Artificial Neural Network Approach for the Identification of Clove Buds Origin Based on Metabolites Composition
This paper examines the use of artificial neural network approach in identifying the origin of clove buds based on metabolites composition. Generally, large data sets are critical for accurate identification. Machine learning with large data sets lead to precise identification based on origins. However, clove buds uses small data sets due to lack of metabolites composition and their high cost of extraction. The results show that backpropagation and resilient propagation with one and two hidden layers identifies clove buds origin accurately. The backpropagation with one hidden layer offers 99.91% and 99.47% for training and testing data sets, respectively. The resilient propagation with two hidden layers offers 99.96% and 97.89% accuracy for training and testing data sets, respectively.
💡 Research Summary
This paper investigates the application of artificial neural networks (ANNs) for classifying the geographic origin of clove buds based on their metabolite profiles. The authors used a relatively small dataset comprising 94 experimental samples, each characterized by concentrations of 47 metabolites. The samples represent four Indonesian regions—Java, Bali, Manado, and Toli‑Toli—with each region encoded as a unique 4‑bit binary label (e.g., Java = 1000).
Data preprocessing was a critical step due to the wide concentration range (10⁻⁴ to 10) and the presence of zero values. The authors applied a logarithmic transformation to all non‑zero measurements and replaced zeros with 10⁻⁵, a value one order of magnitude below the smallest observed concentration, to preserve the biological significance of absent metabolites. Subsequently, each variable was normalized using a z‑score approach (subtracting the mean and dividing by the standard deviation) to ensure equal contribution of all metabolites during training.
Two ANN learning algorithms were evaluated: standard backpropagation (B‑Prop) and resilient propagation (R‑Prop). Network architectures varied in the number of hidden layers and neurons. The input layer always contained 47 neurons (one per metabolite) and the output layer 4 neurons (binary code). For B‑Prop, the authors tested a single hidden layer with 15 neurons (derived from the Shibata‑Ikeda rule Nₕ ≈ √(Nᵢ·Nₒ) ≈ 13, rounded up) and several two‑hidden‑layer configurations (3‑5, 4‑6, 5‑7, 6‑8 neurons). For R‑Prop, only two‑hidden‑layer configurations were examined, using the same neuron counts.
Training employed an 80 %/20 % split of the dataset, repeated 30 times with random selection to obtain robust average performance metrics. Hyperparameters were fixed across experiments: learning rate = 0.9, momentum = 0.1, maximum epochs = 5000, and an error target of 10⁻³. Performance was measured by classification accuracy, mean squared error (MSE), and coefficient of determination (R²).
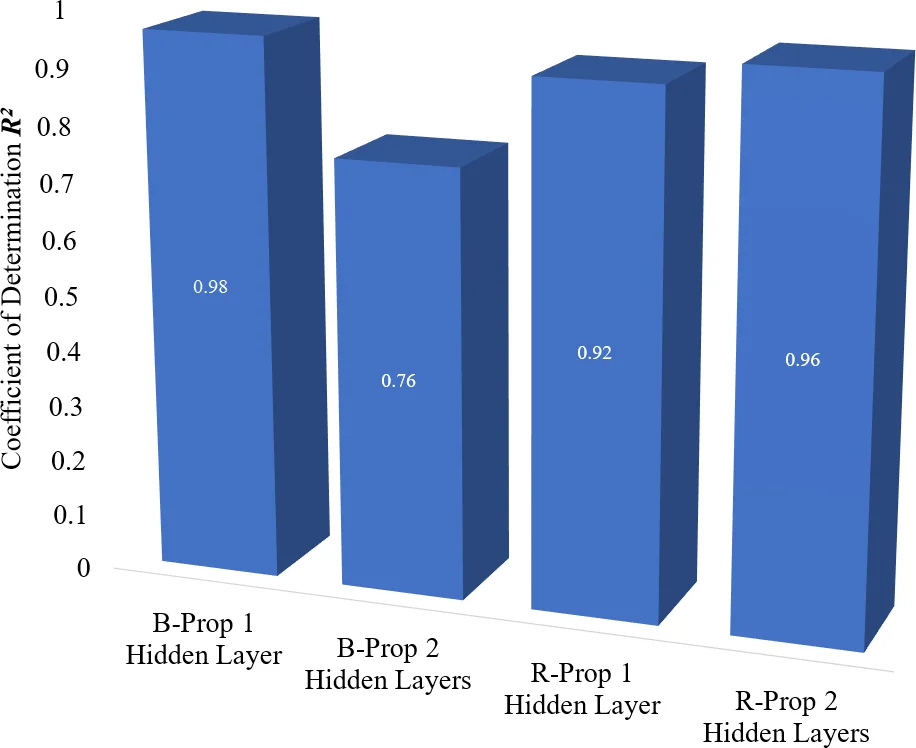
Results showed that the B‑Prop network with a single hidden layer (47‑15‑4) achieved the best overall performance: 99.91 % training accuracy, 99.47 % testing accuracy, MSE = 0.0721/0.0773, and R² ≈ 0.99/0.98. In contrast, increasing the number of hidden layers generally reduced performance; the two‑hidden‑layer B‑Prop models exhibited lower accuracies (down to ~73 % testing) and higher MSEs, confirming that added complexity led to over‑fitting on the limited data.
R‑Prop with two hidden layers performed comparably in training (up to 99.96 % accuracy for the 47‑5‑7‑4 architecture) but lagged in testing (97.89 % accuracy), indicating that while R‑Prop converges quickly, its generalization suffers when data are scarce. The authors attribute this to the algorithm’s aggressive weight‑update scheme, which can amplify noise in small datasets.
The study concludes that even with a modest sample size, ANN models—particularly a simple backpropagation network with one hidden layer—can reliably discriminate clove bud origins based on metabolite fingerprints. The findings underscore the importance of balancing model complexity against data availability; adding hidden layers or neurons beyond a certain point yields diminishing returns and may introduce local‑minimum trapping.
Future work suggested includes expanding the dataset, applying data‑augmentation or cross‑validation techniques, exploring ensemble methods, and benchmarking against alternative classifiers such as support vector machines or random forests. Such extensions would help validate the robustness of the proposed approach and potentially improve its applicability to other food‑authentication problems where metabolomic data are limited.
Comments & Academic Discussion
Loading comments...
Leave a Comment