 1 Foo te , Markus D 1 0 Ra j poot, Nasir 1 0 Yang, Jianhuan 1 5 Kru e cker, Joc hen 1 1 Xu, Jingyun 1 0 Awan, Ruqayya 1 0 Mountney, Peter 1 5 Haskins, Grant 1 1 Jin, Xiance 1 0 Zhou, Xiang Sean 1 0 Rhode, Kawal 1 5 Matuzevieius, D ali us 1 1 Lu, Zheming 1 0 Zhan, Yiqiang 1 0 Ri naldi, Christo ph e r 1 5 Stank e vieius, G. 1 1 Ma, Longhua 1 0 Harder, Martin 1 0 Kur ze ndorfer, Tanj a 1 5 Abanovie, Eldar 1 1 Liu, Cong 1 0 Reda, Fitsum 1 0 Toth, Daniel 1 5 Ino, Fumihiko 1 1 Sonke, J 1 0 Bhatia, Parmeet S 1 0
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment